- 1Hainan Key Laboratory for Sustainable Utilization of Tropical Bioresources, College of Agriculture, Hainan University, Haikou, China
- 2State Key Laboratory of Tree Genetics and Breeding, Research Institute of Forestry, Chinese Academy of Forestry, Beijing, China
- 3Key Laboratory of Plant Germplasm Enhancement and Specialty Agriculture, Wuhan Botanical Garden, Chinese Academy of Sciences, Wuhan, China
As a dehydrin belonging to group II late embryogenesis abundant protein (LEA) family, Arabidopsis Low Temperature-Induced 30 (LTI30)/XERO2 has been shown to be involved in plant freezing stress resistance. However, the other roles of AtLTI30 remain unknown. In this study, we found that the expression of AtLTI30 was largely induced by drought stress and abscisic acid (ABA) treatments. Thereafter, AtLTI30 knockout mutants and overexpressing plants were isolated to investigate the possible involvement of AtLTI30 in ABA and drought stress responses. AtLTI30 knockout mutants were less sensitive to ABA-mediated seed germination, while AtLTI30 overexpressing plants were more sensitive to ABA compared with wild type (WT). Consistently, the AtLTI30 knockout mutants displayed decreased drought stress resistance, while the AtLTI30 overexpressing plants showed improved drought stress resistance compared with WT, as evidenced by a higher survival rate and lower leaf water loss than WT after drought stress. Moreover, manipulation of AtLTI30 expression positively regulated the activities of catalases (CATs) and endogenous proline content, as a result, negatively regulated drought stress-triggered hydrogen peroxide (H2O2) accumulation. All these results indicate that AtLTI30 is a positive regulator of plant drought stress resistance, partially through the modulation of ABA sensitivity, H2O2 and proline accumulation.
Introduction
Plants are exposed to various environmental conditions, however, plants can not change their location to avoid unfavorable circumstance (Shi et al., 2013a,b, 2014a,b). Among multiple stresses, drought stress is one of the most harsh environmental stresses (Seki et al., 2007; Harb et al., 2010; Chan and Shi, 2015). To date, plants have developed sophisticated strategies to counteract sudden environmental changes. Many secondary messengers, including abscisic acid (ABA), and hydrogen peroxide (H2O2), are involved in plant stress transduction (Seki et al., 2007; Yu et al., 2008; Fujii et al., 2009; Cutler et al., 2010; Qin et al., 2011). Both endogenous concentrations and the underlying signaling pathways of ABA and H2O2 play essential roles in plant drought stress responses (Fujii et al., 2009; Cutler et al., 2010; Harb et al., 2010; Qin et al., 2011; Munemasa et al., 2013).
Arabidopsis Low Temperature-Induced 30 (LTI30)/XERO2 belongs to the group II late embryogenes abundant protein (LEA)/dehydrin family (Rouse et al., 1996). The transcript level of AtLTI30/XERO2 can be induced by ABA, cold, dehydration, wounding, and salt stresses (Welin et al., 1994; Rouse et al., 1996; Nylander et al., 2001; Chung and Parish, 2008). Overexpression of AtLTI30/XERO2 enhances freezing stress resistance in Arabidopsis (Puhakainen et al., 2004). AtCBF1, AtCBF2, and AtCBF3 (also known as AtDREB1b, AtDREB1c, and AtDREB1a, respectively) are important transcription factors in plant abiotic stress responses. To date, many stress-responsive genes with C-repeat (CRT)/dehydration-responsive element (DRE) in the promoters have been identified as the direct targets of AtCBFs. These genes include COR (cold regulated), ERD (early responsive to dehydration), KIN (cold inducible), LTI (low-temperature induced), and RD (responsive to dehydration) (Gilmour et al., 1998; Zarka et al., 2003; Cook et al., 2004; Thomashow, 2010). Using multiple combinations of mutations in the promoter of AtLTI30/XERO2, Chung and Parish (2008) found that two of the ACGT and DRE/CRT elements in the promoter of AtLTI30/XERO2 were essential for cold and ABA transcriptional induction of AtLTI30/XERO2. Mouillon et al. (2006) found that the lysine-rich segment of AtLTI30/XERO2 showed sequence similarity with the animal chaperone heat shock protein 90 (HSP90). The conserved segments of AtLTI30 exerted its biological function more locally upon interaction with specific biological targets. Moreover, Eriksson et al. (2011) identified three factors that regulate the lipid interaction of LTI30 in vitro, including the pH dependent His on/off switch, reversal of membrane binding by proteolytic digestion, and phosphorylation by protein kinase C.
Although AtLTI30 has been shown to be involved in plant freezing stress resistance, the other roles of AtLTI30 and the underlying mechanisms remain unknown. In this study, the expression and function of AtLTI30 were characterized in response to drought stress treatment. We investigated the effects of manipulation of AtLTI30 expression on drought stress resistance, as well as the underlying mechanisms. The results indicate that AtLTI30 is a positive regulator of drought stress resistance in Arabidopsis.
Materials And Methods
Plant Materials and Growth Conditions
After stratification at 4°C for 3 days in darkness, Arabidopsis thaliana seeds were sown in soil in a growth chamber, and watered with a nutrient solution twice per week. The growth chamber was controlled at 23°C, with an irradiance of about 150 μmol quanta m-2 s-1, under 65% relative humidity and 16-h light and 8-h dark cycles. The lti30-1 (SALK_114915) and lti30-2 (SALK_016819) mutants were obtained from the Arabidopsis Biological Resource Center (ABRC).
RNA Isolation, Semi-quantitative RT-PCR and Quantitative Real-time PCR
Total RNA was extracted and purified using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) and RQ1 RNase-free DNase (Promega, Madison, WI, USA). First-strand cDNA was synthesized from total RNA using reverse transcriptase (TOYOBO, Osaka city, Japan) as (Shi et al., 2013a,b, 2014a,b, 2015) previously described. Semi-quantitative RT-PCR was performed as Shi et al. (2013a) described using ubiquitin 10 (UBQ10) as the internal control. Quantitative real-time PCR was performed using the CFX96TM Real-Time System (BIO-RAD, Hercules, CA, USA) and the comparative ΔΔCT method with UBQ10 as a reference gene following (Shi et al., 2013a,b, 2014a,b, 2015). The primers of UBQ10 (At4g05320) and LTI30 (At3g50970) are shown in Supplementary Table S1.
Construction of Vectors and Generation of Transgenic Lines
For the pLTI30::β-glucuronidase (GUS) transgenic construction, the promoter region of AtLTI30 was amplified by PCR and inserted into the BamHI site of the pBI101.2 vector with kanamycin resistance. For AtLTI30 overexpressing transgenic construction, the coding region of AtLTI30 was amplified by PCR and inserted into the SmaI/XhoI sites of the pBIM vector with kanamycin resistance under the control of the cauliflower mosaic virus (CaMV) 35S promoter (Yang et al., 2005). The responsible primers for the above vector constructions are shown in Supplementary Table S2. The recombinant constructions were confirmed by DNA sequencing and introduced into wild type (WT) plants of Columbia-0 (Col-0) using Agrobacterium tumefaciens strain GV3101-mediated transformation and the floral dip method (Clough and Bent, 1998). Thereafter the transgenic plants were selected on MS medium using kanamycin resistance, and further confirmed by PCR analysis.
GUS Staining and Quantification of GUS Activity
Glucuronidase staining and quantification of GUS activity were performed as Jefferson et al. (1987) previously described. For GUS staining, proLTI30::GUS transgenic plants were incubated in staining solution (100 mM sodium phosphate buffer, pH 7.5, 0.5 mM K3[Fe(CN)6], and 0.5 mM K4[Fe(CN)6], 10 mM EDTA, 1.0 mM 5-bromo–chloro-3-indolyl-β-D-glucuronide and 0.1% Triton X-100) at 37°C for 8 h, after which the plants were incubated in 70% ethanol to remove chlorophyll. GUS activity was quantified by detecting the conversion of 4-methylumbelliferone from the substrate 4-methylumbelliferyl-β-glucuronide in the same concentration of protein extract, as Jefferson et al. (1987) previously described.
