- Laboratory of Plant Molecular Genetics, Graduate School of Agricultural and Life Sciences, The University of Tokyo, Tokyo, Japan
Membrane traffic between two organelles begins with the formation of transport vesicles from the donor organelle. Dynamin-related proteins (DRPs), which are large multidomain GTPases, play crucial roles in vesicle formation in post-Golgi traffic. Numerous in vivo and in vitro studies indicate that animal dynamins, which are members of DRP family, assemble into ring- or helix-shaped structures at the neck of a bud site on the donor membrane, where they constrict and sever the neck membrane in a GTP hydrolysis-dependent manner. While much is known about DRP-mediated trafficking in animal cells, little is known about it in plant cells. So far, two structurally distinct subfamilies of plant DRPs (DRP1 and DRP2) have been found to participate in various pathways of post-Golgi traffic. This review summarizes the structural and functional differences between these two DRP subfamilies, focusing on their molecular, cellular and developmental properties. We also discuss the molecular networks underlying the functional machinery centering on these two DRP subfamilies. Furthermore, we hope that this review will provide direction for future studies on the mechanisms of vesicle formation that are not only unique to plants but also common to eukaryotes.
Introduction
Eukaryotic cells are distinguished by the presence of internal membrane-bound organelles, including mitochondria, peroxisomes, plastids, and other single membrane-bound organelles [e.g., the endoplasmic reticulum (ER), Golgi apparatus, trans-Golgi network (TGN), plasma membrane (PM), and a series of endosomal compartments]. These single membrane-bound organelles are connected with each other through a membrane trafficking system mediated by vesicular and/or tubular membranous transport carriers. Membrane traffic consists of four sequential processes: the formation of cargo-bearing vesicles or tubules from the donor membrane, targeted delivery of transport carriers, tethering of carriers to target membranes and membrane fusion (Bonifacino and Glick, 2004). Especially, in post-Golgi traffic, the formation of transport vesicles from the donor organelle is accomplished by snipping the neck of the invaginated membrane by dynamin-related proteins (DRPs; McMahon and Mills, 2004; Praefcke and McMahon, 2004). DRPs are large multidomain GTPase that regulate membrane fission, fusion, and tabulation during diverse cellular activities such as endocytosis, cytokinesis, vacuolar sorting, fission and fusion of mitochondria, biogenesis of peroxisomes, and the maintenance of ER morphology (Heymann and Hinshaw, 2009; McNew et al., 2013). This review summarizes recent advances in understanding how DRPs are involved in post-Golgi traffic, focusing on the unique aspects of the plant system.
Dynamin, Membrane-Scission Catalyst in Animals
Presently, the best characterized DRPs are animal dynamins that act in post-Golgi clathrin-mediated traffic (Praefcke and McMahon, 2004; Ferguson and De Camilli, 2012). During clathrin-coated vesicle (CCV) formation, dynamin is thought to assemble into helical or ring shaped-structures at the neck of clathrin-coated membrane buds (Takei et al., 1995), and constrict to sever the bud neck membrane in a GTP hydrolysis-dependent manner (Sweitzer and Hinshaw, 1998; Macia et al., 2006). Much progress has been made in dynamin function using in vitro analyses (Chappie and Dyda, 2013). Purified dynamin assembles into a ring and a spiral-shaped structure with 40 ∼ 50 nm outer diameter (Hinshaw, 2000; Faelber et al., 2011; Ford et al., 2011). The intramolecular conformational change of dynamin with the activation of its GTPase reduces the dynamin helix diameter (Stowell et al., 1999; Danino et al., 2004). However, attempts to directly observe dynamin-spiral formation and constriction in living cells have been unsuccessful thus far.
On the basis of its sequence, dynamin harbors five distinct domains (Figure 1): an amino terminal GTPase domain, whose activation causes the intramolecular conformational change of dynamin (Ford et al., 2011), a middle domain, which mediates the intermolecular interaction between dynamins during self-assembly (Ramachandran et al., 2007), GTPase effector domain (GED), which stimulates the GTPase activity (Sever et al., 1999), a pleckstrin homology domain (PH domain) that may participate in the generation of membrane curvature and the breakdown of the lipid bilayer through the binding of acidic phospholipids and phosphatidyl inositol-4,5-bisphosphate (PI4,5P2) on the donor membrane (Ferguson et al., 1994; Zheng et al., 1996; Schmid and Frolov, 2011), and a carboxy-terminal proline-rich domain (PRD) harboring an array of PXXP amino acid motifs, which interact with many Src homology 3 (SH3) domain-containing proteins to localize dynamin at vesicle formation sites (Shpetner et al., 1996; Grabs et al., 1997). The former three domains (GTPase domain, middle domain and GED) are conserved among almost all DRP proteins. However, DRPs with a domain configuration similar to that of dynamin, which also harbor additional domains, have been found only in Metazoans and Embryophyta (Chanez et al., 2006; Miyagishima et al., 2008; Heymann and Hinshaw, 2009).
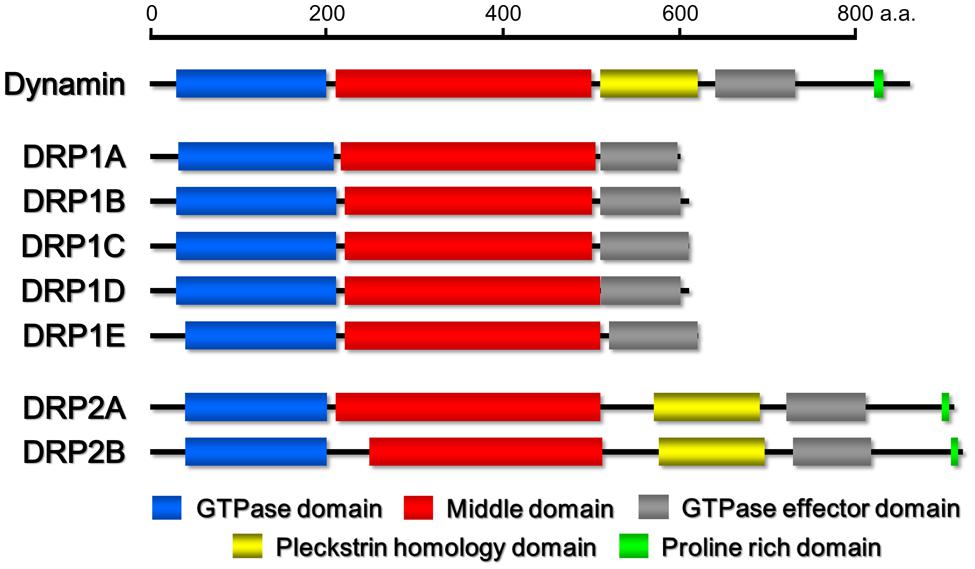
FIGURE 1. Schematic drawings of the domain organizations of Homo sapiens dynamin and Arabidopsis thaliana DRP1 and DRP2 family members. The domains were identified by the pfam program (http://pfam.sanger.ac.uk/).
X-ray crystallographic studies have provided insights into the spatial structure and disposition of each domain within the dynamin spiral polymer (Chappie et al., 2011; Faelber et al., 2011; Ford et al., 2011). The GTPase domain is placed at the outside of the spiral and interacts with the GTPase domain of dynamin in the adjacent turns of dynamin spiral. The PRD may protrude outward from the dynamin spiral structure (Ferguson and De Camilli, 2012). In contrast, the PH domain sits at the inside of the spiral, which is known as the “foot” region. This location is consistent with the expected function of the PH domain. The middle domain and the N-terminal region of the GED interact to form an intramolecular helical “stalk” region which is located between the GTPase and PH domains and is responsible for the dimerization of dynamin. This dimer in which the two GTPase domains are oriented in opposite directions is the basic unit for dynamin polymerization. The C-terminal region of the GED associates with the N- and C-terminal helix of the GTPase domain to assemble into a helix bundle, referred to as “neck” region, which modulates the GTPase activity of dynamin in vitro (Chappie et al., 2009).
