- Translational Research Laboratory of Transgenic Rice, Department of Botany, University of Calcutta, Kolkata, India
Small RNAs including micro RNAs (miRNA) play an indispensable role in cell signaling mechanisms. Generally, miRNAs that are 20–24 nucleotides long bind to specific complementary transcripts, attenuating gene expression at the post-transcriptional level or via translational inhibition. In plants, miRNAs have emerged as the principal regulator of various stress responses, including low nutrient availability. It has been reported that miRNAs are vital for maintaining nutrient homeostasis in plants by regulating the expression of transporters that are involved in nutrient uptake and mobilization. The present review highlights the role of various miRNAs in several macro- or micronutrient deficiencies in plants. Understanding the regulation of different transporters by miRNAs will aid in elucidating the underlying molecular signal transduction mechanisms during nutritional stress. Recent findings regarding nutrient related-miRNAs and their gene regulation machinery may delineate a novel platform for improving the nutritional status of cereal grains or crop biofortification programs in the future.
Introduction
Plants acquire mineral ions from the soil, which they allocate to various compartments following long- or short-distance transport. The underlying molecular mechanisms of uptake, transport, and loading of mineral ions into storage organs depend on the differential expression of various transporters based on mineral availability (Sperotto et al., 2014). The deficiency of mineral ions in the soil elicits signaling responses in plants, and interactions among signaling molecules and transporter or carrier genes can facilitate ion transport (Vigani et al., 2013a, b). If a plant is under nutrient starvation, the information is immediately transmitted to the genetic material via signaling molecules to maintain nutrient homeostasis. The scarcity of nutrients activates transporters or alters the architecture and growth of roots to enhance mineral uptake (Jung and McCouch, 2013). Signaling molecules interact with specific nucleotide sequences of mRNA to alter gene expression.
Small regulatory RNAs are considered the most ubiquitous signaling molecules that regulate post-transcriptional gene expression. These small RNAs bind with specific mRNA that exhibit their perfect complementary bases and attenuate gene expression (Bartel, 2004). In plants, two major types of small RNAs, small interfering RNAs (siRNAs) and microRNAs (miRNAs), are likely associated with the silencing of gene expression. siRNAs, which are small 20- to 24-nucleotide regulatory RNAs, are primarily derived from endogenous genomic regions or exogenously supplied duplexes of nucleic acid, as used in RNAi-mediated gene silencing technology (Jamalkandi and Masoudi-Nejad, 2009; Khraiwesh et al., 2012). Dicer-like enzymes (DCL in Arabidopsis) cleave the double-stranded duplex (with hairpin loop) to form short, perfect duplexes, which are loaded onto the RNA-induced silencing complex (RISC). The sense strand of the RISC complex recognizes the target gene or transcript and binds to the perfect complementary nucleotide sequences and regulates the gene expression level at post transcriptional or transcriptional levels (Schauer et al., 2002). However, miRNAs, 20–24 nucleotides in length, are derived from endogenous primary transcripts upon processing. The processed mature miRNA is incorporated into the RISC complex and directs the RISC complex toward degradation or the translational inhibition of target mRNAs (Lee et al., 2004; Vazquez et al., 2010).
In plants, miRNAs are primarily associated with several physiological phenomena, such as growth, development, organogenesis, and responses to various biotic and abiotic stresses (Chen, 2004; Sunkar and Zhu, 2004; Kidner and Martienssen, 2005; Lu et al., 2008; Zhang et al., 2008; Kruszka et al., 2012). Recently, several novel miRNAs were reported for the uptake and transport of nutrient minerals in plants (Jones-Rhoades and Bartel, 2004; Fischer et al., 2013; Kehr, 2013). In the present review, current knowledge of the different classes of nutrient responsive-miRNAs and of their functions in nutrient homeostasis was extensively studied, which may aid in understanding the role of miRNAs as master regulators of nutrient loading in plants. The regulation of the differential expression of transporter genes by miRNAs may reveal a novel paradigm for crop biofortification or mineral bioavailability in cereal grains in the future.
miRNAs in Nutrient Homeostasis
miRNA and Phosphorus Nutrition
Inorganic phosphate (Pi), an essential mineral for plant growth and development is often a limiting factor in plant nutrition. Despite the adequate amount of phosphorus (P) in cultivated land, plants can uptake very small amounts of phosphorus due to its low availability. Plants synthesize several classes of nucleases and phosphatases to solubilize external Pi or to release Pi from organic substances and to upregulate the gene expression of certain exporter/importer transport proteins to acclimatize to phosphate starvation (Poirier and Bucher, 2002; Abel, 2011). Recent studies have identified the involvement of small RNAs, particularly miRNAs, in the differential regulation of phosphate-related gene expression in Arabidopsis, rice, wheat, barley, maize, soybean, white lupin, and tomato, etc. (Table 1; Pant et al., 2009; Gu et al., 2010; Lundmark et al., 2010; Zhu et al., 2010; Hackenberg et al., 2013; Pei et al., 2013; Xu et al., 2013; Zhao et al., 2013a). In Arabidopsis, the deprivation of Pi is rapidly transmitted to the shoots and induces the synthesis of miR399a-f families. miR399 binds to the five complementary sites of the phosphate over accumulator2 (PHO2) transcript, which is in the 5′ untranslated region (UTR), 200–400 bp upstream and induces mRNA cleavage (Pant et al., 2008). PHO2, an essential transporter for phosphate mobilization, is associated with E2 ubiquitin conjugating enzyme. The reduction of PHO2 helps to accumulate high amounts of Pi in the shoots. The role of miR399 and its biosynthesis regulation have been well demonstrated. The myeloblastosis (MYB) transcription factors like phosphate starvation response1 (PHR1) binding sites, such as P1BS, PHO, and PHO-like, are present upstream of the promoter region of miR399 (Bari et al., 2006; Zeng et al., 2010; Xu et al., 2013). The expression of miR399f is also regulated by the binding of MYB2 transcription factor at the cis-acting elements of the MIR399f promoter, as reported in Arabidopsis (Baek et al., 2013). Notably, long non-coding RNA, another signaling molecule, which is induced by phosphate starvation1 (IPS1), plays an inhibitory role by protecting the PHO2 transcript and interferes with miR399-mediated PHO2 gene regulation (Franco-Zorrilla et al., 2007; Hu et al., 2011; Huang et al., 2011).
