- 1Energy, Mining and Environment, National Research Council Canada, Montréal, QC, Canada
- 2Institut de Recherche en Biologie Végétale – Jardin Botanique de Montréal and Université de Montréal, Montréal, QC, Canada
The goal of microbiome engineering is to manipulate the microbiome toward a certain type of community that will optimize plant functions of interest. For instance, in crop production the goal is to reduce disease susceptibility, increase nutrient availability increase abiotic stress tolerance and increase crop yields. Various approaches can be devised to engineer the plant–microbiome, but one particularly promising approach is to take advantage of naturally evolved plant–microbiome communication channels. This is, however, very challenging as the understanding of the plant–microbiome communication is still mostly rudimentary and plant–microbiome interactions varies between crops species (and even cultivars), between individual members of the microbiome and with environmental conditions. In each individual case, many aspects of the plant–microorganisms relationship should be thoroughly scrutinized. In this article we summarize some of the existing plant–microbiome engineering studies and point out potential avenues for further research.
Introduction
Virtually every plant part is colonized by microorganisms, including bacteria, archaea, fungi, collectively designated as the plant–microbiome or phytomicrobiome. Depending on the plant part it colonizes, the phytomicrobiome is often referred to as endophytic (inside plant parts), epiphytic (on aboveground plant parts), or rhizospheric (in the soil closely associated to the roots) (Kowalchuk et al., 2010; Lakshmanan et al., 2014). Microorganisms are a key component of the plant, often inextricable from their host and the plant–microbiome is thought to function as a metaorganism or holobiont (Bosch and McFall-Ngai, 2011; Vandenkoornhuyse et al., 2015). The biomass and composition of the microbiome strongly affects the interactions between plants and their environments (Ryan et al., 2009). The rhizosphere harbors a largely increased bacterial abundance and activity, not only as compared to other plant compartments (Smalla et al., 2001; Kowalchuk et al., 2002), but also when compared to bulk soil. However, bacterial diversity in the rhizosphere is generally lower than in the bulk soil (Marilley and Aragno, 1999) and microbial community composition is very different (Smalla et al., 2001; Kowalchuk et al., 2002; Griffiths et al., 2006; Kielak et al., 2008; Bulgarelli et al., 2012; Peiffer et al., 2013), suggesting a strongly selective environment. As the microbial density, diversity, and activity in the endosphere (the microbial habitat inside both above- and belowground plant organs) and phyllosphere (the aboveground plant surfaces) are generally lower than in the rhizosphere, the focus of this contribution will be mainly on the rhizosphere.
There is ample evidence that shows that the plant–microbe relationship is critical to health, productivity and the overall condition of the plant (Baudoin et al., 2003; Chaparro et al., 2012, 2014; Marasco et al., 2012; Adesemoye and Egamberdieva, 2013; Ziegler et al., 2013; Glick, 2014). There are different kinds of interactions between plants and microbes, spanning the whole spectrum from beneficial to pathogenic, and the outcome of the interaction between a plant and a microbe can vary among this spectrum depending on plant species, nutrient conditions, etc. (Figure 1). The goal of plant–microbiome engineering is to push this interaction toward enhanced beneficial outcomes for the plant. Many microbially mediated functions are important to enhance beneficial outcome, including nutrient cycling, mineralization of soil organic matter, induction of disease resistance and response to abiotic stresses such as drought and salinity (Marasco et al., 2012; Zolla et al., 2013). The plant–microbiome interactions are complex and often depend on plant species/cultivar, soil type and environmental conditions such as biotic/abiotic stress, climatic conditions, and anthropogenic effects. Different soils as well as different environmental stresses (e.g., nutrient deficiencies, metal toxicity, pathogen attack, etc.) have been shown to trigger plant-species-dependent physiological responses and consequently different exudation patterns (Bais et al., 2006; Oburger et al., 2013). Microbes in the rhizosphere can also influence the plant exudation, as for example, when antimicrobial-resistant Pseudomonas block the production of plant antimicrobial compounds (Hartmann et al., 2009). One interesting avenue for rhizosphere microbiome engineering is to harness these variations in exudation patterns to enhance the beneficial rhizosphere microbiome.
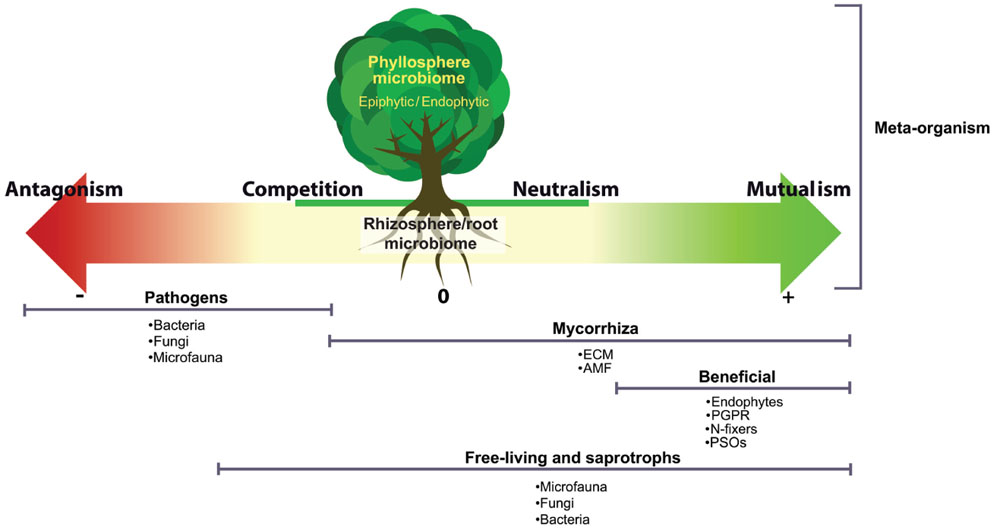
FIGURE 1. The different interactions taking place within the plant–microbiome meta-organism. A vast spectrum of microorganisms are involved in these interactions: ectomycorrhiza (ECM), arbuscular mycorrhizal fungi (AMF), plant growth promoting rhizobacteria (PGPR), phosphate-solubilizing organisms (PSOs), endophytes, epiphytes, and microfaunal organisms.
Signaling in the Rhizosphere
A variety of direct and indirect interactions take place in the rhizosphere such as plant–plant, microbe–microbe, and plant–microbe, as well as the interaction with the other eukaryotic micro-, meso-, and macro-soil inhabitants (Tarkka et al., 2008; De-la-Peña et al., 2012). In view of the complexity of these interactions, knowledge of the chemical communication between all members is essential to unravel how microbial populations coordinate their behavior and interact with the plant roots. Numerous literature reviews have addressed the many different molecules and mechanisms that coordinates the establishment of specific symbiotic interactions in the rhizosphere with the potential to enhance plant growth and productivity (Mabood et al., 2008; Pieterse et al., 2009; Berendsen et al., 2012; Miller and Oldroyd, 2012; Sugiyama and Yazaki, 2012; Bakker et al., 2013; Morel and Castro-Sowinski, 2013; Oldroyd, 2013). However, the understanding of the interactions between the plants and the microbiome as a whole is still rudimentary as the diversity of organisms, molecules, and mechanisms of interaction involved is staggering. Nevertheless, the signaling compounds that make part of this complex rhizosphere interaction have the potential to improve plant functions of interest if understood and harnessed.
Plants have been found to release 5–20% of net photosynthetically fixed C into the rhizosphere (Marschner, 1995). These rhizodeposits include inorganic (CO2 from cell respiration and H+ efflux) and a variety of complex organic compounds like sloughed-off cells and tissue, intact root border cells, mucilage (polysaccharides) and proteins, all of them classified as high molecular weight compounds. Also, part of the rhizodeposits are the insoluble and soluble low molecular weight (LMW) organic compounds, collectively known as root exudates, which are actively or passively released by growing roots. Root exudates can be classified in different classes such as sugars, amino acids, and amides, organic acids, as well as aromatic and phenolic acids (Bais et al., 2006, 2008; De-la-Peña et al., 2012; Oburger et al., 2013; Zhang et al., 2015). This complex cocktail of root-secreted molecules mediate the interactions occurring in the rhizosphere (Bakker et al., 2012; Berendsen et al., 2012; Lakshmanan et al., 2014; Qiu et al., 2014) and acts as chemical attractants and repellants to shape the root microbiome (Walker et al., 2003; Berendsen et al., 2012; Ellouze et al., 2012). From the plant point of view, the goal of shaping the rhizosphere microbiome is to attract preferred partners like plant growth promoting microorganisms through the exudation of specific carbon compounds that can be used as feed and to deter pathogens or unwanted competitors for nutrients through the exudation of antimicrobial compounds such as volatiles or proteins (Bais et al., 2006; Lioussanne et al., 2008; Badri et al., 2009; Hartmann et al., 2009; De-la-Peña et al., 2012; Dangl et al., 2013). Plant exudates are also involved in coping with herbivores, encouraging beneficial symbioses, changing the chemical and physical properties of the soil, and inhibiting the growth of competing plant species (Ping and Boland, 2004; Badri et al., 2009; Morel and Castro-Sowinski, 2013).
