- 1State Key Laboratory of Tree Genetics and Breeding, Key Laboratory of Tree Breeding and Cultivation of the State Forestry Administration, Research Institute of Forestry, Chinese Academy of Forestry, Beijing, China
- 2Collaborative Innovation Center of Sustainable Forestry in Southern China, Nanjing Forestry University, Nanjing, China
Salix psammophila is a desert shrub willow that has extraordinary adaptation to abiotic stresses and plays an important role in maintaining local ecosystems. Moreover, S. psammophila is regarded as a promising biomass feedstock because of its high biomass yields and short rotation coppice cycle. However, few suitable reference genes (RGs) for quantitative real-time polymerase chain reaction (qRT-PCR) constrain the study on normalization of gene expression in S. psammophila until now. Here, we investigated the expression stabilities of 14 candidate RGs across tissue types and under four abiotic stress treatments, including heat, cold, salt, and drought treatments. After calculation of PCR efficiencies, three different software, NormFinder, geNorm, and BestKeeper were employed to analyze systematically the qRT-PCR data, and the outputs were merged by RankAggreg software. The optimal RGs selected for gene expression analysis were EF1α (Elongation factor-1 alpha) and OTU (OTU-like cysteine protease family protein) for different tissue types, UBC (Ubiquitin-conjugating enzyme E2) and LTA4H (Leukotriene A-4 hydrolase homolog) for heat treatment, HIS (Histone superfamily protein H3) and ARF2 (ADP-ribosylation factor 2) for cold treatment, OTU and ACT7 (Actin 7) for salt treatment, UBC and LTA4H for drought treatment. The expression of UBC, ARF2, and VHAC (V-type proton ATPase subunit C) varied the least across tissue types and under abiotic stresses. Furthermore, the relative genes expression profiles of one tissue-specific gene WOX1a (WUSCHEL-related homeobox 1a), and four stress-inducible genes, including Hsf-A2 (Heat shock transcription factors A2), CBF3 (C-repeat binding factor 3), HKT1 (High-Affinity K+ Transporter 1), and GST (Glutathione S-transferase), were conducted to confirm the validity of the RGs in this study. These results provided an important RGs application guideline for gene expression characterization in S. psammophila.
Introduction
Salix psammophila (Salix, Salicaceae), an important desert shrub willow, distributes in arid and semi-arid desert regions of Northern China, including Mu Us sandy land and Kubuqi desert in Inner Mongolia, Yulin in Shaanxi Province, Yanchi in Ningxia Province, and others. With extraordinary adaptation to environmental stresses (e.g., water deficit, extreme temperature and ion toxicity or deficiency) and strong ability in wind-breaking and sand-fixation, S. psammophila plays a significant role in maintaining local ecosystems (Bao and Zhang, 2012; Huang et al., 2015). Moreover, as a promising biomass feedstock for biofuels and bioenergy, S. psammophila has been applied in paper making, production of bio-based chemicals and hydrochar, such as carboxylic acids, phenolic derivatives, furan compounds, solid fuel, and adsorbents (Li et al., 2013; Yang et al., 2014). Owing to its remarkable advantages and value, the interest in the study of discovery of stress- and development-related genes on S. psammophila is stimulated for some researches.
Understanding the gene expression patterns will be of great help in the systematic elucidation the gene mechanism in complex regulatory networks. Many methods are used to assess gene expression level. Thereinto, quantitative real-time polymerase chain reaction (qRT-PCR) is the most prevalent method in detecting gene expression level with certain advantages of speed, high sensitivity, convenience, benefits, reproducibility, and accuracy (Bustin, 2002; Bustin et al., 2005; Nolan et al., 2006). However, the accuracy of experimental data obtained by qRT-PCR always was affected by many variable factors, such as RNA quality, reverse transcription efficiency (Bustin, 2002; Udvardi et al., 2008). In order to minimize the experimental errors, reference gene (RG) is used to normalize the experimental data from qRT-PCR (Radonić et al., 2004), so the accuracy of experimental results of qRT-PCR often depends on selected RGs which are appropriate or not.
An ideal RG should be stable expressed in different tissue types, development stages and under different experimental conditions. Then, housekeeping genes involved in basic cellular processes are used as representative of the traditional internal RGs in many studies, such as ACT (Actin), TUB (Tubulin), 18S (18S ribosomal RNA), UBC (Ubiquitin-conjugating enzyme), EF1α (Elongation factor-1 alpha) (Fiume and Fletcher, 2012; Mousavi et al., 2013; Sakuraba et al., 2014; Tian H. et al., 2015; Xu et al., 2015; Liu et al., 2016). However, many studies also show that housekeeping genes have been not stably expressed under any tissues and all experimental conditions (Schmittgen and Zakrajsek, 2000; Jain et al., 2006; Hong et al., 2008; Wang et al., 2014; Xu et al., 2016). For example, the expression of 18S rRNA was stable in long-duration salt treatment, but it was variable in drought treatment in Populus euphratica (Wang et al., 2014). EF1α was an ideal RG in different tissues and abiotic stresses, but it was not suitable in the different stages and hormone treatment in Brachypodium distachyon (Hong et al., 2008). The expression of ACT2/7 and ACT11 were unstable under stress treatments in soybean (Yim et al., 2015). UBC2 was also unstable in abiotic stress, organ and tissue in watermelon (Kong et al., 2014a).
Recent research found that many new RGs were better than traditional genes as ideal genes, e.g., PP2A (protein phosphatase 2A) in buffalo grass (Li et al., 2014) and hybrid roses (Klie and Debener, 2011), CUL1 (CULLIN 1) and APT3 (ADENINE PHOSPHORIBOSYL TRANSFERASE 3) in Marchantia polymorpha (Saint-Marcoux et al., 2015), Bic-C2, F-box protein 2 and VPS-like in soybean (Yim et al., 2015), RPL (ribosomal protein L) and RPS15 (cytosolic ribosomal protein S15) in melon fruits (Kong et al., 2016). Therefore, it is important to select suitable RGs under different conditions in various species.
Many studies of RGs expression have been reported in Arabidopsis thaliana (Remans et al., 2008), rice (Jain et al., 2006), potato (Nicot et al., 2005), tomato (Expósito-Rodríguez et al., 2008), watermelon (Kong et al., 2014a), melon (Kong et al., 2014b), poplar (Wang et al., 2014), sorghum (Reddy et al., 2016), tea plant (Wu et al., 2016), tree peony (Li et al., 2016), Eucalyptus (Oliveira et al., 2012), and so on. Nevertheless, no study on validating of RGs has been conducted in Salix genus until now. To data, Actin and Tubulin are the most widely used as RGs to analyze gene expression profiling in various tissues and under different abiotic stress in Salix matsudana Koidz (Liu M. et al., 2014; Song et al., 2015; Yang et al., 2015) and Salix suchowensis (Zhang et al., 2015). However, recent studies had demonstrated that both of ACT and TUB were poorly used as RG for qRT-PCR in developmental stages of leaf and hormone stimuli in tea plant (Wu et al., 2016), and ACT was not the optimal RG in melon fruits (Kong et al., 2016). TUB was unstable in different tissues and abiotic stress in sorghum (Reddy et al., 2016). So it is doubtful to use ACT or TUB as RG in the Salix study. Recently, with the completion of whole-genome sequence of Salix purpurea and Salix suchowensis, the researches of gene function in Salix are rapidly launched. Thus, the systematic exploration of RGs under different experimental conditions for accurate transcript normalization is important and requisite for the future research in Salix genus.
In this study, 14 endogenous genes were selected as candidate RGs, and their expression stabilities were detected across tissue types and under different abiotic stress treatments in S. psammophila. Three software (NormFinder, geNorm, and BestKeeper) were employed to analyze systematically the qRT-PCR data, and the outputs were merged using RankAggreg software. In order to validate the suitability of these RGs, the relative genes expression profiles of one tissue-specific gene, WOX1a (WUSCHEL-related homeobox 1) (Nakata et al., 2012; Liu B. et al., 2014) and four stress-inducible genes, Hsf-A2 (Heat shock transcription factors A2) (Zhang et al., 2015), CBF3 (C-repeat binding factor 3) (Hu et al., 2013), HKT1 (High-Affinity K+ Transporter 1) (Waters et al., 2013), and GST (Glutathione S-transferase) (Roxas et al., 1997) were conducted to confirm the validity of the RGs in this study. This work provide a list of suitable RGs for future gene expression studies in S. psammophila.
Materials and Methods
Plant Materials and Stress Treatments
Two-month-old seedlings of S. psammophila clones were cultured using Hoagland nutrient solution and grown in a growth chamber under long-day conditions (16 h light/8 h dark) at 25/22°C (day/night). Various tissues, including leaf (L), primary stem (PS), secondary stem (SS), root (R), female catkin (FC), and male catkin (MC) were collected from the S. psammophila seedlings. For stress treatments, the seedlings were subjected to abiotic stresses under heat (42°C), cold (4°C), salt (200 mM NaCl), and 22% (w/v) polyethylene glycol (PEG 6000)-simulated drought treatments. Leaves from individuals were harvested at six selected time points (0, 6, 12, 24, 48, and 96 h) during each stress treatments, frozen immediately in liquid nitrogen, and stored at -80°C for further analysis. Three biological replicates were performed in different pots for each treatment.
Total RNA Isolation and cDNA Synthesis
RNA of different tissues and different stress treatments were extracted using the RNeasy Plant Mini Kit (Qiagen, Germany) according to the manufacturer’s instructions. RNA concentration were determined by NanoDrop8000 (Thermo Scientific, USA). Only the RNA sample with an absorbance ratio at OD260/280 between 1.9 and 2.2 and OD260/230 greater than 2.0 were used for further analysis and the RNA integrity were verified by 1.0% agarose gel-electrophoresis. First-strand cDNA synthesis was carried out with approximately 4 μg RNA using the SuperScript III reverse transcription kit (Invitrogen, USA) according to the manufacturer’s procedure, and the final product was diluted 40-fold for experiments.