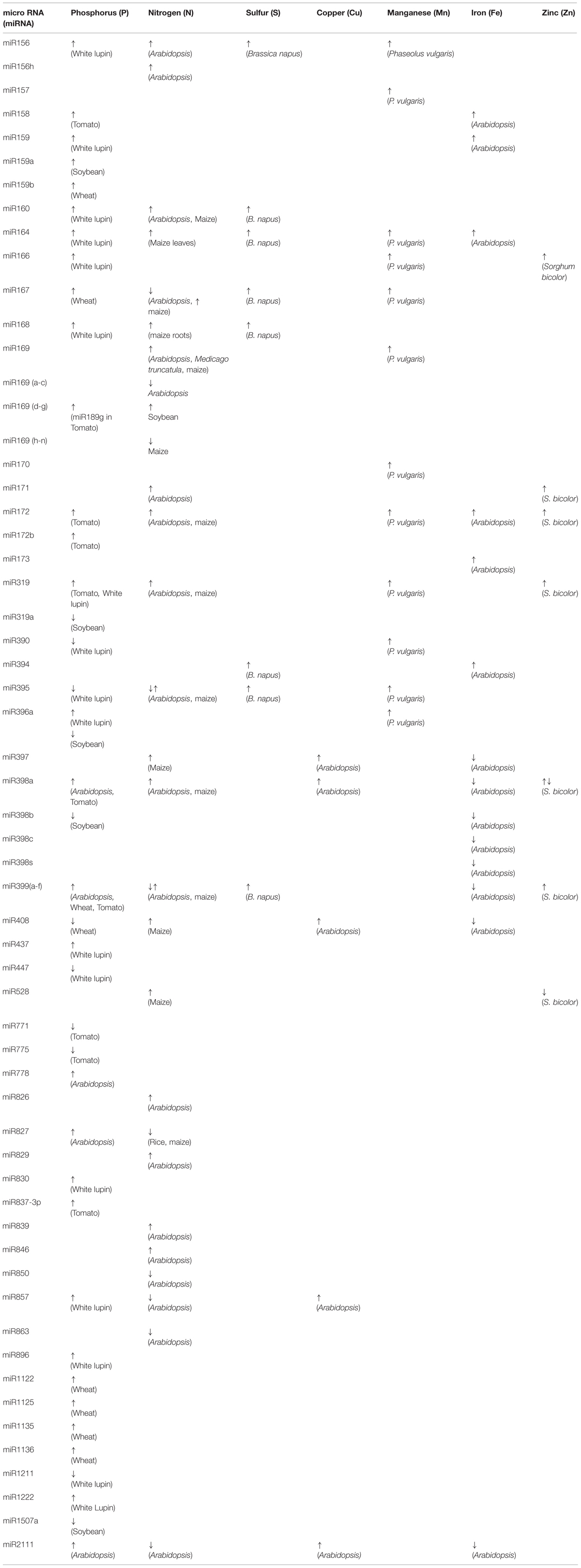
TABLE 1. Differentially expressed miRNAs reported under different nutrient/metal (N, P, S, Cu, Mn, Fe, Zn) deprivation in various plant species.
In Arabidopsis, phosphate limitation also increases the amounts of three other miRNAs: miR778, miR827, and miR2111. Intriguingly, within 3 h of Pi re-addition, the abundance of miR2111, and miR778 rapidly reduce by approximately twofold. Two copies of miR2111 are present in the Arabidopsis genome, and both loci show similar phosphate response activities (Pant et al., 2009). Recently, the dual responses of miR827, miR399, miR2111, and miR827 have been elucidated. The activity of miR2111 is reversed under nitrogen (N)-starvation compared to phosphate (P)-starvation (Liang et al., 2012). The miR827 and miR399 have also been identified to function in N-starvation by targeting the nitrogen limitation adaptation (NLA) gene and are thought to increase the expression of the PHO2 transporter. In lower abundances of nitrates, the nla mutant has been found to play an important role in phosphorus homeostasis by accumulating excessive Pi. A similar condition has been observed for miR827-overexpressing Arabidopsis plants and supports the miR827-mediated gene regulation of phosphate transporters (Kant et al., 2011). In rice, Pi accumulation followed by a perturbation of phosphorus mobilization in old leaves is also manifested by miR827 (Wang et al., 2012). Notably, the expression of miR398a is regulated by P, carbon (C) and N limitation, indicating a more general role in mineral homeostasis. Repression due to C limitation was also shown to be correlated with the induction by sucrose (Dugas and Bartel, 2008). Furthermore, soybean roots, miR159a has been found to be up regulated during P deficiency, whereas down regulation of miR319a, miR396a, miR398b, miR1507a has been shown. The differential regulation of miRNAs during P-starvation depends on the frequency of phosphorus responsive motifs (P responsive motifs) in the promoter of the miRNA genes. The number of P-responsive motifs in the cis-acting region of miRNA genes is reported to be higher than those of non-responding miRNA genes (Zeng et al., 2010).
miRNAs in Nitrogen Uptake and Transport
Nitrogen, an essential constituent for nucleic acids, protein, chlorophyll, etc., plays a crucial role in plant growth and development. N transport depends on the external acquisition of N by roots from the soil, except the mechanism of biological N fixation in legumes. Plants can adapt in N-limiting soil conditions by up- or down-regulating a specific group of exporter or importer proteins. The differential regulation of transporters is selectively controlled by several small miRNA families. In Arabidopsis, depending on the occurrence at N-starvation, the miRNA-responsive populations can be categorized into two groups by Solexa high-throughput sequencing. N-starvation-induced (NSI) miRNA families include several members, such as miR156, miR169, miR171, miR160, miR319, miR826, miR829, miR839, and miR846, whereas miR167, miR172, miR399, miR395, miR850, miR857, miR863, and miR827 are recognized as N-starvation-suppressed (NSS) miRNA group members (Liang et al., 2012; Table 1). Furthermore, 15 and 14 miRNA families have been identified to be responsive in N-limiting conditions in rice and maize, respectively (Xu et al., 2011; Nischal et al., 2012). The miR156 family in Arabidopsis has been found at the highest abundance, and miR156h is thought to be the most important among the three members of the miR156 family. On the contrary, miR172 is negatively regulated by miR156 and inhibits the reproductive phase by prolonging the juvenile period. In N-starvation, the induction of miRNA160 inhibits lateral root development, whereas miR170 hastens the growth of primary roots by targeting auxin response factor (ARF16/17) and SCL6 regulatory proteins, respectively (Liang et al., 2012). In contrast, the perturbation of miR167 biogenesis in N-limiting condition attenuates the expression of ARF6/8, which in turn facilitates the development of lateral and adventitious roots (Jones-Rhoades and Bartel, 2004; Gifford et al., 2008).
In Medicago truncatula, miR169 and miR172 play a pivotal role in nodule development by regulating the expression of the HAP2 and AP2 genes. The lower abundances of miR169 during N-limitation upregulates HAP2 gene expression and the subsequent differentiation of nodule primordial by maintaining low N in the roots (Pant et al., 2009). However, different members of the miR169 family, such as miR169a, miR169bc, miR169d-g, and miR169h-n, have been reported in different plant species of Arabidopsis, soybean, and maize, etc. They are actively associated with the up-regulation of nitrate transporters during N-starvation (Xu et al., 2011; Zhao et al., 2011, 2012, 2013b; Liang et al., 2012; Wang et al., 2013b). Under low N availability, the expression of miR169d-g has been shown to be increased. miR169d-g exhibits a similar pattern of expression during P and sulfur (S) deficiencies, whereas other members are expressed differently (Liang et al., 2012). Very recently, miR172 was found to be exclusively expressed in the nodules of soybeans, targeting Arabidopsis homologous gene APETALA2-related transcription factors in response to P-starvation (Yan et al., 2013).
