- Molecular and Developmental Complexity Group, Unidad de Genómica Avanzada, Langebio-Cinvestav, Irapuato, México
Since the development of agriculture, humans have relied on the cultivation of plants to satisfy our increasing demand for food, natural products, and other raw materials. As we understand more about plant development, we can better manipulate plants to fulfill our particular needs. Auxins are a class of simple metabolites that coordinate many developmental activities like growth and the appearance of functional structures in plants. Computational modeling of auxin has proven to be an excellent tool in elucidating many mechanisms that underlie these developmental events. Due to the complexity of these mechanisms, current modeling efforts are concerned only with single phenomena focused on narrow spatial and developmental contexts; but a general model of plant development could be assembled by integrating the insights from all of them. In this perspective, we summarize the current collection of auxin-driven computational models, focusing on how they could come together into a single model for plant development. A model of this nature would allow researchers to test hypotheses in silico and yield accurate predictions about the behavior of a plant under a given set of physical and biochemical constraints. It would also provide a solid foundation toward the establishment of plant engineering, a proposed discipline intended to enable the design and production of plants that exhibit an arbitrarily defined set of features.
Auxin Modeling in An Intracellular Context
A change in auxin concentration induces a change in the transcriptional program of the cell. Auxin response is carried out by the Auxin Response Factor (ARF) family of transcriptional regulators. Due to the variety of ARFs present in plants, the auxin signal can be interpreted differently by cells that belong to different lineages and tissues (Rademacher et al., 2011). Taking all of them into account is necessary to accurately predict a plant’s phenotype, but this would require the simulation of a very complex regulation network. It has been shown, however, that this complexity can be considerably reduced if we identify the dominant players of the network and design a simplified model accordingly (Vernoux et al., 2011).
Auxin is transported from cell to cell through the action of active transporters that are present at the cell’s membrane, such as those belonging to the AUX/LAX and PIN families. This enables many important developmental events (Blilou et al., 2005). These transporter proteins are commonly found polarized toward a preferred region of the membrane, as described in a mathematical model by Kleine-Vehn et al. (2011). The location of these transporters correlates with the amount of auxin present in adjacent cells. Several hypotheses have been proposed to explain this and they have been tested through computational modeling (Wabnik et al., 2011).
A relatively simple model could account for both auxin response and transporter polarization to yield predictions on the distribution of auxin across a larger tissue. Other noteworthy phenomena that could be considered at this scale are: the diffusion of auxin within the cell (Lituiev et al., 2013); auxin transport between the cytoplasm and the nucleus (Herud et al., 2016); the feedback between auxin concentration and the amount of protein regulators like Aux/IAAs (Middleton et al., 2010) and membrane transporters like LAX3 (Mellor et al., 2015); and other mechanisms that are involved with auxin homeostasis (Mellor et al., 2016). All these examples could be integrated into a single model that describes the behavior of auxin from a cellular point of view.
Auxin Maxima
On a broader scale, as auxin travels across tissues, it tends to accumulate around localized spots which accrue a significantly higher concentration. Auxin maxima, as they are commonly known, usually precede the appearance of organs and other morphological structures, and consequently have been one of the most important traits that computational auxin models aim to recreate.
At the tip of Arabidopsis thaliana roots, a maximum of auxin maintains the identity of a group of cells known as the root stem cell niche (RSCN). A model by Grieneisen et al. (2007) reproduces the auxin dynamics in this region and considers auxin transport due to efflux membrane transporters (i.e., PIN proteins). The model represents the root as a 2-dimensional lattice of square cells which have different permeability values on their sides. With a similar setup, Tian et al. (2014) showed that the presence of WOX5, involved in auxin biosynthesis, is necessary for the appearance of this auxin maximum. Another study by Band et al. (2014) makes use of a realistic layout of cells and further extends the model by accounting for the effect of auxin influx transporters (i.e., AUX1/LAX).
It is crucial to consider the properties of auxin transporters when modeling the auxin flow that travels through a group of cells (Mitchison, 2015). Other mechanisms that regulate transporters themselves need to be considered as well. In A. thaliana, ICR1 and a family of related proteins control the deposition of PIN proteins on the cell membrane (Hazak et al., 2010). Their activity is dependent on the amount of auxin present within the cell and they are able to alter the resulting auxin distribution pattern of the tissue to which they belong (Hazak et al., 2014).
Regardless of the completeness of a single cellular model of auxin development, auxin maxima are emergent spatial features which cannot be seen if cells are considered as isolated entities. Their location, timing, and other physiological properties are the result of a complex system of cells interacting together and these characteristics are heavily influenced by the geometrical arrangement of neighboring cells and the phenomena associated with auxin transport from cell to cell.
Patterning and Growth
Auxin coordinates the development of many different structures that a plant needs to function, thrive, and reproduce (Vanneste and Friml, 2009). This is accomplished mainly through mechanisms that involve cell elongation, division, and differentiation; all of which are studied through computational modeling.
The distribution of cells found at the A. thaliana root tip can be explained by a model where cell division is controlled by a biological clock that depends on the amount of auxin present (Barrio et al., 2013), as well as specific developmental rules that seek to, among other things, balance out the total strain exerted by the growing tissue (De Vos et al., 2014).
