- 1Department of Plant Pathology and Microbiology, Center for Plant Responses to Environmental Stresses, Iowa State University, Ames, IA, USA
- 2Bioinformatics and Computational Biology Graduate Program, Iowa State University, Ames, IA, USA
- 3Corn Insects and Crop Genetics Research Unit, U.S. Department of Agriculture-Agricultural Research Service, Iowa State University, Ames, IA, USA
- 4Department of Statistics, Iowa State University, Ames, IA, USA
Plants have evolved complex regulatory mechanisms to control a multi-layered defense response to microbial attack. Both temporal and spatial gene expression are tightly regulated in response to pathogen ingress, modulating both positive and negative control of defense. BLUFENSINs, small knottin-like peptides in barley, wheat, and rice, are highly induced by attack from fungal pathogens, in particular, the obligate biotrophic fungus, Blumeria graminis f. sp. hordei (Bgh), causal agent of barley powdery mildew. Previous research indicated that Blufensin1 (Bln1) functions as a negative regulator of basal defense mechanisms. In the current report, we show that BLN1 and BLN2 can both be secreted to the apoplast and Barley stripe mosaic virus (BSMV)-mediated overexpression of Bln2 increases susceptibility of barley to Bgh. Bimolecular fluorescence complementation (BiFC) assays signify that BLN1 and BLN2 can interact with each other, and with calmodulin. We then used BSMV-induced gene silencing to knock down Bln1, followed by Barley1 GeneChip transcriptome analysis, to identify additional host genes influenced by Bln1. Analysis of differential expression revealed a gene set enriched for those encoding proteins annotated to nuclear import and the secretory pathway, particularly Importin α1-b and Sec61 γ subunits. Further functional analysis of these two affected genes showed that when silenced, they also reduced susceptibility to Bgh. Taken together, we postulate that Bln1 is co-opted by Bgh to facilitate transport of disease-related host proteins or effectors, influencing the establishment of Bgh compatibility on its barley host.
Introduction
Obligate fungal biotrophs, i.e., pathogens that require their host to survive, are a major threat to crop production worldwide. To establish biotrophy, the fungus must penetrate cell walls, suppress defense, and establish haustoria for nutrient acquisition (Dodds et al., 2004; Micali et al., 2011; Mentlak et al., 2012). In general, these pathogens interfere with recognition at the host plasma membrane or secrete effector proteins, often through feeding structures termed haustoria, into the plant cell cytosol that alter resistance signaling or the downstream manifestation of resistance responses. Many cloned effectors are small proteins of unknown function containing a signal for secretion into the apoplast; how these effectors gain entry into host cells and contribute to pathogen colonization has been a major focus to understand the underlying mechanisms determining pathogenicity (Rovenich et al., 2014; Stotz et al., 2014).
The host responds with an integrated multi-layer defense system. Typically, pathogen-associated molecular patterns (PAMPs) trigger the initial activation of non-specific, innate immune responses, currently termed PAMP Triggered Immunity (PTI) (Macho and Zipfel, 2014). These include the transcription of thousands of stress-related genes, as well as production of antimicrobial metabolites and peptides during early stages of pathogen invasion. A second layer, designated Effector-Triggered Immunity (ETI) generally follows gene-for-gene interactions, in which specific resistance (R) proteins initiate a signal cascade when they recognize, either directly or indirectly, corresponding effectors delivered by the pathogen (Bent and Mackey, 2007; Jacob et al., 2013; Cesari et al., 2014).
Host factors that are activated and recruited by pathogen effectors interfere with different layers of the plant defense response. These plant factors are either called negative regulators of plant defense or susceptibilty factors, which are co-opted by the pathogen to optimize growth and parasitism; both are encoded by susceptibility (S) genes (Vogel et al., 2002, 2004; Hückelhoven et al., 2013; Lapin and Van Den Ackerveken, 2013; Van Schie and Takken, 2014). Mutation of an S gene has the potential to alter the plants susceptibility and lead to resistance, an important feature that is often used in breeding. For example, the cell wall has long been recognized as a major barrier against pathogen infection (Bellincampi et al., 2014; Malinovsky et al., 2014). PMR5 and PMR6 are two potential susceptibility factors identified in Arabidopsis. Mutations in pmr5 [defective in a gene encoding a predicted endoplasmic reticulum (ER) protein] and pmr6 (defective in a cell wall-degrading pectate lyase-like gene) genes both affect pectin composition of the cell wall, thus increasing Arabidopsis resistance to powdery mildew (Vogel et al., 2002, 2004). Other examples include xa5, encoding a subunit of transcription factor IIA (Iyer and McCouch, 2004; Jiang et al., 2006), and xa13, encoding a plasma membrane protein and essential for pollen development (Chu et al., 2006). Both loss-of-function mutants to bacterial blight have been used successfully in rice cultivation (Iyer-Pascuzzi and McCouch, 2007).
A classic case in barley is the well-characterized Mlo gene that encodes a transmembrane protein, which negatively regulates penetration resistance to powdery mildew (Büschges et al., 1997). Loss of function mlo mutants result in durable and broad-spectrum resistance, which has been wildly adapted for cultivation in Europe (Büschges et al., 1997; Panstruga, 2005; Acevedo-Garcia et al., 2014). MLO2, an Arabidopsis thaliana homolog of the barley S-gene Mlo, was found to be the target of the Pseudomonas syringae effector HopZ2 (Lewis et al., 2012).
BAX INHIBITOR-1 (BI-1) inhibits BAX-induced PCD in yeast and Arabidopsis; additionally, BI-1 modulates cell-wall-associated defense and contributes to establishing full compatibility of barley with the obligate biotrophic fungus, Blumeria graminis f. sp. hordei (Bgh), causal agent of powdery mildew disease (Eichmann et al., 2010). Interestingly, overexpression of BI-1 was found to negatively regulate penetration resistance mediated by mlo and almost restored the penetration efficiency (PE) of Bgh to wild-type levels (Hückelhoven et al., 2003), suggesting these genes have important roles in a complex interconnected network. The MLO protein in barley negatively regulates the actin-dependent resistance pathway, and the actin cytoskeleton is thought to contribute to the establishment of effective barriers at the cell periphery against fungal access (Miklis et al., 2007). The RAC/ROP family G-protein RACB, another potential host susceptibility factor, is also involved in the modulation of actin reorganization and cell polarity in the interaction of barley with Bgh (Opalski et al., 2005).
We previously reported the discovery of the monocot-specific Blufensin family of cysteine-rich, peptides, which negatively impact plant defense (Meng et al., 2009). The Bln1 and Bln2 transcripts are highly upregulated in response to infection by a wide array of fungal pathogens, including Blumeria, Puccinia, Cochliobolus, and Fusarium spp., as compared to uninfected control plants. The genes that encode these peptides are so far unique to the cereal grain crops barley, wheat, and rice, and the resulting proteins are similar to knottins, a diverse family of proteins characterized by a unique disulfide through disulfide knot (Gracy et al., 2008).
In the work described herein, we used BLN-GFP fusion constructs to demonstrate that BLN1 and BLN2 can be secreted into the apoplast. Bimolecular fluorescence complementation (BiFC) assays (Kerppola, 2006) suggest that BLN1 and BLN2 interact with calmodulin, as well as each other. Barley stripe mosaic virus (BSMV)-mediated Bln overexpression increased susceptibility of barley to Bgh. BSMV–Virus Induced Gene Silencing (VIGS) coupled with a Barley1 GeneChip transcriptome analysis, identified additional genes in the Blufensin1 (Bln1) network. These candidates appear to have key roles in R-gene mediated and innate immunity networks, thus, the functional identification of their precise roles will be a significant step in understanding plant defense.
Results
BLN1 and BLN2 can be Secreted into the Apoplast
In previous research, BSMV-VIGS of Bln1 decreased barley susceptibility to Bgh in compatible interactions. Likewise, single cell transient overexpression of Bln1 significantly increased accessibility toward virulent Bgh. Moreover, silencing of Bln1 in plants harboring the Mildew locus o (Mlo) susceptibility factor decreased accessibility to Bgh, suggesting BLN1 functions in parallel with or upstream of MLO to modulate penetration resistance (Meng et al., 2009).
Computational analysis of the BLN1 and BLN2 signal peptides (SP) predicted that BLN could be secreted into the apoplast, and thus, may act as ligands to generate a signal transduction cascade, influencing Bgh accessibility (Meng et al., 2009). To test this hypothesis, six different Bln-GFP fusion constructs were assembled for bombardment into onion epidermal cells, [BLN1 or 2 minus SP (35S:BLN1/2–SP), BLN1 or 2 plus SP (35S:BLN1/2 + SP), and BLN1 or 2 SP only (35S:BLN1/2 SP only)] (Figure 1A).
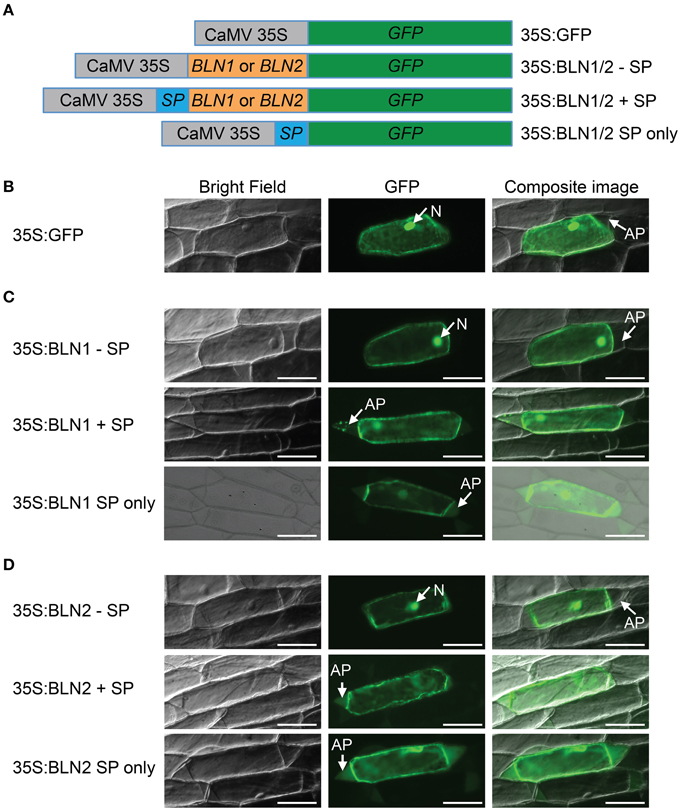
Figure 1. Subcellular localization of BLN1 and BLN2. (A) Schematic diagram of expression constructs. Gray boxes represent Cauliflower Mosaic Virus (CaMV) 35S promoter; the green, orange, and the blue boxes represent coding regions for GFP, mature BLN1/BLN2, and BLN1/BLN2 signal peptides, respectively. The CaMV 35S promoter was used to drive gene expression. The GFP coding sequence is fused to Bln1 and Bln2 without signal peptide-coding region (35S:BLN1/2—SP), to Bln1 and Bln2 with signal peptide-coding region (35S:BLN1/2 + SP), and to BLN1/2 signal peptide-coding region only (35S:BLN1/2 SP only). The construct harboring GFP coding sequence alone was used as a subcellular localization control. (B) Microscopic observation of GFP signal in onion epidermal cells after plasmolysis. GFP signal was observed in the cytoplasm and nucleus region in cells expressing GFP alone. No GFP signal was observed in apoplast region. (C,D) Microscopic observation of GFP signal in onion epidermal cells after plasmolysis. GFP signal was observed in apoplast when GFP was fused to full-length BLN1 or BLN2 with signal peptide (35S:BLN1 + SP, 35S:BLN2 + SP), as well as in cells expressing GFP fused to signal peptides from BLN1 and BLN2 (35S:BLN1 SP only, 35S:BLN2 SP only). By contrast, no GFP signal was observed in the apoplastic region in cells expressing GFP fused to BLN1 or BLN2 without signal peptides from BLN1 and BLN2 (35S:BLN1-SP, 35S:BLN2-SP). Left column: bright field images; middle column: fluorescence microscopic images of GFP; right column: composite images of the GFP and bright light images. AP, apoplast; N, Nucleus. Bar = 100 μm.
Because GFP is unstable at low pH, to visualize its expression in the apoplast, onion epidermal cells were treated with 20 mM Pipes-KOH (pH 7.0) to neutralize the pH according to Genovesi et al. (2008) (see Materials and Methods). The pH 7.0 medium neutralizes the normally acidic apoplast, facilitating the visualization of GFP-mediated fluorescence. As illustrated in Figures 1C,D (middle panel), GFP fluorescence was detected in the apoplast, cytoplasm and nuclei of plasmolysed cells when transformed with the full-length Bln1 or Bln2 ORFs fused with GFP. Similar results were obtained when constructs harboring GFP fused with coding sequences for signal peptides from BLN1 or BLN2 (Figures 1C,D, lower panel). By contrast, GFP fluorescence was found only in the cytoplasm or the nucleus when onion epidermal cells were bombarded with constructs absent the signal peptides (Figures 1C,D, upper panel), similar to the GFP-only control (Figure 1B); no visible fluorescent signal was observed in the apoplastic region. The above results indicate that the BLN1 and BLN2 signal peptides can direct protein secretion, and both BLN1 and BLN2 can be secreted from the cytoplasm into the apoplast.