miRNAs in Sulfur Homeostasis
Sulfur (S), an indispensable inorganic mineral, is mainly taken up by roots in the form of sulfate from the soil. The S is assimilated into cysteine, methionine, glutathione, glucosinolate compounds, and various Fe–S proteins, cofactors, and lipoic acids, which are associated with both primary and secondary metabolism during stress (Rausch and Wachter, 2005). In Arabidopsis, sulfate is transported through xylem or phloem via cell-specific transporters such as sulfate transporters1;1 (SULTR1;1), SULTR2;1, and SULTR2;2. The expression of the transporters is predominantly regulated by miR395 depending on S starvation or abundance (Kawashima et al., 2009). Intriguingly, sulfate limitation induces the expression of miR395 and its low affinity sulfate transporter SULTR2;1, in contrast to the inhibitory effect of sulfate deficiency. SULTR2;1 is primarily confined to the xylem parenchyma, whereas miR395 is highly abundant in the phloem parenchyma and plays crucial role in sulfate remobilization between leaves during sulfate deficiency (Liang and Yu, 2010). The restriction of SULTR2;1 expression by miR395 in the xylem parenchyma facilitates the translocation of sulfate ions from the roots to the shoots. In addition, S deficiency leads to the elevated synthesis of SULFUR LIMITATION1 (SLIM1) protein in the roots, which in turn activates various sulfate transporters to enhance S uptake (Liang et al., 2010; Kawashima et al., 2011). The role of miR395 has also been elucidated in S assimilation by suppressing the expression of ATP sulfurylase genes, such as APS1, APS3, and APS4, which catalyze the first step of S assimilation (Matthewman et al., 2012). The expression levels of miR156, miR160, miR164, miR167, miR168, and miR394 are also modulated by S deprivation, as observed in Brassica napus (Huang et al., 2010).
miRNAs in Copper Homeostasis
Copper (Cu) is an essential micronutrient that serves primarily as a cofactor of metabolic enzymes and protein complexes in the electron transport chain. It is an integral member of plastocyanin, which actively participates in the electron transport of chloroplast grana during photosynthesis. Cu plays an important role against oxidative stress responses by acting as a cofactor of Copper/Zinc superoxide dismutase (CSD). During Cu limitation, the induction of miR398 down regulates CSD1, CSD2, and Cu chaperones for superoxide dismutase SOD1 (CCS1) gene expression (Sunkar et al., 2006; Beauclair et al., 2010). CCS1 is a chaperone protein that delivers the Cu ions to CSD1 and CSD2 apoprotein. Under Cu-deficient conditions, Cu/Zn superoxide function is replaced by iron (Fe) superoxide dismutase due to the low availability of Cu. The subunit of cytochrome C oxidase, the inner membrane protein of mitochondria encoded by CYCLO MONOOXYGENASE is also repressed by miR398 expression in Arabidopsis (Abdel-Ghany and Pilon, 2008). Other families of miRNAs, such as miR397, miR408, and miR857, have been found to be up regulated during Cu starvation, which in turn suppresses the expression of laccase and plastocyanin genes. The three laccase genes (LAC), such as LAC3, LAC12, and LAC13, are down regulated by miR408, and the mRNA of LAC2, LAC4, and LAC17 are degraded by miR397. miR857 is primarily responsible for targeting LAC7 transcripts. Notably, miRNAs related to Cu homeostasis facilitate plastocyanin biosynthesis by reducing the biosynthesis of non-essential Cu enzymes, thus ensuring Cu homeostasis by altering Cu availability among various groups of proteins (Gifford et al., 2008).
miRNA in Other Mineral Homeostasis
Manganese (Mn), Fe, and Zinc (Zn) are essential minerals for plant growth and nutrition. Several miRNAs have been found to be up regulated during Mn starvation and are also associated with other mineral stresses. In Phaseolus vulgaris, miR319, miR169, miR396, miR170, miR164, miR390, miR395, miR166, miR172, miR157, miR156, and miR167 are up regulated during Mn toxicity and attenuate the expression of a wide group of genes, including various transcription factors such as TEOSINTE-LIKE1, CYCLOIDEA, PROLIFERATING CELL FACTOR1 (TCP), HAPLESS (HAP2), SCARECROW-LIKE, NO APICAL MERISTEM (NAC), Arabidopsis transcription activation factor, CUP SHAPED COTYLEDON, serine threonine protein kinase, and APETALA2, etc. (Valdes-Lopez et al., 2010).
Iron and Zn are indispensable micronutrients for plants, as they are the major cofactors for several key metabolic enzymes, including Fe-S cluster proteins and ferredoxin molecules (Couturier et al., 2013; Forieri et al., 2013). Furthermore, the inadequate amounts of Fe and Zn that are stored in the edible parts of cereal grains play immense roles in human nutrition. The bioavailability of minerals in grains is directly associated with the uptake, transport, and loading of mineral ions (Aung et al., 2013). Recently, the miRNA-mediated regulation of Fe-related transporters and storage proteins were elucidated in Arabidopsis. miR398, one of the key activators for CSD gene expression during Cu deficiency, is also regulated by Fe deficiency but in an opposite manner. Fe deficiency reduces the expression of miR397, miR398a, miR398b, miR398c, miR398s, miR399, miR408, and miR2111, whereas Cu deficiency increases their expression and in turn regulates the expression of CSD1 and CSD2 (Buhtz et al., 2010; Waters et al., 2012). Therefore, the Cu-Fe interrelationship is another novel finding regarding the study of gene expression during Fe homeostasis. In addition, eight miRNAs from five families, including miR159, miR164, miR172, miR173, and miR394, were previously identified as Fe-responsive families from the small RNA library population in Arabidopsis. Intriguingly, the Fe deficiency responsive cis-acting elements1 and 2 (IDE1/IDE2) were found within the promoters of twenty-four miRNA genes in Arabidopsis and resemble the Fe-responsive gene families that are regulated during Fe deficiencies (Kong and Yang, 2010). However, the roles of other miRNAs in Fe transport and storage have not been clearly established.
The Zn deficiency in Sorghum bicolor aggravates the upregulation of several miRNA families, such as miR166, miR171, miR172, miR398, miR399, and miR319, which in turn target many gene family members including transporters (Li et al., 2013). Interestingly, two miRNA family members were found to be involved in the regulation of the expression of the CSD gene family but in an opposite manner. The upregulation of miR398 reduces the gene expression of CSD in the roots, whereas attenuated miR528 elevate the level of CSD transcripts in the seeds and the leaves. Furthermore, miRNAs maintain nutrient homeostasis by possessing an endogenous signal for the transport of micro and macronutrients (Liu et al., 2009; Marín-González and Suárez-López, 2012; Kehr, 2013). The up- or downregulation of diverse miRNAs during other metal stresses, such as Al, Cd, Hg, and Cd stress, have also been reported and likely play a role in the adaptive mechanisms of plants by regulating the expression of various stress-related genes (Zeng et al., 2012; Zhou et al., 2012; Zhang et al., 2013).
miRNAs in Systemic Mobility and Long-Distance Transport
After uptake by the roots, nutrients are allocated to the various storage parts of plants following long-distance transport. Xylem and phloem contribute to long-distance transport, and phloem-mediated communication plays an important role during nutrient stress. Phloem not only preserves the source-sink relationship but it also ensures cell-to-cell signal communication during different biotic and abiotic stress responses. Increasing evidence suggests that phloem-specific mRNAs coupled with small RNAs act as signaling molecules in different physiological responses, including nutrient transport under low nutrient conditions in several plant species (Varkonyi-Gasic et al., 2010; Kehr, 2013). Recently, microarrays of B. napus have revealed the presence of a specific set of phloem sap-specific miRNAs that are accumulated during S and Cu deficiency and are distinct in the roots, leaves, and inflorescence axis (Buhtz et al., 2010). miR395, which is known as sulfur deficiency responsive-miRNA, accumulates in phloem sap with miR399 and miR2111. miR399 and miR2111, the phosphate starvation responsive miRNAs, have also been found in phloem sap under Cu-deficient conditions (Abdel-Ghany and Pilon, 2008; Pant et al., 2008; Buhtz et al., 2010). miR399d, the member of miRNA399 family, exhibits long-distance transport from the shoots to the roots via phloem, conjugated with small RNA binding proteins, exemplified by Cucurbita maxima phloem small RNA binding protein 1 (CmPSRP1) and C. maxima phloem protein 16 (CmPP16; Pant et al., 2009). Furthermore, grafting experiments in the Arabidopsis mutant, hen-1-1, corroborates the mobility of miR399 and miR395 from the shoots to the roots via phloem, thus transmitting signals during nutrient deficiency. Interestingly, the translocation of miR395 was found to down-regulate only APS4 but not APS1 or AtSULTR2;1. On the other hand, miR158 in phloem sap appears to play an important role in nutrient transport by targeting lipase and xyloglucan fucosyltransferase genes during Fe deficiency, whereas miR172 was found to play an essential role in tuber formation and is considered as a phloem-specific signaling intermediate for plant growth and development (Martin et al., 2009; Kasai et al., 2010).
