- 1State Key Laboratory of Systematic and Evolutionary Botany, Institute of Botany, Chinese Academy of Sciences, Beijing, China
- 2University of Chinese Academy of Sciences, Beijing, China
- 3State Key Laboratory of Genetic Engineering and Collaborative Innovation Center for Genetics and Development, Ministry of Education Key Laboratory of Biodiversity Sciences and Ecological Engineering, Institute of Plant Biology, Institute of Biodiversity Science, Center for Evolutionary Biology, School of Life Sciences, Fudan University, Shanghai, China
- 4Shanghai Key Lab of Intelligent Information Processing and School of Computer Science, Fudan University, Shanghai, China
- 5Institutes of Biomedical Sciences, Fudan University, Shanghai, China
Meiosis is a special type of cell division process necessary for the sexual reproduction of all eukaryotes. The ever expanding meiosis research calls for an effective and specialized database that is not readily available yet. To fill this gap, we have developed a knowledge database MeioBase (http://meiosis.ibcas.ac.cn), which is comprised of two core parts, Resources and Tools. In the Resources part, a wealth of meiosis data collected by curation and manual review from published literatures and biological databases are integrated and organized into various sections, such as Cytology, Pathway, Species, Interaction, and Expression. In the Tools part, some useful tools have been integrated into MeioBase, such as Search, Download, Blast, Comparison, My Favorites, Submission, and Advice. With a simplified and efficient web interface, users are able to search against the database with gene model IDs or keywords, and batch download the data for local investigation. We believe that MeioBase can greatly facilitate the researches related to meiosis.
Introduction
Meiosis is a specialized cell division process essential for all sexually reproducing organisms. During meiosis, a single round of DNA replication is followed by two successive rounds of nuclear division, meiosis I and meiosis II. Meiosis I is unique and involves the segregation of homologous chromosomes (homologs), whereas meiosis II is similar to mitosis and results in the segregation of sister chromatids. The function of meiosis is to generate four haploid gametes, which are able to develop into germ cells. Fertilization of the germ cells, then, can restore the offspring to the chromosome number and complexity level of their parents (Zickler and Kleckner, 1998; Hamant et al., 2006). Meiosis not only ensures the stability of chromosome numbers between generations, but also provides genetic materials for biodiversity.
Studies of meiosis have been carried out extensively for over 100 years (Hamant et al., 2006). Chromosome behaviors in some species have been examined in detail by using cytological approaches (Orr-Weaver, 1995; Zickler and Kleckner, 1999; Ma, 2005; Birchler and Han, 2013). Over the last two decades, much efforts have been devoted to understanding the genetic basis and molecular mechanisms of meiosis in model species, such as nematode (Caenorhabditis elegans), budding yeast (Saccharomyces cerevisiae), Arabidopsis (Arabidopsis thaliana), rice (Oryza sativa), and maize (Zea mays). Genes regulating meiosis, especially those involved in homologous chromosome paring, synapsis, recombination and separation in prophase I, have also been cloned and characterized in terms of their functions (Hollingsworth et al., 1990; Sym et al., 1993; Keeney et al., 1997; Klimyuk and Jones, 1997; Yang et al., 1999; Li et al., 2004; Tang et al., 2014). Recent advances in transcriptomics, protein-protein interactions (PPIs), and phylogenetic analyses of genes and gene families related to meiosis have improved our understanding of this complex process dramatically (Kee and Keeney, 2002; Wang et al., 2004; Lin et al., 2006; Vignard et al., 2007; Chen et al., 2010; Tang et al., 2010; Yang et al., 2011; Dukowic-Schulze et al., 2014).
The ever expanding studies of meiosis and the data accumulated, which are scattered in tremendously diverse literatures and a few of large databases, such as NCBI, Ensemble, and TAIR, call for an integrative and encyclopedia-like platform for meiosis (Hamant et al., 2006; Handel and Schimenti, 2010; Luo et al., 2014). Recently, two databases related to reproductive development, GermOnline and Plant Male Reproduction Database (PMRD), have been established. GermOnline is a cross-species microarray expression database focusing on germline development, reproduction and cancer (Lardenois et al., 2010). PMRD is a comprehensive resource for browsing and retrieving knowledge about genes and mutants related to plant male reproduction (Cui et al., 2012). Notwithstanding, neither databases provide comprehensive information about meiosis, because data related to meiosis is heavily fragmented. Therefore, researchers are in great need of effective tools or databases to quickly obtain precise meiotic data from the exponentially increasing amount of information.