BSMV-VOX: A New BSMV-mediated Overexpression System for Functional Analysis of Bln1 and Bln2
As described above, we developed a bombardment based BSMV-VIGS system for high-throughput silencing of candidate genes involved in interactions with the barley powdery mildew fungus (Meng et al., 2009). To complement these gene-silencing studies, we further developed BSMV as a transient overexpression system (BSMV-VOX) for functional analysis in both host and pathogen. To generate the expected cleavage products from the artificial fusion proteins, a 54-nucleotide sequence encoding the 18 amino-acid foot and mouth virus peptide (FMDV-2A) was inserted in front of the 5′ end of BSMV:γ ORF B. The ORF encoding GFP was inserted between the StuI and BamH1 sites before FMDV 2A as a visible marker to monitor overexpression (Figure 2A). These GFP and BSMV:γB coding regions are fused in-frame via the FMDV 2A coding sequence. The FMDV 2A peptide mediates the primary cis-“cleavage” of the FMDV polyprotein in a cascade of processing events that ultimately generate the mature FMDV proteins. Subsequently, FMDV 2A efficiently generates the expected cleavage products from the artificial fusion proteins in cells (Furler et al., 2001). The BSMV-mediated overexpression construct (pBSMV-OEx) was then co-bombarded with BSMV:α and BSMV:β to barley cultivar Black Hulless, which is susceptible to BSMV. Overexpression of GFP (pBSMV-OEx:GFP) was used to examine the efficacy of overexpression with this approach. Microscopic observation showed that all leaves with BSMV infected stripe and mosaic symptoms also exhibit green fluorescence as detected by UV microscopy (Figure 2B), signifying the robustness of BSMV-VOX system for transient gene overexpression. In addition, due to the systemic infection of BSMV, the BSMV-VOX system results in transient gene overexpression throughout BSMV-infected barley leaves (Lee et al., 2012), as compared to single-cell-overexpression in epidermal cells (Meng et al., 2009). Therefore, the infection phenotypes can be observed by the naked eye, hyphal growth and associated symptoms can be quantified digitally, and the target gene expression levels can be assayed by quantitative real-time reverse transcriptase PCR (qRT-PCR), in the absence of stable transformation.
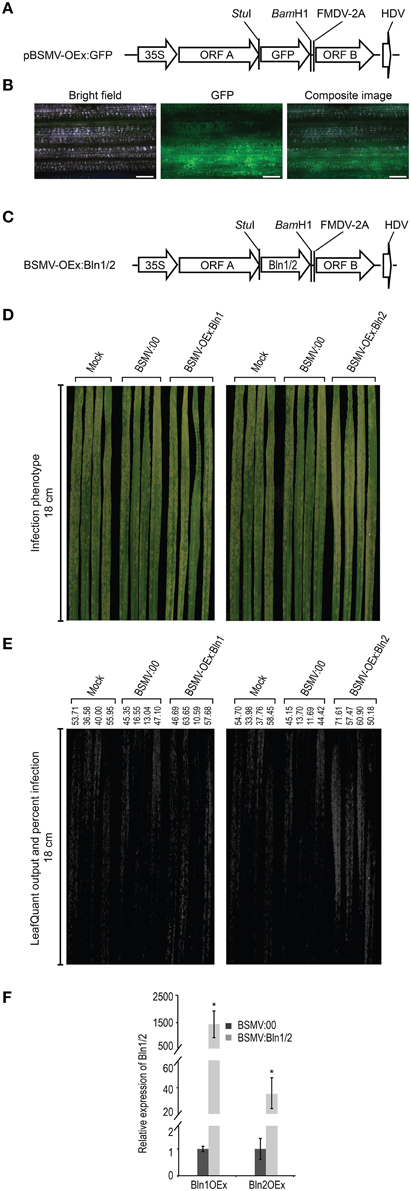
Figure 2. BSMV-mediated over-expression of Bln1/2. (A) Schematic diagram of BSMV-mediated GFP over-expression (VOX) construct. The coding sequence for the fusion protein of GFP and foot- and-mouth disease virus (FMDV)-2A self-cleavage peptide was digested by BglII and KpnI, ligated into the construct BSMV: γ, which harbors the BSMV γ-subgenome. Subsequent digestion with BglII and KpnI, resulted in BSMV-mediated GFP over-expression construct pBSMV-OEx:GFP. (B) Microscopic observation of GFP signal (middle column) in barley leaves (black hull-less) co-bombarded with BSMV-mediated GFP over-expression construct pBSMV-OEx:GFP, BSMV:α, and BSMV:β. The efficacy of the observed GFP signal is 100% in leaves that showed BSMV infection symptom. Bar = 100 μm. (C) Schematic diagram of BSMV-mediated BLN1/2 over-expression construct pBSMV-OEx:Bln1/2. PCR amplified fragments for Bln1/2 coding regions were digested by StuI and BamH1 and inserted into StuI and BamH1 digested pBSMV-OEx:GFP, generating constructs pBSMV-OEx:Bln1 and pBSMV-OEx:Bln2. (D) Phenotype of Bgh infected leaves treated with buffer (Mock), BSMV:00 control, and pBSMV-OEx:Bln1 (Bln1 over-expression) (left column); buffer (Mock), BSMV:00 control, and Bln2 over-expression (pBSMV-OEx:Bln2) (right column). (E) LeafQuant infection phenotype images and quantification of Bgh hyphal growth on leaves treated with buffer (mock), BSMV:00 control, and over-expression for Bln1 and Bln2 (See Table 1). (F) Quantitative RT-PCR analyses for Bln1 and Bln2 levels in leaves treated by BSMV-mediated over-expression Bln1OEx and BlnOEx. Bars represent standard error calculated from at least four independent plants for each treatment from two replicate experiments shown in this figure. The average Bln1 and Bln2 levels in BSMV:00 control were set to 1.00 (* designates that p < 0.05).
Overexpression of Bln2 Increases Susceptibility in Compatible Interactions
Our newly developed BSMV-VOX system (described above) was adopted to further corroborate the function of Bln genes in barley immunity to Bgh. Full-length Bln1-1 and Bln2 ORFs were substituted in place of GFP in the BSMV-VOX vector, pBSMV-OEx:GFP, to create the expression constructs BSMV-OEx:Bln1 and BSMV-OEx:Bln2 (Figure 2C). pBSMV-OEx:Bln1 or pBSMV-OEx:Bln2 plasmids were then co-bombarded with the BSMV:α and BSMV:β separately into 7 day old Black Hulless seedlings. After 7 days, sap from BSMV infected barley leaves was used to mechanically inoculate barley cultivar HOR11358 (Mla9). Twelve days after overexpression, plants were subsequently challenged with the virulent Bgh isolate 5874 (avra9). Control bombardments were performed with the BSMV:00 construct (see Materials and Methods). Systemic overexpression of Bln2 in whole barley leaves significantly increased susceptibility in compatible barley-Bgh interactions (p = 0.0194). Although one can observe a small increase in Bgh colony proliferation on Bln1-OEx barley leaves, this did not result in a significant difference in quantifiable growth (Figures 2D,E, Table 1). This contrasts with our previous result using transient single-cell-overexpression in barley epidermal cells (Meng et al., 2009), and may be due to the differential resistance of barley genotypes to BSMV (Hein et al., 2005), which could further influence the phenotypic effects of BSMV-mediated overexpression, as opposed to single cell bombardment assays (which contain no BSMV).
Transcript accumulation of Bln1 and Bln2 was assayed to monitor the level of gene overexpression. Third leaves of BSMV-treated plants were used for qRT-PCR assays at 24 HAI with Bgh. Barley Actin mRNA was used as an internal quantitative control for all samples. Results of qRT-PCR demonstrated the distinct induction of Bln1 and Bln2 transcripts in Bgh inoculated leaves that harbored overexpression constructs as compared to BSMV:00 inoculated plants (Figure 2F).
Interaction of BLN1 and BLN2
Next, we were interested to see if the BLN1 and BLN2 secreted small peptides could physically interact as BLN complexes to facilitate cellular signaling. To test this hypothesis, bimolecular fluorescent complementation (BiFC) assays (Kerppola, 2006) were performed to test the interaction between BLN1 and BLN2. Bln1 and Bln2 full-length open reading frames were fused to both N-terminal and C-terminal halves of yellow fluorescent protein (YFP), respectively, and co-expressed in onion epidermal cells. As shown in Figure 3A and Table 2, the interaction of N-terminal BLN2 and C-terminal BLN1 re-comprised YFP activity. We did not observe YFP fluorescence in tests with the reciprocal (N-terminal BLN1 and C-terminal BLN2) constructs, implying some conformational constraints on successful interactions (Table 2). This may be due to the orientation of the GFP tag in relation to the interacting interface. This non-reciprocity was also observed in BiFC interaction experiments among Bgh effector proteins and barley small heat shock proteins (Ahmed et al., 2015).
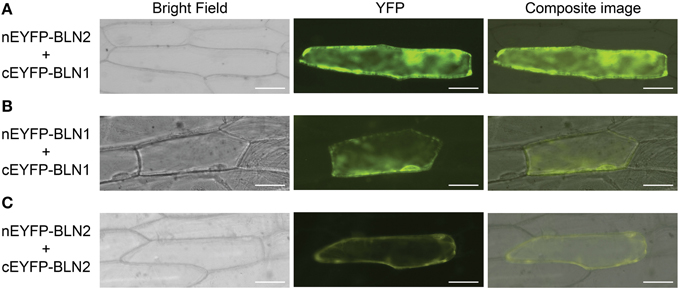
Figure 3. Bimolecular fluorescence complementation (BiFC) assay for BLN1 and BLN2 interactions. The Bln1 and Bln2 full length ORFs were PCR-amplified and cloned into EcoRI-BamHI sites of pSAT4-nEYFP-C1 or pSAT4-cEYFP-C1 to generate pSAT4-nEYFP-Bln1, pSAT4-nEYFP-Bln2, pSAT4-cEYFP-Bln1 pSAT4-cEYFP-Bln2 respectively. The designated plasmid combinations were co-bombarded into onion epidermal cells. The bright field and fluorescent images were taken using the Zeiss Axiovert 100 microscope equipped with the appropriate YFP filter. Statistical analysis of YFP count data is presented in Table 2. (A) Microscopic observation for interaction between BLN1 and BLN2. (B) Microscopic observation for BLN1 self-interaction. (C) Microscopic observation for BLN2 self-interaction.
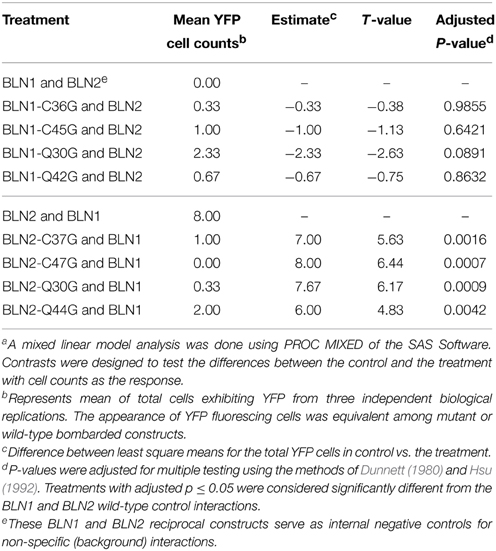
Table 2. Mixed linear analysis of mutations in the BLN1 and BLN2 IQ domain and cysteines and their effect on forming heteroduplexesa.
DISULFIND software (Ceroni et al., 2006) predicted that the cysteines in BLN1 and BLN2 form disulfide bonds; these are expected to stabilize knottin protein structures, which may be critical for interactions with other proteins (Combelles et al., 2008; Gracy et al., 2008). To test if these two conserved cysteines may be involved in the interaction interface between BLN1 and BLN2 (Kerppola, 2006), Cys36 and Cys45 in BLN1 and Cys37 and Cys47 in BLN2 were mutated to Gly and the resulting BiFC constructs were co-bombarded into onion epidermal cells. As shown in Table 2, the average number of observed fluorescent cells from three independent replications was significantly reduced (adjusted p < 0.0016). The reciprocal construct described above, as well as each of the BLN1 and BLN2 site-directed mutants also serve as negative controls for non-specific interactions (Kerppola, 2006). Interestingly, co-bombardment of constructs harboring BLN1 fused to the N-terminal and C-terminal halves of YFP also showed YFP activity (Figure 3B); similar results were also observed for BLN2 (Figure 3C). These results suggest that BLN family members can not only interact with each other, but also dimerize or polymerize with themselves. Even so, mutations in conserved residues may compromise protein stability, thus, this preliminary result should be viewed with caution without direct evidence that the mutant proteins actually accumulate.