In the A. thaliana RSCN, asymmetric cell division of the Cortex and Endodermis Initial (CEI) is controlled by the active form of the SCR/SHR protein complex (Di Laurenzio et al., 1996; Sabatini et al., 2003; Cui et al., 2007). RBR protein binds to, and inactivates, the SCR/SHR complex, unless prevented by CYCD6;1, which is expressed only in the CEI due to the existing auxin maximum at the RSCN. As cells move away from the auxin maximum and are, thus, exposed to a lower auxin concentration, RBR can deactivate SCR/SHR and prevent further asymmetric division of the daughter cells (Cruz-Ramírez et al., 2012).
Also in the root, the PLT family of transcription factors follows a concentration gradient that increases toward the tip and controls the growth of cells at the elongation zone. While the presence of PLT is relatively uniform and symmetric, the auxin gradient present is sensitive to changes in the physical orientation of the root structure. A model by Mähönen et al. (2014) describes how, through the integration of these two signals, the plant can effect different elongation rates on different regions of the root, allowing it to carry out an adequate gravitropic response.
Phyllotaxis is a particular pattern of organization, resembling a spiral, that is exhibited by many biological structures such as leaves, and the characteristic arrangement of seeds in a sunflower. Early studies involving auxin and its relationship with phyllotaxis have been carried out by Jönsson et al. (2006) and Smith et al. (2006). These studies locate auxin transporters within a given cell depending on the auxin concentration present in adjacent cells. This results in the repeated appearance of auxin maxima yielding a pattern that heavily resembles a phyllotactic arrangement. The phyllotactic pattern has also been found to be involved, and modeled, in the organization of floral primordia (van Mourik et al., 2012); and also influencing the development of leaves and their resulting shape (Chitwood et al., 2012).
Many other examples of computational auxin models applied to the study of patterning events exist. Jones et al. (2009) made use of a mathematical model to estimate the amount of auxin present in root hair cells; Bilsborough et al. (2011) showed how the appearance of auxin maxima spots along the edge of leaves gives rise to their resulting serration pattern; Fujita et al. (2011) explored the variety of arrangements arising from the shoot meristem of plants; Mirabet et al. (2012) studied the extent to which the phyllotactic pattern is vulnerable to naturally occurring fluctuations; Péret et al. (2013) proposed a mechanism that explains how lateral roots may emerge from the primary root structure; and Fàbregas et al. (2015) described a model that recreates the position and number of vascular bundles that appear in the developing shoot.
All these models further push the spatial context of auxin simulations into a larger realm that allows the study of the intended physiological function of biological structures. They are also built up from a common set of premises, particularly similar in regard to how they implement auxin transport between cells. The variety of biological functions that had been reproduced using this relatively small set of rules provides a strong argument in favor of a general model of plant development that relies on auxin as a common integrating signal.
Canalization Models
Plants rely on specialized structures that irrigate tissues with water and nutrients in order to function properly. The organization and function of these vascular tissues is controlled by a patterning event that involves auxin and its transport. When auxin flows through a tissue, the auxin transporters in its composing cells reorient toward the direction of the broader auxin flux. This leads to the establishment of preferential canals where auxin is transported along, a phenomenon known as canalization.
Early models involving canalization were proposed by Feugier et al. (2005), Rolland-Lagan and Prusinkiewicz (2005), Fujita and Mochizuki (2006a,b), Prusinkiewicz et al. (2009), and Wabnik et al. (2010), who all showed that the venation patterns can be produced through stable and self-organizing mechanisms. With different locations for auxin sources and sinks, an extensive variety of vascular arrangements can be generated. Cell division also needs to be carefully controlled in cells that are part of the emerging canal, so as to maintain the continuity of the venation pattern (Lee et al., 2014).
Canalization models are needed to incorporate into exhaustive models of plant development, as they explain how a plant can promote and maintain its functional structures. A study by O’Connor et al. (2014), takes a step in this direction, showing how vascularization and organ initiation are closely related events carried out by the concerted action of distinct PIN proteins. This study also extends the validity of the canonical auxin/PIN mechanism into a broader phylogenetic context, since they used Brachypodium distachyon as the model organism.
Structural Mechanics
Mechanical stress across tissues plays an important role during the life of a plant. This stress can be perceived by cells as it travels through their internal structure and it is known to induce change in the organization of the cell cytoskeleton (Hamant et al., 2008) and alter the polarization of auxin transporters (Heisler et al., 2010). Complementary to that, developmental effects resulting from auxin response alter the shape and other structural properties of a growing plant, hinting at a system where auxin and mechanical forces continuously interact with each other.
Auxin can induce changes in the pH of the apoplastic space, as thoroughly explained by a model from Steinacher et al. (2012). When the pH on the environment of a cell wall drops, its structural strength is diminished, allowing it to change its shape and yield to other forces like turgor pressure. This is a mechanism that allows the plant to control the anisotropic growth of tissues through the presence of varying concentrations of auxin (Sassi et al., 2014).
Mechanical forces are part of another theoretical layer that would greatly expand the scope of computational models of plant development. Unfortunately, accurate measurements on the mechanical properties of plant cells and tissues are still scarce. Nevertheless, recent efforts are producing valuable parameters; for example, Beauzamy et al. (2015) estimated the cell wall’s stiffness and turgor pressure at the A. thaliana shoot meristem.
