- 1Plant Genomics Laboratory, Department of Life Sciences, Presidency University, Kolkata, India
- 2Division of Plant Biology, Bose Institute, Kolkata, India
Bamboos are an important member of the subfamily Bambusoideae, family Poaceae. The plant group exhibits wide variation with respect to the timing (1–120 years) and nature (sporadic vs. gregarious) of flowering among species. Usually flowering in woody bamboos is synchronous across culms growing over a large area, known as gregarious flowering. In many monocarpic bamboos this is followed by mass death and seed setting. While in sporadic flowering an isolated wild clump may flower, set little or no seed and remain alive. Such wide variation in flowering time and extent means that the plant group serves as repositories for genes and expression patterns that are unique to bamboo. Due to the dearth of available genomic and transcriptomic resources, limited studies have been undertaken to identify the potential molecular players in bamboo flowering. The public release of the first bamboo genome sequence Phyllostachys heterocycla, availability of related genomes Brachypodium distachyon and Oryza sativa provide us the opportunity to study this long-standing biological problem in a comparative and functional genomics framework. We identified bamboo genes homologous to those of Oryza and Brachypodium that are involved in established pathways such as vernalization, photoperiod, autonomous, and hormonal regulation of flowering. Additionally, we investigated triggers like stress (drought), physiological maturity and micro RNAs that may play crucial roles in flowering. We also analyzed available transcriptome datasets of different bamboo species to identify genes and their involvement in bamboo flowering. Finally, we summarize potential research hurdles that need to be addressed in future research.
Introduction
Flowering is one of the most important adaptations in the evolution of land plants. Numerous studies have been performed on annual, herbaceous model plants from dicotyledonous (Arabidopsis, Antirrhinum) and monocotyledonous (Oryza) groups to identify and characterize important floral pathway genes (Putterill et al., 2004; Colasanti and Coneva, 2009). However, the majority of commercially important plants are perennial and there remains a gap in translating knowledge gained from annual, model plants to perennial plants. Therefore, increasing research attention is being paid to perennial plants. While poplar (Jansson and Douglas, 2007) and white spurge have emerged as model perennial dicotyledonous plants (Anderson et al., 2007), research on perennialism remains elusive in monocots.
Bamboos are an important member of subfamily Bambusoideae, family Poaceae (Kellogg, 2015). Wide variations exist across bamboo species with respect to the flowering time, ranging from annual flowering to flowering after 120 years of vegetative growth (Janzen, 1976). There are even species for which the flowering time is not yet known. Variations in flowering time are not only diverse among species, but also at the population level. For instance, in the case of gregarious flowering all the individuals of a species growing over a wide geographical area bloom within a brief interval of time, and then all die after flowering (Nadgauda et al., 1997; Bhattacharya et al., 2009; Marchesini et al., 2009; Austin and Marchesini, 2012; Chaubey et al., 2013; Xie et al., 2016). In contrast, for sporadic flowering only a few culms of a population flower at a time (Ramanayake and Yakandawala, 1998; Bhattacharya et al., 2006; Xie et al., 2016). Such a wide variation in flowering time and extent indicates that the plant group serves as a repository for a wide range of genes and expression patterns that support such a life style. The ecological consequences of bamboo flowering, such as changes in dynamics of neighboring plant populations (Sertse et al., 2011), and impacts on endangered animals that depend on bamboo shoots (Reid et al., 1991; Azad-Thakur and Firake, 2014) have been topics of active research over decades. In comparison, the molecular aspects of bamboo flowering remain at a nascent stage. Studies have been conducted to characterize a limited number of flowering genes in different bamboo species such as MADS18 from Dendrocalamus latiflorus (Bo et al., 2005), FLOWERING LOCUS T (FT) from P. meyeri (Hisamoto et al., 2008), TERMINAL FLOWER 1 (TFL1) like gene from Bambusa oldhamii (Zeng et al., 2015), FRIGIDA (FRI) from P. violascens (Liu et al., 2015), MADS1 and MADS2 from P. praecox (Lin et al., 2009), 10 genes related to floral transition and meristem identity in D. latiflorus (Wang et al., 2014) and 16 MADS box genes from B. edulis (Shih et al., 2014). Such targeted approaches are being complemented by high-throughput approaches, namely, de novo transcriptome sequencing and suppression subtractive hybridization (Lin et al., 2010; Liu et al., 2012; Zhang et al., 2012; Peng et al., 2013; Gao et al., 2014; Ge et al., 2016; Wysocki et al., 2016; Zhao et al., 2016).
