- Key Laboratory of Molecular Cytogenetics and Genetic Breeding of Heilongjiang Province, College of Life Science and Technology, Harbin Normal University, Harbin, China
The AP2/ERF superfamily is a large, plant-specific transcription factor family that is involved in many important processes, including plant growth, development, and stress responses. Using Medicago truncatula genome information, we identified and characterized 123 putative AP2/ERF genes, which were named as MtERF1–123. These genes were classified into four families based on phylogenetic analysis, which is consistent with the results of other plant species. MtERF genes are distributed throughout all chromosomes but are clustered on various chromosomes due to genomic tandem and segmental duplication. Using transcriptome, high-throughput sequencing data, and qRT-PCR analysis, we assessed the expression patterns of the MtERF genes in tissues during development and under abiotic stresses. In total, 87 MtERF genes were expressed in plant tissues, most of which were expressed in specific tissues during development or under specific abiotic stress treatments. These results support the notion that MtERF genes are involved in developmental regulation and environmental responses in M. truncatula. Furthermore, a cluster of DREB subfamily members on chromosome 6 was induced by both cold and freezing stress, representing a positive gene regulatory response under low temperature stress, which suggests that these genes might contribute to freezing tolerance to M. truncatula. In summary, our genome-wide characterization, evolutionary analysis, and expression pattern analysis of MtERF genes in M. truncatula provides valuable information for characterizing the molecular functions of these genes and utilizing them to improve stress tolerance in plants.
Introduction
On a global scale, plant growth, and development are threatened by various abiotic stresses, such as extreme temperatures, drought, and high salinity. Due to environmental conditions continuously accumulating, plants are being confronted with increasingly serious challenges to their survival. Plants employ complex regulatory mechanisms to adapt to environmental stresses, undergoing physiological and biochemical changes in response to unfavorable conditions (Zhu, 2002; Chinnusamy et al., 2004; Katagiri, 2004). It is notable that plants contain numerous genes encoding transcription factors (TFs), which regulate the expression of downstream genes by binding to their cis-acting elements (Singh et al., 2002; Yamaguchi-Shinozaki and Shinozaki, 2006; Le Hir and Bellini, 2013). TFs play important roles in plant growth, development and responses to environmental stress by directly responding to stress or regulating the expression of downstream target genes, confirming their critical roles in plant life cycles. However, only a few TF families have been characterized outside of well-studied model plant systems, such as rice and Arabidopsis. The APETALA2/ethylene-responsive element binding factor (AP2/ERF) superfamily is one of the largest groups of TFs in plants. These TFs contain at least one AP2 domain. Based on the number of AP2 domains and other DNA binding domains, AP2/ERF TFs are classified into four families, including the AP2, ERF, RAV, and Soloist families (Cao et al., 2001; Sakuma et al., 2002; Mizoi et al., 2012). AP2 family members contain a double, tandemly repeated AP2 domain, while ERF family members contain a single AP2 domain. RAV family members have a single AP2 domain and an additional B3 domain, i.e., a DNA-binding domain commonly found in other TFs (Nakano et al., 2006; Mizoi et al., 2012; Li et al., 2015).
AP2/ERF TFs regulate a number of biological processes, such as plant growth, development, and responses to stress (Mizoi et al., 2012; Matías-Hernández et al., 2014). AP2 family members regulate the development of various plant tissues. For example, CRL5 is expressed in the stem region and regulates the crown root initiation process in rice (Kitomi et al., 2011). Some members of the AP2 family regulate reproductive development in Arabidopsis, including flower, ovule, and sepal development (Kunst et al., 1989; Krizek, 2009). AP2 TFs are also involved in fruit development and the ripening process in tomato and grapevine (Chung et al., 2010; Licausi et al., 2010; Sharma et al., 2010). The ERF family consists of two subfamilies (ERF and DREB), which participate in many developmental and stress response processes. ERF subfamily members, which bind to GCC-boxes, are involved in hormone signaling pathways, such as the ethylene, jasmonic acid, and salicylic acid pathways, which are important for plant development and stress responses (Fujimoto et al., 2000; Oñate-Sánchez and Singh, 2002; Andriankaja et al., 2007; Mantiri et al., 2008a). DREB subfamily members bind to dehydration-responsive element/C-repeat (DRE/CRT) elements, which are present in stress-responsive genes, including RD (responsive to desiccation, RD29) and COR (cold-regulated, COR15) genes (Cao et al., 2001; Xu et al., 2011). Therefore, many members of the DREB subfamily improve the stress tolerance of various plants under different environmental stress, including cold, drought, salinity stress, and so on.
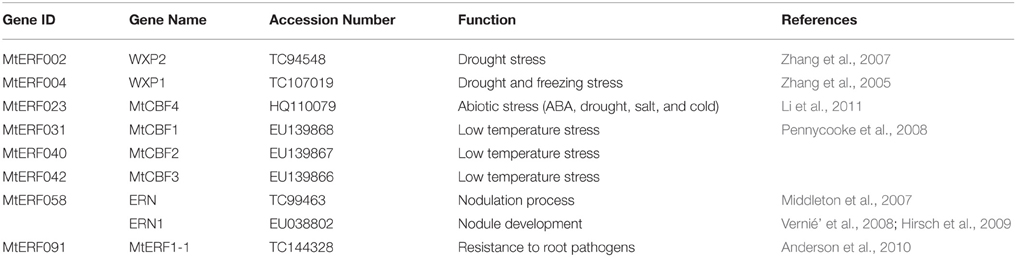
Since the release of the whole-genome sequences of many plant species, the AP2/ERF superfamily has been successfully identified and investigated in plants including Arabidopsis, rice (Nakano et al., 2006), grapevine (Licausi et al., 2010), poplar (Zhuang et al., 2008), and soybean (Zhang et al., 2008). Genome-wide analysis of AP2/ERF TFs has helped elucidate their regulatory functions in plant growth, development and especially stress responses. Medicago truncatula is an annual, diploid legume plant. Due to its features, such as its small genome, self-pollination, high genetic transformation efficiency, nitrogen fixation, and so on, this plant has been chosen as a model legume for molecular genetic and genomic analyses. To date, only 37 AP2/ERF genes were identified based on EST sequences, which is far fewer than the number identified in any other plant species (Zhang et al., 2013b). The functions of AP2/ERF superfamily members in M. truncatula, especially those in responses to abiotic stress, have hardly been reported. ERN and ERN1, with an AP2 domain, were isolated from M. truncatula. Many studies have demonstrated that they have a regulation function in the process of nodulation, as do ERN2 and ERN3 (Middleton et al., 2007; Vernié et al., 2008; Hirsch et al., 2009). MtSERF1, regulated by the plant hormones auxin and cytokinin, plays an important role in somatic embryogenesis (Mantiri et al., 2008a,b). MtERF1-1, a member of the AP2/ERF B3 subgroup, has been shown to mediate resistance to root pathogens in M. truncatula (Anderson et al., 2010). WXP1 and WXP2 can improve drought tolerance in transgenic alfalfa by increasing leaf wax accumulation (Zhang et al., 2005, 2007). Pennycooke et al. (2008) isolated MtCBF1-3 from M. truncatula, and demonstrated their action against low temperature stress. MtCBF4 has been shown to respond to abiotic stresses, including cold, drought, salt, and ABA (Li et al., 2011). Interestingly, 12 potential MtCBF genes clustering on chromosome 6 have been shown to play a major role in tolerance to freezing by QTL mapping (Tayeh et al., 2013). However, their mechanism of doing so, has not yet been discovered. Their expression profiles during freezing remain to be determined. Recently, a draft of the M. truncatula genome sequence was completed and released (Young et al., 2011). A number of gene families have been analyzed based on this genome information, such as ARF and CCCH families, which were promoting Medicago Genus and legume genetic research (Zhang et al., 2013a; Gujaria-Verma et al., 2014; Shen et al., 2015).
In this study, we performed a comprehensive analysis of the AP2/ERF superfamily in M. truncatula, including phylogenetic analysis, chromosomal localization, gene duplication analysis, and expression profiling. We also characterized the functions of these TFs in the abiotic stress response via transcriptome analysis. The results of this study will be helpful for future investigations aimed at the functional characterization of these AP2/ERF TFs and their utilization for the genetic improvement of legumes.
Materials and Methods
Identification and Classification of the AP2/ERF Genes in Medicago truncatula
M. truncatula genome and proteins sequences were downloaded from the JCVI website (M. truncatula Genome Project v4.0, http://www.jcvi.org/medicago/; Young et al., 2011), and Arabidopsis AP2/ERF gene sequences were downloaded from the DATF database (http://datf.cbi.pku.edu.cn; Guo et al., 2005). These Arabidopsis AP2/ERF sequences were utilized for BLAST (Altschul et al., 1990) searches against the M. truncatula genome sequence with the parameters of expected values ≤1E-3 and more than 80% coverage. All BLAST hits were retrieved and searched using the Hidden Markov Model (HMM) profile of the AP2 domain (PF002701), which was downloaded from the Pfam website (pfam.sanger.ac.uk; Finn et al., 2014). The AP2/ERF sequences were confirmed based on the presence of an AP2 domain, and all of the putative AP2/ERF proteins were aligned to Arabidopsis AP2/ERF proteins to classify them into different groups, as described by Nakano et al. (2006). Furthermore, all of the annotation information about putative AP2/ERF genes was retrieved from the M. truncatula genome website, and the number and distribution of introns in AP2/ERF genes were investigated using M. truncatula genome annotation information.