Pathway Crosstalk
Further extending the influence of auxin throughout plant development, auxin is known to interact with other hormonal pathways. For example, cytokinins (CK) are known to have a particularly close relationship with auxin. The auxin/CK antagonistic interplay has been studied by Muraro et al. (2011), who predict a dynamic which can switch from a bistable equilibrium system to a system that exhibits oscillatory changes, leading to different developmental consequences for the plant. This model was used to recreate the architecture of the A. thaliana root by Muraro et al. (2013). The relationship between auxin and CK also is involved with other physiological features like miRNA regulation (Muraro et al., 2014; el-Showk et al., 2015) and the geometrical distribution of the developing cells (De Rybel et al., 2014).
Auxin and brassinosteroids (BRs) work together in the formation of vascular tissue inside the A. thaliana shoot. This tissue is directly specified by the presence of regularly spaced auxin maxima regions along the shoot. However, with different levels of BRs, the cells at the inflorescence stem change in number and size, which influences the resulting pattern of auxin maxima and, consequently, the number of vascular bundles that appear (Ibañes et al., 2009). Strigolactone is another hormone that interacts with auxin, as it is known to promote the dissociation of PIN1 from the membrane and, thus, alter the effective rate of auxin transport. This is linked to developmental features like shoot branching in A. thaliana (Shinohara et al., 2013).
Sometimes, multiple hormones can interact with auxin to produce a single phenotype. For instance, at the A. thaliana root, the observed auxin distribution pattern is known to be affected by a complex regulation network involving CK, ethylene, and the PLS protein (Liu et al., 2010; Moore et al., 2015).
The above are all prime examples of how auxin models could be extended to incorporate other regulation mechanisms that are important for plant development. A computational model intended to reproduce as much as we know about the metabolism of plants, should consider auxin as a starting point, due to the extensive work that has been published and is available to date.
Tropic Response
Plants can sense external stimuli from their environment and react to it by changing their developmental program. For example, regarding the gravitropic response in the A. thaliana root, the signal originates at the root tip, increasing the local auxin concentration in the tissue following the orientation of the gravitropic stimulus. This pulse, then, travels through the root to deliver the signal to the elongation zone, where the appropriate response is carried out (Mähönen et al., 2014). Band et al. (2012) devised a mathematical model to estimate the speed of auxin redistribution in response to changes in the gravity vector. Swarup et al. (2005) showed how the auxin pulse is transported through the epidermis, due to the presence of PIN and AUX/LAX membrane transporters.
A physical deformation exerted over a tissue is enough to trigger a tropic response from the plant, as it is known that roots are more prone to spawn lateral roots in regions that were previously bent. A convincing explanation for this is given in Laskowski et al. (2008), which predicts that modifying the shape of the root at the meristematic zone leads to an increase in auxin toward the outer region of the curve. This causes an increased accumulation of AUX1 transporters in pericycle cells and induces their reprogramming into lateral root founder cells.
Tropic response in computational models provides an environmental layer which greatly influences the development of a plant. Auxin is known to play a fundamental role in the plant’s response to gravity, light, humidity, and other environmental cues (Retzer et al., 2014). It would be reasonable, then, to consider auxin a fundamental signal that enables this environmental layer to interact with the remaining parts in a model of plant development.
Toward A Single Model of Plant Development
All computational models considered in this perspective are shown in Figure 1, along with their properties. The vast majority of them implement auxin diffusion and transport as the core mechanism that drives the computational simulation. Only a few computational models, to date, have considered the effects of a dynamically growing tissue together with its structural properties (e.g., Hamant et al., 2008; Barrio et al., 2013; Sassi et al., 2014).
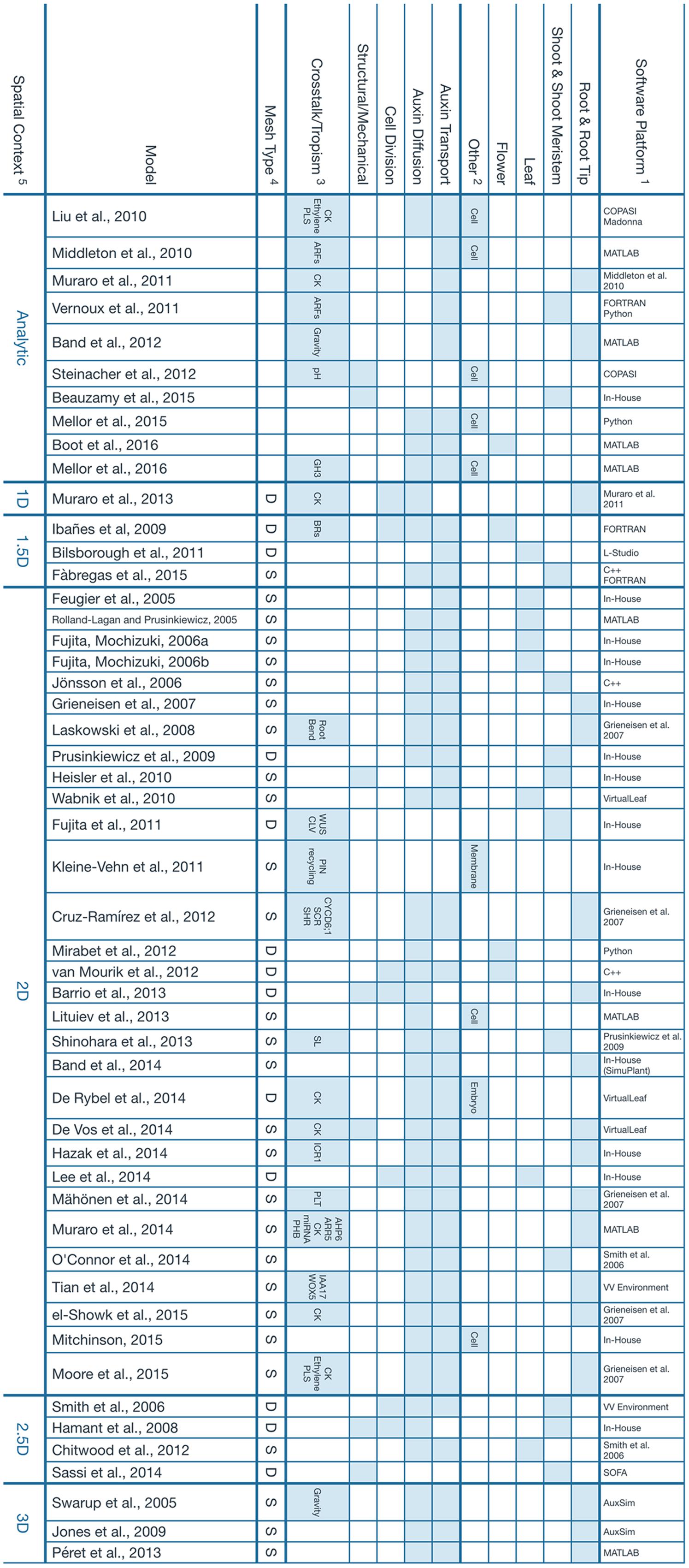