Conserved Groups of miRNAs in Nutrient Homeostasis
microRNAs play an crucial role in nutrient homeostasis by altering gene expression in plants. A single miRNA family has been reported to take part in different nutrient homeostasis conditions, thus playing orchestrated roles as signaling intermediates in several metabolic pathways. For example, two families of miRNAs, the miR169 and miR172 families are exclusively found in nodules and are involved in N, P, and Mn stresses. In addition, elevated levels of miR167 and miR395 during N and S starvation have been reported. These findings can be attributed to the fact that some common transcription factors activated by N and S-stress responses are responsible for the biosynthesis of these two miRNAs. The up-regulation of miR319 and miR396 during N and Mn starvation also supports the hypothesis (Valdes-Lopez et al., 2010). In this review, based on the abundances during various nutritional stresses, many families of miRNAs can be categorized into four conserved groups (Table 2). In different plant species, miR164, miR172, miR398, and miR399 are involved in the homeostasis of five different nutrients, thus representing the highest conserved group. miR156, miR167, miR395, miR319, miR408, and miR2111 are classified as highly conserved depending on their up-regulation during the four types of nutrient stress responses. miR160, miR168, miR166, miR397, and miR857 are categorized as moderately conserved (abundance frequencies three times), and miR158, miR159, miR169, miR170, miR171, miR528, miR390, miR396, miR394, and miR827 belong to the least conserved group (twice abundance frequencies). A particular miRNA can regulate different nutritional homeostasis conditions by up- or down-regulating the expression of various target genes (Table 2), which suggests a common signaling role of miRNAs in the regulation of diverse nutritional stress responses. Other unique miRNAs also have been identified in some plant species during particular nutrient stress responses, and their roles in the homeostasis of other nutrients should be investigated in the future.
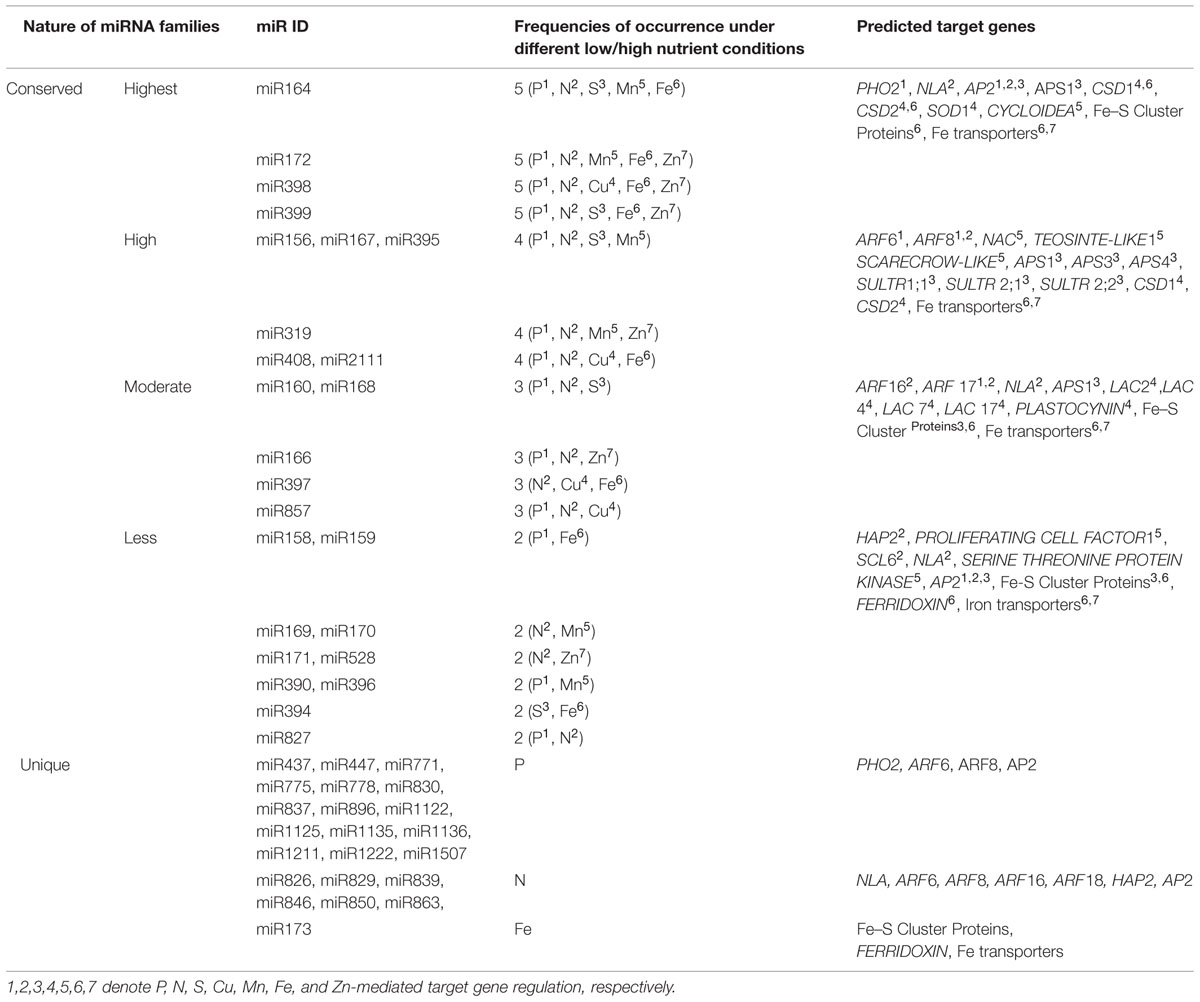
TABLE 2. Categorization of “conserved” and “unique” miRNAs under different nutrient stress and their predicted target genes (1,2,3,4,5,6,7 denote P, N, S, Cu, Mn, Fe, and Zn-mediated target gene regulation, respectively).
Concluding Remarks and Future Perspectives
Major research endeavors have focused on the genetic regulation of P and N transporters under the respective nutrient stress conditions. Future approaches for miRNA-mediated regulation of nutrient transporters and other metabolic enzymes and their implementation in future biotechnological research are summarized in Table 3. Because phytate is an important source of inorganic phosphorus, the role of miRNAs in phosphate metabolism, including inositol phosphate or phytic acid biosynthesis is a promising arena for future research. Nitrate metabolism and the N to C ratio determine the biomass of cereals. The roles of miRNA in the regulation of nitrate transporters and metabolism, including several enzymes such as aspartate amino transferase, glutamine synthase, and glutamate dehydrogenase, should be more extensively investigated. Regarding metal homeostasis, future investigations on differentially expressed-miRNAs and their regulatory roles in various Fe and Zn transporters may aid in the development of a novel platform for Fe and Zn loading in cereal grains. Fe and Zn, two important dietary nutrients, are found only in small amount in the consumable parts of cereal grains. To increase the content of these metals in milled grain, various biotechnological strategies have been utilized (Paul et al., 2013; Wang et al., 2013a; Borrill et al., 2014; Khan et al., 2014). However, the role of miRNAs in regulating specific transporters or transcription factors in Fe nutrition has not been studied extensively to date. The Fe-related gene regulation mechanism is important for understanding Fe nutrition and may elucidate the clear scenario of gene regulation during nutrient homeostasis.
