- 1Department of Pediatrics, Affiliated Hospital of Guangdong Medical College, Zhanjiang, China
- 2State Key Laboratory of Agrobiotechnology, College of Biological Sciences, China Agricultural University, Beijing, China
- 3National Center for Nanoscience and Technology, Beijing, China
Abscisic acid (ABA) plays a key role in many developmental processes and responses to adaptive stresses in plants. Recently, a new family of nucleocytoplasmic PYR/PYL/RCAR (PYLs) has been identified as bona fide ABA receptors. PYLs together with protein phosphatases type-2C (PP2Cs), Snf1 (Sucrose-non-fermentation 1)-related kinases subfamily 2 (SnRK2s) and downstream substrates constitute the core ABA signaling network. Generally, PP2Cs inactivate SnRK2s kinases by physical interaction and direct dephosphorylation. Upon ABA binding, PYLs change their conformations and then contact and inhibit PP2Cs, thus activating SnRK2s. Here, we reviewed the recent progress in research regarding the structures of the core signaling pathways of ABA, including the (+)-ABA, (−)-ABA and ABA analogs pyrabactin as well as 6AS perception by PYLs, SnRK2s mimicking PYLs in binding PP2Cs. PYLs inhibited PP2Cs in both the presence and absence of ABA and activated SnRK2s. The present review elucidates multiple ABA signal perception and transduction by PYLs, which might shed light on how to design small chemical compounds for improving plant performance in the future.
Introduction
Abscisic Acid
Abscisic acid (ABA) was discovered half a century ago (Addicott and Lyon, 1969; Milborrow, 1974; Cutler et al., 2010). Briefly, several groups isolated from different plant tissues plant growth regulators that could promote leaf abscission (Ohkuma et al., 1963) and seed dormancy (Cornforth et al., 1965b) and inhibit growth (Bennet-Clark and Kefford, 1953) and embryo germination (Cornforth et al., 1965b). Chemical analyses demonstrated that the activities of the isolated endogenous extracts were exerted by the same compound, which was ultimately named abscisic acid (Cornforth et al., 1965b; Milborrow, 1967).
ABA is an important sesquiterpenoid phytohormone that is derived from isopentenyl pyrophosphate (Nambara and Marion-Poll, 2005), and its chemical structure was finally confirmed by spectroscopic methods (Ohkuma et al., 1965) and chemical synthesis (Cornforth et al., 1965a). ABA is a pivotal regulator in plants and coordinates a complex regulatory network enabling plants to cope with abiotic stresses, such as drought, salinity, and temperature fluctuations (Verslues et al., 2006; Cutler et al., 2010; Kim et al., 2010; Miyakawa et al., 2013). Generation of ABA by cleavage of ABA conjugates (Lee et al., 2006) or de novo ABA biosynthesis (Nambara and Marion-Poll, 2005) significantly increases the ABA content under abiotic stress and thus regulates gene expression to assist plants in adapting to adverse environmental conditions (Hetherington, 2001; Schroeder et al., 2001). ABA also plays a key role in plant growth and development under non-stress conditions, including during embryo, seed and seedling development (Finkelstein et al., 2002) and seed dormancy (Finkelstein et al., 2008).
Identification of ABA Receptors
There are usually three common features of a receptor and its ligand: high affinity, high specificity, and a saturable and reversible interaction. Conventional genetic screening is beneficial to plant hormone research. Many key components that are involved in hormone signaling pathways were identified by screening of Arabidopsis mutants with increased or decreased sensitivity to a hormone (Santner and Estelle, 2009). However, such genetic screening failed to identify ABA receptors. This failure is mainly attributed to functional redundancy or pleiotropic effects, including embryo or gamete lethality for ABA receptors (Santiago et al., 2012). On the other hand, biochemical techniques were widely used to identify ABA-binding proteins. ABAP1 in barley aleurone was first reported to be an ABA binding protein (Razem et al., 2004). However, the homologous FCA in Arabidopsis, an RNA-binding protein that is responsible for flowering time (Razem et al., 2006), was unsuccessful in reproducing the ABA-binding ability using radioligand binding assays (Risk et al., 2008). The filter-based ligand-binding assay that was adopted in the FCA might be artificial due to non-specific binding (Risk et al., 2008). Then, the Mg-chelatase H subunit CHLH/GUN5/ABAR from Arabidopsis (Shen et al., 2006) as well as its homologue from Vicia faba (Zhang et al., 2002) were identified as ABA-binding proteins. The overexpression of either the full-length (Shen et al., 2006) or the C-terminal half of CHLH in Arabidopsis showed a hypersensitivity to ABA (Wu et al., 2009). However, the homologous CHLH protein in barley refused to bind ABA (Muller and Hansson, 2009). CHLH affected ABA signaling in stomatal guard cells, but no obvious ABA binding was detected using radioligand binding assays (Tsuzuki et al., 2011). Further experiments are required to determine the functions of CHLH in the ABA signaling pathway (Figure 1). In addition to the above two proteins, pharmacological evidence suggested that GTG1/GTG2 (Pandey et al., 2009) and GCR2 (Liu et al., 2007) were also ABA-binding proteins. However, Arabidopsis gtg1/gtg2 double mutants only slightly impaired the sensitivity to ABA in seed germination and stomatal responses (Pandey et al., 2009). The subsequent measurements were unable to detect the binding of ABA to GCR2 (Risk et al., 2009).
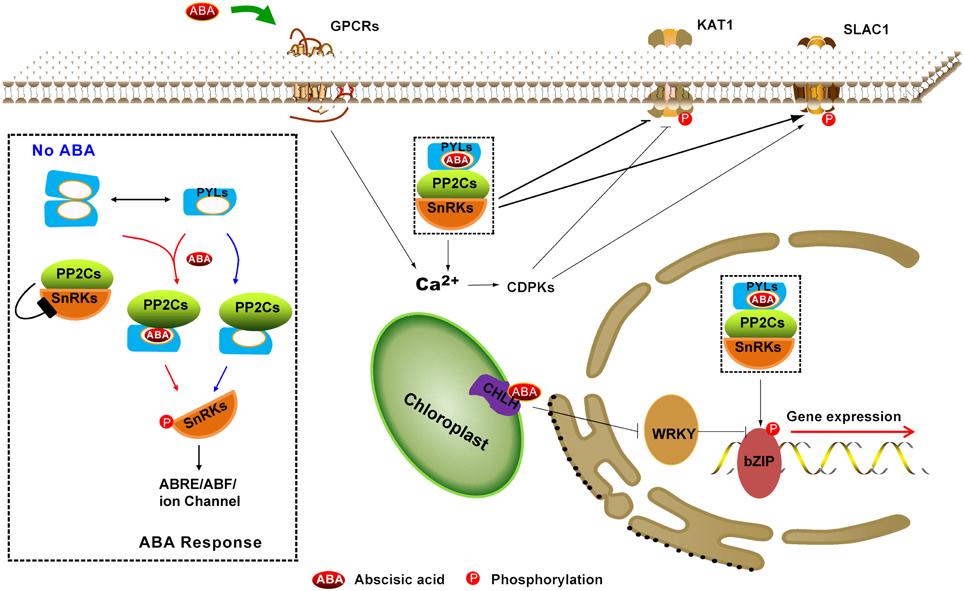
Figure 1. Summary of the ABA perception and signaling pathway. ABA receptors include nucleocytoplasmic PYR/PYL/RCARs (PYLs), most likely the plastid-localized Mg-chelatase H subunit (CHLH/GUN5/ABAR) and plasma membrane–localized GPCR type G-proteins (GPCRs). A core signaling pathway (black dotted rectangle enlarged in left) consists of PYLs, PP2Cs, SnRK2s, and downstream substrates, which control gene expression (long ABA signaling pathway) and stomatal closure (short ABA signaling pathway). Dimeric PYLs cannot bind to PP2Cs in the absence of ABA; thus, SnRK2s activity is inhibited by PP2Cs. Nevertheless, a subfamily of monomeric PYLs exhibits constitutive inhibitory activity on PP2Cs. In the presence of ABA, PYLs contact and inhibit PP2Cs, leading to the activation of SnRK2s through autophosphorylation. Then, the activated SnRK2 targets ABA-responsive element binding factors (such as bZIP) to regulate gene expression in the nucleus and the cation channel SLOW ANION CHANNEL-ASSOCIATED 1 (SLAC1) and POTASSIUM CHANNEL IN ARABIDOPSIS THALIANA 1 (KAT1) to cause stomatal closure in cytoplasm. The solid line with double-headed arrows indicates the equilibrium between dimeric PYLs and monomeric PYLs. The solid line with an arrow indicates direct positive interactions. The solid line with a bar indicates repression.
Our knowledge of ABA receptors was not clear until the major breakthrough of PYR/PYL/RCAR (hereafter referred to as PYLs) in 2009. Chemical genetics was used to find PYRABACTIN RESISTANCE 1 (PYR1) mutants that were insensitive to the synthetic selective ABA agonist pyrabactin (Park et al., 2009). Meanwhile, the function of the family as an ABA receptor was confirmed by a yeast two-hybrid assay using the ABI1/2 or HAB1 as bait (Ma et al., 2009). Nine independent members of the PYLs family were identified as the major in vivo interactors of ABI1 (Nishimura et al., 2010). PYL8 plays a non-redundant role in the regulation of root ABA sensitivity (Antoni et al., 2013) and promotes lateral root growth by enhancing the MYB77-dependent transcription of auxin-responsive genes (Zhao et al., 2014). In addition to PYLs, there may be other intracellular and extracellular ABA receptors that need to be identified (Cutler et al., 2010; Klingler et al., 2010).
The Core Signaling Network in the ABA Response
Recent studies have revealed the core ABA signaling components including ABA receptors (PYLs), type 2C protein phosphatases (PP2Cs), protein kinases [Snf1 (Sucrose-non-fermentation 1)-related kinases subfamily 2, SnRK2s] and downstream targets (Fujii et al., 2009; Umezawa et al., 2009) (Figure 1). When plants are challenged by various abiotic and biotic stresses, the endogenous ABA content increases and then exquisitely initiates some cellular signaling network to switch on adaptive responses and to regulate numerous developmental processes. ABA binds to PYLs and then the binary complex physically interacts with PP2Cs. The PYLs-PP2Cs heterodimer precludes substrate SnRK2s binding to PP2Cs, and thus stimulates SnRK2s kinase activity, which was formerly inhibited by PP2Cs (Yoshida et al., 2006; Park et al., 2009; Santiago et al., 2009b; Umezawa et al., 2009) (Figure 1). Activated SnRK2s can target NADPH oxidases (Sirichandra et al., 2009) and ion channels, such as the SLOW ANION CHANNEL-ASSOCIATED 1 (SLAC1) (Geiger et al., 2009; Sirichandra et al., 2009; Vahisalu et al., 2010) and the K+ channel in Arabidopsis Thaliana 1 (KAT1) (Sato et al., 2009), to control stomatal closure. In addition, activated SnRK2s can also target ABA-INSENSITIVE 5 (ABI5) (Nakashima et al., 2009a) and ABA-responsive element (ABRE) binding protein (AREB)/ABRE-binding factor (ABF) as well as transcription factors (TFs) (Furihata et al., 2006; Fujii and Zhu, 2009; Fujii et al., 2009; Fujita et al., 2009, 2013; Nakashima et al., 2009a) to regulate ABRE-dependent gene expression (Fujita et al., 2011, 2013) in seeds or vegetative tissues. So far, 14 members of PYLs, six to nine group-A PP2Cs, three subclass III SnRK2s and four to nine group-A bZIP TFs are involved in the core ABA signaling pathway (Klingler et al., 2010; Hauser et al., 2011; Takezawa et al., 2011), which was successfully reconstituted in vitro (Fujii et al., 2009) (Figure 1).
Since 2009, a rapidly growing body of literature has supported a double negative regulatory system that orchestrates PYLs as bona fide ABA receptors, PP2Cs as key negative regulators and SnRK2s as positive regulators, which play a critical role in ABA signaling networks (Cutler et al., 2010; Hubbard et al., 2010; Melcher et al., 2010b; Raghavendra et al., 2010; Umezawa et al., 2010; Joshi-Saha et al., 2011; Santiago et al., 2012; Miyakawa et al., 2013). As ABA receptors, PYLs have witnessed abundant structural and functional data from independent research groups (Melcher et al., 2009; Miyazono et al., 2009; Nishimura et al., 2009; Santiago et al., 2009a; Yin et al., 2009; Hao et al., 2010, 2011; Peterson et al., 2010; Yuan et al., 2010; Sun et al., 2012; Zhang et al., 2012, 2013; Li et al., 2013). In this review, we will predominantly focus on the structural data regarding the perception and recognition of (+)-ABA, (−)-ABA or ABA analogs by PYLs, their inhibitory interaction with PP2Cs, the autoactivation of SnRK2s through autophosphorylation and PP2Cs inhibiting SnRK2s through physical binding as well as mutual packing of their catalytic sites and dephosphorylation.
Structures of PYLs Receptores
Apo-Form of PYLs
There are 14 PYLs family members, named PYR1 and PYL1-PYL13 in Arabidopsis, that belong to the START superfamily (Iyer et al., 2001). Although several structural homologs, such as the pollen allergen Bet V 1α and CSBP from Vigna radiata, were previously known, it remained unclear how receptors perceive the ABA molecule. To date, the crystal structures of PYR1 (Nishimura et al., 2009; Santiago et al., 2009a), PYL1 (Miyazono et al., 2009), PYL2 (Melcher et al., 2009; Yin et al., 2009), PYL3 (Zhang et al., 2012, 2013), PYL5 (Zhang et al., 2013), PYL9 (Zhang et al., 2013; Nakagawa et al., 2014), PYL10 (Hao et al., 2011; Sun et al., 2012), and PYL13 (Li et al., 2013) have been reported (Figure 2A). From the published apo-PYLs structures, we know that all PYLs share a highly similar helix-grip structure that is characterized by a seven-stranded β-sheet that is flanked by two α-helices. In addition, a particular feature of PYLs receptors is an α-helix in the N termini.