The main aim of this article is to consider the current status of molecular understanding of bamboo flowering from the perspective of comparative genomics and transcriptomics. We queried the only sequenced genome of a temperate bamboo, P. heterocycla syn. P. edulis, to identify marker genes in established floral pathways (e.g., photoperiodic, vernalization, hormonal, and autonomous) and the influence of additional factors such as drought stress and physiological maturity. P. edulis is a diploid, temperate bamboo with chromosome number 2n = 48 and having a genome size of 2.075 Gb (Gui et al., 2007; Peng et al., 2013). In addition, we also explored transcriptome datasets of available bamboo taxa to assess their possible role in bamboo flowering. Finally, we have identified challenges that need to be overcome to understand what triggers bamboo flowering, the genetic controls of flowering, and the effects of gregarious monocarpic flowering cycles on bamboo evolution.
Bamboo Genes Related to Established Flowring Pathways
Depending on the nature of environmental or endogenous cues, flowering pathways can be broadly classified into vernalization (cold responsive), photoperiodic (day length responsive), autonomous (endogenous factors) and hormonal pathways.
Vernalization Pathway
In the model monocot Oryza the important vernalization genes are VERNALIZATION 1 (VRN1), VERNALIZATION INSENSITIVE LIKE 2, and 3 (VIL 2, 3). An additional vernalization sensitive gene VRN2 was isolated from Triticum (Dubcovsky et al., 2006), while its Brachypodium homolog BdVRN2L is vernalization insensitive (Ream et al., 2014). BLAST analyses have identified multiple copies of OsVRN1, OsVIL2, and OsVIL3 homologs in P. heterocycla genome, but the homolog of VRN2 remained undetected (Table 1). In order to understand their possible involvement in bamboo flowering, all available floral transcriptomes were searched. VRN1 was detected in the shoot tissue specific EST library of B. oldhamii (Lin et al., 2010), while VIN3 was identified from the floral transcriptomes of P. heterocycla (Peng et al., 2013) and D. latiflorus (Zhang et al., 2012). Another important vernalization gene, At.FLC, performs cold-mediated suppression of the floral activator At.FT during the seasonal transition from fall to winter (Michaels and Amasino, 1999). However, during prolonged cold exposure in winter, FLC activity is gradually down-regulated by VRN1, VRN2, and VIN3 so that flowering is delayed until spring (Levy et al., 2002; Sung and Amasino, 2004). It was believed that FLC-like genes are absent in monocot plants (Choi et al., 2011), but recently two major FLC clades, namely, MADS37 and MADS51 genes, were identified in the temperate grass Brachypodium distachyon (Ruelens et al., 2013). Our BLAST analyses, however, could not detect MADS37 or MADS51 homologs in P. heterocycla at the set criterion of e−40, identity ≥50% and length coverage ≥60% of the query sequence (Table 1).