Phylogenetic and Conserved Motif Analysis of the AP2/ERF Genes
Multiple alignments of candidate AP2/ERF protein sequences were carried out using ClustalW with default parameters (Thompson et al., 2002). Unrooted phylogenetic trees of all AP2/ERF proteins were generated with MEGA (V4.0) using the neighbor-joining (NJ) method with the following parameters: Poisson correction, pair-wise deletion and 1000 bootstrap replicates (Tamura et al., 2007). Conserved motifs in M. truncatula AP2/ERF TFs were identified using the motif finding tool MEME (Multiple EM for Motif Elicitation, V4.8.1; Bailey et al., 2006). MEME searching was performed across MtERF protein sequencing using the following parameters: (1) optimum motif width was set to ≥10 and ≤200; (2) the maximum number of motifs was set to identify 25 motifs; (3) occurrences of a single motif distributed among the sequences with model: zero or one per sequence (-mod zoops).
Chromosomal Localization and Gene Duplication Analysis of the AP2/ERF Genes
Positional information about all of the AP2/ERF genes was investigated, and diagrams of their chromosome locations in M. truncatula were drawn using the Circos software (http://circos.ca/; Krzywinski et al., 2009), revealing duplications between AP2/ERF genes in M. truncatula. If two genes with similarities of more than 85% were separated by four or fewer gene loci, they were identified as tandem duplications (TD). Others were identified as segmental duplications (SD), separated by more than five genes. In addition, duplications between the AP2/ERF genes were also identified and complemented using the PGDD database (http://chibba.agtec.uga.edu/duplication/; Lee et al., 2013). Duplicated genes between different chromosomes or loci were linked with colored lines in the diagrams using the Circos as previously described.
In silico Expression Analysis of the AP2/ERF Genes during Plant Development
Genome-wide transcriptome data from M. truncatula in different tissues during development were downloaded from the NCBI short read archive database (SRA database; http://www.ncbi.nlm.nih.gov, Accession numbers SRX099057–SRX099062). The transcriptome data were derived from six tissues, including roots, nodules, blades, buds, seedpods, and flowers. All transcriptome data were mapped to the M. truncatula genome using the TopHat (Trapnell et al., 2009), and the expression of MtERF genes was evaluated using the software Cufflinks as previously described (Trapnell et al., 2012). The expression data were analyzed and clustered using hierarchical cluster programs HCLUST of R and CLUSTERGRAM of Matlab (MathWorks, R2012a).
Plant Material and Stress Treatments
Seeds of M. truncatula (cv. Jemalong A17) were germinated and transferred to a mixture of perlite and sand (3:1, V/V). All seedlings were grown in a growth chamber (Conviron E15) at a temperature of 18 (night) and 24°C (day), a relative humidity of 60–80% and a 14/10 h photoperiod (daytime, 06:00–20:00). The seedlings were irrigated with half-strength Hoagland solution once every 2 days, and after 8 weeks, they were randomly divided into six groups for stress treatments. For cold stress (B group) and freezing stress (C group) treatment, the seedlings were transferred into another chamber with the temperature set at 4 or −8°C, respectively. For drought stress (D group) and salt stress (E group) treatment, the seedlings were treated with 300 mM mannitol or 200 mM NaCl solution, respectively. For ABA treatment (F group), the seedlings' leaves were sprayed with 100 μM ABA solution. Control (untreated, A group) and treated (B–F groups) seedlings were harvested at 3 h after treatment. For each group, five randomly chosen whole seedlings were pooled to form a biological replicate. All plant samples were frozen in liquid nitrogen and stored at −80°C until use.
Transcriptome Analysis of the Response of the AP2/ERF Genes to Abiotic Stress
Total RNA was extracted from six samples (one biological replicate sample per group) using the RNeasy Plant Mini Kit (Qiagen, Valencia, CA) following the manufacturer's instructions. The integrity of the RNA was assessed by formaldehyde agarose gel electrophoresis. Total RNA was quantified using a NanoDrop ND-1000 spectrophotometer (Thermo Fisher Scientific, Wilmington, DE, USA) and a Bioanalyzer 2100 (Agilent Technologies, CA). RNA Integrity Number (RIN) values were greater than 8.0 for all samples. Purified RNA samples were sent to BGI-Shenzhen Ltd. (Shenzhen, China) for construction of pair-end cDNA libraries and Illumina sequencing of abiotic stress-treated samples. Processing of raw data, removal of adapter sequences, base-calling, and quality value calculations were performed to produce clean data. Clean reads from six samples were mapped to the M. truncatula genome, and splice junctions were mapped using the TopHat. MtERF gene expression across six treatment samples (groups A–F) were evaluated using the Cufflinks software, and clustered using hierarchical cluster programs HCLUST of R and CLUSTERGRAM of Matlab (Mathworks, R2012a). Compared to controls, MtERF genes with fold changes ≥2 or ≤0.5 were identified as differentially expressed in response to abiotic stresses. The expression of MtERF genes was analyzed and visualized using the package ggplot2 of R platform.
qRT-PCR Validation of the AP2/ERF Genes Response to Abiotic Stress