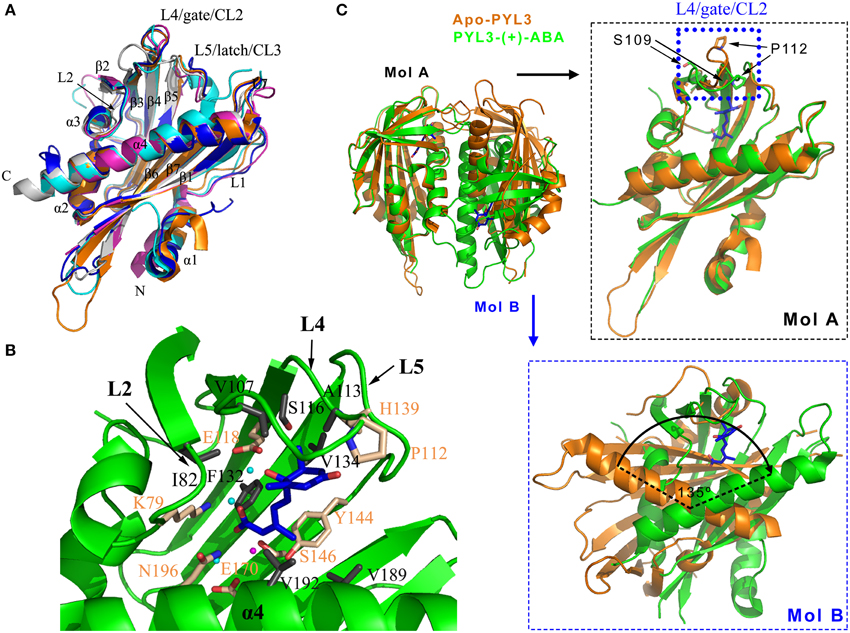
Figure 2. (+)-ABA induced conformation changes of PYLs. (A) PYR1 (PDB ID: 3K90, cyan), PYL1 (PDB ID: 3KAY, magenta), PYL2 (PDB ID: 3KDH, blue), PYL3 (PDB ID: 3KLX, orange), and PYL10 (PDB ID: 3R6P, gray) are shown in cartoon. (B) The ligand-binding pocket and the residues partaken in the hydrophobic and polar interactions of (+)-ABA-bound PYL3 (PDB ID: 4DSC, green). The ligand-binding pocket is encompassed by the L2, L4, L5 loops and α4 helix. The involved hydrophobic and polar residues were shown in gray and wheat sticks, respectively. Three water molecules involved in hydrogen bonds and one Mg2+ are shown in cyan and magenta spheres, respectively. 2D map of these interactions were seen in Figures S1, S2. (C) Superposition of cis-dimeric apo-PYL3 and trans-dimeric PYL3-(+)-ABA. The P112 residue moves toward the pocket to close the “gate” loop, whereas the S109 residue is flipped outward the cavity in response to (+)-ABA.
(+)-ABA-Bound PYLs
Based on ABA-bound structures, such as PYR1 (Nishimura et al., 2009; Santiago et al., 2009a), PYL1 (Miyazono et al., 2009), PYL2 (Melcher et al., 2009; Yin et al., 2009), PYL3 (Zhang et al., 2012), PYL9 (Zhang et al., 2013), and PYL10 (Hao et al., 2011; Sun et al., 2012), These structural architectures share a common ligand-binding pocket. The L2 loop between the α3 helix and β2 strand, the L4 loop (also referred to as CL2 or “gate” loop) between the β3 and β4 strands, the L5 loop (also referred to as CL3 or “latch” loop) (Melcher et al., 2009; Yin et al., 2009) between the β5 and β6 strands, and the C-terminal helix α4 encompass the entrance of the ligand-binding pocket (Figure 2B, Figure S2), which is very important for ABA binding. PYLs possess a large internal cavity in which the ABA molecule sits by a combination of ionic bonds, hydrophobic interactions and water-mediated hydrogen bonds (Figure 2B, Figures S1, S2). The carboxyl of ABA forms a salt bridge with the amine group of lysine (PYR1 K59, PYL1 K86, PYL2 K64, PYL3 K79, PYL9 K63, and PYL10 K56) as well as a water-mediated hydrogen bond network with several side chains of polar residues. The addition of a bulky group to the carboxylic group of ABA is likely to interfere with the binding to PYLs. In this scenario, the carboxylic group of ABA coupled to the amino group of a 10-atom spacer arm of a sepharose resin was utilized to identify ABA binding proteins and an ABA receptor such as CHLH (Zhang et al., 2002; Shen et al., 2006). Such approach probably cripples the interaction between the carboxylate group of ABA and ABA binding proteins or receptor. Therefore, it is suggested that ABA binding by CHLH must be further confirmed (Santiago et al., 2012). Moreover, the hydroxyl group and the ketone group of ABA also interact with the polar side chains of PYLs through water-mediated hydrogen bonds. In addition, the pentadienoic acid moieties and the cyclohexene contact hydrophobically with many inward-facing apolar side chains in the PYLs' cavity. These residues that are involved in binding ABA are strictly conserved in PYLs.
The superposition of the apo-PYLs and ABA-bound PYLs made it possible to depict the ABA-induced conformational changes. Here, the superposition of the ABA-bound and apo-PYL3 shows obvious conformational changes in the two conserved loops that flank the entrance to the ABA binding pocket and the C-terminal α-helix (Melcher et al., 2009; Nishimura et al., 2009; Santiago et al., 2009a; Yin et al., 2009; Zhang et al., 2012). When ABA binds to the receptor, the P112 residue of PYL3 on the “gate” loop (PYR1 P88, PYL1 P115, and PYL2 P92) moves toward the pocket to close the “gate” loop, whereas the S109 residue on the “gate” loop (PYR1 S85, PYL1 S112, and PYL2 S89) is flipped outward the cavity (Figure 2C right panel). In addition, the imidazole group of the H139 residue on the “latch” loop (PYR1 H115, PYL1 H142, and PYL2 H119) orientates inward the cavity to contact ABA. In addition, the α-helix α4 in the C termini moves slightly toward ABA to facilitate the closure of the “gate” and “latch” loops. These altered conformations create a new surface that is favorable for binding to PP2Cs, which in turn lock the “gate” and “latch” loops into the closed conformation (discussed below). Thus, ABA perception by PYLs allosterically regulates the conformation of the “gate” and “latch” loops to switch the ABA signal transduction.
The structural changes by ABA perception that were mentioned above only considered one protomer of PYLs. Interestingly, based on the oligomeric state of apo-PYLs, the results from gel filtration chromatography, small angle X-ray scattering, static light scattering or analytical ultracentrifugation convincingly showed that PYL4-10, except for uncharacterized PYL7, are monomers, whereas PYR1 (Nishimura et al., 2009; Santiago et al., 2009a) and PYL1-3 are homodimers in solution (Dupeux et al., 2011b; Hao et al., 2011; Zhang et al., 2012). For homodimeric PYR1 and PYL1-2, the relative orientation of one protomer with respect to the other is slightly changed in response to ABA. As a result, a significant rearrangement of the interface is generated, leading to a diminished number of van der Waals contacts and hydrogen bonds, and consequently a weakening of the dimer interface (Nishimura et al., 2009; Yin et al., 2009). Similarly, cis-homodimer of PYL3 has also been observed in the apo-PYL3 structure (Figure 2C). However, ABA-bound PYL3 transforms into a trans-homodimer by one protomer rotation of almost 135° compared to the cis-homodimer (Figure 2C lower panel). The “gate” and “latch” loops in the trans-homodimer are more exposed in the solvent (Zhang et al., 2012). These data suggest that the binding of ABA could influence the dimer interface and the relative orientation of the two protomers from dimeric PYLs, which may be used to strictly regulate the ABA-dependent switching of signal transduction by PYLs (Umezawa et al., 2010).
The Stereospecificity of PYLs to (+/−)-ABA Stereoisomers
ABA contains one optical center at C1′, and the form of S-(+)-ABA exists in nature [hereafter referred to as (+)-ABA]. Another feature is the side chain of the ABA molecule, which contains two double bonds that are conjugated to the carboxylic acid. The configuration of the double bond that is adjacent to the ring is trans, while that proximal to the acid group is cis (Figure 3A). Upon exposure to UV light, biologically active 2-cis,4-trans ABA would be isomerized to the inactive form 2-trans,4-trans ABA (Cutler et al., 2010). In order to study the importance of the S-(+)-ABA stereoisomer and its biological activity, the unnatural but bioactive stereoisomer R-(−)-ABA was synthesized [hereafter referred to as (−)-ABA] (Lin et al., 2005; Zaharia et al., 2005) (Figure 3A). Both of the stereoisomers showed obviously different activities in many aspects, such as stomatal closure, but nonetheless a comparable activity in seed germination (Nambara et al., 2002; Xie et al., 2005). Microarray and genetic studies showed that the in vivo function of R-(−)-ABA also requires the signaling pathway of (+)-ABA (Xie et al., 2005; Huang et al., 2007). Two mechanisms were hypothesized to explain the bioactivity of (−)-ABA: one mechanism was that the same site was occupied between these two ABA stereoisomers by flipping the cyclohexene plane (Milborrow, 1974), and the other mechanism was the dual selectivity of ABA receptors (Nambara et al., 2002).
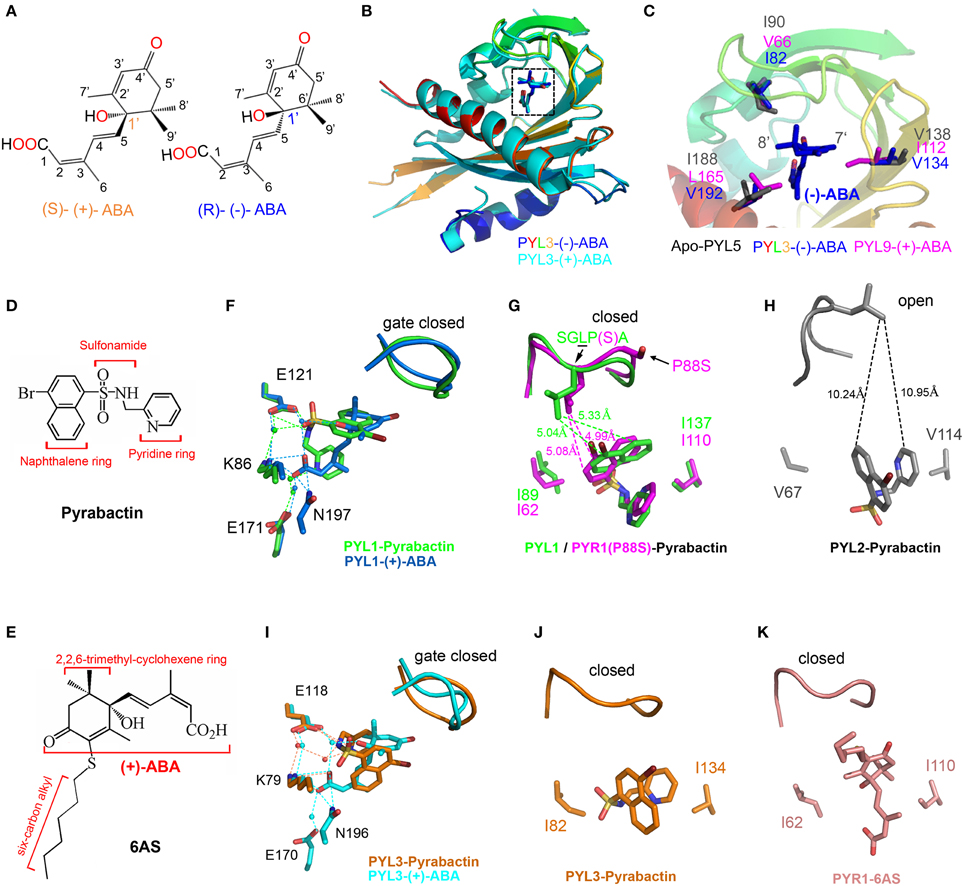
Figure 3. Structural basis for the selective activation of ABA receptors by different ligands. (A) Chemical structures of (+)-ABA and (−)-ABA. (B) Superposition of PYL3-(−)-ABA (PDB ID: 4JDA, rainbow) and PYL3-(+)-ABA (PDB ID: 4DSC, cyan). (−)-ABA and (+)-ABA were shown in blue and cyan sticks, respectively. (C) Superposition of PYL3-(−)-ABA (rainbow cartoon), PYL9-(+)-ABA (magenta sticks), and apo-PYL5 (gray sticks) indicates that the major variant residues underlie the stereospecificity of PYLs to (−)-ABA. Two bulk side chains of I112 and L165 in PYL9 seriously clash with the 7′ and 8′ methyl groups of (−)-ABA, respectively. The chemical structures of pyrabactin (D) and 6AS (E). (F–K) Superposition of PYLs-ligands. Superposition of PYL1-pyrabactin (PDB ID: 3NEF, green) with PYL1-(+)-ABA (PDB ID: 3JRS, marine) shows the basis of pyrabactin as an agonist for PYL1, inducing the closure of the “gate” loop just as (+)-ABA. There are two residues determining the agonism and antagonism of pyrabactin for PYLs [PYR1(P88S)-pyrabactin (PDB ID: 3NJO, magenta) and PYL2-pyrabactin (PDB ID: 3NR4, gray)]. In particular, the superposition of PYL3-pyrabactin (PDB ID: 3OJI, orange) with PYL3-(+)-ABA (PDB ID: 4DSC, cyan) indicates that the “gate” loop of PYL3 closes in response to pyrabactin, which is not compatible for the insertion of the conserved tryptophan of PP2Cs. Therefore, pyrabactin is an antagonist for PYL3 (Zhang et al., 2012). The PYR1-6AS (PDB ID: 3WG8, wheat) structure shows that 6AS, mimicking (+)-ABA, induces the same conformation of the “gate” loop, but its long 3′ alkyl chains impedes PP2Cs docking; thus, 6AS is also an antagonist for PYLs (Takeuchi et al., 2014). All dotted lines represented the distance between two resides in angstrom units.