Selection of Candidate RGs and Primers Design
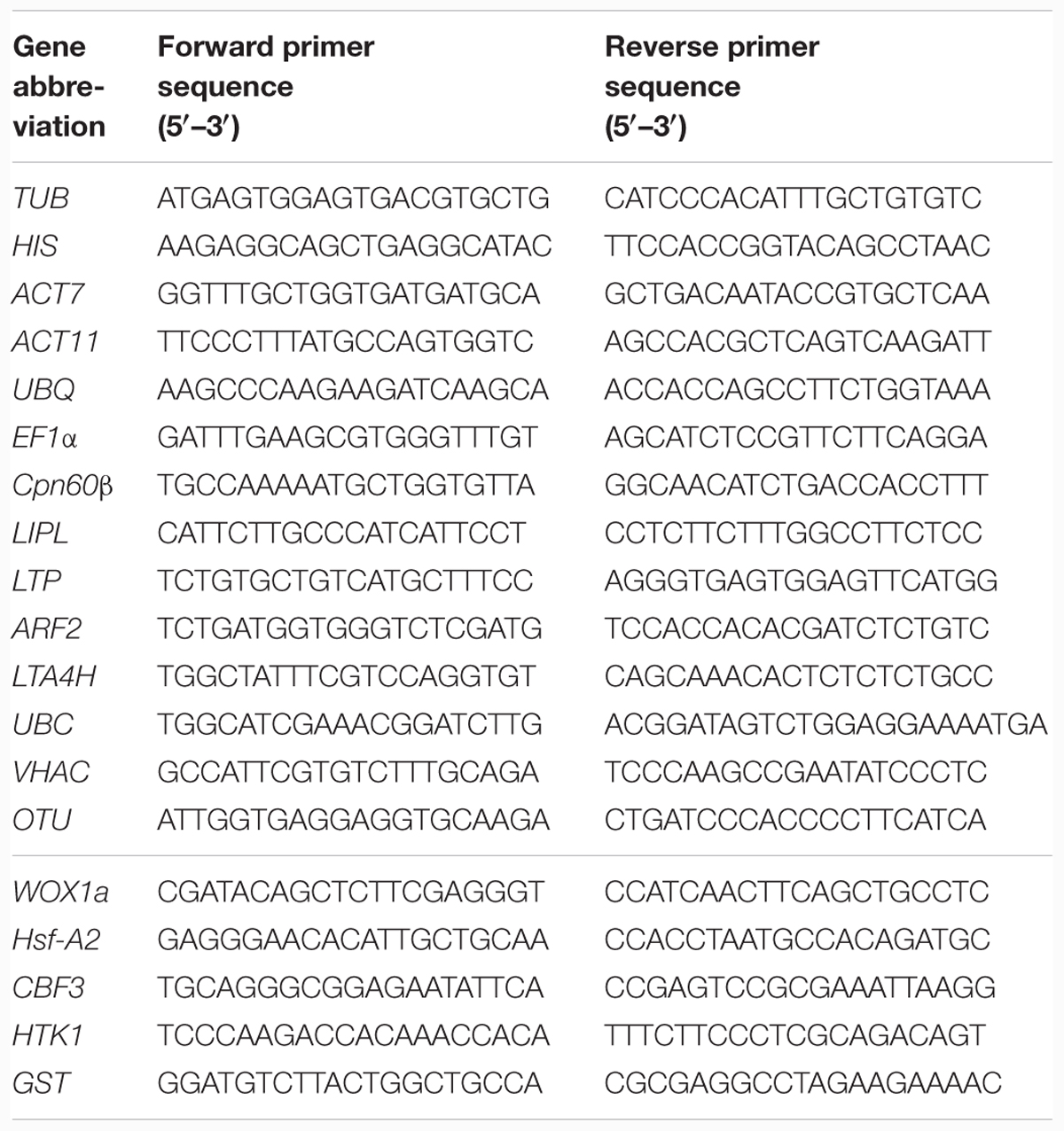
Here, we selected a total of 14 potential candidate RGs, including six novel RGs [LIPL (Lipocalin-like gene), LTP (Seed storage/lipid transfer protein), ARF2 (ADP-ribosylation factor 2), LTA4H (Leukotriene A-4 hydrolase homolog), VHAC (V-type proton ATPase subunit C) and OTU (OTU-like cysteine protease family protein)] and eight traditional RGs [TUB (Tubulin beta chain), HIS (Histone superfamily protein H3), ACT7 (Actin 7), ACT11 (Actin 11), UBC (Ubiquitin-conjugating enzyme E2), UBQ (Ubiquitin family), EF1α (Elongation factor-1 alpha) and Cpn60β (60-kDa chaperoninβ-subunit)] (Table 1) (Wang et al., 2014; Yim et al., 2015; Ma et al., 2016; Wu et al., 2016). Among of them, ARF2, LTA4H, VHAC, and OTU were selected as candidate RGs because they exhibited stable expression in the drought stress based on our transcript data (Unpublished). In addition, the traditional RGs and the two novel genes (LTP and LTPL) were chosen based on the previous studies. The sequence of these candidate RGs were obtained from transcript data. The Primers were designed using Primer3 software1 in the specific region of sequences, with melting temperatures of 58–62°C, 20–23 bp primer length and amplified product size of 150–250 bp. All the primer sequences in this study were listed in Table 2.
qRT-PCR Conditions
To verify the specificity of all the primer sets, PCR was performed using pooled cDNA as templates, and the PCR products were examined by 2% (w/v) agarose gel electrophoresis. The amplicons should appear as a single band with the correct size. qRT-PCR reactions were conducted on LightCycler® 480 Real Time PCR System (Roche, Germany). The 20 μl reaction system contained 10 μl of SYBR Premix Ex TaqTM (TaKaRa, China), 2 μl of cDNA template, 0.8 μl of each primer, and 6.4 μl ddH2O. The PCR conditions were conducted by the manufacturer (pre-incubation of 5 min at 95°C, amplification of 45 cycles of 95°C for 10 s, 60°C for 10 s and 72°C for 10 s, melting curve analysis by heating the PCR products from 65 to 95°C, finally by cooling at 40°C for 30 s). The final threshold cycle (Ct) values were the mean of three values for each treatment and four technical replicates.
Data Analysis of Gene Expression Stability
Expression levels of the 14 RGs in all samples were determined by their cycle threshold values (Ct) which using the formula: 2-ΔCt, in which ΔCt = each corresponding Ct value – minimum Ct value. Standard curves were generated using dilutions of the mixed cDNA template (1, 1/5, 1/25, 1/125, 1/625/, 1/3125). The correlation coefficients (R2 values) and slope (S) values can be obtained from standard curves. And the PCR efficiency (E) was derived from the expression [5(1/-S) - 1] × 100% (Han et al., 2012).
Three software algorithms, geNorm (Vandesompele et al., 2002), NormFinder (Andersen et al., 2004), and BestKeeper (Pfaffl et al., 2004) were used to rank the stability of the selected RGs across all the experimental sets. The methods were performed according to Han et al. (2012). The RankAggreg package of R program v.3.2.32 (Pihur et al., 2009) was used to merge the stability measurements obtained from the three software algorithms and establish a consensus rank of RGs according to the method described by Wang et al. (2014).
Validation of Reference Genes
In order to validate the applicability of the identified RGs, the expression levels of five target genes, including one tissue-specific gene WOX1a and four stress-inducible genes, including Hsf-A2 for heat stress, CBF for cold stress, HKT1 for salt stress, and GST for drought stress were analyzed using the most and least stable RGs. The average Ct value was calculated from three biological replicates and four technical replicates and used for relative expression analyses. The relative gene expression data were calculated according to the 2-ΔΔCt method and presented as fold change (Schmittgen and Livak, 2008).
Results
Verification of Amplicons, PCR Specificity, and Amplification Efficiency
A total of 14 candidate RGs were selected for qRT-PCR analysis. Gene name, the homologous gene accession in S. purpurea and S. suchowensis, amplicon length, amplification efficiency (E) and correlation coefficients (R2) were listed in Table 1. The amplified products were also further analyzed by 2% agarose gel electrophoresis and only one band of the expected size was detected, no primer dimers or non-specific amplification in each experiment (Supplementary Figure S1A). For PCR specificity detection, the presence of a single peak in the melting curve was obtained after amplification (Supplementary Figure S1B). Amplification efficiency of PCR reactions varied from 89.13% for Cpn60β to 119.11% for VHAC, and correlation coefficients of the standard curve ranged from 0.9873 for VHAC to 0.9996 for EF1α (Table 1).
Expression Profiles of Candidate Reference Genes
The RNA from different tissues and stress treatments were reverse transcribed into cDNA which was used as templates for qRT-PCR. The Ct values were obtained from each reaction with 14 primer pairs. Lower Ct values mean higher expression abundance, and the higher Ct values mean lower expression abundance. The average Ct values ranged from 17.72 to 29.76 in different tissues and four treatments (Figure 1A); EF1α had the highest expression level with Ct of 17.72 cycle, whereas LTP was the lowest expression abundance with Ct values as high as 29.76 cycle.