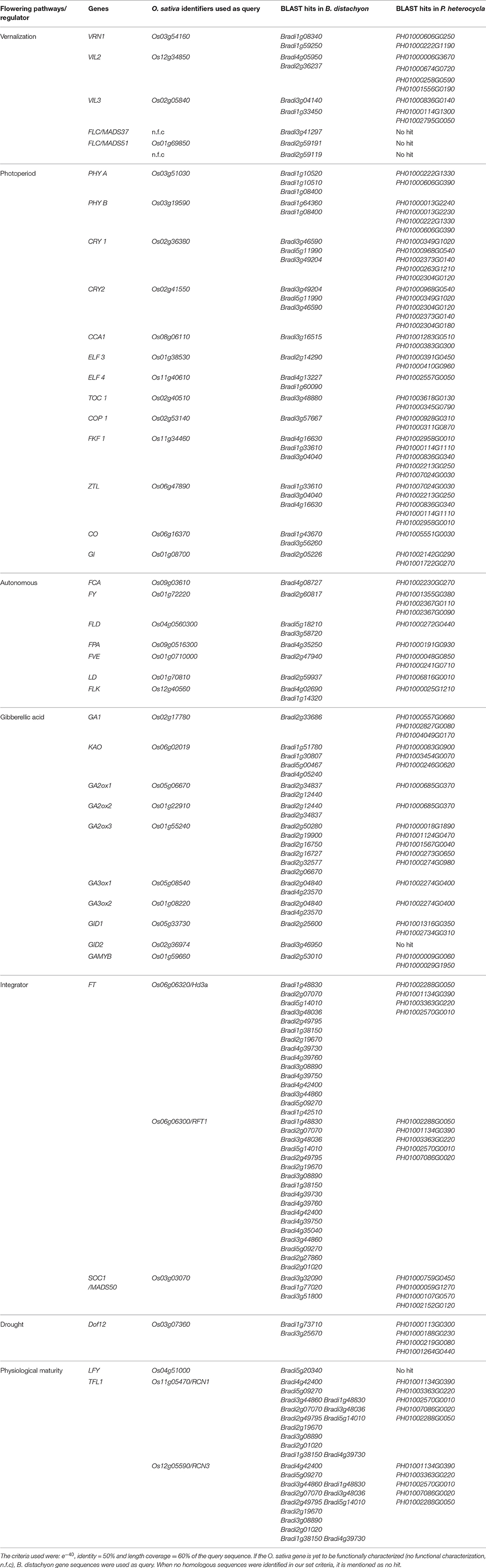
Table 1. Identification of important flowering gene homologs in the model temperate grass- Brachypodium distachyon and temperate bamboo- Phyllostachys heterocycla using Oryza sativa amino acid sequences as query in BLAST-P analyses.
Photoperiodic Pathway
In the photoperiodic pathway, the circadian rhythm of light and dark periods plays a major role in flower initiation. In Oryza a series of genes that include PHYTOCHROMES A and B (PHYA and PHYB), CRYPTOCHROMES 1 and 2 (CRY1 and CRY2), CIRCADIAN CLOCK ASSOCIATED 1 (CCA1), EARLY FLOWERING 4 (ELF4), TIMING OF CAB EXPRESSION 1 (TOC1), CONSTITUTIVE PHOTOMORPHOGENIC 1 (COP1), EARLY FLOWERING 3 (ELF3), GIGANTEA (GI), FLAVIN-BINDING KELCH REPEAT F BOX 1(FKF1) and ZEITLUPE (ZTL) receive the circadian signal and transfer it to CONSTANS (CO) for further downstream regulation. Our BLAST analyses identified at least one one homologous copy of each of these genes in the queried P. heterocycla genome (Table 1). ESTs homologous to CRY1, CRY2, PHY, FKF1, COP1, ELF3, ELF4, GI, CCA1, and CO were found in the floral transcriptomes of P. edulis, B. oldhamii, and D. latiflorus, suggesting their role in bamboo flower induction (Lin et al., 2010; Zhang et al., 2012; Peng et al., 2013; Gao et al., 2014). The transcriptional expression level of CO varied across libraries. For instance, it was low in P. edulis and correlated with the presence of L1 and GYPSY transposable elements in the regulatory region of the gene (Peng et al., 2013). On the other hand, a high level of CO expression was obtained in the floral tissues of D. latiflorus (Zhang et al., 2012). CO, along with the CCAAT box binding factor (NFY), bind to the CCAAT box of FT promoter and result in flowering (Ben-Naim et al., 2006). Therefore, the co-expression of CO and FT (i.e., CO-FT regulon) plays a crucial role in the regulation of flowering time. Our BLAST analyses identified 5 FT-like and 1 CO-like homologs in P. heterocycla (Table 1). Similarly, single or multiple FT copies have been identified and characterized in D. latiflorus, P. meyeri, and P. violascens (Hisamoto and Kobayashi, 2007, 2013; Hisamoto et al., 2008; Wang et al., 2014; Guo et al., 2015). Detailed expression analysis of PmFT revealed that its expression is primarily restricted to leaves, but highest during full bloom (Hisamoto and Kobayashi, 2013). Expression of the two FT genes and their functional diversification was reported in P. violascens (Guo et al., 2015). PvFT1 is expressed in leaves and induces flowering, while PvFT2 possibly plays a role in floral organogenesis. Another important floral integrator, SUPRESSOR OF OVEREXPRESSION OF CONSTANS 1 (SOC1), was identified by our BLAST analyses (Table 1) and was also expressed in the floral transcriptomes of P. edulis, Guadua inermis, Otatea acuminate, Lithachne pauciflora, and P. aurea (Peng et al., 2013; Wysocki et al., 2016).