Total RNA was isolated using the total RNA kit (Tiangen, Beijing, China) and then reverse transcribed into cDNA using the PrimeScript RT reagent Kit (Toyobo, Shanghai, China). qRT–PCR was performed using ABI 7300 Real-time Detection System (Applied Biosystems, USA) with SYBR Premix Ex TaqTM II (Toyobo, Shanghai, China). The PCR conditions were set as follows: 95°C for 2 min; 40 cycles of 95°C for 30 s and 55°C for 30 s; and 72°C for 1 min, and the experiments were repeated three biological replicates. The ΔΔCT method was used to calculate relative expression levels of MtERF genes using GAPDH as reference gene. Primers of nine MtERF genes (randomly selected from DREB subfamily) and GAPDH gene used for qRT-PCR detection are listed in Table S1.
Results
Identification of the AP2/ERF TFs in Medicago truncatula
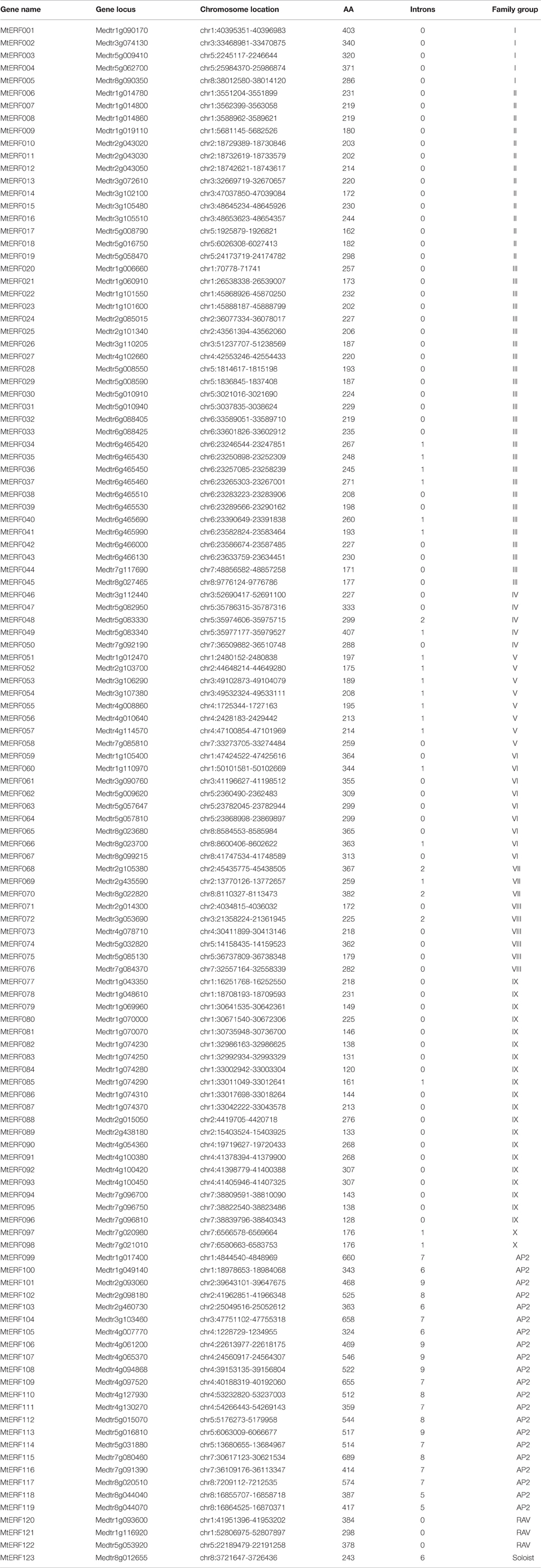
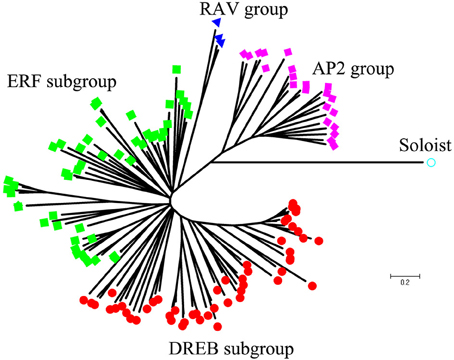
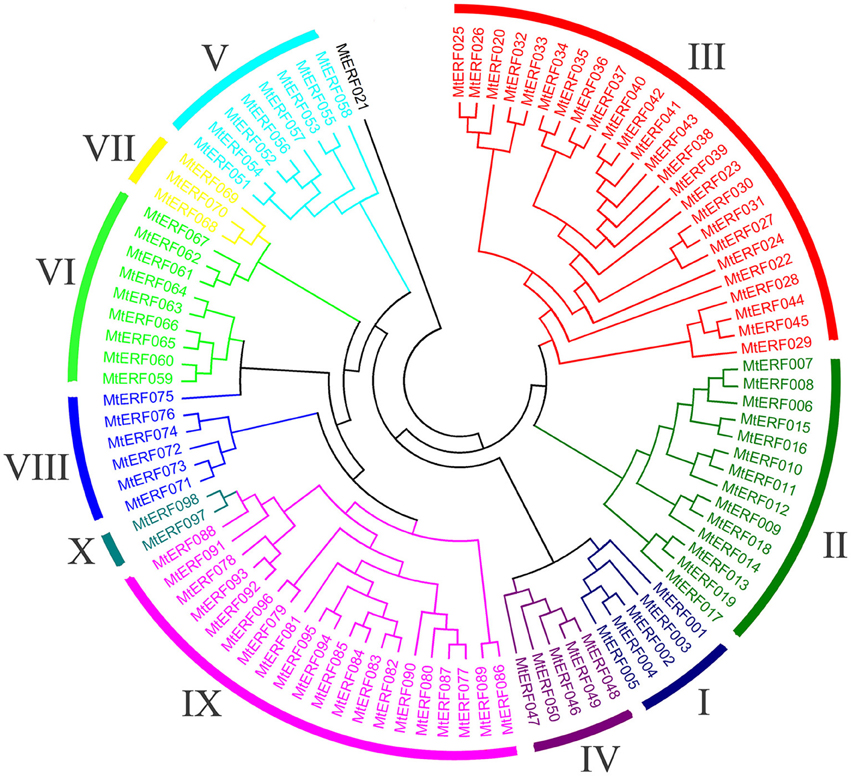
Using homology searches and domain confirmation, we identified 123 putative AP2/ERF TF genes in M. truncatula, designated as MtERF001 to MtERF123, and we determined that these AP2/ERF genes encode putative proteins ranging from 120 to 689 aa in length (see Table 1). Among these TF genes, 98 genes with a single AP2/ERF domain were assigned to the ERF family, and based on the similarity of their encoded amino acid sequences, these genes were further classified into two subfamilies: 50 genes were identified as DREB subfamily members and 48 genes were identified as ERF subfamily members. Of the remaining MtERF genes, 21 genes were grouped into the AP2 family due to their tandemly repeated double AP2/ERF domain, and three genes were classified as RAV family members, as they encode proteins containing a single AP2/ERF domain together with a B3 domain. MtERF123 is homologous to the Arabidopsis Soloist gene (At4g13040) and was therefore designated as Soloist, as shown in Figure 1. As Nakano et al. (2006) previously described, the genes of the ERF family can be subdivided into 10 groups according to their similarity to Arabidopsis ERF sequences, as shown in Figure 2. The DREB subfamily includes group I–IV, containing five, 14, 26, and five members, respectively, while the ERF subfamily consisted of group V–X, with eight, nine, three, six, 21, and two members, respectively. By BLAST search, nine AP2/ERF genes previously identified and totally characterized were also confirmed in the present study, see Table 2. The AP2/ERF family in M. truncatula has relatively few members compared with other plants, such as Arabidopsis (147), soybean (148), grapevine (149), rice (180), and poplar (202) (Nakano et al., 2006; Zhang et al., 2008; Zhuang et al., 2008; Licausi et al., 2010). The numbers of AP2 and RAV family members varies little among species, ranging from 18 to 29 and from three to six, respectively, which contributes little to the reduction in the total number identified in M. truncatula. This small number may arise from the reduced number of ERF family members, as there are 98 members of the ERF family in M. truncatula and 122, 122, 169, and 145 members in Arabidopsis, grapevine, soybean, rice, and poplar, respectively.
Phylogenetic and Conserved Motif Analysis of the AP2/ERF TFs in Medicago truncatula
To determine the evolutionary relationships between AP2/ERF family proteins in M. truncatula, a phylogenetic tree was constructed based on alignment of full-length sequences of MtERF proteins. The phylogenetic tree confirms that MtERF TFs could be classified into four groups, shown as Figure 1, which is consistent with the classification results obtained using homology searches as described above. The phylogenetic relationships of the 98 ERF family members were assessed in depth. The results show that these genes (except MtERF21) could be divided into 10 groups, as described by Nakano et al. (2006). As shown in Figure 2, groups I–IV were identified as DREB subfamily members, while groups V–X were characterized as ERF subfamily members.
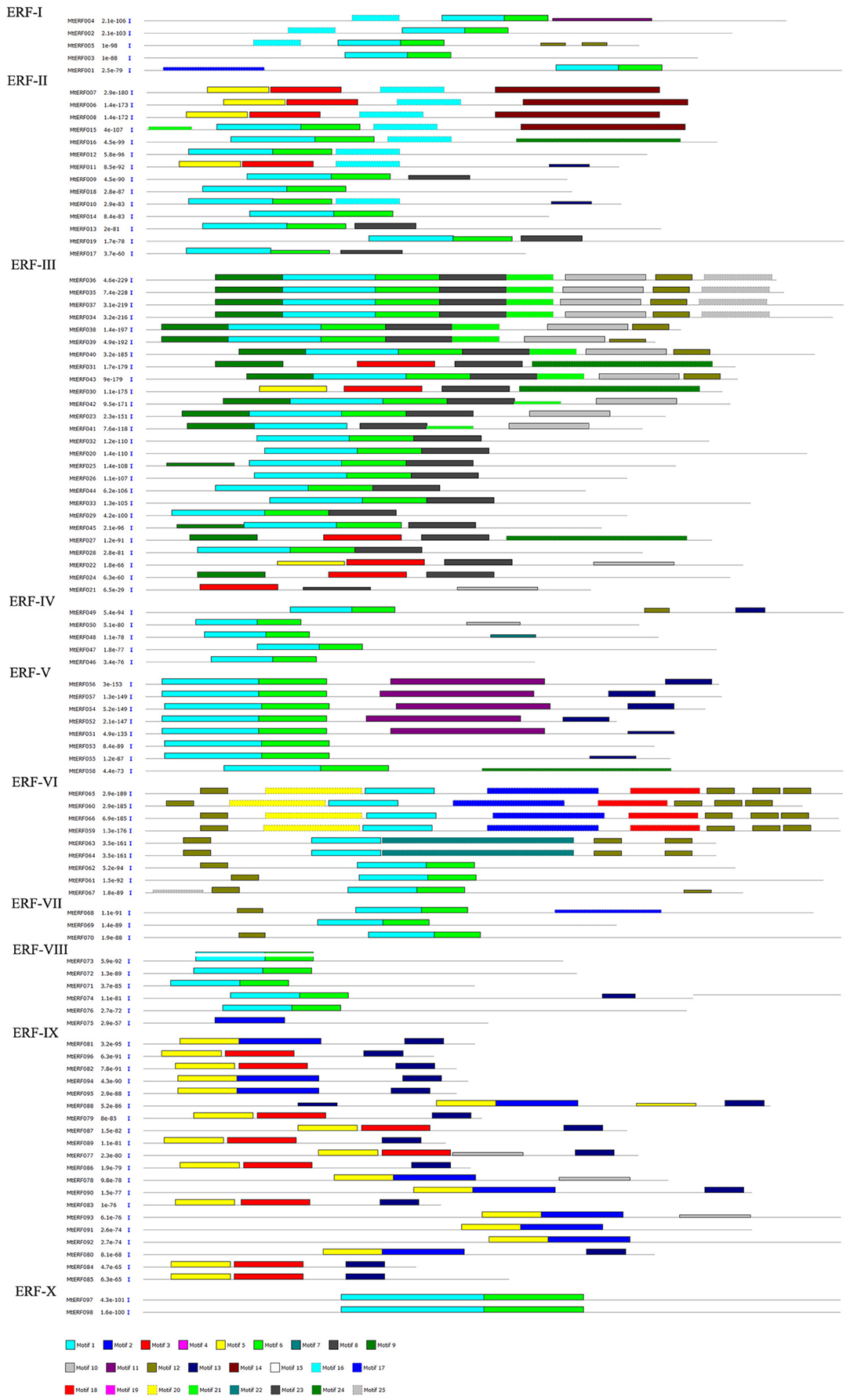
The conserved motifs in AP2/ERF family proteins in M. truncatula were investigated using MEME, revealing a total of 25 conserved motifs (designated motifs 1–25), as shown in Figures 3, 4. Motifs 1–6 were found to be similar to the AP2/ERF domain region, while the remaining motifs corresponded to regions outside of the AP2/ERF domain region, which are distributed in specific clades in the phylogenetic tree. Proteins in the same group or subgroup contain similar motifs, while the motifs are divergent among different groups or subgroups. For example, motif 23 is only present in each member of the RAV family, which indicates that this motif is specific to the RAV family. Similarly, motif 19 is shared by members of the AP2 family, motif 16 is present in groups ERF-I and II, and the ERF II group contains motif 14 while the ERF I group does not. Motifs 8–10 are specific to the ERF III group, and motif 8 is shared by each member of the ERF III group. Finally, while motif 9 and 10 are shared by most members (16/26, 62%), motif 11 and 13 are conserved in the ERF V group and motif 12 is shared by groups ERF VI and VII. These results indicate that most motifs are distributed among specific groups, which is correlated with their functional divergence (see Figures S1–S4).