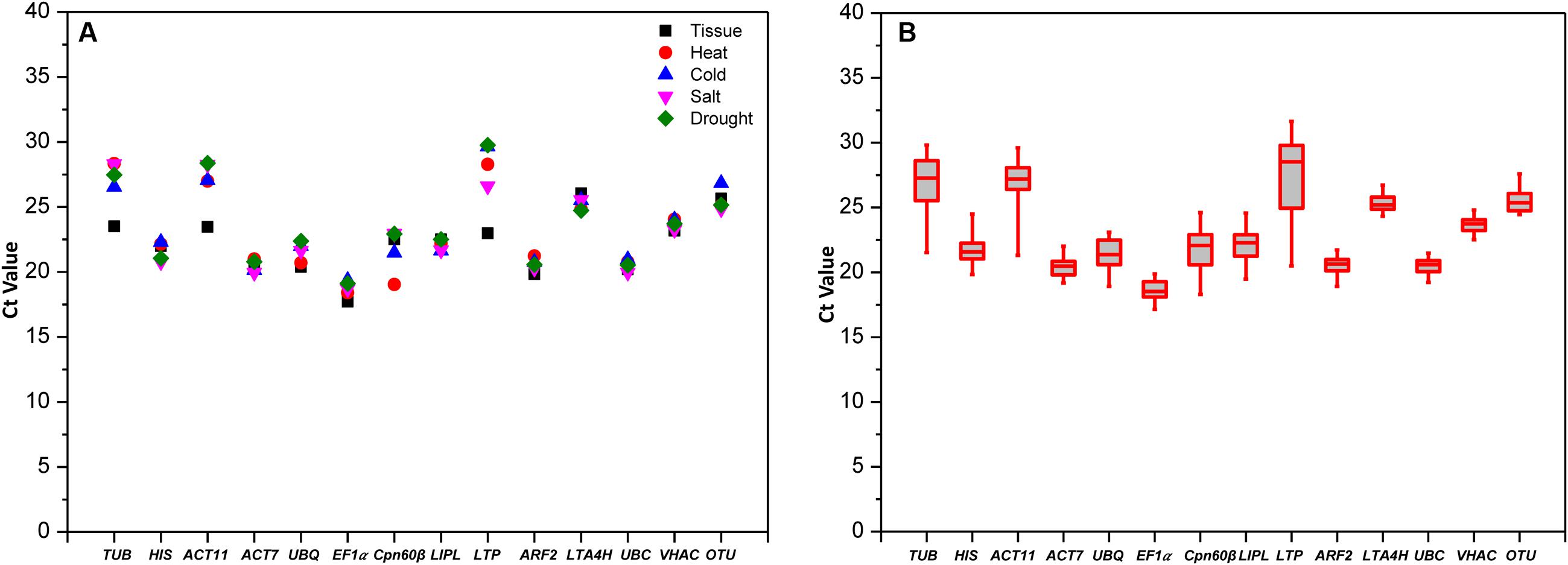
FIGURE 1. qRT-PCR Ct values of the 14 candidate RGs in all experimental samples. (A) Mean values in different treatments. (B) Box-whisker plot showing the Ct variation among 30 test samples. A line across the box depicts the median. In each box, the upper and lower edges indicate the 25th and 75th percentiles.
The Ct values data distribution of genes in all tested samples are shown as a box-plot in Figure 1B, the more narrow range of Ct values indicates that the gene is more stability in the conditions. The Ct data of UBC (Ct values from 19.22 to 21.48) with most concentration means it was the most stable gene in all the samples, and the followed stable genes were VHAC and LTA4H with Ct values from 22.52 to 24.81 and from 24.33 to 26.72, respectively. The greatest variation three genes were displayed by LTP, ACT11, and TUB (more than six Ct values), while the Ct values of LTP were the most scattered (Ct values from 20.50 to 31.64). These results indicated that the selected genes showed variable expression levels in different tissues and under four abiotic stresses, so it was necessary to screen out the best RG for target gene expression normalization.
Statistical Analysis of qRT-PCR Data Using Three Bioinformatics Programs
In order to detect the stabilities of 14 candidate RGs, the three Excel-based software (NormFinder, geNorm, and BestKeeper) were used for statistical analysis. Data from different tissues and four treatments were analyzed separately, and then added together. The analysis results were as follows:
geNorm Analysis
geNorm was used to rank the RGs by calculating gene expression stability value M based on the average pairwise expression ratio (Vandesompele et al., 2002). M-value is negatively correlated with gene stability which means the lower M-value of gene was considered as more stable gene. In this study, M-value setting 1.5 which recommends by geNorm Program was used to identify RGs with stable expression (Vandesompele et al., 2002). As shown in Figure 2, the value of all the 14 RGs were all below 1.5 in different tissues and treatments. In different tissues, EF1α, OTU, and UBC showed highly stable expression with the M-value below 0.5, whereas Cpn60β was the least stable gene with the M-value 1.38 (Figure 2A). In heat, cold and salt treatments, 8 of 14 genes were below 0.5 (Figures 2B–D). In drought treatment (Figure 2E), 11 of 14 genes were below 0.5. In heat and drought treatment, UBC and LTA4H were the most stable genes and the least stable gene was LTP (Figures 2B,E), while the LTP was also the least stable gene in salt treatment and the most stable gene were OTU and ACT7 in salt treatment (Figure 2D). In cold treatment, HIS and VHAC were stably expressed (Figure 2C). Overall, all of the tested genes except LTP had low M-value s of less than 1.5, and UBC and ARF2 were the most stable genes in all the samples (Figure 2F).
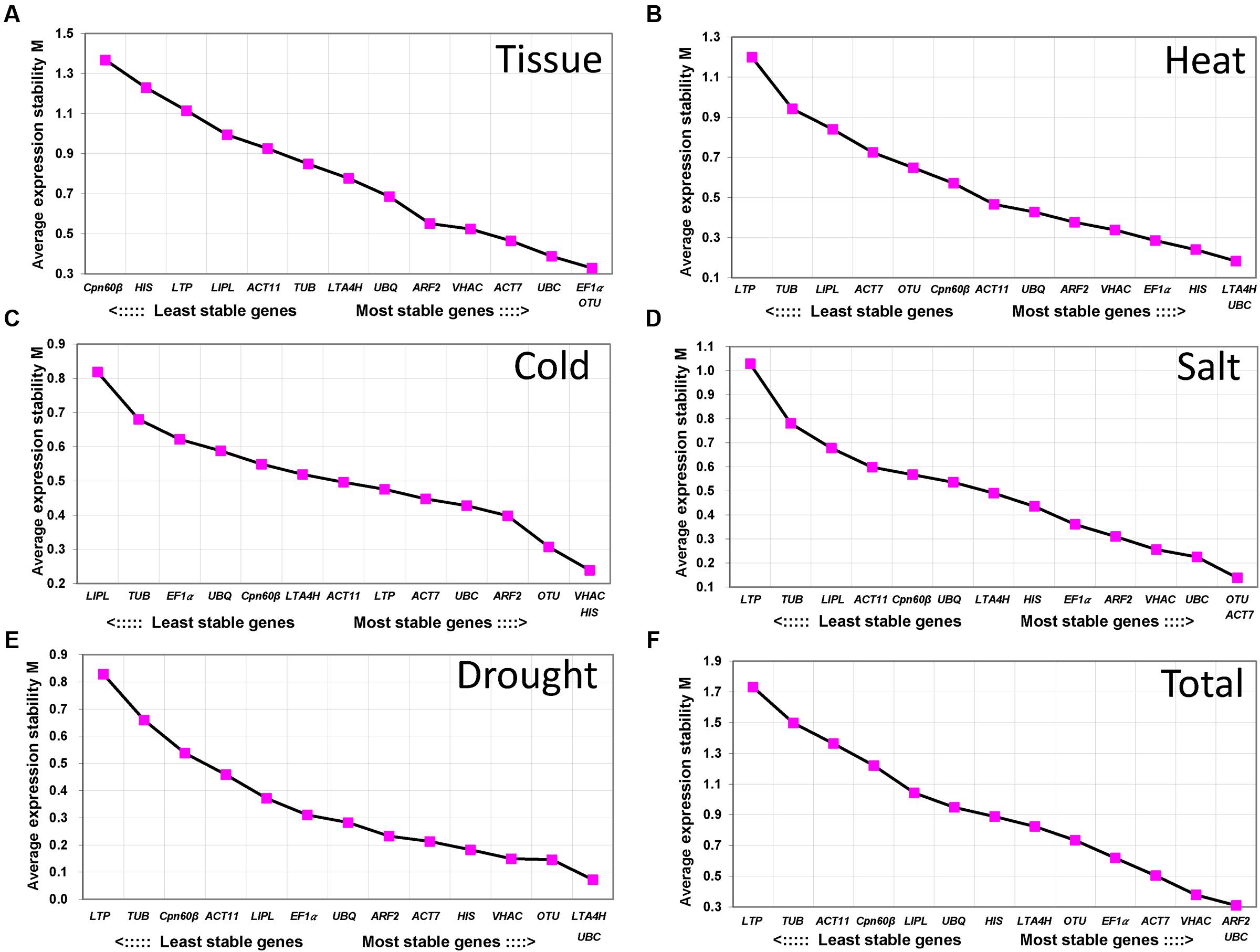
FIGURE 2. Average expression stability values (M) and ranking of the candidate RGs calculated using geNorm. M-value s and ranking of the candidate RGs in tissue (A), heat stress (B), cold stress (C), salt stress (D), drought stress (E), and total sample (F). A lower value of the average expression stability indicates more stable expression.
In some experiments, one RG can’t meet the requirement to effective normalize the gene expression and two or more RGs were required. In present study, the pairwise variation (Vn/Vn+1, n corresponds to the number of RGs) was calculated by geNorm program to evaluate the optimal number of the RGs required for accurate normalization (Vandesompele et al., 2002; Han et al., 2012). The value of Vn/Vn+1 generally set 0.15 as the threshold limit. When Vn/Vn+1 < 0.15, it means the number of RGs less than or equal to value of n is sufficient to use as RGs. In the subsets of tissues and treatments, all the V2/3 value were below 0.15, which suggested that two RGs would be enough used for normalization (Figure 3). Thus, the combined use of the two most stable RGs would be effective for normalizing gene expression studies.