Generally, PYLs show a greater affinity for (+)-ABA than for (−)-ABA (Melcher et al., 2009; Miyazono et al., 2009). Although PYL9 shows a stringent preference for the natural (+)-CityABA, PYL5 can bind (−)-ABA with a Kd 8-fold higher than for (+)-ABA (Santiago et al., 2009b). (−)-ABA promoted the binding affinity of PYL2-4 but not PYR1 to HAB1 (Park et al., 2009). (−)-ABA could be structurally modeled into the PYL2 pocket without any steric clash (Yin et al., 2009). The initial 1.8 Å resolution structure was determined with mixed enantiomorphic (+/−)-ABA, suggesting that the chiral difference was accommodated within the ABA receptor binding pocket by the flipping of the ABA ring by 180° (Nishimura et al., 2009).
From the recent report, PYL5 showed the strongest binding affinity to (−)-ABA among all of the tested PYLs. PYL9 was a stringently exclusive (+)-ABA receptor (Zhang et al., 2013). PYL3 was a dual receptor to both ABA enantiomers. The structures of apo-PYL5, PYL3-(−)-ABA and PYL9-(+)-ABA were determined to elucidate the mechanism of stereospecificity of PYLs to ABA enantiomers (Zhang et al., 2013). Interestingly, the entire profile of (−)-ABA-bound PYL3 resembles that of PYL3-(+)-ABA, with a closed “gate” loop and a trans-dimeric rearrangement. The superimposition of these structures showed that the binding orientation of (−)-ABA in the PYLs pocket is obviously different from that of (+)-ABA (Figure 3B), which may deny the “flip” hypothesis. Structural and biochemical investigations showed that the major variable residues surrounding the mono-methyl and di-methyl groups of the ABA cyclohexene ring might underlay the preference of PYL binding to ABA enantiomers, such as steric hindrance by the two bulk side chains of I112 and L165 in PYL9 (Figure 3C). Moreover, the hydrophobic interaction through indirect interaction with the 8′, 9′ methyl groups of (−)-ABA (Figure 3C) also contributes to the stereospecificity of PYLs to ABA enantiomers because the V66I mutation increases the inhibitory ability of PYL9 to PP2Cs in the presence of (−)-ABA. The relative stereo specificity of different PYLs and their ability to bind alternative ligands could be used to explore the link between biochemical activities and physiological responses and may also provide useful tools to manipulate ABA signaling in both experimental and agricultural contexts (Hubbard et al., 2010).
Pyrabactin as an Agonist/Antagonist of PYLs
Due to functional redundancy or pleiotropic effects (McCourt, 1999), the selective analogs of ABA are required to illuminate the function of a certain ABA receptor or find other unknown ABA receptors. Many analogs of ABA were designed mainly by substituting the 7′, 8′, or 9′-carbon atoms (Zaharia et al., 2005). Unlike ABA structurally, pyrabactin (Figure 3D), which was employed to find the first ABA receptor PYR1 by chemical genetics (Park et al., 2009), is a selective ABA agonist in seed germination and stomatal closure (Zhao et al., 2007; Puli and Raghavendra, 2012). Several years later, the structures of PYR1 and PYL1-3 complexed with pyrabactin were reported to clarify the selective activation of PYLs (Hao et al., 2010; Melcher et al., 2010b; Peterson et al., 2010; Yuan et al., 2010; Zhang et al., 2012). Pyrabactin as an agonist binds to PYL1 and PYR1 in a productive mode, whereas as an antagonist binds to PYL2 or PYL3 in a non-productive mode, indicating that pyrabactin can adopt different conformations in the conserved pocket of PYLs.
Structural analyses have revealed that pyrabactin lies in the cavity of PYL1 or PYR1 to induce the closure of the “gate” loop as ABA does (Hao et al., 2010; Peterson et al., 2010). Although pyrabactin structurally does not resemble ABA, several key interactions are conserved between PYL1-(+)-ABA and PYL1-pyrabactin (Hao et al., 2010; Melcher et al., 2010b) (Figure 3F). The pyridyl nitrogen of pyrabactin is located in the position of the carboxylate oxygen of ABA, forming water-mediated hydrogen bonds with the K86 and E171 in PYL1 (Figure 3F). In addition, the amine group of pyrabactin occupies the position of the hydroxyl group of ABA, which forms hydrogen bonds with N197 and E121 in PYL1. The sulfonamide of pyrabactin also forms hydrogen bonds with E121. In addition, the naphthalene ring of pyrabactin imitates the 2,6,6-trimethylcyclohexene ring of ABA, forming hydrophobic contacts with the “gate” loop to promote closure (Figure 3G). Moreover, the PYL1-pyrabactin-ABI1 complex structure reveals that pyrabactin inhibits PP2C activity in a productive mode (Melcher et al., 2010b).
In contrast, the binding of pyrabactin to the pocket of PYL2 does not provoke the closure of the “gate” loop (Melcher et al., 2010b; Peterson et al., 2010; Yuan et al., 2010) (Figure 3H). The orientation of pyrabactin occupying the pocket of PYL2 is rotated by approximately 90° compared to that in the pocket of PYR1 or PYL1. The naphthalene ring cannot induce the closure of the PYL2 “gate” loop because it is too far away to interact with each other (Figure 3H). The orientation of pyrabactin is positioned by the two small and hydrophobic residues V114 and V67 in PYL2, which correspond to I110 and I62 in PYR1 and I137 and I89 in PYL1, respectively (Figure 3G). Compared to the wild type, the V67I, V114I, and V67I/V114I mutants of PYL2 showed an increasing inhibition onto PP2Cs in response to pyrabactin (Peterson et al., 2010; Yuan et al., 2010). In addition, the I62V, I110V, and I62V/I110V mutants of PYR1 achieved only partial PP2C inhibition in response to pyrabactin (Peterson et al., 2010). These two smaller valine residues in PYL2 make the naphthalene ring deep in the pocket and far away from the “gate” loop, which results in pyrabactin as an antagonist binding to PYL2.
Intriguingly, the situation in PYL3 is totally different. The orientations of the naphthalene and pyridine ring of pyrabactin are significantly rotated compared to those in other PYLs-pyrabactin complexes. Moreover, the sulfonamide group moves to F81 and does not form a hydrogen bond with K79, while the conserved lysine residue in other PYLs-pyrabactin complexes forms a hydrogen bond with the pyridyl nitrogen to locate pyrabactin inside of the binding pocket. Pyrabactin can induce the closure of the “gate” in PYL3, but the “gate” loop moves further toward the binding pocket and tightly closes the “latch” (Figures 3I,J). The “gate” loop in PYL3 binds to pyrabactin tighter, which gives rise to the more compact space between the “gate” loop and the binding pocket compared to that in PYL1, PYL2, or PYR1. Therefore, there is no enough space between the “gate” loop and the “latch” loop for the insertion of conserved tryptophan residue from PP2Cs. Taken together, pyrabactin works as an antagonist for PYL3 (Figures 3I,J) (Zhang et al., 2012).
These different orientations of pyrabactin in PYLs either provide or impair interactions with the “gate” loop, inhibiting PP2Cs in a productive or non-productive mode, respectively. In addition, AM1, a small-molecule ABA mimic, acts as a potent activator of several members of PYLs (Cao et al., 2013), whereas 6AS, an ABA analog containing a six-carbon alkylsulfanyl that is linked to ABA's 3′ ring, was designed and confirmed as a potent ABA antagonist (Figures 3E,K) (Takeuchi et al., 2014). These reported structural data shed light on the concept of ABA receptor agonism and antagonism and are useful in the design of selective PYLs analogs.
ABA-Dependent Inhibition of PP2Cs by PYLs Receptors
Architecture of the Ternary Complexes PYLs-(+)-ABA-PP2C
Four crystal structures of the ternary complexes PYL1-ABA-ABI1, PYL2-ABA-HAB1, PYR1-ABA-HAB1, and PYL3-ABA-HAB1 have been described (Melcher et al., 2009; Miyazono et al., 2009; Yin et al., 2009; Dupeux et al., 2011a; Zhang et al., 2012). PP2Cs dock into the ABA-bound PYLs, in which the major interface comprises the closed “gate” loop, the “latch” loop and the C-terminal helix (Figure 4A). These structures elucidate the mechanism by which PYLs change their conformation upon ABA binding to inhibit the phosphatase activity of PP2Cs in an ABA-dependent manner. In these structures, the PP2Cs catalytic cores (residues: ABI1 125-429, HAB1 172-511) adopt a fold with two central five-stranded β-sheets that are sandwiched by two pairs of α-helices, and the catalytic site is located at the edge of the two central β-sheets (Figure 4A). The N-terminal portion of the PP2Cs perhaps regulates PP2Cs activity via the recruitment of other interacting partners, but its structure has not been determined.
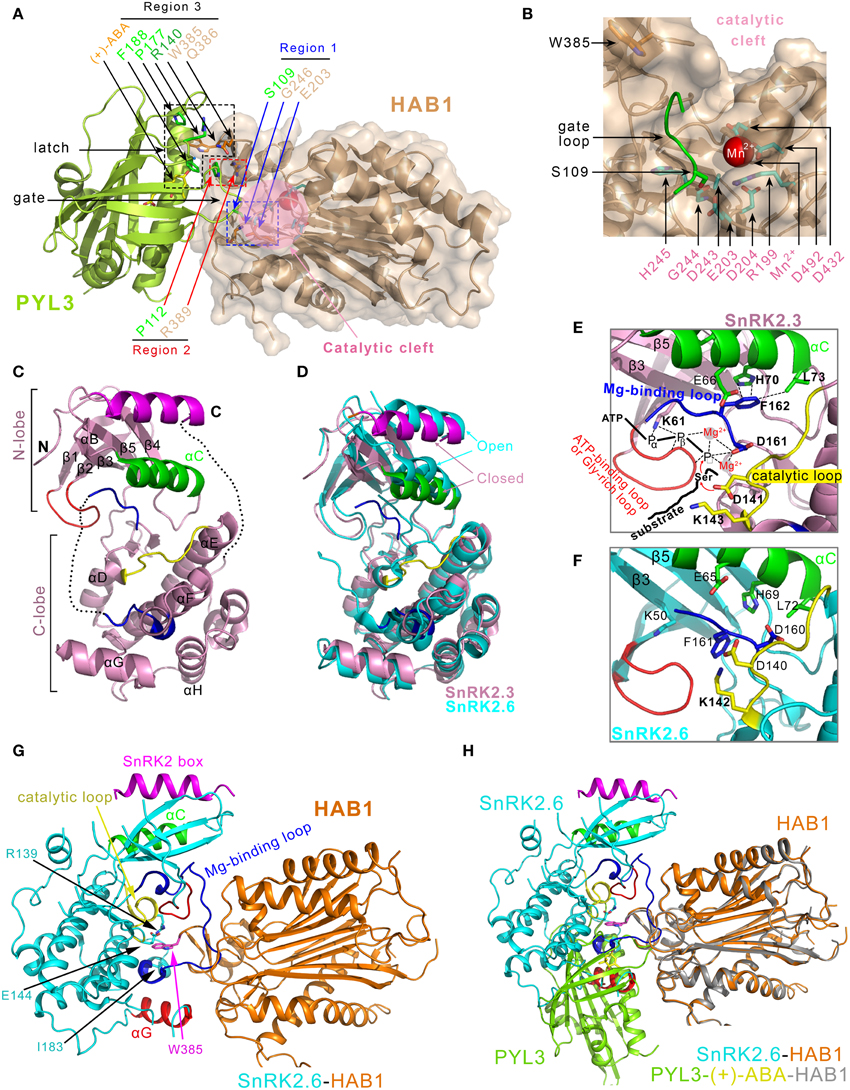
Figure 4. Structural characterizations of the PYLs-(+)-ABA-HAB1 and SnRK2s-PP2Cs complex. (A) Overview of three contact regions between PYL3 and HAB1 (PDB ID: 4DS8). The active site of HAB1 is shown in the pink circle. The PP2C contacts the S109 in the “gate” loop of PYL3 through its E203 and G246 in the active site cleft (Region 1), and contacts the P112 in the “gate” loop of PYL3 through its R389 residue (Region 2). The W385 and Q386 residues of PP2C (Region 3) contact several elements surrounding the entrance of the PYL3 cavity, including R140, P177, and F188 of PYL3 and the ketone group of ABA. (B) Details of the phosphatase active center, which consists of residues R199, E203, D204, D243, G244, H245, D432, and D492. Mn2+ is marked in a magenta sphere. (C) Structural characterization of SnRK2.3 (PDB ID: 3UC3) (Ng et al., 2011). Helix αC is displayed in green, the ATP-binding loop (Ng et al., 2011)/Gly-rich loop (Yunta et al., 2011) is displayed in red, the SnRK2 box (Ng et al., 2011)/DI domain (Yunta et al., 2011) is shown in magenta, the Mg2+ binding loop/activation loop are shown in blue, and the catalytic loop is colored yellow. (D) Superposition of the SnRK2.3 (see panel C) and SnRK2.6 (PDB ID: 3UC4, cyan) structures, which manifests that SnRK2.3 and SnRK2.6 adopt closed and open conformations, respectively. (E) SnRK2.3 adopts a partially active conformation. ATP, Mg2+ and substrate are introduced into apo-SnRK2.3 to illustrate active kinase conformational features. The residues that are involved in intra-molecular interaction network in the active sites are shown (Ng et al., 2011). (F) SnRK2.6 adopts an inactive conformation. E65, D160, and the catalytic D140 are far from the active site (Ng et al., 2011). (G) Overview of the SnRK2.6-HAB1 (PDB ID: 3UJG) complex structure. Three regions in SnRK2.6 are responsible for interaction with HAB1: the catalytic loop (yellow), the helix αG (red) and the Mg2+-binding loop (blue). In addition, R139, E144, and I183 (magenta) of SnRK2.6 establish interactions with the conserved W385 in HAB1 (cyan). (H) Overlay of the interaction surfaces from the SnRK2.6-HAB1 and PYL3-(+)-ABA-HAB1 complexes.