The quality and amount of root exudates are highly dynamic in time and space and they depend on the plant species/cultivars, the physiological stage of the plant (Chaparro et al., 2013, 2014), presence or absence of neighbors, plant nutritional status, mechanical impedance (Bengough and Mullins, 1990), sorption characteristics of the soil, and the microbial activity in the rhizosphere (De-la-Peña et al., 2012; Oburger et al., 2013). Plant productivity, nutrient allocation, and tissue chemistry can also vary significantly depending on the identity of neighboring individuals, suggesting that the effects of a given plant host on the soil microbiome may be substantially mediated by the community context of that host (Bakker et al., 2012, 2013). Although very complex and still not well understood, exudation has therefore the potential to highly influence plant performance, health, and competiveness. Some studies have started analyzing the composition of plant root exudates (Phillips et al., 2008; Oburger et al., 2013; Ziegler et al., 2013), but the diversity of the compounds involved and the complexity of the soil matrix makes comprehensive analysis difficult to perform.
Many microorganisms also secrete signaling compounds in the rhizosphere. According to their functions and characteristics, these compounds have been categorized into: phytohormones, extracellular enzymes, organic acids, surface factors [compounds recognized by the host plant that activate an immune response via high-affinity cell surface pattern-recognition receptors (PRR), e.g., flagellins and lipopolysaccharides in Pseudomonas (Ping and Boland, 2004; Dangl et al., 2013)], antibiotics and volatile signals. Plant-associated bacteria produce and utilize diffusible quorum sensing (QS) molecules (e.g., N-acyl-homoserine lactones, AHLs) to signal to each other and to regulate their gene expression (Berendsen et al., 2012). Bacterially produced AHLs have been shown to affect root development of Arabidopsis (Ortiz-Castro et al., 2008) and have been suggested to elicit a phenomenon known as induced systemic resistance (ISR) which allows the plants to endure pathogen attacks that could be lethal without the presence of these bacterial factors. The effect of this mechanism is systemic, e.g., root inoculation with many different plant growth promoting rhizobacteria (PGPR) such as Pseudomonas, Burkholderia, and Bacillus sp. results in the entire plant being non-susceptible to pathogens (Schuhegger et al., 2006; Choudhary et al., 2007; Tarkka et al., 2008), further highlighting the importance of AHLs in cross-kingdom signaling in the rhizosphere. Plant can also exploit this microbial communication system to manipulate gene expression in their associated microbial communities. For instance, some plant-associated bacteria have LuxR-like proteins that are stimulated by plant signals (Soto et al., 2006; Ferluga and Venturi, 2009). Certain bacteria have the capacity to quench signals by degrading various plant- and microbial-produced compounds in the rhizosphere such as quorum sensing signals (Tarkka et al., 2008) and other compounds, like ethylene, that might have negatively affected plants (Bais et al., 2006).
Many of the plant response implicate the intervention of the plant immune (system systemic acquired resistance or SAR) consisting of two interconnected levels of receptors, one outside and one inside the plant cell, that govern recognition of microbes and response to infections. The first level of the plant immune system is governed by extracellular surface PRRs that are activated by recognition of evolutionarily conserved pathogen or microbial-associated molecular patterns (PAMPs or MAMPs). Activation of PRRs leads to intracellular signaling, transcriptional reprogramming, and biosynthesis of a complex output response. This response limits microbial colonization (Dangl et al., 2013) and shapes the soil microbial community in the rhizosphere by selective feeding of beneficial microorganisms and by excreting substances with antimicrobial potential such as root volatiles or root proteins which acts as the primer barriers of plant defense (Bais et al., 2006; Badri et al., 2009; Hartmann et al., 2009; De-la-Peña et al., 2012; Dangl et al., 2013).
How Can we Engineer the Rhizosphere Microbiome?
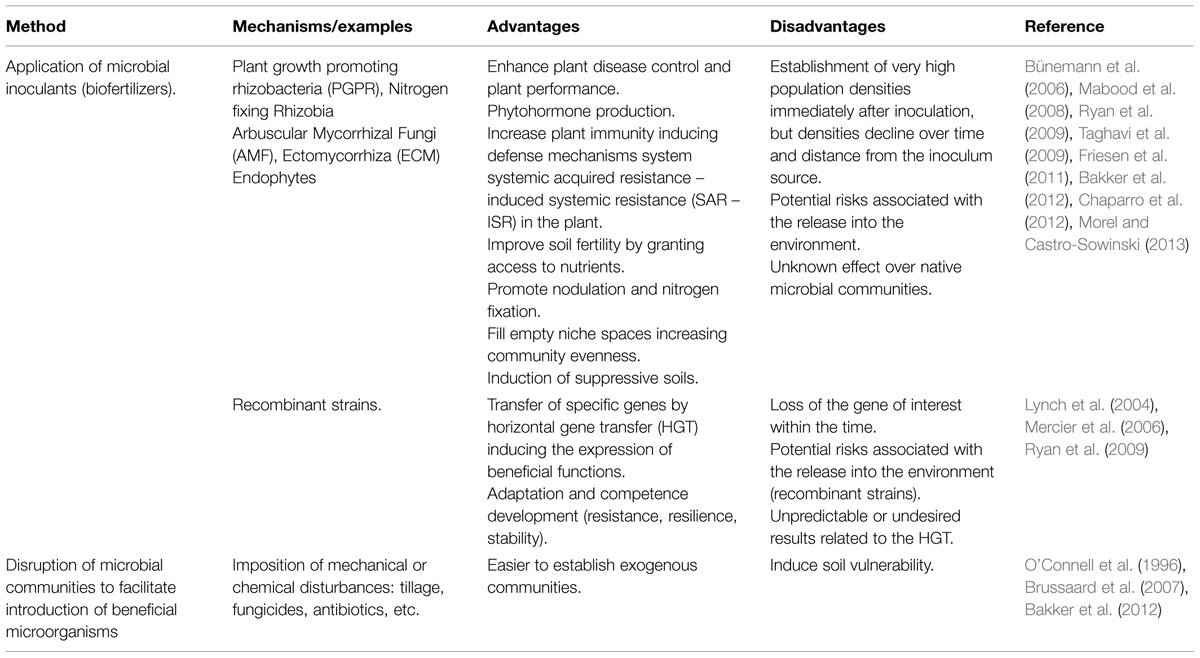
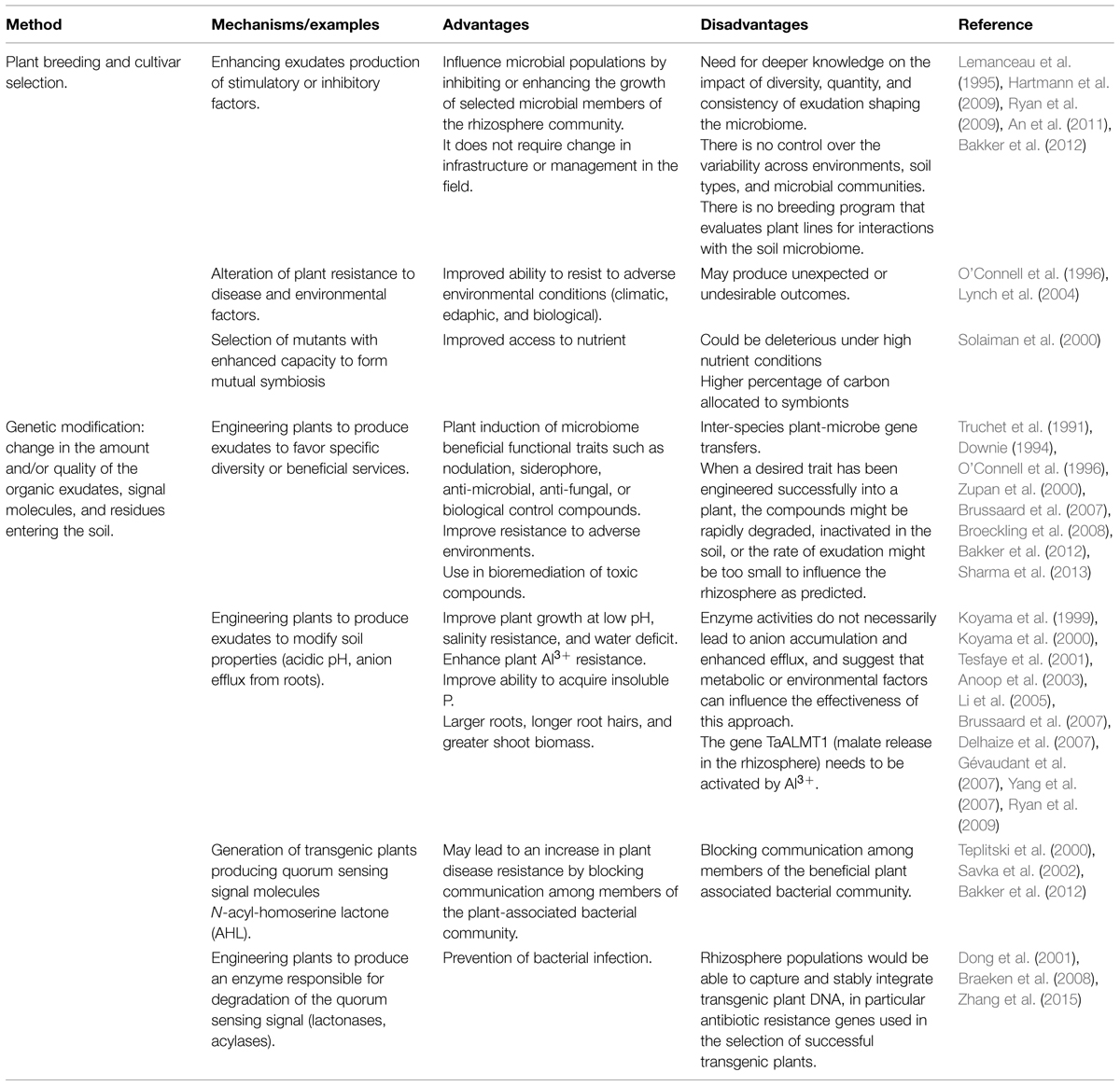
The efforts to elucidate rhizosphere interactions have often been directed to the potential of single plant root exudates to affect single bacteria or fungi. The clear limitation of this type of approach is the removal of the organism from any context that would give relevance to interspecies interactions. The high diversity of the root- and microbe-secreted molecules involved in rhizosphere interactions suggests that studying the direct influence of a single compound on the microbiome might be impossible or not realistic in nature (Ziegler et al., 2013). Rhizosphere engineering in the environment is still a major challenge even though some studies showed some promising results (Tables 1–3). A vast diversity of approaches based on inter-kingdom communication has been utilized in laboratory, greenhouse, and field experiments in order to favor beneficial services to the plants while minimizing inputs requirements. Three potential routes are suggested below in Table 1, the microbiome approach, Table 2, the plant approach, and Table 3, the meta-organism approach. In these tables, we review more generally rhizosphere engineering efforts, but below we will more specifically focus on rhizosphere microbiome engineering studies that took advantage of signalisation channels.