FIGURE 1. Main features of the computational models presented. 1Software tools that were used to build the model or previous model that a particular work was based on. The term In-House is used when, to our best effort, we could not find information regarding the implementation of a particular model. We assume it had to be a unique tool that was developed strictly for the scope of the referenced study. 2Other tissues or organs where the simulation takes place. The umbrella term Cell is used for intracellular dynamics that, in principle, should apply to any kind of tissue. 3Crosstalk and tropisms, but also additional signals or pathways that are modeled as a result of, or coupled to, Auxin expression. 4This property describes the nature of the mesh used to represent the tissue involved in the simulation. The letter S represents a static mesh established since the beginning of the model and that remains fixed throughout the whole simulation. A typical example is the lattice of square cells that is used to depict a 2D slice of the root. The letter D is used for a dynamic mesh that can deform and evolve as the simulation goes on. It can be used to represent cells which continuously change in size and geometry. 5The spatial context refers to the topology of the space where the simulation takes place. Analytical solutions are considered to be dimensionless. 1.5D refers to a 1D system describing a feature that exists normally in a 2D space, for instance, the edge of a growing leaf. Similarly, a 2.5D model is essentially a 2D model projected onto a 3D space, like the surface of the developing shoot meristem. Additional references for the software tools presented: AuxSim: Kramer, 2004; Berkeley Madonna: http://berkeleymadonna.com; COPASI: Hoops et al., 2006; L-Studio: http://algorithmicbotany.org/virtual_laboratory/; SOFA: Faure et al., 2012; VirtualLeaf: Merks et al., 2011; VV Environment: Smith et al., 2004.
The corpus of algorithms that the field of computational modeling of auxin has produced is enough to justify the creation of an integral model of plant development. This unified model would account, at least, for all the phenomena reviewed previously. In summary, this is how, we perceive such a model coming to existence. A single cellular layer integrates signals from physical phenomena (i.e., mechanical force, environmental stimuli) and regulatory pathways to ultimately define the metabolism of auxin and its transport. The cellular layer is then used to predict the resulting pattern of auxin present in a multi-cellular tissue which, in turn, signals the reprogramming of cells and the appearance of new organs as well as the vasculature needed to sustain them.
Studies that establish the physical constraints associated with auxin would further improve the validity of this model (Kramer et al., 2007, 2011; Beauzamy et al., 2015; Kramer and Ackelsberg, 2015; Boot et al., 2016). And, many existing software tools that deal with the acquisition of data (Schmidt et al., 2014; Barbier de Reuille et al., 2015) and its visualization (Band et al., 2014) would prove to be useful as well.
Due to its complexity, a model of this nature would have to come from a large collaborative effort among the interested scientific community. We identify two issues that could prevent this from happening, the lack of interoperation between the existing models and the difficulty of extending them with new features. This leads to most efforts producing solitary software packages that are rarely used beyond the scope of their particular study (Figure 1). These issues could be mitigated by the establishment of standard modeling conventions and tools, and we anticipate the need for a scientific consortium to coordinate the work of the parties concerned.
With a unified model of plant development, the scientific community would be able to evaluate in silico the phenotypical outcome of a given set of initial constraints. Furthermore, it would pave the way for the rational design of new biological structures and functions, a powerful paradigm that, we introduce here as plant engineering.
Plant engineering is a discipline concerned with the application of our current body of knowledge regarding plant development to the design, improvement, and creation of plants to satisfy particular sets of requirements. While this is still a vision set in the far future, we firmly believe that the first step toward it is to establish a common and exhaustive model of plant development built over the foundation laid out by computational modeling of auxin.
Author Contributions
AM-T and AC-R conceived the layout and scope of the manuscript. AM-T wrote the manuscript with guidance and further editing by AC-R.