There are three characteristic interaction regions between the ABA receptors (PYL1-3) and the PP2Cs catalytic core (Figure 4A). First, the conserved serine residue in the “gate” loop (PYL1 S112, PYL2 S89, and PYL3 S109) establishes hydrogen bonds with the backbone amide of glycine (ABI1 G180 and HAB1 G246) and the metal-stabilizing carboxylic group of glutamic acid (ABI1 E142 and HAB1 E203) of PP2Cs (Figure 4A region 1 and Figure 4B). The involved glycine and glutamic acids are located at the catalytic site of PP2Cs; therefore, the catalytic site cleft of PP2Cs is blocked by the closed “gate” loop of PYLs, which is responsible for the inhibitory effect on the phosphatase activity. In addition, a conserved proline residue in the “gate” loop (PYL1 P116, PYL2 P93, and PYL3 P112) stacks with the guanidinium group of a conserved arginine residue in PP2Cs (ABI1 R304 and HAB1 R389) (Figure 4A region 2). This interaction promotes the “gate” loop enclosure and the insertion to the active site of PP2Cs. In addition, the indole ring of a conserved tryptophan residue (ABI1 W300 and HAB1 W385) inserts between the “gate” and “latch” loops of the ABA-bound PYLs (Figure 4A region 3). A representative water-mediated hydrogen network is established among the indole imine group of tryptophan, the ketone group of ABA, the backbone carbonyl of proline in the “gate” loop and the guanidinium group of arginine in the “latch” loop. Therefore, the phosphatase activity of PP2Cs is inhibited by interacting with ABA-bound PYLs. Furthermore, these PP2C-induced conformational changes of PYLs and the interaction between the conserved tryptophan of PP2C and ABA further stabilize the binding of ABA to PYLs and thus decrease the rate of ABA dissociation from the PYLs (Ma et al., 2009; Nishimura et al., 2009; Santiago et al., 2009b; Cutler et al., 2010). Consistently, the overexpression of the HAB1W385A mutant in Arabidopsis leads to reduced ABA sensitivity (Dupeux et al., 2011a).
PYR1, PYL1, PYL2, and PYL3 are homodimers, while the PYLs-PP2Cs complexes are heterodimers in solution. The homodimeric interface of PYLs has a dramatic overlap with the heterodimeric interface of PYLs-PP2Cs. Therefore, the homodimeric PYLs must be dissociated into monomeric PYLs to bind to PP2Cs. ABA-binding could induce a conformational rearrangement of dimeric PYLs, which decreases the interface area and promotes the dissociation of dimeric PYLs (Yin et al., 2009; Zhang et al., 2012). It was postulated that PYLs exist in vivo as inactive homodimers and are incapable of binding or inhibiting PP2Cs (Yin et al., 2009). Therefore, the dimeric nature of PYLs may play some roles in the regulation of PYLs in plants, such as reducing the basal interaction between PYLs and PP2Cs.
Dominant Insensitivity of abi1-1, abi2-1, and hab1-1 to ABA
The missense mutations abi1-1 (G180D) and abi2-1 (G168D) were isolated more than 20 years ago by genetic screenings in Arabidopsis to study the ABA signaling (Koornneef et al., 1984; Leung et al., 1994, 1997; Meyer et al., 1994; Rodriguez et al., 1998). ABI1 and ABI2 were negative regulators of ABA signaling, but both abi1-1 and abi2-1 mutants showed insensitive responses to ABA (Gosti et al., 1999; Merlot et al., 2001; Saez et al., 2004, 2006). The ABI1-1 protein cannot bind PYLs in the presence of ABA (Ma et al., 2009; Park et al., 2009) but retains the normal capacity to interact with SnRK2s (Umezawa et al., 2009; Vlad et al., 2009). The ternary complex structures of PYLs-(+)-ABA-PP2Cs showed that the conserved serine in the “gate” loop forms a hydrogen bond with the conserved glycine in the active site of PP2Cs (Figures 4A,B). The bulkier aspartic acid, which is substituted for the glycine, would disrupt the hydrogen bond, clash with the “gate” loop, and in turn impair the binding of PP2Cs (Yin et al., 2009).
ABA-Independent Inhibition of PP2Cs by PYLs Receptores
ABA-Responsive Inhibition of PP2Cs
ABA first enters the hydrophobic pocket of PYLs, and then the ABA-bound PYLs bind to the downstream substrate PP2Cs, called the induced pathway (Ma et al., 2009; Park et al., 2009; Cutler et al., 2010). This mechanism is convincingly confirmed by the crystal structures of four PYLs-ABA-PP2Cs ternary complexes (Melcher et al., 2009; Miyazono et al., 2009; Yin et al., 2009; Dupeux et al., 2011a; Zhang et al., 2012). However, both yeast two-hybrid and in planta bimolecular fluorescence complementation experiments demonstrated that PYL9 and PYL5, in contrast to PYR1 and PYL1-4, show constitutive interaction with PP2Cs (Ma et al., 2009; Park et al., 2009; Santiago et al., 2009b; Szostkiewicz et al., 2010). ABI1 can be co-purified with PYL5–12 from Arabidopsis plants (Nishimura et al., 2010). Biochemical assays in vitro also showed that PYR1 and PYL1-3 inhibit PP2Cs (ABI1, HAB1, HAB2, and PP2CA) in an ABA-dependent manner, whereas PYL5-6 and PYL8-10 can inhibit PP2Cs even in the absence of ABA (Hao et al., 2011). Therefore, PYLs contact and inhibit PP2Cs, independent of ABA, called the constitutive pathway.
Recently, crystal structures of apo-PYL10 and the binary complex PYL10-HAB1 (Figure 5A) revealed the molecular mechanism by which a subclass of PYLs, represented by PYL10, inhibits PP2Cs even in the absence of ABA (Hao et al., 2011). The alignment of all PYLs primary sequences, particularly the residues that contact the “gate” loop, showed that the bulkier hydrophobic residue L79 in PYL10 provided a platform to dock the hydrophobic L83 in the “gate” loop, thus facilitating a closed conformation of the “gate” loop (Figure 5B). Nevertheless, the corresponding residues in all of the other PYLs except for PYL13 are occupied by a smaller valine. Structural-guided biochemical assays indicated that PYLs in a monomeric state are the first prerequisites for the ABA-independent inhibition of PP2Cs. Secondly, the residues guarding the entrance to the cavity of these PYLs should be bulky and hydrophobic (Hao et al., 2011). The apo-PYL10 with the “gate” loop in closed conformation was also determined, which builds a dynamic equilibrium model between the open and closed conformations of PYL10 (Sun et al., 2012). Comparing the structures of apo-PYL10 and other known apo-PYLs, the “gate” loop of the apo-PYL10 displayed the strongest tendency to form a closure conformation (Figure 5). Thus, ligand-free PYL10 is able to adopt a compatible conformation for PP2C recognition in the absence of ABA.
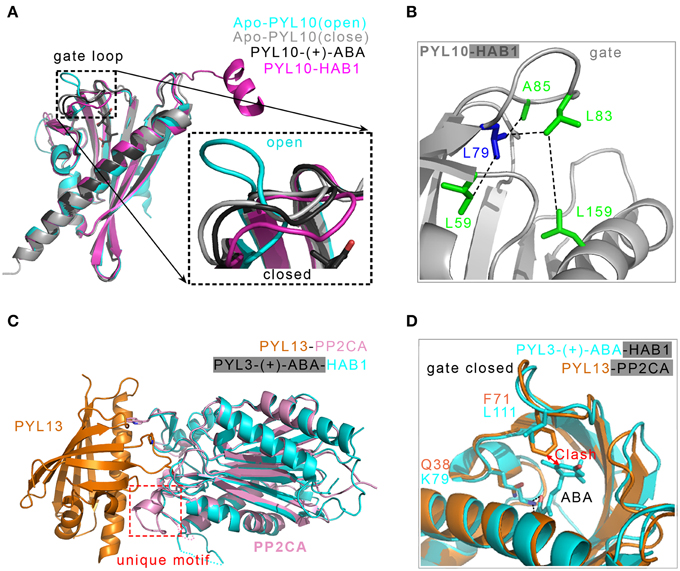
Figure 5. Structural basis of the ABA-independent inhibition of PP2Cs by both PYL13-PP2CA and PYL10-HAB1 complexes. (A) Structural superposition of two apo-PYL10 (PDB ID: 3RT2 and 3UQH), PYL10-(+)-ABA (PDB ID: 3R6P) and PYL10-HAB1 (PDB ID: 3RT0). A putative two-conformation dynamical equilibrium may exist (Sun et al., 2012); therefore, PYL10 displays a high tendency to inhibit PP2Cs in the absence of (+)-ABA (Hao et al., 2011). (B) Structural analysis of the constitutive inhibition of PP2Cs by PYL10. The bulkier hydrophobic residue L79 in PYL10, demarcating the β3 strand and the “gate” loop, provides a platform to dock the hydrophobic L83 in the “gate” loop and close it (Hao et al., 2011). (C) Two PP2Cs from PYL13-PP2CA (PDB ID: 4N0G) and PYL3-(+)-CityABA-HAB1 (PDB ID: 4DS8) are superimposed, the PYL3-(+)-ABA in PYL3-(+)-ABA-HAB1 is not displayed. The PP2CA-unique motif supplies a new interface with PYL13 (Li et al., 2013). (D) Two major events are responsible for the ABA-irresponsive and constitutive inhibition of PP2CA by PYL13. The F71 in the “gate” loop collides with the cyclohexene plane, and the Q38 in PYL13 substituting for the conserved lysine loses the essential salt bonds with the carboxylic group of (+)-ABA (Li et al., 2013). Two PP2Cs including HAB1 and PP2CA are not shown.
However, based on the two abovementioned principles for the constitutive inhibition of PP2Cs, the PYL2V87L/I88K double- mutation, incorporating PYL10's unique “gate” sequence into the low basal-activity receptor PYL2, only partially augments the constitutive inhibition ability onto PP2Cs (Hao et al., 2011). To create constitutively active receptors, a recent study performed site-saturation mutagenesis at 39 residues in PYR1 that were involved in contacting ABA or PP2Cs, and triple or quadruple mutations (PYR1H60P/V83F/F159V, PYR1H60P/V83F/M158I/F159V) with full activation were finally obtained and were indistinguishable from ABA-saturated PYR1 (Mosquna et al., 2011). These mutations would be useful tools for studying the functions of individual PYLs in vivo (Mosquna et al., 2011; Miyakawa et al., 2013).
Although some monomeric PYLs can bind to PP2Cs in the absence of ABA, the inhibitory efficiency is obviously lower compared to that in the presence of ABA. The constitutive interaction between PYLs and PP2Cs, as well as endogenous ABA content alteration, is perhaps orchestrated to regulate ABA signaling for plant survival and growth (Santiago et al., 2009b; Hao et al., 2011).
ABA-Irresponsive Inhibition of PP2Cs
There are 14 members of the PYLs family (Ma et al., 2009; Park et al., 2009). In vivo reconstitution assays in Arabidopsis protoplasts indicated that all of the PYLs except for PYL13 function as ABA receptors in the ABA signaling pathway (Fujii et al., 2009). It was reported that PP2Cs could physically interact with PYL13 (Joshi-Saha et al., 2011). Recently, one group reported that PYL13 can modulate the ABA pathway by interacting with and inhibiting both other PYLs and PP2Cs (Zhao et al., 2013). Nevertheless, PYL13 has an ABA receptor function by interacting with PP2C in early plant development (Fuchs et al., 2014). The PYL13-PP2CA complex structure elucidated the molecular basis for the specific interaction between PP2CA and PYL13 (Li et al., 2013). PYL13-PP2CA shares the common binding features of other PYLs and PP2Cs in the presence of (+)-ABA. In addition, the unique motif of PP2CA, including a short helix and a loop, is not found in ABI1 and ABI2 and is different from that in HAB1 and HAB2. This unique motif supplies a new interface with PYL13 mainly through direct and water-mediated hydrogen bonds (Figure 5C). The known structures demonstrated that a conserved lysine residue in the PYLs pocket is required to anchor the carboxylate group of ABA (Melcher et al., 2009; Miyazono et al., 2009; Nishimura et al., 2009; Santiago et al., 2009a; Yin et al., 2009; Zhang et al., 2012). However, the alignment of all of the PYLs sequences showed that the conserved lysine is substituted with the Q38 at the corresponding position of PYL13, which cripples the essential link with ABA. In addition, structural superimposition showed that there would be a steric clash between the aromatic ring of the F71 residue in PYL13 and the hydrophobic moiety of ABA. This clash can be completely abrogated by the residue leucine at the corresponding position of other PYLs (Li et al., 2013) (Figure 5D). Neither the Q38K nor F71L single mutation could endow PYL13 with response to ABA. The double mutation Q38K/F71L converted PYL13 into an ABA-dependent inhibitor to all of the tested PP2Cs, including PP2CA, ABI1, HAB1, and HAB2. Therefore, the lack of the conserved lysine in the pocket and the replacement of leucine with phenylalanine in the “gate” loop account for the ABA irresponsiveness of PYL13 (Li et al., 2013) (Figure 5D).