Chromosomal Locations and Duplication of the AP2/ERF TFs in Medicago truncatula
The 123 AP2/ERF TF genes are distributed throughout the eight chromosomes of M. truncatula; their physical locations on chromosomes are shown in Figure 5. Each M. truncatula chromosome contains some AP2/ERF genes in numbers ranging from 10 to 27. Chromosomes 1 and 5 have the highest number of AP2/ERF genes (27 and 21 genes, respectively), while chromosomes 7 and 8 contain the fewest genes (11 and 10, respectively). The AP2/ERF genes are not randomly distributed on each chromosome, as there are some gene clusters “hot regions” on the chromosomes. For example, chromosome 6 contains 10 AP2/ERF genes (MtERF34–43) in a short chromosome region (~390 kb), and chromosomes 1, 2, and 5 contain similar gene clusters, as shown in Figure 5. In addition, using gene duplication analysis, we identified 38 pairs of gene duplications, which arose from tandem duplications and segmental duplications. Tandem duplications produced MtERF gene clusters or hot regions, such as the MtERF34–43 cluster on chromosome 6 and the MtERF10–12 cluster on chromosome 2. Segmental duplication produced many homologous AP2/ERF genes on different chromosomes, which expanded the numbers of MtERF genes from different groups. For example, MtERF52, 54, 56, and 57 from group ERF-V are distributed on different chromosomes (MtERF52 on chromosome 2, MtERF54 on chromosome 3, MtERF56, and 57 on chromosome 4), which are products of genome segmental duplication.
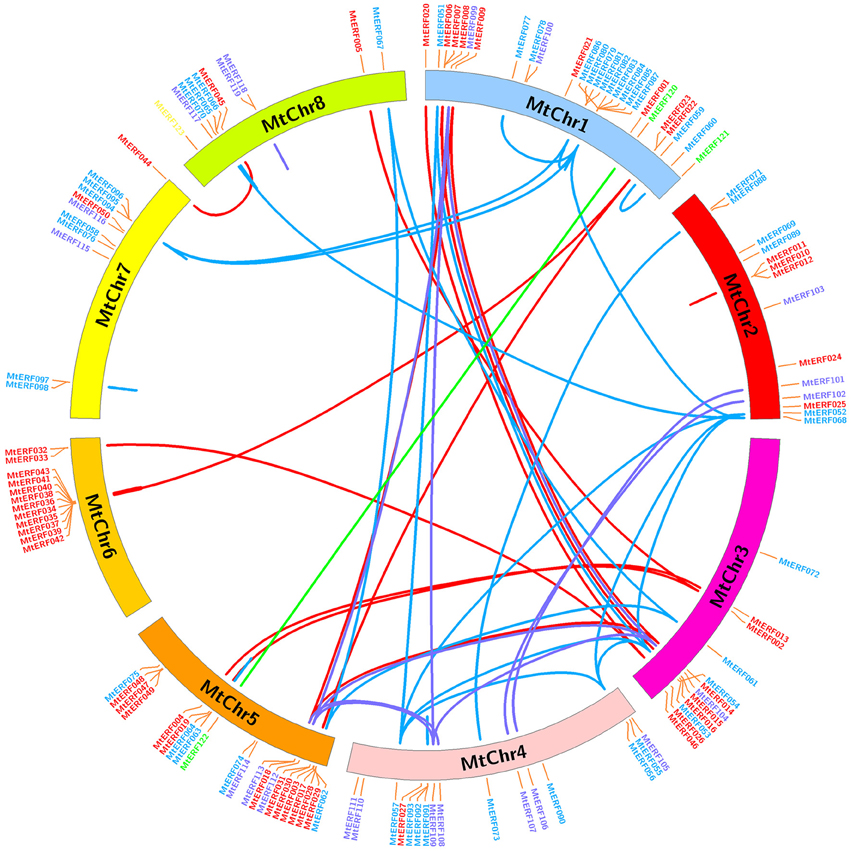
Figure 5. Chromosomal distribution and expansion analysis of MtERF genes in Medicago truncatula. Red lines show duplications between members of the DREB subfamily, blue lines show duplications between members of the ERF subfamily, purple lines show duplications between members of the AP2 family and green lines show duplications between members of the RAV subfamily.
Expression Profiles of MtERF Genes in M. truncatula Tissues
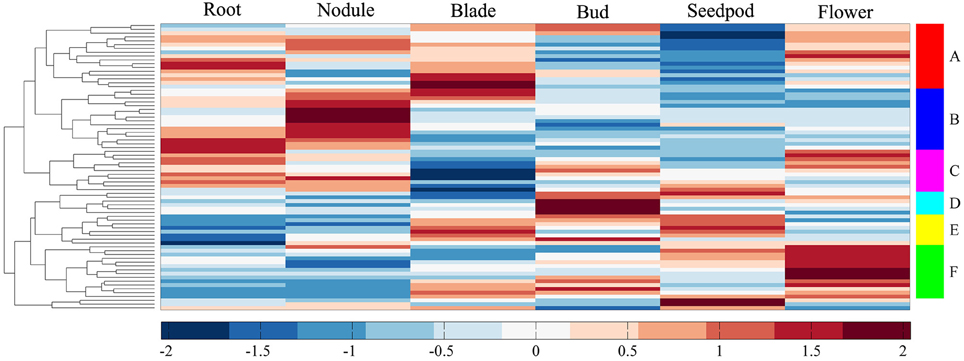
We investigated the expression profiles of MtERF genes in various tissues using high-throughput sequencing data from NCBI, including root, nodule, blade, bud, seedpod, and flower tissues, revealing that 75 MtERF genes were expressed in at least one of six tissues. Of these, the expression of 46 genes was detected in root tissue, 46 genes in nodule tissue, 42 genes in blade tissue, 47 genes in bud tissue, 36 genes in seedpod tissue, and 52 genes in flower tissue (see Figure 6A). To further elucidate the transcription patterns of MtERF genes, their expression patterns were clustered across six tissues, as shown in Figure 7. Among these MtERF genes, group A (17 MtERF genes) were highly expressed across all six tissues, while the others exhibited tissue-specific profiles. For example, group B (16 MtERF genes) were highly expressed in root and nodule tissues and not in other tissues. Similarly, group D (six MtERF genes) were specifically expressed in buds, group E (eight MtERF genes) in seedpod and blades and group E (14 MtERF genes) in flowers.
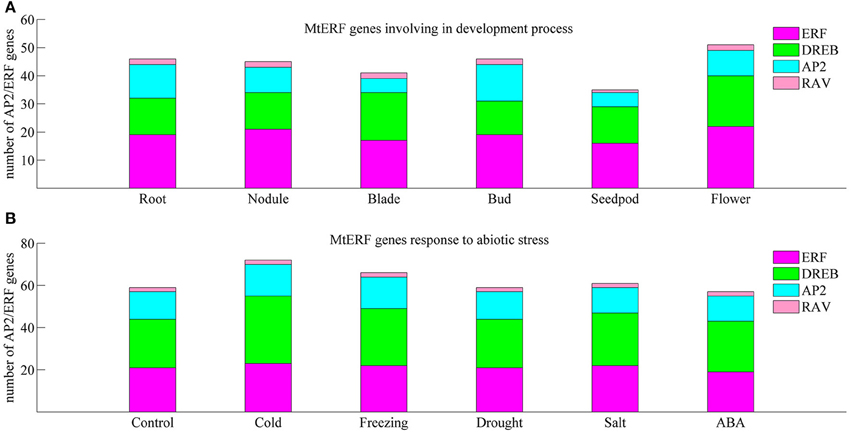
Figure 6. Number of differentially expressed MtERF genes involved in tissue development (A) and stress responses (B).
Recently, researchers have demonstrated that microRNA miR172 plays an important role by targeting AP2 TFs in the root nodule symbiosis of legumes (Reynoso et al., 2013; Wang et al., 2014; Nova-Franco et al., 2015). To determine potential regulatory mechanisms of miRNAs and AP2 TFs in the nodulation process of M. truncatula, we submitted M. truncatula miRNA genes (download from miRBase, Van Peer et al., 2014) and MtERF genes to the psRNATarget website (Dai and Zhao, 2011) for the identification of potential target sites. There were four MtERF genes (MtERF101, 106, 108, and 113) of the miR172 family with unambiguously identified cleavage sites (see Figure 8A, and Table S2). Based MtERF genes expression from high-through data, we found that four target MtERF genes had low expression in nodules. Indeed, two of them (MtERF108 and MtERF113) were not expressed at all (Figure 8B).
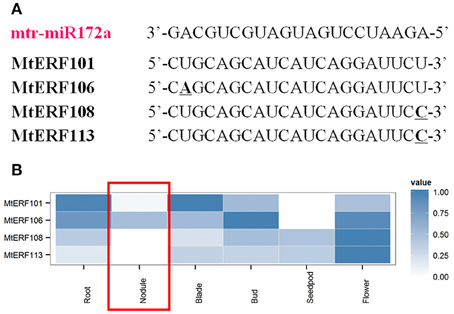
Figure 8. Putative miR172 target sites in mRNAs of AP2 family genes. (A) miRNA172 cleavage sites in MtERF genes; (B) expressional profiles of MtERF genes in tissue development.