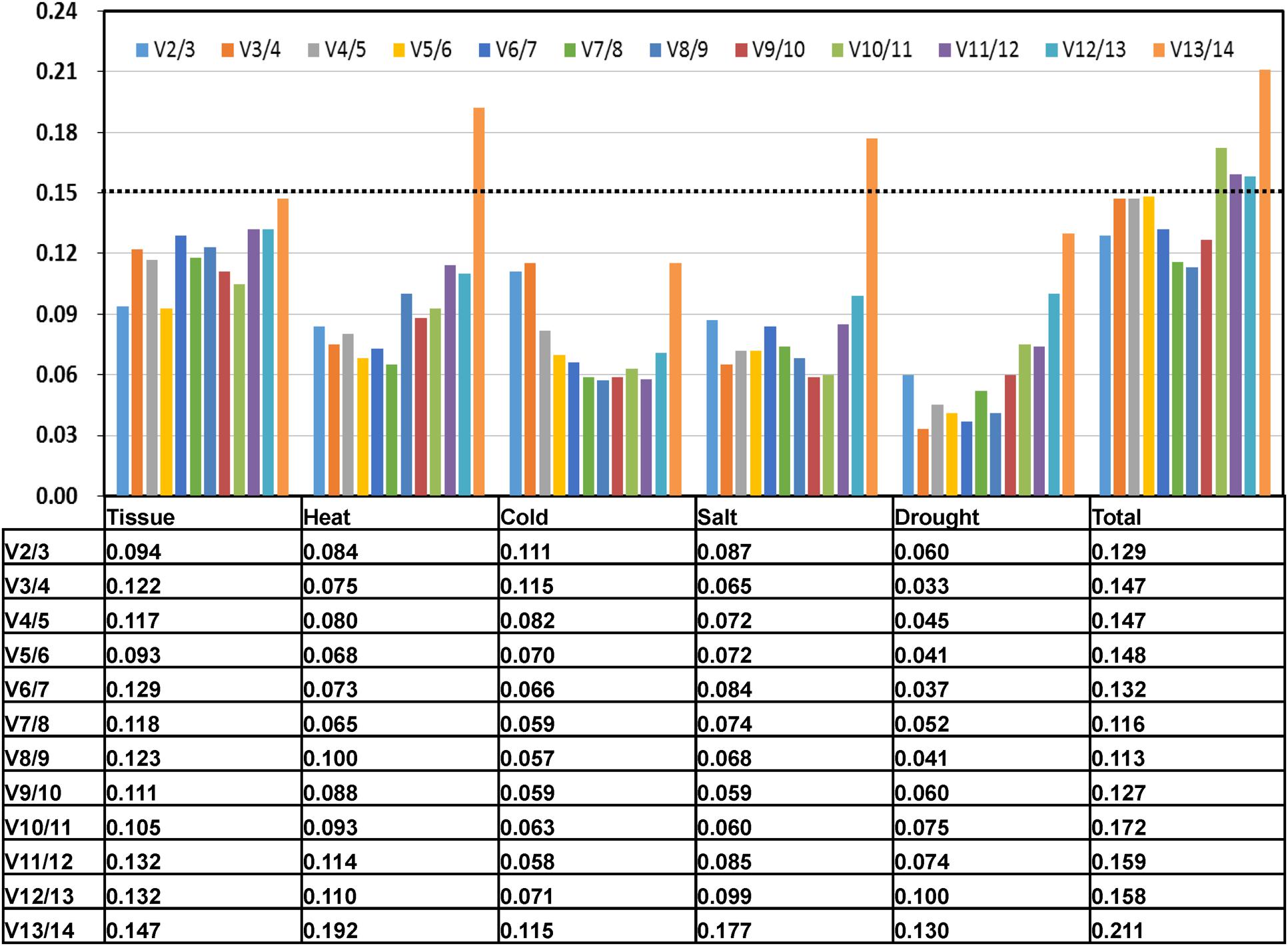
FIGURE 3. PV to determine the optimal number of control genes for accurate normalization. The pairwise variation (Vn/Vn+1) was analyzed between the NFn and NFn+1 using geNorm software, where n is the number of genes involved in the NF.
NormFinder Analysis
NormFinder is also used to evaluate the stability of tested genes, which based on calculating the stability value for each gene (Andersen et al., 2004). The lowest stability value indicates the most stably expressed gene. The stability values of the tested genes are analyzed by NormFinder and the results were shown in Table 3. In different tissues, the most stable gene was EF1α (stability value 0.0560) and followed by OTU (0.1269), UBC (0.1364), and ACT7 (0.2222). In heat treatment, the UBC (0.0647), LTA4H (0.0818), and EF1α (0.1113) had the most stable expression. In cold treatment, the three most stable genes were HIS (0.0416), ARF2 (0.1061), and OTU (0.1189), while the most stable genes were OTU (0.0937), VHAC (0.1179), and ACT7 (0.1466) in salt treatment. In cold and salt treatment, the expression levels of TUB was the most unstable. In drought treatment, VHAC (0.0293), ACT7 (0.0579), and ARF2 (0.0739) had stable expression. Finally, UBC (0.1272), ARF2 (0.1542), and VHAC (0.1695) were the stable RGs, and the LTP (0.3978) was the least stable RG in total.
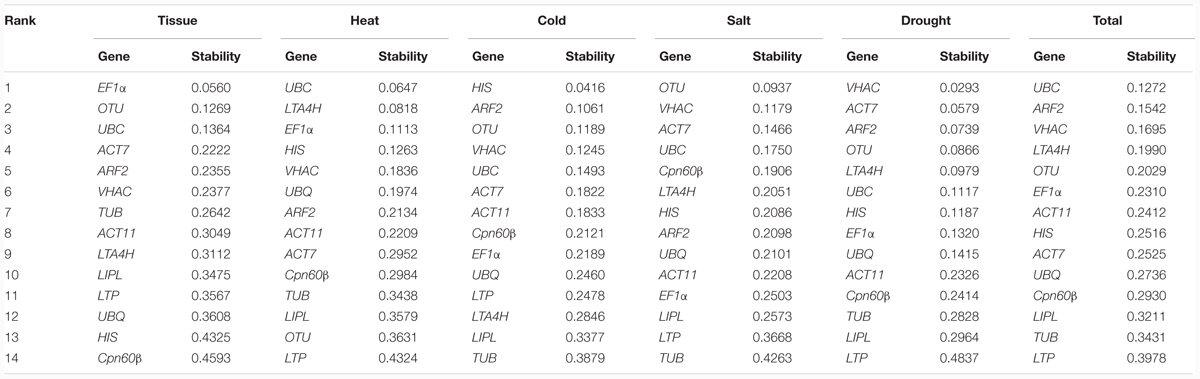
TABLE 3. Rankings of candidate RGs for normalization and their expression stability values calculated using the NormFinder program.
BestKeeper Analysis
BestKeeper program determines the stability ranking of the RGs based on calculating the coefficient of variance (CV) and the standard deviation (SD) of the Ct values (Pfaffl et al., 2004). The rankings of BestKeeper analysis were shown in the Table 4. For the different tissues, the EF1α (1.56 ± 0.28) was identified as the most stable gene in different tissues, UBC (0.55 ± 0.11) in heat treatment, ARF2 (1.10 ± 0.23) in cold treatment, OTU (0.82 ± 0.20) in salt treatment, and HIS (0.40 ± 0.08) in drought treatment. In total, VHAC (1.92 ± 0.45) was the most stable gene and followed by UBC (2.29 ± 0.47), while the LTP (9.61 ± 2.64) was the least stable gene.
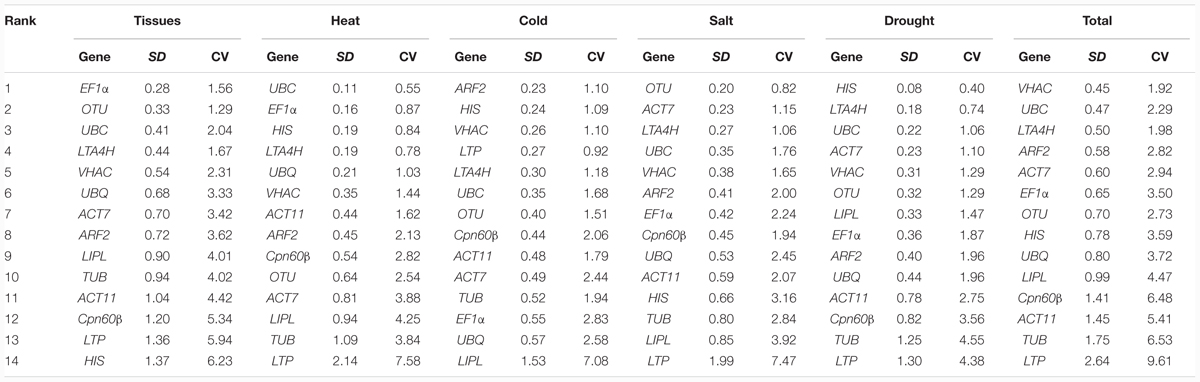
TABLE 4. Rankings of candidate RGs in order of their expression stability as calculated by BestKeeper.