Although PYL13 and PYL10 show a high basal activity of inhibiting PP2Cs, both are different in response to ABA. PYL13 does not bind to ABA because of steric hindrance and the absence of the conserved lysine residue. PYL10 has the capacity to interact with ABA, and the potent constitutive inhibition of PYL10 onto PP2Cs can be further enhanced by ABA application. In addition, PYL13 antagonized PYL10 in the ABA-independent inhibition of PP2Cs because they were the only two PYLs with a leucine at the demarcation point of the β3 strand and the “gate” loop, facilitating the closure of the “gate” loop (Hao et al., 2011; Li et al., 2013).
The constitutive inhibition of PP2Cs can be achieved by the ABA-irresponsive receptor (such as PYL13) or ABA-responsive receptor (such as PYL10). These two types of receptors together with ABA-dependent receptors (such as PYR1 and PYL1-3) may provide a quantitative, fine-tuned and sensitive regulation in the ABA signaling network. Based on ABA-independent mechanisms, constitutively active receptors may be engineered and can be useful in the dissection of individual receptor function in vivo and in the generation of transgenic crops surviving environmental stresses (Hao et al., 2011; Mosquna et al., 2011; Li et al., 2013).
Structural Basis for SnRK2s Kinases in the ABA Signaling Pathway
Basal Activity and Autoactivation of SnRK2s Kinases
The SnRK2s family members were identified from wheat (Anderberg and Walker-Simmons, 1992), fava bean (Li et al., 2000) and Arabidopsis (Mustilli et al., 2002; Yoshida et al., 2002). The SnRK2s family contains 10 members in Arabidopsis, among which the subclass III SnRK2s (SRK2D/SnRK2.2, SRK2E/OST1/SnRK2.6 and SRK2I/SnRK2.3) are strongly responsive to ABA (Yoshida et al., 2006). Several snrk2s mutants, such as the decuple mutant snrk2.1/2/3/4/5/6/7/8/9/10 (=srk2g/d/e/a/h/e/f/c/j/b), the triple mutant snrk2.2/3/6 (=srk2d/e/i) and the septuple mutant snrk2.1/4/5/7/8/9/10 (=srk2g/a/h/f/c/j/b), completely preclude ABA responses, exhibiting conspicuously decreased tolerance to drought and dramatically increased insensitivity to ABA (Fujii and Zhu, 2009; Fujita et al., 2009) and a loss of dormancy and an ascent of seed ABA content compared to those of the wild-type (Fujita et al., 2009; Nakashima et al., 2009a; Fujii et al., 2011). The physiological functions of SnRK2s include transcriptional regulation, rapid regulations of stomatal closure (Pilot et al., 2001; Mustilli et al., 2002; Yoshida et al., 2002; Negi et al., 2008; Vahisalu et al., 2008; Geiger et al., 2009; Sato et al., 2009; Sirichandra et al., 2009; Chen et al., 2010; Kim et al., 2010), and response to ABA in seeds and vegetative tissues (Fujii et al., 2007). SnRK2s can also be activated by hyperosmotic stress (Mikolajczyk et al., 2000; Monks et al., 2001; Umezawa et al., 2004) in an activation pattern that is different from that of ABA (Boudsocq et al., 2004; Kobayashi et al., 2004). In particular, SnRK2.6 is important for the ABA-induced stomata closure in response to drought. SnRK2.2 and SnRK2.3 are predominantly responsible for the inhibition of seed germination and seedling growth in response to ABA. Subclass III SnRK2s contain a well-conserved catalytic domain and two conserved motifs. One motif is a highly acidic C-terminal segment that is termed ABA box, and the other one is a SnRK2 box that is important for kinase activity.
Upon ABA binding, PYLs undergo conformational changes to contact and inhibit PP2Cs (Melcher et al., 2009; Miyazono et al., 2009; Nishimura et al., 2009; Santiago et al., 2009a; Yin et al., 2009; Zhang et al., 2012). The inhibited PP2Cs result in the autoactivation of SnRK2s, which allows SnRK2s to relay the ABA signal (Cutler et al., 2010; Hubbard et al., 2010; Ng et al., 2011; Soon et al., 2012). Recently, the crystal structures of Arabidopsis SnRK2.3 and SnRK2.6 were reported (Ng et al., 2011) (Figures 4C–F). SnRK2.3 and SnRK2.6 kinases displayed a partially active state of phosphorylation-independent activity that can be activated by ABA intervention. Moreover, this partially active state can be extremely augmented by the phosphorylation of the activation loop by both intermolecularly (in trans) and intramolecularly (in cis) (Lochhead, 2009). Both SnRK2.3 and SnRK2.6 structures have the canonical kinase folds that are similar to those of AMPK and the yeast homolog Snf1 (Nayak et al., 2006; Chen et al., 2009; Littler et al., 2010) (Figures 4C,D). Compared to SnRK2.3, the SnRK2.6 structure seems to be more stable and contains more structural elements, including parts of the activation loop and the linker between the kinase domain and the SnRK2 box (Figure 4D). A noticeable feature is the SnRK2 box, containing a single α-helix that establishes extensive interactions with the αC helix. The intramolecular SnRK2s box-αC helix interaction, structurally resembling the intermolecular stabilization of the αC helix in Cdk2 by the helix α5 in cyclin (Jeffrey et al., 1995; Pavletich, 1999), is important for kinase activity. The active conformation structure of Pim-1, complexed with substrate peptide and the ATP analog AMP-PNP (Bullock et al., 2005), was superimposed with SnRK2.6 or SnRK2.3 to gain further insight into the mechanism of autophosphorylation. The detailed information about the two-step activation mechanism of SnRK2s kinases is also seen in the reference (Ng et al., 2011).
Simultaneously, two crystal structures of SnRK2.6 mutants (D160A and D160A/S175D) were reported, displaying an open inactive conformation and showing that the ABA-independent regulation motif (DI) stabilizes the conformation of the catalytically essential kinase αC helix (Yunta et al., 2011). The crystal structures of two catalytically inactive SnRK2.6 mutants are similar to the open, inactive conformation of the wild type (Ng et al., 2011).
The structures in combination with biochemical studies of SnRK2.6 and SnRK2.3 clarify the molecular basis of autophosphorylation activation of SnRK2s, which provides a complete structural framework for understanding the ABA-independent and -dependent regulations for a double-negative regulatory system (PYLs—|PP2C—|SnRK2).
SnRK2s Mimicking PYLs Dock into PP2Cs
There are several lines of evidence of the interaction between SnRK2s and PP2Cs. Yeast two-hybrid analysis demonstrated that ABI1 physically contacted SnRK2.6 (Yoshida et al., 2006). Group A PP2Cs directly inactivate subclass III SnRK2s in vitro by dephosphorylating multiple serine/threonine residues in the kinase activation loop (Umezawa et al., 2009). For example, HAB1 dephosphorylates the kinase within this activation domain to repress the kinase activity (Belin et al., 2006; Boudsocq et al., 2007; Vlad et al., 2009). The S175 residue of SnRK2.6 was identified as a target site of PP2Cs by screening substrates of HAB1 or HABG246D (Vlad et al., 2009). Collectively, in the absence of ABA, SnRK2s are inactivated by PP2Cs (Ma et al., 2009; Park et al., 2009; Umezawa et al., 2009; Yin et al., 2009), which physically bind SnRK2s kinases and dephosphorylate a key serine in the kinase activation loop (Belin et al., 2006; Boudsocq et al., 2007; Vlad et al., 2009).
Recently, the solved SnRK2.6-HAB1 complex structure showed the mutual packing of both the kinase and phosphatase active sites, which formed the major binding interface (Soon et al., 2012) (Figure 4G). SnRK2.6 contributes three separate regions within the kinase domain for binding to HAB1. First, the activation loop inserts deeply into the catalytic cleft of HAB1 and mimics the “gate” loop of PYLs. In addition, the region near residues R139, I183 and E144 of SnRK2.6 emulates the cleft that is formed by the “gate” and “latch” loops of PYLs. The conserved tryptophan (such as HAB1 W385) from PP2Cs inserts into this region. In addition, the SnRK2.6 αG helix binds to the region near the PYLs interaction site in HAB1 (Figure 4G). The SnRK2s-PP2Cs binding interface largely overlaps with that of PYLs-PP2Cs, and the HAB1 structure in the SnRK2.6-HAB1 complex is nearly identical to that in the PYLs-HAB1 complex (Figure 4H). Thus, there is a marked similarity in PP2Cs recognition by SnRK2s and ABA receptors. Both SnRK2.6 and PYLs use a similar gate-and-lock mechanism to recognize PP2Cs (Soon et al., 2012) (Figure 4H). In addition to the kinase domain, the highly acidic C-terminal ABA box of SnRK2s is critical for interaction with PP2Cs (Boudsocq et al., 2004; Kobayashi et al., 2004; Yoshida et al., 2006). However, the ABA box has no clear electron density in the SnRK2.6-HAB1 complex structure. Mutagenesis and hydrogen/deuterium exchange experiments indicated that the negatively charged ABA box interacts primarily with the positively charged PP2Cs surface (Soon et al., 2012).
The SnRK2.6–HAB1 structure suggests a two-step mechanism by which HAB1 completely inactivates SnRK2.6. The first step is mediated by the catalytic activity of HAB1, which dephosphorylates S175 in the activation loop, thus reducing SnRK2.6 activity to the basal level. The second step is the physical inhibition of the SnRK2.6 kinase domain by HAB1 (Soon et al., 2012).
In Arabidopsis, there are 14 members of PYLs, nine members of PP2Cs and 10 members of SnRK2s, and differential binding between PP2Cs and PYLs, PP2Cs and SnRK2s is important for regulating the physiological responses of Arabidopsis to adapt to stress conditions.
Application of the Studies of ABA Receptors for Agriculture
Plant growth can be severely and constantly challenged by adverse environmental stresses, such as drought, salinity, and temperature fluctuations. Drought is one of the major abiotic stresses, and high salinity is the most severe environmental stress. Both of these stresses are closely related and interfere with plant growth in the overlapped mechanisms. Freezing injury leads to physical damages to tissues at temperatures below 0°C due to the formation of ice crystals in plant cells. These abiotic stresses adversely affect plant growth and development, reduce productivity and cause significant crop losses (Thomashow, 1999; Marris, 2008; Battisti and Naylor, 2009; Hubbard et al., 2010; Nakashima and Yamaguchi-Shinozaki, 2013; Roychoudhury et al., 2013).
Given these disadvantages of stresses, plants in response to environmental cues are being exploited to improve crop yield and reduce economic losses. First, the up- or down-regulation of some ABA signaling factors may improve tolerance to environmental stresses. Transgenic plants that over-express AREB/ABFs, such as DREB1A and OsbZIP23, showed improved drought tolerance (Xiang et al., 2008; Nakashima et al., 2009b; Fujita et al., 2011). In addition, the molecular mechanisms and wealth of structural information of the core ABA signaling pathway supply an approach of engineering the components, such as PYLs or PP2Cs. A recent study showed that the triple mutant (PYR1H60P/V83F/F159V) and the quadruple mutant (PYR1H60P/V83F/M158I/F159V), which were indistinguishable from ABA-saturated PYR1, stabilized their agonist-bound conformation to activate ABA signaling in vivo (Mosquna et al., 2011). The engineered HAB1W385A mutant could modulate ABA signaling in vivo through the constitutive inactivation of the kinase SnRK2.6 even in the presence of ABA and PYLs (Dupeux et al., 2011a). In addition, the synthetic ABA analogs mimic ABA, which could control plant growth and development. (−)-ABA (Walker-Simmons et al., 1992; Nambara et al., 2002; Huang et al., 2007; Sirichandra et al., 2009; Zhang et al., 2013) and pyrabactin or its analogs (Park et al., 2009; Cutler et al., 2010; Hao et al., 2010; Melcher et al., 2010a; Peterson et al., 2010; Yuan et al., 2010; Mosquna et al., 2011; Zhang et al., 2012) had profound effects on plant growth, such as seed germination (Zhao et al., 2007). Last but not least, exploiting other plant growth regulators, ABA synthesis or an equivalent compound may prevent a decline in crop production and agricultural economy due to environmental stress.