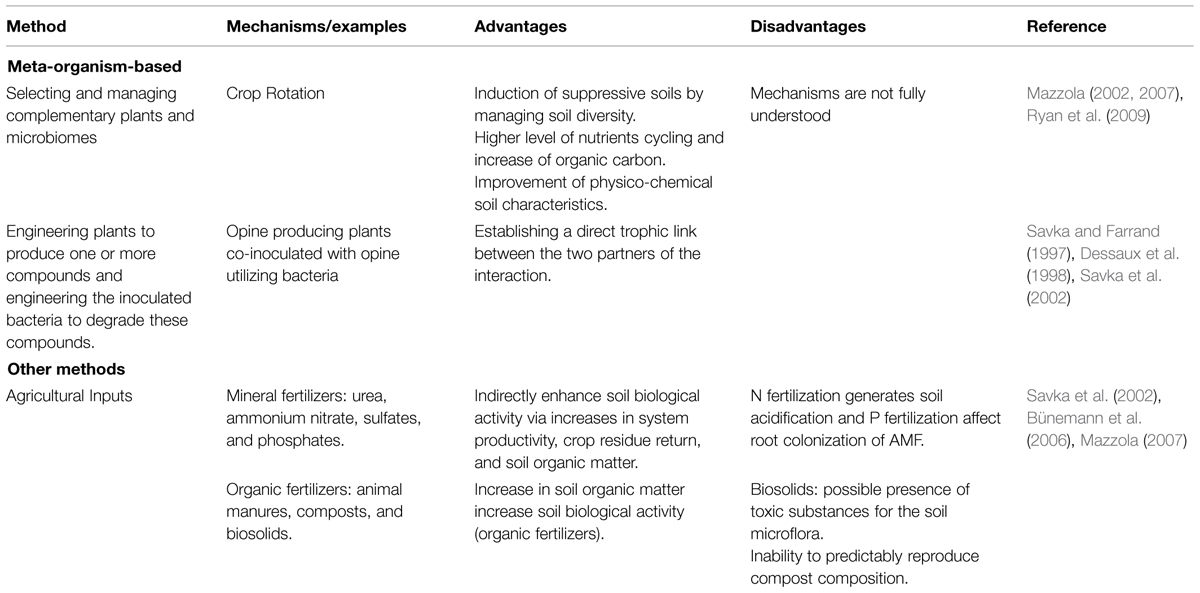
TABLE 3. Selected meta-organism-based methods and other complementary methods used to engineer the rhizosphere microbiome.
The Microbiome Route
Many of the bacteria in the rhizosphere are currently unable to grow in the laboratory and culture-based methods are often inadequate for qualitative analysis of the rhizosphere microbiome. As a consequence, culture-independent approaches such as metagenomics, metatranscriptomics, metaproteomics, and metabolomics have been the approaches of choice when investigating the rhizosphere microbiome (De-la-Peña et al., 2012; Bell et al., 2014a,b; Yergeau et al., 2014; Zhang et al., 2015). However, many rhizosphere microbiome engineering approaches require having microbial isolates at hand, and further efforts to increase the cultivability of rhizosphere microorganisms will be needed. Even though cultured microorganisms show certain functional capacity of their own, it is not clear yet how they behave once they are introduced in a new environmental niche as in some cases they have been shown to be out-competed by the indigenous microbial population (Ryan et al., 2009). The persistence and functionality of these isolates after inoculation need to be further assessed in order to ascertain positive impacts when used as a strategy to manipulate the rhizosphere microbiome (Stefani et al., 2015). Colonization and dominance of specific microbial species in the rhizosphere is critical for both pathogenic and beneficial soil microbes and will have an impact on disease incidence. Although a general increase in the abundance of microbes is always noted in the rhizosphere as compared to bulk soil, the community structure and functional consequences associated to this increase are poorly understood (Bais et al., 2006; Bakker et al., 2012). An increase in the abundance, activity, or diversity of soil organisms is generally viewed as positive (Bünemann et al., 2006), maximizing overall microbial activity or niche saturation which results in competitive exclusion of pathogens, higher levels of nutrient cycling and increased community stability (Figure 2). In that regard, the main microbial strategy to enhance the rhizosphere microbiome include the direct inoculation of microorganisms, focussing on co-inoculation with several strains or mixed cultures of arbuscular mycorrhizal fungi (AMF), ectomycorrhizal fungi (ECM), PGPR and endophytes, enabling combined niche exploitation, cross-feeding, enhancement of one organism’s colonization ability, modulating plant growth, achieving niche saturation and competitive exclusion of pathogens (Ping and Boland, 2004; Bünemann et al., 2006; Ryan et al., 2009). Equally important as the recruitment of the adequate microbiome for the plant, is the activation of its specific functions. Quorum sensing (QS) is the mechanism used to regulate distinct microbial activities (biofilm formation, virulence, symbiosis, antibiotic production, conjugation) and is essential for within-species communication as well as for the crosstalk between species which defines if the relationship with the host plant is synergic or antagonist (Soto et al., 2006; Hartmann et al., 2009; Straight and Kolter, 2009).
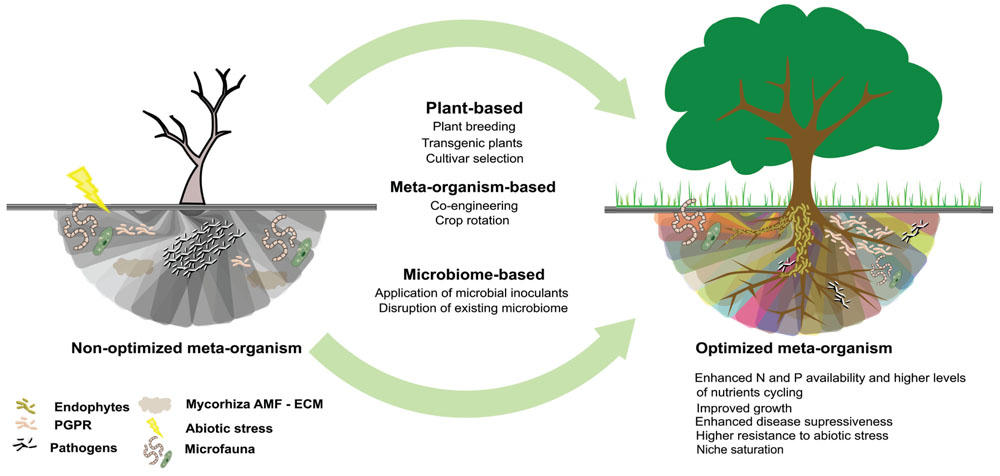
FIGURE 2. Different approaches to rhizosphere microbiome engineering used to bring the microbiome from a low diversity and vulnerable state, with limited functions and productivity to a diverse and resilient state with high functional redundancy and consistent functioning across variable environments and increased resistance to pathogen invasion.