Interactions between BLN Family Members and Calmodulin
Calmodulin (CaM) plays a pivotal role in controlling an abundance of Ca2+-based cellular signaling events (Berridge et al., 2003) and functions in response to changes in cellular calcium levels by interacting with various targets, including those in plant immunity (Yamniuk and Vogel, 2004; Du et al., 2009). These targets include IQ (isoleucine-glutamine; consensus sequence = [FILV]Qxxx[RK]Gxxx[RK])- (Rhoads and Friedberg, 1997; Bahler and Rhoads, 2002) and partial-IQ-motif containing proteins (Houdusse and Cohen, 1995; Munshi et al., 1996; Sienaert et al., 2002). As shown in Figure 4, sequence alignments indicate that BLN family members contain partial IQ motifs. Moreover, previous results from silencing and overexpression experiments suggested that BLN family members possess an S-gene function somewhat similar to MLO, a calmodulin (CaM)-binding protein in plant defense (Kim et al., 2002). Thus, we were interested to investigate if BLN1 or BLN2 could physically interact with CaM. Both Bln1 and Bln2 full-length open reading frames were fused to the N-terminal half of YFP, CaM was fused to C-terminal half, and co-expressed in onion epidermal cells. As illustrated in Figures 5A,B and Table 3, significant YFP fluorescence was observed, indicating a possible interaction of BLN1/BLN2 with CaM. To further examine the function of glutamine residues in the BLN1 and BLN2 partial IQ motifs, Gln30 and Gln42 in BLN1, and Gln30 and Gln44 in BLN2 were mutated to Gly. Quantification for fluorescent cells indicated that these mutations significantly reduced the numbers of observed interactions between BLN and CaM (adjusted p < 5.43E-04), suggesting that these two glutamine residues in BLN1 and BLN2 are necessary to facilitate the full-strength interaction with CaM (Table 3). Interestingly, the Gln mutations in BLN2 also negatively impacted BLN1-BLN2 interactions (Table 2, adjusted p < 0.0042).
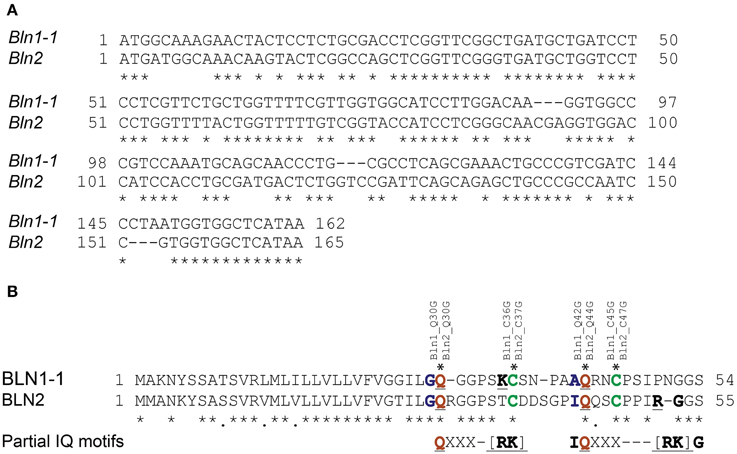
Figure 4. Alignment of Bln1 and Bln2 with the positions of conserved cysteines and potential partial IQ motifs. (A) Nucleotide alignment of Bln1 and Bln2 open reading frames. A “*” indicates conserved nucleotides. (B) Amino acid alignment of BLN1 and BLN2. The conserved glutamines (Q) in the partial IQ motifs are shown in red (with blue adjacent), cysteines (C) are shown in green. The conserved amino acids in potential partial IQ motifs (compared to the complete consensus [FILV]Qxxx[RK]Gxxx[RK]) are underlined. For BLN1, Q to G mutations were inserted at amino acid residues 30 and 42, and C to G mutations at residues 36 and 45. For BLN2, Q to G mutations were inserted at amino acid residues 30 and 44, and C to G mutations at residues 37 and 47.
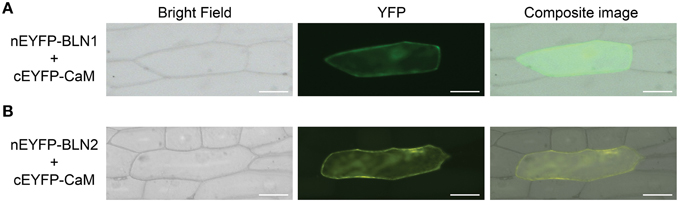
Figure 5. Bimolecular fluorescence complementation (BiFC) assay for interaction between BLN1/2 and CaM. The Bln1 and Bln2 full length ORFs were PCR-amplified and cloned into EcoRI-BamHI sites of EcoRI-BamHI sites of pSAT4-nEYFP-C1 or pSAT4-cEYFP-C1 to generate pSAT4-nEYFP-Bln1, pSAT4-nEYFP-Bln2, pSAT4-cEYFP-CaM respectively. The designated plasmid combinations were co-bombarded into onion epidermal cells. The bright-field and fluorescent images were taken using the Zeiss Axiovert 100 microscope equipped with appropriate YFP filter. Statistical analysis of YFP count data is presented in Table 3. (A) Microscopic observation for interaction between BLN1 and CaM. (B) Microscopic observation for interaction between BLN2 and CaM.
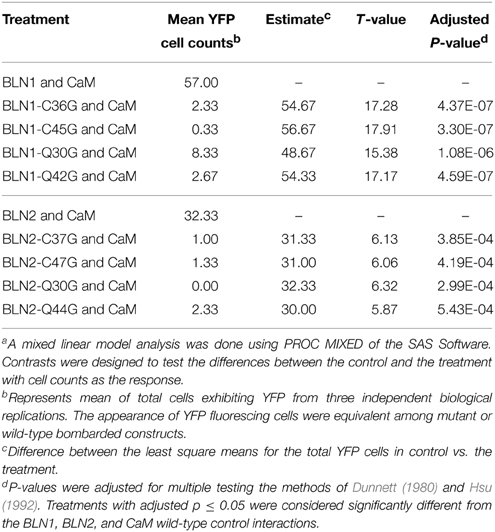
Table 3. Mixed linear analysis of mutations in the BLN1 and BLN2 IQ domain and cysteines and their effect on binding with CaMa.
Both BLN1 and BLN2 are cysteine-rich small peptides (Meng et al., 2009). These cysteines are positioned in or close to the partial IQ motifs (Figure 4). It is predicted that these cysteines may form inter- or intra-molecular disulfide bonds to maintain a structure supporting protein-protein interactions and cysteines in CaM targets are important for CaM-target interactions (Moore et al., 1999). To investigate the possible function of these two cysteines in the interaction between BLN and CaM, the Cys to Gly site-directed mutants described above were used in pairings with CaM, resulting in significantly reduced fluorescence activity (adjusted p < 4.19E-04) (Table 3). These data indicate that these cysteines play a role, either directly or indirectly, in the interaction between BLN protein and CaM.
Identification of Bln-mediated Response Pathways
The data presented above, combined with previous functional studies (Meng et al., 2009), indicate that the monocot-specific, BLN small secreted peptides negatively regulate barley-Bgh interactions. To identify genes influenced by Bln1 function, we took a mutational approach and used the BSMV-VIGS system to knock down Bln1 (Contig12219_at). We then performed Barley1 GeneChip expression profiling on the silenced plants to discover additional genes that impact Bln1-mediated regulation of immunity.
Figure 6 illustrates the basic matrix of the experiment. Key contrasts were designed to compare differences in transcript accumulation among Bln1-silenced plants relative to BSMV:00 (empty vector) controls. Comparison of BSMV:00 to mock (buffer-treated) controls enabled us to detect possible confounding effects of BSMV. To account for background-specific differences, we utilized two host genotypes; both compatible with our Bgh 5874 isolate and previously demonstrated to be good hosts for BSMV-VIGS experiments (Hein et al., 2005; Meng et al., 2009; Meng and Wise, 2012).

Figure 6. Barley1 GeneChip expression profiling design and phenotypes upon Bgh infection. Transcript profiling was based a split-plot design with 5 replications as blocks, Bgh treatment as the whole-plot factor, and all combinations of genotype [cv. Clansman (Mla13) and cv. HOR11358 (Mla9)] and VIGS treatment [Buffer control (mock), BSMV:00 (empty vector), and BSMV:Bln1248] as the split-plot factor for a total of 60 Barley1 GeneChip hybridizations. Seven-day-old plants were treated with Mock, BSMV:00, and BSMV:Bln1248. Twelve days after buffer and BSMV treatments, plants were inoculated with the compatible Bgh isolate 5874 (AVRa1, AVRa6, avra9, avra13), or non-inoculated. Leaves were harvested at 32 h after Bgh inoculation. Five leaves for each treatment were used for phenotyping 7 days after Bgh inoculation.
We performed five independent biological replications of a split-plot experimental design (shown in Figure 6) with replications as blocks, Bgh treatment as the whole-plot factor, and all combinations of genotype [Clansman (Mla13), HOR11358 (Mla9)] and VIGS treatment [Buffer control (mock), BSMV:00 (empty vector), BSMV:Bln1248] as the split-plot factor for a total of 60 GeneChip hybridizations. Ten seedlings were used as a split-plot experimental unit. Twelve days after VIGS treatment, plants were transferred to a growth chamber where half of the plants in each replication were challenged with the compatible Bgh isolate 5874; the other half remained un-inoculated. At 32 h after inoculation (HAI), 5 of the 10 leaves from each treatment were harvested for RNA isolation; this timepoint has the highest differential Bln1 transcript accumulation in prior experiments (Meng et al., 2009), and is after initial establishment of the perihaustorial interface (Caldo et al., 2004). The remaining 5 leaves were used to document infection phenotypes 7 days after inoculation (representative experiments shown in Figure 6).
To interrogate the GeneChip data, we conducted mixed linear model analyses of the normalized signal intensities for each of the 22,840 Barley1 probe sets (Caldo et al., 2004, see Materials and Methods). Using a stringent threshold p < 0.0001 and false discovery rate (FDR) < 5%, 47 genes were suppressed or induced in BSMV:Bln1248 silenced plants as compared to the BSMV:00 controls in the HOR11358 (Mla9) background. Using the same threshold criteria, 48 genes were similarly affected in the Clansman (Mla13) background (Figure 7). Many of the genes affected by silencing Bln1 in HOR11358, as opposed to Clansman, had dissimilar annotations; this could be due to genotype-specific silencing, genotype-specific probe-set efficiency, or it could reflect the threshold p-value we selected (i.e., genes in one background may still be significant, but at a less conservative threshold). Nevertheless, six of these genes were suppressed in common, including the Bln1 target, represented by Barley1 Contig12219_at [p = 4.66E-19(HOR11358)/p = 8.30E-06(Clansman)] (Figure 8A; Supplemental Table S1). Bln2 (Contig26496_at; p = 4.99E-06) was suppressed along with Bln1 in the HOR11358 (Mla9) background (Figure 8B), but was not significant at the selected threshold p < 0.0001 in the Clansman (Mla13) background.
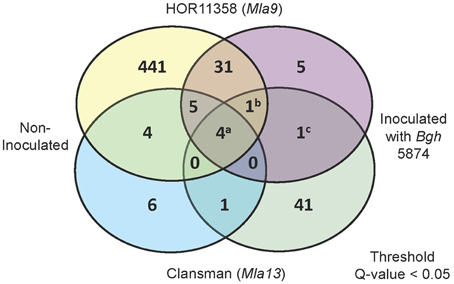
Figure 7. Summary of differentially expressed genes in Bln1-silenced leaves as compared to BSMV:00 controls. Venn diagram represents differentially expressed genes (Threshold: q < 0.05) with and without inoculation with Bgh for both HOR11358 (Mla9; upper circles) and Clansman (Mla13; lower circles) background plants. Contrasts were assessed between empty vector (BSMV:00) and Bln1 silenced (BSMV:Bln1248) plants. Superscripts (a) Contig12608_at, HS16M03u_x_at, HV10C01u_s_at, Contig3615_at (Importin α-1b); (b) Contig12219_at (Bln1); (c) Contig3680_s_at (Sec61 γ).