Funding
Funding for this perspective was provided by Consejo Nacional de Ciencia y Tecnología.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We appreciate the work of all people involved with computational models of auxin and apologize to the ones whose work was not included due to space constraints. We thank Mrs. Collen Beard for her help in proofreading the final version of this manuscript.
References
Band, L. R., Wells, D. M., Fozard, J. A., Ghetiu, T., French, A. P., Pound, M. P., et al. (2014). Systems analysis of auxin transport in the Arabidopsis root apex. Plant Cell 26, 862–875. doi: 10.1105/tpc.113.119495
Band, L. R., Wells, D. M., Larrieu, A., Sun, J., Middleton, A. M., French, A. P., et al. (2012). Root gravitropism is regulated by a transient lateral auxin gradient controlled by a tipping-point mechanism. Proc. Natl. Acad. Sci. U.S.A. 109, 4668–4673. doi: 10.1073/pnas.1201498109
Barbier de Reuille, P., Routier-Kierzkowska, A.-L., Kierzkowski, D., Bassel, G. W., Schüpbach, T., Tauriello, G., et al. (2015). MorphoGraphX: a platform for quantifying morphogenesis in 4D. Elife 4:5864. doi: 10.7554/eLife.05864
Barrio, R. A., Romero-Arias, J. R., Noguez, M. A., Azpeitia, E., Ortiz-Gutiérrez, E., Hernández-Hernández, V., et al. (2013). Cell patterns emerge from coupled chemical and physical fields with cell proliferation dynamics: the Arabidopsis thaliana root as a study system. PLoS Comput. Biol. 9:e1003026. doi: 10.1371/journal.pcbi.1003026
Beauzamy, L., Louveaux, M., Hamant, O., and Boudaoud, A. (2015). Mechanically, the shoot apical meristem of Arabidopsis behaves like a shell inflated by a pressure of about 1 MPa. Front. Plant Sci. 6:1038. doi: 10.3389/fpls.2015.01038
Bilsborough, G. D., Runions, A., Barkoulas, M., Jenkins, H. W., Hasson, A., Galinha, C., et al. (2011). Model for the regulation of Arabidopsis thaliana leaf margin development. Proc. Natl. Acad. Sci. U.S.A. 108, 3424–3429. doi: 10.1073/pnas.1015162108
Blilou, I., Xu, J., Wildwater, M., Willemsen, V., Paponov, I., Friml, J., et al. (2005). The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433, 39–44. doi: 10.1038/nature03184
Boot, K. J. M., Hille, S. C., Libbenga, K. R., Peletier, L. A., Van Spronsen, P. C., Van Duijn, B., et al. (2016). Modelling the dynamics of polar auxin transport in inflorescence stems of Arabidopsis thaliana. J. Exp. Bot. 67, 649–666. doi: 10.1093/jxb/erv471
Chitwood, D. H., Headland, L. R., Ranjan, A., Martinez, C. C., Braybrook, S. A., Koenig, D. P., et al. (2012). Leaf asymmetry as a developmental constraint imposed by auxin-dependent phyllotactic patterning. Plant Cell 24, 2318–2327. doi: 10.1105/tpc.112.098798
Cruz-Ramírez, A., Díaz-Triviño, S., Blilou, I., Grieneisen, V. A., Sozzani, R., Zamioudis, C., et al. (2012). A bistable circuit involving SCARECROW-RETINOBLASTOMA integrates cues to inform asymmetric stem cell division. Cell 150, 1002–1015. doi: 10.1016/j.cell.2012.07.017
Cui, H., Levesque, M. P., Vernoux, T., Jung, J. W., Paquette, A. J., Gallagher, K. L., et al. (2007). An evolutionarily conserved mechanism delimiting SHR movement defines a single layer of endodermis in plants. Science 316, 421–425. doi: 10.1126/science.1139531
De Rybel, B., Adibi, M., Breda, A. S., Wendrich, J. R., Smit, M. E., Novak, O., et al. (2014). Integration of growth and patterning during vascular tissue formation in Arabidopsis. Science 345, 1255215–1255215. doi: 10.1126/science.1255215
De Vos, D., Vissenberg, K., Broeckhove, J., and Beemster, G. T. S. (2014). Putting theory to the test: which regulatory mechanisms can drive realistic growth of a root? PLoS Comput. Biol. 10:e1003910. doi: 10.1371/journal.pcbi.1003910
Di Laurenzio, L., Wysocka-Diller, J., Malamy, J. E., Pysh, L., Helariutta, Y., Freshour, G., et al. (1996). The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 86, 423–433. doi: 10.1016/S0092-8674(00)80115-4
el-Showk, S., Help-Rinta-Rahko, H., Blomster, T., Siligato, R., Marée, A. F. M., Mähönen, A. P., et al. (2015). Parsimonious model of vascular patterning links transverse hormone fluxes to lateral root initiation: auxin leads the way, while cytokinin levels out. PLoS Comput. Biol. 11:e1004450. doi: 10.1371/journal.pcbi.1004450