Enhancing the rhizosphere and root endosphere microbiome often leads to an improvement of beneficial plant functional traits as the microbes are able to expand the plant biochemical capabilities or alter existing pathways (Table 1). For instance, PGPR promote plant growth by acting as biofertilizer and entering in symbiosis with their host plants, endosymbiotic rhizobia (Bradirhizobium, Mesorhizobium, Rhizobium, Sinorhizobium, etc) and free-living diazotrophs (Azospirillum, Acetobacter, Herbaspirillum, Azoarcus, and Azotobacter, etc) fix atmospheric nitrogen, mycorrhiza recover N from NH4 and NO3 (Toussaint et al., 2004; Adesemoye et al., 2008; Mabood et al., 2008; Ryan et al., 2009), and phosphate solubilizing bacteria, AMF, ECM, and siderophore producers increase availability of many nutrients such as phosphorous (P), iron (Fe), cooper (Cu), Cadmiun (Cd), and zinc (Zn) (Savka et al., 2002; Mabood et al., 2008). Rhizobacteria also act as biocontrol agents, for instance, Pseudomonads, Bacillus, and Streptomyces produce antibiotics (DAPG, phenazine, hydrogene cyanide, oligomycin, etc), bacteriocines (Nisin) and antifungal compounds (pyoluteorin, phenazines, and phoroglucinols; Ping and Boland, 2004; Mabood et al., 2008; Paulin et al., 2009; Ryan et al., 2009).
Another strategy is to induce plant metabolic activities by modulating phytohormone synthesis by microbes (phytostimulation). As we mention before, microorganisms are capable of altering plant physiological pathways since they are able to produce all plant hormone identified to date (Friesen et al., 2011) or synthetize compounds that can mimic their actions (Table 1). Using microbes to exploit the plant hormonal system could improve plant growth and root development, leading to higher yields. Phytohormones produced by microorganisms such as auxins (indole-3-acetic acid), gibberelins and citokinins mirror the action of jasmonic acid (JA) which is critical for plant defense against herbivory, plant responses to poor environmental conditions (abiotic and biotic stress tolerance), regulation of signals exchange and nodulation (Hause and Schaarschmidt, 2009) and is involved in the signaling pathway against necrotrophic pathogens (Stein et al., 2008). Crosstalk mediated by salicylic acid, JA and ethylene activate plant SAR and induce systemic resistance (ISR) reducing phytotoxic microbial communities (Ping and Boland, 2004; van Loon et al., 2006; Mabood et al., 2008). Production of 1-aminocyclopropane-1-carboxylate (ACC) deaminase by rhizosphere microorganism is another characteristic that can have a high impact on plant health as this enzyme degrades the ethylene precursor ACC, thereby reducing ethylene levels in the plant. When present in high concentrations ethylene can lead to plant growth inhibition or even death, but in lower amounts ethylene can also help the plant respond to a wide range of environmental stresses (Ryan et al., 2009; Glick, 2014).
Inoculation of recombinant strains is another strategy to enhance plant performance. In some cases, recombinant strains can resolve problems related to rapid decrease in population density and short persistence (Ryan et al., 2009), and as reported by Taghavi et al. (2005), could result in the enhancement of many members of the endogenous population by the transmission of genetic information via horizontal gene transfer (HGT). Even though very promising, the release of recombinant strains to the environment needs to be thoroughly assessed in order to evaluate the potential risks associated. To maximize the effects of inoculations, the disruption of existing microbial communities by fungicide application, crop rotation or tilling can be used to favor the selection of the appropriate microbiome for specific crops in order to establish biological functions in the rhizosphere (O’Connell et al., 1996; Savka et al., 2002; Bakker et al., 2012; Table 1). It is also essential to understand the evolution, organization, and structure of the rhizosphere microbial community throughout plant developmental stages and the way they naturally manipulate the composition of the rhizosphere microbiome, promoting, for instance, suppressive soils (Baker and Cook, 1974; Ryan et al., 2009; Mendes et al., 2011) or particular microbial functions in the rhizosphere like nutrient cycling or resistance to abiotic stress. In addition to specific microbial taxa or functions, community-wide characteristics can also be the target of microbiome engineering efforts as rhizosphere microbiome richness and evenness is linked to higher resilience to disruption, stress, and diseases. Increased microbial richness often results in greater community-level trait diversity and/or functional redundancy, which leads to more consistent functioning across variable environments (Loreau et al., 2001) Because rare members of the microbiome may be unable to effectively perform important functions, high evenness of the microbiome is also very important (Figure 2; Bünemann et al., 2006; Badri and Vivanco, 2009; Bakker et al., 2012; Qiu et al., 2014).
The Plant Route
Plant-based strategies to improve plant productivity through the selection of a more adapted microbiome include the manipulation of plants characteristics of interest mainly by two different approaches: plant breeding (cultivar selection) and specific genetic modifications (Table 2). Using plant breeding to select for a specific microbiome is an interesting avenue, as the technique has mainly focused on improving yields, plant resistance to pests/diseases and other plant physiological traits (Ryan et al., 2009). When microbiome selection was included in plant breeding programs, very specific functions or taxa were targeted. For example, Neal et al. (1970, 1973) used chromosomal substitution between two lines of wheat to improve resistance to root-rot while preserving beneficial populations of rhizosphere bacteria, and Mazzola (2002) compared wheat cultivars for their capacity to stimulate disease suppression by enhancing populations of specific antagonist (Pseudomonads) against Rhizoctonia solani.
Choosing a naturally occurring plant species or cultivar with a high capacity to recruit a beneficial microbiome or to promote the “suppressiveness” of soils is an alternative option that has been explored (Neal et al., 1973; Mazzola, 2002; Bakker et al., 2012). For instance, in the rhizosphere of plants growing in contaminated soils, the host plant exudes specialized antimicrobials and signaling molecules (i.e., flavonoids, salicylic acid, and phytoalexins), carbon and nitrogen compounds that promotes the expression of hydrocarbon degradation genes such as the genes coding for alkane hydroxylases (responsible for the aerobic degradation of aliphatic hydrocarbons) and several genes coding for enzymes implicated in the metabolism of aromatic compounds (Yergeau et al., 2014; Pagé et al., 2015). This recruitment was shown to be cultivar-specific, with native cultivar having an increased capacity to recruit beneficial ECM when growing in highly contaminated soils (Bell et al., 2014a), suggesting that native cultivars can better communicate with indigenous soil microorganisms. Accumulating evidence supports the feasibility of creating biotic soil environments that promote root health using selected plant genotypes. Ellouze et al. (2013) showed in the semiarid grasslands of North America, certain chickpea cultivars can select a more beneficial microbiome for the subsequent wheat plants and were associated with the antagonist species Penicillium canescens.
Using plants as selective agents to enrich beneficial microbial functions in soil implies the inclusion of other variables such as soil type and properties: different soil types not only shape the microbial communities, but also impact plant physiology, which in turn could alter interactions with soil microbes. The creation of genetically modified plants with enhanced ability to harbor particular exudation patterns to change soil properties have already been investigated (Table 2). Li et al. (2005); Gévaudant et al. (2007), and Yang et al. (2007) have worked to manipulate rhizosphere pH using transgenic lines of Nicotiana tabacum and Arabidopsis plants, transformed to over express a modified H+ATP-ase protein (PMA4 in tobacco and AVP1 pyrophosphatase in Arabidopsis) generating phenotypes such as increased H+-efflux from roots, more acidic rhizosphere, improved growth at low pH, improved salinity resistance (tobacco lines), plant mineral nutrition (P mineralization), and exhibiting enhanced resistance to water deficits (AVP1).
Plants can also be genetically modified to alter soil organic anion efflux and transportation from roots by (1) engineering plants with a greater capacity to synthesize organic anions or (2) engineering plants with a greater capacity to transport organic anions out of the cell. The organic anions malate and citrate have been studied as they are commonly release in response to nutrient deficiency and mineral stress. Koyama et al. (1999) and Koyama et al. (2000) reported that transgenic plants with higher ability to excrete citrate from the roots grew better on P-limited soil than the wild type, suggesting crop plants with an enhanced ability to use Al-phosphate and therefore an enhanced ability to grow in acid soils and superior Al tolerance. In order to address the toxicity of the Al3+ in acidic soils, a very common problem in agriculture, Tesfaye et al. (2001), Anoop et al. (2003), and Delhaize et al. (2007) have also reported the use of transgenic plants (Medicago sativa, Brassica napus, and Hordeum vulgare) expressing genes coding for ALMT (Al3+-activated malate transporter) and MATE (multi-drug and toxin extrusion) membrane proteins, a strategy to improve the P efficiency of plants and Al3+ resistance (Table 2).