Consensus List and Validation of the Stability of S. psammophila Reference Genes
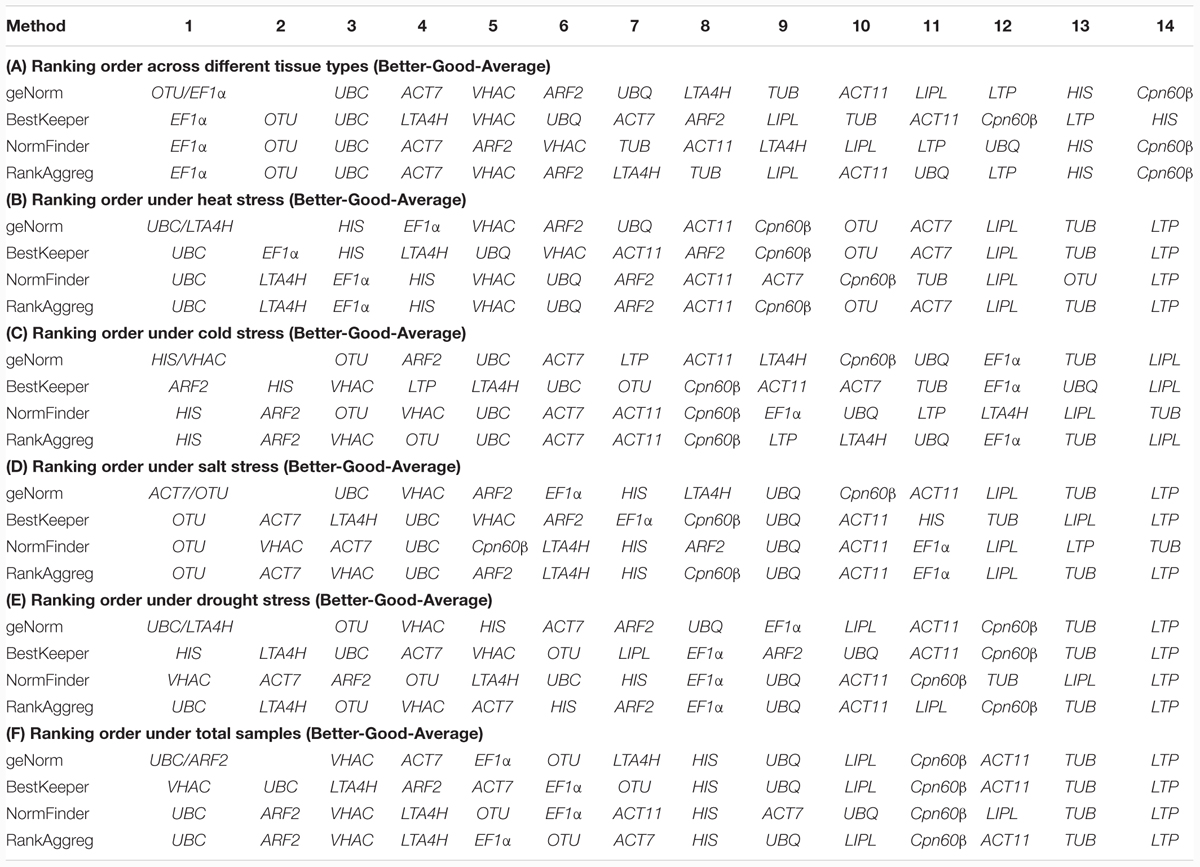
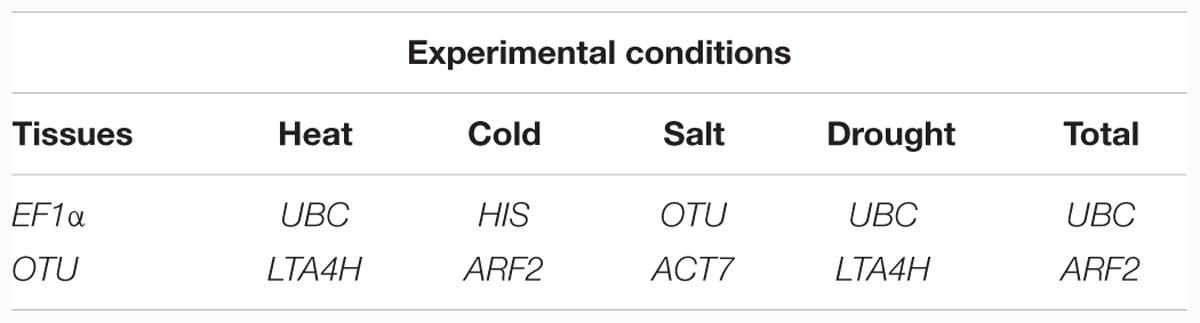
In this study, three different software programs were used to analyze the stability of the 14 tested genes. However, different software use the different algorithm in gene ranking, so the results in ranking patterns showed little differences. In order to provide a comprehensive result, the RankAggreg software (Najafpanah et al., 2013) was used to rank an optimal lists of RGs. According to the analysis of RankAggreg (Figure 4; Table 5), the best RG for normalization was EF1α for different tissue types, UBC for heat and drought treatment, HIS for cold treatment, and OTU for salt treatment. When considering all the sample, the UBC was the best choice as the optimal RG (Figures 4A–F). The comprehensive ranking list by RankAggreg was shown in Table 5. Based on the optimal number of the RGs calculated by geNorm and the raking list by RankAggreg, the best combination of RGs in different subsets were shown in Table 6. The results shown that UBC and LTA4H were the best combination as RGs in heat and drought stress, while UBC and ARF2 in total samples. In addition, the best combination included EF1α and OTU in tissues, HIS and ARF2 in cold stress, OTU and ACT7 in salt stress.
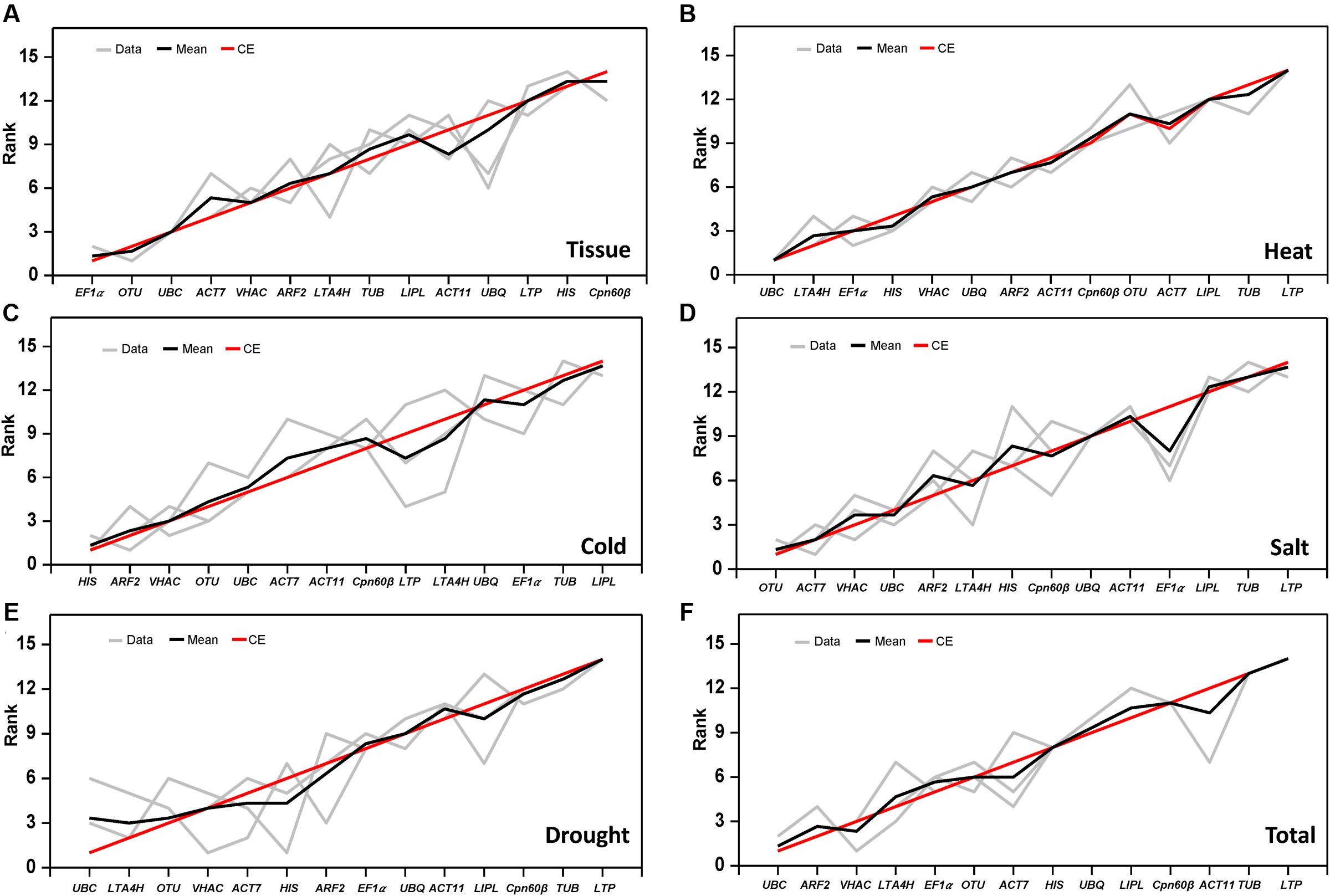
FIGURE 4. Rank aggregation of gene lists using the Monte Carlo algorithm. The solution of the rank aggregation in tissue (A), heat stress (B), cold stress (C), salt stress (D), drought stress (E), and total sample (F) are shown in a plot in which genes are ordered based on their rank position according to each stability measurement (gray lines). The mean rank position of each gene is shown in black lines, as well as the model computed by the Monte Carlo algorithm (red lines).
Reference Gene Validation
In order to assess the effect of the selected RG on the outcome of a practical experiment, the transcription levels of target genes were evaluated using the most and least stable RGs (Figure 5). WOX1a was reported high expression abundance in leaf in A. thaliana and poplar (Nakata et al., 2012; Liu M. et al., 2014), and it was detected for different tissues. The expression of WOX1a was high expression abundance in leaf when using both of EF1α and OTU or only EF1α as RG, but high expressed in female catkin when normalized by the least stable gene Cpn60β (Figure 5A). The expression level of Hsf-A2 (Zhang et al., 2015), CBF3 (Hu et al., 2013), HKT1 (Waters et al., 2013), GST (Roxas et al., 1997) were detected in heat, cold, salt, and drought stress, respectively. For example, in drought stress treatment, when the two stable RGs UBC and LTA4H or only UBC were/was used for normalization, the expression levels of HKT1 gradually increased from 0 h to 24 h and reached maximum at 24 h and subsequently decreased at 48 and 96 h (Figure 5E). However, when LTP was used as RG for normalization, different expression patterns were generated that the expression levels of HKT1 peaked at 6 h and decreased at 12 h. Meantime, the expression pattern of the other target genes are also obvious different in the expression tendency when different RGs were used to normalize the data in heat, cold and salt (Figures 5B–D). These results confirmed the importance of using suitable gene as RG in different experiments.
