- 1Department of Botany, University of British Columbia, Vancouver, BC, Canada
- 2The Michael Smith Laboratories, University of British Columbia, Vancouver, BC, Canada
Plants lack the adaptive immune system possessed by mammals. Instead they rely on innate immunity to defend against pathogen attacks. Genomes of higher plants encode a large number of plant immune receptors belonging to different protein families, which are involved in the detection of pathogens and activation of downstream defense pathways. Plant immunity is tightly controlled to avoid activation of defense responses in the absence of pathogens, as failure to do so can lead to autoimmunity that compromises plant growth and development. Many autoimmune mutants have been reported, most of which are associated with dwarfism and often spontaneous cell death. In this review, we summarize previously reported Arabidopsis autoimmune mutants, categorizing them based on their functional groups. We also discuss how their obvious morphological phenotypes make them ideal tools for epistatic analysis and suppressor screens, and summarize genetic screens that have been carried out in various autoimmune mutant backgrounds.
Introduction
The ability to detect and respond to pathogens is essential for the survival of multicellular organisms. Plants have developed sophisticated immune systems to combat microbial pathogens. Recognition of Pathogen-Associated Molecular Patterns (PAMPs), components important for pathogen lifestyles such as bacterial flagellin, by Pattern Recognition Receptors (PRRs) leads to activation of PAMP-triggered immunity (PTI) (Boller and Felix, 2009). Most PRRs belong to the transmembrane receptor-like kinase (RLK) or receptor-like protein (RLP) families (Monaghan and Zipfel, 2012). Unlike RLKs, which contain extracellular, transmembrane, and intracellular kinase domains, RLPs lack the intracellular kinase domain.
In response to pathogens that deliver effector proteins to promote virulence by avoiding detection by the host cells or inhibiting PTI, plants have evolved resistance (R) proteins, which detect effectors and trigger an immune response termed effector triggered immunity (ETI). R proteins recognize effectors either by directly binding to the effectors or indirectly sensing modifications to host proteins caused by the effectors (Jones and Dangl, 2006). Most R proteins contain a central nucleotide-binding domain (NB) and C-terminal leucine-rich repeats (LRRs), with either a Toll-interleukin 1-like receptor (TIR) or a coiled-coil (CC) domain at their N termini.
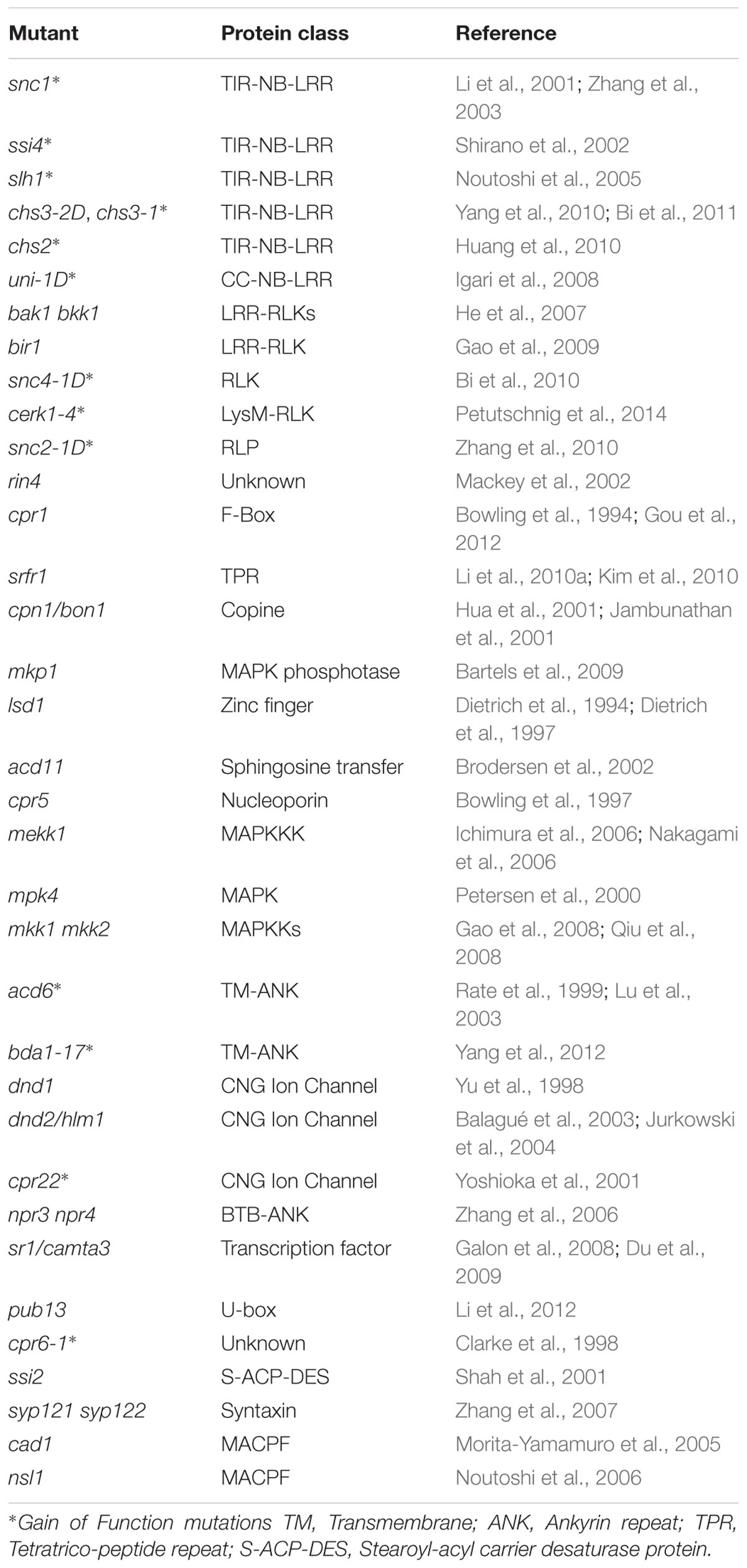
Plant immunity is under tight negative control to avoid activation in the absence of pathogens. Immune receptors are generally maintained at inactive states and activated only upon detection of pathogens. Loss-of-function mutations in negative regulators and gain-of-function mutations in plant immune receptors often lead to autoimmunity. The typical phenotypes of autoimmune mutants include dwarfism, elevated salicylic acid levels, constitutive expression of defense genes and enhanced disease resistance to pathogens, and in some cases also spontaneous lesion formation. They have been instrumental in assisting our studies of plant immunity. Previous reviews have highlighted lesion mimic mutants (Bruggeman et al., 2015; Rodriguez et al., 2016). Here we are focusing on Arabidopsis autoimmune mutants (Table 1) and their application in studying plant immunity.
Autoimmunity Caused By Gain-Of-Function Mutations in NB-LRR Proteins
Autoimmunity in a number of Arabidopsis mutants is caused by mutations in TIR-NB-LRR proteins. Among them, suppressor of npr1-1, constitutive (snc) 1 is one of the most well studied (Li et al., 2001). A single amino acid change in the linker region between the NB and LRR domains leads to over-accumulation of the SNC1 protein and activation of defense responses (Zhang et al., 2003). snc1 exhibits typical autoimmune phenotypes, but does not have spontaneous cell death. Both SA-dependent and SA-independent defense pathways contribute to the enhanced pathogen resistance in snc1. Unlike snc1, suppressor of salicylic acid insensitive (ssi) 4 is another gain-of-function mutant of a TIR-NB-LRR protein that displays both dwarfism and spontaneous cell death (Shirano et al., 2002).