Fàbregas, N., Formosa-Jordan, P., Confraria, A., Siligato, R., Alonso, J. M., Swarup, R., et al. (2015). Auxin influx carriers control vascular patterning and xylem differentiation in Arabidopsis thaliana. PLoS Genet. 11:e1005183. doi: 10.1371/journal.pgen.1005183
Faure, F., Duriez, C., Delingette, H., Allard, J., Gilles, B., Marchesseau, S., et al. (2012). “SOFA: a multi-model framework for interactive physical simulation,” in Soft Tissue Biomechanical Modeling for Computer Assisted Surgery, ed. Y. Payan (Berlin: Springer). doi: 10.1007/8415_2012_125
Feugier, F. G., Mochizuki, A., and Iwasa, Y. (2005). Self-organization of the vascular system in plant leaves: inter-dependent dynamics of auxin flux and carrier proteins. J. Theor. Biol. 236, 366–375. doi: 10.1016/j.jtbi.2005.03.017
Fujita, H., and Mochizuki, A. (2006a). Pattern formation of leaf veins by the positive feedback regulation between auxin flow and auxin efflux carrier. J. Theor. Biol. 241, 541–551. doi: 10.1016/j.jtbi.2005.12.016
Fujita, H., and Mochizuki, A. (2006b). The origin of the diversity of leaf venation pattern. Dev. Dyn. 235, 2710–2721. doi: 10.1002/dvdy.20908
Fujita, H., Toyokura, K., Okada, K., and Kawaguchi, M. (2011). Reaction-diffusion pattern in shoot apical meristem of plants. PLoS ONE 6:e18243. doi: 10.1371/journal.pone.0018243
Grieneisen, V. A., Xu, J., Marée, A. F. M., Hogeweg, P., and Scheres, B. (2007). Auxin transport is sufficient to generate a maximum and gradient guiding root growth. Nature 449, 1008–1013. doi: 10.1038/nature06215
Hamant, O., Heisler, M. G., Jönsson, H., Krupinski, P., Uyttewaal, M., Bokov, P., et al. (2008). Developmental patterning by mechanical signals in Arabidopsis. Science 322, 1650–1655. doi: 10.1126/science.1165594
Hazak, O., Bloch, D., Poraty, L., Sternberg, H., Zhang, J., Friml, J., et al. (2010). A Rho scaffold integrates the secretory system with feedback mechanisms in regulation of auxin distribution. PLoS Biol. 8:e1000282. doi: 10.1371/journal.pbio.1000282
Hazak, O., Obolski, U., Prat, T., Friml, J., Hadany, L., and Yalovsky, S. (2014). Bimodal regulation of ICR1 levels generates self-organizing auxin distribution. Proc. Natl. Acad. Sci. U.S.A. 111, E5471–E5479. doi: 10.1073/pnas.1413918111
Heisler, M. G., Hamant, O., Krupinski, P., Uyttewaal, M., Ohno, C., Jönsson, H., et al. (2010). Alignment between PIN1 polarity and microtubule orientation in the shoot apical meristem reveals a tight coupling between morphogenesis and auxin transport. PLoS Biol. 8:e1000516. doi: 10.1371/journal.pbio.1000516
Herud, O., Weijers, D., Lau, S., and Jürgens, G. (2016). Auxin responsiveness of the MONOPTEROS-BODENLOS module in primary root initiation critically depends on the nuclear import kinetics of the Aux/IAA inhibitor BODENLOS. Plant J. 85, 269–277. doi: 10.1111/tpj.13108
Hoops, S., Sahle, S., Gauges, R., Lee, C., Pahle, J., Simus, N., et al. (2006). COPASI – a complex pathway simulator. Bioinformatics 22, 3067–3074. doi: 10.1093/bioinformatics/btl485
Ibañes, M., Fàbregas, N., Chory, J., and Caño-Delgado, A. I. (2009). Brassinosteroid signaling and auxin transport are required to establish the periodic pattern of Arabidopsis shoot vascular bundles. Proc. Natl. Acad. Sci. U.S.A. 106, 13630–13635. doi: 10.1073/pnas.0906416106
Jones, A. R., Kramer, E. M., Knox, K., Swarup, R., Bennett, M. J., Lazarus, C. M., et al. (2009). Auxin transport through non-hair cells sustains root-hair development. Nat. Cell Biol. 11, 78–84. doi: 10.1038/ncb1815
Jönsson, H., Heisler, M. G., Shapiro, B. E., Meyerowitz, E. M., and Mjolsness, E. (2006). An auxin-driven polarized transport model for phyllotaxis. Proc. Natl. Acad. Sci. U.S.A. 103, 1633–1638. doi: 10.1073/pnas.0509839103
Kleine-Vehn, J., Wabnik, K., Martinière, A., Łangowski,Ł, Willig, K., Naramoto, S., et al. (2011). Recycling, clustering, and endocytosis jointly maintain PIN auxin carrier polarity at the plasma membrane. Mol. Syst. Biol. 7:540. doi: 10.1038/msb.2011.72
Kramer, E. M. (2004). PIN and AUX/LAX proteins: their role in auxin accumulation. Trends Plant Sci. 9, 578–582. doi: 10.1016/j.tplants.2004.10.010
Kramer, E. M., and Ackelsberg, E. M. (2015). Auxin metabolism rates and implications for plant development. Front. Plant Sci. 6:150. doi: 10.3389/fpls.2015.00150