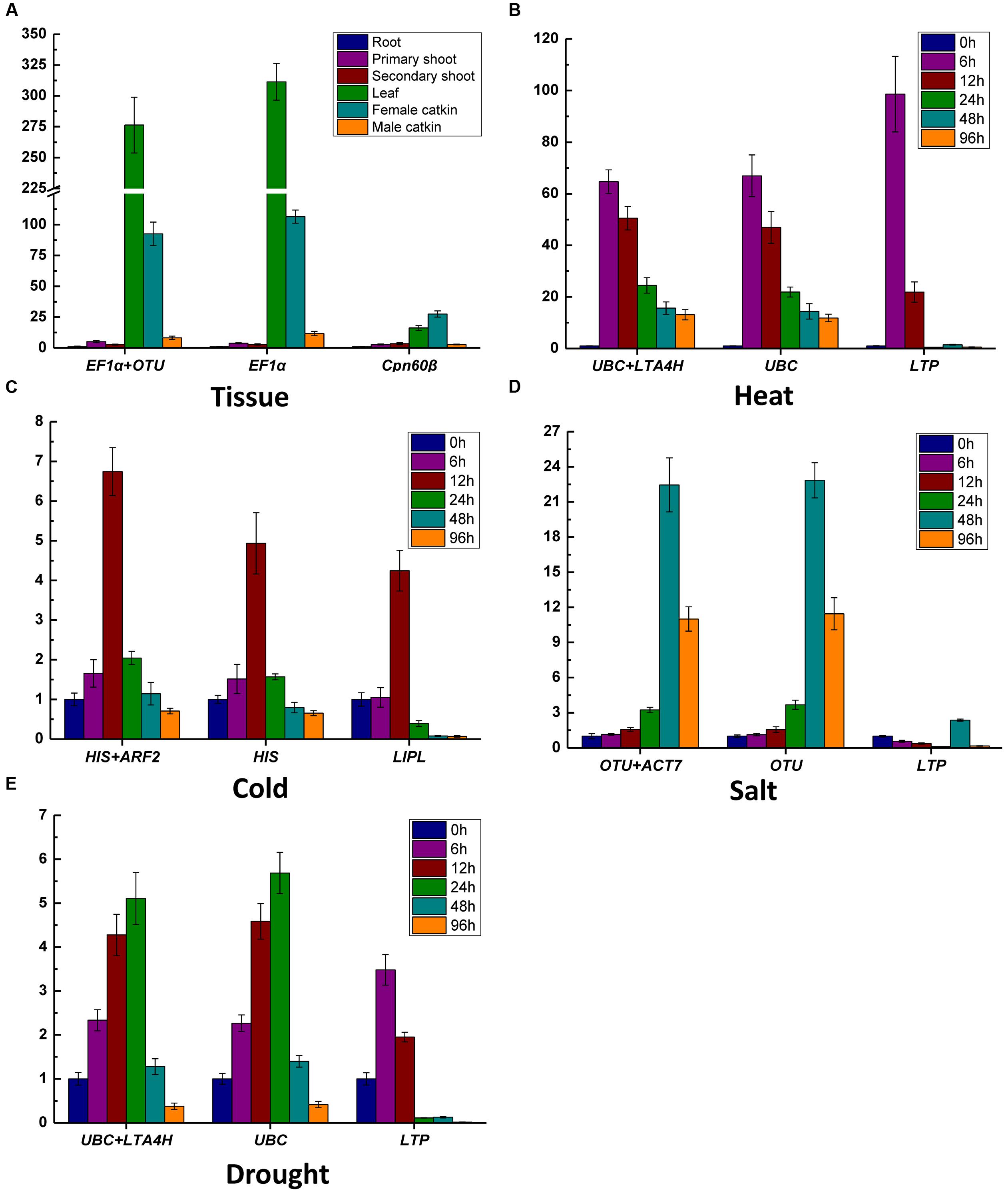
FIGURE 5. Validation of the identified reference genes. The relative expression levels of WOX1a (A), Hsf-A2 (B), CBF3 (C), HKT1 (D), and GST (E) were normalized by the most stably RGs and the least stable RGs in the different samples. The transcript abundance of WOX1a in root was used as control in (A), and the transcript abundances of four genes (Hsf-A2, CBF3, HKT1, and GST) at 0 h were set to be the control in (B–E), individually. Values shown are relative expression levels ±SE (n = 3 experiments).
Discussion
Presently, qRT-PCR as a revolutionary technology has been widely used in detecting the gene expression profiling (Bustin, 2002; Radonić et al., 2004). But unsuitable RGs which used to normalize the experimental data in qRT-PCR will cause the deviation of results. So the aim of this study was looking for the ideal RGs with a stable expression levels in various expression conditions in S. psammophila.
Here, 14 candidate RGs including eight traditional genes and six novel genes were screened across different tissue types as well as under four abiotic stress treatments. UBC, VHAC and LTA4H with narrow Ct values were more stable, while the house-keeping genes TUB and ACT11 with Ct values were scattered and reach to more than six cycles showing great expression variation (Figure 1). It was consistent with the recent studies that the expression of traditional RGs were not always stable in some experiments (Schmittgen and Zakrajsek, 2000; Jain et al., 2006; Hong et al., 2008; Wang et al., 2014; Kong et al., 2016; Xu et al., 2016).
The optimal number of stable RGs depends on accurate calculation following certain experimental conditions. In our study, two stable RGs can ensure the accuracy to detect the genes expression in different tissues and four stress experiment followed the analysis of geNorm program (Figure 3). While in pearl millet, two RGs were enough for individual stress and developmental tissues and three RGs were need in multiple stress (Shivhare and Lata, 2016). In watermelon, nine RGs were required for normalization of target genes for organ and tissue, but two RGs were enough for biotic stress (Kong et al., 2014a). In tung tree, two RGs would be sufficient to normalize the expression of the target gene in different tissues, but at least five RGs are required to ensure the accuracy of the expression in all the sample (Han et al., 2012). It also proved five RGs for salicylic acid (SA) stress and three RGs for heat and gibberellin (GA) stress conditions in carrot (Tian C. et al., 2015). Four RGs were needed for data normalization in melon leaf and root (Kong et al., 2014b), and five RGs were needed for data normalization in melon fruits (Kong et al., 2016).
Three different software packages including geNorm, NormFinder, and BestKeeper were employed to evaluate gene expression stability in our analysis. The different calculation algorithms of the three software lead to the divergence in the gene ranking, especially in the top ranked gene (Table 5). For instance, under drought stress UBC and LTA4H were identified as the best RG by geNorm, while HIS was the best RG predicated by BestKeeper, and VHAC was identified as the most ideal RG by NormFinder program. The difference ranking by three program was also presented in other studies (Kong et al., 2014a; Kanakachari et al., 2016; Li et al., 2016; Wu et al., 2016). In order to summarize the ranking of our dataset analysis, the RankAggreg software was used to merge the data (Wang et al., 2014). The best combination of RGs was get based on the analysis of geNorm and RankAggreg (Figure 4; Tables 5 and 6). Thereinto, three house-keeping genes (UBC, EF1α, and HIS) were the good choice as RGs in this study. These genes are also often considered as reliable RGs in other studies. For example, UBC was one of the most stable genes in salt and cold stress in Lycoris aurea (Ma et al., 2016) and it also showed high stability in the different cultivars of Tree Peony (Li et al., 2016). EF1α was one of the best RG which has been confirmed in pearl millet (Shivhare and Lata, 2016), rice (Jain et al., 2006), Petunia hybrid (Mallona et al., 2010), and P. euphratica (Wang et al., 2014).
In the present study, six new RGs (OTU, LTA4H, ARF2, VHAC, LIPL, and LTP) were detected as the candidate RGs. Based on the results, we found OTU, LTA4H, ARF2, and VHAC were the ideal choice for RGs in special experimental condition. Among of them, OTU gene family exist in various organisms, and involved in the de-ubiquitination signaling (Frias-Staheli et al., 2007), but it is the first time to be reported as a stable RG in plant study. LTA4H is one of the best choice under heat stress and drought stress. It is a bifunctional zinc enzyme which generates the inflammatory mediator leukotriene B4 (LTB4) and involved in many defense mechanism in human, like as inflammation, immune responses and others (Haeggstrom, 2000; Thunnissen et al., 2001), The structure and characterization also were studied in Saccharomyces cerevisiae (Kull et al., 1999), but the understanding of the function of LTA4H in plant is limited. ARF proteins were defined by their ability to act as cofactors in the cholera toxin-catalyzed ADP-ribosylation of G proteins and play role in membrane transport, maintenance of organelle integrity, and the activation of phospholipase D (Bhamidipati et al., 2000). VHAC is part of V-type proton ATPase (Beyenbach and Wieczorek, 2006) and plays as a regulatory linker protein between the V-ATPase and the actin-based cytoskeleton (Marshansky and Futai, 2008). And the recent study reported that VHAC involved in the drought stress (Gao et al., 2011). These four novel genes were the suitable RGs because of the stable expression in different experiment conditions, but the limited information about these genes were known in plant and the further study will be necessary. In our study, other two novel genes (LIPL and LTP) were studied as the candidate RGs. These two genes had been studied as RGs in P. euphratica in which LTP was the most stable gene during cold, but not for other treatment (Wang et al., 2014). It also revealed the RGs was different in species and experimental conditions.