Quorum sensing has been targeted by creating transgenic plants that would be able to mimic or interrupt bacterial QS signals by producing enzymes responsible for their degradation (acylases and lactonases). These modifications allowed these plants to defend themselves more efficiently against some pathogenic bacteria (Savka et al., 2002; Braeken et al., 2008; Ryan et al., 2009; Bakker et al., 2012). Many other studies have focused on the genetic manipulation of plants in order to modify the key exudates to favor the establishment of the desired plant–microbiome (Table 2). Genetic engineering provides unique opportunities to modulate plant–microbe signaling, to diversify exudation, to encourage diverse microbiomes or to stimulate beneficial microbial functions in the rhizosphere. However, despite these efforts, large-scale breeding/genetic improvement programs rarely take into account the plant–microbiome signaling channels during the development of new plant lines.
The Meta-Organism Route
The microbiome and the plant are highly dependent on each other as the microbiome contribute a significant portion of the secondary genome of the host plant, highlighting that the plant an its microbiome might function as a meta-organism or holobiont (Lakshmanan et al., 2014). Taking into account the meta-organism and trying to optimize the whole system instead of each of the part separately is a promising avenue for rhizosphere microbiome engineering. This is the case of the “opine concept” that combined engineering plants to produce specific exudates together with the inoculation of engineered microbes that are able to degrade this substrate, resulting in the colonization of the rhizosphere by a specific population (Table 3). It was also observed that opine production by transgenic plants led, in the long term, to the selection of bacterial populations adapted to the rhizosphere that can maintain themselves at high concentrations, even after removal of the transgenic plants (O’Connell et al., 1996; Savka et al., 2002; Ryan et al., 2009). These strategies (using specific metabolic resources) are highly specific, focusing on interactions between, for example, opine-producing plants, and members of the microbiome responsible for functions such as nodulation and N-fixation. However, this “opine” approach does not take into account others species that could have important functions or fill niche to reduce pathogen vulnerability.
In order to amplify the spectrum of diversity and ecological services to the crops, another strategy to shift rhizosphere microbiome is crop rotation. This strategy could be optimized by taking into account the whole plant–microbiome metaorganism. Indeed, plants are cultured in turns bringing their associated microbiome and generating beneficial allopathy. Crop history and cultivar selection stimulate specific rhizobacterial populations that complements each other developing a beneficial synergy between the cultures. Sturz et al. (1998) reported evidence for the role of bacterial endophytes resulting from the intercrop alternation of red clover and potato promoting plant growth and yield in the potato crops. Mazzola (2007) stated that most common and effective scheme to modify the rhizosphere has been the use of crop rotation. As increased plant diversity can enhance microbial community biomass, mixed cropping systems will generate a more diverse microbial community and thus should be more resilient to pathogen invasion. Disease control is achieved as the plant host for certain pathogen is absent resulting in the diminished viability of this pathogen. Some other advantages of these rhizosphere microbiome engineering approaches are an increase of soil organic carbon, a higher level of nutrients cycling and an improvement of physico-chemical soil characteristics (O’Connell et al., 1996; Sturz et al., 1998; Bakker et al., 2012).
Phytoremediation is another application where harnessing the plant–microbiome holobiont could significantly improve processes (Bell et al., 2014b). In the rhizosphere of contaminated soils, microbes increase the recycling and dissolving of mineral nutrients and the synthesis of amino acids, vitamins, auxins, and gibberellins that stimulate plant growth. These highly competitive populations seem to be selected by the host plant via exudation of specialized antimicrobials and signaling molecules (e.g., flavonoids, salicylic acid, and phytoalexins), carbon and nitrogen compounds, resulting in the degradation or transformation of contaminants due to both increased microbial activity and plant intervention (Marihal and Jagadeesh, 2013; Yergeau et al., 2014). Microorganisms also facilitate the uptake of contaminants and plant resistance to pollutant stress (Taghavi et al., 2005; Nadeem et al., 2013; Bell et al., 2015).
Finally, we should briefly mention organic agricultural inputs as a complementary strategy not included in the abovementioned categories (microbe, plant, or meta-organisms), but that also results in the modification of the rhizosphere microbiome. Organic agriculture aims at limiting or preventing the exposure of plants, microbes, and humans to unnecessary hazards such as pesticides, herbicides, insecticides, and fungicides. Organic fertilizer such as animal manure, biosolids, and compost has been proposed as a resource to amend crops but some disadvantage should be taken into account, namely, increase salinity, presence of active therapeutic agents (manure and sludge waste), heavy metals such as zinc, cooper, and cadmium (industrial biosolids) and residues of synthetic molecules like pesticides, herbicides etc, (green wastes or compost; Table 3). An important aspect to consider when applying any agricultural inputs is to target the increase in the levels of soil organic matter that in turn, will also increase soil biological activity (Savka et al., 2002; Bünemann et al., 2006; Mazzola, 2007).
Conclusion and Outlook
The microbiome is emerging as a fundamental plant trait, resulting in beneficial or detrimental effects on plant growth, health, productivity, and functions. This delicate balance is controlled by complex chemical signals interplay between the plant and its microbiome. Further research aiming at understanding this interplay at the community level is needed to fully understand the factors controlling microbiome assemblage and its feedback to the plant host. New ‘omics tools will undoubtedly help attaining that goal, but at the same time further efforts to cultivate the rhizosphere microbiome will also be needed to reach a deeper mechanistic understanding of it. Based on the engineering efforts detailed in this contribution, further research will hopefully result in methods to purposefully, reliably, and sustainably engineer plant–microbiomes. A full optimization of the plant–microbiome meta-organism should result, among others, in a more sustainable agriculture, reduced greenhouse gas emissions, and increased rates of soil decontamination.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Adesemoye, A. O., and Egamberdieva, D. (2013). “Beneficial effects of plant growth-promoting rhizobacteria on improved crop production: prospects for developing economies,” in Bacteria in Agrobiology: Crop Productivity, eds D. K. Maheshwari, M. Saraf, and A. Aeron (Berlin: Springer), 45–63.
Adesemoye, A. O., Torbert, H. A., and Kloepper, J. W. (2008). Enhanced plant nutrient use efficiency with PGPR and AMF in an integrated nutrient management system. Can. J. Microbiol. 54, 876–886. doi: 10.1139/W08-081
An, M., Zhou, X., Wu, F., Ma, Y., and Yang, P. (2011). Rhizosphere soil microorganism populations and community structures of different watermelon cultivars with differing resistance to Fusarium oxysporum f. sp. niveum. Can. J. Microbiol. 57, 355–365. doi: 10.1139/w11-015
Anoop, V. M., Basu, U., Mccammon, M. T., Mcalister-Henn, L., and Taylor, G. J. (2003). Modulation of citrate metabolism alters aluminum tolerance in yeast and transgenic canola overexpressing a mitochondrial citrate synthase. Plant Physiol. 132, 2205–2217. doi: 10.1104/pp.103.023903
Badri, D. V., and Vivanco, J. M. (2009). Regulation and function of root exudates. Plant Cell Environ. 32, 666–681. doi: 10.1111/j.1365-3040.2009.01926.x
Badri, D. V., Weir, T. L., Van Der Lelie, D., and Vivanco, J. M. (2009). Rhizosphere chemical dialogues: plant–microbe interactions. Curr. Opin. Biotechnol. 20, 642–650. doi: 10.1016/j.copbio.2009.09.014
Bais, H., Broeckling, C., and Vivanco, J. (2008). “Root exudates modulate plant—microbe interactions in the rhizosphere,” in Secondary Metabolites in Soil Ecology, ed. P. Karlovsky (Berlin: Springer), 241–252. doi: 10.1007/978-3-540-74543-3_11
Bais, H. P., Weir, T. L., Perry, L. G., Gilroy, S., and Vivanco, J. M. (2006). The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol. 57, 233–266. doi: 10.1146/annurev.arplant.57.032905.105159
Baker, K., and Cook, R. J. (1974). Biological Control of Plant Pathogens. New York: W.H. Freeman and Company.
Bakker, M., Manter, D., Sheflin, A., Weir, T., and Vivanco, J. (2012). Harnessing the rhizosphere microbiome through plant breeding and agricultural management. Plant Soil 360, 1–13. doi: 10.1007/s11104-012-1361-x
Bakker, P. A. H. M., Berendsen, R. L., Doornbos, R. F., Wintermans, P. C. A., and Pieterse, C. M. J. (2013). The rhizosphere revisited: root microbiomics. Front. Plant Sci. 4:165. doi: 10.3389/fpls.2013.00165
Baudoin, E., Benizri, E., and Guckert, A. (2003). Impact of artificial root exudates on the bacterial community structure in bulk soil and maize rhizosphere. Soil Biol. Biochem. 35, 1183–1192. doi: 10.1016/S0038-0717(03)00179-2
Bell, T. H., Cloutier-Hurteau, B., Al-Otaibi, F., Turmel, M.-C., Yergeau, E., Courchesne, F., et al. (2015). Early rhizosphere microbiome composition is related to the growth and Zn uptake of willows introduced to a former landfill. Environ. Microbiol. doi: 10.1111/1462-2920.12900 [Epub ahead of print].