Determination of ABA Sensitivity
For ABA sensitivity assay, different genotypes of Arabidopsis seeds were sterilized with 70% (v/v) ethyl alcohol, 5% (w/v) NaClO, and deionized water. The seeds were stratified at 4°C for 3 days, thereafter were sown on Murashige and Skoog (MS) medium plates containing different concentrations of ABA. Germination ratios as seen with emerged radicals and green cotyledons were scored after 10 days in the growth chamber. Stomatal aperture in Arabidopsis leaves was determined as Shi et al. (2013a) described.
Plant Drought Stress Treatment and Drought Stress Resistance Assay
For drought stress, 14-day-old Arabidopsis plants in the soil were subjected to control (well-watered) and drought stress (withheld water) conditions for another 21 days. More than three pots of each variety (27 plants) were used in each biological repeat, and all these pots with plants were rotated daily during drought stress to minimize the environment effect. The survival rate of stressed plants was recorded after re-watered for 4 days later than 21 days of drought stress treatment. Relative in vitro leaf water loss rate was expressed as percent change in leaf fresh weight (FW) as (Shi et al., 2013a,b, 2014a,b, 2015) described.
Quantification of Hydrogen Peroxide (H2O2) and Catalase (CAT) Activity
The concentration of H2O2 in plant leaves was determined using the titanium sulfate method, and CAT (EC 1.11.1.6) activity was determined using the enzyme assay kit as (Shi et al., 2013a,b, 2014a,b, 2015) previously reported.
The Determination of Proline Content
Quantification of endogenous proline content was performed as Shi et al. (2013a,b) previously described. Briefly, endogenous proline in plant leaves was extracted using 3% (w/v) sulfosalicylic acid, and the red solution at the absorbance of 520 nm was determined by adding the mixtures of ninhydrin reagent and glacial acetic acid to the extractions.
Statistical Analysis
All experiments were performed with at least three biological repeats, and plant leave samples in each biological repeat were mixture samples of at least 10 plants per genotype. Student’s t-test and analysis of variance (AVOVA) were used to analysis the significant difference, and asterisk symbols (∗) indicate the significant differences of p < 0.05 in comparison to WT.
Results
The Expression Pattern of AtLTI30
Using proLTI30::GUS transgenic plants, we found that AtLTI30 was widely expressed in leaves, stems, flowers, primary roots and lateral roots (Figures 1A–E). Moreover, we found that the GUS activities of proLTI30::GUS transgenic plants were significantly increased after dehydration stress and ABA treatments for 1, 3, and 6 h through GUS activity assay (Figure 2A). The GUS result is consistent with previous studies that ABA and dehydration induced the transcript level of AtLTI30 using northern blot (Welin et al., 1994; Rouse et al., 1996; Nylander et al., 2001; Chung and Parish, 2008). Moreover, we also found that the transcript levels of AtLTI30 are induced by both dehydration and ABA treatments (Figures 2B,C) using the publicly available microarray data (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi) (Winter et al., 2007). These results indicate the possible link between AtLTI30 and these stress treatments, and suggest the possible involvement of AtLTI30 in drought stress responses in Arabidopsis.
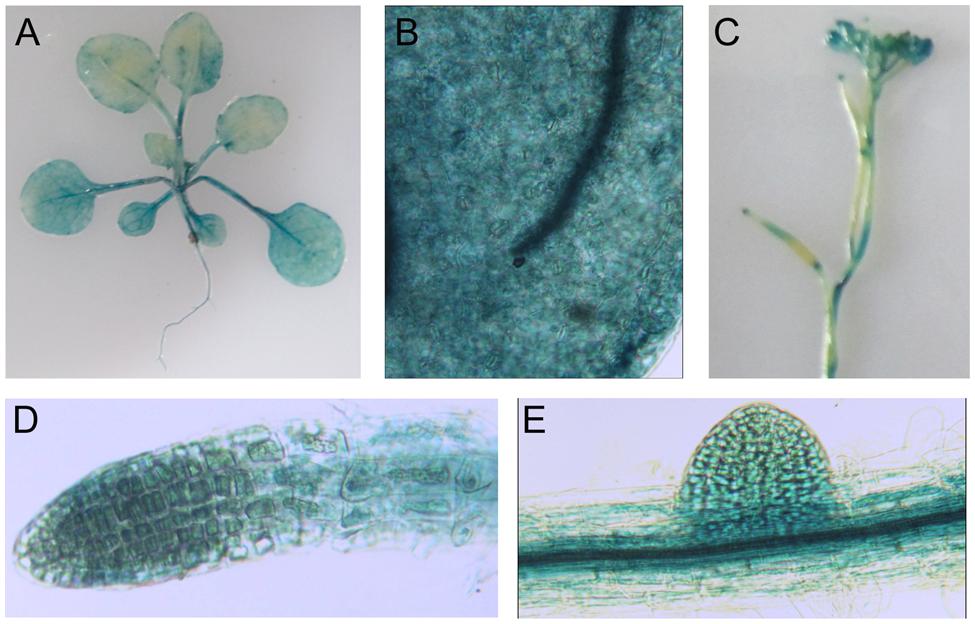
FIGURE 1. Glucuronidase staining of proLTI30::GUS transgenic plants in different organs. (A) GUS staining of 21-day-old of proLTI30::GUS transgenic plants. (B–E) GUS staining of proLTI30::GUS transgenic plants in leaves (B), flowers (C), primary root (D), and lateral root (E).
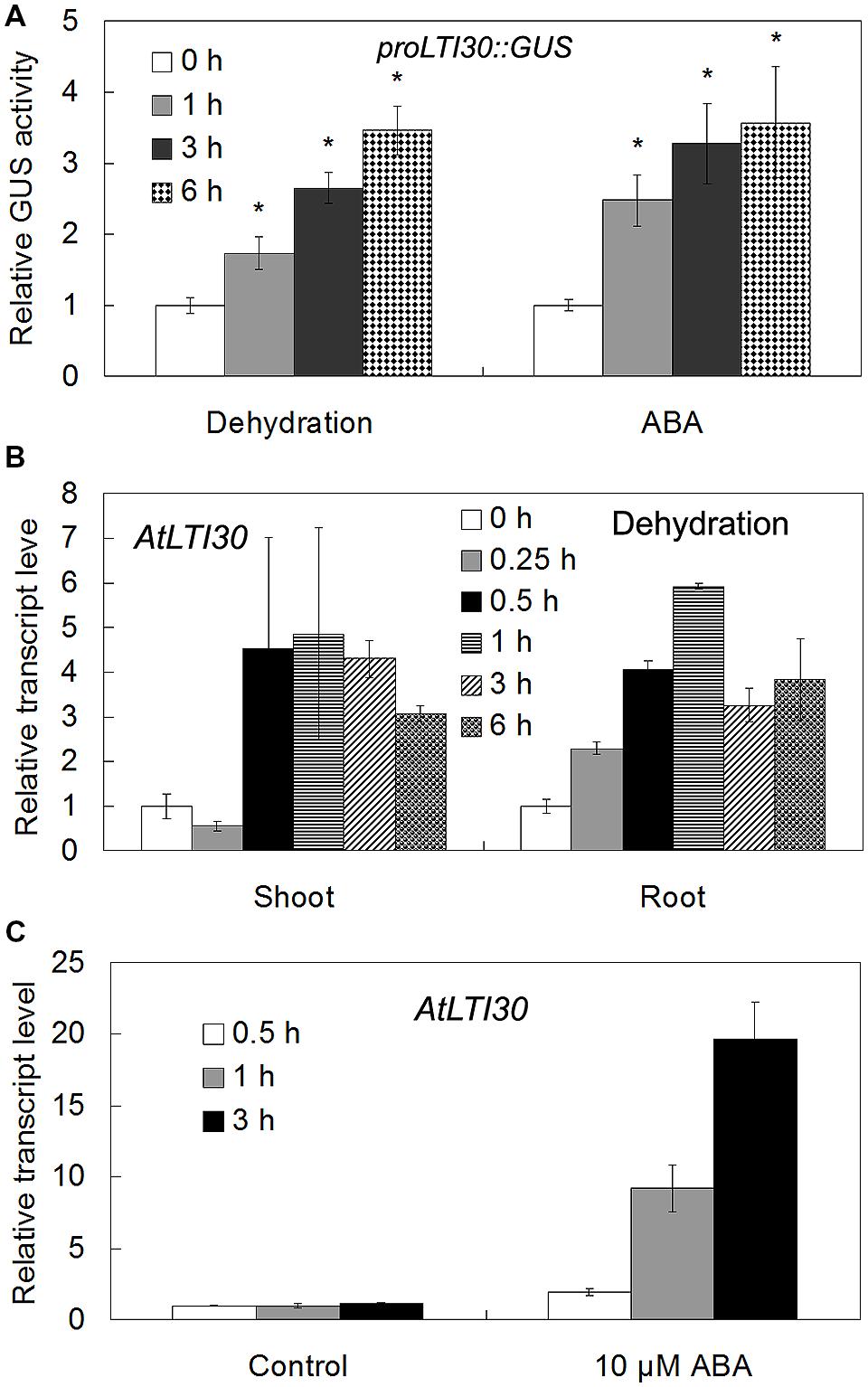
FIGURE 2. The expression levels of AtLTI30 in response to different stress treatments. (A) The GUS activities of proLTI30::GUS transgenic plants in response to different stress treatments. For the assay, 14-day-old of proLTI30::GUS seedlings were subjected to dehydration (un-covering the plate in the growth chamber), or transferred to fresh MS liquid medium containing 50 μM ABA, and samples were collected at 0, 1, 3, and 6 h after treatments. The GUS activity of proLTI30::GUS at 0 h of treatment was normalized as 1.0. The results shown are the means ± SDs of three biological repeats, and asterisk symbols (∗) indicate the significant difference of p < 0.05 compared to the expression at 0 h of treatment. (B,C) The transcript levels of AtLTI30 in response to dehydration (B) and ABA (C) using the publicly available microarray data (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi).