Kramer, E. M., Frazer, N. L., and Baskin, T. I. (2007). Measurement of diffusion within the cell wall in living roots of Arabidopsis thaliana. J. Exp. Bot. 58, 3005–3015. doi: 10.1093/jxb/erm155
Kramer, E. M., Rutschow, H. L., and Mabie, S. S. (2011). AuxV: a database of auxin transport velocities. Trends Plant Sci. 16, 461–463. doi: 10.1016/j.tplants.2011.05.003
Laskowski, M., Grieneisen, V. A., Hofhuis, H., ten Hove, C. A., Hogeweg, P., Marée, A. F. M., et al. (2008). Root system architecture from coupling cell shape to auxin transport. PLoS Biol. 6:2721–2735. doi: 10.1371/journal.pbio.0060307
Lee, S. W., Feugier, F. G., and Morishita, Y. (2014). Canalization-based vein formation in a growing leaf. J. Theor. Biol. 353, 104–120. doi: 10.1016/j.jtbi.2014.03.005
Lituiev, D. S., Krohn, N. G., Müller, B., Jackson, D., Hellriegel, B., Dresselhaus, T., et al. (2013). Theoretical and experimental evidence indicates that there is no detectable auxin gradient in the angiosperm female gametophyte. Development 140, 4544–4553. doi: 10.1242/dev.098301
Liu, J., Mehdi, S., Topping, J., Tarkowski, P., and Lindsey, K. (2010). Modelling and experimental analysis of hormonal crosstalk in Arabidopsis. Mol. Syst. Biol. 6, 3425–3441. doi: 10.1038/msb.2010.26
Mähönen, A. P., ten Tusscher, K., Siligato, R., Smetana, O., Díaz-Triviño, S., Salojärvi, J., et al. (2014). PLETHORA gradient formation mechanism separates auxin responses. Nature 515, 125–129. doi: 10.1038/nature13663
Mellor, N., Bennett, M. J., and King, J. R. (2016). GH3-mediated auxin conjugation can result in either transient or oscillatory transcriptional auxin responses. Bull. Math. Biol. 78, 210–234. doi: 10.1007/s11538-015-0137-x
Mellor, N., Péret, B., Porco, S., Sairanen, I., Ljung, K., Bennett, M., et al. (2015). Modelling of Arabidopsis LAX3 expression suggests auxin homeostasis. J. Theor. Biol. 366, 57–70. doi: 10.1016/j.jtbi.2014.11.003
Merks, R. M. H., Guravage, M., Inzé, D., and Beemster, G. T. S. (2011). VirtualLeaf: an open-source framework for cell-based modeling of plant tissue growth and development. Plant Physiol. 155, 656–666. doi: 10.1104/pp.110.167619
Middleton, A. M., King, J. R., Bennett, M. J., and Owen, M. R. (2010). Mathematical modelling of the Aux/IAA negative feedback loop. Bull. Math. Biol. 72, 1383–1407. doi: 10.1007/s11538-009-9497-4
Mirabet, V., Besnard, F., Vernoux, T., and Boudaoud, A. (2012). Noise and robustness in phyllotaxis. PLoS Comput. Biol. 8:e1002389. doi: 10.1371/journal.pcbi.1002389
Mitchison, G. (2015). The shape of an auxin pulse, and what it tells us about the transport mechanism. PLoS Comput. Biol. 11:e1004487. doi: 10.1371/journal.pcbi.1004487
Moore, S., Zhang, X., Mudge, A., Rowe, J. H., Topping, J. F., Liu, J., et al. (2015). Spatiotemporal modelling of hormonal crosstalk explains the level and patterning of hormones and gene expression in Arabidopsis thaliana wild-type and mutant roots. New Phytol. 207, 1110–1122. doi: 10.1111/nph.13421
Muraro, D., Byrne, H., King, J., and Bennett, M. (2013). The role of auxin and cytokinin signalling in specifying the root architecture of Arabidopsis thaliana. J. Theor. Biol. 317, 71–86. doi: 10.1016/j.jtbi.2012.08.032
Muraro, D., Byrne, H., King, J., Voß, U., Kieber, J., and Bennett, M. (2011). The influence of cytokinin-auxin cross-regulation on cell-fate determination in Arabidopsis thaliana root development. J. Theor. Biol. 283, 152–167. doi: 10.1016/j.jtbi.2011.05.011
Muraro, D., Mellor, N., Pound, M. P., Help, H., Lucas, M., Chopard, J., et al. (2014). Integration of hormonal signaling networks and mobile microRNAs is required for vascular patterning in Arabidopsis roots. Proc. Natl. Acad. Sci. U.S.A. 111, 857–862. doi: 10.1073/pnas.1221766111
O’Connor, D. L., Runions, A., Sluis, A., Bragg, J., Vogel, J. P., Prusinkiewicz, P., et al. (2014). A division in PIN-mediated auxin patterning during organ initiation in grasses. PLoS Comput. Biol. 10:e1003447. doi: 10.1371/journal.pcbi.1003447
Péret, B., Middleton, A. M., French, A. P., Larrieu, A., Bishopp, A., Njo, M., et al. (2013). Sequential induction of auxin efflux and influx carriers regulates lateral root emergence. Mol. Syst. Biol. 9:699. doi: 10.1038/msb.2013.43
Prusinkiewicz, P., Crawford, S., Smith, R. S., Ljung, K., Bennett, T., Ongaro, V., et al. (2009). Control of bud activation by an auxin transport switch. Proc. Natl. Acad. Sci. U.S.A. 106, 17431–17436. doi: 10.1073/pnas.0906696106