Bell, T. H., El-Din Hassan, S., Lauron-Moreau, A., Al-Otaibi, F., Hijri, M., Yergeau, E., et al. (2014a). Linkage between bacterial and fungal rhizosphere communities in hydrocarbon-contaminated soils is related to plant phylogeny. ISME J. 8, 331–343. doi: 10.1038/ismej.2013.149
Bell, T. H., Joly, S., Pitre, F. E., and Yergeau, E. (2014b). Increasing phytoremediation efficiency and reliability using novel omics approaches. Trends Biotechnol. 32, 271–280. doi: 10.1016/j.tibtech.2014.02.008
Bengough, A. G., and Mullins, C. E. (1990). Mechanical impedance to root growth: a review of experimental techniques and root growth responses. J. Soil Sci. 41, 341–358. doi: 10.1111/j.1365-2389.1990.tb00070.x
Berendsen, R. L., Pieterse, C. M. J., and Bakker, P. A. H. M. (2012). The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486. doi: 10.1016/j.tplants.2012.04.001
Bosch, T. C. G., and McFall-Ngai, M. J. (2011). Metaorganisms as the new frontier. Zoology 114, 185–190. doi: 10.1016/j.zool.2011.04.001
Braeken, K., Daniels, R., Ndayizeye, M., Vanderleyden, J., and Michiels, J. (2008). “Quorum sensing in bacteria-plant interactions,” in Molecular Mechanisms of Plant and Microbe Coexistence, eds C. Nautiyal and P. Dion (Berlin: Springer), 265–289. doi: 10.1007/978-3-540-75575-3_11
Broeckling, C. D., Broz, A. K., Bergelson, J., Manter, D. K., and Vivanco, J. M. (2008). Root exudates regulate soil fungal community composition and diversity. Appl. Environ. Microbiol. 74, 738–744. doi: 10.1128/aem.02188-07
Brussaard, L., De Ruiter, P. C., and Brown, G. G. (2007). Soil biodiversity for agricultural sustainability. Agric. Ecosystems Environ. 121, 233–244. doi: 10.1016/j.agee.2006.12.013
Bulgarelli, D., Rott, M., Schlaeppi, K., Ver Loren Van Themaat, E., Ahmadinejad, N., Assenza, F., et al. (2012). Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95. doi: 10.1038/nature11336
Bünemann, E. K., Schwenke, G. D., and Van Zwieten, L. (2006). Impact of agricultural inputs on soil organisms—a review. Soil Res. 44, 379–406. doi: 10.1071/SR05125
Chaparro, J. M., Badri, D. V., Bakker, M. G., Sugiyama, A., Manter, D. K., and Vivanco, J. M. (2013). Root Exudation of phytochemicals in Arabidopsis follows specific patterns that are developmentally programmed and correlate with soil microbial functions. PLoS ONE 8:e55731. doi: 10.1371/journal.pone.0055731
Chaparro, J. M., Badri, D. V., and Vivanco, J. M. (2014). Rhizosphere microbiome assemblage is affected by plant development. ISME J. 8, 790–803. doi: 10.1038/ismej.2013.196
Chaparro, J., Sheflin, A., Manter, D., and Vivanco, J. (2012). Manipulating the soil microbiome to increase soil health and plant fertility. Biol. Fertil. Soils 48, 489–499. doi: 10.1007/s00374-012-0691-4
Choudhary, D. K., Prakash, A., and Johri, B. N. (2007). Induced systemic resistance (ISR) in plants: mechanism of action. Indian J. Microbiol. 47, 289–297. doi: 10.1007/s12088-007-0054-2
Dangl, J. L., Horvath, D. M., and Staskawicz, B. J. (2013). Pivoting the plant immune system from dissection to deployment. Science 341, 746–751. doi: 10.1126/science.1236011
De-la-Peña, C., Badri, D., and Loyola-Vargas, V. (2012). “Plant root secretions and their interactions with neighbors,” in Secretions and Exudates in Biological Systems, eds J. M. Vivanco and F. Baluška (Berlin: Springer), 1–26. doi: 10.1007/978-3-642-23047-9_1
Delhaize, E., Gruber, B. D., and Ryan, P. R. (2007). The roles of organic anion permeases in aluminium resistance and mineral nutrition. FEBS Lett. 581, 2255–2262. doi: 10.1016/j.febslet.2007.03.057
Dessaux, Y., Petit, A., Farrand, S., and Murphy, P. (1998). “Opines and opine-like molecules involved in plant-rhizobiaceae interactions,” in The Rhizobiaceae, eds H. Spaink, A. Kondorosi, and P. J. Hooykaas (Dordrecht: Springer), 173–197. doi: 10.1007/978-94-011-5060-6_9
Dong, Y.-H., Wang, L.-H., Xu, J.-L., Zhang, H.-B., Zhang, X.-F., and Zhang, L.-H. (2001). Quenching quorum-sensing-dependent bacterial infection by an N-acyl homoserine lactonase. Nature 411, 813–817. doi: 10.1038/35081101
Downie, J. A. (1994). Signalling strategies for nodulation of legumes by rhizobia. Trends Microbiol. 2, 318–324. doi: 10.1016/0966-842X(94)90448-0
Ellouze, W., Hamel, C., Cruz, A. F., Ishii, T., Gan, Y., Bouzid, S., et al. (2012). Phytochemicals and spore germination: at the root of AMF host preference? Appl. Soil Ecol. 60, 98–104. doi: 10.1016/j.apsoil.2012.02.004
Ellouze, W., Hamel, C., Vujanovic, V., Gan, Y., Bouzid, S., and St-Arnaud, M. (2013). Chickpea genotypes shape the soil microbiome and affect the establishment of the subsequent durum wheat crop in the semiarid North American Great Plains. Soil Biol. Biochem. 63, 129–141. doi: 10.1016/j.soilbio.2013.04.001
Ferluga, S., and Venturi, V. (2009). OryR Is a LuxR-Family protein involved in interkingdom signaling between pathogenic Xanthomonas oryzae pv. oryzae and Rice. J. Bacteriol. 191, 890–897. doi: 10.1128/jb.01507-08
Friesen, M. L., Porter, S. S., Stark, S. C., Von Wettberg, E. J., Sachs, J. L., and Martinez-Romero, E. (2011). Microbially mediated plant functional traits. Annu. Rev. Ecol. Evol. Syst. 42, 23–46. doi: 10.1146/annurev-ecolsys-102710-145039
Gévaudant, F., Duby, G., Von Stedingk, E., Zhao, R., Morsomme, P., and Boutry, M. (2007). Expression of a constitutively activated plasma membrane H+-ATPase alters plant development and increases salt tolerance. Plant Physiol. 144, 1763–1776. doi: 10.1104/pp.107.103762
Glick, B. R. (2014). Bacteria with ACC deaminase can promote plant growth and help to feed the world. Microbiol. Res. 169, 30–39. doi: 10.1016/j.micres.2013.09.009
Griffiths, R. I., Bailey, M. J., Mcnamara, N. P., and Whiteley, A. S. (2006). The functions and components of the Sourhope soil microbiota. Appli. Soil Ecol. 33, 114–126. doi: 10.1016/j.apsoil.2006.03.007
Hartmann, A., Schmid, M., Tuinen, D., and Berg, G. (2009). Plant-driven selection of microbes. Plant Soil 321, 235–257. doi: 10.1007/s11104-008-9814-y
Hause, B., and Schaarschmidt, S. (2009). The role of jasmonates in mutualistic symbioses between plants and soil-born microorganisms. Phytochemistry 70, 1589–1599. doi: 10.1016/j.phytochem.2009.07.003
Kielak, A., Pijl, A. S., Van Veen, J. A., and Kowalchuk, G. A. (2008). Differences in vegetation composition and plant species identity lead to only minor changes in soil-borne microbial communities in a former arable field. FEMS Microbiol. Ecol. 63, 372–382. doi: 10.1111/j.1574-6941.2007.00428.x
Kowalchuk, G. A., Buma, D., De Boer, W., Klinkhamer, P. L., and Van Veen, J. (2002). Effects of above-ground plant species composition and diversity on the diversity of soil-borne microorganisms. Antonie Van Leeuwenhoek 81, 509–520. doi: 10.1023/A:1020565523615
Kowalchuk, G. A., Yergeau, E., Leveau, J. H. J., Sessitch, A., and Bailey, M. (2010). “Plant-associated microbial communities,” in Environmental Molecular Microbiology, eds W.-T. Liu and J. K. Jansson (Norwich, FC: Caister Academic Press), 133–147.