Isolation of AtLTI30 Knockout Mutants and Overexpressing Plants
To further reveal the in vivo role of AtLTI30, we isolated the T-DNA mutants with T-DNA insertion in the extron of AtLTI30 [lti30-1 (SALK_114915) and lti30-2 (SALK_016819)] (Figures 3A–C), and constructed AtLTI30 overexpressing transgenic plants (Figures 3B,C). Using semi-quantitative RT-PCR analysis, the transcript of AtLTI30 was un-detectable in lti30-1 and lti30-2 mutants (Figure 3B). Consistently, quantitative real-time PCR analysis also showed that the transcript level of AtLTI30 was largely inhibited in lti30-1 and lti30-2 mutants, with about 30% of AtLTI30 transcripts compared with WT plants (Figure 3C). Moreover, the AtLTI30 overexpressing transgenic plants displayed significantly higher AtLTI30 transcripts than WT plants, with 15–25-folds higher AtLTI30 transcripts compared with WT, and the homozygous transgenic plants (OX-2 and OX-3) were chosen for further analysis (Figures 3B,C).
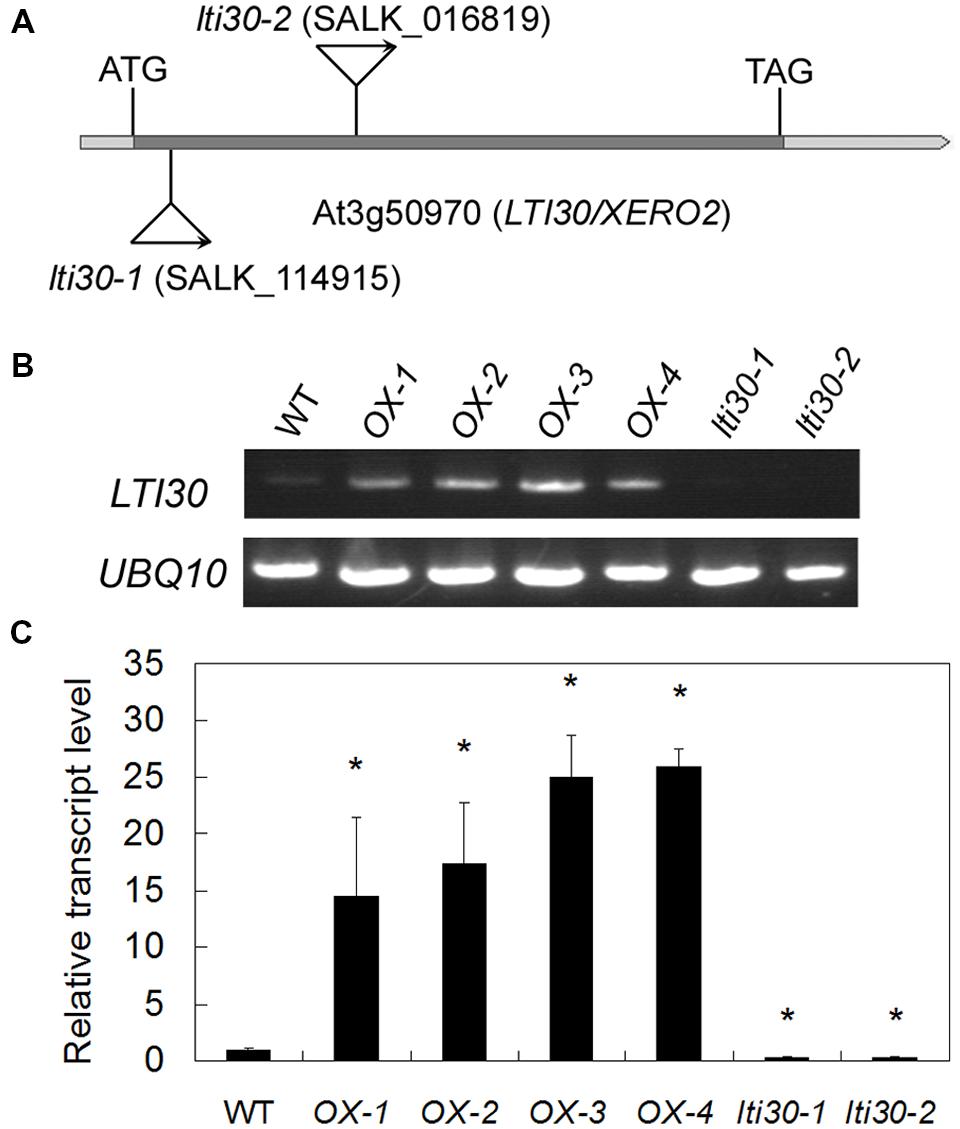
FIGURE 3. Characterization of lti30 knockout mutants and AtLTI30 overexpressing plants. (A) Schematic diagrams of the T-DNA position of lti30-1 and lti30-2 mutants. (B,C) The transcript level of AtLTI30 in lti30 knockout mutants and AtLTI30 overexpressing plants assayed by semi-quantitative RT-PCR (B) and quantitative real-time PCR (C). The expression of UBQ10 was used as an internal control. The results shown are the means ± SDs of three biological repeats, and asterisk symbols (∗) indicate the significant difference of p < 0.05 in comparison to wild type (WT).
AtLTI30 Positively Regulates Plant Sensitivity to ABA
Since the expression of AtLTI30 was increased after ABA treatment, the responses of the WT, lti30 knockout mutants, and AtLTI30 overexpressing plants to ABA were further compared. Germination of AtLTI30 overexpressing plant seeds was severely inhibited after ABA treatment, as shown with less emerged radical, less green cotyledon compared with those of WT (Figures 4A–C). On the contrary, lti30-1 and lti30-2 mutants showed more emerged radicals and more green cotyledons than those of WT (Figures 4A–C). These results indicate that modulation of AtLTI30 expression positively affects ABA sensitivity in seed germination stage.