Dynamin-Related Proteins in Land Plants
Based on phylogenetic analyses and the domain-structure predictions, the genomes of most land plants contain six types of DRPs: DRP1–DRP4, DRP5A, and DRP5B (Hong et al., 2003a; Miyagishima et al., 2008). Over the last two decades, much progress has been made in elucidating the functions of most of the plant DRPs (the function of DRP4 is still unclear). DRP3 is conserved in a wide range of Eukaryota and is involved in mitochondrial fission (Fujimoto et al., 2009). DRP5B is conserved in a wide range of Archaeplastida and is involved in plastid division (Gao et al., 2003; Miyagishima et al., 2003). Both DRP3 and DRP5B are also involved in fission and biogenesis of peroxisomes (Mano et al., 2004; Zhang and Hu, 2010). DRP5A, which closely resembles DRP5B in sequence and domain structure, is distributed in Viridiplantae, Amoebozoa, and Heterolobosea, in which it appears to participate in cytokinesis including cell plate formation (Miyagishima et al., 2008), although molecular functions of DRP5A is unclear. Thus, the functions of DRP5A and DRP5B appear to be different despite their close structural similarity. DRP1 and DRP2 are found in Viridiplantae and Embryophyta, respectively (Lopez-Bautista et al., 2003; Miyagishima et al., 2008). The phylogenetic distribution of DRP1 is wider than that of DRP2 (Hong et al., 2003a). Both proteins function in several types of post-Golgi traffic pathways: clathrin-mediated endocytosis (CME; Konopka et al., 2008; Fujimoto et al., 2010; Taylor, 2011) and cell plate formation (Hong et al., 2003b; Kang et al., 2003a; Fujimoto et al., 2008). DRP2 also participates in vacuolar trafficking (Jin et al., 2001). Interestingly, the overall domain organization of Embryophyta-specific DRP2s is similar to that of animal dynamins (Hong et al., 2003a; Figure 1). DRP1 lacks a PH domain and PRD (Hong et al., 2003a; Figure 1). Despite the similarity in the domain organizations of DRP2 and animal dynamin, the GTPase domain of animal dynamin is more similar to the GTPase domain of DRP1 than to that of DRP2 (e.g., animal dynamin has 62% identity to Arabidopsis thaliana DRP1A and only 29% identity to A. thaliana DRP2B; Figure 2).
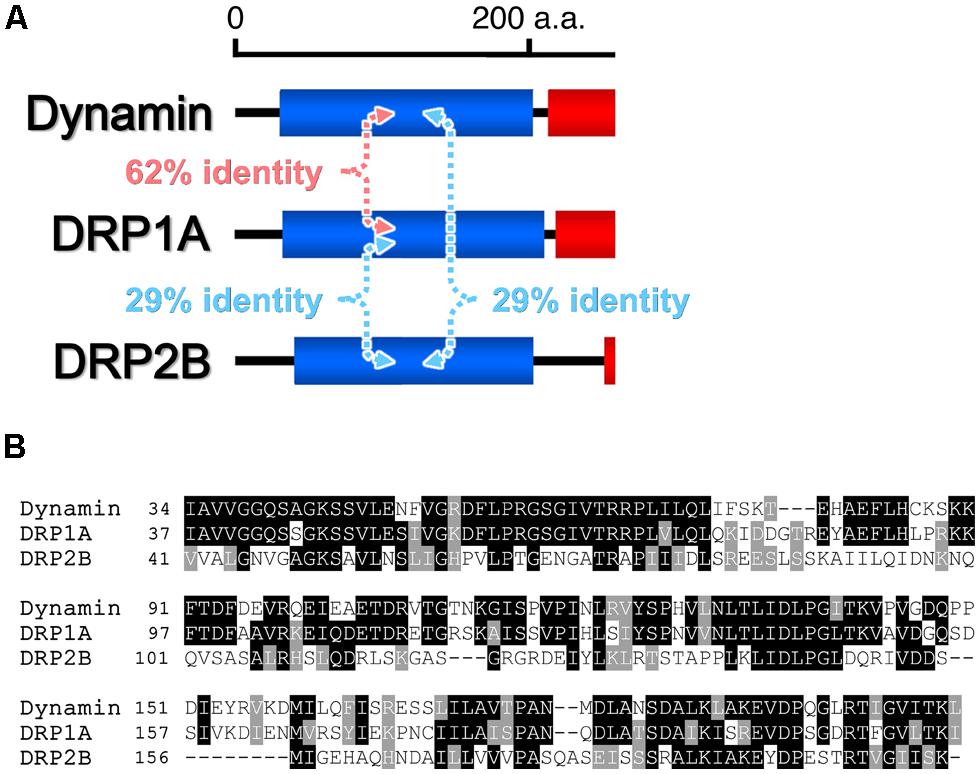
FIGURE 2. Comparisons of the amino acid sequences of the GTPase domains of H. sapiens dynamin and Arabidopsis thaliana DRP1A and DRP2B. (A) Pairwise amino acid sequence identities. (B) Clustal alignment of the sequences. Identical and similar amino acid residues are outlined in black and gray, respectively.
In Planta Functions of DRP1 and DRP2
Viridiplantae-specific DRP1 was originally identified as a dynamin-related GTPase, called phragmoplastin, that was associated with the formation of the cell plate in Glycine max (Gu and Verma, 1996). The gene was first sequenced in A. thaliana (Dombrowski and Raikhel, 1995). A. thaliana has five DRP1s (DRP1A–DRP1E; Kang et al., 2001; Hong et al., 2003a; Figure 1) that share a high degree (63–82%) of amino acid sequence identity. Experiments with DRP1 mutants showed that the five DRP1s have distinct in planta roles, mainly as a result of differences in their spatio-temporal expression patterns and levels (Bednarek and Backues, 2010). Loss-of-function mutants of these members exhibit pleiotropic developmental and cellular phenotypes: drp1a and/or drp1e null mutants have defects of cell plate formation in root and arrest of embryo development, which suggests that DRP1A and DRP1E participate in cytokinetic membrane trafficking or remodeling (Otegui et al., 2001; Kang et al., 2003a; Figure 3). drp1a null mutant also takes up less of an endocytic marker (FM4-64; Collings et al., 2008), does not restrict a cytokinesis-related Qa-SNARE protein (KNOLLE) to the division plane (Boutte et al., 2010) and has an altered cell wall composition (Collings et al., 2008). In addition, drp1c mutant pollen grains and in drp1a mutant stigmatic papillae also have aberrant PM invaginations and defects in cell expansion (Kang et al., 2003a,b). These mutant phenotypes suggest that DRP1 is involved in endocytic and/or recycling trafficking of PM lipids and proteins (Figure 3).