Koyama, H., Kawamura, A., Kihara, T., Hara, T., Takita, E., and Shibata, D. (2000). Overexpression of mitochondrial citrate synthase in Arabidopsis thaliana improved growth on a phosphorus-limited soil. Plant Cell Physiol. 41, 1030–1037. doi: 10.1093/pcp/pcd029
Koyama, H., Takita, E., Kawamura, A., Hara, T., and Shibata, D. (1999). Over expression of mitochondrial citrate synthase gene improves the growth of carrot cells in Al-Phosphate medium. Plant Cell Physiol. 40, 482–488. doi: 10.1093/oxfordjournals.pcp.a029568
Lakshmanan, V., Selvaraj, G., and Bais, H. P. (2014). Functional soil microbiome: belowground solutions to an aboveground problem. Plant Physiol. 166, 689–700. doi: 10.1104/pp.114.245811
Lemanceau, P., Corberand, T., Gardan, L., Latour, X., Laguerre, G., Boeufgras, J., et al. (1995). Effect of two plant species, flax (Linum usitatissinum L.) and tomato (Lycopersicon esculentum Mill.), on the diversity of soilborne populations of fluorescent pseudomonads. Appl. Environ. Microbiol. 61, 1004–1012.
Li, J., Yang, H., Ann Peer, W., Richter, G., Blakeslee, J., Bandyopadhyay, A., et al. (2005). Arabidopsis H+-PPase AVP1 regulates auxin-mediated organ development. Science 310, 121–125. doi: 10.1126/science.1115711
Lioussanne, L., Jolicoeur, M., and St-Arnaud, M. (2008). Mycorrhizal colonization with Glomus intraradices and development stage of transformed tomato roots significantly modify the chemotactic response of zoospores of the pathogen Phytophthora nicotianae. Soil Biol. Biochem. 40, 2217–2224. doi: 10.1016/j.soilbio.2008.04.013
Loreau, M., Naeem, S., Inchausti, P., Bengtsson, J., Grime, J. P., Hector, A., et al. (2001). Biodiversity and ecosystem functioning: current knowledge and future challenges. Science 294, 804–808. doi: 10.1126/science.1064088
Lynch, J. M., Benedetti, A., Insam, H., Nuti, M. P., Smalla, K., Torsvik, V., et al. (2004). Microbial diversity in soil: ecological theories, the contribution of molecular techniques and the impact of transgenic plants and transgenic microorganisms. Biol. Fertil. Soils 40, 363–385. doi: 10.1007/s00374-004-0784-9
Mabood, F., Jung, W., and Smith, D. (2008). “Signals in the underground: microbial signaling and plant productivity,” in Molecular Mechanisms of Plant and Microbe Coexistence, eds C. Nautiyal and P. Dion (Berlin: Springer), 291–318. doi: 10.1007/978-3-540-75575-3_12
Marasco, R., Rolli, E., Ettoumi, B., Vigani, G., Mapelli, F., Borin, S., et al. (2012). A drought resistance-promoting microbiome is selected by root system under desert farming. PLoS ONE 7:e48479. doi: 10.1371/journal.pone.0048479
Marihal, A. K., and Jagadeesh, K. S. (2013). “Plant–Microbe interaction: a potential tool for enhanced bioremediation,” in Plant Microbe Symbiosis: Fundamentals and Advances, ed. N. K. Arora (New Delhi: Springer), 395–410. doi: 10.1007/978-81-322-1287-4_15
Marilley, L., and Aragno, M. (1999). Phylogenetic diversity of bacterial communities differing in degree of proximity of Lolium perenne and Trifolium repens roots. Appl. Soil Ecol. 13, 127–136. doi: 10.1016/S0929-1393(99)00028-1
Mazzola, M. (2002). Mechanisms of natural soil suppressiveness to soilborne diseases. Antonie Van Leeuwenhoek 81, 557–564. doi: 10.1023/A:1020557523557
Mazzola, M. (2007). Manipulation of rhizosphere bacterial communities to induce suppressive soils. J. Nematol. 39, 213–220.
Mendes, R., Kruijt, M., De Bruijn, I., Dekkers, E., Van Der Voort, M., Schneider, J. H. M., et al. (2011). Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332, 1097–1100. doi: 10.1126/science.1203980
Mercier, A., Kay, E., and Simonet, P. (2006). “Horizontal gene transfer by natural transformation in soil environment,” in Nucleic Acids and Proteins in Soil, eds P. Nannipieri and K. Smalla (Berlin: Springer), 355–373. doi: 10.1007/3-540-29449-X_15
Miller, J. B., and Oldroyd, G. D. (2012). “The role of diffusible signals in the establishment of rhizobial and mycorrhizal symbioses,” in Signaling and Communication in Plant Symbiosis, eds S. Perotto and F. Baluška (Berlin: Springer), 1–30.
Morel, M., and Castro-Sowinski, S. (2013). “The complex molecular signaling network in microbe–plant interaction,” in Plant Microbe Symbiosis: Fundamentals and Advances, ed. N. K. Arora (New Delhi: Springer), 169–199. doi: 10.1007/978-81-322-1287-4_6
Nadeem, S., Naveed, M., Zahir, Z., and Asghar, H. (2013). “Plant–microbe interactions for sustainable agriculture: fundamentals and recent advances,” in Plant Microbe Symbiosis: Fundamentals and Advances, ed. N. K. Arora (New Delhi: Springer), 51–103. doi: 10.1007/978-81-322-1287-4_2
Neal, J. L. Jr., Atkinson, T. G., and Larson, R. I. (1970). Changes in the rhizosphere microflora of spring wheat induced by disomic substitution of a chromosome. Can. J. Microbiol. 16, 153–158. doi: 10.1139/m70-027
Neal, J. L. Jr., Larson, R., and Atkinson, T. G. (1973). Changes in rhizosphere populations of selected physiological groups of bacteria related to substitution of specific pairs of chromosomes in spring wheat. Plant Soil 39, 209–212. doi: 10.1007/BF00018061
Oburger, E., Dell‘Mour, M., Hann, S., Wieshammer, G., Puschenreiter, M., and Wenzel, W. W. (2013). Evaluation of a novel tool for sampling root exudates from soil-grown plants compared to conventional techniques. Environ. Exp. Bot. 87, 235–247. doi: 10.1016/j.envexpbot.2012.11.007
O’Connell, K. P., Goodman, R. M., and Handelsman, J. (1996). Engineering the rhizosphere: expressing a bias. Trends Biotechnol. 14, 83–88. doi: 10.1016/0167-7799(96)80928-0
Oldroyd, G. E. D. (2013). Speak, friend, and enter: signalling systems that promote beneficial symbiotic associations in plants. Nat. Rev. Micro. 11, 252–263. doi: 10.1038/nrmicro2990
Ortiz-Castro, R., Martinez-Trujillo, M., and Lopez-Bucio, J. (2008). N-acyl-L-homoserine lactones: a class of bacterial quorum-sensing signals alter post-embryonic root development in Arabidopsis thaliana. Plant Cell Environ. 31, 1497–1509. doi: 10.1111/j.1365-3040.2008.01863.x
Pagé, A. P., Yergeau, E., and Greer, C. W. (2015). Salix purpurea stimulates the expression of specific bacterial xenobiotic degradation genes in a soil contaminated with hydrocarbons. PLoS ONE 10:e0132062. doi: 10.1371/journal.pone.0132062
Paulin, M. M., Novinscak, A., St-Arnaud, M., Goyer, C., Decoste, N. J., Privé, J.-P., et al. (2009). Transcriptional activity of antifungal metabolite-encoding genes phlD and hcnBC in Pseudomonas spp. using qRT-PCR. FEMS Microbiol. Ecol. 68, 212–222. doi: 10.1111/j.1574-6941.2009.00669.x
Peiffer, J. A., Spor, A., Koren, O., Jin, Z., Tringe, S. G., Dangl, J. L., et al. (2013). Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc. Natl. Acad. Sci. U.S.A. 110, 6548–6553. doi: 10.1073/pnas.1302837110
Phillips, R. P., Erlitz, Y., Bier, R., and Bernhardt, E. S. (2008). New approach for capturing soluble root exudates in forest soils. Funct. Ecol. 22, 990–999. doi: 10.1111/j.1365-2435.2008.01495.x
Pieterse, C. M. J., Leon-Reyes, A., Van Der Ent, S., and Van Wees, S. C. M. (2009). Networking by small-molecule hormones in plant immunity. Nat. Chem. Biol. 5, 308–316. doi: 10.1038/nchembio.164
Ping, L., and Boland, W. (2004). Signals from the underground: bacterial volatiles promote growth in Arabidopsis. Trends Plant Sci. 9, 263–266. doi: 10.1016/j.tplants.2004.04.008
Qiu, M., Li, S., Zhou, X., Cui, X., Vivanco, J., Zhang, N., et al. (2014). De-coupling of root–microbiome associations followed by antagonist inoculation improves rhizosphere soil suppressiveness. Biol. Fertil. Soils 50, 217–224. doi: 10.1007/s00374-013-0835-1