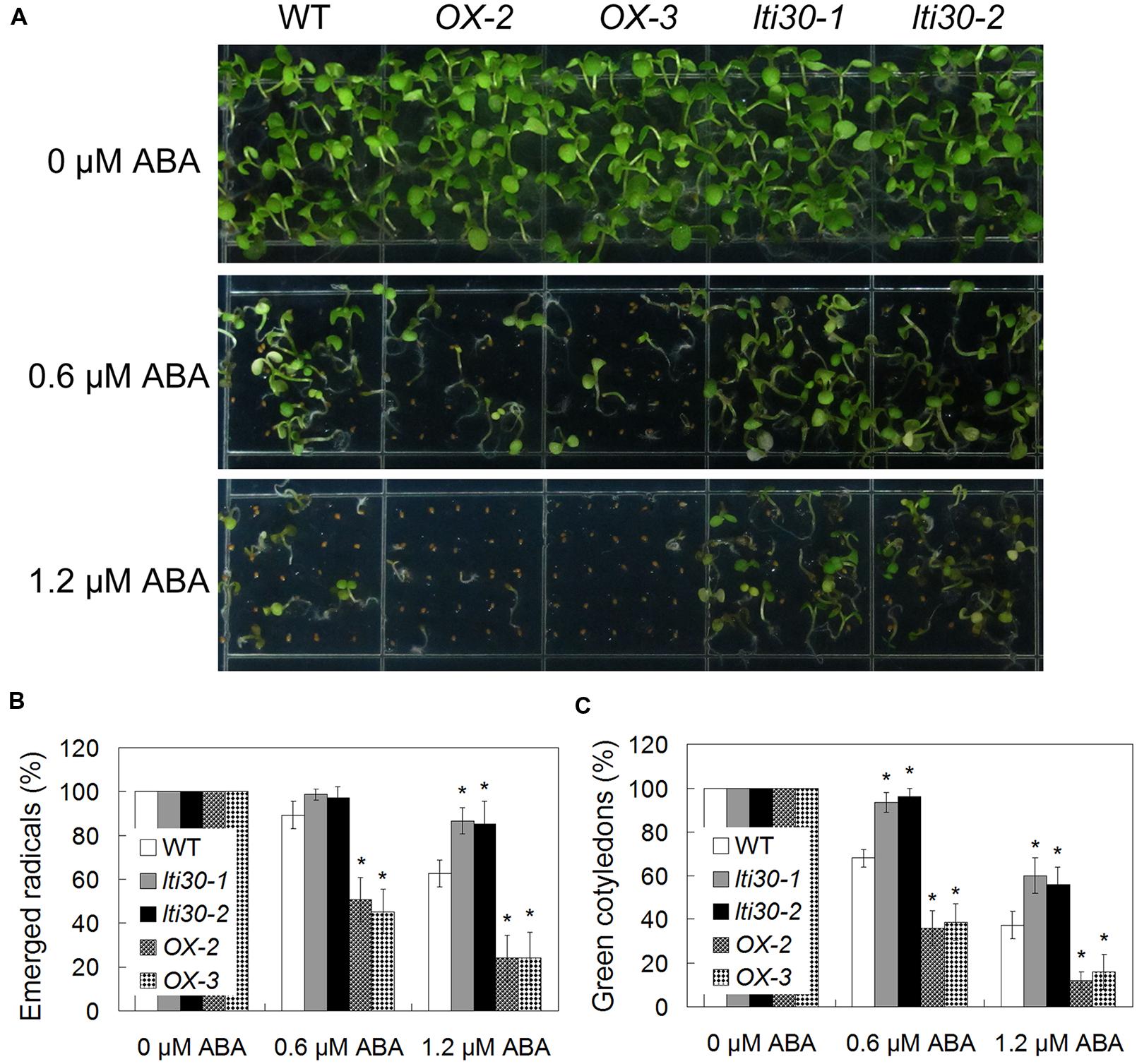
FIGURE 4. Modulation of AtLTI30 expression positively affects plant sensitivity to ABA. (A) The photograph showing 10-day-old of Arabidopsis plant seedlings after grown on MS plate containing different concentrations of ABA. (B,C) Germination rates of plant seeds on MS plate containing different concentrations of ABA that were determined as emerged radicals (B) and green cotyledons (C). The results shown are the means ± SDs of three biological repeats, and asterisk symbols (∗) indicate the significant difference of p < 0.05 in comparison to WT.
AtLTI30 Positively Regulates Drought Stress Resistance
To test whether AtLTI30 regulates drought stress resistance, 14-day-old WT, lti30 mutants, and AtLTI30 overexpressing plants in pots were subjected to drought stress by withholding water for 21 days and then re-watering the plants for 4 days. After the drought stress treatment, the lti30-1 and lti30-2 mutants displayed significantly lower survival rate, while AtLTI30 overexpressing plants exhibited higher survival rate compared with WT (Figures 5A,B). Consistently, the lti30-1 and lti30-2 mutants displayed significantly higher leave water loss rate, while AtLTI30 overexpressing plants showed significantly lower leaf water loss rate from 2 to 8 h after detachment in comparison to WT plants (Figure 5C). These results indicate that AtLTI30 positively regulates drought stress resistance.
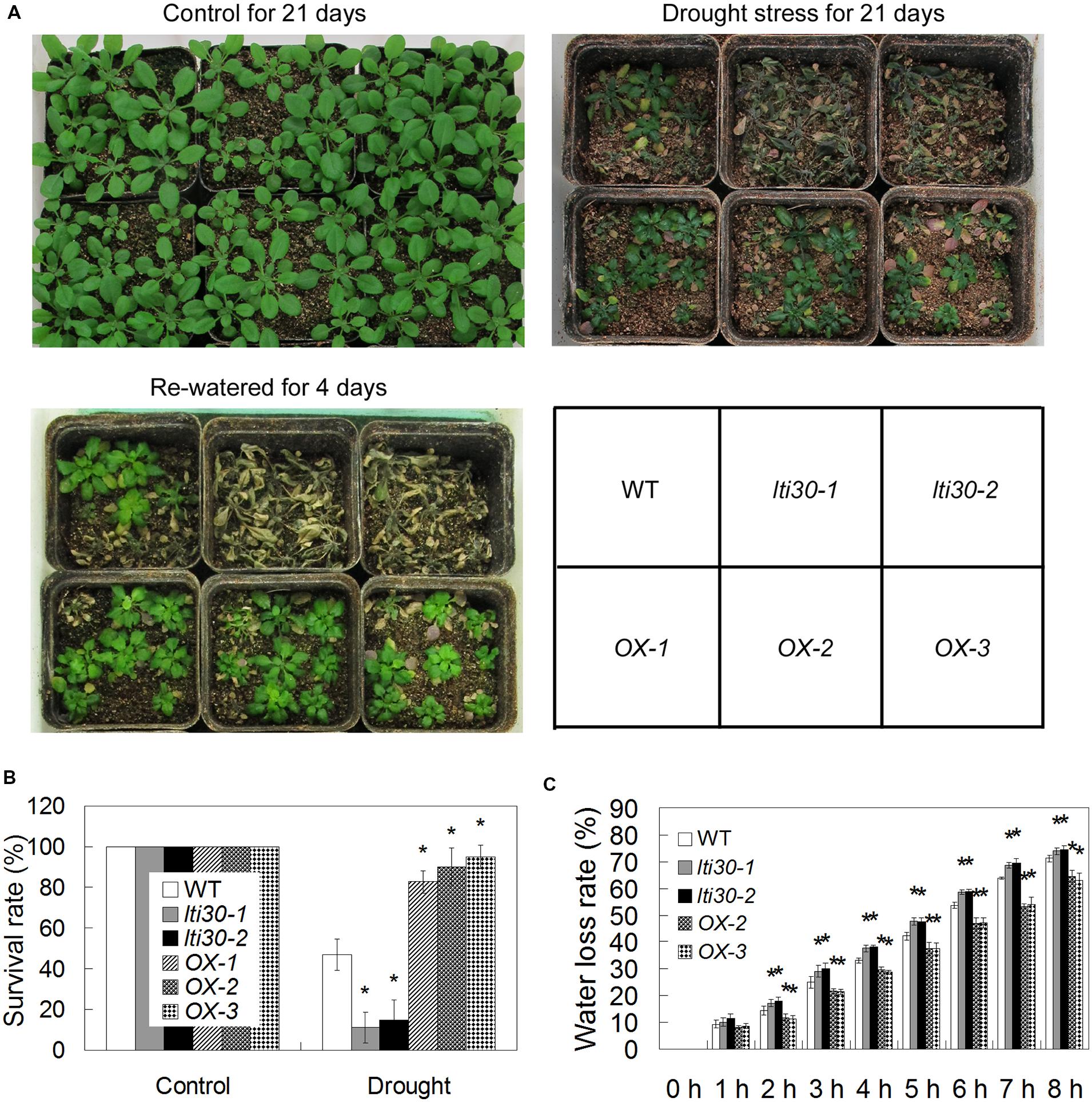
FIGURE 5. AtLTI30 positively regulates drought stress resistance in Arabidopsis. (A) The photograph showing 14-day-old Arabidopsis plants with well-watered for 21 days, withheld water for 21 days, and re-watered for 4 days after withheld water for 21 days. (B) The survival rate of different genotypes after recovery for 4 days after 21 days of drought stress treatment. (C) Relative leaf water loss rate in vitro of different lines. The results shown are the means ± SDs of three biological repeats, and asterisk symbols (∗) indicate the significant difference of p < 0.05 in comparison to WT.