Here, by collecting and integrating all sorts of information related to meiosis, as well as including and developing powerful tools for search, comparison and analysis, we established a comprehensive and specialized database for meiosis, MeioBase. It will not only serve the meiosis research community, but also help any users who need an easy and efficient access to various kinds of information related to meiosis.
Database Structure and Web Interface
The database intends to provide all necessary resources and tools for meiosis-related researches (Figure 1). In the Resources part, information related to meiosis are categorized and integrated into five major sections (i.e., Cytology, Pathway, Species, Interaction, and Expression). In the Tools part, useful functions for searching, analyzing, uploading, and downloading data are included in seven sections (i.e., Search, Download, Blast, Comparison, My Favorites, Submission, and Advice). This structure can provide a centralized and user-friendly web portal for meiosis-related studies.
The website is built on a Linux, Apache, MySQL, and PHP (LAMP) stack. MySQL is used for storage, maintenance, and operation of the database, and 44 data sheets have been designed to form a network storing all the data (Figure 2). The front–end interface is implemented in PHP, which is a popular scripting language for dynamic web page. A well-defined and packaged JavaScript called jQuery is used to enhance the interface of the website and improve user experience. A navigation tool bar containing search box and links, such as Home, Blast, Submission, Download, and Help, are also included in each web page.
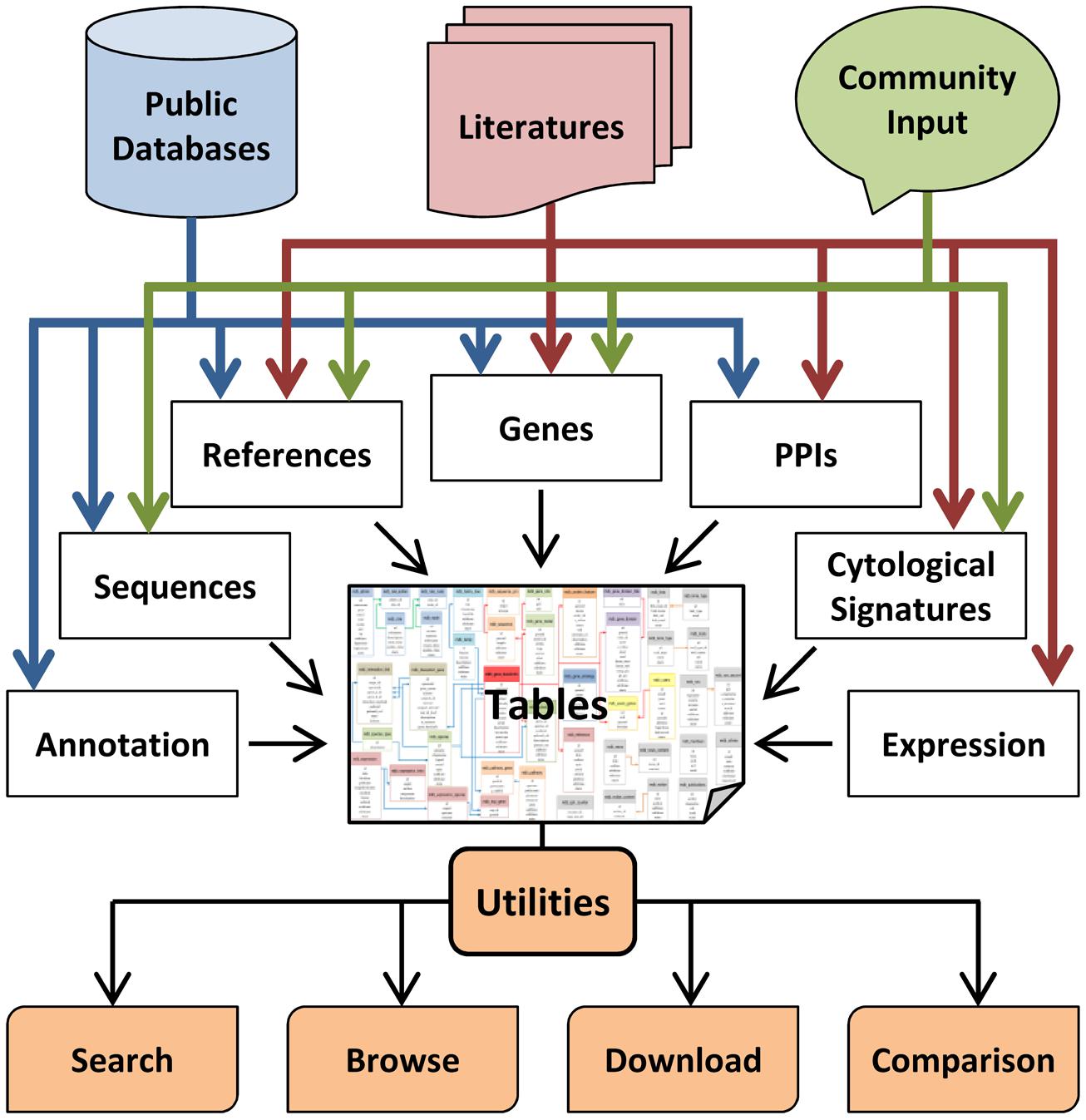
FIGURE 2. The flowchart for construction of MeioBase. MeioBase is a comprehensive database for browsing and retrieving knowledge about meiosis, meiotic genes, and related data. MeioBase brings together three main sources of knowledge: (1) basic information about genes from public databases; (2) detailed curation of meiosis-related studies from the literatures; (3) public contributions from research community and other users. All of the information is stored in relational database tables that could be accessed by the utilities of MeioBase through any web browser.