Expression Responses of MtERF Genes to Abiotic Stress
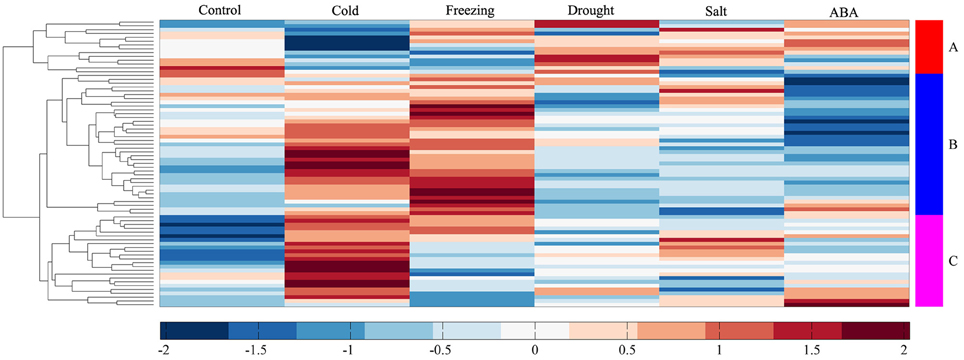
To investigate the molecular functions of MtERF genes in response to abiotic stress, we performed RNA-seq to detect the expression levels of AP2/ERF TF genes under different stresses, including cold, freezing, drought, salt and ABA. In total, more than 135 M clean reads from control (23,357,742), cold (20,437,484), freezing (23,469,784), drought (22,672,350), salt (21,993,278), and ABA (23,441,046) treated libraries were obtained, respectively; all clean data were submitted to the NCBI SRA database (Accession numbers: SRX1056987–92). After mapping these reads to the M. truncatula genome, we determined that the expression of 75 MtERF genes was detected in at least one library. As shown in Figure 6B, 60 MtERF genes were expressed in the control library, 73 MtERFs under cold stress, 67 MtERFs under freezing stress, 60 MtERFs under drought stress, 62 MtERFs under salt stress, and 58 MtERFs under ABA treatment. Based on expressional profiles of MtERF genes responses to abiotic stresses, they were clustered into three groups, as shown in Figure 9. Among these MtERF genes, group A (14 MtERF genes) were highly expressed by most abiotic stresses, except cold stress. While group B (37 MtERF genes) were highly expressed in response to cold and/or freezing stress, but not in response to other stresses. Group C (24 MtERF genes) were specifically expressed in response to cold stress, fewer in freezing stress and others. Compared to the control library, we identified 48 MtERF genes that were differentially expressed under at least one stress condition, as shown in Figure S5. A total of 35 MtERF genes were differentially expressed under cold stress, while 29 MtERF genes were differentially expressed under freezing stress. A total of 12, 12, and 23 MtERF genes were differentially expressed under drought stress, salt stress and ABA stress, respectively, which was consistent with the MtERF genes clustering results. Notably, four MtERF genes (MtERF022, MtERF023, MtERF043, and MtERF073) were up-regulated under all stress conditions, suggesting that they play important roles in the response of M. truncatula to abiotic stress.
To further confirm the RNA-seq results of these MtERF genes to abiotic stresses, qRT-PCR was performed for nine MtERF genes from the DREB subfamily under abiotic stresses. The expression patterns of most of the MtERF genes in the qRT-PCR analysis were consistent with RNA-Seq analysis, but the magnitude of the fold changes varied between RNA-seq and qRT-PCR experiments (Figure 10). The means of the correlation coefficients of the qRT-PCR validations and the RNA-seq results for the MtERF genes were as high as 0.83, implying that our RNA-seq results were highly reliable.
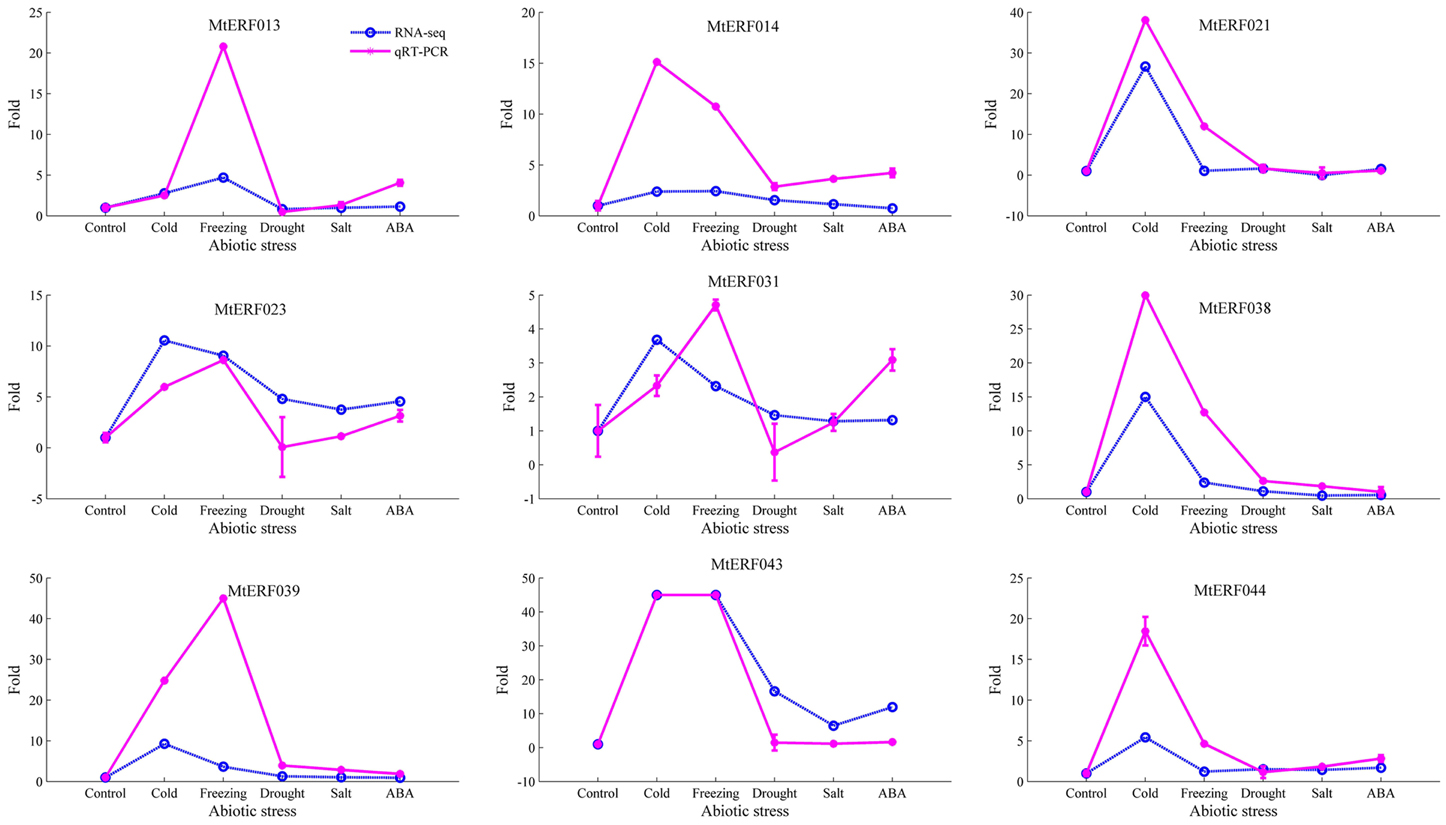
Figure 10. qRT-PCR validation of MtERF genes in the response to abiotic stress. Fold changes more than 45 were set as 45 for good plot (including two genes, MtERF039, and 043).
Discussion
In this study, we performed a comprehensive search for AP2/ERF TF genes throughout the Medicago genome; 123 MtERF genes were identified and characterized. Previously, Zhang et al. (2013b) identified 37 AP2/ERF genes using EST sequences, whereas we characterized 123 AP2/ERF members from the ERF, AP2, and RAV families. Compared to other plants, the number of AP2/ERF genes in Medicago is slightly lower than that in Arabidopsis (148), soybean (147), cucumber (131), and grapevine (149) and much lower than that in rice (180) and poplar (202). This is most likely due to the lower number of DREB and ERF subfamily members, as there are 98 members in Medicago and 122 in Arabidopsis, 120 in soybean, 122 in grapevine, 145 in rice, and 169 in poplar. By contrast, the number of AP2 and RAV family members is highly conserved among higher plants. For example, 21 members of the AP2 family were identified in this study, while 18, 26, 20, 29, and 26 AP2 genes were identified in Arabidopsis, soybean, grapevine, rice, and poplar, respectively.
In general, TFs always harbor some important conserved domains and motifs for their regulatory function. In MtERFs, 25 motifs were identified based on the MEME results; six motifs (motifs1–6) related to the AP2 domain. Among these motifs, motif1 harbored the largest region of the AP2 domain, including the whole of the β-sheet region and part of the α-helix region (G[VI]R, Gx4E, WLG, and AYD elements, as described by Nakano et al., 2006). The other five motifs were short AP2 domains. Motif4 and motif5 contained G[VI]R and Gx4E elements, while motif2, motif3, and motif6 contained WLD and AYD elements. All MtERFs contained at least one of the six motifs, indicating that the AP2 domain is highly conserved in MtERF genes. Besides the AP2 domain-related motifs, the other 19 motifs outside the AP2 domain were present in group-specific distributions. For example, motif17 was specifically present in the ERF VI group (also called the ERF B5 group), and contained (L/F) DLN (L/F) xP residues. It was identified as an ERF-associated amphiphilic repression (EAR) motif, essential for repression function (Ohta et al., 2001). Meanwhile, a SP[TV]SVL motif was characterized. It had the potential to be phosphorylated by mitogen activated protein kinase (MAPK) (Nakano et al., 2006). Motif17 may have specific repression functions by virtue of the phosphorylation of the ERF V group. Motif13 contained a unique “EDLL” motif, and was present in the ERF V and IX groups. The “EDLL” motif was previously characterized as a transcriptional activation domain in AP2/ERF TF (Tiwari et al., 2012), implying its participation in the activating function of the ERF V and IX groups (previously called the B6 and B3 groups, respectively). Motif8 was highly conserved in all members of the ERF III group and in four members of the ERF II group (in total, including 30 out of the 45 members of the DREB subfamily). It contained two conserved residue features: LPRP and D[IV]QAA. They have been identified as being essential regulation signatures of the response to various stresses in plants (Albrecht et al., 2001; Qu and Zhu, 2006). Their expression response to abiotic stresses has been shown by RNA-seq analysis. However, except for the three motifs 8, 13 and 17, and the six AP2 domain related motifs (motifs 1–6), the function of the other 16 conserved motifs (65% of the total) identified in present study is uncertain. Their roles need to be further characterized.