The autoimmune phenotypes in sensitive to low humidity (slh)1 (an allele of RRS1) and chilling sensitive (chs) 3-2D are caused by mutations in atypical TIR-NB-LRR proteins (Noutoshi et al., 2005; Bi et al., 2011). RESISTANT TO RALSTONIA SOLANACEARUM 1 (RRS1) and CHS3 contain extra C-terminal domains proposed to function as integrated decoys for pathogen effectors. RRS1 has a WRKY DNA-binding domain at its C-terminus. In slh1, a single amino acid insertion in the DNA-binding domain causes activation of defense responses (Noutoshi et al., 2005). CHS3 contains a LIM domain at its C-terminus. A missense mutation close to the LIM domain in chs3-2D leads to extreme dwarfism and activation of defense responses (Bi et al., 2011). It is likely that mutations in slh1 and chs3-2D are sensed by the TIR-NB-LRR part of the protein, which triggers immune activation.
A second gain-of-function allele of CHS3, chs3-1, gives rise to a truncated CHS3 without part of the LIM domain (Yang et al., 2010). The phenotype of chs3-1 is temperature dependent and manifested when grown at 16°C or lower. Another chilling sensitive mutant, chs2-1, also contains a gain-of-function mutation in a TIR-NB-LRR protein (Huang et al., 2010). A single amino acid substitution in the NB domain of RECOGNITION OF PERONOSPORA PARASITICA 4 (RPP4) causes constitutive defense activation when the mutant is grown at 16°C or lower. CHS1 encodes a truncated TIR-NB protein. A missense mutation in CHS1 results in activation of cell death and defense responses at low temperature (Wang et al., 2013; Zbierzak et al., 2013).
uni-1D is the only known mutant of a CC-NB-LRR protein that causes autoimmunity (Igari et al., 2008). It carries three amino acid substitutions in the LRR domain. In addition to constitutive defense activation, uni-1D has a variety of developmental defects associated with increased cytokinin accumulation. It is unclear how activation of UNI leads to modifications in the cytokinin pathway.
Autoimmunity Caused By Mutations in RLKS/RLPS
Arabidopsis Brassinosteroid Insensitive 1-associated receptor kinase 1 (BAK1) is a critical component of PTI which functions as co-receptor of multiple PRRs (Liebrand et al., 2014). Interestingly, knocking out both BAK1 and its close homolog of BAK1-like 1 (BKK1) leads to strong autoimmune phenotypes (He et al., 2007). Defense responses are not constitutively activated in the bak1 and bkk1 single mutants, but uncontrolled spreading of necrosis occurs in bak1 knockout mutant plants upon infection by necrotrophic pathogens (Kemmerling et al., 2007). The mechanism of how loss-of-function of BAK1 and BKK1 activates plant immunity remains to be determined.
Several members of the BAK1-interacting RLK (BIR) subfamily have been shown to associate with BAK1 in vivo and serve as regulators of immunity (Gao et al., 2009; Halter et al., 2014). The bir1-1 knockout mutant displayed extreme dwarfism, spontaneous cell death and constitutive defense responses (Gao et al., 2009). Loss of BIR2 function leads to enhanced PTI responses, but does not cause autoimmunity (Halter et al., 2014).
Gain-of-function mutations in two RLKs were shown to cause autoimmunity. In snc4-1D, a single amino acid substitution in the kinase domain of SNC4 leads to extreme dwarfism and constitutive defense responses (Bi et al., 2010). CHITIN ELICITOR RECEPTOR KINASE (CERK) 1 is an RLK involved in perception of chitin and bacterial peptidoglycan (Miya et al., 2007; Wan et al., 2008). In cerk1-4, a single amino acid change in the CERK1 ectodomain results in deregulated cell death upon pathogen challenge (Petutschnig et al., 2014).
snc2-1D is the only reported gain-of-function RLP mutant which causes autoimmunity in Arabidopsis (Zhang et al., 2010). Loss of function of SNC2 results in enhanced susceptibility to the non-pathogenic bacterium P.s.t. DC3000 hrcC, suggesting that SNC2 plays an important role in PTI. The alteration of a G to R in a conserved motif of the transmembrane helix in snc2-1D may disrupt the interaction between SNC2 and its negative regulator.
Autoimmunity Caused By Mutations in Other Types of Protein
Mutations in a large number of genes encoding proteins not in the NB-LRR or RLK/RLP families were also found to result in autoimmunity. In some cases, the autoimmune phenotypes are caused by loss-of-function of negative regulators or gain-of-function of positive regulators of plant immunity. But most often, they are due to activation of immunity mediated by specific NB-LRR immune receptors, probably as targets of pathogen effectors that are monitored or guarded by Resistance proteins. One well-studied example is RPM1 INTERACTING PROTEIN (RIN) 4, which is guarded by two NB-LRR proteins RESISTANT TO P. SYRINGAE (RPS) 2 and RESISTANCE TO P. SYRINGAE PV MACULICOLA (RPM) 1 (Axtell and Staskawicz, 2003; Mackey et al., 2003). Loss of RIN4 triggers defense activation mediated by RPS2 and RPM1 (Belkhadir et al., 2004).
Loss-of-function mutations in several genes cause activation of SNC1-mediated immune responses. CONSTITUTIVE EXPRESSER OF PR GENES (CPR) 1 encodes an F-box protein that regulates the turnover of SNC1 (Cheng et al., 2011; Gou et al., 2012). Mutations in CPR1 cause elevated SNC1 and RPS2 protein levels and activation of defense responses. SUPPRESSOR OF RPS4-RLD (SRFR) 1 is also involved in regulating SNC1 protein levels. Mutations in SRFR1 result in increased SNC1 level and activation of SNC1-mediated immunity (Kim et al., 2010; Li et al., 2010a). Knockout mutants of BONZAI (BON) 1 and MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE (MKP) 1 also exhibit autoimmune phenotypes that are dependent on SNC1 (Yang and Hua, 2004; Bartels et al., 2009). How loss of BON1 and MKP1 leads to activation of SNC1-mediated immunity remains to be determined.
The autoimmune phenotypes of two lesion-mimic mutants, lesion simulating disease (lsd) 1 and accelerated cell death (acd) 11, are also caused by activation of NB-LRR protein mediated immunity. The spontaneous cell death phenotype in lsd1 is dependent on ACTIVATED DISEASE RESISTANCE (ADR) 1, ADR1-L1, and ADR1-L2, which function as “helper NB-LRRs” that assist other NB-LRR proteins to transduce defense signals (Bonardi et al., 2011). LAZARUS (LAZ) 5, a TIR-NB-LRR protein, is required for the cell death and constitutive defense phenotype in acd11 (Palma et al., 2010). The constitutive defense response in another lesion-mimic mutant, cpr5, was also shown to associate with activation of NB-LRR protein mediated immunity (Wang et al., 2014).
Arabidopsis MAPK/ERK KINASE KINASE (MEKK) 1, MAP KINASE KINASE (MKK) 1/MKK2, and MAP KINASE (MPK) 4 form a MAPK cascade that plays critical roles in plant immunity (Gao et al., 2008; Qiu et al., 2008). Loss-of-function of MEKK1, MKK1/MKK2, or MPK4 leads to activation of defense responses (Ichimura et al., 2006; Nakagami et al., 2006; Suarez-Rodriguez et al., 2007). The autoimmune phenotypes of mekk1, mkk1 mkk2, and mpk4 mutants are dependent on the CC-NB-LRR protein SUMM2, suggesting that the MEKK1-MKK1/MKK2-MPK4 cascade is monitored by SUMM2, which can sense its disruption by pathogen effectors (Zhang et al., 2012).