Conclusion and Perspectives
The accumulating evidence indicate that PYLs function as bona fide ABA receptors, converging all aspects of ABA signaling. PYLs-mediated ABA signaling could play a crucial role in favoring stress adaptation and growth development for plants. ABA perception by PYLs receptors is orchestrated by PP2Cs and SnRK2s to establish a double-negative regulatory system for core ABA signaling, which controls ABA signaling in rapid stomatal closure responses for guard cells, as well as in long distance at the transcriptional level for seeds and vegetative tissues in response to water deficit. The detailed structural analyses, including ABA binding to PYLs, ABA-bound PYLs inhibiting PP2Cs and the ABA-induced inhibition of PP2Cs leading to the autoactivation of SnRK2 kinases, explain the gate-latch-lock mechanism (Melcher et al., 2009) and the two-step activation mechanism of SnRK2s kinases (Ng et al., 2011). In addition, the structural examination of SnRK2.6-HAB1 binary complex elucidated the two-step inactivation mechanism of SnRK2s by PP2Cs (Soon et al., 2012). These structural results agree with a substantial body of findings from biochemistry, molecular biology, and genetics. Although substantial progress has occurred in recent years, many questions remain. For example, why does the ABA signaling pathway in Arabidopsis involve 14 members in the PYLs family? What is the structural mechanism of individual receptors in their sensitivity and specialty in response to ABA? How can we exploit new selective ABA agonists and antagonists to ameliorate crop and ornamental plants? Finally, this review is just the beginning of understanding the ABA signaling pathway, and more work is required to gain global mechanistic insight into the complete signaling network.
Summary
This review summarizes the structural information of multiple regulatory mechanisms and ABA signal transduction pathways by PYLs. This structural information elucidates the ABA-responsive and constitutive inhibition mechanisms, signaling pathways of ABA and ABA analog perception by PYLs and provides a basis for the further design of selective ABA analogs and agricultural application.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We apologize for not being able to review all relevant work due to space restrictions. No conflict of interest declared. This work was supported by the National Basic Research Program of China (973 Program, 2011CB965304) and National Natural Science Foundation of China (31222032, 31370720) to ZZC, and by the National Natural Science Foundation of China (31300602), Starting Research Fund from Guangdong Medical College (XB1341) and Medical Research Foundation of Guangdong (B2013307) to XLZ.
Supplementary Material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/journal/10.3389/fpls.2015.00088/abstract
Abbreviations
ABA, Abscisic acid; ABI1/2, ABA insensitive 1/2; ABRE, ABA-responsive element; ABF, ABRE-binding factor; TFs, transcription factors; HAB1/2, Homology to ABA 1/2; PYR1, Pyrabactin resistance 1; PYL, PYR1-like; RCAR1, Regulatory component of ABA receptor 1; SnRK2, SNF1-related protein kinase 2; PP2C, 2C-type protein phosphatase.
References
Addicott, F. T., and Lyon, J. L. (1969). Physiology of abscisic acid and related substances. Ann. Rev. Plant Physiol. 20, 139–164. doi: 10.1146/annurev.pp.20.060169.001035
Anderberg, R. J., and Walker-Simmons, M. K. (1992). Isolation of a wheat cDNA clone for an abscisic acid-inducible transcript with homology to protein kinases. Proc. Natl. Acad. Sci. U.S.A. 89, 10183–10187. doi: 10.1073/pnas.89.21.10183
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Antoni, R., Gonzalez-Guzman, M., Rodriguez, L., Peirats-Llobet, M., Pizzio, G. A., Fernandez, M. A., et al. (2013). PYRABACTIN RESISTANCE1-LIKE8 plays an important role for the regulation of abscisic acid signaling in root. Plant Physiol. 161, 931–941. doi: 10.1104/pp.112.208678
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Battisti, D. S., and Naylor, R. L. (2009). Historical warnings of future food insecurity with unprecedented seasonal heat. Science 323, 240–244. doi: 10.1126/science.1164363
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Belin, C., de Franco, P. O., Bourbousse, C., Chaignepain, S., Schmitter, J. M., Vavasseur, A., et al. (2006). Identification of features regulating OST1 kinase activity and OST1 function in guard cells. Plant Physiol. 141, 1316–1327. doi: 10.1104/pp.106.079327
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Bennet-Clark, T. A., and Kefford, N. P. (1953). Chromatography of the growth substances in plant extracts. Nature 171, 645–647. doi: 10.1038/171645a0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Boudsocq, M., Barbier-Brygoo, H., and Lauriere, C. (2004). Identification of nine sucrose nonfermenting 1-related protein kinases 2 activated by hyperosmotic and saline stresses in Arabidopsis thaliana. J. Biol. Chem. 279, 41758–41766. doi: 10.1074/jbc.M405259200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Boudsocq, M., Droillard, M. J., Barbier-Brygoo, H., and Lauriere, C. (2007). Different phosphorylation mechanisms are involved in the activation of sucrose non-fermenting 1 related protein kinases 2 by osmotic stresses and abscisic acid. Plant Mol. Biol. 63, 491–503. doi: 10.1007/s11103-006-9103-1
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Bullock, A. N., Debreczeni, J., Amos, A. L., Knapp, S., and Turk, B. E. (2005). Structure and substrate specificity of the Pim-1 kinase. J. Biol. Chem. 280, 41675–41682. doi: 10.1074/jbc.M510711200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Cao, M., Liu, X., Zhang, Y., Xue, X., Zhou, X. E., Melcher, K., et al. (2013). An ABA-mimicking ligand that reduces water loss and promotes drought resistance in plants. Cell Res. 23, 1043–1054. doi: 10.1038/cr.2013.95
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chen, L., Jiao, Z. H., Zheng, L. S., Zhang, Y. Y., Xie, S. T., Wang, Z. X., et al. (2009). Structural insight into the autoinhibition mechanism of AMP-activated protein kinase. Nature 459, 1146–1149. doi: 10.1038/nature08075
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chen, Y. H., Hu, L., Punta, M., Bruni, R., Hillerich, B., Kloss, B., et al. (2010). Homologue structure of the SLAC1 anion channel for closing stomata in leaves. Nature 467, 1074–1080. doi: 10.1038/nature09487
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Cornforth, J. W., Milborrow, B. V., and Ryback, G. (1965a). Synthesis of (+/-)-Abscisin II. Nature 206, 715–715. doi: 10.1038/206715a0
Cornforth, J. W., Milborrow, B. V., Ryback, G., and Wareing, P. F. (1965b). Chemistry and physiology of ‘dormins’ in sycamore: identity of sycamore ‘dormin’ with abscisin II. Nature 205, 1269–1270. doi: 10.1038/2051269b0
Cutler, S. R., Rodriguez, P. L., Finkelstein, R. R., and Abrams, S. R. (2010). Abscisic acid: emergence of a core signaling network. Annu. Rev. Plant Biol. 61, 651–679. doi: 10.1146/annurev-arplant-042809-112122
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Dupeux, F., Antoni, R., Betz, K., Santiago, J., Gonzalez-Guzman, M., Rodriguez, L., et al. (2011a). Modulation of abscisic acid signaling in vivo by an engineered receptor-insensitive protein phosphatase type 2C allele. Plant Physiol. 156, 106–116. doi: 10.1104/pp.110.170894
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Dupeux, F., Santiago, J., Betz, K., Twycross, J., Park, S. Y., Rodriguez, L., et al. (2011b). A thermodynamic switch modulates abscisic acid receptor sensitivity. EMBO J. 30, 4171–4184. doi: 10.1038/emboj.2011.294
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Finkelstein, R., Reeves, W., Ariizumi, T., and Steber, C. (2008). Molecular aspects of seed dormancy. Annu. Rev. Plant Biol. 59, 387–415. doi: 10.1146/annurev.arplant.59.032607.092740
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Finkelstein, R. R., Gampala, S. S., and Rock, C. D. (2002). Abscisic acid signaling in seeds and seedlings. Plant Cell 14(Suppl.), S15–S45.
Fuchs, S., Tischer, S. V., Wunschel, C., Christmann, A., and Grill, E. (2014). Abscisic acid sensor RCAR7/PYL13, specific regulator of protein phosphatase coreceptors. Proc. Natl. Acad. Sci. U.S.A. 111, 5741–5746. doi: 10.1073/pnas.1322085111
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fujii, H., Chinnusamy, V., Rodrigues, A., Rubio, S., Antoni, R., Park, S. Y., et al. (2009). In vitro reconstitution of an abscisic acid signalling pathway. Nature 462, 660–664. doi: 10.1038/nature08599
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fujii, H., Verslues, P. E., and Zhu, J. K. (2007). Identification of two protein kinases required for abscisic acid regulation of seed germination, root growth, and gene expression in Arabidopsis. Plant Cell 19, 485–494. doi: 10.1105/tpc.106.048538
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fujii, H., Verslues, P. E., and Zhu, J. K. (2011). Arabidopsis decuple mutant reveals the importance of SnRK2 kinases in osmotic stress responses in vivo. Proc. Natl. Acad. Sci. U.S.A. 108, 1717–1722. doi: 10.1073/pnas.1018367108
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fujii, H., and Zhu, J. K. (2009). Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc. Natl. Acad. Sci. U.S.A. 106, 8380–8385. doi: 10.1073/pnas.0903144106
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fujita, Y., Fujita, M., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2011). ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 124, 509–525. doi: 10.1007/s10265-011-0412-3
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fujita, Y., Nakashima, K., Yoshida, T., Katagiri, T., Kidokoro, S., Kanamori, N., et al. (2009). Three SnRK2 protein kinases are the main positive regulators of abscisic acid signaling in response to water stress in Arabidopsis. Plant Cell Physiol. 50, 2123–2132. doi: 10.1093/pcp/pcp147
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fujita, Y., Yoshida, T., and Yamaguchi-Shinozaki, K. (2013). Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants. Physiol. Plant. 147, 15–27. doi: 10.1111/j.1399-3054.2012.01635.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Furihata, T., Maruyama, K., Fujita, Y., Umezawa, T., Yoshida, R., Shinozaki, K., et al. (2006). Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Proc. Natl. Acad. Sci. U.S.A. 103, 1988–1993. doi: 10.1073/pnas.0505667103
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Geiger, D., Scherzer, S., Mumm, P., Stange, A., Marten, I., Bauer, H., et al. (2009). Activity of guard cell anion channel SLAC1 is controlled by drought-stress signaling kinase-phosphatase pair. Proc. Natl. Acad. Sci. U.S.A. 106, 21425–21430. doi: 10.1073/pnas.0912021106
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Gosti, F., Beaudoin, N., Serizet, C., Webb, A. A., Vartanian, N., and Giraudat, J. (1999). ABI1 protein phosphatase 2C is a negative regulator of abscisic acid signaling. Plant Cell 11, 1897–1910. doi: 10.1105/tpc.11.10.1897
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hao, Q., Yin, P., Li, W., Wang, L., Yan, C., Lin, Z., et al. (2011). The Molecular Basis of ABA-Independent Inhibition of PP2Cs by a Subclass of PYL Proteins. Mol. Cell 42, 662–672. doi: 10.1016/j.molcel.2011.05.011
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hao, Q., Yin, P., Yan, C., Yuan, X., Li, W., Zhang, Z., et al. (2010). Functional mechanism of the abscisic acid agonist pyrabactin. J. Biol. Chem. 285, 28946–28952. doi: 10.1074/jbc.M110.149005
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hauser, F., Waadt, R., and Schroeder, J. I. (2011). Evolution of abscisic acid synthesis and signaling mechanisms. Curr. Biol. 21, R346–R355. doi: 10.1016/j.cub.2011.03.015
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hetherington, A. M. (2001). Guard cell signaling. Cell 107, 711–714. doi: 10.1016/S0092-8674(01)00606-7
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Huang, D., Jaradat, M. R., Wu, W., Ambrose, S. J., Ross, A. R., Abrams, S. R., et al. (2007). Structural analogs of ABA reveal novel features of ABA perception and signaling in Arabidopsis. Plant J. 50, 414–428. doi: 10.1111/j.1365-313X.2007.03056.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hubbard, K. E., Nishimura, N., Hitomi, K., Getzoff, E. D., and Schroeder, J. I. (2010). Early abscisic acid signal transduction mechanisms: newly discovered components and newly emerging questions. Genes Dev. 24, 1695–1708. doi: 10.1101/gad.1953910
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Iyer, L. M., Koonin, E. V., and Aravind, L. (2001). Adaptations of the helix-grip fold for ligand binding and catalysis in the START domain superfamily. Proteins 43, 134–144. doi: 10.1002/1097-0134(20010501)43:2<134::AID-PROT1025>3.0.CO;2-I
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Jeffrey, P. D., Russo, A. A., Polyak, K., Gibbs, E., Hurwitz, J., Massague, J., et al. (1995). Mechanism of CDK activation revealed by the structure of a cyclinA-CDK2 complex. Nature 376, 313–320. doi: 10.1038/376313a0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Joshi-Saha, A., Valon, C., and Leung, J. (2011). Abscisic acid signal off the STARting block. Mol. Plant 4, 562–580. doi: 10.1093/mp/ssr055
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kim, T. H., Bohmer, M., Hu, H., Nishimura, N., and Schroeder, J. I. (2010). Guard cell signal transduction network: advances in understanding abscisic acid, CO2, and Ca2+ signaling. Annu. Rev. Plant Biol. 61, 561–591. doi: 10.1146/annurev-arplant-042809-112226