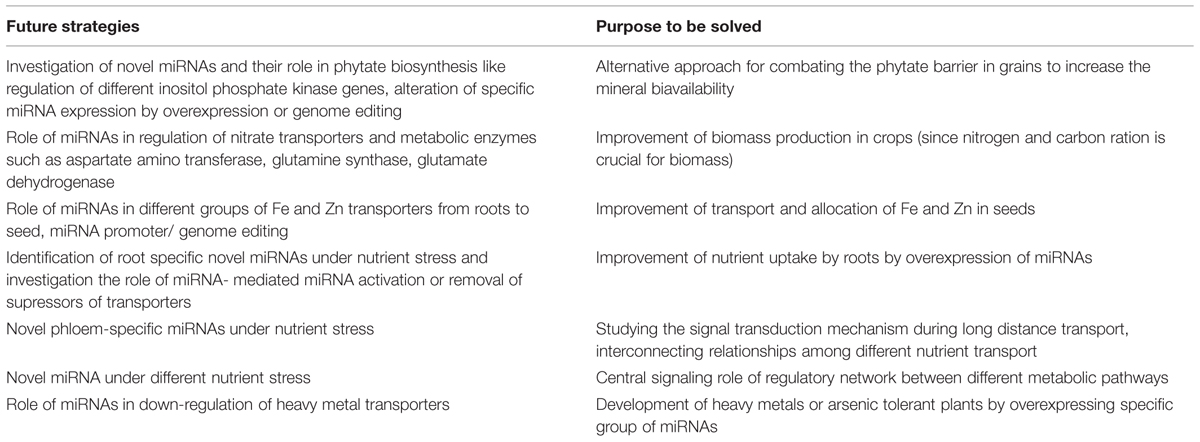
TABLE 3. Some probable future strategies for improvement of plant nutrition associated with miRNAs research.
microRNAs-mediated signal transduction during low/high nutrient stress is a fascinating topic of plant nutrition research. The alteration of nutrient levels in soil can trigger specific signaling molecules that act as repressors of target nutrient responsive-miRNAs. The decreased accumulation of miRNAs subsequently stabilizes the expression of transporters (Figure 1). On contrary, the optimal conditions or higher amounts of nutrients can trigger a specific group of miRNAs/small RNAs that directly affect the transporter (as exemplified by the phosphate transporter) or induce other miRNAs that suppress the expression of repressor genes. Therefore, the differential expression of miRNAs and their regulation under nutrient stress provide valuable information. The discovery of phloem-specific novel miRNAs during nutrient starvation and their cell-to-cell transmission will lead to a better understanding of the interrelationship among different nutrients. The identification of promoter regions of specific up- or down-regulated miRNAs that are responsive to micronutrient stresses and the subsequent development of knock-out mutants by inducing mutation in cis-acting elements using targeted genome-editing technologies, such as transcription activator-like effector nuclease (TALEN) or clustered regularly interspaced short palindromic repeats -CRISPR-associated 9 (CRISPR-Cas9) techniques, may lead to essential crop-improvement strategies in the future.
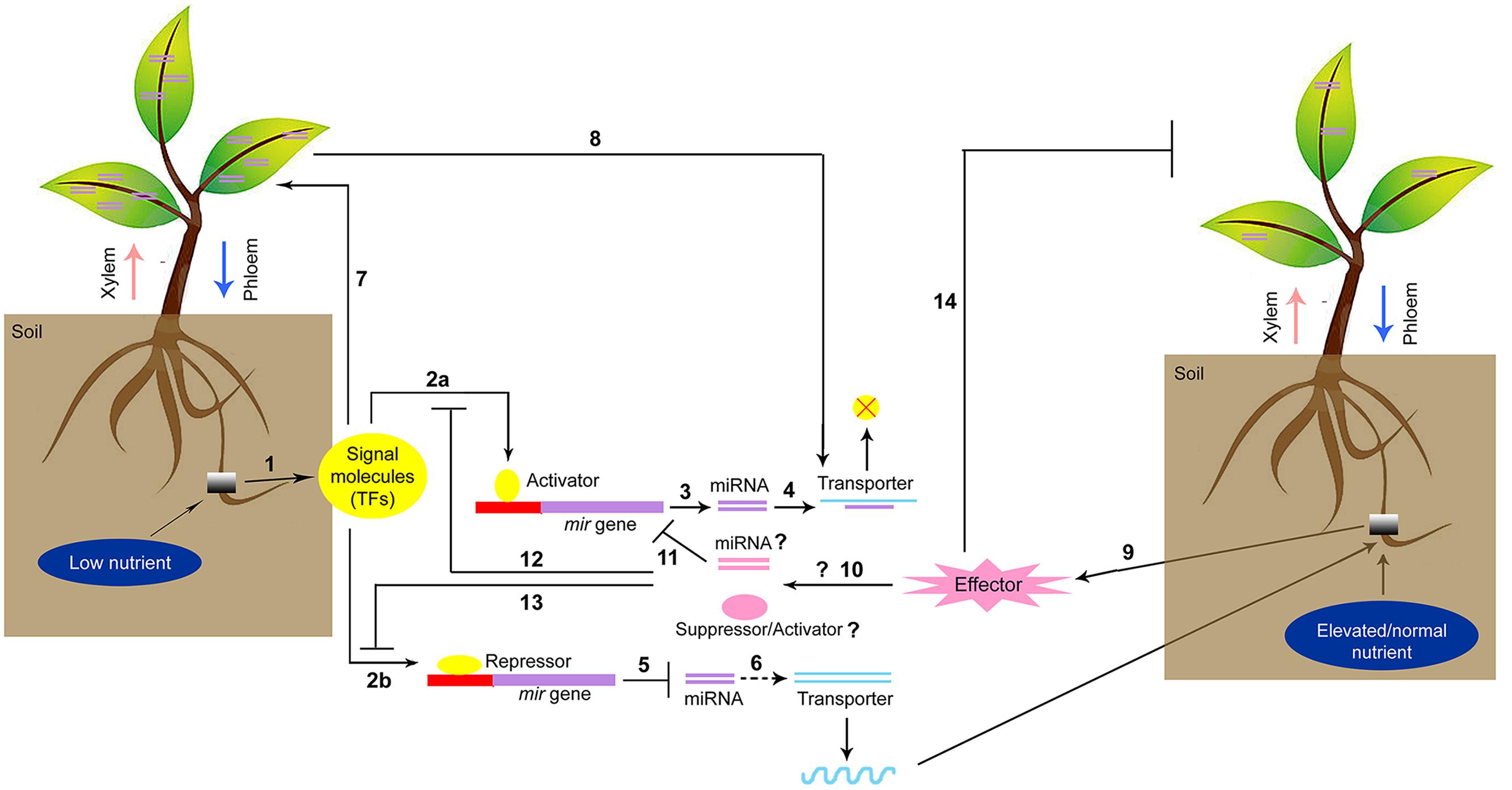
FIGURE 1. Proposed model of miRNA-mediated signaling network for regulation of nutrient homeostasis. (1) Low nutrient in soil upregulates signaling molecules which can act as an activator of nutrient stress responsive miRNA biosynthesis following interaction with mir promoter (2a) or repress the miRNA biosynthesis (2b) Activation of miRNA (3) attenuates the transporter gene expression (4) Reduction of miRNA biosynthesis (5) further stabilizes transporter gene expression (6) and more nutrients are transported. Signal molecules can induce phloem specific small or miRNA biosynthesis (7) which in turn regulates the mobility of nutrients through phloem by activating the transporters (8) Elevated or ambient condition of nutrients can induce other signal molecules which act as an effector (9) and facilitate the biosynthesis of other groups of miRNA or small RNAs (10) that interact with low nutrient stress responsive-miRNA in a opposite way (11) stimulating the specific group of transporters. The effector molecules may act as suppressor against the low nutrient stress responsive-activator (12) or inducer to mitigate the activity of low nutrient stress responsive-activator (13) Effector molecules can also reduce the biosynthesis of phloem specific miRNA or small RNA families and the phloem mediated transport is regulated.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We record our special thanks to the Department of Biotechnology (DBT) of Government of India and University Grants Commission (UGC) for financial support for different research projects in the laboratories.