To validate the suitability of the RGs we identified, the expression of WOX1a, Hsf-A2, CBF3, HKT1, and GST were detected in various tissues and abiotic stress using different RGs. The date once again demonstrated that RGs play a key role in normalizing the data of the qRT-PCR, and the inappropriate RGs may lead to incorrect results for the target genes.
Conclusion
The expression stability of 14 candidate RGs were detected across tissue types and under four abiotic stress treatments using four software in S. psammophila. And the optimum RG and the best combination of RGs were identified in different experiment conditions. To the best of our knowledge, it is the first study to identify the suitable RGs for normalizing the gene expression studies using qRT-PCR in S. psammophila. And it provides new information which will be useful to analyze the expression profiles and function of target genes involved in the development and stress tolerance in S. psammophila.
Author Contributions
JL and HJ performed most of the experiments and wrote the manuscript. XH designed the experiments and edited the manuscript. JZ and PS helped in data collection, sample preparation, and RNA extraction. ML and JH coordinated the project, conceived, and designed the experiments. All authors read and approved the final manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by National Non-profit Institute Research Grant of CAF (CAFYBB2014ZX001-4) and (TGB2013009) to JH. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2016.01505
Footnotes
References
Andersen, C. L., Jensen, J. L., and Ørntoft, T. F. (2004). Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250. doi: 10.1158/0008-5472.CAN-04-0496
Bao, Y., and Zhang, G. (2012). Study of adsorption characteristics of methylene blue onto activated carbon made by Salix psammophila. Energy Procedia 16, 1141–1146. doi: 10.1016/j.egypro.2012.01.182
Beyenbach, K. W., and Wieczorek, H. (2006). The V-type H+ ATPase: molecular structure and function, physiological roles and regulation. J. Exp. Biol. 209, 577–589. doi: 10.1242/jeb.02014
Bhamidipati, A., Lewis, S. A., and Cowan, N. J. (2000). ADP ribosylation factor-like protein 2 (Arl2) regulates the interaction of tubulin-folding cofactor D with native tubulin. J. Cell. Biol. 149, 1087–1096. doi: 10.1083/jcb.149.5.1087
Bustin, S. A. (2002). Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems. J. Mol. Endocrinol. 29, 23–39. doi: 10.1677/jme.0.0290023
Bustin, S. A., Benes, V., Nolan, T., and Pfaffl, M. W. (2005). Quantitative real-time RT-PCR-a perspective. J. Mol. Endocrinol. 34, 597–601. doi: 10.1677/jme.1.01755
Expósito-Rodríguez, M., Borges, A. A., Borges-Pérez, A., and Pérez, J. A. (2008). Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol. 8:131. doi: 10.1186/1471-2229-8-131
Fiume, E., and Fletcher, J. C. (2012). Regulation of Arabidopsis embryo and endosperm development by the polypeptide signaling molecule CLE8. Plant Cell 24, 1000–1012. doi: 10.1105/tpc.111.094839
Frias-Staheli, N., Giannakopoulos, N. V., Kikkert, M., Taylor, S. L., Bridgen, A., Paragas, J., et al. (2007). Ovarian tumor domain-containing viral proteases evade ubiquitin-and ISG15-dependent innate immune responses. Cell Host Microbe 2, 404–416. doi: 10.1016/j.chom.2007.09.014
Gao, C. Q., Wang, Y. C., Jiang, B., Liu, G. F., Yu, L. L., Wei, Z. G., et al. (2011). A novel vacuolar membrane H+-ATPase c subunit gene (ThVHAc1) from Tamarix hispida confers tolerance to several abiotic stresses in Saccharomyces cerevisiae. Mol. Biol. Rep. 38, 957–963. doi: 10.1007/s11033-010-0189-9
Haeggstrom, J. Z. (2000). Structure, function, and regulation of leukotriene A4 hydrolase. Am. J. Respir. Crit. Care Med. 161, S25–S31. doi: 10.1164/ajrccm.161
Han, X. J., Lu, M. Z., Chen, Y. C., Zhan, Z. Y., Cui, Q. Q., and Wang, Y. D. (2012). Selection of reliable reference genes for gene expression studies using real-time PCR in tung tree during seed development. PLoS ONE 7:e43084. doi: 10.1371/journal.pone.0043084
Hong, S. Y., Seo, P. J., Yang, M. S., Xiang, F. N., and Park, C. M. (2008). Exploring valid reference genes for gene expression studies in Brachypodium distachyon by real-time PCR. BMC Plant Biol. 8:112. doi: 10.1186/1471-2229-8-112
Hu, Y. R., Jiang, L. Q., Wang, F., and Yu, D. Q. (2013). Jasmonate regulates the inducer of CBF expression–c-repeat binding factor/DRE binding factor1 cascade and freezing tolerance in Arabidopsis. Plant Cell 25, 2907–2924. doi: 10.1105/tpc.113.112631
Huang, J., Zhou, Y., Yin, L., Wenninger, J., Zhang, J., Hou, G., et al. (2015). Climatic controls on sap flow dynamics and used water sources of Salix psammophila in a semi-arid environment in northwest China. Environ. Earth Sci. 73, 289–301. doi: 10.1007/s12665-014-3505-1
Jain, M., Nijhawan, A., Tyagi, A. K., and Khurana, J. P. (2006). Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR. Biochem. Biophys. Res. Commun. 345, 646–651. doi: 10.1016/j.bbrc.2006.04.140
Kanakachari, M., Solanke, A. U., Prabhakaran, N., Ahmad, I., Dhandapani, G., Jayabalan, N., et al. (2016). Evaluation of suitable reference genes for normalization of qPCR gene expression studies in Brinjal (Solanum melongena L.) during fruit developmental stages. Appl. Biochem. Biotechnol. 178, 433–450. doi: 10.1007/s12010-015-1884-8
Klie, M., and Debener, T. (2011). Identification of superior reference genes for data normalisation of expression studies via quantitative PCR in hybrid roses (Rosa hybrida). BMC Res. Notes 4:518. doi: 10.1186/1756-0500-4-518
Kong, Q., Gao, L., Cao, L., Liu, Y., Saba, H., Huang, Y., et al. (2016). Assessment of suitable reference genes for quantitative gene expression studies in melon fruits. Front. Plant Sci. 7:1178. doi: 10.3389/fpls.2016.01178
Kong, Q., Yuan, J., Gao, L., Zhao, S., Jiang, W., Huang, Y., et al. (2014a). Identification of suitable reference genes for gene expression normalization in qrt-pcr analysis in watermelon. PLoS ONE 9:e90612. doi: 10.1371/journal.pone.0090612
Kong, Q., Yuan, J., Niu, P., Xie, J., Jiang, W., Huang, Y., et al. (2014b). Screening suitable reference genes for normalization in reverse transcription quantitative real-time PCR analysis in melon. PLoS ONE 9:e87197. doi: 10.1371/journal.pone.0087197
Kull, F., Ohlson, E., and Haeggström, J. Z. (1999). Cloning and characterization of a bifunctional leukotriene A4 hydrolase from Saccharomyces cerevisiae. J. Biol. Chem. 274, 34683–34690. doi: 10.1074/jbc.274.49.34683
Li, C., Yang, X., Zhang, Z., Zhou, D., Zhang, L., Zhang, S., et al. (2013). Hydrothermal liquefaction of desert shrub Salix psammophila to high value-added chemicals and hydrochar with recycled processing water. BioResources 8, 2981–2997. doi: 10.15376/biores.8.2.2981-2997
Li, J., Han, J. G., Hu, Y. H., and Yang, J. (2016). Selection of reference Genes for quantitative real-time PCR during flower development in Tree Peony (Paeonia suffruticosa Andr.). Front. Plant Sci. 7:516. doi: 10.3389/fpls.2016.00516
Li, W., Qian, Y. Q., Han, L., Liu, J. X., and Sun, Z. Y. (2014). Identification of suitable reference genes in buffalo grass for accurate transcript normalization under various abiotic stress conditions. Gene 547, 55–62. doi: 10.1016/j.gene.2014.06.015
Liu, B., Wang, L., Zhang, J., Li, J. B., Zheng, H. Q., Chen, J., et al. (2014). WUSCHEL-related Homeobox genes in Populus tomentosa: diversified expression patterns and a functional similarity in adventitious root formation. BMC Genomics 15:296. doi: 10.1186/1471-2164-15-296
Liu, M., Qiao, G. R., Jiang, J., Han, X. J., Sang, J., and Zhuo, R. Y. (2014). Identification and expression analysis of salt-responsive genes using a comparative microarray approach in Salix matsudana. Mol. Biol. Rep. 41, 6555–6568. doi: 10.1007/s11033-014-3539-1
Liu, X. R., Pan, T., Liang, W. Q., Gao, L., Wang, X. J., Li, H. Q., et al. (2016). Overexpression of an orchid (Dendrobium nobile) SOC1/TM3-Like ortholog, DnAGL19, in Arabidopsis regulates HOS1-FT expression. Front. Plant Sci. 7:99. doi: 10.3389/fpls.2016.00099
Ma, R., Xu, S., Zhao, Y. C., Xia, B., and Wang, R. (2016). Selection and validation of appropriate reference genes for quantitative real-time PCR analysis of gene expression in Lycoris aurea. Front. Plant Sci. 7:536. doi: 10.3389/fpls.2016.00536
Mallona, I., Lischewski, S., Weiss, J., Hause, B., and Egea-Cortines, M. (2010). Validation of reference genes for quantitative real-time PCR during leaf and flower development in Petunia hybrida. BMC Plant Biol. 10:4. doi: 10.1186/1471-2229-10-4