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Klingler, J. P., Batelli, G., and Zhu, J. K. (2010). ABA receptors: the START of a new paradigm in phytohormone signalling. J. Exp. Bot. 61, 3199–3210. doi: 10.1093/jxb/erq151
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kobayashi, Y., Yamamoto, S., Minami, H., Kagaya, Y., and Hattori, T. (2004). Differential activation of the rice sucrose nonfermenting1-related protein kinase2 family by hyperosmotic stress and abscisic acid. Plant Cell 16, 1163–1177. doi: 10.1105/tpc.019943
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Koornneef, M., Reuling, G., and Karssen, C. M. (1984). The isolation and characterization of abscisic acid-insensitive mutants of Arabidopsis thaliana. Physiol. Plant. 61, 377–383. doi: 10.1111/j.1399-3054.1984.tb06343.x
Lee, K. H., Piao, H. L., Kim, H. Y., Choi, S. M., Jiang, F., Hartung, W., et al. (2006). Activation of glucosidase via stress-induced polymerization rapidly increases active pools of abscisic acid. Cell 126, 1109–1120. doi: 10.1016/j.cell.2006.07.034
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Leung, J., Bouvier-Durand, M., Morris, P. C., Guerrier, D., Chefdor, F., and Giraudat, J. (1994). Arabidopsis ABA response gene ABI1: features of a calcium-modulated protein phosphatase. Science 264, 1448–1452. doi: 10.1126/science.7910981
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Leung, J., Merlot, S., and Giraudat, J. (1997). The Arabidopsis ABSCISIC ACID-INSENSITIVE2 (ABI2) and ABI1 genes encode homologous protein phosphatases 2C involved in abscisic acid signal transduction. Plant Cell 9, 759–771. doi: 10.1105/tpc.9.5.759
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Li, J., Wang, X.-Q., Watson, M. B., and Assmann, S. M. (2000). Regulation of abscisic acid-induced stomatal closure and anion channels by guard cell AAPK kinase. Science 287, 300–303. doi: 10.1126/science.287.5451.300
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Li, W., Wang, L., Sheng, X., Yan, C., Zhou, R., Hang, J., et al. (2013). Molecular basis for the selective and ABA-independent inhibition of PP2CA by PYL13. Cell Res. 23, 1369–1379. doi: 10.1038/cr.2013.143
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lin, B. L., Wang, H. J., Wang, J. S., Zaharia, L. I., and Abrams, S. R. (2005). Abscisic acid regulation of heterophylly in Marsilea quadrifolia L: effects of R-(−) and S-(+) isomers. J. Exp. Bot. 56, 2935–2948. doi: 10.1093/jxb/eri290
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Littler, D. R., Walker, J. R., Davis, T., Wybenga-Groot, L. E., Finerty, P. J. Jr., Newman, E., et al. (2010). A conserved mechanism of autoinhibition for the AMPK kinase domain: ATP-binding site and catalytic loop refolding as a means of regulation. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 66, 143–151. doi: 10.1107/S1744309109052543
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Liu, X., Yue, Y., Li, B., Nie, Y., Li, W., Wu, W.-H., et al. (2007). A G protein–coupled receptor is a plasma membrane receptor for the plant hormone abscisic acid. Science 315, 1712–1716. doi: 10.1126/science.1135882
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lochhead, P. A. (2009). Protein kinase activation loop autophosphorylation in cis: overcoming a Catch-22 situation. Sci. Signal. 2:pe4. doi: 10.1126/scisignal.254pe4
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ma, Y., Szostkiewicz, I., Korte, A., Moes, D., Yang, Y., Christmann, A., et al. (2009). Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324, 1064–1068. doi: 10.1126/science.1172408
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Marris, E. (2008). Water: more crop per drop. Nature 452, 273–277. doi: 10.1038/452273a
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
McCourt, P. (1999). Genetic analysis of hormone signaling. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 219–243. doi: 10.1146/annurev.arplant.50.1.219
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Melcher, K., Ng, L. M., Zhou, X. E., Soon, F. F., Xu, Y., Suino-Powell, K. M., et al. (2009). A gate-latch-lock mechanism for hormone signalling by abscisic acid receptors. Nature 462, 602–608. doi: 10.1038/nature08613
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Melcher, K., Xu, Y., Ng, L. M., Zhou, X. E., Soon, F. F., Chinnusamy, V., et al. (2010a). Identification and mechanism of ABA receptor antagonism. Nat. Struct. Mol. Biol. 17, 1102–1108. doi: 10.1038/nsmb.1887
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Melcher, K., Zhou, X. E., and Xu, H. E. (2010b). Thirsty plants and beyond: structural mechanisms of abscisic acid perception and signaling. Curr. Opin. Struct. Biol. 20, 722–729. doi: 10.1016/j.sbi.2010.09.007
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Merlot, S., Gosti, F., Guerrier, D., Vavasseur, A., and Giraudat, J. (2001). The ABI1 and ABI2 protein phosphatases 2C act in a negative feedback regulatory loop of the abscisic acid signalling pathway. Plant J. 25, 295–303. doi: 10.1046/j.1365-313x.2001.00965.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Meyer, K., Leube, M. P., and Grill, E. (1994). A protein phosphatase 2C involved in ABA signal transduction in Arabidopsis thaliana. Science 264, 1452–1455. doi: 10.1126/science.8197457
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mikolajczyk, M., Awotunde, O. S., Muszynska, G., Klessig, D. F., and Dobrowolska, G. (2000). Osmotic stress induces rapid activation of a salicylic acid-induced protein kinase and a homolog of protein kinase ASK1 in tobacco cells. Plant Cell 12, 165–178. doi: 10.1105/tpc.12.1.165
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Milborrow, B. V. (1967). The identification of (+)-abscisin II [(+)-dormin] in plants and measurement of its concentrations. Planta 76, 93–113. doi: 10.1007/BF00385456
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Milborrow, B. V. (1974). The chemistry and physiology of abscisic acid. Annu. Rev. Plant Physiol. 25, 259–307. doi: 10.1146/annurev.pp.25.060174.001355
Miyakawa, T., Fujita, Y., Yamaguchi-Shinozaki, K., and Tanokura, M. (2013). Structure and function of abscisic acid receptors. Trends Plant Sci. 18, 259–266. doi: 10.1016/j.tplants.2012.11.002
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Miyazono, K., Miyakawa, T., Sawano, Y., Kubota, K., Kang, H. J., Asano, A., et al. (2009). Structural basis of abscisic acid signalling. Nature 462, 609–614. doi: 10.1038/nature08583
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Monks, D. E., Aghoram, K., Courtney, P. D., Dewald, D. B., and Dewey, R. E. (2001). Hyperosmotic stress induces the rapid phosphorylation of a soybean phosphatidylinositol transfer protein homolog through activation of the protein kinases SPK1 and SPK2. Plant Cell 13, 1205–1219. doi: 10.1105/tpc.13.5.1205
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mosquna, A., Peterson, F. C., Park, S. Y., Lozano-Juste, J., Volkman, B. F., and Cutler, S. R. (2011). Potent and selective activation of abscisic acid receptors in vivo by mutational stabilization of their agonist-bound conformation. Proc. Natl. Acad. Sci. U.S.A. 108, 20838–20843. doi: 10.1073/pnas.1112838108
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Muller, A. H., and Hansson, M. (2009). The barley magnesium chelatase 150-kd subunit is not an abscisic acid receptor. Plant Physiol. 150, 157–166. doi: 10.1104/pp.109.135277
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mustilli, A. C., Merlot, S., Vavasseur, A., Fenzi, F., and Giraudat, J. (2002). Arabidopsis OST1 protein kinase mediates the regulation of stomatal aperture by abscisic acid and acts upstream of reactive oxygen species production. Plant Cell 14, 3089–3099. doi: 10.1105/tpc.007906
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nakagawa, M., Kagiyama, M., Shibata, N., Hirano, Y., and Hakoshima, T. (2014). Mechanism of high-affinity abscisic acid binding to PYL9/RCAR1. Genes Cells 19, 386–404. doi: 10.1111/gtc.12140
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nakashima, K., Fujita, Y., Kanamori, N., Katagiri, T., Umezawa, T., Kidokoro, S., et al. (2009a). Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/SnRK2.3, involved in ABA signaling are essential for the control of seed development and dormancy. Plant Cell Physiol. 50, 1345–1363. doi: 10.1093/pcp/pcp083
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nakashima, K., Ito, Y., and Yamaguchi-Shinozaki, K. (2009b). Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol. 149, 88–95. doi: 10.1104/pp.108.129791
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nakashima, K., and Yamaguchi-Shinozaki, K. (2013). ABA signaling in stress-response and seed development. Plant Cell Rep. 32, 959–970. doi: 10.1007/s00299-013-1418-1
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nambara, E., and Marion-Poll, A. (2005). Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 56, 165–185. doi: 10.1146/annurev.arplant.56.032604.144046
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nambara, E., Suzuki, M., Abrams, S., McCarty, D. R., Kamiya, Y., and McCourt, P. (2002). A screen for genes that function in abscisic acid signaling in Arabidopsis thaliana. Genetics 161, 1247–1255.
Nayak, V., Zhao, K., Wyce, A., Schwartz, M. F., Lo, W. S., Berger, S. L., et al. (2006). Structure and dimerization of the kinase domain from yeast Snf1, a member of the Snf1/AMPK protein family. Structure 14, 477–485. doi: 10.1016/j.str.2005.12.008
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Negi, J., Matsuda, O., Nagasawa, T., Oba, Y., Takahashi, H., Kawai-Yamada, M., et al. (2008). CO2 regulator SLAC1 and its homologues are essential for anion homeostasis in plant cells. Nature 452, 483–486. doi: 10.1038/nature06720
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ng, L. M., Soon, F. F., Zhou, X. E., West, G. M., Kovach, A., Suino-Powell, K. M., et al. (2011). Structural basis for basal activity and autoactivation of abscisic acid (ABA) signaling SnRK2 kinases. Proc. Natl. Acad. Sci. U.S.A. 108, 21259–21264. doi: 10.1073/pnas.1118651109
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nishimura, N., Hitomi, K., Arvai, A. S., Rambo, R. P., Hitomi, C., Cutler, S. R., et al. (2009). Structural mechanism of abscisic acid binding and signaling by dimeric PYR1. Science 326, 1373–1379. doi: 10.1126/science.1181829
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nishimura, N., Sarkeshik, A., Nito, K., Park, S. Y., Wang, A., Carvalho, P. C., et al. (2010). PYR/PYL/RCAR family members are major in-vivo ABI1 protein phosphatase 2C-interacting proteins in Arabidopsis. Plant J. 61, 290–299. doi: 10.1111/j.1365-313X.2009.04054.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ohkuma, K., Lyon, J. L., Addicott, F. T., and Smith, O. E. (1963). Abscisin II, an abscission-accelerating substance from young cotton fruit. Science 142, 1592–1593. doi: 10.1126/science.142.3599.1592
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ohkuma, K. A. F., Smith, O. E, and Thiessen, W. E. (1965). The structure of abscisin II. Tetrahedron Lett. 6, 2529–2535. doi: 10.1016/S0040-4039(01)84019-3
Pandey, S., Nelson, D. C., and Assmann, S. M. (2009). Two novel GPCR-type G proteins are abscisic acid receptors in Arabidopsis. Cell 136, 136–148. doi: 10.1016/j.cell.2008.12.026
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Park, S. Y., Fung, P., Nishimura, N., Jensen, D. R., Fujii, H., Zhao, Y., et al. (2009). Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324, 1068–1071. doi: 10.1126/science.1173041
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Pavletich, N. P. (1999). Mechanisms of cyclin-dependent kinase regulation: structures of Cdks, their cyclin activators, and Cip and INK4 inhibitors. J. Mol. Biol. 287, 821–828. doi: 10.1006/jmbi.1999.2640
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Peterson, F. C., Burgie, E. S., Park, S. Y., Jensen, D. R., Weiner, J. J., Bingman, C. A., et al. (2010). Structural basis for selective activation of ABA receptors. Nat. Struct. Mol. Biol. 17, 1109–1113. doi: 10.1038/nsmb.1898
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Pilot, G., Lacombe, B., Gaymard, F., Cherel, I., Boucherez, J., Thibaud, J. B., et al. (2001). Guard cell inward K+ channel activity in arabidopsis involves expression of the twin channel subunits KAT1 and KAT2. J. Biol. Chem. 276, 3215–3221. doi: 10.1074/jbc.M007303200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Puli, M. R., and Raghavendra, A. S. (2012). Pyrabactin, an ABA agonist, induced stomatal closure and changes in signalling components of guard cells in abaxial epidermis of Pisum sativum. J. Exp. Bot. 63, 1349–1356. doi: 10.1093/jxb/err364
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Raghavendra, A. S., Gonugunta, V. K., Christmann, A., and Grill, E. (2010). ABA perception and signalling. Trends Plant Sci. 15, 395–401. doi: 10.1016/j.tplants.2010.04.006
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Razem, F. A., El-Kereamy, A., Abrams, S. R., and Hill, R. D. (2006). The RNA-binding protein FCA is an abscisic acid receptor. Nature 439, 290–294. doi: 10.1038/nature04373
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Razem, F. A., Luo, M., Liu, J. H., Abrams, S. R., and Hill, R. D. (2004). Purification and characterization of a barley aleurone abscisic acid-binding protein. J. Biol. Chem. 279, 9922–9929. doi: 10.1074/jbc.M311064200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Risk, J. M., Day, C. L., and Macknight, R. C. (2009). Reevaluation of abscisic acid-binding assays shows that G-Protein-Coupled Receptor2 does not bind abscisic Acid. Plant Physiol. 150, 6–11. doi: 10.1104/pp.109.135749