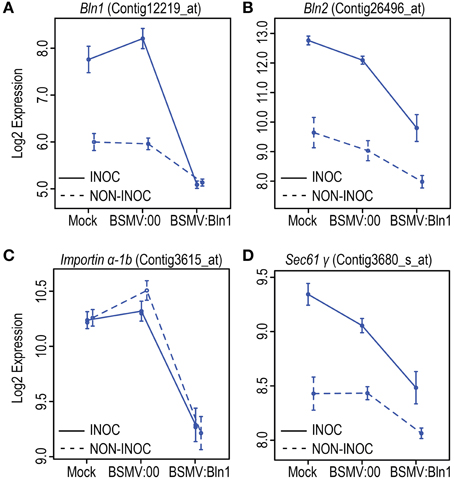
Figure 8. Log2 expression level results from Barley1 GeneChip profiling of Bln1, Bln2, Importin α-1b, and Sec61 γ in Bln1-silenced leaves. Barley stripe mosaic virus (BSMV)-Virus Induced Gene silencing was performed on HOR11358 (Mla9) plants as described in Figure 6. Twelve days after buffer and BSMV treatments, plants were inoculated with the compatible Bgh isolate 5874 (AVRa1, AVRa6, avra9, avra13). Log2 expression levels for Bln1 (A), Bln2 (B), Importin α-1b (C), and Sec61 γ (D) are plotted in graphs. INOC, inoculation; NON-INOC, Non-inoculated. Error bars represent the standard errors (SE).
The experiment also yielded many genes that were influenced by infection with BSMV:00, in addition to BSMV:Bln1248 (Supplemental Table S1). This may be due to an overlap in general defense gene functions, or may represent strictly BSMV-dependent responses. Although one must be cautious regarding overlap in general defense-gene functions, BSMV has been shown not to interfere with infection of Blumeria graminis f. sp. tritici, the causal agent of powdery mildew in wheat (Tufan et al., 2011). However, to specifically understand Bln1-regulated targets, we restricted our follow-up functional assays to genes with no confounding effects (Mock vs. BSMV:00 p > 0.01).
In addition to the newly identified genes, we also discovered that Bln2 (Contig26496_at) was suppressed in Bln1 silenced plants (Figure 8B). Sequence comparison of Bln1 and Bln2 indicates that these two genes share significant identity (Figure 4; Meng et al., 2009). Although the Bln1 silencing construct was designed across a 3′ divergent region, the two Bln family members share 13 contiguous nucleotides, indicating that Bln2 might be unintentionally silenced (Jackson et al., 2003).
Functional Characterization of Conserved Genes Suppressed Upon Bln1 Silencing
Analysis of the cohorts described above should provide mechanistic clues to the function of Bln1 in innate immunity. For example, the most significant candidate from this comparison is Barley1 Contig3615_at [p = 4.85E-11(HOR11538)/p = 1.0E-8(Clansman)], representing the gene encoding Importin subunit α-1b, which is suppressed in BSMV:Bln1248 silenced plants (Figure 8C). Importin subunit α-1b localizes to the perinuclear region of the cytoplasm, where it binds specifically to substrates containing a nuclear localization signal (NLS) and promotes docking of these substrates to the nuclear envelope for subsequent import (Jiang et al., 2001). A homolog of Importin α is also involved in innate immunity in Arabidopsis (Palma et al., 2005). Silencing of Bln1 also results in the suppression of genes encoding components in the protein secretory pathway, including Sec61 γ, represented by Contig3680_at [p = 2.89E-05(HOR11538)/p = 9.31E-05(Clansman)] (Figure 8D). Sec61 γ protein is a component of the SEC61 complex that is a conserved protein-conducting channel for secretory protein translocation across the endoplasmic reticulum (ER) membrane (Osborne et al., 2005). Also represented in the common set of six is a gene encoding a putative cysteine protease inhibitor (HV10C01u_s_at). This protein is similar to maize CC9, an apoplastic cysteine protease inhibitor that suppresses host immunity to Ustilago maydis (Van Der Linde et al., 2012). Another gene in the conserved set (Contig12608_at) encodes a protein of unknown function but with a classic nuclear localization signal (Dinkel et al., 2014). Genes represented by Contig12219_at (Bln1), Contig3680_at (Sec61 γ), HS16M03u_x_at (unknown), and HV10C01u_s_at (cysteine protease inhibitor) are induced by Bgh infection, whereas, Contig3615_at (Importin α-1b) and Contig12608_at (NLS protein) are not. Nevertheless, transcript accumulation of all six is suppressed in Bln1-silenced plants (Figure 7).
Based on these predicted annotations, we then selected a sub-set for functional analysis via BSMV-mediated gene silencing. Both Importin α-1b (Contig3615_at) and Sec61 γ (Contig3680_s_at), were introduced into the BSMV-VIGS system as BSMV:Imp α-1b319 and BSMV:Sec61γ319, respectively, and plants were subjected to silencing as described above. To alleviate off-target silencing, the chosen genes were aligned to the barley genome resource (Mayer et al., 2012). Subsequently, unique, single-copy regions of each target gene were used to design BSMV-VIGS primers (Supplemental Table S2), and each construct was bombarded in at least two independent replicates of 10 plants each. As shown in Figure 9, both of these genes impact powdery mildew development, as demonstrated by significantly less fungal colonies and hyphal growth on the surface of epidermal cells, as compared to BSMV:00 controls (Figures 9A,B, Table 1). qRT-PCR on RNA isolated from the silenced leaves as well as BSMV:00 controls confirmed that transcript accumulation for the target genes was suppressed (Figure 9C). These data suggest that both Importin α-1b and Sec61 γ play negative roles in barley innate immunity to Bgh.
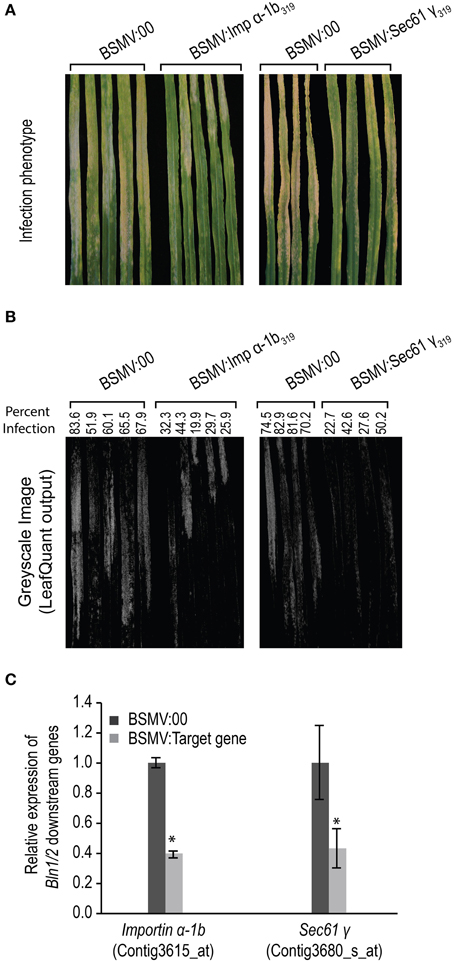
Figure 9. Infection phenotype of Blumeria graminis f. sp. hordei (Bgh) for leaves silenced for Importin α-1b and Sec61 γ. Gene silencing was performed on HOR11358 (Mla9) plants and mediated by Barley stripe mosaic virus (BSMV). Seven-day-old plants were treated with buffer and BSMV. Twelve days after buffer and BSMV treatments, plants were inoculated with compatible Bgh isolate 5874 (AVRa1, AVRa6, avra9, avra13). The infection phenotypes were photographed 7 days after inoculation and hyphal growth area for each leaf was quantified by LeafQuant-VIGS software (Table 1). RNA samples for gene expression analysis were prepared from the same quantified leaves and gene levels were analyzed by quantitative reverse-transcriptase polymerase chain reaction analysis on transcript accumulation. The amount of RNA in each reaction was normalized using primers specific for barley Actin. The 2−ΔCT method was used to calculate the target gene expression for each individual silencing construct as compared with BSMV:00 treated plants (Schmittgen and Livak, 2008). Fold change due to silencing is calculated by dividing the expression mean value for the targeted gene in silenced plants by the mean value measured in BSMV:00-treated plants (* designates that p < 0.05). Bars represent standard error calculated from at least five independent plants from two replicate experiments. The average value in BSMV:00 samples was set to 1.0. (A) Infection phenotype of Bgh on Importin α-1b and Sec61 γ silenced leaves. (B) LeafQuant outputs and quantification fungal infection. (C) Relative expression of Importin α-1b and Sec61 γ in BMSV induced gene silenced leaves.
Discussion
BLUFENSIN1 (BLN1) and BLUFENSIN2 (BLN2), two members in a monocot-specific family of cysteine-rich peptides, are barley susceptibility factors to powdery mildew (Meng et al., 2009). Both BLN1 and BLN2 reveal structural and sequence similarities to knottins, small disulfide-rich proteins characterized by a unique “disulfide through disulfide knot” (Combelles et al., 2008). Bln family members are highly upregulated upon fungal infection and BLN proteins can be secreted into the apoplast. Although silencing Bln does not break Mla [Nucleotide binding, Leucine rich repeat (NLR)]-mediated resistance, knockdown of Bln1 increases barley innate immune responses and overexpression renders the barley host supersusceptible in compatible interactions.
Based on these observations, we postulate that BLN family members are potential signal molecules co-opted by Bgh effectors to bypass innate immune systems and colonize the host. In turn, interactors or partners of BLN would also be expected to play key roles in mediating the plant immune response. This hypothesis is supported by the observed interaction between BLN1 and BLN2, and barley calmodulin (CaM) (Figure 5). CaM, as a universal Ca2+ sensor, plays essential roles in regulating numerous intracellular processes, including plant defense (Kim et al., 2002; Reddy et al., 2011; Bender and Snedden, 2013). The BLN protein may function as a ligand to interact with CaM and change its conformation to alter downstream CaM signaling (Figure 10). Alternatively, BLN may interact with barley partner(s) or Bgh effector(s) to favor basic compatibility.
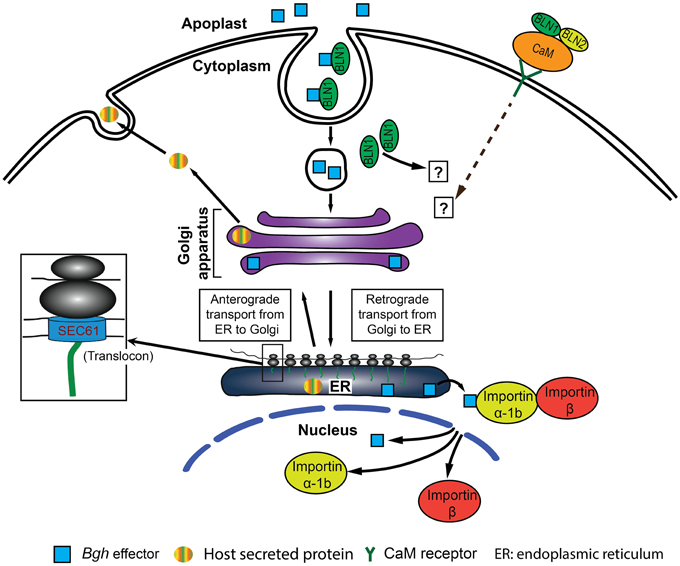
Figure 10. Predictive model for BLN1 and BLN2 function in the barley immune system. (1) BLN (green filled circle) can function as a ligand and binds with effectors in the apoplast to promote barley-Bgh compatible interactions. (2) The BLN-effector complex then can enter plant cells by endocytic routes, and move to the ER region through a retrograde secretory pathway. (3) Subsequently, effectors could be released into the cytoplasm through the SEC61 pore-forming complex and picked up by Importin α-1b and translocated into the nucleus by the Importin complex to regulate gene expression. (4) The level of BLN1-effector complex may be the positive force regulating the expression of Importin α-1b and Sec61 γ to titrate the BLN-effector level. Alternatively, BLN binds CaM, a Ca2+ sensor, in the apoplast and changes CaM conformation to alternate the interaction of CaM with its receptor in the plasma membrane (Cui et al., 2005; Wang et al., 2009) to positively regulate the state-steady levels of Importin α-1b and Sec61 γ. In both cases, BLN, Importin α-1b, and Sec61 γ proteins are positive regulators for barley-Bgh compatible interactions (or negative regulators for innate immunity). Silencing of Bln1 results in the suppression of Importin α-1b and Sec61 γ, impairing protein anterograde and retrograde trafficking as well as translocation into the nucleus required for barley-Bgh compatible interaction.