Resources
Cytological features of meiosis have been described in detail for many species, which laid a solid foundation for the study of the molecular basis of meiosis (Zickler and Kleckner, 1999; Ma, 2005). Here, in the Cytology section, we provide an overview of the cytological process of meiosis, with special emphasis on the chromosome behaviors at different stages. To help understand the conservation and variability of the meiotic process during evolution, cytological features in model species are summarized and compared. Important advances related to meiosis can also be retrieved through clicking the Updated Advances links from the overview page.
During meiotic prophase I, several critical events related to meiotic chromosome structure and interaction occur, including pairing, synapsis, recombination, and segregation. Consistent with this, genes regulating meiosis have also been grouped into pathways or networks, each of which corresponds to one of the meiotic events (Gerton and Hawley, 2005; Chang et al., 2011). For this reason, we have established the Pathway section, in which genes with similar or related functions, as well as the complex regulatory relationships between them, are visualized by diagrams.
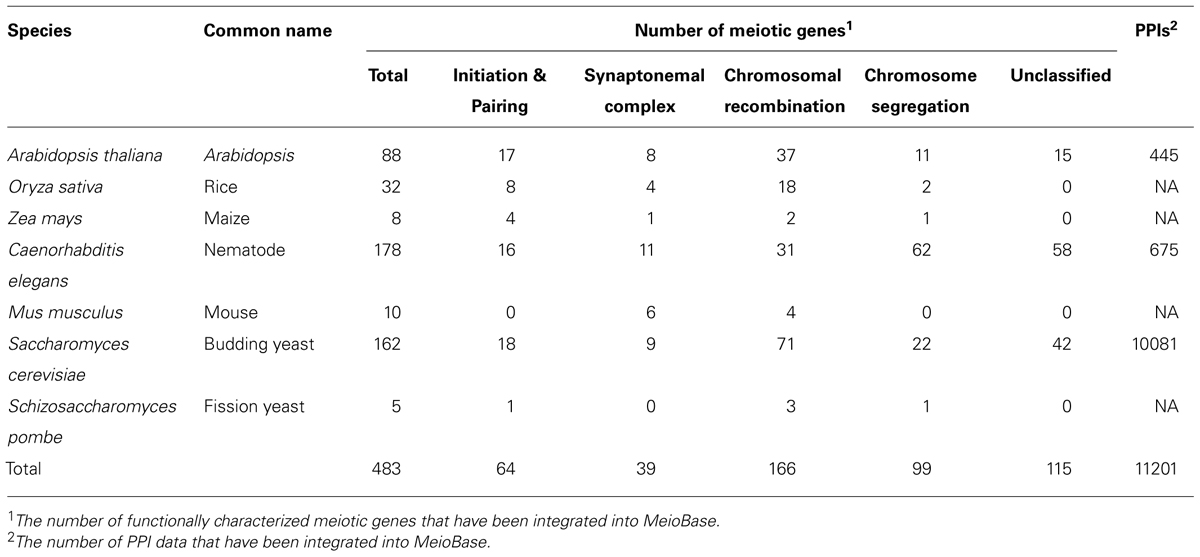
During the last 20 years, many genes involved in meiosis have been discovered and functionally characterized. By literature mining, 483 meiotic genes have been collected (Table 1). In the Species section, users can find the genes of a certain species and go into the details. In the Gene Detailed Information page, we have integrated useful information from references, databases and web servers. Data are organized by different aspects, such as General Information, Featured Domains, Protein Signatures, Gene Ontology, Protein Sequence, and References. General Information contains species name, gene name, gene family name, a brief description, gene model ID, and the coding sequence (CDS) of the gene. Featured Domains provides protein domains predicted by Pfam. Protein Signatures and Gene Ontology provide more information of the functions of the proteins. References includes literatures with PMIDs in PubMed (Figure 3).