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Risk, J. M., Macknight, R. C., and Day, C. L. (2008). FCA does not bind abscisic acid. Nature 456, E5–E6. doi: 10.1038/nature07646
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Rodriguez, P. L., Leube, M. P., and Grill, E. (1998). Molecular cloning in Arabidopsis thaliana of a new protein phosphatase 2C (PP2C) with homology to ABI1 and ABI2. Plant Mol. Biol. 38, 879–883. doi: 10.1023/A:1006012218704
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Roychoudhury, A., Paul, S., and Basu, S. (2013). Cross-talk between abscisic acid-dependent and abscisic acid-independent pathways during abiotic stress. Plant Cell Rep. 32, 985–1006. doi: 10.1007/s00299-013-1414-5
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Saez, A., Apostolova, N., Gonzalez-Guzman, M., Gonzalez-Garcia, M. P., Nicolas, C., Lorenzo, O., et al. (2004). Gain-of-function and loss-of-function phenotypes of the protein phosphatase 2C HAB1 reveal its role as a negative regulator of abscisic acid signalling. Plant J. 37, 354–369. doi: 10.1046/j.1365-313X.2003.01966.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Saez, A., Robert, N., Maktabi, M. H., Schroeder, J. I., Serrano, R., and Rodriguez, P. L. (2006). Enhancement of abscisic acid sensitivity and reduction of water consumption in Arabidopsis by combined inactivation of the protein phosphatases type 2C ABI1 and HAB1. Plant Physiol. 141, 1389–1399. doi: 10.1104/pp.106.081018
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Santiago, J., Dupeux, F., Betz, K., Antoni, R., Gonzalez-Guzman, M., Rodriguez, L., et al. (2012). Structural insights into PYR/PYL/RCAR ABA receptors and PP2Cs. Plant Sci. 182, 3–11. doi: 10.1016/j.plantsci.2010.11.014
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Santiago, J., Dupeux, F., Round, A., Antoni, R., Park, S. Y., Jamin, M., et al. (2009a). The abscisic acid receptor PYR1 in complex with abscisic acid. Nature 462, 665–668. doi: 10.1038/nature08591
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Santiago, J., Rodrigues, A., Saez, A., Rubio, S., Antoni, R., Dupeux, F., et al. (2009b). Modulation of drought resistance by the abscisic acid receptor PYL5 through inhibition of clade A PP2Cs. Plant J. 60, 575–588. doi: 10.1111/j.1365-313X.2009.03981.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Santner, A., and Estelle, M. (2009). Recent advances and emerging trends in plant hormone signalling. Nature 459, 1071–1078. doi: 10.1038/nature08122
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Sato, A., Sato, Y., Fukao, Y., Fujiwara, M., Umezawa, T., Shinozaki, K., et al. (2009). Threonine at position 306 of the KAT1 potassium channel is essential for channel activity and is a target site for ABA-activated SnRK2/OST1/SnRK2.6 protein kinase. Biochem. J. 424, 439–448. doi: 10.1042/BJ20091221
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Schroeder, J. I., Kwak, J. M., and Allen, G. J. (2001). Guard cell abscisic acid signalling and engineering drought hardiness in plants. Nature 410, 327–330. doi: 10.1038/35066500
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Shen, Y. Y., Wang, X. F., Wu, F. Q., Du, S. Y., Cao, Z., Shang, Y., et al. (2006). The Mg-chelatase H subunit is an abscisic acid receptor. Nature 443, 823–826. doi: 10.1038/nature05176
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Sirichandra, C., Gu, D., Hu, H. C., Davanture, M., Lee, S., Djaoui, M., et al. (2009). Phosphorylation of the Arabidopsis AtrbohF NADPH oxidase by OST1 protein kinase. FEBS Lett. 583, 2982–2986. doi: 10.1016/j.febslet.2009.08.033
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Soon, F. F., Ng, L. M., Zhou, X. E., West, G. M., Kovach, A., Tan, M. H., et al. (2012). Molecular mimicry regulates ABA signaling by SnRK2 kinases and PP2C phosphatases. Science 335, 85–88. doi: 10.1126/science.1215106
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Sun, D., Wang, H., Wu, M., Zang, J., Wu, F., and Tian, C. (2012). Crystal structures of the Arabidopsis thaliana abscisic acid receptor PYL10 and its complex with abscisic acid. Biochem. Biophys. Res. Commun. 418, 122–127. doi: 10.1016/j.bbrc.2011.12.145
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Szostkiewicz, I., Richter, K., Kepka, M., Demmel, S., Ma, Y., Korte, A., et al. (2010). Closely related receptor complexes differ in their ABA selectivity and sensitivity. Plant J. 61, 25–35. doi: 10.1111/j.1365-313X.2009.04025.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Takeuchi, J., Okamoto, M., Akiyama, T., Muto, T., Yajima, S., Sue, M., et al. (2014). Designed abscisic acid analogs as antagonists of PYL-PP2C receptor interactions. Nat. Chem. Biol. 10, 477–482. doi: 10.1038/nchembio.1524
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Takezawa, D., Komatsu, K., and Sakata, Y. (2011). ABA in bryophytes: how a universal growth regulator in life became a plant hormone? J. Plant Res. 124, 437–453. doi: 10.1007/s10265-011-0410-5
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Thomashow, M. F. (1999). Plant cold acclimation: freezing tolerance genes and regulatory mechanisms. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 571–599. doi: 10.1146/annurev.arplant.50.1.571
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Tsuzuki, T., Takahashi, K., Inoue, S., Okigaki, Y., Tomiyama, M., Hossain, M. A., et al. (2011). Mg-chelatase H subunit affects ABA signaling in stomatal guard cells, but is not an ABA receptor in Arabidopsis thaliana. J. Plant Res. 124, 527–538. doi: 10.1007/s10265-011-0426-x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Umezawa, T., Nakashima, K., Miyakawa, T., Kuromori, T., Tanokura, M., Shinozaki, K., et al. (2010). Molecular basis of the core regulatory network in ABA responses: sensing, signaling and transport. Plant Cell Physiol. 51, 1821–1839. doi: 10.1093/pcp/pcq156
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Umezawa, T., Sugiyama, N., Mizoguchi, M., Hayashi, S., Myouga, F., Yamaguchi-Shinozaki, K., et al. (2009). Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 106, 17588–17593. doi: 10.1073/pnas.0907095106
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Umezawa, T., Yoshida, R., Maruyama, K., Yamaguchi-Shinozaki, K., and Shinozaki, K. (2004). SRK2C, a SNF1-related protein kinase 2, improves drought tolerance by controlling stress-responsive gene expression in Arabidopsis thaliana. Proc. Natl. Acad. Sci. U.S.A. 101, 17306–17311. doi: 10.1073/pnas.0407758101
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Vahisalu, T., Kollist, H., Wang, Y. F., Nishimura, N., Chan, W. Y., Valerio, G., et al. (2008). SLAC1 is required for plant guard cell S-type anion channel function in stomatal signalling. Nature 452, 487–491. doi: 10.1038/nature06608
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Vahisalu, T., Puzorjova, I., Brosche, M., Valk, E., Lepiku, M., Moldau, H., et al. (2010). Ozone-triggered rapid stomatal response involves the production of reactive oxygen species, and is controlled by SLAC1 and OST1. Plant J. 62, 442–453. doi: 10.1111/j.1365-313X.2010.04159.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Verslues, P. E., Agarwal, M., Katiyar-Agarwal, S., Zhu, J., and Zhu, J. K. (2006). Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status. Plant J. 45, 523–539. doi: 10.1111/j.1365-313X.2005.02593.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Vlad, F., Rubio, S., Rodrigues, A., Sirichandra, C., Belin, C., Robert, N., et al. (2009). Protein phosphatases 2C regulate the activation of the Snf1-related kinase OST1 by abscisic acid in Arabidopsis. Plant Cell 21, 3170–3184. doi: 10.1105/tpc.109.069179
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Walker-Simmons, M. K., Anderberg, R. J., Rose, P. A., and Abrams, S. R. (1992). Optically pure abscisic Acid analogs-tools for relating germination inhibition and gene expression in wheat embryos. Plant Physiol. 99, 501–507. doi: 10.1104/pp.99.2.501
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Wu, F. Q., Xin, Q., Cao, Z., Liu, Z. Q., Du, S. Y., Mei, C., et al. (2009). The magnesium-chelatase H subunit binds abscisic acid and functions in abscisic acid signaling: new evidence in Arabidopsis. Plant Physiol. 150, 1940–1954. doi: 10.1104/pp.109.140731
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Xiang, Y., Tang, N., Du, H., Ye, H., and Xiong, L. (2008). Characterization of OsbZIP23 as a key player of the basic leucine zipper transcription factor family for conferring abscisic acid sensitivity and salinity and drought tolerance in rice. Plant Physiol. 148, 1938–1952. doi: 10.1104/pp.108.128199
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Xie, Z., Zhang, Z. L., Zou, X., Huang, J., Ruas, P., Thompson, D., et al. (2005). Annotations and functional analyses of the rice WRKY gene superfamily reveal positive and negative regulators of abscisic acid signaling in aleurone cells. Plant Physiol. 137, 176–189. doi: 10.1104/pp.104.054312
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yin, P., Fan, H., Hao, Q., Yuan, X., Wu, D., Pang, Y., et al. (2009). Structural insights into the mechanism of abscisic acid signaling by PYL proteins. Nat. Struct. Mol. Biol. 16, 1230–1236. doi: 10.1038/nsmb.1730
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yoshida, R., Hobo, T., Ichimura, K., Mizoguchi, T., Takahashi, F., Aronso, J., et al. (2002). ABA-activated SnRK2 protein kinase is required for dehydration stress signaling in Arabidopsis. Plant Cell Physiol. 43, 1473–1483. doi: 10.1093/pcp/pcf188
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yoshida, R., Umezawa, T., Mizoguchi, T., Takahashi, S., Takahashi, F., and Shinozaki, K. (2006). The regulatory domain of SRK2E/OST1/SnRK2.6 interacts with ABI1 and integrates abscisic acid (ABA) and osmotic stress signals controlling stomatal closure in Arabidopsis. J. Biol. Chem. 281, 5310–5318. doi: 10.1074/jbc.M509820200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yuan, X., Yin, P., Hao, Q., Yan, C., Wang, J., and Yan, N. (2010). Single amino acid alteration between valine and isoleucine determines the distinct pyrabactin selectivity by PYL1 and PYL2. J. Biol. Chem. 285, 28953–28958. doi: 10.1074/jbc.M110.160192
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yunta, C., Martinez-Ripoll, M., Zhu, J. K., and Albert, A. (2011). The structure of Arabidopsis thaliana OST1 provides insights into the kinase regulation mechanism in response to osmotic stress. J. Mol. Biol. 414, 135–144. doi: 10.1016/j.jmb.2011.09.041
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zaharia, L. I., Walker-Simmon, M., Rodríguez, C., and Abrams, S. (2005). Chemistry of abscisic acid, abscisic acid catabolites and analogs. J. Plant Growth Regul. 24, 274–284. doi: 10.1007/s00344-005-0066-2
Zhang, D. P., Wu, Z. Y., Li, X. Y., and Zhao, Z. X. (2002). Purification and identification of a 42-kilodalton abscisic acid-specific-binding protein from epidermis of broad bean leaves. Plant Physiol. 128, 714–725. doi: 10.1104/pp.010531
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhang, X., Jiang, L., Wang, G., Yu, L., Zhang, Q., Xin, Q., et al. (2013). Structural insights into the abscisic acid stereospecificity by the ABA receptors PYR/PYL/RCAR. PLoS ONE 8:e67477. doi: 10.1371/journal.pone.0067477
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhang, X., Zhang, Q., Xin, Q., Yu, L., Wang, Z., Wu, W., et al. (2012). Complex structures of the abscisic acid receptor PYL3/RCAR13 reveal a unique regulatory mechanism. Structure 20, 780–790. doi: 10.1016/j.str.2012.02.019
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhao, Y., Chan, Z., Xing, L., Liu, X., Hou, Y. J., Chinnusamy, V., et al. (2013). The unique mode of action of a divergent member of the ABA-receptor protein family in ABA and stress signaling. Cell Res. 23, 1380–1395. doi: 10.1038/cr.2013.149
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhao, Y., Chow, T. F., Puckrin, R. S., Alfred, S. E., Korir, A. K., Larive, C. K., et al. (2007). Chemical genetic interrogation of natural variation uncovers a molecule that is glycoactivated. Nat. Chem. Biol. 3, 716–721. doi: 10.1038/nchembio.2007.32
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhao, Y., Xing, L., Wang, X., Hou, Y. J., Gao, J., Wang, P., et al. (2014). The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes. Sci. Signal. 7, ra53. doi: 10.1126/scisignal.2005051
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Keywords: abscisic acid, PYR/PYL/RCAR, PP2Cs, SnRK2s, ABA analogs, crystal structures
Citation: Zhang XL, Jiang L, Xin Q, Liu Y, Tan JX and Chen ZZ (2015) Structural basis and functions of abscisic acid receptors PYLs. Front. Plant Sci. 6:88. doi: 10.3389/fpls.2015.00088
Received: 06 December 2014; Accepted: 02 February 2015;
Published online: 19 February 2015.
Edited by:
Maria Hrmova, The Australian Centre for Plant Functional Genomics, AustraliaCopyright © 2015 Zhang, Jiang, Xin, Liu, Tan and Chen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xing L. Zhang, Department of Pediatrics, Affiliated Hospital of Guangdong Medical College, Zhanjiang 524001, China e-mail:Y2hldW5neGluZ2xpYW5nQGdtYWlsLmNvbQ==;
Zhong Z. Chen, State Key Laboratory of Agrobiotechnology, College of Biological Sciences, China Agricultural University, Beijing 100193, China e-mail:Y2hlbnpob25nemhvdUBjYXUuZWR1LmNu
 Xing L. Zhang
Xing L. Zhang Lun Jiang
Lun Jiang Qi Xin
Qi Xin Yang Liu
Yang Liu Jian X. Tan1
Jian X. Tan1 Zhong Z. Chen
Zhong Z. Chen