Nuclear Import and the Secretory Pathway—Pathogen Effector Import Systems in Host Plants
Negative regulation of the basal defense pathway prevents unchecked potentiation of the response and deleterious effects on normal cell functions (Ge et al., 2007). Forty-seven and forty-eight genes were identified in HOR11358 (Mla9) and Clansman (Mla13), respectively, that are significantly differentially expressed (p < 0.0001) when Bln1 is silenced (Figure 7, Supplemental Table S1). Six of these genes were suppressed in common between the two backgrounds. Of these, Importin α-1b and Sec61 γ are involved in protein trafficking, the CC9 homolog (HV10C01u_s_at) is a putative apoplastic cysteine protease inhibitor, and finally, the gene represented by Contig12608_at, encodes an unknown protein with a classic nuclear localization signal (Dinkel et al., 2014). Silencing two (Importin α-1b and Sec61 γ) of the conserved set of six genes significantly reduced host susceptibility in compatible interactions of barley and Bgh (Figure 9), and CC9 suppresses host immunity to Ustilago maydis in maize (Van Der Linde et al., 2012), suggesting that transcription of these plant genes is essential for the fungus to successfully colonize host cells.
It has become evident that the interaction of pathogen effectors and resistance proteins in the nucleus is critical to R-gene-mediated resistance (Burch-Smith et al., 2007; Shen et al., 2007; Tameling and Baulcombe, 2007; Wirthmueller et al., 2007; Liu and Coaker, 2008; Tameling et al., 2010). Data presented here indicates that translocation of protein into the nucleus is a key step in innate immunity as well. Importin α in pepper was shown to interact with AvrBs3, a type III-secreted effector from Xanthomonas campestris pv. vesicatoria, both in yeast and in vitro through a nuclear localization signal (NLS) (Szurek et al., 2001). Given that the gene encoding Importin subunit α-1b (Contig3615_at) is significantly down-regulated in Bln1-silenced plants, we postulate that a function of BLN1, even in the absence of Bgh infection (Figure 8C), could be to sustain transcript accumulation of Importin. Induction of BLN1 by Bgh may enhance the translocation of select effectors from the apoplast to the nucleus, which might be necessary for the fungus to colonize its barley host.
Such a scenario, though not reported before in cereal-fungal interactions, is not without precedent. In the interaction of Arabidopsis with Agrobacterium, a historically important pathogen most widely know for its role in plant transformation (McCullen and Binns, 2006), multiple Importin α proteins interact with both Agrobacterium VirD2 and VirE2. However, Importin α-4 appears to be the most crucial isoform for transfer of Vir proteins to the plant cell nucleus (Bhattacharjee et al., 2008).
It should also be noted that although two full-length Importin α isoforms have been identified in barley (Importin α-1a and Importin α-1b, represented by probe sets Contig4129_at and Contig3615_at, respectively), only expression of Importin α-1b is significantly affected by Bln1 silencing. This indicates that Importin α-1b is specifically involved in BLN-mediated resistance to barley powdery mildew.
Nonetheless, Importin α-1b in barley and Importin α3 in Arabidopsis (Palma et al., 2005) have opposite effects on plant defense against pathogens. This suggests that different isoforms of Importin α have substrate-specific recognition to differentially regulate plant immunity. Substrate-specific recognition of Importin has also been observed for nuclear import of proteins involved in rice photomorphogenesis (Jiang et al., 2001) and for neural differentiation of mouse embryonic stem cells (Goldfarb et al., 2004; Yasuhara et al., 2007, 2013), signifying the diverse temporal and spatial regulation influenced by Importin isoforms in plant and animal systems.
Silencing of Bln1 also results in the coordinate suppression of genes encoding components in the protein secretory pathway, including the SEC61 complex. The SEC61 complex is a conserved protein-conducting channel for translocation across the endoplasmic reticulum (ER) membrane (Osborne et al., 2005; Kelkar and Dobberstein, 2009; Park and Rapoport, 2012) and is required for induction of systemic aquired resistance to Pseudomonas syringae pv. maculicola in Arabidopsis (Wang et al., 2005). Recently, the Sec61 β subunit in barley has been shown to be an ER protein transporting pore that is required for host susceptibility to powdery mildew (Zhang et al., 2013).
Does BLN1 Drive Protein Trafficking?
The protein secretory pathway starts with insertion of protein into the SEC61 translocon complex and involves a series of steps by which proteins are transported between organelles in anterograde or retrograde directions. Protein trafficking into the nucleus includes the interaction of transported targets with the Importin complex. The secretory pathway plays a vital role in plant disease resistance (Kwon et al., 2008a,b; Rojo and Denecke, 2008; Wang and Dong, 2011). Down-regulation of multiple components in the secretory pathway in Bln1 silenced plants point to a role of Bln1 in the regulation of this pathway. One possibility is that BLN1 may interact with host-secreted proteins required for pathogen-host interaction. Silencing of Bln1 may reduce the amount of these BLN1 interacting host-secreted proteins, resulting in down-regulation of anterograde protein transport. Alternatively, interaction of BLN1 with Bgh or host secreted proteins could signal the entry of BLN interactors into the host cell via the retrograde pathway (Spooner et al., 2006; Johannes and Popoff, 2008; Dong et al., 2013; Drerup and Nechiporuk, 2013; Koyuncu et al., 2013). Silencing of Bln1 reduces these interactions and therefore release the demand for components involved in protein trafficking pathway, such as Sec61 γ and Importin α-1b. A third scenario may involve BLN1 interacting with a specific signal molecule, such as the Ca2+ sensor CaM, to positively regulate transcript accumulation of genes encoding components involved in protein trafficking, such as Sec61 γ and Importin α-1b. Knock down of Bln1 would suppress the expression of genes encoding these components (Figure 10). All three hypotheses are supported by the observation that Bln family members regulate the expression of genes implicated in protein trafficking, overexpression of Bln renders barley more susceptible to Bgh (Figure 2), and that silencing of Bln1, Importin α-1b, and Sec61 γ increases barley innate immunity (Figures 6, 9). Thus, BLN proteins may modulate protein transport in barley-Bgh interactions and BLN levels influence protein trafficking in infected barley cells.
Summary: Are BLN1 and BLN2-potential Host-targeting Signals for Bgh?
In the interaction between barley and Bgh, transfer of signals may be expected to occur between host and pathogen during formation of the perihaustorial interface (Dodds et al., 2004, 2006).
From an evolutionary standpoint, genes in redundant networks with an incredible level of buffering capacity imply minimal selective pressures acting on these genes. One example was discussed by Xu et al. (2006), where Atwrky18/Atwrky40 and Atwrky18/Atwrky60 double mutants were more resistant to Pseudomonas syringae DC3000 but more susceptible to Botrytis cinerea infection, but single Atwrky mutants behaved similar to wild-type plants. BLN1 and BLN2 are potentially part of a redundant set of negative regulators for plant defense.
Both BLN1 and BLN2 are highly induced upon Bgh inoculation and transient overexpression increases barley plants susceptibility; these results are consistent with the silencing results. We further identified that BLN proteins could be secreted into the apoplast and interact with each other and CaM in plant cells. Thus, BLN could act as potential host-targeting signals for Bgh to colonize in plant cells; the interactions between BLN and Bgh effectors could fine-tune sets of protein pathways, which might be involved in transporting fungal proteins into the host cells.
Materials and Methods
Plant Materials and Fungal Isolates
Seedlings of barley lines Black Hulless, CI 16151 (Mla6), HOR11358 (Mla9), and Clansman (Mla13) were used for functional analysis. Virus infected barley was maintained in a growth chamber with a 16 h photoperiod with light intensity at 550 μmol m−2 s−1 and a daytime temperature of 24°C and dark temperature of 20°C. Bgh isolate 5874 (AVRa1, AVRa6, avra9, avra13) was propagated on H. vulgare cv. Manchuria (CI 2330) in a controlled growth chamber at 18°C (16 h light/8 h darkness).
BLN1 and BLN2 Subcellular Localization
Total RNA was extracted from CI 16151 (Mla6) plants 20 hai with Bgh isolate 5874 (AVRa6) according to the method of Caldo et al. (2004). First-strand cDNA was synthesized using 2 μg of total RNA, oligo(dT)20 primer and Superscript reverse transcriptase III (Invitrogen, Carlsbad, CA). Subsequently, first strand cDNA was used as the template to amplify Bln1 and Bln2 coding sequences with/without signal peptides or the signal peptide regions; Primers were designed according to Bln1 EST sequence (GeneBank Accession no. is FJ156737) and Bln2 EST sequence (GeneBank Accession no. is FJ156745) and listed in Supplemental Table S2.
BLN-GFP chimeric constructs were made using overlapping PCR. First, full length Bln1 was amplified using primer pair Bln1-NcoN_pf1 and Bln1-C_pr1; GFP was amplified with pEGFP (Clontech Laboratories, Inc., Mountain View, CA) as template and primer pair GFP-FKS_pf1 and GFP-Bam_pr1. The final PCR was performed using PCR products of the previous two reactions as template and primer pair Bln1-NcoN_pf1 and GFP-Bam_pr1. The final PCR product was digested with NcoI and BamHI and ligated into similarly treated pTRL2 (Restrepo et al., 1990) to yield p35S:BLN1+SP, harboring coding regions for full-length BLN1 and GFP. A similar strategy was adopted to make the Bln1 signal peptide-GFP construct, p35S:BLN1_SP only, as well as Bln1-GFP construct absent the Bln1 signal peptide, p35S:BLN1-SP. When making corresponding Bln2-GFP constructs (p35S:BLN2+SP, p35S:BLN2_SP only and p35S:BLN2-SP), PCR products were digested with NcoI and SmaI and inserted into similarly treated pBLN1+SP.
These constructs were delivered into onion epidermal cells by using biolistic PDS-1000/he system (Bio-Rad, Hercules, CA, USA) as described by Elling et al. (2007). After bombardment, epidermal peels were incubated for 24 h in the dark at 20–25°C. Plasmolysis of onion epidermal cells was attained by soaking the peels in 1 M sucrose solution for 20 min. A Zeiss Axio Imager M.1 microscope (Zeiss, Inc., Thornwood, NY) was used for observation. At least three independent replicate experiments were conducted.
To visualize GFP in the apoplast, onion epidermal cells were incubated in agar medium supplemented with 3% sucrose (Murashige and Skoog, 1962) for 20 h, and transferred to 20 mM Pipes-KOH (pH 7.0) for 4 h on half-concentrated Murashige and Skoog (1962) agar medium supplemented with 1% sucrose (Genovesi et al., 2008). The pH 7.0 medium neutralizes the normally acidic apoplast in order to observe the fluorescence patterns.
Overexpression of Bln1 and Bln2 by Using the BSMV system
To introduce a foot- and-mouth disease virus (FMDV)-2A self-cleavage peptide and GFP for expression of foreign gene, pBPMV-IA-V5 (Zhang et al., 2010) was used as template with primer pair BS3-G4F1 and BS3-G4R1 for PCR to produce DNA fragment A, which was used as template with primer pair BS3-G4F2 and BS3-G4R2 to produce DNA fragment B. BSMV:γ (Meng et al., 2009) was used as template with primer pair BSMV-R3-F3 and BS3-4Rev for PCR to produce DNA fragment C. DNA fragments B and C were then used for overlapping PCR with primer pair BSMV-R3-F3 and BS3-G4R2 for PCR to produce DNA fragment D. Product D was digested with BglII and KpnI and ligated into BSMV: γ digested by BglII and KpnI to produce pBSMV-OEx:GFP.
To make Bln1 and Bln2 overexpression constructs, Bln1 and Bln2 cDNA described above were used as templates and primer pairs BSBln1Ov_pf1/BSBln1Ov_pr1 or BSBln2Ov_pf1/BSBln2Ov_pr1 were used respectively (Supplemental Table S2). PCR fragments contained an introduced StuI and BamH1 recognition sites at the 5′ and 3′ ends, respectively and were inserted into the StuI and BamH1 sites of pBSMV-OEx:GFP, the resulting vectors were designated as pBSMV-OEx:Bln1 and pBSMV-OEx:Bln2, respectively.
DNA bombardment and subsequent virion mechanical infection on HOR11358 (Mla9) plants was performed according to Meng et al. (2009). A Zeiss Axio Imager M.1 microscope (Zeiss, Inc., Thornwood, NY) was used for observation the GFP. At least two independent replicate experiments were performed.
BLN1 and BLN2 Interactions Via Bimolecular Fluorescence Complementation (BiFC)
Site-directed mutagenesis was carried out on Bln1 and Bln2 full length ORFs using QuickChange™ site-directed mutagenesis kit (Stratagene, La Jolla, CA). Primers used for amplification and mutagenesis are listed in Supplemental Table S2. From the BLN1 start codon, amino acid residues 30 and 42 were changed from Q to G and residues 36 and 45 were changed from C to G. For BLN2, amino acid residues 30 and 44 were changed from Q to G and residues 37 and 47 were changed from C to G. The resulting products contained EcoRI and BamHI restriction sites respectively, and cloned into EcoRI-BamHI sites of pSAT4-nEYFP-C1 or pSAT4-cEYFP-C1 to generate pSAT4-nEYFP-Bln1, pSAT4-nEYFP-Bln2, pSAT4-cEYFP-Bln1, and pSAT4-cEYFP-Bln2, respectively. For coexpression, particle bombardment was performed using onion epidermal cells. Gold particles (1.6 μm diameter) (Bio-Rad) were washed with 100% ethanol and coated with 1.25 μg of each DNA using standard procedures. cDNA-coated gold particles were bombarded at 1100 p.s.i. and 9 cm distance using a Biolistic Particle Delivery System PDS-1000/He (Bio-Rad). Bombarded tissues were incubated at 25°C in darkness for ~24 h before being assayed for YFP activity. The bright-field and fluorescent images were taken using the Zeiss Axiovert 100 microscope with appropriate YFP filter.