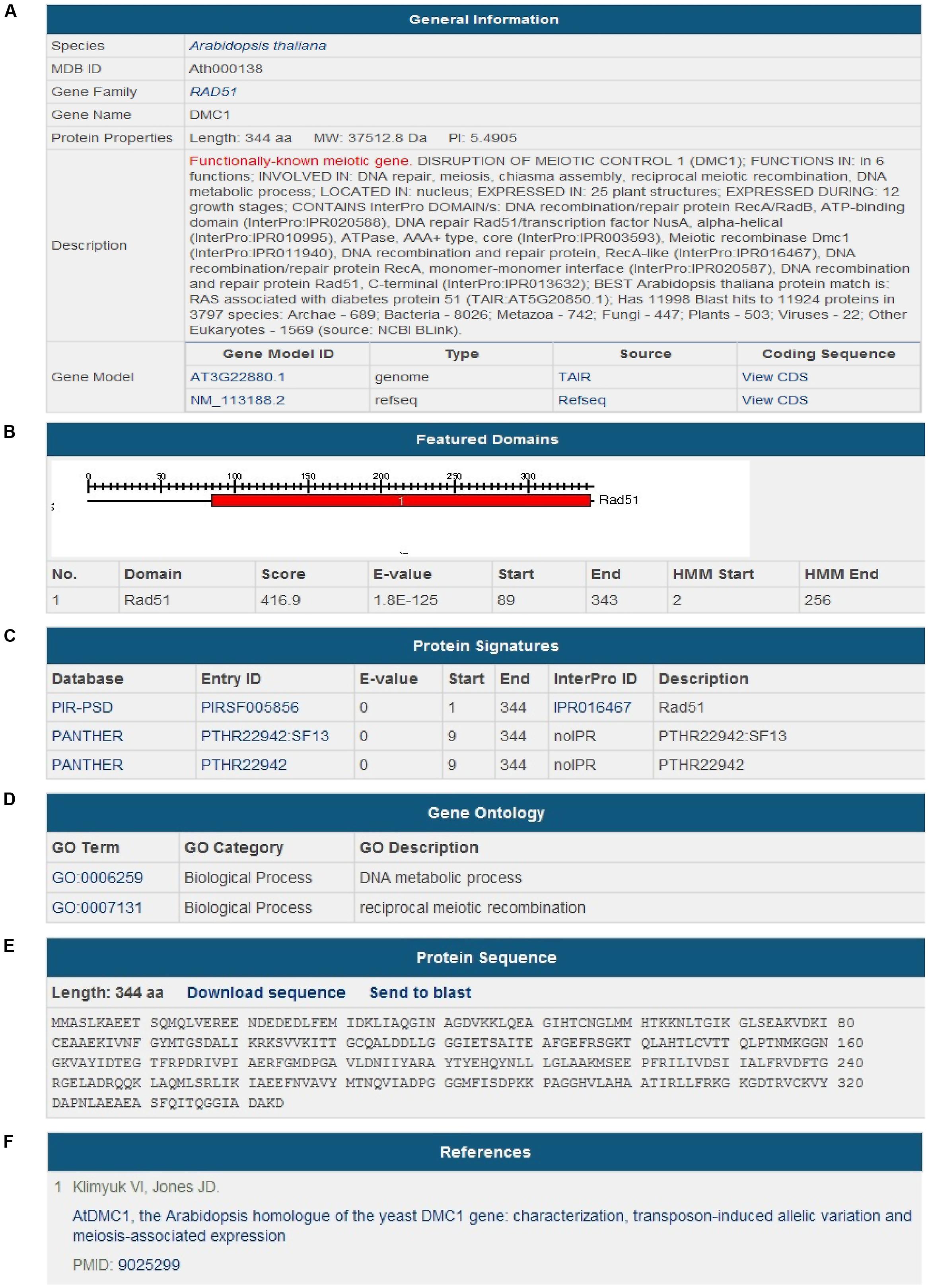
FIGURE 3. Screenshot of gene information. The Gene Detailed Information page of a meiotic gene consists of six aspects. (A) General Information; (B) Featured Domains; (C) Protein Signatures; (D) Gene Ontology; (E) Protein Sequence; (F) References.
We also include 11,201 pieces of PPI data of the collected meiotic genes of Arabidopsis, nematode, and budding yeast in MeioBase, and display them in the Interaction section (Table 1). The PPI data are retrieved from 283 literatures and 11 databases, such as BIND, BioGRID, DIP, IntAct, and STRING. Users can search for partners of the proteins of interest, for which corresponding sources and references are provided.
To provide more comprehensive information on meiosis, references related to gene expression patterns are collected and displayed in the Expression section. Until now, important references of various species, such as Arabidopsis, rice, maize, petunia, wheat, mouse, rat, budding yeast, and fission yeast, have been listed in this section and more expression data of meiosis are being collected.
In addition to the aforementioned five sections, MeioBase provides many other resources in the Links section, such as the commonly used databases, powerful web servers for molecular and genomic analysis, and experimental protocols for meiosis research, etc. An introduction to our “Plant Meiosis Project” can be found in the Project section. Important progresses and events on meiosis research are available in the News section. Release notes of MeioBase are announced in the Notice section. To provide an overview of our database, we also include a detailed introduction in the Help section.
Tools
MeioBase provides various ways to retrieve the data that users are interested in. By using the search box, users can search genes by gene names, gene model IDs or MeioBase IDs. In the Search section, users can search genes not only by inputting their model IDs but also by providing keywords describing them. PPI data can also be searched with keywords. All data can be downloaded in the Download section, which links to the FTP site.