ACD6 and BIAN DA (BDA) 1 are two structurally related transmembrane proteins with an N-terminal ankyrin-repeat domain (Lu et al., 2003; Yang et al., 2012). acd6-1 and bda1-17D are both gain-of-function mutations located in the transmembrane regions, which cause spontaneous cell death and constitutive defense responses. ACD6 positively regulates the abundance of several PRRs (Tateda et al., 2015). It likely functions as a critical regulator of PTI. BDA1 is required for the autoimmunity of snc2-1D, suggesting that it is a positive regulator of SNC2-mediated defense responses (Yang et al., 2012).
Mutations in multiple genes in the cyclic nucleotide-gated ion channel (CNGC) family have been shown to cause autoimmune phenotypes, highlighting the potential importance of calcium channels in plant immune regulation. Loss-of-function mutations in DEFENSE NO DEATH (DND) 1 (CNGC2) and DND2 (CNGC4) results in constitutive defense responses in the absence of cell death (Clough et al., 2000; Balagué et al., 2003; Jurkowski et al., 2004), suggesting that they play critical roles in negative regulation of plant defense. The spontaneous cell death and constitutive defense responses in the semi-dominant mutant cpr22 are caused by a novel chimeric protein generated by a deletion that fused CNGC11 and CNGC12 (Yoshioka et al., 2006). Knockout analysis showed that both CNGC11 and CNGC12 function in positive regulation of pathogen resistance (Yoshioka et al., 2006).
Mutations in several other negative regulators of plant immunity were shown to cause activation of defense responses. Double knockout mutants of NON-EXPRESSER OF PR GENES (NPR) 3 and NPR4 exhibit constitutive defense gene expression and enhanced resistance to pathogens (Zhang et al., 2006). T-DNA insertion mutants of CALMODULIN-BINDING TRANSCRIPTION ACTIVATOR (CAMTA) 3 displayed enhanced disease resistance and dwarfism when grown at low temperatures (Du et al., 2009), whereas a gain-of-function mutation in CAMTA3 leads to compromised immune responses (Jing et al., 2011; Nie et al., 2012). PLANT U-BOX (PUB) 13 is a E3 ubiquitin ligase involved in degradation of FLAGELLIN-SENSITIVE2 and the ABA co-receptor ABI1 (Lu et al., 2011; Kong et al., 2015). Knockout mutant plants of pub13 exhibit spontaneous cell death and enhanced pathogen resistance (Li et al., 2012).
In some autoimmune mutants, the mechanism of how defense responses are constitutively activated is unclear. cpr6-1 is a semi-dominant mutant with constitutive defense responses due to a mutation of unknown nature (Clarke et al., 1998). Loss-of-function of the stearoyl-ACP desaturase SSI2 activates NPR1-independent defense responses (Shah et al., 2001; Kachroo A. et al., 2003). The link between altered fatty acid metabolism and activation of immunity remains to be determined. Double knockout mutant plants of SYNTAXIN OF PLANTS (SYP) 121 and SYP122 display a lesion-mimic phenotype and enhanced pathogen resistance (Zhang et al., 2007). Loss-of-function mutations in two membrane attack complex/perforin proteins, CONSTITUTIVELY ACTIVATED CELL DEATH (CAD) 1 and NECROTIC SPOTTED LESIONS (NSL) 1, result in spontaneous cell death and constitutive defense response (Morita-Yamamuro et al., 2005; Noutoshi et al., 2006). The exact roles of CAD1 and NSL1 in immunity are unclear.
Applications of Autoimmune Mutants in Genetic Analysis of Plant Immunity
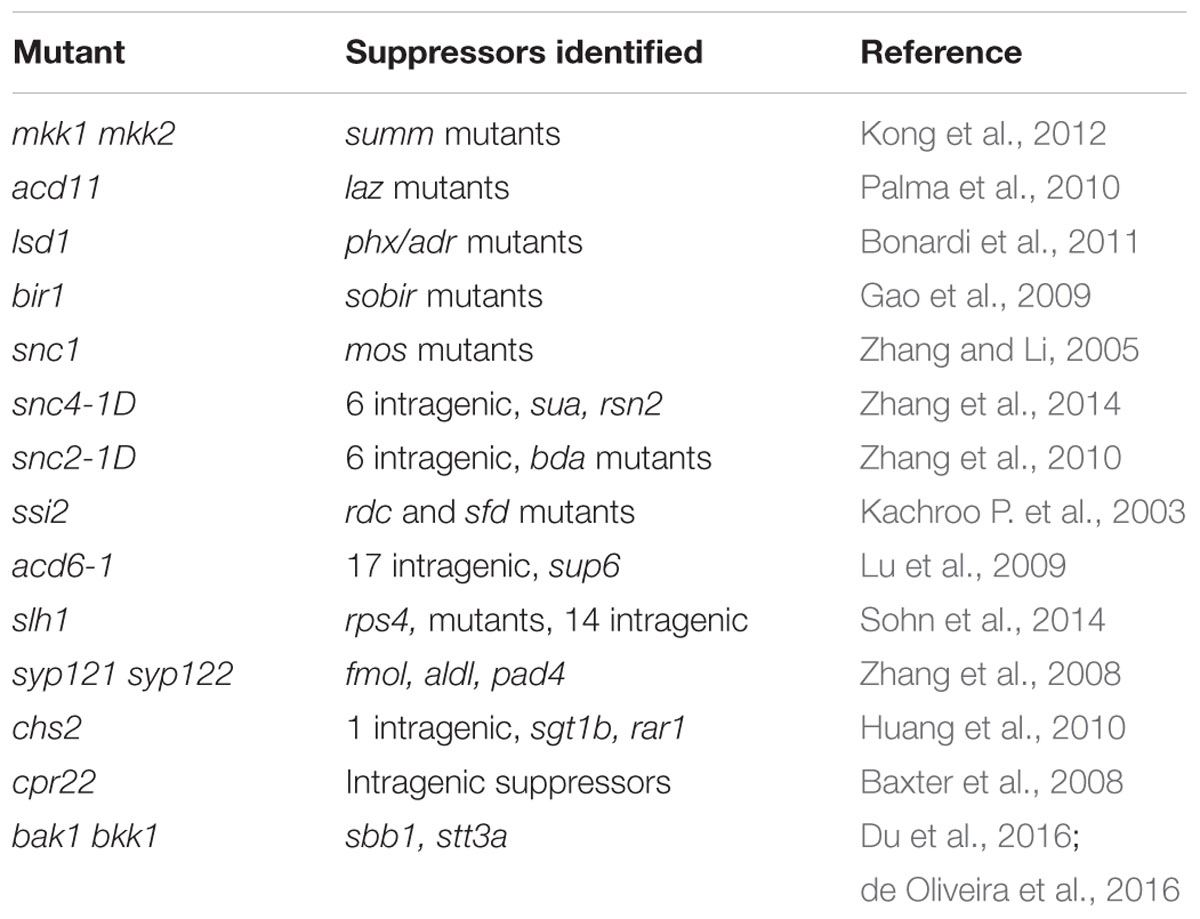
One obvious feature of autoimmune mutants is the distinct morphological phenotypes such as dwarfism and spontaneous cell death. They are often used in epistasis analysis to test genetic interactions. Because large populations of plants can be screened using these morphological phenotypes as readout, a number of suppressor mutant screens have been carried out on various autoimmune mutants (Table 2). Mutants obtained from a suppressor screen can be either intragenic or extragenic suppressors. Intragenic suppressor mutations can be useful for structure-function analysis, whereas extragenic suppressors are valuable for identifying components required for the autoimmune phenotypes in the original mutant.