Modulation of AtLTI30 Expression Affects H2O2 Accumulation
Oxidative burst especially H2O2 accumulation occurs following drought stress in plants. We further investigated the effects of AtLTI30 expression on H2O2 accumulation and associated antioxidant defense enzyme activity during the drought stress treatments. During the period between 0 and 15 days of drought stress, H2O2 burst was significantly displayed in WT, the AtLTI30 knockout mutants and overexpressing plants (Figure 6A). In comparison to WT plants, the AtLTI30 knockout mutants showed higher levels of H2O2 at 0, 5, 10, and 15 days of drought stress, while the AtLTI30 overexpressing plants displayed relatively lower levels of H2O2 at these periods (Figure 6A).
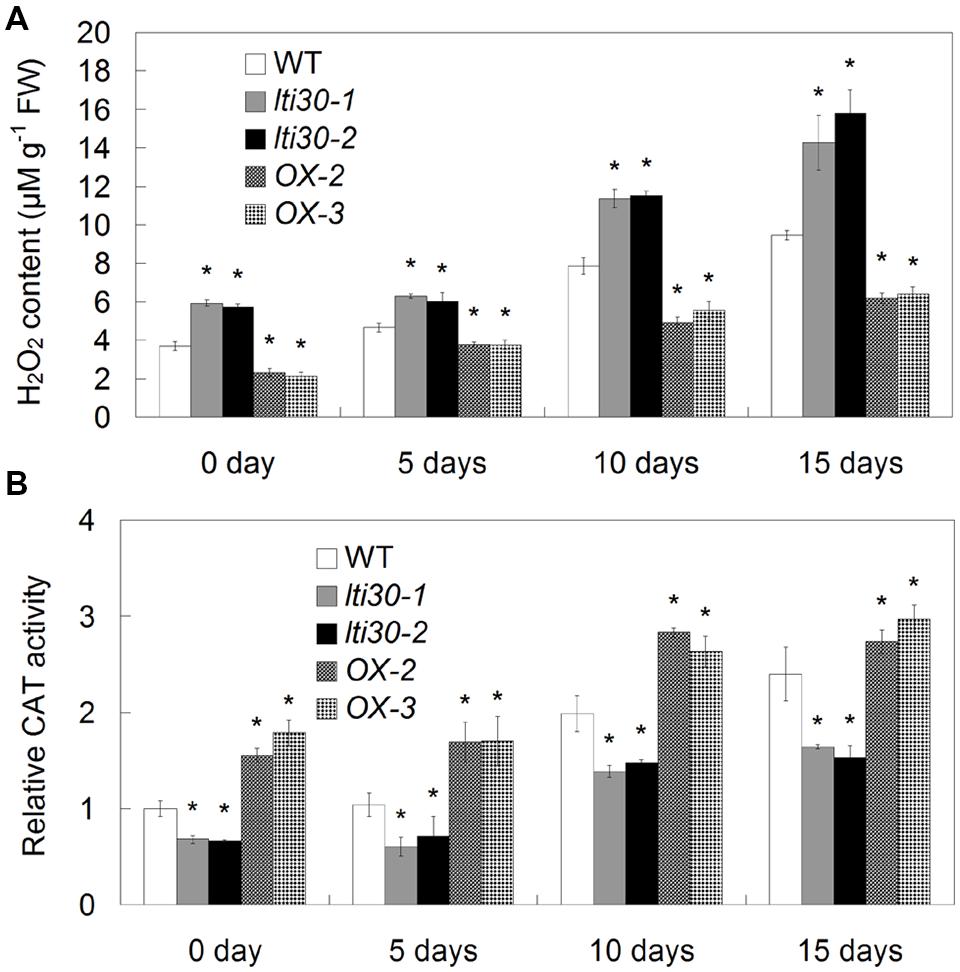
FIGURE 6. Quantifications of H2O2 content (A) and CAT activity (B) in different lines during drought stress treatments. For the assays, 14-d-old plant leaves were subject to drought stress by withheld water for 0, 5, 10, and 15 days. The results shown are the means ± SDs of three biological repeats, and asterisk symbols (∗) indicate the significant difference of p < 0.05 in comparison to WT.
In accordance with the H2O2 burst, the AtLTI30 knockout mutants exhibited lower activities of AtCATs under both control and drought stress conditions, while the AtLTI30 overexpressing plants showed relatively higher activities of AtCATs in comparison to WT (Figure 6B). Therefore, these results indicate that the AtLTI30 positively regulates the activities of AtCATs, and negatively regulates H2O2 accumulation during drought stress treatment.
AtLTI30 Positively Regulates Drought Stress Resistance
During the period between 0 and 15 days of drought stress, the endogenous proline content gradually increased in WT, AtLTI30 knockout mutants and overexpressing plants (Figure 7). In comparison to WT plants, the AtLTI30 knockout mutants exhibited lower proline contents at 0, 5, 10, and 15 days of drought stress, while the AtLTI30 overexpressing plants displayed higher levels of proline at these periods (Figure 7).
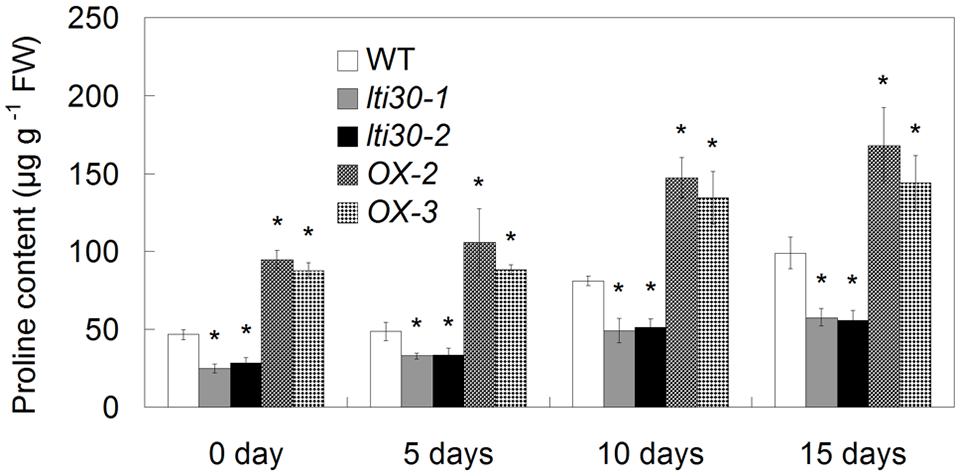
FIGURE 7. Modulation of AtLTI30 expression affects the endogenous proline level. For the assays, 14-d-old plant leaves were subject to drought stress by withheld water for 0, 5, 10, and 15 days. The results shown are the means ± SDs of three biological repeats, and asterisk symbols (∗) indicate the significant difference of p < 0.05 in comparison to WT.