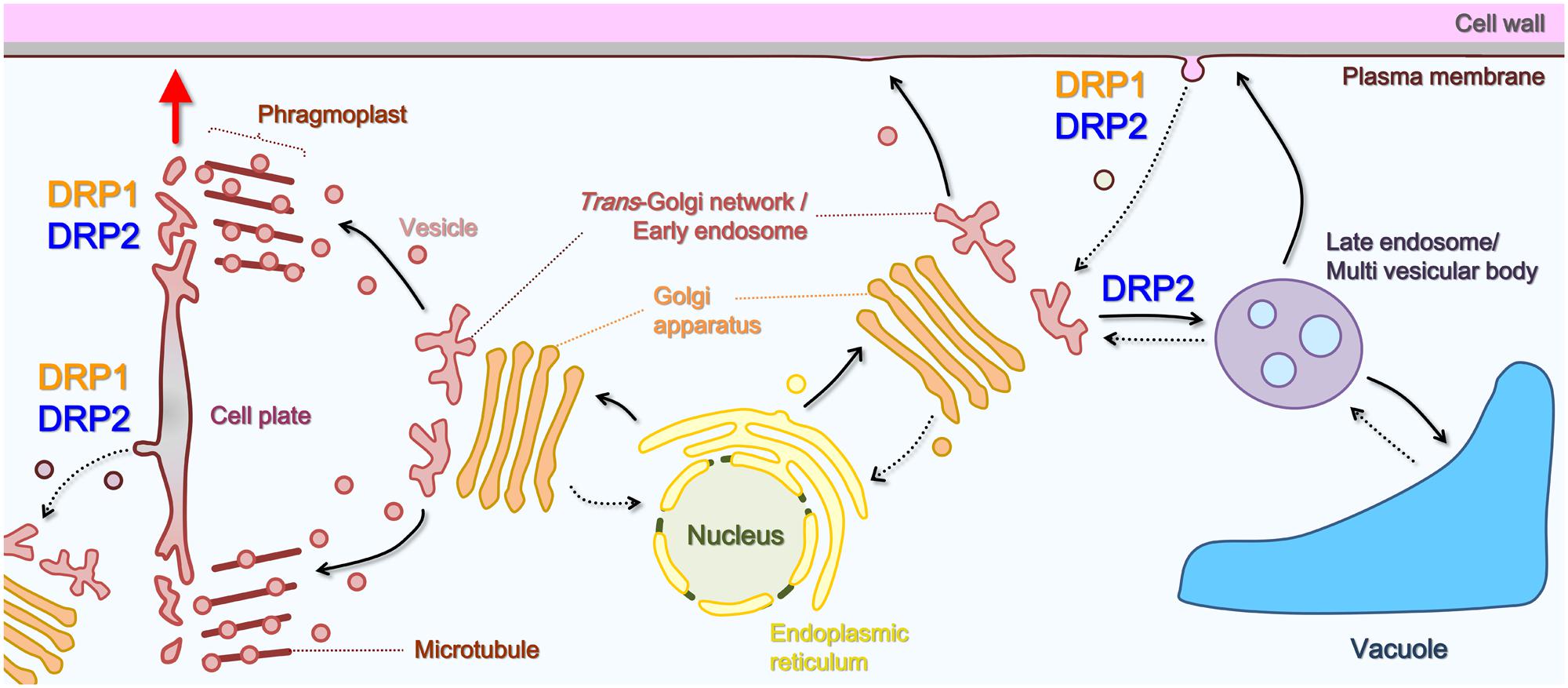
FIGURE 3. Schematic illustration of the roles of DRP1 and DRP2 in post-Golgi trafficking pathways. Solid black arrows, secretory or vacuolar pathways; dotted black arrows, endocytic or retrograde pathway; red arrow, direction of cell plate expansion.
On the other hand, genes encoding DRP2 (DRP2A and DRP2B) were initially identified in the A. thaliana genome (Mikami et al., 2000; Figure 1). DRP2s appear to be unique to embryophytes. DRP2A and DRP2B have high sequence identity (92%). They are expressed ubiquitously but most strongly around root apical meristems and vascular bundles (our unpublished results). No loss-of-function phenotypes have been detected in drp2a or drp2b mutants under laboratory conditions, which suggests that the two proteins have functional redundancy (Backues et al., 2010; Taylor, 2011). However, the double mutation of drp2a and drp2b caused defects in male and female gametogenesis with aberrant Golgi cisterna and alteration of cell wall composition and structure (Backues et al., 2010), which was also detected in a loss-of-function mutant of one of three members of the DRP2 subfamily in Oryza sativa (Hirano et al., 2010; Xiong et al., 2010). Moreover, the transient and inducible expression of mutated DRP2A and/or DRP2B, which harbor an amino acid substitution resulting in the loss of GTPase activity, have dominant-negative effects on the TGN-to-vacuole trafficking of some soluble and membrane cargoes (Jin et al., 2001) and on the uptake of FM4-64 from the PM (Taylor, 2011). These loss-of-function phenotypes imply that DRP2 participates in multiple pathways of post-Golgi trafficking (Figure 3).
Subcellular Localization and Dynamics of DRP1 and DRP2
Observations with confocal microscopy clearly showed that DRP1 is localized to the cell plate, in agreement with the finding that it is required for cell plate formation (Gu and Verma, 1996, 1997; Kang et al., 2003a,b; Collings et al., 2008). During cytokinesis, DRP1 mainly localizes to the newly synthesized edge region of the forming cell plate (Hong et al., 2003b; Fujimoto et al., 2008). Otegui et al. (2001) examined the localization of antibody-labeled A. thaliana DRP1A in dividing endosperm cells with high resolution electron tomography. They detected antibody-labeled spiral-shaped structures constricting the membranous tubular networks in the region where the cells were dividing. Interestingly, a large amount of CCVs and buds were also observed just inside the leading edge of the cell plate, which may be involved in removing and recycling excess membrane materials (Samuels et al., 1995; Otegui et al., 2001; Segui-Simarro et al., 2004). These findings suggest DRP1 is involved in membrane remodeling and vesicle formation at the forming cell plate (Figure 3).
Total internal reflection fluorescence microscopy (TIRFM) is an optical technique for observing fluorescence in the cellular surface layer very close to the cover glass (100–400 nm from the cover glass; Toomre and Manstein, 2001). In animal and yeast cells, TIRFM has provided live imaging of endocytosis-related molecules that act near the PM, such as dynamin and clathrin (Merrifield et al., 2002; Rappoport and Simon, 2003). In plant cells surrounded by a cell wall, the fine localizations and dynamics of DRP1 and DRP2 was also visualized using a TIRFM-related technique called variable angle epifluorescence microscopy (VAEM) or variable incidence angle fluorescence microscopy (VIAFM). In VIAFM images of A. thaliana root epidermal cells and cultured cells, fluorescently labeled DRP1 forms discrete and mobile foci whose diameters are 200–500 nm and partially co-localizes with fluorescent fusions of the clathrin light chain (CLC; Fujimoto et al., 2007; Konopka et al., 2008; Konopka and Bednarek, 2008; Fujimoto et al., 2010), which is similar to those of animal dynamin (Merrifield et al., 2002). The localizations and dynamics of DRP1 foci at the PM are affected by several compounds. These include some endocytic inhibitors (Konopka et al., 2008; Fujimoto et al., 2010), Tyrphostin A23, which perturbs the interaction between the cargo and the AP-2 clathrin adaptor complex (Banbury et al., 2003) and fenpropimorph, which inhibits the biosynthesis of sterols that mediate the assembly of the endocytic machinery and its cargo at the PM (He et al., 2003; Pichler and Riezman, 2004; Schrick et al., 2004). These results suggest that DRP1 is involved in the formation of endocytic vesicles (Figure 3).
Confocal and electron microscopy studies of fluorescently labeled DRP2 have shown that DRP2 has a broader localization than DRP1. Although DRP2, like DRP1, localizes to the cell plate and the PM, it also localizes to other post-Golgi organelles such as Golgi/TGN/endosomes (Jin et al., 2001; Lam et al., 2002; Fujimoto et al., 2008; Taylor, 2011). Moreover, VIAFM observations using A. thaliana root epidermal and cultured cells revealed that fluorescently labeled DRP2 also forms into discrete foci that partially co-localize with fluorescently labeled CLCs (Fujimoto et al., 2010). These observations suggest that DRP2 participates in multiple post-Golgi trafficking pathways including endocytosis (Figure 3). Interestingly, in a time-course confocal laser scanning microscopy (CLSM) analysis, fluorescently labeled DRP2 and DRP1 closely co-localized at the newly formed edge and the central matured region of the cell plate throughout cytokinesis (Fujimoto et al., 2008). Moreover, a VIAFM analysis showed that fluorescently labeled DRP2 and DRP1 largely co-localize with each other in discrete foci and assemble/disassemble together at the PM (Fujimoto et al., 2010). These spatiotemporal relationships between DRP1 and DRP2 raise the hypothesis that these two structurally different DRPs act coordinately in membrane remodeling and CCV formation during cytokinesis and endocytosis (Figure 4).