One very useful feature of autoimmune mutants is that the dwarf phenotype is often dependent on environmental conditions such as temperature and humidity. For example, the dwarf phenotype of bon1 and snc4-1D can be fully suppressed and the dwarf phenotype of snc2-1D, mkk1 mkk2, and bir1-1 can be partially suppressed when grown at 28°C (Hua et al., 2001; Gao et al., 2009; Bi et al., 2010; Zhang et al., 2010; Zhang et al., 2012). The dwarf phenotype of several other mutants such as ssi4, cpr22, and slh1 can be suppressed by high humidity (Yoshioka et al., 2001; Zhou et al., 2004; Noutoshi et al., 2005). Temperature and humidity sensitivity are frequently observed in the same mutants, suggesting a common element of these phenotypes, although the molecular mechanisms behind these conditional phenotypes are not clear yet. For mutants with extreme dwarf morphology, seeds of a mutagenized mutant population can be obtained by growing plants at high temperature or high humidity. Mutant screens can be subsequently carried out in the next generation at regular growth conditions. The conditional features of the autoimmune mutants allow screening of large mutagenized populations, enabling systematic discovery.
Analysis of suppressors of autoimmune mutants has been very effective for investigating the molecular basis of the respective constitutive defense responses. For example, studies on suppressor of mkk1 mkk (summ) 2 revealed that the autoimmune phenotypes in mekk1, mkk1 mkk2, and mpk4 are due to activation of immune responses mediated by the CC-NB-LRR protein SUMM2 (Zhang et al., 2012). Characterization of suppressor mutants of acd11 revealed that loss of ACD11 results in activation of immune responses mediated by the TIR-NB-LRR protein LAZ5 (Palma et al., 2010). Analysis of suppressor mutants of lsd1 revealed that ADR1, ADR1-L1, and ADR1-L2 are required for the lesion mimic phenotype of lsd1 (Bonardi et al., 2011), suggesting that NB-LRR-mediated defense responses are activated in the mutant. Studies on suppressors of bir1-1 revealed that loss of BIR1 leads to activation of immune responses mediated by RLKs BAK1 and SUPPRESSOR OF BIR1 (SOBIR) 1 (Gao et al., 2009; Liu et al., 2016).
Suppressors of autoimmune mutants have been very useful for studying the biogenesis of plant immune receptors and dissecting signaling pathways downstream of plant immune receptors. For example, analysis of suppressors of snc1 revealed that SNC1 is regulated at both transcriptional and post-transcriptional levels and highlights the importance of nucleocytoplasmic trafficking and RNA processing in plant immunity (Palma et al., 2005, 2007; Zhang and Li, 2005; Zhang et al., 2005; Cheng et al., 2009; Germain et al., 2010; Li et al., 2010b; Xu et al., 2011, 2012; Xia et al., 2013). The identification of MODIFIER OF SNC1 (MOS) 5 suggests that protein ubiquitination plays an important role in immunity mediated by NB-LRR proteins (Goritschnig et al., 2007). From the snc1 suppressor screen, the transcriptional repressor TOPLESS RELATED 1 was identified as a critical plant immune regulator that suppresses the expression of negative regulators of immunity (Zhu et al., 2010). Identification of set domain group 8 (sdg8) as a suppressor of acd11 revealed that the expression of LAZ5 is regulated by histone modifications (Palma et al., 2010). Studies on snc4-1D suppressor mutants revealed critical roles of alternative splicing in immunity mediated by the RLKs SNC4 and CERK1 (Zhang et al., 2014).
Analysis of suppressor mutants of snc2-1D identified BDA1 as a critical regulator of PTI (Yang et al., 2012). It also identified WRKY DNA-BINDING PROTEIN (WRKY) 70 as a critical regulator of the SA-independent defense pathway (Zhang et al., 2010). The identification of gtp binding protein beta (agb) 1 as a suppressor of bir1-1 and subsequent studies revealed that heterotrimeric G-proteins serve as a converging point for immunity activated by multiple RLKs (Liu et al., 2013).
In addition to suppressor screens, autoimmune mutants can also be used for enhancer screens. For example, a genetic screen looking for enhancers of snc1 identified many muse (mutants, snc1-enhancing) mutants (Huang et al., 2013). Analysis of the muse mutants lead to the identification of several components involved in proteasome-mediated degradation of SNC1 (Huang S. et al., 2014; Huang Y. et al., 2014; Huang et al., 2016). It also revealed a crucial role of N-terminal acetylation on the turnover of SNC1 (Xu et al., 2015).
Concluding Remarks
Mutant analyses have been instrumental in the mechanistic studies of plant immunity. A wide range of autoimmune mutants has been identified in Arabidopsis. In some of these mutants, the mechanism of defense activation is unclear. Interpretation of data obtained from them is often difficult, especially when the mutations occur in proteins involved in general biological processes. However, the autoimmune phenotypes of many mutants are caused by activation of defense responses mediated by specific plant immune receptors. Their distinct phenotypes facilitate easy and fast identification of suppressor or enhancer mutants, enabling new discoveries and allowing researchers to study signaling mechanisms in plant immunity.
Author Contributions
All authors listed, have made substantial, direct and intellectual contribution to the work, and approved it for publication.
Funding
This work is funded by Natural Sciences and Engineering Research Council of Canada.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors apologize to researchers whose work could not be included in the review due to space constraints. Works in the Zhang and Li labs are supported by NSERC-Discovery program.
References
Axtell, M. J., and Staskawicz, B. J. (2003). Initiation of RPS2-specified disease resistance in Arabidopsis is coupled to the AvrRpt2-directed elimination of RIN4. Cell 112, 369–377. doi: 10.1016/S0092-8674(03)00036-9
Balagué, C., Lin, B., Alcon, C., Flottes, G., Malmstrom, S., Kohler, C., et al. (2003). HLM1, an essential signaling component in the hypersensitive response, is a member of the cyclic nucleotide-gated channel ion channel family. Plant Cell 15, 365–379. doi: 10.1105/tpc.006999