Blast search against all the data in MeioBase is provided to facilitate users of finding similar sequence of a given sequence. In the Comparison section, users can add any two genes in the gene list and compare them. Users can also add any ten genes into the My Favorites section for fast checking afterward.
Moreover, we have integrated sections specifically for contributing meiosis data or suggestions to MeioBase. In the Submission section, users can first download and fill in a customized excel file as directions with meiotic genes, PPI data, and other data not yet included in the database, and then upload it to the database. After checking the uploaded data, we will add the qualified ones into MeioBase timely. In Advice section in homepage and at the foot of every page, users could ask any questions or give us comments or suggestions about this database. We will appreciate every user for improving MeioBase and reply as soon as possible.
Future Plans
MeioBase is the first web database providing comprehensive information on meiosis. It is only a start of establishing a large and well-known database on meiosis. We will reiterate the process of database structure and user interface development to enhance the data content and functionality. The major data content enhancement will come from elaboration of the gene annotation and incorporation of more meiotic genes in other species, various expression data from references and other databases, PPI networks of different species, and other vital pathways during meiosis.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Yongjie Sui for assistance in developing web interface, Dr. Zhihao Cheng and Prof. Zhukuan Cheng for help in checking meiosis data, and Dr. Hong Luo, Dr. Zhe Li, Prof. Jingchu Luo, Prof. Fangpu Han, and Prof. Ji Yang for helpful discussions, and two anonymous reviewers for valuable suggestions. This work was supported by grants from the Ministry of Sciences and Technology of China (2011CB944604).