References
Abdel-Ghany, S. E., and Pilon, M. (2008). MicroRNA-mediated systemic down-regulation of copper protein expression in response to low copper availability in Arabidopsis. J. Biol. Chem. 283, 15932–15945. doi: 10.1074/jbc.M801406200
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Abel, S. (2011). Phosphate sensing in root development. Curr. Opin. Plant Biol. 14, 303–309. doi: 10.1016/j.pbi.2011.04.007
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Aung, M. S., Masuda, H., Kobayashi, T., Nakanishi, H., Yamakawa, T., and Nishizawa, N. K. (2013). Iron biofortification of Myanmar rice. Front. Plant Sci. 4:158. doi: 10.3389/fpls.2013.00158
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Baek, D., Kim, M. C., Chun, H. J., Kang, S., Park, H. C., Shin, G., et al. (2013). Regulation of miR399f transcription by AtMYB2 affects phosphate starvation responses in Arabidopsis. Plant Physiol. 161, 362–373. doi: 10.1104/pp.112.205922
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bari, R., Pant, B. D., Stitt, M., and Scheible, W. R. (2006). PHO2, microRNA399, and PHR1 define a phosphate-signaling pathway in plants. Plant Physiol. 141, 988–999. doi: 10.1104/pp.106.079707
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bartel, D. P. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297. doi: 10.1016/S0092-8674(04)00045-5
Beauclair, L., Yu, A., and Bouche, N. (2010). microRNA directed cleavage and translational repression of the copper chaperone for superoxide dismutase mRNA in Arabidopsis. Plant J. 62, 454–462. doi: 10.1111/j.1365-313X.2010.04162.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Borrill, P., Connorton, J. M., Balk, J., Miller, A. J., Sanders, D., and Uauy, C. (2014). Biofortification of wheat grain with iron and zinc: integrating novel genomic resources and knowledge from model crops. Front. Plant Sci. 5:53. doi: 10.3389/fpls.2014.00053
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Buhtz, A., Pieritz, J., Springer, F., and Kehr, J. (2010). Phloem small RNAs, nutrient stress responses, and systemic mobility. BMC Plant Biol. 10:64. doi: 10.1186/1471-2229-10-64
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Chen, X. (2004). A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022–2025. doi: 10.1126/science.1088060
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Couturier, J., Touraine, B., Briat, J.-F., Gaymard, F., and Rouhier, N. (2013). The iron-sulfur cluster assembly machineries in plants: current knowledge and open questions. Front. Plant Sci. 4:259. doi: 10.3389/fpls.2013.00259
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dugas, D. V., and Bartel, B. (2008). Sucrose induction of Arabidopsis miR398 represses two Cu/Zn superoxide dismutases. Plant Mol. Biol. 67, 403–417. doi: 10.1007/s11103-008-9329-1
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fischer, J. J., Beatty, P. H., Good, A. G., and Muench, D. G. (2013). Manipulation of microRNA expression to improve nitrogen use efficiency. Plant Sci. 210, 70–81. doi: 10.1016/j.plantsci.2013.05.009
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Forieri, I., Wirtz, M., and Hell, R. (2013). Toward new perspectives on the interaction of iron and sulfur metabolism in plants. Front. Plant Sci. 4:357. doi: 10.3389/fpls.2013.00357
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Franco-Zorrilla, J. M., Valli, A., Todesco, M., Mateos, I., Puga, M. I., Rubio-Somoza, I., et al. (2007). Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 39, 1033–1037. doi: 10.1038/ng2079
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gifford, M. L., Dean, A., Gutierrez, R. A., Coruzzi, G. M., and Birnbaum, K. D. (2008). Cell-specific nitrogen responses mediate developmental plasticity. Proc. Natl. Acad. Sci. U.S.A. 105, 803–808. doi: 10.1073/pnas.0709559105
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gu, M., Xu, K., Chen, A., Zhu, Y., Tang, G., Xu, G., et al. (2010). Expression analysis suggests potential roles of microRNAs for phosphate and arbuscular mycorrhizal signaling in Solanum lycopersicum. Physiol. Plant. 138, 226–237. doi: 10.1111/j.1399-3054.2009.01320
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hackenberg, M., Huang, P. J., Huang, C. Y., Shi, B. J., Gustafson, P., and Langridge, P. A. (2013). Comprehensive expression profile of microRNAs and other classes of non-coding small RNAs in barley under phosphorous-deficient and -sufficient conditions. DNA Res. 20, 109–125. doi: 10.1093/dnares/dss037
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hu, B., Zhu, C., Li, F., Tang, J., Wang, Y., Lin, A., et al. (2011). LEAF TIP NECROSIS1 plays a pivotal role in the regulation of multiple phosphate starvation responses in rice. Plant Physiol. 156, 1101–1115. doi: 10.1104/pp.110.170209
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Huang, C. Y., Shirley, N., Genc, Y., Shi, B., and Langridge, P. (2011). Phosphate utilization efficiency correlates with expression of low-affinity phosphate transporters and noncoding RNA, IPS1, in barley. Plant Physiol. 156, 1217–1229. doi: 10.1104/pp.111.178459
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Huang, S. Q., Xiang, A. L., Che, L. L., Chen, S., Li, H., Song, J. B., et al. (2010). A set of miRNAs Brassica napus in response to sulphate deficiency and cadmium stress. Plant Biotechol. J. 8, 887–899. doi: 10.1111/j.1467-7652.2010.00517
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jamalkandi, S. A., and Masoudi-Nejad, A. (2009). Reconstruction of Arabidopsis thaliana fully integrated small RNA pathway. Funct. Integr. Genomics 9, 419–432. doi: 10.1007/s10142-009-0141-z
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jones-Rhoades, M. W., and Bartel, D. P. (2004). Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol. Cell. 14, 787–99. doi: 10.1016/j.molcel.2004.05.027
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jung, J. K. H., and McCouch, S. (2013). Getting to the roots of it: genetic and hormonal control of root architecture. Front. Plant Sci. 4:186. doi: 10.3389/fpls.2013.00186
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kant, S., Peng, M., and Rothstein, S. J. (2011). Genetic regulation by NLA and microRNA827 for maintaining nitrate-dependent phosphate homeostasis in Arabidopsis. PLoS Genet. 7:e1002021. doi: 10.1371/journal.pgen.1002021
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kasai, A., Kanehira, A., and Harada, T. (2010). miR172 can move longdistances in Nicotiana benthamiana. Open Plant Sci. J. 4, 1–6.