Bartels, S., Anderson, J. C., Gonzalez Besteiro, M. A., Carreri, A., Hirt, H., Buchala, A., et al. (2009). MAP kinase phosphatase1 and protein tyrosine phosphatase1 are repressors of salicylic acid synthesis and SNC1-mediated responses in Arabidopsis. Plant Cell 21, 2884–2897. doi: 10.1105/tpc.109.067678
Baxter, J., Moeder, W., Urquhart, W., Shahinas, D., Chin, K., Christendat, D., et al. (2008). Identification of a functionally essential amino acid for Arabidopsis cyclic nucleotide gated ion channels using the chimeric AtCNGC11/12 gene. Plant J. 56, 457–469. doi: 10.1111/j.1365-313X.2008.03619.x
Belkhadir, Y., Nimchuk, Z., Hubert, D. A., Mackey, D., and Dangl, J. L. (2004). Arabidopsis RIN4 negatively regulates disease resistance mediated by RPS2 and RPM1 downstream or independent of the NDR1 signal modulator and is not required for the virulence functions of bacterial type III effectors AvrRpt2 or AvrRpm1. Plant Cell 16, 2822–2835. doi: 10.1105/tpc.104.024117
Bi, D., Cheng, Y. T., Li, X., and Zhang, Y. (2010). Activation of plant immune responses by a gain-of-function mutation in an atypical receptor-like kinase. Plant Physiol. 153, 1771–1779. doi: 10.1104/pp.110.158501
Bi, D., Johnson, K., Huang, Y., Zhu, Z., Li, X., and Zhang, Y. (2011). Mutations in an atypical TIR-NB-LRR-LIM resistance protein confers autoimmunity. Front. Plant Sci. 2:71. doi: 10.3389/fpls.2011.00071
Boller, T., and Felix, G. (2009). A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 60, 379–406. doi: 10.1146/annurev.arplant.57.032905.105346
Bonardi, V., Tang, S., Stallmann, A., Roberts, M., Cherkis, K., and Dangl, J. L. (2011). Expanded functions for a family of plant intracellular immune receptors beyond specific recognition of pathogen effectors. Proc. Natl. Acad. Sci. U.S.A. 108, 16463–16468. doi: 10.1073/pnas.1113726108
Bowling, S. A., Clarke, J. D., Liu, Y., Klessig, D. F., and Dong, X. (1997). The cpr5 mutant of Arabidopsis expresses both NPR1-dependent and NPR1-independent resistance. Plant Cell 9, 1573–1584. doi: 10.2307/3870444
Bowling, S. A., Guo, A., Cao, H., Gordon, A. S., Klessig, D. F., and Dong, X. (1994). A mutation in Arabidopsis that leads to constitutive expression of systemic acquired resistance. Plant Cell 6, 1845–1857. doi: 10.2307/3869912
Brodersen, P., Petersen, M., Pike, H. M., Olszak, B., Skov, S., Ødum, N., et al. (2002). Knockout of Arabidopsis accelerated-cell-death11 encoding a sphingosine transfer protein causes activation of programmed cell death and defense. Genes Dev. 16, 490–502. doi: 10.1101/gad.218202
Bruggeman, Q., Raynaud, C., Benhamed, M., and Delarue, M. (2015). To die or not to die? Lessons from lesion mimic mutants. Front. Plant Sci. 6:24. doi: 10.3389/fpls.2015.00024
Cheng, Y. T., Germain, H., Wiermer, M., Bi, D., Xu, F., Garcia, A. V., et al. (2009). Nuclear pore complex component MOS7/Nup88 is required for innate immunity and nuclear accumulation of defense regulators in Arabidopsis. Plant Cell 21, 2503–2516. doi: 10.1105/tpc.108.064519
Cheng, Y. T., Li, Y., Huang, S., Huang, Y., Dong, X., Zhang, Y., et al. (2011). Stability of plant immune-receptor resistance proteins is controlled by SKP1-CULLIN1-F-box (SCF)-mediated protein degradation. Proc. Natl. Acad. Sci. U.S.A. 108, 14694–14699. doi: 10.1073/pnas.1105685108
Clarke, J. D., Liu, Y., Klessig, D. F., and Dong, X. (1998). Uncoupling PR gene expression from NPR1 and bacterial resistance: characterization of the dominant Arabidopsis cpr6-1 mutant. Plant Cell 10, 557–569. doi: 10.1105/tpc.10.4.557
Clough, S. J., Fengler, K. A., Yu, I. C., Lippok, B., Smith, R. K. Jr., and Bent, A. F. (2000). The Arabidopsis dnd1 “defense, no death” gene encodes a mutated cyclic nucleotide-gated ion channel. Proc. Natl. Acad. Sci. U.S.A. 97, 9323–9328. doi: 10.1073/pnas.150005697
de Oliveira, M. V., Xu, G., Li, B., de Souza Vespoli, L., Meng, X., Chen, X., et al. (2016). Specific control of Arabidopsis BAK1/SERK4-regulated cell death by protein glycosylation. Nat. Plants 2:15218. doi: 10.1038/nplants.2015.218
Dietrich, R. A., Delaney, T. P., Uknes, S. J., Ward, E. R., Ryals, J. A., and Dangl, J. L. (1994). Arabidopsis mutants simulating disease resistance response. Cell 77, 565–577. doi: 10.1016/0092-8674(94)90218-6
Dietrich, R. A., Richberg, M. H., Schmidt, R., Dean, C., and Dangl, J. L. (1997). A novel zinc finger protein is encoded by the Arabidopsis LSD1 gene and functions as a negative regulator of plant cell death. Cell 88, 685–694. doi: 10.1016/S0092-8674(00)81911-X
Du, J., Gao, Y., Zhan, Y., Zhang, S., Wu, Y., Xiao, Y., et al. (2016). Nucleocytoplasmic trafficking is essential for BAK1-and BKK1-mediated cell-death control. Plant J. 85, 520–531. doi: 10.1111/tpj.13125
Du, L., Ali, G. S., Simons, K. A., Hou, J., Yang, T., Reddy, A. S., et al. (2009). Ca(2+)/calmodulin regulates salicylic-acid-mediated plant immunity. Nature 457, 1154–1158. doi: 10.1038/nature07612
Galon, Y., Nave, R., Boyce, J. M., Nachmias, D., Knight, M. R., and Fromm, H. (2008). Calmodulin-binding transcription activator (CAMTA) 3 mediates biotic defense responses in Arabidopsis. FEBS Lett. 582, 943–948. doi: 10.1016/j.febslet.2008.02.037
Gao, M., Liu, J., Bi, D., Zhang, Z., Cheng, F., Chen, S., et al. (2008). MEKK1, MKK1/MKK2 and MPK4 function together in a mitogen-activated protein kinase cascade to regulate innate immunity in plants. Cell Res. 18, 1190–1198. doi: 10.1038/cr.2008.300
Gao, M., Wang, X., Wang, D., Xu, F., Ding, X., Zhang, Z., et al. (2009). Regulation of cell death and innate immunity by two receptor-like kinases in Arabidopsis. Cell Host Microbe 6, 34–44. doi: 10.1016/j.chom.2009.05.019
Germain, H., Qu, N., Cheng, Y. T., Lee, E., Huang, Y., Dong, O. X., et al. (2010). MOS11: a new component in the mRNA export pathway. PLoS Genet. 6:e1001250. doi: 10.1371/journal.pgen.1001250
Goritschnig, S., Zhang, Y., and Li, X. (2007). The ubiquitin pathway is required for innate immunity in Arabidopsis. Plant J. 49, 540–551. doi: 10.1111/j.1365-313X.2006.02978.x
Gou, M. Y., Shi, Z. Y., Zhu, Y., Bao, Z. L., Wang, G. Y., and Hua, J. (2012). The F-box protein CPR1/CPR30 negatively regulates R protein SNC1 accumulation. Plant J. 69, 411–420. doi: 10.1111/j.1365-313X.2011.04799.x
Halter, T., Imkampe, J., Mazzotta, S., Wierzba, M., Postel, S., Bucherl, C., et al. (2014). The leucine-rich repeat receptor kinase BIR2 is a negative regulator of BAK1 in plant immunity. Curr. Biol. 24, 134–143. doi: 10.1016/j.cub.2013.11.047