References
Birchler, J. A., and Han, F. (2013). Meiotic behavior of small chromosomes in maize. Front. Plant Sci. 4:505. doi: 10.3389/fpls.2013.00505
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chang, F., Wang, Y., Wang, S., and Ma, H. (2011). Molecular control of microsporogenesis in Arabidopsis. Curr. Opin. Plant Biol. 14, 66–73. doi: 10.1016/j.pbi.2010.11.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chen, C., Farmer, A. D., Langley, R. J., Mudge, J., Crow, J. A., May, G. D.,et al. (2010). Meiosis-specific gene discovery in plants: RNA-Seq applied to isolated Arabidopsis male meiocytes. BMC Plant Biol. 10:280. doi: 10.1186/1471-2229-10-280
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Cui, X., Wang, Q., Yin, W., Xu, H., Wilson, Z. A., Wei, C.,et al. (2012). PMRD: a curated database for genes and mutants involved in plant male reproduction. BMC Plant Biol. 12:215. doi: 10.1186/1471-2229-12-215
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Dukowic-Schulze, S., Sundararajan, A., Mudge, J., Ramaraj, T., Farmer, A. D., Wang, M.,et al. (2014). The transcriptome landscape of early maize meiosis. BMC Plant Biol. 14:118. doi: 10.1186/1471-2229-14-118
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Gerton, J. L., and Hawley, R. S. (2005). Homologous chromosome interactions in meiosis: diversity amidst conservation. Nat. Rev. Genet. 6, 477–487. doi: 10.1038/nrg1614
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hamant, O., Ma, H., and Cande, W. Z. (2006). Genetics of meiotic prophase I in plants. Annu. Rev. Plant Biol. 57, 267–302. doi: 10.1146/annurev.arplant.57.032905.105255
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Handel, M. A., and Schimenti, J. C. (2010). Genetics of mammalian meiosis: regulation, dynamics and impact on fertility. Nat. Rev. Genet. 11, 124–136. doi: 10.1038/nrg2723
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hollingsworth, N. M., Goetsch, L., and Byers, B. (1990). The HOP1 gene encodes a meiosis-specific component of yeast chromosomes. Cell 61, 73–84. doi: 10.1016/0092-8674(90)90216-2
Kee, K., and Keeney, S. (2002). Functional interactions between SPO11 and REC102 during initiation of meiotic recombination in Saccharomyces cerevisiae. Genetics 160, 111–122.
Keeney, S., Giroux, C. N., and Kleckner, N. (1997). Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88, 375–384. doi: 10.1016/S0092-8674(00)81876-0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Klimyuk, V. I., and Jones, J. D. (1997). AtDMC1, the Arabidopsis homologue of the yeast DMC1 gene: characterization, transposon-induced allelic variation and meiosis-associated expression. Plant J. 11, 1–14. doi: 10.1046/j.1365-313X.1997.11010001.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lardenois, A., Gattiker, A., Collin, O., Chalmel, F., and Primig, M. (2010). GermOnline 4.0 is a genomics gateway for germline development, meiosis and the mitotic cell cycle. Database 2010:baq030. doi: 10.1093/database/baq030
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Li, W., Chen, C., Markmann-Mulisch, U., Timofejeva, L., Schmelzer, E., Ma, H.,et al. (2004). The Arabidopsis AtRAD51 gene is dispensable for vegetative development but required for meiosis. Proc. Natl. Acad. Sci. U.S.A. 101, 10596–10601. doi: 10.1073/pnas.0404110101
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lin, Z., Kong, H., Nei, M., and Ma, H. (2006). Origins and evolution of the recA/RAD51 gene family: evidence for ancient gene duplication and endosymbiotic gene transfer. Proc. Natl. Acad. Sci. U.S.A. 103, 10328–10333. doi: 10.1073/pnas.0604232103
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Luo, Q., Li, Y., Shen, Y., and Cheng, Z. (2014). Ten years of gene discovery for meiotic event control in rice. J. Genet. Genomics 41, 125–137. doi: 10.1016/j.jgg.2014.02.002