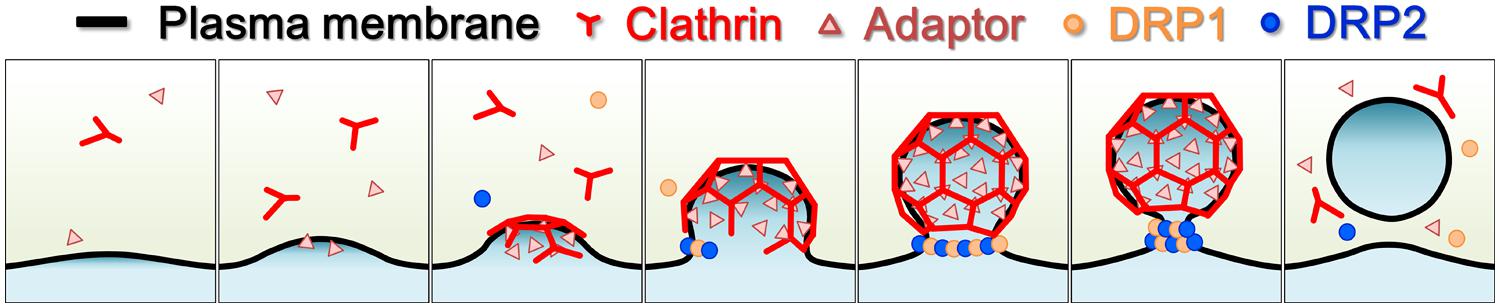
FIGURE 4. Schematic illustration of endocytic clathrin-coated vesicle formation in land plants. Light yellow, cytosol; black line, plasma membrane; red, clathrin and clathrin coat; pink triangles, clathrin adaptors; orange dots, DRP1; blue dots, DRP2.
Molecular Network Underlying the Functional Machinery for DRP1 and DRP2
As described above, dynamin in vitro polymerizes into a ring- and a spiral-shaped structure. However, in vivo, dynamin has been reported to form a functional complex with various proteins for membrane curvature sensing/generating, cargo concentrating/selecting and so on (McMahon and Boucrot, 2011). Dynamin also associates with the membrane lipids to localize to the vesicle formation site where it snips the budding membrane at its neck (Schmid and Frolov, 2011).
Genomic and phylogenetic analyses indicate that plants rarely have obvious orthologs of dynamin-interacting proteins (Holstein, 2002; Chen et al., 2011; De Craene et al., 2012; Gadeyne et al., 2014). However, these studies also show that many plant proteins contain domains for membrane curvature sensing/generation, membrane lipid association and cargo selection. In animals, such domains are found in protein–protein interaction networks centering on dynamin (McMahon and Boucrot, 2011). For example, an A. thaliana protein (VAN3) contains a Bin/amphiphysin/Rvs (BAR) domain, which recognizes and/or induces membrane curvature (Suetsugu et al., 2010). VAN3 has been shown to interact with DRP1 via yeast-two hybrid (Y2H) and co-immunoprecipitation (Co-IP; Koizumi et al., 2005; Sawa et al., 2005). VAN3, which localizes to TGN and PM (Naramoto et al., 2009, 2010) also has two other domains. One is an ADP-ribosylation factor-GTPase activating protein (ARF-GAP) domain that regulates vesicle formation in post-Golgi traffic (Spang et al., 2010). The other is a PH domain, which associates with a membrane phospholipid, phosphatidylinositol 4-phosphate (PI4P) that accumulates at TGN and PM (Koizumi et al., 2005; Simon et al., 2014). Fluorescently labeled DRP1 and VAN3 co-localize at the TGN in A. thaliana protoplasts, and a drp1a mutant has a vascularization defect that is also observed in a van3 mutant (Koizumi et al., 2005; Sawa et al., 2005). These findings imply that DRP1 also participates in multiple trafficking pathways around TGN. Moreover, DRP1 has been shown to interact with a putative trafficking cargo protein, the auxin efflux carrier PIN1, in A. thaliana via Co-IP, bimolecular fluorescence complementation (BiFC) and fluorescence resonance energy transfer (FRET; Mravec et al., 2011). DRP1 also binds to two other cargo proteins, cell plate-specific callose synthase and UDP-glucose transferase in G. max via Co-IP and Y2H (Hong et al., 2001a,b). The interactions between DRP1 and trafficking cargos raise the possibility that DRPs, which are unique to plants, function in vesicle formation, such as in selecting or concentrating the vesicle’s cargo or in sensing the progression of vesicle formation.
Some proteins have been identified as interacting partners of DRP2 via Y2H screening with the C-terminal half of A. thaliana DRP2A. One of them is AtSH3P3, which was shown to bind to the PRD of DRP2A (Lam et al., 2002). AtSH3P3 has a SH3 domain that is known to be a PRD-binding motif (Li, 2005; Kaneko et al., 2008). SH3 domains are also found in animal amphiphysins, which participate in CME through their BAR-domain (Daumke et al., 2014). The A. thaliana genome encodes three SH3-domain containing proteins. Although these proteins lack a BAR-domain, they are thought to be involved in clathrin-mediated trafficking (Lam et al., 2001). DRP2 has another proline rich region in the GED, which interacts with g-adaptin (Lam et al., 2002). g-Adaptin is a subunit of the AP1-clathrin adaptor complex that participates in late secretory and vacuolar traffic around the TGN (Park et al., 2013; Teh et al., 2013; Wang et al., 2013). The C-terminal half of A. thaliana DRP2A also interacts with yeast Sec13 homolog, AtSeh1, which specifically binds to the PH domain of DRP2, apparently to regulate its phosphoinositide interaction (Lee et al., 2006). The TPLATE complex, which is also referred to as TSET, is an endocytic clathrin adaptor that is conserved across a wide range of eukaryotes (Hirst et al., 2014). Co-IP assays have shown that some subunits of this complex interact with both DRP1 and DRP2 in A. thaliana (Gadeyne et al., 2014). This strengthens the possibility that DRP2 participates in CME in a manner similar to that of DRP1. Additional protein–protein interaction screening will be needed to elucidate the functions of DRP1 and DRP2.
Most DRP family proteins undergo homo-polymerization (Praefcke and McMahon, 2004). Y2H assays have shown that DRP1 and DRP2 interact, which indicates that structurally distinct DRPs can form hetero-polymers (Hong et al., 2003b; Fujimoto et al., 2010). As mentioned above, DRP3 and DRP5B are involved in the division of mitochondria and plastids, respectively. A Co-IP assay in A. thaliana plants raised the possibility that DRP3 and DRP5B form a heteromeric complex that is involved in the fission of peroxisomes (Zhang and Hu, 2010). However, the hetero-polymerization of DRPs belonging to different subfamilies has not been reported in other eukaryotic lineages. Further studies using Co-IP, BiFC, or FRET are needed to verify the heteropolymerization of DRP1 and DRP2.
Phosphoinositide within the cytosolic side leaflet of each post-Golgi organelle membrane participates in the assembly and disassembly of membrane traffic machinery (Krauss and Haucke, 2007; Mayinger, 2012). PI4, 5P2, which is the main binding target of the PH domain of dynamin (Ferguson et al., 1994; Zheng et al., 1996; Schmid and Frolov, 2011), functions in the recruitment and assembly of dynamin to the site of vesicle formation (Antonescu et al., 2011; Ferguson and De Camilli, 2012). Although DRP1 lacks a PH domain and other domains known to associated with phosphoinositides, liposome-binding in vitro studies suggest that A. thaliana DRP1A interacts with three kinds of phosphoinositides (Backues and Bednarek, 2010). One is phosphatidylinositol 3-phosphate (PI3P), which is localized in endosomes/prevacuolar compartments (PVCs) and the vacuole (Vermeer et al., 2006; Simon et al., 2014). Another is PI4P, which accumulates mainly in the TGN, PM and the newly synthesized part of the cell plate (Vermeer et al., 2009; Simon et al., 2014). The third is PI5P, whose subcellular localization is unclear (Krauss and Haucke, 2007). In contrast to DRP1, A. thaliana DRP2 has a PH domain. Protein–lipid overlay assays and liposome sedimentation assays suggest that the PH domain interacts with PI3P (Lee et al., 2002; Backues and Bednarek, 2010), PI4P (Lam et al., 2002; Lee et al., 2002; Backues and Bednarek, 2010), and PI4, 5P2 (Lam et al., 2002) which is localized specifically to the PM (Simon et al., 2014). These multiple phosphoinositide interactions of DRP2 are consistent with its wide distribution in post-Golgi organelles. However, the effects of multiple phosphoinositide interactions of DRP1, which has a narrower distribution than DRP2, on its subcellular localization remain unclear. Further studies are needed to understand the changes in the subcellular localizations of these two DRPs and phosphoinositide during developmental and physiological acclimation.