Discussion
As a dehydrin belonging to the group II LEA family, AtLTI30/XERO2 is widely involved in plant freezing stress resistance (Rouse et al., 1996; Nylander et al., 2001). Not only is its transcription induced under cold stress treatment, but its overexpression confers improved freezing stress resistance in Arabidopsis (Rouse et al., 1996; Nylander et al., 2001; Puhakainen et al., 2004). Moreover, the common expression between CBF1/2/3 and AtLTI30/XERO2, and between ABA insensitive 5 (ABI5) and AtLTI30/XERO2, indicated that AtLTI30/XERO2 may be a direct target of both AtCBFs and AtABI5 (Chung and Parish, 2008). Using multiple combinations of mutations in the promoter of AtLTI30/XERO2, they also found that two of the ACGT and CRT/DRE elements are essential for both ABA and cold transcriptional induction of AtLTI30/XERO2 (Chung and Parish, 2008). However, the possible involvement of AtLTI30 in ABA signaling and the in vivo role of AtLTI30 in plant drought stress response remains unknown.
To date, there are three effective methods to improve plant drought stress resistance. The first is to screen and identify drought tolerant varieties, and the second method is the exogenous application of multiple small molecules such as ABA, nitric oxide (NO), polyamines, hydrogen sulfide (H2S), and melatonin (Shi et al., 2013a, 2014a, 2015; Chan and Shi, 2015). The third method is the investigation of new genes that confer plant drought stress resistance and genetic breeding (Miao et al., 2006; Yu et al., 2008; Qin et al., 2011; Zhang et al., 2013). In this study, the new roles of AtLTI30 in drought stress resistance may provide a useful candidate gene for drought tolerant crop genetic breeding.
In response to drought stress, plant endogenous ABA synthesis and the following ABA-responsive genes are rapidly and largely activated. Briefly, with an increase in the endogenous ABA level, ABA receptors (Pyrabactin Resistance (PYR)/PYR1-Like (PYL)/Regulatory Components of ABA Receptor (RCAR)) disrupt the interaction between type 2C protein phosphatases (PP2Cs) and sucrose non-fermenting 1 (SNF1)-related protein kinases 2 (SnRK2s) by competitively interacting with PP2Cs. Thereafter, these interaction prevent PP2Cs-mediated dephosphorylation of SnRK2s and the activation of the SnRK2s, leading to the transcriptional activation of ABA-responsive genes (Fujii et al., 2009; Cutler et al., 2010; Harb et al., 2010). Moreover, ABA also induces the accumulation of H2O2, and both ABA and ABA-induced H2O2 play important roles in plant drought stress response, especially in the modulation of stomatal closure (Zhang et al., 2001; Miao et al., 2006; Munemasa et al., 2013). In accordance with previous studies, which showed that ABA and dehydration induced the transcription level of AtLTI30, as evidenced by northern blot analysis (Welin et al., 1994; Rouse et al., 1996; Nylander et al., 2001; Chung and Parish, 2008), the expression of AtLTI30 was significantly increased after ABA and drought stress treatments using proLTI30::GUS transgenic plants and the publicly available microarray data (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi) (Figure 2). Together with the association among ABA, H2O2 and drought stress, these results indicated the possible role of the AtLTI30 in the process. After identifying the AtLTI30 knockout mutants and the AtLTI30 overexpressing plants (Figure 3), we found that the AtLTI30 positively regulated plant sensitivity to ABA (Figure 4). In accordance with the ABA insensitive phenotype, the lti30-1 and lti30-2 mutants showed decreased drought stress resistance, as evidenced by a higher water loss rate and lower survival rate in comparison to WT (Figure 5). On the contrary, the AtLTI30 overexpressing plants were more sensitive to ABA and exhibited improved drought stress resistance (Figure 5). These results suggest that AtLTI30 may function in drought stress response in an ABA-dependent pathway. However, the stomatal response of the AtLTI30 knockout mutants and the AtLTI30 overexpressing plants displayed no significant difference in comparison to WT plants under mock, ABA and H2O2 conditions (Supplementary Figure S1). This result indicated that modulation of AtLTI30 expression has no significant effect on stomatal response in Arabidopsis.
Reactive oxygen species (ROS) including H2O2, superoxide anions (O2⋅-), singlet oxygen (1O2) and hydroxyl radical (OH-) plays pivotal roles in plant drought stress responses. On one hand, H2O2 is key secondary messenger in drought stress perception and transduction (Zhang et al., 2001; Miao et al., 2006; Munemasa et al., 2013; Wang et al., 2013). On the other hand, as toxic by-products of physiological metabolism, H2O2 accumulation is rapidly and largely increased under drought stress conditions, and overproduction of H2O2 triggers serious oxidative damage and decreased drought resistance (Miller et al., 2010; Mittler et al., 2011). To cope with stress-triggered ROS overproduction and oxidative stress, plants have developed complex defense systems including both enzymatic and non-enzymatic antioxidants. Among the enzymatic enzymes, CAT catalyzes the decomposition of H2O2 into H2O and O2 and plays an essential role in controlling ROS homeostasis. In Arabidopsis, the AtCATs transcripts can be largely induced by various stress treatments including ABA, drought, salt, cold, and oxidative stresses (Du et al., 2008; Mhamdi et al., 2010; Hu et al., 2011). The interactions among nucleoside diphosphate kinase 2 (NDPK2), CAT2 or CAT3 and Salt Overly Sensitive 2 (SOS2) indicate the relationship between H2O2 and abiotic stress response (Verslues et al., 2007). Together with previous studies showing the interaction of SOS2 and other SnRK3s with ABI1 and ABI2 (Guo et al., 2002; Ohta et al., 2003) and the importance of H2O2-dependent inactivation of ABI1 and ABI2 in ABA signaling (Miao et al., 2006), CAT2 and CAT3 may occur in the same protein complex as ABI1 and ABI2, indicating the possible involvement of AtCATs in the ABA signaling pathway. In this study, the positive effect of AtLTI30 expression on the activities of AtCATs may be directly related to drought stress-induced ROS accumulation (Figure 6), as well as AtLTI30-mediated drought stress resistance. Additionally, H2O2 is an important secondary messenger in ABA signal transduction (Miao et al., 2006; Cutler et al., 2010). Thus, the effects of AtLTI30 expression on ABA sensitivity, H2O2 accumulation and drought resistance further suggest the dual cross-talks among these pathways.