AP2/ERF genes in various plant species are differentially expressed in different tissues, indicating that they play important roles in plant tissue development. Genes in the ERF subfamily play many diverse roles, such as functioning in the response to hormonal stimuli and regulating developmental processes in various angiosperms. Meanwhile, members of the AP2 family participate in the regulation of developmental processes, such as flower development and meristem determinacy (Krizek, 2009). In the current study, 75 MtERF genes were found to be expressed in at least one tissue based on high-throughput sequencing data analysis, including DREB (26/50, 52%), ERF (31/48, 64.58%), AP2 (15/21, 71.43%) and RAV (2/3, 66.67%), genes and one Soloist gene (1/1, 100%). Except for two families with fewer members (RAV and Soloist), most members of the ERF and AP2 families participate in the regulation of tissue determinacy. When we compared the expression of MtERF genes in these two families, we found that ERF genes had higher expression levels than AP2 genes, which may be the result of the higher intron content of the AP2 family. Each member of the AP2 family has five to nine introns, while most ERF genes lack introns (74/98) and others contain only one or two introns. Due to the small number of introns, genes in the ERF family respond more quickly and are expressed at higher levels than AP2 family genes. Of the six tissues examined in Medicago, MtERF genes were most highly expressed in flower tissue, which is consistent with previous reports (Krizek, 2009; Matías-Hernández et al., 2014).
Like other leguminous plants, M. truncatula has established a symbiotic relationship with nitrogen fixing rhizobial bacteria, resulting in the formation of specialized lateral organs, called nodules. Nodulation is a complex developmental process involving many molecular signals constituting a genetic regulatory network. TFs, including GRAS, AP2/ERF, NF-Y, and others, have been shown to have important roles in controlling the expression of early nodulation genes (ENODs) and in regulating the later steps of the rhizobial symbiotic interaction (Combier et al., 2006; Andriankaja et al., 2007; Middleton et al., 2007; Vernié et al., 2008; Hirsch et al., 2009; Zanetti et al., 2010; Soyano et al., 2013; Laloum et al., 2014; Baudin et al., 2015). The ERF V group, previously called the B6 group in other studies, contains eight members, four of which (MtERF 52, 53, 55, and 58) were highly expressed in nodule tissue (Figure S6). MtERF 55 and 58 were only expressed in nodule tissue. This strongly suggests that these genes play unique and important roles in regulating root development and the symbiotic associations with rhizobia. We found that MtERF58 was homologous to ERN and ERN1 (Table 2). Previous researchers have demonstrated that the ERN gene (similar to ERN1), which is homologous to RAP2.11, interacts with GRAS factors in the regulation of NF-elicited gene transcription rhizobial infection (Andriankaja et al., 2007; Middleton et al., 2007; Vernié et al., 2008). MtERF55 is homologous to PtaERF003, which has been identified as promoting root development in Populus (Trupiano et al., 2013). These results confirm the potential of the ERF V group for the regulation of nodulation. Their functions in the development of other tissues need further investigation.
Nodulation is a complex process in M. truncatula, involving the ERF V group and other AP2/ERF genes, with different regulation patterns. As previously mentioned, many researchers have found that miRNA miR172 is also involved in the nodule development of legumes (Wang et al., 2014; Nova-Franco et al., 2015). It regulates the nodulation process by targeting AP2/ERF TFs, and repressing their expressions. In so doing, it is a negative regulator of root nodulation. In the present study, four AP2/ERF genes (Figure 8A), belonging to the AP2 family, were identified as potential targets of miR172 with unambiguous target sites. Based on expression analysis in silico, we found that these MtERF genes had low levels of expression in nodules, implying that they were potentially down-regulated by miR172 in the nodulation process. These results show the divergent functions of MtERF genes in nodulation. MtERF58, with high levels of expression, is a positive regulator of nodulation in M. truncatula; as confirmed in many studies. However, members of the AP2 family (MtERF 101, 106, 108, 113) were cleaved by miR172, and have low levels of expression in nodules. They may be important repressors of nodulation. This is well-characterized in soybeans and in common beans (Wang et al., 2014; Nova-Franco et al., 2015). However, its function in M. truncatula needs further confirmation.
When adapting to various environmental conditions, plants employ many TF families involved in regulating a wide range of defense responses to environmental clues. AP2/ERF superfamily TFs with ERF domains can bind to GCC-box elements or DRE motifs, thereby regulating gene expression in response to biotic or abiotic stress (Fujimoto et al., 2000; Cao et al., 2001). As previously shown, MtCBF1-3 isolated from M. truncatula, has one AP2 domain, and have key functions in cold acclimatization in M. truncatula and the related species, M. falcate (Pennycooke et al., 2008; Table 2). Similarly, Li et al. (2011) identified the role of MtCBF4 in responding to abiotic stresses, including cold, drought, and salt. Studies in which it was over-expressed conferred improved drought and salt tolerance. Notably, Tayeh et al. (2013) identified a major role for QTL (Mt-FTQTL6) in tolerance against freezing, demonstrated 40% of the phenotypic variation by QTL mapping, and characterized 12 potential MtCBF genes clustering in the Mt-FTQTL6 region. However, the expression profiles of these MtCBF genes in response to freezing stress are still poorly understood, and their function in freezing tolerance in M. truncatula is unknown. In this study, we analyzed the expression of MtERF genes under different stress conditions via transcriptome sequencing. We identified 75 MtERF genes that were expressed under various stress treatments, among which 63 were also expressed during tissue development, while 12 were specifically expressed under stress treatment, as shown in Figure S7. Compared to the control samples, 48 MtERF genes were found to be differentially expressed in response to abiotic stress (see Figure S5), and most MtERF genes were induced by abiotic stress. These MtERF genes include DREB (23/48, 47.92%), ERF (15/48, 31.25%), AP2 (7/48, 14.58%), RAV (2/48, 4.17%), and one Soloist (1/48, 2.08%) gene, the DREB and ERF subfamilies are the largest groups that were responsive to abiotic stress. We found that the DREB and ERF subfamilies were more sensitive to abiotic stress, while ERF and AP2 members were active in tissue development. Among these MtERF genes, 43 MtERF genes were responsive to cold or freezing stress (see Figure S8), while 30 MtERF genes were responsive to salt, drought or ABA-induced stress, as shown in Figure S9. A total of 25 MtERF genes commonly respond to abiotic stress. These include MtERF22, 23, 43, amongst others. MtERF23, which was called MtCBF4 in the report by Li et al. (2011), was previously identified as being induced by cold, drought, salt, and ABA stresses. Our RNA-seq data and qRT-PCR experiments confirmed its high expression under abiotic stresses, (see Figure 10). Of the MtERF genes, 21 were specifically induced or repressed by low temperature stresses, including the previously reported MtCBF 1-3 (Pennycooke et al., 2008). In our study, these were called MtERF 31, 40, and 42, respectively. Most of them are from the DREB subfamily (see Figure S5). Their expression profiles indicate that MtERF genes play more important roles in regulating the response to abiotic stresses M. truncatula, especially DREB subfamily critical function to low temperature stress. Interestingly, 10 MtERF genes (MtERF34–43) are clustered on chromosome 6, and they were tandemly duplicated within an ~393 kb region, which was previously identified by Tayeh et al. (2013). In the present study, we identified and characterized 10 MtERF genes, belonging to the DREB subfamily. Two previously identified potential MtCBF genes were excluded because the aligning region with Arabidopsis ERF genes was less than 80%. Moreover, our RNA-seq profiles demonstrated that six of these MtERF genes (also called MtCBF genes) were highly up-regulated under both cold and freezing stress (see Figure S5), and qRT-PCR had confirmed their high expression in cold and freezing stresses (MtERF38, 39, and 43, see Figure 10). The expression profiles suggested important and complementary potential roles of the MtERF gene cluster on chromosome 6 under abiotic stresses, especially its major contribution to freezing tolerance in M. truncatula.
Conclusions
In summary, we identified 123 MtERF genes from the M. truncatula genome sequence. We investigated the classification, evolution, and tissue-specific expression of these MtERF genes, revealing that MtERF genes broadly participate in the regulation of plant tissue development. Meanwhile, we identified 48 candidate MtERF genes that may be involved in abiotic stress responses, their expression profiles were confirmed by qRT-PCR experiment. In particular, a tandem array of MtERF genes on chromosome 6 was identified and found to function in the response to cold and freezing stress, as revealed by RNA-seq analysis. The results of this study will be useful for identifying and characterizing these genes. Further functional analyses of these genes will be performed in the future to enable them to be used for transgenic applications.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by grants from the Natural and Science Foundation of China (Nos. 31302019 and 31470571), the China Postdoctoral Science Foundation (2015M571430), the MOST 863 project (2013AA102607-5), and the Heilongjiang Province Postdoctoral Science Foundation (No. LBH-Z14126).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2015.01247
Figure S1. Conserved motifs identified from members of the DREB subfamily in Medicago truncatula.
Figure S2. Conserved motifs identified from members of the ERF subfamily in Medicago truncatula.
Figure S3. Comparison of amino acid sequences of the AP2/ERF domains in the DREB subfamily.
Figure S4. Comparison of amino acid sequences of the AP2/ERF domains in the ERF subfamily.