He, K., Gou, X., Yuan, T., Lin, H., Asami, T., Yoshida, S., et al. (2007). BAK1 and BKK1 regulate brassinosteroid-dependent growth and brassinosteroid-independent cell-death pathways. Curr. Biol. 17, 1109–1115. doi: 10.1016/j.cub.2007.05.036
Hua, J., Grisafi, P., Cheng, S. H., and Fink, G. R. (2001). Plant growth homeostasis is controlled by the Arabidopsis BON1 and BAP1 genes. Genes Dev. 15, 2263–2272. doi: 10.1101/gad.918101
Huang, S., Chen, X., Zhong, X., Li, M., Ao, K., Huang, J., et al. (2016). Plant TRAF proteins regulate NLR immune receptor turnover. Cell Host Microbe 19, 204–215. doi: 10.1016/j.chom.2016.01.005
Huang, S., Monaghan, J., Zhong, X., Lin, L., Sun, T., Dong, O. X., et al. (2014). HSP90s are required for NLR immune receptor accumulation in Arabidopsis. Plant J. 79, 427–439. doi: 10.1111/tpj.12573
Huang, Y., Minaker, S., Roth, C., Huang, S., Hieter, P., Lipka, V., et al. (2014). An E4 ligase facilitates polyubiquitination of plant immune receptor resistance proteins in Arabidopsis. Plant Cell 26, 485–496. doi: 10.1105/tpc.113.119057
Huang, X., Li, J., Bao, F., Zhang, X., and Yang, S. (2010). A gain-of-function mutation in the Arabidopsis disease resistance gene RPP4 confers sensitivity to low temperature. Plant Physiol. 154, 796–809. doi: 10.1104/pp.110.157610
Huang, Y., Chen, X., Liu, Y., Roth, C., Copeland, C., McFarlane, H. E., et al. (2013). Mitochondrial AtPAM16 is required for plant survival and the negative regulation of plant immunity. Nat. Commun. 4:2558. doi: 10.1038/ncomms3558
Ichimura, K., Casais, C., Peck, S. C., Shinozaki, K., and Shirasu, K. (2006). MEKK1 is required for MPK4 activation and regulates tissue-specific and temperature-dependent cell death in Arabidopsis. J. Biol. Chem. 281, 36969–36976. doi: 10.1074/jbc.M605319200
Igari, K., Endo, S., Hibara, K., Aida, M., Sakakibara, H., Kawasaki, T., et al. (2008). Constitutive activation of a CC-NB-LRR protein alters morphogenesis through the cytokinin pathway in Arabidopsis. Plant J. 55, 14–27. doi: 10.1111/j.1365-313X.2008.03466.x
Jambunathan, N., Siani, J. M., and McNellis, T. W. (2001). A humidity-sensitive Arabidopsis copine mutant exhibits precocious cell death and increased disease resistance. Plant Cell 13, 2225–2240. doi: 10.1105/tpc.13.10.2225
Jing, B., Xu, S., Xu, M., Li, Y., Li, S., Ding, J., et al. (2011). Brush and spray: a high-throughput systemic acquired resistance assay suitable for large-scale genetic screening. Plant Physiol. 157, 973–980. doi: 10.1104/pp.111.182089
Jones, J. D., and Dangl, J. L. (2006). The plant immune system. Nature 444, 323–329. doi: 10.1038/nature05286
Jurkowski, G. I., Smith, R. K. Jr., Yu, I. C., Ham, J. H., Sharma, S. B., Klessig, D. F., et al. (2004). Arabidopsis DND2, a second cyclic nucleotide-gated ion channel gene for which mutation causes the “defense, no death” phenotype. Mol. Plant Microbe Interact. 17, 511–520. doi: 10.1094/MPMI.2004.17.5.511
Kachroo, A., Lapchyk, L., Fukushige, H., Hildebrand, D., Klessig, D., and Kachroo, P. (2003). Plastidial fatty acid signaling modulates salicylic acid–and jasmonic acid–mediated defense pathways in the Arabidopsis ssi2 mutant. Plant Cell 15, 2952–2965. doi: 10.1105/tpc.017301
Kachroo, P., Kachroo, A., Lapchyk, L., Hildebrand, D., and Klessig, D. F. (2003). Restoration of defective cross talk in ssi2 mutants: role of salicylic acid, jasmonic acid, and fatty acids in SSI2-mediated signaling. Mol. Plant Microbe Interact. 16, 1022–1029. doi: 10.1094/MPMI.2003.16.11.1022
Kemmerling, B., Schwedt, A., Rodriguez, P., Mazzotta, S., Frank, M., Qamar, S. A., et al. (2007). The BRI1-associated kinase 1, BAK1, has a brassinolide-independent role in plant cell-death control. Curr. Biol. 17, 1116–1122. doi: 10.1016/j.cub.2007.05.046
Kim, S. H., Gao, F., Bhattacharjee, S., Adiasor, J. A., Nam, J. C., and Gassmann, W. (2010). The Arabidopsis resistance-like gene SNC1 is activated by mutations in SRFR1 and contributes to resistance to the bacterial effector AvrRps4. PLoS Pathog. 6:e1001172. doi: 10.1371/journal.ppat.1001172
Kong, L., Cheng, J., Zhu, Y., Ding, Y., Meng, J., Chen, Z., et al. (2015). Degradation of the ABA co-receptor ABI1 by PUB12/13 U-box E3 ligases. Nat. Commun. 6:8630. doi: 10.1038/ncomms9630
Kong, Q., Qu, N., Gao, M., Zhang, Z., Ding, X., Yang, F., et al. (2012). The MEKK1-MKK1/MKK2-MPK4 kinase cascade negatively regulates immunity mediated by a mitogen-activated protein kinase kinase kinase in Arabidopsis. Plant Cell 24, 2225–2236. doi: 10.1105/tpc.112.097253
Li, W., Ahn, I. P., Ning, Y., Park, C. H., Zeng, L., Whitehill, J. G., et al. (2012). The U-Box/ARM E3 ligase PUB13 regulates cell death, defense, and flowering time in Arabidopsis. Plant Physiol. 159, 239–250. doi: 10.1104/pp.111.192617
Li, X., Clarke, J. D., Zhang, Y., and Dong, X. (2001). Activation of an EDS1-mediated R-gene pathway in the snc1 mutant leads to constitutive, NPR1-independent pathogen resistance. Mol. Plant Microbe Interact. 14, 1131–1139. doi: 10.1094/MPMI.2001.14.10.1131
Li, Y., Li, S., Bi, D., Cheng, Y. T., Li, X., and Zhang, Y. (2010a). SRFR1 negatively regulates plant NB-LRR resistance protein accumulation to prevent autoimmunity. PLoS Pathog. 6:e1001111. doi: 10.1371/journal.ppat.1001111
Li, Y., Tessaro, M., Li, X., and Zhang, Y. (2010b). Regulation of the expression of plant resistance gene SNC1 by a protein with a conserved BAT2 domain. Plant Physiol. 153, 1425–1434. doi: 10.1104/pp.110.156240
Liebrand, T. W., van den Burg, H. A., and Joosten, M. H. (2014). Two for all: receptor-associated kinases SOBIR1 and BAK1. Trends Plant Sci. 19, 123–132. doi: 10.1016/j.tplants.2013.10.003
Liu, J., Ding, P., Sun, T., Nitta, Y., Dong, O., Huang, X., et al. (2013). Heterotrimeric G proteins serve as a converging point in plant defense signaling activated by multiple receptor-like kinases. Plant Physiol. 161, 2146–2158. doi: 10.1104/pp.112.212431
Liu, Y., Huang, X., Li, M., He, P., and Zhang, Y. (2016). Loss-of-function of Arabidopsis receptor-like kinase BIR1 activates cell death and defense responses mediated by BAK1 and SOBIR1. New Phytol. 212, 637–645. doi: 10.1111/nph.14072
Lu, D., Lin, W., Gao, X., Wu, S., Cheng, C., Avila, J., et al. (2011). Direct ubiquitination of pattern recognition receptor FLS2 attenuates plant innate immunity. Science 332, 1439–1442. doi: 10.1126/science.1204903