BLN Interactions with Calmodulin Via Bimolecular Fluorescence Complementation (BiFC)
Calmodulin full length ORFs were PCR-amplified using primers listed in Supplemental Table S2. The resulting product containing EcoRI and BamHI restriction sites was cloned into EcoRI-BamHI sites of pSAT4-cEYFP-C1 to generate pSAT4-cEYFP-CaM. This was combined with Bln1 or Bln2 pSAT4 constructs as described above for bombardment and microscopic observation. At least three independent replicate experiments were conducted.
Statistical Analysis of YFP Cell Count Data
All cells exhibiting YFP were counted for each of the 3 independent biological replications. A mixed linear model analysis of observed YFP cell count data was conducted using PROC MIXED in SAS software. The model used total YFP cell count as the response and included random effect for replications. Four contrasts in SAS software were used to compare different controls (Bln1 and CaM, Bln2 and CaM, Bln1 and Bln2, Bln2 and Bln1) with their respective treatments. The least square means were correlated and when the p-values were adjusted using the Dunnett (1980) method, PROC MIXED used the factor-analytic covariance approximation described in Hsu (1992).
Target Synthesis and GeneChip Hybridization
Total RNA was isolated using a hot (60°C) phenol/guanidine thiocyanate method described by Caldo et al. (2004). Trizol-like reagent was made from 38% saturated phenol (pH 4.3), 0.8M guanidine thiocyanate, 0.4M ammonium thiocyanate, 0.1M sodium acetate (pH 5.0) and 5% glycerol (Fisher Scientific, Pittsburg, PA). RNA was purified further using RNeasy columns (Qiagen, Valencia, CA). Probe synthesis, labeling, and hybridization to Barley1 GeneChip probe arrays (Affymetrix #900515; Close et al., 2004) were performed using One Cycle and GeneChip IVT labeling protocols based on the Affymetrix manual (Affymetrix, Santa Clara, CA) at the Iowa State University GeneChip Core facility.
Normalization and Mixed Linear Model Analysis of Barley1 Genechip Data
As described previously (Caldo et al., 2004; Wise et al., 2007), we conducted mixed linear model analyses of the normalized signal intensities for each of the 22,840 Barley1 probe sets (Wolfinger et al., 2001). RMA normalization and data transformation was done using package affy in BioConductor/R. Mixed linear model analysis was conducted using PROC MIXED in SAS software. The model used RMA normalized expression values as the response, replication, inoculation and treatment (genotype*vector) as fixed factors, and replication*inoculation and replication*treatment as random factors. Contrasts in SAS software (SAS Institute Inc., Cary, NC, U.S.A.) were used to compare transcript levels between treatments (Mock vs. BSMV:00; BSMV:00 vs. BSMV:Bln1248) of a specific genotype [HOR11538 (Mla9) or Clansman (Mla13)] after infection with Bgh 5874. Q-values were estimated using the smoother method described in Storey and Tibshirani (2003). From these analyses, we expected to identify sets of genes involved in Bln1-mediated compatibility or incompatibility, and also genes that are perturbed in response to BSMV infection, in both Bgh inoculated vs. non-inoculated reference plants.
BSMV-VIGS
Inserts for BSMV-VIGS were amplified by PCR using primers that add PacI and NotI restriction sites to the 5′ end and 3′ end, respectively (Supplemental Table S2). These sites enable ligation of the fragment in antisense orientation into the BSMV:γ vector. Silencing experiments were performed as described previously (Meng et al., 2009; Xi et al., 2009; Meng and Wise, 2012; Xu et al., 2014). Plants were maintained for 12 days in a growth chamber (Percival Scientific, Perry, IA, USA) with 16 h of light at 24°C (550 μmol m−2 s−1) and 8 h darkness at 20°C. Plants were then inoculated with Bgh isolate 5874 (avra9) conidiospores [compatible interaction with HOR 11358 (Mla9)] and maintained in a growth chamber 16 h of light/ 8 h of darkness at 18°C. The infection phenotype was monitored for 7 days.
Quantitative Real-time PCR
Barley leaves were pulverized in liquid nitrogen and total RNA extracted using Trizol-like reagent (Caldo et al., 2004). Genomic DNA was degraded by RNase-free DNase I (Ambion, Austin, TX, U.S.A.). SuperScript III reverse transcriptase (Invitrogen, Carlsbad, CA, USA) was used to synthesize first strand cDNA using 2 μg total RNA and oligo(dT)20 primer. This cDNA was used as a template for qRT-PCR to determine expression of various target genes to barley Actin. The qRT-PCR was performed using a Bio-Rad iCycler (Bio-Rad, Hercules, CA, USA). Conditions for 20 μL reactions using PerfeCTa® SYBR® Green FastMix® for iQ (Quanta Biosciences, Gaithersburg, MD) were 95°C for 3 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 1 min, then a melt curve was determined by starting at 55°C for 10 s and then increasing by 0.5°C every 10 s for 80 cycles. Three technical replicates for each biological sample in addition to four or five biological samples per treatment were included in each experiment. Target gene expression was calculated using the 2−ΔCT method for the BSMV:target gene and BSMV:00-treated plants. The fold change due to silencing was calculated by dividing the expression value for each BSMV:target gene treated leaves by the mean value measured in BSMV:00 treated plants (Schmittgen and Livak, 2008).
Imaging of Bgh Infection Phenotypes
At seven days after powdery mildew inoculation, third leaves of BSMV treated plants were randomly selected and photographed at high resolution (2592 pixels × 3456 pixels, i.e., 9 Megapixel at 4:3 aspect ratio) using a Canon PowerShot SX110 IS and the Vidpro professional Photo and Video LED light kit model Z-96K. The leaves were set on black felt for uniform, high contrast background. Subsequent images were analyzed using an in-house pattern-recognition software, designated LeafQuant-VIGS, developed using MathWorks® MATLAB® 7.14 and Image Processing Toolbox™ 8.0. Starting with high-resolution RGB images of each leaf, LeafQuant-VIGS first defines the edges, then detects the background and converts it to uniformly true black, converts the high-resolution color RGB image to an 8-bit gray-scale image with 256 shades of gray, and outputs histograms of the hyphal distribution per leaf, which then reports mean, median, and quantiles of the results as a csv (comma separated values) file for further processing (Whigham et al., 2015). Because elongating secondary hyphae (ESH), an indicator of functional haustoria (Ellingboe, 1972), are white and the barley leaf is green, these differences can be used to quantify fungal growth in terms of percent infection. LeafQuant-VIGS software is offered under MIT license via Github at http://git.io/leafquant.
Analysis of LeafQuant-VIGS Data
A separate linear model analysis of the LeafQuant-VIGS data was conducted for each silencing and over-expression construct using the package MULTCOMP in R programming language (Hothorn et al., 2008). All models used percent infection as the response. The model was used to compare the mean percent infection of BSMV:00 treated leaves to the mean percent infection for mock and BSMV:construct treated leaves. P-values were adjusted for multiple testing using the Dunnett method (Dunnett, 1955).
Data Access
All detailed data and data from expression profiling have been deposited as Accession number BB101 in PLEXdb (http://plexdb.org/) (Dash et al., 2012). Files can be downloaded as batch files in MAGE-ML, CSV, CEL, DAT, or expression data formats at the Download Center or downloaded as individual CEL, CHP, DAT, or EXP files under “browse experiments.” Data has also been deposited as Accession number GSE61644 at NCBI-GEO.
Author Contributions
Conceived and designed the experiments: YM, WX, DN, and RW. Performed the experiments: YM, WX, GF, and RW. Analyzed the data: WX, YM, PS, GF, DN, and RW. Contributed reagents/materials/ analysis tools: YM, WX, PS, GF, DN, and RW. Wrote and edited the paper: WX, YM, PS, GF, and RW.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Matt Moscou for initial analysis of the Barley1 GeneChip dataset. This research was supported by National Science Foundation Plant Genome grants 05-00461, 09-22746, and 13-39348. This article is a joint contribution of the Iowa Agriculture and Home Economics Experiment Station and the Corn Insects and Crop Genetics Research Unit, USDA-Agricultural Research Service. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture or the National Science Foundation.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2015.00409/abstract
References
Acevedo-Garcia, J., Kusch, S., and Panstruga, R. (2014). Magical mystery tour: MLO proteins in plant immunity and beyond. New Phytol. 204, 273–281. doi: 10.1111/nph.12889
Ahmed, A. A., Pedersen, C., Schultz-Larsen, T., Kwaaitaal, M., Jørgensen, H. J., and Thordal-Christensen, H. (2015). The barley powdery mildew candidate secreted effector protein CSEP0105 inhibits the chaperone activity of a small heat shock protein. Plant Physiol. 168, 321–333. doi: 10.1104/pp.15.00278
Bahler, M., and Rhoads, A. (2002). Calmodulin signaling via the IQ motif. FEBS Lett. 513, 107–113. doi: 10.1016/S0014-5793(01)03239-2
Bellincampi, D., Cervone, F., and Lionetti, V. (2014). Plant cell wall dynamics and wall-related susceptibility in plant-pathogen interactions. Front. Plant Sci. 5:228. doi: 10.3389/fpls.2014.00228
Bender, K. W., and Snedden, W. A. (2013). Calmodulin-related proteins step out from the shadow of their namesake. Plant Physiol. 163, 486–495. doi: 10.1104/pp.113.221069
Bent, A. F., and Mackey, D. (2007). Elicitors, effectors, and R genes: the new paradigm and a lifetime supply of questions. Annu. Rev. Phytopathol. 45, 399–436. doi: 10.1146/annurev.phyto.45.062806.094427
Berridge, M. J., Bootman, M. D., and Roderick, H. L. (2003). Calcium signalling: dynamics, homeostasis and remodelling. Nat. Rev. Mol. Cell Biol. 4, 517–529. doi: 10.1038/nrm1155
Bhattacharjee, S., Lee, L.-Y., Oltmanns, H., Cao, H., Veena, C. J., and Gelvin, S. B. (2008). IMPa-4, an Arabidopsis importin α isoform, is preferentially involved in Agrobacterium-mediated plant transformation. Plant Cell 20, 2661–2680. doi: 10.1105/tpc.108.060467
Burch-Smith, T. M., Schiff, M., Caplan, J. L., Tsao, J., Czymmek, K., and Dinesh-Kumar, S. P. (2007). A novel role for the TIR domain in association with pathogen-derived elicitors. PLoS Biol. 5:e68. doi: 10.1371/journal.pbio.0050068
Büschges, R., Hollricher, K., Panstruga, R., Simons, G., Wolter, M., Frijters, A., et al. (1997). The barley Mlo Gene: a novel control element of plant pathogen resistance. Cell 88, 695–705. doi: 10.1016/S0092-8674(00)81912-1
Caldo, R. A., Nettleton, D., and Wise, R. P. (2004). Interaction-dependent gene expression in Mla-specified response to barley powdery mildew. Plant Cell 16, 2514–2528. doi: 10.1105/tpc.104.023382
Ceroni, A., Passerini, A., Vullo, A., and Frasconi, P. (2006). DISULFIND: a disulfide bonding state and cysteine connectivity prediction server. Nucleic Acids Res. 34, W177–W181. doi: 10.1093/nar/gkl266
Cesari, S., Bernoux, M., Moncuquet, P., Kroj, T., and Dodds, P. N. (2014). A novel conserved mechanism for plant NLR protein pairs: the ‘integrated decoy’ hypothesis. Front. Plant Sci. 5:606. doi: 10.3389/fpls.2014.00606
Chu, Z., Yuan, M., Yao, J., Ge, X., Yuan, B., Xu, C., et al. (2006). Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev. 20, 1250–1255. doi: 10.1101/gad.1416306
Close, T. J., Wanamaker, S. I., Caldo, R. A., Turner, S. M., Ashlock, D. A., Dickerson, J. A., et al. (2004). A new resource for cereal genomics: 22K barley GeneChip comes of age. Plant Physiol. 134, 960–968.