Kawashima, C. G., Matthewman, C. A., Huang, S., Lee, B. R., Yoshimoto, N., Koprivova, A., et al. (2011). Interplay of SLIM1 and miR395 in the regulation of sulfate assimilation Arabidopsis. Plant J. 66, 863–876. doi: 10.1111/j.1365-313X.2011.04547.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kawashima, C. G., Yoshimoto, N., Maruyama-Nakashita, A., Tsuchiya, Y. N., Saito, K., and Takahashi, H. (2009). Sulphur starvation induces the expression of microRNA-395 and one of its target genes but in different cell types. Plant J. 57, 313–321. doi: 10.1111/j.1365-313X.2008.03690.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kehr, J. (2013). Systemic regulation of mineral homeostasis by micro RNAs. Front. Plant Sci. 4:145. doi: 10.3389/fpls.2013.00145
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Khan, M. A., Castro-Guerrero, N., and Mendoza-Cozatl, D. G. (2014). Moving toward a precise nutrition: preferential loading of seeds with essential nutrients over non-essential toxic elements. Front. Plant Sci. 5:51. doi: 10.3389/fpls.2014.00051
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Khraiwesh, B., Zhu, J. K., and Zhu, J. (2012). Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim. Biophys. Acta 1819, 137–148. doi: 10.1016/j.bbagrm.2011.05.001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kidner, C. A., and Martienssen, R. A. (2005). The developmental role of microRNA in plants. Curr. Opin. Plant Biol. 8, 38–44. doi: 10.1016/j.pbi.2004.11.008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kong, W. W., and Yang, Z. M. (2010). Identification of iron-deficiency responsive microRNA genes and cis-elements in Arabidopsis. Plant Physiol. Biochem. 48, 153–159. doi: 10.1016/j.plaphy.2009.12.008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kruszka, K., Pieczynski, M., Windels, D., Bielewicz, D., Jarmolowski, A., Kulinska-Szweykowska, Z., et al. (2012). Role of micro RNAs and other sRNAs of plants in their changing environments. J. Plant Physiol. 169, 1664–1672. doi: 10.1016/j.jplph.2012.03.009
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, Y., Kim, M., Han, J., Yeom, K. H., Lee, S., Baek, S. H., et al. (2004). MicroRNA genes are transcribed by polymerase II. EMBO J. 23, 4051–4060. doi: 10.1038/sj.emboj.7600385
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Li, Y., Zhang, Y., Shi, D., Liu, X., Qin, J., Ge, Q., et al. (2013). Spatial-temporal analysis of zinc homeostasis reveals the response mechanisms to acute zinc deficiency in Sorghum bicolor. New Phytol. 200, 1102–1115. doi: 10.1111/nph.12434
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Liang, G., He, H., and Yu, D. (2012). Identification of nitrogen starvation-responsive miRNAs in Arabidopsis thaliana. PLoS ONE 7:e48951. doi: 10.1371/journal.pone.0048951
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Liang, G., Yang, F., and Yu, D. (2010). MicroRNA395 mediates regulation of sulfate accumulation and allocation in Arabidopsis thaliana. Plant J. 62, 1046–1057. doi: 10.1111/j.1365-313X.2010.04216.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Liang, G., and Yu, D. (2010). Reciprocal regulation among and miR395 posttranscriptionally APS and SULTR2; 1 Arabidopsis thaliana. Plant Signal. Behav. 5, 1257–1259. doi: 10.1111/j.1365-313X
Liu, T. Y., Chang, C. Y., and Chiou, T. J. (2009). The long-distance signaling of mineral macronutrients. Curr. Opin. Plant Biol. 12, 312–319. doi: 10.1016/j.pbi.2009.04.004
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lu, S., Sun, Y. H., and Chiang, V. L. (2008). Stress-responsive microRNAs in Populus. Plant J. 55, 131–151. doi: 10.1111/j.1365-313X.2008.03497.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lundmark, M., Korner, C. J., and Nielsen, T. H. (2010). Global analysis of microRNA in Arabidopsis in response to phosphate starvation as studied by locked nucleic acid-based microarrays. Physiol. Plant. 140, 57–68. doi: 10.1111/j.1399-3054.2010.01384.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Marín-González, E., and Suárez-López, P. (2012). “And yet it moves”: cell-to-cell and long-distance signaling by plant microRNAs. Plant Sci. 196, 18–30. doi: 10.1016/j.plantsci.2012.07.009
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Martin, A., Adam, H., Diaz-Mendoza, M., Zurczak, M., Gonzalez-Schain, N., and Suarez-Lopez, P. (2009). Graft-transmissible induction of potato tuberization by the microRNA miR172. Development 136, 2873–2881.
Matthewman, C. A., Kawashima, C. G., Huska, D., Csorba, T., Dalmay, T., and Kopriva, S. (2012). miR395 is a general component of the sulfate assimilation regulatory network in Arabidopsis. FEBS Lett. 586, 3242–3248. doi: 10.1016/j.febslet.2012.06.044
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nischal, L., Mohsin, M., Khan, I., Kardam, H., Wadhwa, A., Abrol, Y. P., et al. (2012). Identification and comparative analysis of microRNAs associated with low-N tolerance in rice genotypes. PLoS ONE 7:e50261. doi: 10.1371/journal.pone.0050261
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Pant, B. D., Buhtz, A., Kehr, J., and Scheible, W. R. (2008). MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J. 53, 731–738. doi: 10.1111/j.1365-313X.2007.03363.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Pant, B. D., Musialak-lange, M., Nuc, P., May, P., Buhtz, A., Kehr, J., et al. (2009). Identification of nutrient-responsive Arabidopsis and rapeseed microRNAs by comprehensive real-time polymerase chain reaction profiling and small RNA sequencing. Plant Physiol. 150, 1541–1555. doi: 10.1104/pp.109.139139
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Paul, S., Ali, N., Sarkar, S. N., Datta, S. K., and Datta, K. (2013). Loading and bioavailability of iron in cereal grains. Plant Cell Tiss. Organ Cult. 113, 363–373. doi: 10.1007/s11240-012-0286-7
Pei, L., Jin, Z., Li, K., Yin, H., Wang, J., and Yang, A. (2013). Identification and comparative analysis of low phosphate tolerance associated microRNAs in two maize genotypes. Plant Physiol. Biochem. 70, 221–234. doi: 10.1016/j.plaphy.2013.05.043
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Poirier, Y., and Bucher, M. (2002). “Phosphate transport and homeostasis in Arabidopsis,” in the Arabidopsis, eds Somerville R., and Meyerowitz E. M. (Rockvilli, MD: American Society of Plant Biologists), 1.