Lu, H., Rate, D. N., Song, J. T., and Greenberg, J. T. (2003). ACD6, a novel ankyrin protein, is a regulator and an effector of salicylic acid signaling in the Arabidopsis defense response. Plant Cell 15, 2408–2420. doi: 10.1105/tpc.015412
Lu, H., Salimian, S., Gamelin, E., Wang, G., Fedorowski, J., LaCourse, W., et al. (2009). Genetic analysis of acd6-1 reveals complex defense networks and leads to identification of novel defense genes in Arabidopsis. Plant J. 58, 401–412. doi: 10.1111/j.1365-313X.2009.03791.x
Mackey, D., Belkhadir, Y., Alonso, J. M., Ecker, J. R., and Dangl, J. L. (2003). Arabidopsis RIN4 is a target of the type III virulence effector AvrRpt2 and modulates RPS2-mediated resistance. Cell 112, 379–389. doi: 10.1016/S0092-8674(03)00040-0
Mackey, D., Holt, B. F., Wiig, A., and Dangl, J. L. (2002). RIN4 interacts with Pseudomonas syringae type III effector molecules and is required for RPM1-mediated resistance in Arabidopsis. Cell 108, 743–754. doi: 10.1016/S0092-8674(02)00661-X
Miya, A., Albert, P., Shinya, T., Desaki, Y., Ichimura, K., Shirasu, K., et al. (2007). CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 104, 19613–19618. doi: 10.1073/pnas.0705147104
Monaghan, J., and Zipfel, C. (2012). Plant pattern recognition receptor complexes at the plasma membrane. Curr. Opin. Plant Biol. 15, 349–357. doi: 10.1016/j.pbi.2012.05.006
Morita-Yamamuro, C., Tsutsui, T., Sato, M., Yoshioka, H., Tamaoki, M., Ogawa, D., et al. (2005). The Arabidopsis gene CAD1 controls programmed cell death in the plant immune system and encodes a protein containing a MACPF domain. Plant Cell Physiol. 46, 902–912. doi: 10.1093/pcp/pci095
Nakagami, H., Soukupova, H., Schikora, A., Zarsky, V., and Hirt, H. (2006). A Mitogen-activated protein kinase kinase kinase mediates reactive oxygen species homeostasis in Arabidopsis. J. Biol. Chem. 281, 38697–38704. doi: 10.1074/jbc.M605293200
Nie, H., Zhao, C., Wu, G., Wu, Y., Chen, Y., and Tang, D. (2012). SR1, a calmodulin-binding transcription factor, modulates plant defense and ethylene-induced senescence by directly regulating NDR1 and EIN3. Plant Physiol. 158, 1847–1859. doi: 10.1104/pp.111.192310
Noutoshi, Y., Ito, T., Seki, M., Nakashita, H., Yoshida, S., Marco, Y., et al. (2005). A single amino acid insertion in the WRKY domain of the Arabidopsis TIR-NBS-LRR-WRKY-type disease resistance protein SLH1 (sensitive to low humidity 1) causes activation of defense responses and hypersensitive cell death. Plant J. 43, 873–888. doi: 10.1111/j.1365-313X.2005.02500.x
Noutoshi, Y., Kuromori, T., Wada, T., Hirayama, T., Kamiya, A., Imura, Y., et al. (2006). Loss of necrotic spotted lesions 1 associates with cell death and defense responses in Arabidopsis thaliana. Plant Mol. Biol. 62, 29–42. doi: 10.1007/s11103-006-9001-6
Palma, K., Thorgrimsen, S., Malinovsky, F. G., Fiil, B. K., Nielsen, H. B., Brodersen, P., et al. (2010). Autoimmunity in Arabidopsis acd11 is mediated by epigenetic regulation of an immune receptor. PLoS Pathog. 6:e1001137. doi: 10.1371/journal.ppat.1001137
Palma, K., Zhang, Y., and Li, X. (2005). An importin alpha homolog, MOS6, plays an important role in plant innate immunity. Curr. Biol. 15, 1129–1135. doi: 10.1016/j.cub.2005.05.022
Palma, K., Zhao, Q., Cheng, Y. T., Bi, D., Monaghan, J., Cheng, W., et al. (2007). Regulation of plant innate immunity by three proteins in a complex conserved across the plant and animal kingdoms. Genes Dev. 21, 1484–1493. doi: 10.1101/gad.1559607
Petersen, M., Brodersen, P., Naested, H., Andreasson, E., Lindhart, U., Johansen, B., et al. (2000). Arabidopsis map kinase 4 negatively regulates systemic acquired resistance. Cell 103, 1111–1120. doi: 10.1016/S0092-8674(00)00213-0
Petutschnig, E. K., Stolze, M., Lipka, U., Kopischke, M., Horlacher, J., Valerius, O., et al. (2014). A novel Arabidopsis CHITIN ELICITOR RECEPTOR KINASE 1 (CERK1) mutant with enhanced pathogen-induced cell death and altered receptor processing. New Phytol. 204, 955–967. doi: 10.1111/nph.12920
Qiu, J. L., Zhou, L., Yun, B. W., Nielsen, H. B., Fiil, B. K., Petersen, K., et al. (2008). Arabidopsis mitogen-activated protein kinase kinases MKK1 and MKK2 have overlapping functions in defense signaling mediated by MEKK1, MPK4, and MKS1. Plant Physiol. 148, 212–222. doi: 10.1104/pp.108.120006
Rate, D. N., Cuenca, J. V., Bowman, G. R., Guttman, D. S., and Greenberg, J. T. (1999). The gain-of-function Arabidopsis acd6 mutant reveals novel regulation and function of the salicylic acid signaling pathway in controlling cell death, defenses, and cell growth. Plant Cell 11, 1695–1708. doi: 10.2307/3871047
Rodriguez, E., El Ghoul, H., Mundy, J., and Petersen, M. (2016). Making sense of plant autoimmunity and ’negative regulators’. FEBS J. 283, 1385–1391. doi: 10.1111/febs.13613
Shah, J., Kachroo, P., Nandi, A., and Klessig, D. F. (2001). A recessive mutation in the Arabidopsis SSI2 gene confers SA-and NPR1-independent expression of PR genes and resistance against bacterial and oomycete pathogens. Plant J. 25, 563–574. doi: 10.1046/j.1365-313x.2001.00992.x
Shirano, Y., Kachroo, P., Shah, J., and Klessig, D. F. (2002). A gain-of-function mutation in an Arabidopsis Toll Interleukin1 receptor-nucleotide binding site-leucine-rich repeat type R gene triggers defense responses and results in enhanced disease resistance. Plant Cell 14, 3149–3162. doi: 10.1105/tpc.005348
Sohn, K. H., Segonzac, C., Rallapalli, G., Sarris, P. F., Woo, J. Y., Williams, S. J., et al. (2014). The nuclear immune receptor RPS4 is required for RRS1 SLH1-dependent constitutive defense activation in Arabidopsis thaliana. PLoS Genet. 10:e1004655. doi: 10.1371/journal.pgen.1004655
Suarez-Rodriguez, M. C., Adams-Phillips, L., Liu, Y., Wang, H., Su, S. H., Jester, P. J., et al. (2007). MEKK1 is required for flg22-induced MPK4 activation in Arabidopsis plants. Plant Physiol. 143, 661–669. doi: 10.1104/pp.106.091389
Tateda, C., Zhang, Z., and Greenberg, J. T. (2015). Linking pattern recognition and salicylic acid responses in Arabidopsis through ACCELERATED CELL DEATH6 and receptors. Plant Signal. Behav. 10:e1010912. doi: 10.1080/15592324.2015.1010912
Wan, J., Zhang, X. C., Neece, D., Ramonell, K. M., Clough, S., Kim, S. Y., et al. (2008). A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20, 471–481. doi: 10.1105/tpc.107.056754