Combelles, C., Gracy, J., Heitz, A., Craik, D. J., and Chiche, L. (2008). Structure and folding of disulfide-rich miniproteins: insights from molecular dynamics simulations and MM-PBSA free energy calculations. Proteins 73, 87–103. doi: 10.1002/prot.22054
Cui, S. J., Guo, X. Q., Chang, F., Cui, Y. W., Ma, L. G., Sun, Y., et al. (2005). Apoplastic calmodulin receptor-like binding proteins in suspension-cultured cells of Arabidopsis thaliana. J. Biol. Chem. 280, 31420–31427. doi: 10.1074/jbc.M501349200
Dash, S., Van Hemert, J., Hong, L., Wise, R. P., and Dickerson, J. A. (2012). PLEXdb: gene expression resources for plants and plant pathogens. Nucleic Acids Res. 40, D1194–D1201. doi: 10.1093/nar/gkr938
Dinkel, H., Van Roey, K., Michael, S., Davey, N. E., Weatheritt, R. J., Born, D., et al. (2014). The eukaryotic linear motif resource ELM: 10 years and counting. Nucleic Acids Res. 42, D259–D266. doi: 10.1093/nar/gkt1047
Dodds, P. N., Lawrence, G. J., Catanzariti, A.-M., Ayliffe, M. A., and Ellis, J. G. (2004). The Melampsora lini AvrL567 avirulence genes are expressed in haustoria and their products are recognized inside plant cells. Plant Cell 16, 755–768. doi: 10.1105/tpc.020040
Dodds, P. N., Lawrence, G. J., Catanzariti, A. M., Teh, T., Wang, C. I., Ayliffe, M. A., et al. (2006). Direct protein interaction underlies gene-for-gene specificity and coevolution of the flax resistance genes and flax rust avirulence genes. Proc. Natl. Acad. Sci. U.S.A. 103, 8888–8893. doi: 10.1073/pnas.0602577103
Dong, B., Kakihara, K., Otani, T., Wada, H., and Hayashi, S. (2013). Rab9 and retromer regulate retrograde trafficking of luminal protein required for epithelial tube length control. Nat. Commun. 4:1358. doi: 10.1038/ncomms2347
Drerup, C. M., and Nechiporuk, A. V. (2013). JNK-interacting protein 3 mediates the retrograde transport of activated c-Jun N-terminal kinase and lysosomes. PLoS Genet. 9:e1003303. doi: 10.1371/journal.pgen.1003303
Du, L. Q., Ali, G. S., Simons, K. A., Hou, J. G., Yang, T. B., Reddy, A. S. N., et al. (2009). Ca2+/calmodulin regulates salicylic-acid-mediated plant immunity. Nature 457, 1154–U1116. doi: 10.1038/nature07612
Dunnett, C. W. (1955). A multiple comparison procedure for comparing several treatments with a control. J. Am. Statist. Ass. 50, 1096–1121.
Dunnett, C. W. (1980). Pairwise multiple comparisons in the unequal variance case. J. Am. Statist. Ass. 75, 796–800.
Eichmann, R., Bischof, M., Weis, C., Shaw, J., Lacomme, C., Schweizer, P., et al. (2010). BAX INHIBITOR-1 is required for full susceptibility of barley to powdery mildew. Mol. Plant Microbe Interact. 23, 1217–1227. doi: 10.1094/MPMI-23-9-1217
Elling, A. A., Davis, E. L., Hussey, R. S., and Baum, T. J. (2007). Active uptake of cyst nematode parasitism proteins into the plant cell nucleus. Int. J. Parasitol. 37, 1269–1279. doi: 10.1016/j.ijpara.2007.03.012
Ellingboe, A. H. (1972). Genetics and physiology of primary infection by Erysiphe graminis. Phytopathology 62, 401–406. doi: 10.1094/Phyto-62-401
Furler, S., Paterna, J. C., Weibel, M., and Bueler, H. (2001). Recombinant AAV vectors containing the foot and mouth disease virus 2A sequence confer efficient bicistronic gene expression in cultured cells and rat substantia nigra neurons. Gene Ther. 8, 864–873. doi: 10.1038/sj.gt.3301469
Ge, X., Li, G.-J., Wang, S.-B., Zhu, H., Zhu, T., Wang, X., et al. (2007). AtNUDT7, a negative regulator of basal immunity in Arabidopsis, modulates two distinct defense response pathways and is involved in maintaining redox homeostasis. Plant Physiol. 145, 204–215. doi: 10.1104/pp.107.103374
Genovesi, V., Fornalé, S., Fry, S. C., Ruel, K., Ferrer, P., Encina, A., et al. (2008). ZmXTH1, a new xyloglucan endotransglucosylase/hydrolase in maize, affects cell wall structure and composition in Arabidopsis thaliana. J. Exp. Bot. 59, 875–889. doi: 10.1093/jxb/ern013
Goldfarb, D. S., Corbett, A. H., Mason, D. A., Harreman, M. T., and Adam, S. A. (2004). Importin α: a multipurpose nuclear-transport receptor. Trends Cell Biol. 14, 505–514. doi: 10.1016/j.tcb.2004.07.016
Gracy, J., Le-Nguyen, D., Gelly, J.-C., Kaas, Q., Heitz, A., and Chiche, L. (2008). KNOTTIN: the knottin or inhibitor cystine knot scaffold in 2007. Nucl. Acids Res. 36, D314–D319. doi: 10.1093/nar/gkm939
Hein, I., Barciszewska-Pacak, M., Hrubikova, K., Williamson, S., Dinesen, M., Soenderby, I. E., et al. (2005). Virus-Induced Gene Silencing-based functional characterization of genes associated with powdery mildew resistance in barley. Plant Physiol. 138, 2155–2164. doi: 10.1104/pp.105.062810
Hothorn, T., Bretz, F., and Westfall, P. (2008). Simultaneous inference in general parametric models. Biom. J. 50, 346–363. doi: 10.1002/bimj.200810425
Houdusse, A., and Cohen, C. (1995). Target sequence recognition by the calmodulin superfamily—Implications from light-chain binding to the regulatory domain of scallop myosin. Proc. Natl. Acad. Sci. U.S.A. 92, 10644–10647. doi: 10.1073/pnas.92.23.10644
Hsu, J. C. (1992). The factor analytic approach to simultaneous inference in the general linear model. J. Comp. Graph. Statist. 1, 151–168.
Hückelhoven, R., Dechert, C., and Kogel, K. H. (2003). Overexpression of barley BAX inhibitor 1 induces breakdown of mlo-mediated penetration resistance to Blumeria graminis. Proc. Natl. Acad. Sci. U.S.A. 100, 5555–5560. doi: 10.1073/pnas.0931464100
Hückelhoven, R., Eichmann, R., Weis, C., Hoefle, C., and Proels, R. K. (2013). Genetic loss of susceptibility: a costly route to disease resistance? Plant Pathol. 62, 56–62. doi: 10.1111/ppa.12103
Iyer, A. S., and McCouch, S. R. (2004). The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol. Plant Microbe Interact. 17, 1348–1354. doi: 10.1094/MPMI.2004.17.12.1348
Iyer-Pascuzzi, A. S., and McCouch, S. R. (2007). Recessive resistance genes and the Oryza sativa-Xanthomonas oryzae pv. oryzae pathosystem. Mol. Plant Microbe Interact. 20, 731–739. doi: 10.1094/MPMI-20-7-0731
Jackson, A. L., Bartz, S. R., Schelter, J., Kobayashi, S. V., Burchard, J., Mao, M., et al. (2003). Expression profiling reveals off-target gene regulation by RNAi. Nat. Biotech. 21, 635–637. doi: 10.1038/nbt831
Jacob, F., Vernaldi, S., and Maekawa, T. (2013). Evolution and conservation of plant NLR functions. Front. Immunol. 4:297. doi: 10.3389/fimmu.2013.00297
Jiang, C.-J., Shoji, K., Matsuki, R., Baba, A., Inagaki, N., Ban, H., et al. (2001). Molecular cloning of a novel Importin alpha homologue from rice, by which constitutive photomorphogenic 1 (COP1) nuclear localization signal (NLS)-protein Is preferentially nuclear imported. J. Biol. Chem. 276, 9322–9329. doi: 10.1074/jbc.M006430200
Jiang, G. H., Xia, Z. H., Zhou, Y. L., Wan, J., Li, D. Y., Chen, R. S., et al. (2006). Testifying the rice bacterial blight resistance gene xa5 by genetic complementation and further analyzing xa5 (Xa5) in comparison with its homolog TFIIAγ1. Mol. Genet. Genomics 275, 354–366. doi: 10.1007/s00438-005-0091-7
Johannes, L., and Popoff, V. (2008). Tracing the retrograde route in protein trafficking. Cell 135, 1175–1187. doi: 10.1016/j.cell.2008.12.009
Kelkar, A., and Dobberstein, B. (2009). Sec61β, a subunit of the Sec61 protein translocation channel at the endoplasmic reticulum, is involved in the transport of Gurken to the plasma membrane. BMC Cell Biol. 10:11. doi: 10.1186/1471-2121-10-11
Kerppola, T. K. (2006). Design and implementation of bimolecular fluorescence complementation (BiFC) assays for the visualization of protein interactions in living cells. Nat. Protoc. 1, 1278–1286. doi: 10.1038/nprot.2006.201
Kim, M. C., Panstruga, R., Elliott, C., Muller, J., Devoto, A., Yoon, H. W., et al. (2002). Calmodulin interacts with MLO protein to regulate defence against mildew in barley. Nature 416, 447–451. doi: 10.1038/416447a
Koyuncu, O. O., Perlman, D. H., and Enquist, L. W. (2013). Efficient retrograde transport of pseudorabies virus within neurons requires local protein synthesis in axons. Cell Host Microbe 13, 54–66. doi: 10.1016/j.chom.2012.10.021
Kwon, C., Bednarek, P., and Schulze-Lefert, P. (2008a). Secretory pathways in plant immune responses. Plant Physiol. 147, 1575–1583. doi: 10.1104/pp.108.121566
Kwon, C., Neu, C., Pajonk, S., Yun, H. S., Lipka, U., Humphry, M., et al. (2008b). Co-option of a default secretory pathway for plant immune responses. Nature 451, 835–840. doi: 10.1038/nature06545
Lapin, D., and Van Den Ackerveken, G. (2013). Susceptibility to plant disease: more than a failure of host immunity. Trends Plant Sci. 18, 546–554. doi: 10.1016/j.tplants.2013.05.005
Lee, W. S., Hammond-Kosack, K. E., and Kanyuka, K. (2012). Barley Stripe Mosaic Virus-mediated tools for investigating gene function in cereal plants and their pathogens: virus-induced gene silencing, host-mediated gene silencing, and virus-mediated overexpression of heterologous protein. Plant Physiol. 160, 582–590. doi: 10.1104/pp.112.203489
Lewis, J., Wan, J., Ford, R., Gong, Y., Fung, P., Nahal, H., et al. (2012). Quantitative Interactor Screening with next-generation Sequencing (QIS-Seq) identifies Arabidopsis thaliana MLO2 as a target of the Pseudomonas syringae type III effector HopZ2. BMC Genomics 13:8. doi: 10.1186/1471-2164-13-8
Liu, J., and Coaker, G. (2008). Nuclear trafficking during plant innate immunity. Mol. Plant 1, 411–422. doi: 10.1093/mp/ssn010
Macho, A. P., and Zipfel, C. (2014). Plant PRRs and the activation of innate immune signaling. Mol. Cell 54, 263–272. doi: 10.1016/j.molcel.2014.03.028
Malinovsky, F. G., Fangel, J. U., and Willats, W. G. T. (2014). The role of the cell wall in plant immunity. Front. Plant Sci. 5:178. doi: 10.3389/fpls.2014.00178
Mayer, K., Waugh, R., Langridge, P., Close, T. J., Wise, R. P., Graner, A., et al. (2012). A physical, genetic and functional sequence assembly of the barley genome. Nature 491, 711–716. doi: 10.1038/nature11543
McCullen, C. A., and Binns, A. N. (2006). Agrobacterium tumefaciens and plant cell interactions and activities required for interkingdom macromolecular transfer. Annu. Rev. Cell Dev. Biol. 22, 101–127. doi: 10.1146/annurev.cellbio.22.011105.102022
Meng, Y., Moscou, M. J., and Wise, R. P. (2009). Blufensin1 negatively impacts basal defense in response to barley powdery mildew. Plant Physiol. 149, 271–285. doi: 10.1104/pp.108.129031
Meng, Y., and Wise, R. P. (2012). HvWRKY10, HvWRKY19, and HvWRKY28 Regulate Mla-triggered immunity and basal defense to barley powdery mildew. Mol. Plant Microbe Interact. 25, 1492–1505. doi: 10.1094/MPMI-04-12-0082-R
Mentlak, T. A., Kombrink, A., Shinya, T., Ryder, L. S., Otomo, I., Saitoh, H., et al. (2012). Effector-mediated suppression of chitin-triggered immunity by Magnaporthe oryzae Is necessary for rice blast disease. Plant Cell 24, 322–335. doi: 10.1105/tpc.111.092957
Micali, C. O., Neumann, U., Grunewald, D., Panstruga, R., and O'Connell, R. (2011). Biogenesis of a specialized plant–fungal interface during host cell internalization of Golovinomyces orontii haustoria. Cell. Microbiol. 13, 210–226. doi: 10.1111/j.1462-5822.2010.01530.x
Miklis, M., Consonni, C., Bhat, R. A., Lipka, V., Schulze-Lefert, P., and Panstruga, R. (2007). Barley MLO modulates actin-dependent and actin-independent antifungal defense pathways at the cell periphery. Plant Phys. 144, 1132–1143. doi: 10.1104/pp.107.098897
Moore, C. P., Zhang, J. Z., and Hamilton, S. L. (1999). A role for cysteine 3635 for RYR1 in redox modulation and calmodulin binding. J. Biol. Chem. 274, 36831–36834. doi: 10.1074/jbc.274.52.36831
Munshi, H. G., Burks, D. J., Joyal, J. L., White, M. F., and Sacks, D. B. (1996). Ca2+ regulates calmodulin binding to IQ motifs in IRS-1. Biochemistry 35, 15883–15889. doi: 10.1021/bi962107y
Murashige, T., and Skoog, F. (1962). A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497. doi: 10.1111/j.1399-3054.1962.tb08052.x
Opalski, K. S., Schultheiss, H., Kogel, K.-H., and Hückelhoven, R. (2005). The receptor-like MLO protein and the RAC/ROP family G-protein RACB modulate actin reorganization in barley attacked by the biotrophic powdery mildew fungus Blumeria graminis f. sp. hordei. Plant J. 41, 291–303. doi: 10.1111/j.1365-313X.2004.02292.x
Osborne, A. R., Rapoport, T. A., and Van Den Berg, B. (2005). Protein translocation by the Sec61/SecY channel. Annu. Rev. Cell Dev. Biol. 21, 529–550. doi: 10.1146/annurev.cellbio.21.012704.133214
Palma, K., Zhang, Y., and Li, X. (2005). An Importin α homolog, MOS6, plays an important role in plant innate immunity. Curr. Biol. 15, 1129–1135. doi: 10.1016/j.cub.2005.05.022
Panstruga, R. (2005). Serpentine plant MLO proteins as entry portals for powdery mildew fungi. Biochem. Soc. Trans. 33, 389–392. doi: 10.1042/bst0330389
Park, E., and Rapoport, T. A. (2012). Mechanisms of Sec61/SecY-mediated protein translocation across membranes. Annu. Rev. Biophys. 41, 21–40. doi: 10.1146/annurev-biophys-050511-102312
Reddy, A. S., Ali, G. S., Celesnik, H., and Day, I. S. (2011). Coping with stresses: roles of calcium- and calcium/calmodulin-regulated gene expression. Plant Cell 23, 2010–2032. doi: 10.1105/tpc.111.084988
Restrepo, M. A., Freed, D. D., and Carrington, J. C. (1990). Nuclear transport of plant potyviral proteins. Plant Cell 2, 987–998. doi: 10.1105/tpc.2.10.987
Rhoads, A. R., and Friedberg, F. (1997). Sequence motifs for calmodulin recognition. Faseb J. 11, 331–340.