Marshansky, V., and Futai, M. (2008). The V-type H+-ATPase in vesicular trafficking: targeting, regulation and function. Curr. Opin. Cell Biol. 20, 415–426. doi: 10.1016/j.ceb.2008.03.015
Mousavi, S. A., Chauvin, A., Pascaud, F., Kellenberger, S., and Farmer, E. E. (2013). GLUTAMATE RECEPTOR-LIKE genes mediate leaf-to-leaf wound signalling. Nature 500, 422–426. doi: 10.1038/nature12478
Najafpanah, M. J., Sadeghi, M., and Bakhtiarizadeh, M. R. (2013). Reference genes selection for quantitative real-time PCR using RankAggreg method in different tissues of Capra hircus. PLoS ONE 8:e83041. doi: 10.1371/journal.pone.0083041
Nakata, M., Matsumoto, N., Tsugeki, R., Rikirsch, E., Laux, T., and Okada, K. (2012). Roles of the middle domain–specific WUSCHEL-RELATED HOMEOBOX genes in early development of leaves in Arabidopsis. Plant Cell 24, 519–535. doi: 10.1105/tpc.111.092858
Nicot, N., Hausman, J. F., Hoffmann, L., and Evers, D. (2005). Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J. Exp. Bot. 56, 2907–2914. doi: 10.1093/jxb/eri285
Nolan, T., Hands, R. E., and Bustin, S. A. (2006). Quantification of mRNA using real-time RT-PCR. Nat. Protoc. 1, 1559–1582. doi: 10.1038/nprot.2006.236
Oliveira, L. A., Breton, M. C., Bastolla, F. M., Camargo, S., Margis, R., Frazzon, J., et al. (2012). Reference genes for the normalization of gene expression in Eucalyptus species. Plant Cell Physiol. 53, 405–422. doi: 10.1093/pcp/pcr187
Pfaffl, M. W., Tichopad, A., Prgomet, C., and Neuvians, T. P. (2004). Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515. doi: 10.1023/B:BILE.0000019559.84305.47
Pihur, V., Datta, S., and Datta, S. (2009). RankAggreg, an R package for weighted rank aggregation. BMC Bioinformatics 10:62. doi: 10.1186/1471-2105-10-62
Radonić, A., Thulke, S., Mackay, I. M., Landt, O., Siegert, W., and Nitsche, A. (2004). Guideline to reference gene selection for quantitative real-time PCR. Biochem. Biophys. Res 313, 856–862. doi: 10.1016/j.bbrc.2003.11.177
Reddy, P. S., Reddy, D. S., Sivasakthi, K., Bhatnagar-Mathur, P., Vadez, V., and Sharma, K. K. (2016). Evaluation of sorghum [Sorghum bicolor (L.)] reference genes in various tissues and under abiotic stress conditions for quantitative real-time PCR data normalization. Front. Plant Sci. 7:529. doi: 10.3389/fpls.2016.00529
Remans, T., Smeets, K., Opdenakker, K., Mathijsen, D., Vangronsveld, J., and Cuypers, A. (2008). Normalisation of real-time RT-PCR gene expression measurements in Arabidopsis thaliana exposed to increased metal concentrations. Planta 227, 1343–1349. doi: 10.1007/s00425-008-0706-4
Roxas, V. P., Smith, R. K., Allen, E. R., and Allen, R. D. (1997). Overexpression of glutathione S-transferase/glutathioneperoxidase enhances the growth of transgenic tobacco seedlings during stress. Nat. Biotechnol. 15, 988–991. doi: 10.1038/nbt1097-988
Saint-Marcoux, D., Proust, H., Dolan, L., and Langdale, J. A. (2015). Identification of reference genes for real-time quantitative PCR experiments in the liverwort Marchantia polymorpha. PLoS ONE 10:e0118678. doi: 10.1371/journal.pone.0118678
Sakuraba, Y., Jeong, J., Kang, M. Y., Kim, J., Paek, N. C., and Choi, G. (2014). Phytochrome-interacting transcription factors PIF4 and PIF5 induce leaf senescence in Arabidopsis. Nat. Commun. 5:4636. doi: 10.1038/ncomms5636
Schmittgen, T. D., and Livak, K. J. (2008). Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 3, 1101–1108. doi: 10.1038/nprot.2008.73
Schmittgen, T. D., and Zakrajsek, B. A. (2000). Effect of experimental treatment on housekeeping gene expression: validation by real-time, quantitative RT-PCR. J. Biochem. Bioph. Methods. 46, 69–81. doi: 10.1016/S0165-022X(00)00129-9
Shivhare, R., and Lata, C. (2016). Selection of suitable reference genes for assessing gene expression in pearl millet under different abiotic stresses and their combinations. Sci. Rep. 6:23036. doi: 10.1038/srep23036
Song, X. X., Fang, J., Han, X. J., He, X. L., Liu, M. Y., Hu, J., et al. (2015). Overexpression of quinone reductase from Salix matsudana Koidz enhances salt tolerance in transgenic Arabidopsis thaliana. Gene 576, 520–527. doi: 10.1016/j.gene.2015.10.069
Thunnissen, M. M. G. M., Nordlund, P., and Haeggström, J. Z. (2001). Crystal structure of human leukotriene A4 hydrolase, a bifunctional enzyme in inflammation. Nat. Struct. Biol. 8, 131–135. doi: 10.1038/84117
Tian, C., Jiang, Q., Wang, F., Wang, G. L., Xu, Z. S., and Xiong, A. S. (2015). Selection of suitable reference genes for qPCR normalization under abiotic stresses and hormone stimuli in carrot leaves. PLoS ONE 10:e0117569. doi: 10.1371/journal.pone.0117569
Tian, H., Guo, H., Dai, X., Cheng, Y., Zheng, K., Wang, X., et al. (2015). An ABA down-regulated bHLH transcription repressor gene, bHLH129 regulates root elongation and ABA response when overexpressed in Arabidopsis. Sci. Rep. 5:17587. doi: 10.1038/srep17587
Udvardi, M. K., Czechowski, T., and Scheible, W. R. (2008). Eleven golden rules of quantitative RT-PCR. Plant Cell 20, 1736–1737. doi: 10.1105/tpc.108.061143
Vandesompele, J., De Preter, K., Pattyn, F., Poppe, B., Van Roy, N., De Paepe, A., et al. (2002). Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, 1–12. doi: 10.1186/gb-2002-3-7-research0034
Wang, H. L., Chen, J. H., Tian, Q. Q., Wang, S., Xia, X. L., and Yin, W. L. (2014). Identification and validation of reference genes for Populus euphratica gene expression analysis during abiotic stresses by quantitative real-time PCR. Physiol. Plant 152, 529–545. doi: 10.1111/ppl.12206
Waters, S., Gilliham, M., and Hrmova, M. (2013). Plant high-affinity potassium (HKT) transporters involved in salinity tolerance: structural insights to probe differences in ion selectivity. Int. J. Mol. Sci. 14, 7660–7680. doi: 10.3390/ijms14047660
Wu, Z. J., Tian, C., Jiang, Q., Li, X. H., and Zhuang, J. (2016). Selection of suitable reference genes for qRT-PCR normalization during leaf development and hormonal stimuli in tea plant (Camellia sinensis). Sci. Rep. 6:19748. doi: 10.1038/srep19748
Xu, M., Xie, W., and Huang, M. (2015). Two WUSCHEL-related HOMEOBOX genes, PeWOX11a and PeWOX11b, are involved in adventitious root formation of poplar. Physiol. Plant 4, 446–456. doi: 10.1111/ppl.12349
Xu, X., Liu, X. M., Chen, S. L., Li, B. H., Wang, X. Y., Fan, C. Q., et al. (2016). Selection of relatively exact reference genes for gene expression studies in flixweed (Descurainia sophia) by quantitative real-time polymerase chain reaction. Pestic. Biochem. Physiol. 127, 59–66. doi: 10.1016/j.pestbp.2015.09.007
Yang, J. L., Chen, Z., Wu, S. Q., Cui, Y., Zhang, L., Dong, H., et al. (2015). Overexpression of the Tamarix hispida ThMT3 gene increases copper tolerance and adventitious root induction in Salix matsudana Koidz. Plant Cell Tissue Organ. Cult. 121, 469–479. doi: 10.1007/s11240-015-0717-3
Yang, X., Lyu, H., Chen, K., Zhu, X., Zhang, S., and Chen, J. (2014). Selective extraction of bio-oil from hydrothermal liquefaction of Salix psammophila by organic solvents with different polarities through multistep extraction separation. BioResources 9, 5219–5233. doi: 10.15376/biores.9.3.5219-5233
Yim, A. K. Y., Wong, J. W. H., Ku, Y. S., Qin, H., Chan, T. F., and Lam, H. M. (2015). Using RNA-Seq data to evaluate reference genes suitable for gene expression studies in Soybean. PLoS ONE 10:e0136343. doi: 10.1371/journal.pone.0136343
Keywords: Salix psammophila, reference genes, quantitative real-time PCR, tissue types, abiotic stresses
Citation: Li J, Jia H, Han X, Zhang J, Sun P, Lu M and Hu J (2016) Selection of Reliable Reference Genes for Gene Expression Analysis under Abiotic Stresses in the Desert Biomass Willow, Salix psammophila. Front. Plant Sci. 7:1505. doi: 10.3389/fpls.2016.01505
Received: 05 August 2016; Accepted: 22 September 2016;
Published: 05 October 2016.
Edited by:
Roger Deal, Emory University, USAReviewed by:
M. Teresa Sanchez-Ballesta, Spanish National Research Council, SpainQiusheng Kong, Huazhong Agricultural University, China
Copyright © 2016 Li, Jia, Han, Zhang, Sun, Lu and Hu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jianjun Hu, aHVqakBjYWYuYWMuY24=
†These authors have contributed equally to this work.
 Jianbo Li
Jianbo Li Huixia Jia
Huixia Jia Xiaojiao Han1
Xiaojiao Han1 Jianjun Hu
Jianjun Hu