Ryan, P., Dessaux, Y., Thomashow, L., and Weller, D. (2009). Rhizosphere engineering and management for sustainable agriculture. Plant Soil 321, 363–383. doi: 10.1007/s11104-009-0001-6
Savka, M. A., Dessaux, Y., Oger, P., and Rossbach, S. (2002). Engineering bacterial competitiveness and persistence in the phytosphere. Mol. Plant Microbe Interact. 15, 866–874. doi: 10.1094/MPMI.2002.15.9.866
Savka, M. A., and Farrand, S. K. (1997). Modification of rhizobacterial populations by engineering bacterium utilization of a novel plant-produced resource. Nat. Biotechnol. 15, 363–368. doi: 10.1038/nbt0497-363
Schuhegger, R., Ihring, A., Gantner, S., Bahnweg, G., Knappe, C., Vogg, G., et al. (2006). Induction of systemic resistance in tomato by N-acyl-L-homoserine lactone-producing rhizosphere bacteria. Plant Cell Environ. 29, 909–918. doi: 10.1111/j.1365-3040.2005.01471.x
Sharma, S., Shukla, K. P., Singh, V., Singh, J., Devi, S., and Tewari, A. (2013). “Plant–Microbe symbiosis: perspectives and applications,” in Plant Microbe Symbiosis: Fundamentals and Advances, ed. N. K. Arora (New Delhi: Springer), 119–145. doi: 10.1007/978-81-322-1287-4_4
Smalla, K., Wieland, G., Buchner, A., Zock, A., Parzy, J., Kaiser, S., et al. (2001). Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: plant-dependent enrichment and seasonal shifts revealed. Appl. Environ. Microbiol. 67, 4742–4751. doi: 10.1128/aem.67.10.4742-4751.2001
Solaiman, M. Z., Senoo, K., Kawaguchi, M., Imaizumi-Anraku, H., Akao, S., Tanaka, K., et al. (2000). Characterization of mycorrhiza formed by Glomus sp. on roots of hypernodulating mutants of Lotus japonicus. J. Plant Res. 113, 443–448. doi: 10.1007/PL00013953
Soto, M. J., Sanjuán, J., and Olivares, J. (2006). Rhizobia and plant-pathogenic bacteria: common infection weapons. Microbiology 152, 3167–3174. doi: 10.1099/mic.0.29112-0
Stefani, F. O. P., Bell, T. H., Marchand, C., De La Providencia, I. E., El Yassimi, A., St-Arnaud, M., et al. (2015). Culture-dependent and -independent methods capture different microbial community fractions in hydrocarbon-contaminated soils. PLoS ONE 10:e0128272. doi: 10.1371/journal.pone.0128272
Stein, E., Molitor, A., Kogel, K.-H., and Waller, F. (2008). Systemic resistance in Arabidopsis conferred by the mycorrhizal fungus piriformospora indica requires jasmonic acid signaling and the cytoplasmic function of NPR1. Plant Cell Physiol. 49, 1747–1751. doi: 10.1093/pcp/pcn147
Straight, P. D., and Kolter, R. (2009). Interspecies chemical communication in bacterial development. Annu. Rev. Microbiol. 63, 99–118. doi: 10.1146/annurev.micro.091208.073248
Sturz, A. V., Christie, B. R., and Matheson, B. G. (1998). Associations of bacterial endophyte populations from red clover and potato crops with potential for beneficial allelopathy. Can. J. Microbiol. 44, 162–167. doi: 10.1139/w97-146
Sugiyama, A., and Yazaki, K. (2012). “Root exudates of legume plants and their involvement in interactions with soil microbes,” in Secretions and Exudates in Biological Systems, eds J. M. Vivanco and F. Baluška (Berlin: Springer), 27–48.
Taghavi, S., Barac, T., Greenberg, B., Borremans, B., Vangronsveld, J., and Van Der Lelie, D. (2005). Horizontal gene transfer to endogenous endophytic bacteria from poplar improves phytoremediation of toluene. Appl. Environ. Microbiol. 71, 8500–8505. doi: 10.1128/aem.71.12.8500-8505.2005
Taghavi, S., Garafola, C., Monchy, S., Newman, L., Hoffman, A., Weyens, N., et al. (2009). Genome survey and characterization of endophytic bacteria exhibiting a beneficial effect on growth and development of poplar trees. Appl. Environ. Microbiol. 75, 748–757. doi: 10.1128/aem.02239-08
Tarkka, M., Schrey, S., and Hampp, R. (2008). “Plant associated soil micro-organisms,” in Molecular Mechanisms of Plant and Microbe Coexistence, eds C. Nautiyal and P. Dion (Berlin: Springer), 3–51. doi: 10.1007/978-3-540-75575-3_1
Teplitski, M., Robinson, J. B., and Bauer, W. D. (2000). Plants secrete substances that mimic bacterial N-Acyl homoserine lactone signal activities and affect population density-dependent behaviors in associated bacteria. Mol. Plant Microbe Interact. 13, 637–648. doi: 10.1094/MPMI.2000.13.6.637
Tesfaye, M., Temple, S. J., Allan, D. L., Vance, C. P., and Samac, D. A. (2001). Overexpression of malate dehydrogenase in transgenic alfalfa enhances organic acid synthesis and confers tolerance to aluminum. Plant Physiol. 127, 1836–1844. doi: 10.1104/pp.010376
Toussaint, J.-P., St-Arnaud, M., and Charest, C. (2004). Nitrogen transfer and assimilation between the arbuscular mycorrhizal fungus Glomus intraradices Schenck & Smith and Ri T-DNA roots of Daucus carota L. in an in vitro compartmented system. Can. J. Microbiol. 50, 251–260. doi: 10.1139/w04-009
Truchet, G., Roche, P., Lerouge, P., Vasse, J., Camut, S., De Billy, F., et al. (1991). Sulphated lipo-oligosaccharide signals of Rhizobium meliloti elicit root nodule organogenesis in alfalfa. Nature 351, 670–673. doi: 10.1038/351670a0
Vandenkoornhuyse, P., Quaiser, A., Duhamel, M., Le Van, A., and Dufresne, A. (2015). The importance of the microbiome of the plant holobiont. New Phytol. 206, 1196–1206. doi: 10.1111/nph.13312
van Loon, L. C., Rep, M., and Pieterse, C. M. J. (2006). Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 44, 135–162. doi: 10.1146/annurev.phyto.44.070505.143425
Walker, T. S., Bais, H. P., Grotewold, E., and Vivanco, J. M. (2003). Root exudation and rhizosphere biology. Plant Physiol. 132, 44–51. doi: 10.1104/pp.102.019661
Yang, H., Knapp, J., Koirala, P., Rajagopal, D., Peer, W. A., Silbart, L. K., et al. (2007). Enhanced phosphorus nutrition in monocots and dicots over-expressing a phosphorus-responsive type I H+-pyrophosphatase. Plant Biotechnol. J. 5, 735–745. doi: 10.1111/j.1467-7652.2007.00281.x
Yergeau, E., Sanschagrin, S., Maynard, C., St-Arnaud, M., and Greer, C. W. (2014). Microbial expression profiles in the rhizosphere of willows depend on soil contamination. ISME J. 8, 344–358. doi: 10.1038/ismej.2013.163
Zhang, Y., Ruyter-Spira, C., and Bouwmeester, H. J. (2015). Engineering the plant rhizosphere. Curr. Opin. Biotechnol. 32, 136–142. doi: 10.1016/j.copbio.2014.12.006
Ziegler, M., Engel, M., Welzl, G., and Schloter, M. (2013). Development of a simple root model to study the effects of single exudates on the development of bacterial community structure. J. Microbiol. Methods 94, 30–36. doi: 10.1016/j.mimet.2013.04.003
Zolla, G., Badri, D. V., Bakker, M. G., Manter, D. K., and Vivanco, J. M. (2013). Soil microbiomes vary in their ability to confer drought tolerance to Arabidopsis. Appl. Soil Ecol. 68, 1–9. doi: 10.1016/j.apsoil.2013.03.007
Keywords: rhizosphere, signaling, beneficial microorganisms, agriculture, plant–microbe interactions
Citation: Quiza L, St-Arnaud M and Yergeau E (2015) Harnessing phytomicrobiome signaling for rhizosphere microbiome engineering. Front. Plant Sci. 6:507. doi: 10.3389/fpls.2015.00507
Received: 13 May 2015; Accepted: 25 June 2015;
Published: 14 July 2015.
Edited by:
Jean-Michel Ané, University of Wisconsin–Madison, USAReviewed by:
Oswaldo Valdes-Lopez, National Autonomous University of Mexico, MexicoThomas Robert Rey, De Sangosse, France
Copyright © 2015 Quiza, St-Arnaud and Yergeau. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Etienne Yergeau, National Research Council Canada, 6100 Royalmount Avenue, Montréal, QC H4P 2R2, Canada,ZXRpZW5uZS55ZXJnZWF1QGNucmMtbnJjLmdjLmNh
 Liliana Quiza
Liliana Quiza Marc St-Arnaud
Marc St-Arnaud Etienne Yergeau
Etienne Yergeau