Rademacher, E. H., Möller, B., Lokerse, A. S., Llavata-Peris, C. I., van den Berg, W., and Weijers, D. (2011). A cellular expression map of the Arabidopsis AUXIN RESPONSE FACTOR gene family. Plant J. 68, 597–606. doi: 10.1111/j.1365-313X.2011.04710.x
Retzer, K., Korbei, B., and Luschnig, C. (2014). “Auxin and Tropisms,” in Auxin and Its Role in Plant Development, eds E. Zažímalová, J. Petrášek, and E. Benková (Vienna: Springer), 361–387. doi: 10.1007/978-3-7091-1526-8_16
Rolland-Lagan, A. G., and Prusinkiewicz, P. (2005). Reviewing models of auxin canalization in the context of leaf vein pattern formation in Arabidopsis. Plant J. 44, 854–865. doi: 10.1111/j.1365-313X.2005.02581.x
Sabatini, S., Heidstra, R., Wildwater, M., and Scheres, B. (2003). SCARECROW is involved in positioning the stem cell niche in the Arabidopsis root meristem. Genes Dev. 17, 354–358. doi: 10.1101/gad.252503
Sassi, M., Ali, O., Boudon, F., Cloarec, G., Abad, U., Cellier, C., et al. (2014). An auxin-mediated shift toward growth isotropy promotes organ formation at the shoot meristem in Arabidopsis. Curr. Biol. 24, 2335–2342. doi: 10.1016/j.cub.2014.08.036
Schmidt, T., Pasternak, T., Liu, K., Blein, T., Aubry-Hivet, D., Dovzhenko, A., et al. (2014). The iRoCS Toolbox – 3D analysis of the plant root apical meristem at cellular resolution. Plant J. 77, 806–814. doi: 10.1111/tpj.12429
Shinohara, N., Taylor, C., and Leyser, O. (2013). Strigolactone can promote or inhibit shoot branching by triggering rapid depletion of the auxin efflux protein PIN1 from the plasma membrane. PLoS Biol. 11:e1001474. doi: 10.1371/journal.pbio.1001474
Smith, C., Prusinkiewicz, P., and Samavati, F. (2004). Local specification of surface subdivision algorithms. 3062, 313–327. doi: 10.1007/978-3-540-25959-6_23
Smith, R. S., Guyomarc’h, S., Mandel, T., Reinhardt, D., Kuhlemeier, C., and Prusinkiewicz, P. (2006). A plausible model of phyllotaxis. Proc. Natl. Acad. Sci. U.S.A. 103, 1301–1306. doi: 10.1073/pnas.0510457103
Steinacher, A., Leyser, O., and Clayton, R. R. (2012). A computational model of auxin and pH dynamics in a single plant cell. J. Theor. Biol. 296, 84–94. doi: 10.1016/j.jtbi.2011.11.020
Swarup, R., Kramer, E. M., Perry, P., Knox, K., Leyser, H. M. O., Haseloff, J., et al. (2005). Root gravitropism requires lateral root cap and epidermal cells for transport and response to a mobile auxin signal. Nat. Cell Biol. 7, 1057–1065. doi: 10.1038/ncb1316
Tian, H., Wabnik, K., Niu, T., Li, H., Yu, Q., Pollmann, S., et al. (2014). WOX5-IAA17 feedback circuit-mediated cellular auxin response is crucial for the patterning of root stem cell niches in Arabidopsis. Mol. Plant 7, 277–289. doi: 10.1093/mp/sst118
van Mourik, S., Kaufmann, K., van Dijk, A. D. J., Angenent, G. C., Merks, R. M. H., and Molenaar, J. (2012). Simulation of organ patterning on the floral meristem using a polar auxin transport model. PLoS ONE 7:e28762. doi: 10.1371/journal.pone.0028762
Vanneste, S., and Friml, J. (2009). Auxin: a trigger for change in plant development. Cell 136, 1005–1016. doi: 10.1016/j.cell.2009.03.001
Vernoux, T., Brunoud, G., Farcot, E., Morin, V., Van den Daele, H., Legrand, J., et al. (2011). The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Mol. Syst. Biol. 7, 508–508. doi: 10.1038/msb.2011.39
Wabnik, K., Govaerts, W., Friml, J., and Kleine-Vehn, J. (2011). Feedback models for polarized auxin transport: an emerging trend. Mol. Biosyst. 7, 2352–2359. doi: 10.1039/c1mb05109a
Keywords: auxin, computational modeling, development, morphodynamics, plants
Citation: Morales-Tapia A and Cruz-Ramírez A (2016) Computational Modeling of Auxin: A Foundation for Plant Engineering. Front. Plant Sci. 7:1881. doi: 10.3389/fpls.2016.01881
Received: 01 September 2016; Accepted: 29 November 2016;
Published: 20 December 2016.
Edited by:
Daniel H. Chitwood, Donald Danforth Plant Science Center, USAReviewed by:
Jan Petrášek, Charles University, CzechiaGerhard Buck-Sorlin, Agrocampus Ouest, France
Copyright © 2016 Morales-Tapia and Cruz-Ramírez. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Alfredo Cruz-Ramírez, YWxmcmVkby5jcnV6QGNpbnZlc3Rhdi5teA==
 Alejandro Morales-Tapia
Alejandro Morales-Tapia Alfredo Cruz-Ramírez
Alfredo Cruz-Ramírez