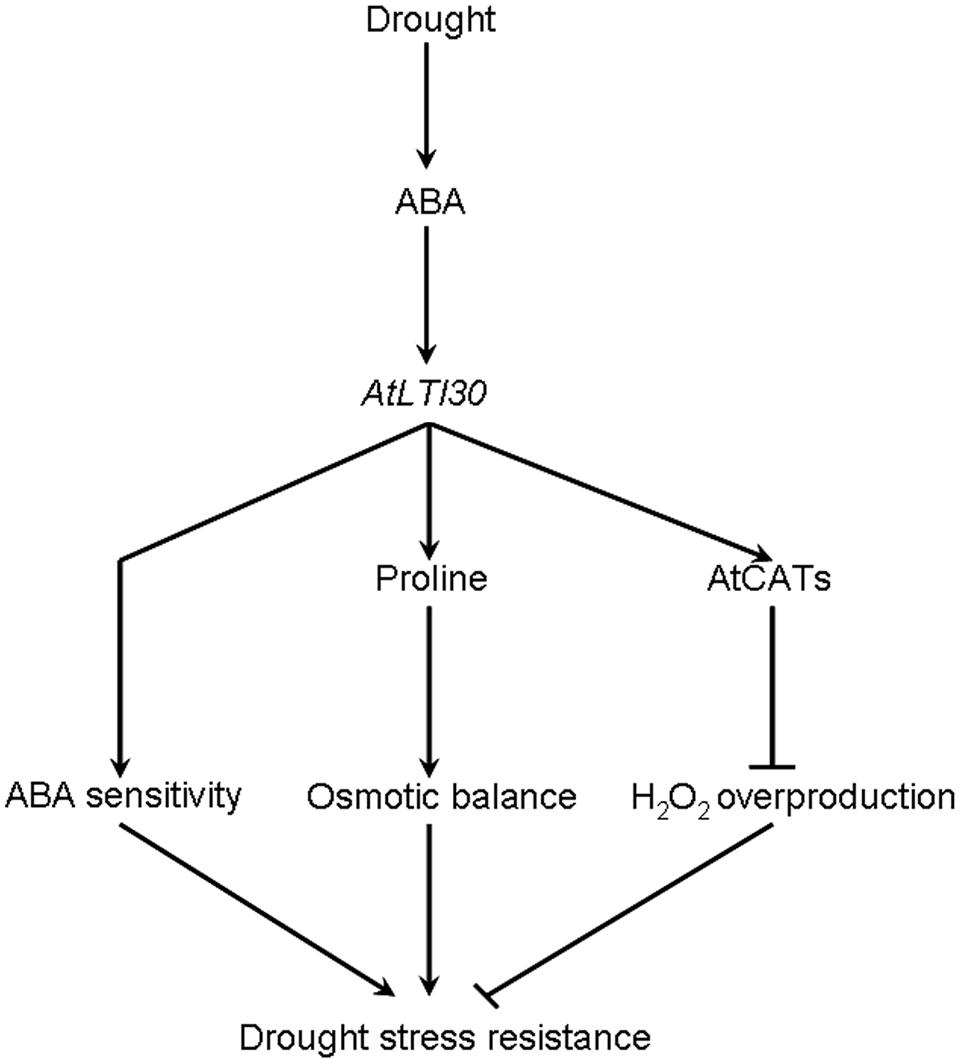
Based on these results, a model for AtLTI30-mediated drought stress response is proposed in this study (Figure 8). In response to drought stress, the endogenous ABA level is rapidly and largely induced, and thereafter induces the expression of AtLTI30. Firstly, overexpression of AtLTI30 conferred ABA sensitivity, which is directly linked with ABA-mediated stress responses. Secondly, overexpression of AtLTI30 up-regulated the activities of AtCATs, leading to less H2O2 accumulation as well as less oxidative damage under the drought stress condition. Moreover, AtLTI30 positively regulated the endogenous level of proline, which functions as an important osmolyte in alleviating osmotic pressure under drought stress conditions (Shi et al., 2013b, 2014a), thereafter resulting in less osmotic pressure in response to drought stress. Thus, the ABA sensitivity, lower H2O2 accumulation and more proline content resulted in enhanced drought stress resistance of AtLTI30 overexpressing plants.
Taken together, these results indicate the possible novel role of AtLTI30 in ABA signaling, ROS accumulation and drought stress resistance. AtLTI30 confers enhanced drought stress resistance in Arabidopsis, by positively regulating ABA sensitivity, CAT activity and proline accumulation, at least partially.
Author Contributors
HS conceived and directed this study, designed and performed the experiments, analyzed the data, wrote and revised the manuscript; YC and YQ performed the experiments, analyzed the data and revised the manuscript; ZC provided suggestions and revised the manuscript. All authors approved the manuscript and the version to be published, and agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This research was supported by the National Natural Science Foundation of China (No.31570249) and the startup funding of Hainan University (No. kyqd1531) to HS, the Basic Research Fund of National Non-profit Research Institutions (No. CAFYBB2012043 and No. RIF2014-03) to YQ, the National Natural Science Foundation of China (No. 31370302) and “the Hundred Talents Program” (No.54Y154761O01076 and No. 29Y329631O0263) to ZC.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2015.00893
References
Chan, Z., and Shi, H. (2015). Improved abiotic stress tolerance of bermudagrass by exogenous small molecules. Plant Signal. Behav. 10:e991577. doi: 10.4161/15592324.2014.991577
Chung, S., and Parish, R. W. (2008). Combinatorial interactions of multiple cis-elements regulating the induction of the Arabidopsis XERO2 dehydrin gene by abscisic acid and cold. Plant J. 54, 15–29. doi: 10.1111/j.1365-313X.2007.03399.x
Clough, S. J., and Bent, A. F. (1998). Floral dip: a simplified method for Agrobacterium mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743. doi: 10.1046/j.1365-313x.1998.00343.x
Cook, D., Fowler, S., Fiehn, O., and Thomashow, M. F. (2004). A prominent role for the CBF cold response pathway in configuring the low-temperature metabolome of Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 101, 15243–15248. doi: 10.1073/pnas.0406069101
Cutler, S. R., Rodriguez, P. L., Finkelstein, R. R., and Abrams, S. R. (2010). Abscisic acid: emergence of a core signaling network. Annu. Rev. Plant Biol. 61, 651–679. doi: 10.1146/annurev-arplant-042809-112122
Du, Y. Y., Wang, P. C., Chen, J., and Song, C. P. (2008). Comprehensive functional analysis of the catalase gene family in Arabidopsis thaliana. J. Integr. Plant Biol. 50, 1318–1326. doi: 10.1111/j.1744-7909.2008.00741.x
Eriksson, S. K., Kutzer, M., Procek, J., Gröbner, G., and Harryson, P. (2011). Tunable membrane binding of the intrinsically disordered dehydrin Lti30, a cold-induced plant stress protein. Plant Cell 23, 2391–2404. doi: 10.1105/tpc.111.085183
Fujii, H., Chinnusamy, V., Rodrigues, A., Rubio, S., Antoni, R., Park, S. Y., et al. (2009). In vitro reconstitution of an abscisic acid signalling pathway. Nature 462, 660–664. doi: 10.1038/nature08599
Gilmour, S. J., Zarka, D. G., Stockinger, E. J., Salazar, M. P., Houghton, J. M., and Thomashow, M. F. (1998). Low temperature regulation of the Arabidopsis CBF family of AP2 transcriptional activators as an early step in cold-induced COR gene expression. Plant J. 16, 433–442. doi: 10.1046/j.1365-313x.1998.00310.x
Guo, Y., Xiong, L. M., Song, C. P., Gong, D. M., Halfter, U., and Zhu, J. K. (2002). A calcium sensor and its interacting protein kinase are global regulators of abscisic acid signaling in Arabidopsis. Dev. Cell 3, 233–244. doi: 10.1016/S1534-5807(02)00229-0
Harb, A., Krishnan, A., Ambavaram, M. M. R., and Pereira, A. (2010). Molecular and physiological analysis of drought stress in Arabidopsis reveals early responses leading to acclimation in plant growth. Plant Physiol. 154, 1254–1271. doi: 10.1104/pp.110.161752
Hu, Y. Q., Liu, S., Yuan, H. M., Li, J., Yan, D. W., Zhang, J. F., et al. (2011). Functional comparison of catalase genes in the elimination of photorespiratory H2O2 using promoter- and 3’-untranslated region exchange experiments in the Arabidopsis cat2 photorespiratory mutant. Plant Cell Environ. 33, 1656–1670. doi: 10.1111/j.1365-3040.2010.02171.x
Jefferson, R. A., Kavanagh, T. A., and Bevan, M. W. (1987). GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907.