Wang, S., Gu, Y., Zebell, S. G., Anderson, L. K., Wang, W., Mohan, R., et al. (2014). A noncanonical role for the CKI-RB-E2F cell-cycle signaling pathway in plant effector-triggered immunity. Cell Host Microbe 16, 787–794. doi: 10.1016/j.chom.2014.10.005
Wang, Y., Zhang, Y., Wang, Z., Zhang, X., and Yang, S. (2013). A missense mutation in CHS1, a TIR-NB protein, induces chilling sensitivity in Arabidopsis. Plant J. 75, 553–565. doi: 10.1111/tpj.12232
Xia, S. T., Cheng, Y. T., Huang, S., Win, J., Soards, A., Jinn, T. L., et al. (2013). Regulation of transcription of nucleotide-binding leucine-rich repeat-encoding genes SNC1 and RPP4 via H3K4 trimethylation. Plant Physiol. 162, 1694–1705. doi: 10.1104/pp.113.214551
Xu, F., Huang, Y., Li, L., Gannon, P., Linster, E., Huber, M., et al. (2015). Two N-terminal acetyltransferases antagonistically regulate the stability of a nod-like receptor in Arabidopsis. Plant Cell 27, 1547–1562. doi: 10.1105/tpc.15.00173
Xu, F., Xu, S. H., Wiermer, M., Zhang, Y. L., and Li, X. (2012). The cyclin L homolog MOS12 and the MOS4-associated complex are required for the proper splicing of plant resistance genes. Plant J. 70, 916–928. doi: 10.1111/j.1365-313X.2012.04906.x
Xu, S., Zhang, Z., Jing, B., Gannon, P., Ding, J., Xu, F., et al. (2011). Transportin-SR is required for proper splicing of resistance genes and plant immunity. PLoS Genet. 7:e1002159. doi: 10.1371/journal.pgen.1002159
Yang, H., Shi, Y., Liu, J., Guo, L., Zhang, X., and Yang, S. (2010). A mutant CHS3 protein with TIR-NB-LRR-LIM domains modulates growth, cell death and freezing tolerance in a temperature-dependent manner in Arabidopsis. Plant J. 63, 283–296. doi: 10.1111/j.1365-313X.2010.04241.x
Yang, S., and Hua, J. (2004). A haplotype-specific resistance gene regulated by BONZAI1 mediates temperature-dependent growth control in Arabidopsis. Plant Cell 16, 1060–1071. doi: 10.1105/tpc.020479
Yang, Y., Zhang, Y., Ding, P., Johnson, K., Li, X., and Zhang, Y. (2012). The ankyrin-repeat transmembrane protein BDA1 functions downstream of the receptor-like protein SNC2 to regulate plant immunity. Plant Physiol. 159, 1857–1865. doi: 10.1104/pp.112.197152
Yoshioka, K., Kachroo, P., Tsui, F., Sharma, S. B., Shah, J., and Klessig, D. F. (2001). Environmentally sensitive, SA-dependent defense responses in the cpr22 mutant of Arabidopsis. Plant J. 26, 447–459. doi: 10.1046/j.1365-313X.2001.2641039.x
Yoshioka, K., Moeder, W., Kang, H. G., Kachroo, P., Masmoudi, K., Berkowitz, G., et al. (2006). The chimeric Arabidopsis CYCLIC NUCLEOTIDE-GATED ION CHANNEL11/12 activates multiple pathogen resistance responses. Plant Cell 18, 747–763. doi: 10.1105/tpc.105.038786
Yu, I. C., Parker, J., and Bent, A. F. (1998). Gene-for-gene disease resistance without the hypersensitive response in Arabidopsis dnd1 mutant. Proc. Natl. Acad. Sci. U.S.A. 95, 7819–7824. doi: 10.1073/pnas.95.13.7819
Zbierzak, A. M., Porfirova, S., Griebel, T., Melzer, M., Parker, J. E., and Dormann, P. (2013). A TIR-NBS protein encoded by Arabidopsis chilling sensitive 1 (CHS1) limits chloroplast damage and cell death at low temperature. Plant J. 75, 539–552. doi: 10.1111/tpj.12219
Zhang, Y., Cheng, Y. T., Bi, D., Palma, K., and Li, X. (2005). MOS2, a protein containing G-patch and KOW motifs, is essential for innate immunity in Arabidopsis thaliana. Curr. Biol. 15, 1936–1942. doi: 10.1016/j.cub.2005.09.038
Zhang, Y., Cheng, Y. T., Qu, N., Zhao, Q., Bi, D., and Li, X. (2006). Negative regulation of defense responses in Arabidopsis by two NPR1 paralogs. Plant J. 48, 647–656. doi: 10.1111/j.1365-313X.2006.02903.x
Zhang, Y., Goritschnig, S., Dong, X., and Li, X. (2003). A gain-of-function mutation in a plant disease resistance gene leads to constitutive activation of downstream signal transduction pathways in suppressor of npr1-1, constitutive 1. Plant Cell 15, 2636–2646. doi: 10.1105/tpc.015842
Zhang, Y., and Li, X. (2005). A putative nucleoporin 96 is required for both basal defense and constitutive resistance responses mediated by suppressor of npr1-1, constitutive 1. Plant Cell 17, 1306–1316. doi: 10.1105/tpc.104.029926
Zhang, Y., Yang, Y., Fang, B., Gannon, P., Ding, P., and Li, X. (2010). Arabidopsis snc2-1D activates receptor-like protein-mediated immunity transduced through WRKY70. Plant Cell 22, 3153–3163. doi: 10.1105/tpc.110.074120
Zhang, Z., Feechan, A., Pedersen, C., Newman, M. A., Qiu, J. L., Olesen, K. L., et al. (2007). A SNARE-protein has opposing functions in penetration resistance and defence signalling pathways. Plant J. 49, 302–312. doi: 10.1111/j.1365-313X.2006.02961.x
Zhang, Z., Lenk, A., Andersson, M. X., Gjetting, T., Pedersen, C., Nielsen, M. E., et al. (2008). A lesion-mimic syntaxin double mutant in Arabidopsis reveals novel complexity of pathogen defense signaling. Mol. Plant 1, 510–527. doi: 10.1093/mp/ssn011
Zhang, Z., Liu, Y., Ding, P., Li, Y., Kong, Q., and Zhang, Y. (2014). Splicing of receptor-like kinase-encoding SNC4 and CERK1 is regulated by two conserved splicing factors that are required for plant immunity. Mol. Plant 7, 1766–1775. doi: 10.1093/mp/ssu103
Zhang, Z., Wu, Y., Gao, M., Zhang, J., Kong, Q., Liu, Y., et al. (2012). Disruption of PAMP-induced MAP kinase cascade by a Pseudomonas syringae effector activates plant immunity mediated by the NB-LRR protein SUMM2. Cell Host Microbe 11, 253–263. doi: 10.1016/j.chom.2012.01.015
Zhou, F., Menke, F. L., Yoshioka, K., Moder, W., Shirano, Y., and Klessig, D. F. (2004). High humidity suppresses ssi4-mediated cell death and disease resistance upstream of MAP kinase activation, H2O2 production and defense gene expression. Plant J. 39, 920–932. doi: 10.1111/j.1365-313X.2004.02180.x
Keywords: Arabidopsis, autoimmunity, suppressor screening, PAMP-triggered immunity (PTI), effector-triggered immunity (ETI), receptor-like kinases, NB-LRR proteins
Citation: van Wersch R, Li X and Zhang Y (2016) Mighty Dwarfs: Arabidopsis Autoimmune Mutants and Their Usages in Genetic Dissection of Plant Immunity. Front. Plant Sci. 7:1717. doi: 10.3389/fpls.2016.01717
Received: 11 September 2016; Accepted: 01 November 2016;
Published: 17 November 2016.
Edited by:
Dingzhong Tang, Institute of Genetics and Developmental Biology (CAS), ChinaCopyright © 2016 van Wersch, Li and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yuelin Zhang, eXVlbGluLnpoYW5nQHViYy5jYQ==
 Rowan van Wersch
Rowan van Wersch Xin Li1,2
Xin Li1,2 Yuelin Zhang
Yuelin Zhang