Rausch, T., and Wachter, A. (2005). Sulfur metabolism: a versatile platform for launching defence operations. Trends Plant Sci. 10, 503–509. doi: 10.1016/j.tplants.2005.08.006
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schauer, S. E., Jacobson, S. E., Meinke, D. W., and Ray, A. (2002). DICER-LIKE1: blind men and elephants in Arabidopsis development. Trends Plant Sci. 7, 487–491. doi: 10.1016/S1360-1385(02)02355-5
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sperotto, R. A., Ricachenevsky, F. K., Williams, L. E., Vasconcelos, M. W., and Menguer, P. K. (2014). From soil to seed: micronutrient movement into and within the plant. Front. Plant Sci. 5:438. doi: 10.3389/fpls.2014.00438
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sunkar, R., Kapoor, A., and Zhu, J. K. (2006). Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 18, 2051–2065. doi: 10.1105/tpc.106.041673
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sunkar, R., and Zhu, J. K. (2004). Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16, 2001–2019. 10.1105/tpc.104.022830
Valdes-Lopez, O., Yang, S. S., Aparicio-Fabre, R., Graham, P. H., Reyes, J. L., Vance, C. P., et al. (2010). MicroRNA expression profile in common bean (Phaseolus vulgaris) under nutrient deficiency stresses and manganese toxicity. New Phytol. 187, 805–818. doi: 10.1111/j.1469-8137.2010.03320.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Varkonyi-Gasic, E., Gould, N., Sandanayaka, M., Sutherland, P., and MacDiarmid, R. M. (2010). Characterisation of microRNAs from apple (Malus domestica ‘Royal Gala’) vascular tissue and phloem sap. BMC Plant Biol. 10:159. doi: 10.1186/1471-222910-159
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Vazquez, F., Legrand, S., and Windels, D. (2010). The biosynthetic pathways and biological scopes of plant small RNAs. Trends Plant Sci. 15, 337–345. doi: 10.1016/j.tplants.2010.04.001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Vigani, G., Morandini, P., and Murgia, I. (2013a). Searching iron sensors in plants by exploring the link among 2′-OG-dependent dioxygenases, the iron deficiency response and metabolic adjustments occurring under iron deficiency. Front. Plant Sci. 4:169. doi: 10.3389/fpls.2013.00169
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Vigani, G., Zochhi, G., Bashir, K., Phillipar, K., and Briat, J. F. (2013b). Cellular iron homeostasis and metabolism in plants. Front. Plant Sci. 4:490. doi: 10.3389/fpls.2013.00490
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, C., Huang, W., Ying, Y., Li, S., Secco, D., Tyerman, S., et al. (2012). Functional characterization of the rice SPXMFS family reveals a key role of OsSPX-MFS1 in controlling phosphate homeostasis in leaves. New Phytol. 196, 139–148. doi: 10.1111/j.1469-8137.2012.04227.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, M., Gruissem, W., and Bhullar, N. K. (2013a). Nicotianamine synthase overexpression positively modulates iron homeostasis-related genes in high iron rice. Front. Plant Sci. 4:156. doi: 10.3389/fpls.2013.00156
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, Y., Zhang, C., Hao, Q., Sha, A., Zhou, R., Zhou, X., et al. (2013b). Elucidation of miRNAs-mediated responses to low nitrogen stress by deep sequencing of two soybean genotypes. PLoS ONE 8:e67423. doi: 10.1371/journal.pone.0067423
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Waters, B. M., Mcinturf, S. A., and Stein, R. J. (2012). Rosette iron deficiency transcript and microRNA profiling reveals links between copper and iron homeostasis in Arabidopsis thaliana. J. Exp. Bot. 63, 5903–5918. doi: 10.1093/jxb/ers239
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Xu, F., Liu, Q., Chen, L., Kuang, J., Walk, T., Wang, J., et al. (2013). Genome-wide identification of soybean microRNAs andtheir targets reveals their organ-specificity and responses to phosphate starvation. BMC Genomics 14:66. doi: 10.1186/1471-2164-14-66
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Xu, Z., Zhong, S., Li, X., Li, W., Rothstein, S. J., Zhang, S., et al. (2011). Genome-wide identification of microRNAs in response to low nitrate availability in maize leaves and roots. PLoS ONE 6:e28009. doi: 10.1371/journal.pone.0028009
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Yan, Z., Hossain, M. S., Wang, J., Valdes-Lopez, O., Liang, Y., Libault, M., et al. (2013). miR172 regulates soybean nodulation. Mol. Plant Microbe Interact. 26, 1371–1377. doi: 10.1094/MPMI-04-13-0111-R
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zeng, H. Q., Zhu, Y. Y., Huang, S. Q., and Yang, Z. M. (2010). Analysis of phosphorus-deficient responsive miRNAs and cis elements from soybean (Glycine max L.). J. Plant Physiol. 167, 1289–1297. doi: 10.1016/j.jplph.2010.04.017
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zeng, Q. Y., Yang, C. Y., Ma, Q. B., Li, X. P., Dong, W. W., and Nian, H. (2012). Identification of wild soybean miRNAs and their target genes responsive to aluminum stress. BMC Plant Biol. 12:182. doi: 10.1186/1471-2229-12-182
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zhang, B., Pan, X., and Stellwag, E. J. (2008). Identification of soybean microRNAs and their targets. Planta 229, 161–182. doi: 10.1007/s00425-008-0818-x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zhang, L. W., Song, J. B., Shu, X. X., Zhang, Y., and Yang, Z. M. (2013). miR395 is involved in detoxification of cadmium in Brassica napus. J. Hazard Mater. 250–251, 204–211. doi: 10.1016/j.jhazmat.2013.01.053
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zhao, M., Ding, H., Zhu, J. K., Zhang, F., and Li, W. X. (2011). Involvement of miR169 in the nitrogen-starvation responses in Arabidopsis. New Phytol. 190, 906–915. doi: 10.1111/j.1469-8137.2011.03647.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zhao, M., Tai, H., Sun, S., Zhang, F., Xu, Y., and Li, W. X. (2012). Cloning and characterization of maize miRNAs involved in responses to nitrogen deficiency. PLoS ONE 7:e29669. doi: 10.1371/journal.pone.0029669
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zhao, X., Liu, X., Guo, C., Gu, J., and Xiao, K. (2013a). Identification and characterization of microRNAs from wheat (Triticum aestivum L.) under phosphorus deprivation. J. Plant Biochem. Biotechnol. 22, 113–123. doi: 10.1007/s13562-012-0117-2
Zhao, Y., Xu, Z., Mo, Q., Zou, C., Li, W., Xu, Y., et al. (2013b). Combined small RNA and degradome sequencing reveals novel miRNAs and their targets in response to low nitrate availability in maize. Ann. Bot. 112, 633–642. doi: 10.1093/aob/mct133
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zhou, Z. S., Song, J. B., and Yang, Z. M. (2012). Genome-wide identification of Brassica napus microRNA and their targets in response to cadmium. J. Exp. Bot. 63, 4597–4613. doi: 10.1093/jxb/ers136
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: miRNA, plant, micronutrient, macronutrient, transporters, nutrient homeostasis
Citation: Paul S, Datta SK and Datta K (2015) miRNA regulation of nutrient homeostasis in plants. Front. Plant Sci. 5:588. doi: 10.3389/fpls.2015.00232
Received: 10 December 2014; Accepted: 23 March 2015;
Published online: 10 April 2015.
Edited by:
Clelia De-la-Peña, Centro de Investigación Científica de Yucatán, MexicoReviewed by:
Stewart Gillmor, Centro de Investigación y de Estudios Avanzados del Instituto Politécnico Nacional, MexicoMaofeng Chai, The Samuel Roberts Noble Foundation, USA
Copyright © 2015 Paul, Datta and Datta. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Karabi Datta, Translational Research Laboratory of Transgenic Rice, Department of Botany, University of Calcutta, 35 Ballygunge Circular Road, Kolkata-700019, WB, Indiaa3JiZGF0dGFAeWFob28uY29t
 Soumitra Paul
Soumitra Paul Swapan K. Datta
Swapan K. Datta