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ma, H. (2005). Molecular genetic analyses of microsporogenesis and microgametogenesis in flowering plants. Annu. Rev. Plant Biol. 56, 393–434. doi: 10.1146/annurev.arplant.55.031903.141717
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Orr-Weaver, T. L. (1995). Meiosis in Drosophila: seeing is believing. Proc. Natl. Acad. Sci. U.S.A. 92, 10443–10449. doi: 10.1073/pnas.92.23.10443
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Sym, M., Engebrecht, J. A., and Roeder, G. S. (1993). ZIP1 is a synaptonemal complex protein required for meiotic chromosome synapsis. Cell 72, 365–378. doi: 10.1016/0092-8674(93)90114-6
Tang, D., Miao, C., Li, Y., Wang, H., Liu, X., Yu, H.,et al. (2014). OsRAD51C is essential for double-strand break repair in rice meiosis. Front. Plant Sci. 5:167. doi: 10.3389/fpls.2014.00167
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Tang, X., Zhang, Z., Zhang, W., Zhao, X., Li, X., Zhang, D.,et al. (2010). Global gene profiling of laser-captured pollen mother cells indicates molecular pathways and gene subfamilies involved in rice meiosis. Plant Physiol. 154, 1855–1870. doi: 10.1104/pp.110.161661
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Vignard, J., Siwiec, T., Chelysheva, L., Vrielynck, N., Gonord, F., Armstrong, S. J.,et al. (2007). The interplay of RecA-related proteins and the MND1-HOP2 complex during meiosis in Arabidopsis thaliana. PLoS Genet. 3:1894–1906. doi: 10.1371/journal.pgen.0030176
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Wang, G., Kong, H., Sun, Y., Zhang, X., Zhang, W., Altman, N.,et al. (2004). Genome-wide analysis of the cyclin family in Arabidopsis and comparative phylogenetic analysis of plant cyclin-like proteins. Plant Physiol. 135, 1084–1099. doi: 10.1104/pp.104.040436
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yang, H., Lu, P., Wang, Y., and Ma, H. (2011). The transcriptome landscape of Arabidopsis male meiocytes from high-throughput sequencing: the complexity and evolution of the meiotic process. Plant J. 65, 503–516. doi: 10.1111/j.1365-313X.2010.04439.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yang, M., Hu, Y., Lodhi, M., Mccombie, W. R., and Ma, H. (1999). The Arabidopsis SKP1-LIKE1 gene is essential for male meiosis and may control homologue separation. Proc. Natl. Acad. Sci. U.S.A. 96, 11416–11421. doi: 10.1073/pnas.96.20.11416
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zickler, D., and Kleckner, N. (1998). The leptotene-zygotene transition of meiosis. Annu. Rev. Genet. 32, 619–697. doi: 10.1146/annurev.genet.32.1.619
Keywords: meiosis, MeioBase, knowledge database, meiotic genes, protein-protein interaction, eukaryotes, sexual reproduction
Citation: Li H, Meng F, Guo C, Wang Y, Xie X, Zhu T, Zhou S, Ma H, Shan H and Kong H (2014) MeioBase: a comprehensive database for meiosis. Front. Plant Sci. 5:728. doi: 10.3389/fpls.2014.00728
Received: 31 October 2014; Accepted: 01 December 2014;
Published online: 16 December 2014.
Edited by:
Dazhong Dave Zhao, University of Wisconsin–Milwaukee, USAReviewed by:
Raphael Mercier, Institut National de la Recherche Agronomique, FranceFangpu Han, Chinese Academy of Sciences, China
Copyright © 2014 Li, Meng, Guo, Wang, Xie, Zhu, Zhou, Ma, Shan and Kong. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hongyan Shan and Hongzhi Kong, State Key Laboratory of Systematic and Evolutionary Botany, Institute of Botany, Chinese Academy of Sciences, Nanxincun 20, Beijing 100093, China e-mail:c2hhbmhvbmd5YW5AaWJjYXMuYWMuY24=;aHprb25nQGliY2FzLmFjLmNu
 Hao Li
Hao Li Fanrui Meng
Fanrui Meng Chunce Guo
Chunce Guo Yingxiang Wang
Yingxiang Wang Xiaojing Xie4
Xiaojing Xie4 Hong Ma
Hong Ma Hongyan Shan
Hongyan Shan Hongzhi Kong
Hongzhi Kong