Perspectives
The above findings show that DRP1 and DRP2 share several properties, including subcellular co-localization, intermolecular interaction, and membrane phospholipid binding similarity. This raises the possibility that DRP1 and DRP2 act together in a heteromeric complex to form the vesicles used in post-Golgi traffic (Figure 4). They also appear to work together in cell plate formation, in contrast to metazoan dynamins which perform this function by themselves. However, this hetero-polymerization of structurally distinct DRPs has so far been observed only in embryophytes. This raises three major questions about the embryophyte-unique hetero-DRP complex. First, how does this heteromeric complex bend or sever the organelle membrane? To answer this question, the effects of DRP1 and DRP2 and their GTPase-defective mutants on the morphology of the lipid bilayer need to be compared both in vitro and in vivo. The tertiary structures of DRP1 and DRP2 also need to be determined. Second, if a heteromeric complex of DRP1 and DRP2 has roles in both membrane remodeling and severing, how does it switch from one role to the other? To answer this question, studies are needed to determine the molecular ratios of DRP1 and DRP2 in the heteromeric complex at the leading edge and in the mature region of the cell plate and in other post-Golgi organelles. DRP1 and DRP2 have different kinetic velocities of GTPase activity (Lam et al., 2002; Sawa et al., 2005), which may be due to a structural difference in the GTPase domain. Third, what is the reason for the evolutionary development of a heteromeric complex in embryophytes? Three studies are needed to answer this question: determine the phenotype caused by the partial-loss of DRP2 function, identify other proteins that interact with DRP2 and clarify how vesicles are formed in Chlorophyta, which do not have DRP2.
In addition to questions about the embryophyte-unique DRP complex, two fundamental questions about DRP assembly remain to be solved. First, does the DRP that is involved in post-Golgi traffic really form a ring or spiral shaped polymers and sever the membrane in a GTP hydrolysis-dependent constriction? Second, if this is the case, do DRP proteins assemble directionally or randomly? As mentioned above, the current action model of DRP is mainly based on electron microscopic observation and biochemical analyses of metazoan dynamins. Recently, some high resolution microscopy techniques that overcome the diffraction barrier in existing light microscopy have become available. These include structured illumination microscopy (SIM; Gustafsson, 2005; Gustafsson et al., 2008), stimulated emission depletion microscopy (STED; Toomre and Bewersdorf, 2010), and super-resolution confocal live imaging microscopy (SCLIM; Ito et al., 2012). These techniques, when applied to fluorescent live cell imaging, have the potential to reveal the fine structure and behavior of the molecular machinery regulating organelle dynamics. They should also help to clarify the mechanisms of DRP assembly and constriction, which are the core functions of DRPs.
Conflict of Interest Statement
The Guest Associate, Dr. Takashi Ueda, declares that, despite being affiliated to the same institution as author(s), the review process was handled objectively and no conflict of interest exists. The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Antonescu, C. N., Aguet, F., Danuser, G., and Schmid, S. L. (2011). Phosphatidylinositol-(4,5)-bisphosphate regulates clathrin-coated pit initiation, stabilization, and size. Mol. Biol. Cell 22, 2588–2600. doi: 10.1091/mbc.E11-04-0362
Backues, S. K., and Bednarek, S. Y. (2010). Arabidopsis dynamin-related protein 1A polymers bind, but do not tubulate, liposomes. Biochem. Biophys. Res. Commun. 393, 734–739. doi: 10.1016/j.bbrc.2010.02.070
Backues, S. K., Korasick, D. A., Heese, A., and Bednarek, S. Y. (2010). The Arabidopsis dynamin-related protein2 family is essential for gametophyte development. Plant Cell 22, 3218–3231. doi: 10.1105/tpc.110.077727
Banbury, D. N., Oakley, J. D., Sessions, R. B., and Banting, G. (2003). Tyrphostin A23 inhibits internalization of the transferrin receptor by perturbing the interaction between tyrosine motifs and the medium chain subunit of the AP-2 adaptor complex. J. Biol. Chem. 278, 12022–12028. doi: 10.1074/jbc.M211966200
Bednarek, S. Y., and Backues, S. K. (2010). Plant dynamin-related protein families DRP1 and DRP2 in plant development. Biochem. Soc. Trans. 38, 797–806. doi: 10.1042/BST0380797
Bonifacino, J. S., and Glick, B. S. (2004). The mechanisms of vesicle budding and fusion. Cell 116, 153–166. doi: 10.1016/S0092-8674(03)01079-1
Boutte, Y., Frescatada-Rosa, M., Men, S., Chow, C. M., Ebine, K., Gustavsson, A.,et al. (2010). Endocytosis restricts Arabidopsis KNOLLE syntaxin to the cell division plane during late cytokinesis. EMBO J. 29, 546–558. doi: 10.1038/emboj.2009.363
Chanez, A. L., Hehl, A. B., Engstler, M., and Schneider, A. (2006). Ablation of the single dynamin of T. brucei blocks mitochondrial fission and endocytosis and leads to a precise cytokinesis arrest. J. Cell Sci. 119, 2968–2974. doi: 10.1242/jcs.03023
Chappie, J. S., Acharya, S., Liu, Y. W., Leonard, M., Pucadyil, T. J., and Schmid, S. L. (2009). An intramolecular signaling element that modulates dynamin function in vitro and in vivo. Mol. Biol. Cell 20, 3561–3571. doi: 10.1091/mbc.E09-04-0318
Chappie, J. S., and Dyda, F. (2013). Building a fission machine – structural insights into dynamin assembly and activation. J. Cell Sci. 126, 2773–2784. doi: 10.1242/jcs.108845
Chappie, J. S., Mears, J. A., Fang, S., Leonard, M., Schmid, S. L., Milligan, R. A.,et al. (2011). A pseudoatomic model of the dynamin polymer identifies a hydrolysis-dependent powerstroke. Cell 147, 209–222. doi: 10.1016/j.cell.2011.09.003
Chen, X., Irani, N. G., and Friml, J. (2011). Clathrin-mediated endocytosis: the gateway into plant cells. Curr. Opin. Plant Biol. 14, 674–682. doi: 10.1016/j.pbi.2011.08.006
Collings, D. A., Gebbie, L. K., Howles, P. A., Hurley, U. A., Birch, R. J., Cork, A. H.,et al. (2008). Arabidopsis dynamin-like protein DRP1A: a null mutant with widespread defects in endocytosis, cellulose synthesis, cytokinesis, and cell expansion. J. Exp. Bot. 59, 361–376. doi: 10.1093/jxb/erm324