Mhamdi, A., Queval, G., Chaouch, S., Vanderauwera, S., Breusegem, F. V., and Noctor, G. (2010). Catalase function in plants: a focus on Arabidopsis mutants as stress-mimic models. J. Exp. Bot. 61, 4197–4220. doi: 10.1093/jxb/erq282
Miao, Y., Lv, D., Wang, P., Wang, X. C., Chen, J., Miao, C., et al. (2006). An Arabidopsis glutathione peroxidase functions as both a redox transducer and a scavenger in abscisic acid and drought stress responses. Plant Cell 18, 2749–2766. doi: 10.1105/tpc.106.044230
Miller, G., Suzuki, N., Ciftci-Yilmaz, S., and Mittler, R. (2010). Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ. 33, 453–467. doi: 10.1111/j.1365-3040.2009.02041.x
Mittler, R., Vanderauwera, S., Suzuki, N., Miller, G., Tognetti, V. B., Vandepoele, K., et al. (2011). ROS signaling: the new wave. Trends Plant Sci. 16, 1360–1385. doi: 10.1016/j.tplants.2011.03.007
Mouillon, J. M., Gustafsson, P., and Harryson, P. (2006). Structural investigation of disordered stress proteins. comparison of full-length dehydrins with isolated peptides of their conserved segments. Plant Physiol. 141, 638–650. doi: 10.1104/pp.106.079848
Munemasa, S., Muroyama, D., Nagahashi, H., Nakamura, Y., Mori, I. C., and Murata, Y. (2013). Regulation of reactive oxygen species-mediated abscisic acid signaling in guard cells and drought tolerance by glutathione. Front. Plant Sci. 4:472. doi: 10.3389/fpls.2013.00472
Nylander, M., Svensson, J., Palva, E. T., and Welin, B. V. (2001). Stress-induced accumulation and tissue-specific localization of dehydrins in Arabidopsis thaliana. Plant Mol. Biol. 45, 263–279. doi: 10.1023/A:1006469128280
Ohta, M., Guo, Y., Halfter, U., and Zhu, J. K. (2003). A novel domain in the protein kinase SOS2 mediates interaction with the protein phosphatase 2C ABI2. Proc. Natl. Acad. Sci. U.S.A. 100, 11771–11776. doi: 10.1073/pnas.2034853100
Puhakainen, T., Hess, M. W., Mäkelä, P., Svensson, J., Heino, P., and Palva, E. T. (2004). Overexpression of multiple dehydrin genes enhances tolerance to freezing stress in Arabidopsis. Plant Mol. Biol. 54, 743–753. doi: 10.1023/B:PLAN.0000040903.66496.a4
Qin, F., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2011). Achievements and challenges in understanding plant abiotic stress responses and tolerance. Plant Cell Physiol. 52, 1569–1582. doi: 10.1093/pcp/pcr106
Rouse, D. T., Marotta, R., and Parish, R. W. (1996). Promoter and expression studies on an Arabidopsis thaliana dehydrin gene. FEBS Lett. 381, 252–256. doi: 10.1016/0014-5793(96)00051-8
Seki, M., Umezawa, T., Urano, K., and Shinozaki, K. (2007). Regulatory metabolic networks in drought stress responses. Curr. Opin. Plant Biol. 10, 296–302. doi: 10.1016/j.pbi.2007.04.014
Shi, H., Ye, T., Chen, F., Cheng, Z., Wang, Y., Yang, P., et al. (2013a). Manipulation of arginase expression modulates abiotic stress tolerance in Arabidopsis: effect on arginine metabolism and ROS accumulation. J. Exp. Bot. 64, 1367–1379. doi: 10.1093/jxb/ers400
Shi, H., Ye, T., Wang, Y., and Chan, Z. (2013b). Arabidopsis ALTERED MERISTEM PROGRAM 1 negatively modulates plant responses to abscisic acid and dehydration stress. Plant Physiol. Biochem. 67, 209–216. doi: 10.1016/j.plaphy.2013.03.016
Shi, H., Ye, T., Han, N., Bian, H., Liu, X., and Chan, Z. (2015). Hydrogen sulfide regulates abiotic stress tolerance and biotic stress resistance in Arabidopsis. J. Integr. Plant Biol. 57, 628–640. doi: 10.1111/jipb.12302
Shi, H., Ye, T., Zhu, J. K., and Chan, Z. (2014a). Constitutive production of nitric oxide leads to enhanced drought stress resistance and extensive transcriptional reprogramming in Arabidopsis. J. Exp. Bot. 65, 4119–4131. doi: 10.1093/jxb/eru184
Shi, H., Wang, X., Ye, T., Chen, F., Deng, J., Yang, P., et al. (2014b). The Cysteine2/Histidine2-type transcription factor ZINC FINGER OF Arabidopsis thaliana 6 modulates biotic and abiotic stress responses by activating salicylic acid-related genes and C-REPEAT-BINDING FACTOR genes in Arabidopsis. Plant Physiol. 165, 1367–1379. doi: 10.1104/pp.114.242404
Thomashow, M. F. (2010). Molecular basis of plant cold acclimation: insights gained from studying the CBF cold response pathway. Plant Physiol. 154, 571–577. doi: 10.1104/pp.110.161794
Verslues, P. E., Batelli, G., Grillo, S., Agius, F., Kim, Y. S., Zhu, J., et al. (2007). Interaction of SOS2 with nucleoside diphosphate kinase 2 and catalases reveals a point of connection between salt stress and H2O2 signaling in Arabidopsis thaliana. Mol. Cell. Biol. 27, 7771–7780. doi: 10.1128/MCB.00429-07
Wang, P., Du, Y., Zhao, X., Miao, Y., and Song, C. P. (2013). The MPK6-ERF6-ROSE7/GCC-box complex modulates oxidative gene transcription and the oxidative response in Arabidopsis thaliana. Plant Physiol. 161, 1392–1408. doi: 10.1104/pp.112.210724
Welin, B. V., Olson, A., Nylander, M., and Palva, E. T. (1994). Characterization and differential expression of dhn/lea/rab-like genes during cold acclimation and drought stress in Arabidopsis thaliana. Plant Mol. Biol. 26, 131–144. doi: 10.1007/BF00039526
Winter, D., Vinegar, B., Nahal, H., Ammar, R., Wilson, G. V., and Provart, N. J. (2007). An “Electronic Fluorescent Pictograph” browser for exploring and analyzing large-scalebiological data sets. PLoS ONE 2:e718. doi: 10.1371/journal.pone.0000718
Yang, L. X., Wang, R. Y., Ren, F., Liu, J., Cheng, J., and Lu, Y. T. (2005). AtGLB1 enhances the tolerance of Arabidopsis to hydrogen peroxide stress. Plant Cell Physiol. 46, 1309–1316. doi: 10.1093/pcp/pci140
Yu, H., Chen, X., Hong, Y. Y., Wang, Y., Xu, P., Ke, S. D., et al. (2008). Activated expression of an Arabidopsis HD-START protein confers drought tolerance with improved root system and reduced stomatal density. Plant Cell 20, 1134–1151. doi: 10.1105/tpc.108.058263
Zarka, D. G., Vogel, J. T., Cook, D., and Thomashow, M. F. (2003). Cold induction of Arabidopsis CBF genes involves multiple ICE (inducer of CBF expression) promoter elements and a cold-regulatory circuit that is desensitized by low temperature. Plant Physiol. 133, 910–918. doi: 10.1104/pp.103.027169
Zhang, S., Qi, Y., Liu, M., and Yang, C. (2013). SUMO E3 ligase AtMMS21 regulates drought tolerance in Arabidopsis thaliana. J. Integr. Plant Biol. 55, 83–95. doi: 10.1111/jipb.12024
Keywords: Low Temperature-Induced 30, abscisic acid, hydrogen peroxide, drought stress, reactive oxygen species, Arabidopsis
Citation: Shi H, Chen Y, Qian Y and Chan Z (2015) Low Temperature-Induced 30 (LTI30) positively regulates drought stress resistance in Arabidopsis: effect on abscisic acid sensitivity and hydrogen peroxide accumulation. Front. Plant Sci. 6:893. doi: 10.3389/fpls.2015.00893
Received: 17 August 2015; Accepted: 08 October 2015;
Published: 20 October 2015.
Edited by:
Zuhua He, Shanghai Institute for Biological Sciences, ChinaReviewed by:
Nabil I. Elsheery, Tanta University, EgyptZhaoqing Chu, Shanghai Chenshan Plant Science Research Center, China
Copyright © 2015 Shi, Chen, Qian and Chan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Haitao Shi, aGFpdGFvc2hpQGhhaW51LmVkdS5jbg==; Zhulong Chan, emh1bG9uZ2NoQHdiZ2Nhcy5jbg==
†These authors have contributed equally to this work.
 Haitao Shi
Haitao Shi Yinhua Chen
Yinhua Chen Yongqiang Qian
Yongqiang Qian Zhulong Chan
Zhulong Chan