Autonomous and Hormonal Pathway
In addition to environmental cues, additional flower inducing factors are present within a plant itself and are called endogenous or autonomous signals. This pathway is well studied in Arabidopsis, but is less characterized in monocot plants (Lee et al., 2005; Abou-Elwafa et al., 2011). The important genes are FLOWERING LOCUS CA (FCA), FLOWERING LOCUS D (FLD), FLOWERING LOCUS KH DOMAIN (FLK), FLOWERING LOCUS PA (FPA), FLOWERING LOCUS VE (FVE), FLOWERING LOCUS Y (FY), and LUMINIDEPENDENS (LD, Simpson, 2004). These genes promote flowering by suppressing FLC expression (Simpson, 2004; Quesada et al., 2005). Our BLAST analyses identified one or more P. heterocycla homologs for the majority of these genes (Table 1), which were reported in the floral transcriptomes of B. oldhamii (Lin et al., 2010), D. latiflorus (Zhang et al., 2012), and P. heterocycla (Peng et al., 2013) and suggest possible roles in bamboo flowering.
The role of gibberellic acid (GA) in the induction of flowering is well established in Oryza (Kwon and Paek, 2016). Many important genes related to GA biosynthesis (ent-KAURENE SYNTHETASE A- GA1, ent-KAURENOIC ACID OXIDASE-KAO, GA 2-OXIDASE-GA2ox, GA3ox) and receptors (GIBBERELLIN INSENSITIVE DWARF1- GID1, GID2) have been characterized (Sakamoto et al., 2004). GID1 and GID2 are responsible for proteasome mediated DELLA degradation and promote flowering through upregulation of GAMYB (Kwon and Paek, 2016). At least one P. heterocycla homolog has been detected for the majority of these genes in our BLAST analyses (Table 1). The possible involvement of GA in bamboo flowering is supported by the identification of GA1, SLY, GID1, GID2, GAMYB ESTs in the floral transcriptome of P. heterocycla (Gao et al., 2014) and D. latiflorus (Zhang et al., 2012).
Possible Physiological and Genetic Factors Regulating Bamboo Flowering
Stress
Increasing evidence suggests a link between stress and bamboo flowering (Rai and Dey, 2012; Peng et al., 2013; Ge et al., 2016). Overall expression level of general stress responsive genes involved in ABA, ethylene, sugar metabolism and Ca+2 dependent signaling pathway were 11.1-fold higher than that of the flowering genes in P. heterocycla (Peng et al., 2013). Particularly, a few members of the DNA binding with one finger (Dof) transcription factor family were highly up-regulated in the floral transcriptome (Imaizumi et al., 2005). For instance, Ph.Dof12 was about 16-fold up-regulated in the flowering tissues of P. heterocycla collected from a drought affected area (Peng et al., 2013). Similarly, 28 unigenes related to Dof3, Dof4, Dof5, Dof12, and Cycling Dof Factors (CDF) were detected in the floral transcriptome of P. edulis (Gao et al., 2014). The Dof family is composed of 15 genes in Phyllostachys and a comprehensive functional characterization of these genes may provide new insights. Particularly, analyzing the enrichment of the drought-responsive cis-elements in their promoter regions could identify candidate genes that are induced under drought conditions.
Physiological Maturity and Micro RNAs
Scientific evidence emerging from research on various perennial plants suggests an important role of TERMINAL FLOWER 1 (TFL1) and microRNAs (miRNAs) in maintaining a long vegetative phase (Huijser and Schmid, 2011). Our BLAST analyses identified five copies of Ph.TFL1 genes in P. heterocycla (Table 1). A functional TFL1 gene was isolated from B. oldhamii and was overexpressed in Arabidopsis (Zeng et al., 2015). The overexpressed lines showed delayed flowering, suggesting that TFL1 may have a role in maintaining vegetative growth. In addition, TFL1 may have an important function in differentiation of bamboo floral organs, as indicated by higher expression of TFL1 in late floral developmental stages relative to early stages in B. oldhamii and D. latiflorus (Wang et al., 2014).