Danino, D., Moon, K. H., and Hinshaw, J. E. (2004). Rapid constriction of lipid bilayers by the mechanochemical enzyme dynamin. J. Struct. Biol. 147, 259–267. doi: 10.1016/j.jsb.2004.04.005
Daumke, O., Roux, A., and Haucke, V. (2014). BAR domain scaffolds in dynamin-mediated membrane fission. Cell 156, 882–892. doi: 10.1016/j.cell.2014.02.017
De Craene, J. O., Ripp, R., Lecompte, O., Thompson, J. D., Poch, O., and Friant, S. (2012). Evolutionary analysis of the ENTH/ANTH/VHS protein superfamily reveals a coevolution between membrane trafficking and metabolism. BMC Genomics 13:297. doi: 10.1186/1471-2164-13-297
Dombrowski, J. E., and Raikhel, N. V. (1995). Isolation of a cDNA encoding a novel GTP-binding protein of Arabidopsis thaliana. Plant Mol. Biol. 28, 1121–1126. doi: 10.1007/BF00032672
Faelber, K., Posor, Y., Gao, S., Held, M., Roske, Y., Schulze, D.,et al. (2011). Crystal structure of nucleotide-free dynamin. Nature 477, 556–560. doi: 10.1038/nature10369
Ferguson, K. M., Lemmon, M. A., Schlessinger, J., and Sigler, P. B. (1994). Crystal structure at 2.2 A resolution of the pleckstrin homology domain from human dynamin. Cell 79, 199–209. doi: 10.1016/0092-8674(94)90190-2
Ferguson, S. M., and De Camilli, P. (2012). Dynamin, a membrane-remodelling GTPase. Nat. Rev. Mol. Cell Biol. 13, 75–88. doi: 10.1038/nrm3266
Ford, M. G., Jenni, S., and Nunnari, J. (2011). The crystal structure of dynamin. Nature 477, 561–566. doi: 10.1038/nature10441
Fujimoto, M., Arimura, S., Mano, S., Kondo, M., Saito, C., Ueda, T.,et al. (2009). Arabidopsis dynamin-related proteins DRP3A and DRP3B are functionally redundant in mitochondrial fission, but have distinct roles in peroxisomal fission. Plant J. 58, 388–400. doi: 10.1111/j.1365-313X.2009.03786.x
Fujimoto, M., Arimura, S., Nakazono, M., and Tsutsumi, N. (2007). Imaging of plant dynamin-related proteins and clathrin around the plasma membrane by variable incidence angle fluorescence microscopy. Plant Biotechnol. 24, 449–455. doi: 10.5511/plantbiotechnology.24.449
Fujimoto, M., Arimura, S., Nakazono, M., and Tsutsumi, N. (2008). Arabidopsis dynamin-related protein DRP2B is co-localized with DRP1A on the leading edge of the forming cell plate. Plant Cell Rep. 27, 1581–1586. doi: 10.1007/s00299-008-0583-0
Fujimoto, M., Arimura, S., Ueda, T., Takanashi, H., Hayashi, Y., Nakano, A.,et al. (2010). Arabidopsis dynamin-related proteins DRP2B and DRP1A participate together in clathrin-coated vesicle formation during endocytosis. Proc. Natl. Acad. Sci. U.S.A. 107, 6094–6099. doi: 10.1073/pnas.0913562107
Gadeyne, A., Sanchez-Rodriguez, C., Vanneste, S., Di Rubbo, S., Zauber, H., Vanneste, K.,et al. (2014). The TPLATE adaptor complex drives clathrin-mediated endocytosis in plants. Cell 156, 691–704. doi: 10.1016/j.cell.2014.01.039
Gao, H., Kadirjan-Kalbach, D., Froehlich, J. E., and Osteryoung, K. W. (2003). ARC5, a cytosolic dynamin-like protein from plants, is part of the chloroplast division machinery. Proc. Natl. Acad. Sci. U.S.A. 100, 4328–4333. doi: 10.1073/pnas.0530206100
Grabs, D., Slepnev, V. I., Songyang, Z., David, C., Lynch, M., Cantley, L. C.,et al. (1997). The SH3 domain of amphiphysin binds the proline-rich domain of dynamin at a single site that defines a new SH3 binding consensus sequence. J. Biol. Chem. 272, 13419–13425. doi: 10.1074/jbc.272.20.13419
Gu, X., and Verma, D. P. (1996). Phragmoplastin, a dynamin-like protein associated with cell plate formation in plants. EMBO J. 15, 695–704.
Gu, X., and Verma, D. P. (1997). Dynamics of phragmoplastin in living cells during cell plate formation and uncoupling of cell elongation from the plane of cell division. Plant Cell 9, 157–169. doi: 10.1105/tpc.9.2.157
Gustafsson, M. G. (2005). Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl. Acad. Sci. U.S.A. 102, 13081–13086. doi: 10.1073/pnas.0406877102
Gustafsson, M. G., Shao, L., Carlton, P. M., Wang, C. J., Golubovskaya, I. N., Cande, W. Z.,et al. (2008). Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys. J. 94, 4957–4970. doi: 10.1529/biophysj.107.120345
He, J. X., Fujioka, S., Li, T. C., Kang, S. G., Seto, H., Takatsuto, S.,et al. (2003). Sterols regulate development and gene expression in Arabidopsis. Plant Physiol. 131, 1258–1269. doi: 10.1104/pp.014605
Heymann, J. A. W., and Hinshaw, J. E. (2009). Dynamins at a glance. J. Cell Sci. 122, 3427–3431. doi: 10.1242/jcs.051714
Hinshaw, J. E. (2000). Dynamin and its role in membrane fission. Annu. Rev. Cell Dev. Biol. 16, 483–519. doi: 10.1146/annurev.cellbio.16.1.483
Hirano, K., Kotake, T., Kamihara, K., Tsuna, K., Aohara, T., Kaneko, Y.,et al. (2010). Rice BRITTLE CULM 3 (BC3) encodes a classical dynamin OsDRP2B essential for proper secondary cell wall synthesis. Planta 232, 95–108. doi: 10.1007/s00425-010-1145-6
Hirst, J., Schlacht, A., Norcott, J. P., Traynor, D., Bloomfield, G., Antrobus, R.,et al. (2014). Characterization of TSET, an ancient and widespread membrane trafficking complex. Elife (Cambridge) 3, e02866. doi: 10.7554/eLife.02866
Holstein, S. E. (2002). Clathrin and plant endocytosis. Traffic 3, 614–620. doi: 10.1034/j.1600-0854.2002.30903.x
Hong, Z., Bednarek, S. Y., Blumwald, E., Hwang, I., Jurgens, G., Menzel, D.,et al. (2003a). A unified nomenclature for Arabidopsis dynamin-related large GTPases based on homology and possible functions. Plant Mol. Biol. 53, 261–265. doi: 10.1023/B:PLAN.0000007000.29697.81
Hong, Z., Geisler-Lee, C. J., Zhang, Z., and Verma, D. P. (2003b). Phragmoplastin dynamics: multiple forms, microtubule association and their roles in cell plate formation in plants. Plant Mol. Biol. 53, 297–312. doi: 10.1023/B:PLAN.0000006936.50532.3a
Hong, Z., Delauney, A. J., and Verma, D. P. (2001a). A cell plate-specific callose synthase and its interaction with phragmoplastin. Plant Cell 13, 755–768. doi: 10.1105/tpc.13.4.755
Hong, Z., Zhang, Z., Olson, J. M., and Verma, D. P. (2001b). A novel UDP-glucose transferase is part of the callose synthase complex and interacts with phragmoplastin at the forming cell plate. Plant Cell 13, 769–779. doi: 10.1105/tpc.13.4.769
Ito, Y., Uemura, T., Shoda, K., Fujimoto, M., Ueda, T., and Nakano, A. (2012). cis-Golgi proteins accumulate near the ER exit sites and act as the scaffold for Golgi regeneration after brefeldin A treatment in tobacco BY-2 cells. Mol. Biol. Cell 23, 3203–3214. doi: 10.1091/mbc.E12-01-0034