Figure S5. Differential expression analysis of MtERF genes involved in the response to abiotic stress.
Figure S6. Expression profile cluster analysis members of MtERF V group in tissue development.
Figure S7. Venn diagram of shared expression MtERF genes between tissues development and stresses response.
Figure S8. Venn diagram of shared expression MtERF genes between cold stress and freezing stress.
Figure S9. Venn diagram of shared expression MtERF genes among salt, drought and ABA stresses.
Table S1. List of qRT-PCR validation primers used in the present study.
Table S2. Summary of miR172 targeting AP2/ERF TF genes in Medicago truncatula.
References
Albrecht, V., Ritz, O., Linder, S., Harter, K., and Kudla, J. (2001). The NAF domain defines a novel protein-protein interaction module conserved in Ca2+-regulated kinases. EMBO J. 20, 1051–1063. doi: 10.1093/emboj/20.5.1051
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990). Basic local alignment search tool. J. Mol. Biol. 215, 403–410. doi: 10.1016/S0022-2836(05)80360-2
Anderson, J. P., Lichtenzveig, J., Gleason, C., Oliver, R. P., and Singh, K. B. (2010). The B-3 ethylene response factor MtERF1-1 mediates resistance to a subset of root pathogens in Medicago truncatula without adversely affecting symbiosis with rhizobia. Plant Physiol. 154, 861–873. doi: 10.1104/pp.110.163949
Andriankaja, A., Boisson-Dernier, A., Frances, L., Sauviac, L., Jauneau, A., Barker, D. G., et al. (2007). AP2-ERF transcription factors mediate Nod factor dependent Mt ENOD11 activation in root hairs via a novel cis-regulatory motif. Plant Cell 19, 2866–2885. doi: 10.1105/tpc.107.052944
Bailey, T. L., Williams, N., Misleh, C., and Li, W. W. (2006). MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 34, W369–W373. doi: 10.1093/nar/gkl198
Baudin, M., Laloum, T., Lepage, A., Rípodas, C., Ariel, F., Frances, L., et al. (2015). A phylogenetically conserved group of nuclear factor-Y transcription factors interact to control nodulation in Legumes. Plant Physiol. 169, 2761–2773. doi: 10.1104/pp.15.01144
Cao, Z. F., Li, J., Chen, F., Li, Y. Q., Zhou, H. M., and Liu, Q. (2001). Effect of two conserved amino acid residues on DREB1A function. Biochemistry (Mosc) 66, 623–627. doi: 10.1023/A:1010251129429
Chinnusamy, V., Schumaker, K., and Zhu, J. K. (2004). Molecular genetic perspectives on cross-talk and specificity in abiotic stress signalling in plants. J. Exp. Bot. 55, 225–236. doi: 10.1093/jxb/erh005
Chung, M. Y., Vrebalov, J., Alba, R., Lee, J., McQuinn, R., Chung, J. D., et al. (2010). A tomato (Solanum lycopersicum) APETALA2/ERF gene, SlAP2a, is a negative regulator of fruit ripening. Plant J. 64, 936–947. doi: 10.1111/j.1365-313X.2010.04384.x
Combier, J.-P., Frugier, F., de Billy, F., Boualem, A., El-Yahyaoui, F., Moreau, S., et al. (2006). MtHAP2-1 is a key transcriptional regulator of symbiotic nodule development regulated by microRNA169 in Medicago truncatula. Genes Dev. 20, 3084–3088. doi: 10.1101/gad.402806
Dai, X., and Zhao, P. X. (2011). psRNATarget: a plant small RNA target analysis server. Nucleic Acids Res. 39, W155–W159. doi: 10.1093/nar/gkr319
Finn, R. D., Bateman, A., Clements, J., Coggill, P., Eberhardt, R. Y., Eddy, S. R., et al. (2014). Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230. doi: 10.1093/nar/gkt1223
Fujimoto, S. Y., Ohta, M., Usui, A., Shinshi, H., and Ohme-Takagi, M. (2000). Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 12, 393–404. doi: 10.1105/tpc.12.3.393
Gujaria-Verma, N., Vail, S. L., Carrasquilla-Garcia, N., Penmetsa, R. V., Cook, D. R., Farmer, A. D., et al. (2014). Genetic mapping of legume orthologs reveals high conservation of synteny between lentil species and the sequenced genomes of Medicago and chickpea. Front. Plant Sci. 5:676. doi: 10.3389/fpls.2014.00676
Guo, A., He, K., Liu, D., Bai, S., Gu, X., Wei, L., et al. (2005). DATF: a database of Arabidopsis transcription factors. Bioinformatics 21, 2568–2569. doi: 10.1093/bioinformatics/bti334
Hirsch, S., Kim, J., Muñoz, A., Heckmann, A. B., Downie, J. A., and Oldroyd, G. E. (2009). GRAS proteins form a DNA binding complex to induce gene expression during nodulation signaling in Medicago truncatula. Plant Cell 21, 545–557. doi: 10.1105/tpc.108.064501
Katagiri, F. (2004). A global view of defense gene expression regulation–a highly interconnected signaling network. Curr. Opin. Plant Biol. 7, 506–511. doi: 10.1016/j.pbi.2004.07.013
Kitomi, Y., Ito, H., Hobo, T., Aya, K., Kitano, H., and Inukai, Y. (2011). The auxin responsive AP2/ERF transcription factor CROWN ROOTLESS5 is involved in crown root initiation in rice through the induction of OsRR1, a type-A response regulator of cytokinin signaling. Plant J. 67, 472–484. doi: 10.1111/j.1365-313X.2011.04610.x
Krizek, B. (2009). AINTEGUMENTA and AINTEGUMENTA-LIKE6 act redundantly to regulate Arabidopsis floral growth and patterning. Plant Physiol. 150, 1916–1929. doi: 10.1104/pp.109.141119
Krzywinski, M., Schein, J., Birol, I., Connors, J., Gascoyne, R., Horsman, D., et al. (2009). Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645. doi: 10.1101/gr.092759.109
Kunst, L., Klenz, J. E., Martinez-Zapater, J., and Haughn, G. W. (1989). AP2 Gene determines the identity of perianth organs in flowers of Arabidopsis thaliana. Plant Cell 1, 1195–1208. doi: 10.1105/tpc.1.12.1195
Laloum, T., Baudin, M., Frances, L., Lepage, A., Billault-Penneteau, B., Cerri, M. R., et al. (2014). Two CCAAT-box-binding transcription factors redundantly regulate early steps of the legume-rhizobia endosymbiosis. Plant J. 79, 757–768. doi: 10.1111/tpj.12587
Le Hir, R., and Bellini, C. (2013). The plant-specific dof transcription factors family: new players involved in vascular system development and functioning in Arabidopsis. Front. Plant Sci. 4:164. doi: 10.3389/fpls.2013.00164
Lee, T. H., Tang, H., Wang, X., and Paterson, A. H. (2013). PGDD: a database of gene and genome duplication in plants. Nucleic Acids Res. 41, D1152–D1158. doi: 10.1093/nar/gks1104
Li, D., Zhang, Y., Hu, X., Shen, X., Ma, L., Su, Z., et al. (2011). Transcriptional profiling of Medicago truncatula under salt stress identified a novel CBF transcription factor MtCBF4 that plays an important role in abiotic stress responses. BMC Plant Biol. 11:109. doi: 10.1186/1471-2229-11-109
Li, M. Y., Xu, Z. S., Huang, Y., Tian, C., Wang, F., and Xiong, A. S. (2015) Genome-wide analysis of AP2/ERF transcription factors in carrot (Daucus carota L.) reveals evolution and expression profiles under abiotic stress. Mol. Genet. Genomics 290, 2049–2061. doi: 10.1007/s00438-015-1061-3
Licausi, F., Giorgi, F. M., Zenoni, S., Osti, F., Pezzotti, M., and Perata, P. (2010). Genomic and transcriptomic analysis of the AP2/ERF superfamily in Vitis vinifera. BMC Genomics 11:719. doi: 10.1186/1471-2164-11-719
Mantiri, F. R., Kurdyukov, S., Chen, S. K., and Rose, R. J. (2008a). The transcription factor MtSERF1 may function as a nexus between stress and development in somatic embryogenesis in Medicago truncatula. Plant Signal Behav. 3, 498–500. doi: 10.4161/psb.3.7.6049
Mantiri, F. R., Kurdyukov, S., Lohar, D. P., Sharopova, N., Saeed, N. A., Wang, X. D., et al. (2008b). The transcription factor MtSERF1 of the ERF subfamily identified by transcriptional profiling is required for somatic embryogenesis induced by auxin plus cytokinin in Medicago truncatula. Plant Physiol. 146, 1622–1636. doi: 10.1104/pp.107.110379
Matías-Hernández, L., Aguilar-Jaramillo, A. E., Marín-González, E., Suárez-López, P., and Pelaz, S. (2014). RAV genes: regulation of floral induction and beyond. Ann. Bot. 114, 1459–1470. doi: 10.1093/aob/mcu069
Middleton, P. H., Jakab, J., Penmetsa, R. V., Starker, C. G., Doll, J., Kalo, P., et al. (2007). An ERF transcription factor in Medicago truncatula that is essential for Nod factor signal transduction. Plant Cell 19, 1221–1234. doi: 10.1105/tpc.106.048264
Mizoi, J., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2012). AP2/ERF family transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta 1819, 86–96. doi: 10.1016/j.bbagrm.2011.08.004
Nakano, T., Suzuki, K., Fujimura, T., and Shinshi, H. (2006). Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 140, 411–432. doi: 10.1104/pp.105.073783