Rojo, E., and Denecke, J. (2008). What is moving in the secretory pathway of plants? Plant Physiol. 147, 1493–1503. doi: 10.1104/pp.108.124552
Rovenich, H., Boshoven, J. C., and Thomma, B. P. H. J. (2014). Filamentous pathogen effector functions: of pathogens, hosts and microbiomes. Curr. Opin. Plant Biol. 20, 96–103. doi: 10.1016/j.pbi.2014.05.001
Schmittgen, T. D., and Livak, K. J. (2008). Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108. doi: 10.1038/nprot.2008.73
Shen, Q. H., Saijo, Y., Mauch, S., Biskup, C., Bieri, S., Keller, B., et al. (2007). Nuclear activity of MLA immune receptors links isolate-specific and basal disease-resistance responses. Science 315, 1098–1103. doi: 10.1126/science.1136372
Sienaert, I., Kasri, N. N., Vanlingen, S., Parys, J. B., Callewaert, G., Missiaen, L., et al. (2002). Localization and function of a calmodulin-apocalmodulin-binding domain in the N-terminal part of the type 1 inositol 1,4,5-trisphosphate receptor. Biochem. J. 365, 269–277. doi: 10.1042/BJ20020144
Spooner, R. A., Smith, D. C., Easton, A. J., Roberts, L. M., and Lord, J. M. (2006). Retrograde transport pathways utilised by viruses and protein toxins. Virol. J. 3:26. doi: 10.1186/1743-422X-3-26
Storey, J. D., and Tibshirani, R. (2003). Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. U.S.A. 100, 9440–9445. doi: 10.1073/pnas.1530509100
Stotz, H. U., Mitrousia, G. K., De Wit, P. J. G. M., and Fitt, B. D. L. (2014). Effector-triggered defence against apoplastic fungal pathogens. Trends Plant Sci. 19, 491–500. doi: 10.1016/j.tplants.2014.04.009
Szurek, B., Marois, E., Bones, U., and Van den Ackerveken, G. (2001). Eukaryotic features of the Xanthomonas type III effector AvrBs3: protein domains involved in transcriptional activation and the interaction with nuclear import receptors from pepper. Plant J. 26, 523–534.
Tameling, W. I. L., and Baulcombe, D. C. (2007). Physical association of the NB-LRR resistance protein Rx with a ran GTPase-activating protein is required for extreme resistance to Potato virus X. Plant Cell 19, 1682–1694. doi: 10.1105/tpc.107.050880
Tameling, W. I. L., Nooijen, C., Ludwig, N., Boter, M., Slootweg, E., Goverse, A., et al. (2010). RanGAP2 mediates nucleocytoplasmic partitioning of the NB-LRR immune receptor Rx in the Solanaceae, thereby dictating Rx function. Plant Cell Online 22, 4176–4194. doi: 10.1105/tpc.110.077461
Tufan, H. A., Stefanato, F. L., McGrann, G. R., Maccormack, R., and Boyd, L. A. (2011). The Barley stripe mosaic virus system used for virus-induced gene silencing in cereals differentially affects susceptibility to fungal pathogens in wheat. J. Plant Physiol. 168, 990–994. doi: 10.1016/j.jplph.2010.11.019
Van Der Linde, K., Hemetsberger, C., Kastner, C., Kaschani, F., Van Der Hoorn, R. A. L., Kumlehn, J., et al. (2012). A maize cystatin suppresses host Immunity by inhibiting apoplastic cysteine proteases. Plant Cell 24, 1285–1300. doi: 10.1105/tpc.111.093732
Van Schie, C. C. N., and Takken, F. L. W. (2014). Susceptibility genes 101: how to be a good host. Annu. Rev. Phytopathol. 52, 551–581. doi: 10.1146/annurev-phyto-102313-045854
Vogel, J. P., Raab, T. K., Schiff, C., and Somerville, S. C. (2002). PMR6, a pectate lyase-like gene required for powdery mildew susceptibility in Arabidopsis. Plant Cell 14, 2095–2106. doi: 10.1105/tpc.003509
Vogel, J. P., Raab, T. K., Somerville, C. R., and Somerville, S. C. (2004). Mutations in PMR5 result in powdery mildew resistance and altered cell wall composition. Plant J. 40, 968–978. doi: 10.1111/j.1365-313X.2004.02264.x
Wang, D., and Dong, X. N. (2011). A highway for war and peace: the secretory pathway in plant-microbe interactions. Mol. Plant 4, 581–587. doi: 10.1093/mp/ssr053
Wang, D., Weaver, N. D., Kesarwani, M., and Dong, X. N. (2005). Induction of protein secretory pathway is required for systemic acquired resistance. Science 308, 1036–1040. doi: 10.1126/science.1108791
Wang, Q. L., Chen, B., Liu, P., Zheng, M. Z., Wang, Y. Q., Cui, S. J., et al. (2009). Calmodulin binds to extracellular sites on the plasma membrane of plant cells and elicits a rise in intracellular calcium concentration. J. Biol. Chem. 284, 12000–12007. doi: 10.1074/jbc.M808028200
Whigham, E., Qi, S., Mistry, D., Surana, P., Xu, R., Fuerst, G. S., et al. (2015). Broadly conserved fungal effector BEC1019 suppresses host cell death and enhances pathogen virulence in powdery mildew of barley (Hordeum vulgare L.). Mol. Plant Microbe Interact. doi: 10.1094/MPMI-02-15-0027-FI. [Epub ahead of print].
Wirthmueller, L., Zhang, Y., Jones, J. D. G., and Parker, J. E. (2007). Nuclear accumulation of the Arabidopsis immune receptor RPS4 is necessary for triggering EDS1-dependent defense. Curr. Biol. 17, 2023–2029. doi: 10.1016/j.cub.2007.10.042
Wise, R. P., Moscou, M. J., Bogdanove, A. J., and Whitham, S. A. (2007). Transcript profiling in host-pathogen interactions. Ann. Rev. Phytopathol. 45, 329–369.
Wolfinger, R. D., Gibson, G., Wolfinger, E. D., Bennett, L., Hamadeh, H., Bushel, P., et al. (2001). Assessing gene significance from cDNA microarray expression data via mixed models. J. Comput. Biol. 8, 625–637. doi: 10.1089/106652701753307520
Xi, L., Moscou, M. J., Meng, Y., Xu, W., Caldo, R. A., Shaver, M., et al. (2009). Transcript-based cloning of RRP46, a regulator of rRNA processing and R gene-independent cell death in barley-powdery mildew interactions. Plant Cell 21, 3280–3295. doi: 10.1105/tpc.109.066167
Xu, W., Meng, Y., and Wise, R. P. (2014). Mla- and Rom1-mediated control of microRNA398 and chloroplast copper/zinc superoxide dismutase regulates cell death in response to the barley powdery mildew fungus. New Phytol. 201, 1396–1412. doi: 10.1111/nph.12598
Xu, X., Chen, C., Fan, B., and Chen, Z. (2006). Physical and functional interactions between pathogen-induced Arabidopsis WRKY18, WRKY40, and WRKY60 transcription factors. Plant Cell 18, 1310–1326. doi: 10.1105/tpc.105.037523
Yamniuk, A. P., and Vogel, H. J. (2004). Calmodulin's flexibility allows for promiscuity in its interactions with target proteins and peptides. Mol. Biotechnol. 27, 33–57. doi: 10.1385/MB:27:1:33
Yasuhara, N., Shibazaki, N., Tanaka, S., Nagai, M., Kamikawa, Y., Oe, S., et al. (2007). Triggering neural differentiation of ES cells by subtype switching of importin-α. Nat. Cell Biol. 9, 72–79. doi: 10.1038/ncb1521
Yasuhara, N., Yamagishi, R., Arai, Y., Mehmood, R., Kimoto, C., Fujita, T., et al. (2013). Importin α subtypes determine differential transcription factor localization in embryonic stem cells maintenance. Dev. Cell 26, 123–135. doi: 10.1016/j.devcel.2013.06.022
Zhang, C., Bradshaw, J. D., Whitham, S. A., and Hill, J. H. (2010). The development of an efficient multipurpose Bean pod mottle virus viral vector set for foreign gene expression and RNA silencing. Plant Physiol. 153, 52–65. doi: 10.1104/pp.109.151639
Keywords: knottin, nuclear import, secretory pathway, powdery mildew, calmodulin, BSMV-VIGS, gene expression, negative regulator
Citation: Xu W, Meng Y, Surana P, Fuerst G, Nettleton D and Wise RP (2015) The knottin-like Blufensin family regulates genes involved in nuclear import and the secretory pathway in barley-powdery mildew interactions. Front. Plant Sci. 6:409. doi: 10.3389/fpls.2015.00409
Received: 03 October 2014; Accepted: 21 May 2015;
Published: 04 June 2015.
Edited by:
Xin Li, University of British Columbia, CanadaReviewed by:
Erik Limpens, Wageningen University, NetherlandsHugo Germain, Université du Québec à Trois-Rivières, Canada
Copyright © 2015 Xu, Meng, Surana, Fuerst, Nettleton and Wise. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Roger P. Wise, Corn Insects and Crop Genetics Research Unit, U.S. Department of Agriculture-Agricultural Research Service and Department of Plant Pathology and Microbiology, Iowa State University, 351 Bessey Hall, Ames, IA 50011–1020, USA,cm9nZXIud2lzZUBhcnMudXNkYS5nb3Y=
†Present Address: Yan Meng, Department of Agriculture, Alcorn State University, Lorman, USA
‡These authors have contributed equally to this work.
 Weihui Xu
Weihui Xu Yan Meng1†‡
Yan Meng1†‡ Priyanka Surana
Priyanka Surana Dan Nettleton
Dan Nettleton Roger P. Wise
Roger P. Wise