Jin, J. B., Kim, Y. A., Kim, S. J., Lee, S. H., Kim, D. H., Cheong, G. W.,et al. (2001). A new dynamin-like protein, ADL6, is involved in trafficking from the trans-Golgi network to the central vacuole in Arabidopsis. Plant Cell 13, 1511–1526. doi: 10.1105/tpc.13.7.1511
Kaneko, T., Li, L., and Li, S. S. (2008). The SH3 domain – a family of versatile peptide- and protein-recognition module. Front. Biosci. 13:4938–4952. doi: 10.2741/3053
Kang, B. H., Busse, J. S., and Bednarek, S. Y. (2003a). Members of the Arabidopsis dynamin-like gene family, ADL1, are essential for plant cytokinesis and polarized cell growth. Plant Cell 15, 899–913. doi: 10.1105/tpc.009670
Kang, B. H., Rancour, D. M., and Bednarek, S. Y. (2003b). The dynamin-like protein ADL1C is essential for plasma membrane maintenance during pollen maturation. Plant J. 35, 1–15. doi: 10.1046/j.1365-313X.2003.01775.x
Kang, B. H., Busse, J. S., Dickey, C., Rancour, D. M., and Bednarek, S. Y. (2001). The Arabidopsis cell plate-associated dynamin-like protein, ADL1Ap, is required for multiple stages of plant growth and development. Plant Physiol. 126, 47–68. doi: 10.1104/pp.126.1.47
Koizumi, K., Naramoto, S., Sawa, S., Yahara, N., Ueda, T., Nakano, A.,et al. (2005). VAN3 ARF-GAP-mediated vesicle transport is involved in leaf vascular network formation. Development 132, 1699–1711. doi: 10.1242/dev.01716
Konopka, C. A., Backues, S. K., and Bednarek, S. Y. (2008). Dynamics of Arabidopsis dynamin-related protein 1C and a clathrin light chain at the plasma membrane. Plant Cell 20, 1363–1380. doi: 10.1105/tpc.108.059428
Konopka, C. A., and Bednarek, S. Y. (2008). Comparison of the dynamics and functional redundancy of the Arabidopsis dynamin-related isoforms DRP1A and DRP1C during plant development. Plant Physiol. 147, 1590–1602. doi: 10.1104/pp.108.116863
Krauss, M., and Haucke, V. (2007). Phosphoinositides: regulators of membrane traffic and protein function. FEBS Lett. 581, 2105–2111. doi: 10.1016/j.febslet.2007.01.089
Lam, B. C., Sage, T. L., Bianchi, F., and Blumwald, E. (2001). Role of SH3 domain-containing proteins in clathrin-mediated vesicle trafficking in Arabidopsis. Plant Cell 13, 2499–2512. doi: 10.1105/tpc.13.11.2499
Lam, B. C., Sage, T. L., Bianchi, F., and Blumwald, E. (2002). Regulation of ADL6 activity by its associated molecular network. Plant J. 31, 565–576. doi: 10.1046/j.1365-313X.2002.01377.x
Lee, M. H., Lee, S. H., Kim, H., Jin, J. B., Kim, D. H., and Hwang, I. (2006). A WD40 repeat protein, Arabidopsis Sec13 homolog 1, may play a role in vacuolar trafficking by controlling the membrane association of AtDRP2A. Mol. Cells 22, 210–219.
Lee, S. H., Jin, J. B., Song, J., Min, M. K., Park, D. S., Kim, Y. W.,et al. (2002). The intermolecular interaction between the PH domain and the C-terminal domain of Arabidopsis dynamin-like 6 determines lipid binding specificity. J. Biol. Chem. 277, 31842–31849. doi: 10.1074/jbc.M204770200
Li, S. S. (2005). Specificity and versatility of SH3 and other proline-recognition domains: structural basis and implications for cellular signal transduction. Biochem. J. 390, 641–653. doi: 10.1042/BJ20050411
Lopez-Bautista, J. M., Waters, D. A., and Chapman, R. L. (2003). Phragmoplastin, green algae and the evolution of cytokinesis. Int. J. Syst. Evol. Microbiol. 53, 1715–1718. doi: 10.1099/ijs.0.02561-0
Macia, E., Ehrlich, M., Massol, R., Boucrot, E., Brunner, C., and Kirchhausen, T. (2006). Dynasore, a cell-permeable inhibitor of dynamin. Dev. Cell 10, 839–850. doi: 10.1016/j.devcel.2006.04.002
Mano, S., Nakamori, C., Kondo, M., Hayashi, M., and Nishimura, M. (2004). An Arabidopsis dynamin-related protein, DRP3A, controls both peroxisomal and mitochondrial division. Plant J. 38, 487–498. doi: 10.1111/j.1365-313X.2004.02063.x
Mayinger, P. (2012). Phosphoinositides and vesicular membrane traffic. Biochim. Biophys. Acta 1821, 1104–1113. doi: 10.1016/j.bbalip.2012.01.002
McMahon, H. T., and Boucrot, E. (2011). Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 12, 517–533. doi: 10.1038/nrm3151
McMahon, H. T., and Mills, I. G. (2004). COP and clathrin-coated vesicle budding: different pathways, common approaches. Curr. Opin. Cell Biol. 16, 379–391. doi: 10.1016/j.ceb.2004.06.009
McNew, J. A., Sondermann, H., Lee, T., Stern, M., and Brandizzi, F. (2013). GTP-dependent membrane fusion. Annu. Rev. Cell Dev. Biol. 29, 529–550. doi: 10.1146/annurev-cellbio-101512-122328
Merrifield, C. J., Feldman, M. E., Wan, L., and Almers, W. (2002). Imaging actin and dynamin recruitment during invagination of single clathrin-coated pits. Nat. Cell Biol. 4, 691–698. doi: 10.1038/ncb837
Mikami, K., Iuchi, S., Yamaguchi-Shinozaki, K., and Shinozaki, K. (2000). A novel Arabidopsis thaliana dynamin-like protein containing the pleckstrin homology domain. J. Exp. Bot. 51, 317–318. doi: 10.1093/jexbot/51.343.317
Miyagishima, S. Y., Kuwayama, H., Urushihara, H., and Nakanishi, H. (2008). Evolutionary linkage between eukaryotic cytokinesis and chloroplast division by dynamin proteins. Proc. Natl. Acad. Sci. U.S.A. 105, 15202–15207. doi: 10.1073/pnas.0802412105
Miyagishima, S. Y., Nishida, K., Mori, T., Matsuzaki, M., Higashiyama, T., Kuroiwa, H.,et al. (2003). A plant-specific dynamin-related protein forms a ring at the chloroplast division site. Plant Cell 15, 655–665. doi: 10.1105/tpc.009373
Mravec, J., Petrasek, J., Li, N., Boeren, S., Karlova, R., Kitakura, S.,et al. (2011). Cell plate restricted association of DRP1A and PIN proteins is required for cell polarity establishment in Arabidopsis. Curr. Biol. 21, 1055–1060. doi: 10.1016/j.cub.2011.05.018
Naramoto, S., Kleine-Vehn, J., Robert, S., Fujimoto, M., Dainobu, T., Paciorek, T.,et al. (2010). ADP-ribosylation factor machinery mediates endocytosis in plant cells. Proc. Natl. Acad. Sci. U.S.A. 107, 21890–21895. doi: 10.1073/pnas.1016260107
Naramoto, S., Sawa, S., Koizumi, K., Uemura, T., Ueda, T., Friml, J.,et al. (2009). Phosphoinositide-dependent regulation of VAN3 ARF-GAP localization and activity essential for vascular tissue continuity in plants. Development 136, 1529–1538. doi: 10.1242/dev.030098
Otegui, M. S., Mastronarde, D. N., Kang, B. H., Bednarek, S. Y., and Staehelin, L. A. (2001). Three-dimensional analysis of syncytial-type cell plates during endosperm cellularization visualized by high resolution electron tomography. Plant Cell 13, 2033–2051. doi: 10.1105/tpc.13.9.2033