Nova-Franco, B., Íñiguez, L. P., Valdés-López, O., Alvarado-Affantranger, X., Leija, A., Fuentes, S. I., et al. (2015). The micro-RNA72c-APETALA2-1 node as a key regulator of the common bean-Rhizobium etli nitrogen fixation symbiosis. Plant Physiol. 168, 273–291. doi: 10.1104/pp.114.255547
Ohta, M., Matsui, K., Hiratsu, K., Shinshi, H., and Ohme-Takagi, M. (2001). Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 13, 1959–1968. doi: 10.1105/tpc.010127
Onãte-Sánchez, L., and Singh, K. B. (2002). Identification of Arabidopsis ethylene-responsive element binding factors with distinct induction kinetics after pathogen infection. Plant Physiol. 128, 1313–1322. doi: 10.1104/pp.010862
Pennycooke, J. C., Cheng, H., and Stockinger, E. J. (2008). Comparative genomic sequence and expression analyses of Medicago truncatula and alfalfa subspecies falcata COLD-ACCLIMATION-SPECIFIC genes. Plant Physiol. 146, 1242–1254. doi: 10.1104/pp.107.108779
Qu, L. J., and Zhu, Y. X. (2006). Transcription factor families in Arabidopsis: major progress and outstanding issues for future research. Curr. Opin. Plant Biol. 9, 544–549. doi: 10.1016/j.pbi.2006.07.005
Reynoso, M. A., Blanco, F. A., Bailey-Serres, J., Crespi, M., and Zanetti, M. E. (2013). Selective recruitment of mRNAs and miRNAs to polyribosomes in response to rhizobia infection in Medicago truncatula. Plant J. 73, 289–301. doi: 10.1111/tpj.12033
Sakuma, Y., Liu, Q., Dubouzet, J. G., Abe, H., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2002). DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem. Biophys. Res. Commun. 290, 998–1009. doi: 10.1006/bbrc.2001.6299
Sharma, M. K., Kumar, R., Solanke, A. U., Sharma, R., Tyagi, A. K., and Sharma, A. K. (2010). Identification, phylogeny, and transcript profiling of ERF family genes during development and abiotic stress treatments in tomato. Mol. Genet. Genomics 284, 455–475. doi: 10.1007/s00438-010-0580-1
Shen, C., Yue, R., Sun, T., Zhang, L., Xu, L., Tie, S., et al. (2015). Genome-wide identification and expression analysis of auxin response factor gene family in Medicago truncatula. Front. Plant Sci. 6:73. doi: 10.3389/fpls.2015.00073
Singh, K., Foley, R. C., and Oñate-Sánchez, L. (2002). Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 5, 430–436. doi: 10.1016/S1369-5266(02)00289-3
Soyano, T., Kouchi, H., Hirota, A., and Hayashi, M. (2013). NODULE INCEPTION directly targets NF-Y subunit genes to regulate essential processes of root nodule development in Lotus japonicus. PLoS Genet. 9:e1003352. doi: 10.1371/journal.pgen.1003352
Tamura, K., Dudley, J., Nei, M., and Kumar, S. (2007). MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599. doi: 10.1093/molbev/msm092
Tayeh, N., Bahrman, N., Sellier, H., Bluteau, A., Blassiau, C., Fourment, J., et al. (2013). A tandem array of CBF/DREB1 genes is located in a major freezing tolerance QTL region on Medicago truncatula chromosome 6. BMC Genomics 14:814. doi: 10.1186/1471-2164-14-814
Thompson, J. D., Gibson, T. J., and Higgins, D. G. (2002). Multiple sequence alignment using ClustalW and ClustalX. Curr. Protoc. Bioinformatics Chapter 2, Unit 2.3. doi: 10.1002/0471250953.bi0203s00
Tiwari, S. B., Belachew, A., Ma, S. F., Young, M., Ade, J., Shen, Y., et al. (2012). The EDLL motif: a potent plant transcriptional activation domain from AP2/ERF transcription factors. Plant J. 70, 855–865. doi: 10.1111/j.1365-313X.2012.04935.x
Trapnell, C., Pachter, L., and Salzberg, S. L. (2009). TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111. doi: 10.1093/bioinformatics/btp120
Trapnell, C., Roberts, A., Goff, L., Pertea, G., Kim, D., Kelley, D. R., et al. (2012). Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578. doi: 10.1038/nprot.2012.016
Trupiano, D., Yordanov, Y., Regan, S., Meilan, R., Tschaplinski, T., Scippa, G. S., et al. (2013). Identification, characterization of an AP2/ERF transcription factor that promotes adventitious, lateral root formation in Populus. Planta 238, 271–282. doi: 10.1007/s00425-013-1890-4
Van Peer, G., Lefever, S., Anckaert, J., Beckers, A., Rihani, A., Van Goethem, A., et al. (2014). miRBase Tracker: keeping track of microRNA annotation changes. Database (Oxford) 2014:bau080. doi: 10.1093/database/bau080
Vernié, T., Moreau, S., de Billy, F., Plet, J., Combier, J. P., Rogers, C., et al. (2008). EFD Is an ERF transcription factor involved in the control of nodule number and differentiation in Medicago truncatula. Plant Cell 20, 2696–2713. doi: 10.1105/tpc.108.059857
Wang, Y., Wang, L., Zou, Y., Chen, L., Cai, Z., Zhang, S., et al. (2014). Soybean miR172c targets the repressive AP2 transcription factor NNC1 to activate ENOD40 expression and regulate nodule initiation. Plant Cell 26, 4782–4801. doi: 10.1105/tpc.114.131607
Xu, Z. S., Chen, M., Li, L. C., and Ma, Y. Z. (2011). Functions and application of the AP2/ERF transcription factor family in crop improvement. J. Integr. Plant Biol. 53, 570–585. doi: 10.1111/j.1744-7909.2011.01062.x
Yamaguchi-Shinozaki, K., and Shinozaki, K. (2006). Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu. Rev. Plant Biol. 57, 781–803. doi: 10.1146/annurev.arplant.57.032905.105444
Young, N. D., Debellé, F., Oldroyd, G. E., Geurts, R., Cannon, S. B., Udvardi, M. K., et al. (2011). The Medicago genome provides insight into the evolution of rhizobial symbioses. Nature 480, 520–524. doi: 10.1038/nature10625
Zanetti, M. E., Blanco, F. A., Beker, M. P., Battaglia, M., and Aguilar, O. M. (2010). A C subunit of the plant nuclear factor NF-Y required for rhizobial infection and nodule development affects partner selection in the common bean–rhizobium etli symbiosis. Plant Cell 22, 4142–4157. doi: 10.1105/tpc.110.079137
Zhang, C., Zhang, H., Zhao, Y., Jiang, H., Zhu, S., Cheng, B., et al. (2013a). Genome-wide analysis of the CCCH zinc finger gene family in Medicago truncatula. Plant Cell Rep. 32, 1543–1555. doi: 10.1007/s00299-013-1466-6
Zhang, G., Chen, M., Chen, X., Xu, Z., Guan, S., Li, L. C., et al. (2008). Phylogeny, gene structures, and expression patterns of the ERF gene family in soybean (Glycine max L.). J. Exp. Bot. 59, 4095–4107. doi: 10.1093/jxb/ern248
Zhang, J. Y., Broeckling, C. D., Blancaflor, E. B., Sledge, M. K., Sumner, L. W., and Wang, Z. Y. (2005). Overexpression of WXP1, a putative Medicago truncatula AP2 domain-containing transcription factor gene, increases cuticular wax accumulation and enhances drought tolerance in transgenic alfalfa (Medicago sativa). Plant J. 42, 689–707. doi: 10.1111/j.1365-313X.2005.02405.x
Zhang, J. Y., Broeckling, C. D., Sumner, L. W., and Wang, Z. Y. (2007). Heterologous expression of two Medicago truncatula putative ERF transcription factor genes, WXP1 and WXP2, in Arabidopsis led to increased leaf wax accumulation and improved drought tolerance, but differential response in freezing tolerance. Plant Mol. Biol. 64, 265–278. doi: 10.1007/s11103-007-9150-2
Zhang, Z., Zhou, Q., Yang, Z., and Jiang, J. (2013b). Discovery AP2/ERF family genes in silico in Medicago truncatula. Afr. J. Biotechnol. 12, 3636–3642. doi: 10.5897/AJB12.2781
Zhu, J. K. (2002). Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 53, 247–273. doi: 10.1146/annurev.arplant.53.091401.143329
Keywords: Medicago truncatula, AP2/ERF transcription factors, abiotic stress, phylogenetic analysis, transcriptome analysis
Citation: Shu Y, Liu Y, Zhang J, Song L and Guo C (2016) Genome-Wide Analysis of the AP2/ERF Superfamily Genes and their Responses to Abiotic Stress in Medicago truncatula. Front. Plant Sci. 6:1247. doi: 10.3389/fpls.2015.01247
Received: 13 June 2015; Accepted: 21 December 2015;
Published: 19 January 2016.
Edited by:
Maria Eugenia Zanetti, Consejo Nacional de Investigaciones Científicas y Técnicas and Universidad Nacional de la Plata, ArgentinaReviewed by:
Asa Ben-Hur, Colorado State University, USAFederico Damian Ariel, Institute of Plant Science Paris-Saclay, Centre National de la Recherche Scientifique, France
Copyright © 2016 Shu, Liu, Zhang, Song and Guo. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yongjun Shu, c3lqdW4yMDAzQDEyNi5jb20=;
Changhong Guo, a2FrdV8yMDA4QDE2My5jb20=
 Yongjun Shu
Yongjun Shu Ying Liu
Ying Liu Lili Song
Lili Song