Park, M., Song, K., Reichardt, I., Kim, H., Mayer, U., Stierhof, Y. D.,et al. (2013). Arabidopsis mu-adaptin subunit AP1M of adaptor protein complex 1 mediates late secretory and vacuolar traffic and is required for growth. Proc. Natl. Acad. Sci. U.S.A. 110, 10318–10323. doi: 10.1073/pnas.1300460110
Pichler, H., and Riezman, H. (2004). Where sterols are required for endocytosis. Biochim. Biophys. Acta 1666, 51–61. doi: 10.1016/j.bbamem.2004.05.011
Praefcke, G. J., and McMahon, H. T. (2004). The dynamin superfamily: universal membrane tubulation and fission molecules? Nat. Rev. Mol. Cell Biol. 5, 133–147. doi: 10.1038/nrm1313
Ramachandran, R., Surka, M., Chappie, J. S., Fowler, D. M., Foss, T. R., Song, B. D.,et al. (2007). The dynamin middle domain is critical for tetramerization and higher-order self-assembly. EMBO J. 26, 559–566. doi: 10.1038/sj.emboj.7601491
Rappoport, J. Z., and Simon, S. M. (2003). Real-time analysis of clathrin-mediated endocytosis during cell migration. J. Cell Sci. 116, 847–855. doi: 10.1242/jcs.00289
Samuels, A. L., Giddings, T. H. Jr., and Staehelin, L. A. (1995). Cytokinesis in tobacco BY-2 and root tip cells: a new model of cell plate formation in higher plants. J. Cell Biol. 130, 1345–1357. doi: 10.1083/jcb.130.6.1345
Sawa, S., Koizumi, K., Naramoto, S., Demura, T., Ueda, T., Nakano, A.,et al. (2005). DRP1A is responsible for vascular continuity synergistically working with VAN3 in Arabidopsis. Plant Physiol. 138, 819–826. doi: 10.1104/pp.105.061689
Schmid, S. L., and Frolov, V. A. (2011). Dynamin: functional design of a membrane fission catalyst. Annu. Rev. Cell Dev. Biol. 27, 79–105. doi: 10.1146/annurev-cellbio-100109-104016
Schrick, K., Fujioka, S., Takatsuto, S., Stierhof, Y. D., Stransky, H., Yoshida, S.,et al. (2004). A link between sterol biosynthesis, the cell wall, and cellulose in Arabidopsis. Plant J. 38, 227–243. doi: 10.1111/j.1365-313X.2004.02039.x
Segui-Simarro, J. M., Austin, J. R. II, White, E. A., and Staehelin, L. A. (2004). Electron tomographic analysis of somatic cell plate formation in meristematic cells of Arabidopsis preserved by high-pressure freezing. Plant Cell 16, 836–856. doi: 10.1105/tpc.017749
Sever, S., Muhlberg, A. B., and Schmid, S. L. (1999). Impairment of dynamin’s GAP domain stimulates receptor-mediated endocytosis. Nature 398, 481–486. doi: 10.1038/19024
Shpetner, H. S., Herskovits, J. S., and Vallee, R. B. (1996). A binding site for SH3 domains targets dynamin to coated pits. J. Biol. Chem. 271, 13–16. doi: 10.1074/jbc.271.1.13
Simon, M. L., Platre, M. P., Assil, S., Van Wijk, R., Chen, W. Y., Chory, J.,et al. (2014). A multi-colour/multi-affinity marker set to visualize phosphoinositide dynamics in Arabidopsis. Plant J. 77, 322–337. doi: 10.1111/tpj.12358
Spang, A., Shiba, Y., and Randazzo, P. A. (2010). Arf GAPs: gatekeepers of vesicle generation. FEBS Lett. 584, 2646–2651. doi: 10.1016/j.febslet.2010.04.005
Stowell, M. H., Marks, B., Wigge, P., and Mcmahon, H. T. (1999). Nucleotide-dependent conformational changes in dynamin: evidence for a mechanochemical molecular spring. Nat. Cell Biol. 1, 27–32. doi: 10.1038/8997
Suetsugu, S., Toyooka, K., and Senju, Y. (2010). Subcellular membrane curvature mediated by the BAR domain superfamily proteins. Semin. Cell Dev. Biol. 21, 340–349. doi: 10.1016/j.semcdb.2009.12.002
Sweitzer, S. M., and Hinshaw, J. E. (1998). Dynamin undergoes a GTP-dependent conformational change causing vesiculation. Cell 93, 1021–1029. doi: 10.1016/S0092-8674(00)81207-6
Takei, K., Mcpherson, P. S., Schmid, S. L., and De Camilli, P. (1995). Tubular membrane invaginations coated by dynamin rings are induced by GTP-gamma S in nerve terminals. Nature 374, 186–190. doi: 10.1038/374186a0
Taylor, N. G. (2011). A role for Arabidopsis dynamin related proteins DRP2A/B in endocytosis; DRP2 function is essential for plant growth. Plant Mol. Biol. 76, 117–129. doi: 10.1007/s11103-011-9773-1
Teh, O. K., Shimono, Y., Shirakawa, M., Fukao, Y., Tamura, K., Shimada, T.,et al. (2013). The AP-1 mu adaptin is required for KNOLLE localization at the cell plate to mediate cytokinesis in Arabidopsis. Plant Cell Physiol. 54, 838–847. doi: 10.1093/pcp/pct048
Toomre, D., and Bewersdorf, J. (2010). A new wave of cellular imaging. Annu. Rev. Cell Dev. Biol. 26, 285–314. doi: 10.1146/annurev-cellbio-100109-104048
Toomre, D., and Manstein, D. J. (2001). Lighting up the cell surface with evanescent wave microscopy. Trends Cell Biol. 11, 298–303. doi: 10.1016/S0962-8924(01)02027-X
Vermeer, J. E., Thole, J. M., Goedhart, J., Nielsen, E., Munnik, T., and Gadella, T. W. Jr. (2009). Imaging phosphatidylinositol 4-phosphate dynamics in living plant cells. Plant J. 57, 356–372. doi: 10.1111/j.1365-313X.2008.03679.x
Vermeer, J. E., Van Leeuwen, W., Tobena-Santamaria, R., Laxalt, A. M., Jones, D. R., Divecha, N.,et al. (2006). Visualization of PtdIns3P dynamics in living plant cells. Plant J. 47, 687–700. doi: 10.1111/j.1365-313X.2006.02830.x
Wang, J. G., Li, S., Zhao, X. Y., Zhou, L. Z., Huang, G. Q., Feng, C.,et al. (2013). HAPLESS13, the Arabidopsis mu1 adaptin, is essential for protein sorting at the trans-Golgi network/early endosome. Plant Physiol. 162, 1897–1910. doi: 10.1104/pp.113.221051
Xiong, G., Li, R., Qian, Q., Song, X., Liu, X., Yu, Y.,et al. (2010). The rice dynamin-related protein DRP2B mediates membrane trafficking, and thereby plays a critical role in secondary cell wall cellulose biosynthesis. Plant J. 64, 56–70. doi: 10.1111/j.1365-313X.2010.04308.x
Zhang, X., and Hu, J. (2010). The Arabidopsis chloroplast division protein DYNAMIN-RELATED PROTEIN5B also mediates peroxisome division. Plant Cell 22, 431–442. doi: 10.1105/tpc.109.071324
Keywords: dynamin, dynamin-related protein, plant, post-Golgi traffic, cytokinesis
Citation: Fujimoto M and Tsutsumi N (2014) Dynamin-related proteins in plant post-Golgi traffic. Front. Plant Sci. 5:408. doi: 10.3389/fpls.2014.00408
Received: 26 June 2014; Accepted: 31 July 2014;
Published online: 04 September 2014.
Edited by:
Takashi Ueda, The University of Tokyo, JapanCopyright © 2014 Fujimoto and Tsutsumi. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Masaru Fujimoto, Laboratory of Plant Molecular Genetics, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan e-mail:YW1mdWppQG1haWwuZWNjLnUtdG9reW8uYWMuanA=
 Masaru Fujimoto
Masaru Fujimoto Nobuhiro Tsutsumi
Nobuhiro Tsutsumi