Long maintenance of the vegetative phase in the majority of bamboos can also be regulated at the post-transcriptional level, such as by miRNAs. In rice miR156 is known to repress flowering by targeting SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SBP/SPL) transcription factor (SPLs, Xiong et al., 2006). Expression of miR156 showed significant down-regulation through the transition from vegetative to flowering stages in P. edulis (Gao et al., 2015). Additional candidates that may have roles are miR164a, miR166a, miR167a, miR535a, miR159a.1, miR164a, and miR168-3-p (Gao et al., 2015; Ge et al., 2016). In contrast, some micro RNAs may play positive roles in bamboo flowering. One such candidate is miR172, which controls flowering time and the formation of floral organs through the regulation of the AP2-like transcription factor (Lee et al., 2014). miR172a showed an increase in expression level during progression from vegetative to the flowering phase in P. edulis (Gao et al., 2015). The expression of other miRNAs such as miR169b, miR395h-5p, and miR529-3p were higher in floral tissues than in vegetative tissues.
Future Challenges
Appropriate Tissue Sampling
Identification of proper tissue stages is critical since the majority of flowering genes are transiently expressed soon before or after floral induction. Unlike Arabidopsis or Oryza, wild bamboo floral tissue stages are not easily traceable. Therefore, tissue culture methods have been tried to induce flowering and to study defined stages of induced floral transcriptomes of B. oldhamii in vitro (Lin et al., 2010). However, this study raised doubt about comparability of the transcription patterns under in vitro conditions vs. naturally occurring flowering. A large unigene set (146,395) generated from the floral transcriptomes of naturally grown D. latiflorus could not detect the important integrator gene FT, although it was detected in the transcriptome of P. edulis. This emphasizes the need to define in vivo floral stages with higher accuracy in order to make data generated by different research groups more comparable. Therefore, we studied the microscopic histology of different flowering stages of wild B. tulda plants and compared them with the external morphology of buds to identify phenotypic markers for specific growth stages (Figure 1). The external morphological features of nodal vegetative buds are indistinguishable from those of early stage inflorescence bud. However, this is one of the most crucial tissue stages with respect to the identification of genes involved in flower induction. Close observation of the early inflorescence bud revealed that it is slightly smaller in size, pale yellow in color, and bulged in the middle (Figures 1A,C). Histological analyses reveal that the shoot apical meristem of the nodal vegetative bud is dome shaped and covered with compactly arranged leaf primordia (Figure 1B). But the early staged inflorescence meristem is slightly smaller in size and triangular in shape (Figure 1D). The middle stage floral bud could be differentiated from the early stage by its elongated shape and bright green color (Figure 1E). Histological analysis revealed that it is composed of one or two floral primordia at the base of the rachis and an undifferentiated inflorescence meristem at the apex (Figure 1F). The late inflorescence bud is easily identifiable from all the other stages by its long and slender shape (Figure 1G). It is composed of three to four visible florets having differentiated anther primordia at the base of the rachis and an undifferentiated apical inflorescence meristem (Figure 1H).
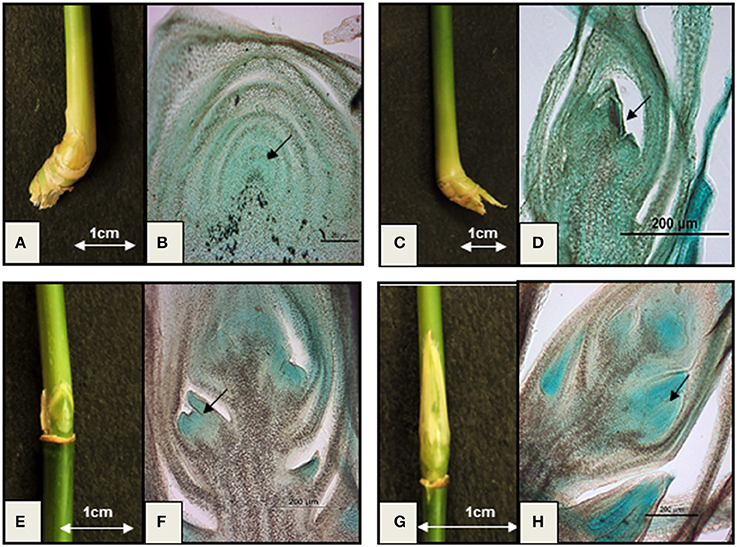
Figure 1. Important vegetative and floral developmental stages of B. tulda. (A) External morphology of nodal vegetative bud (~0.6 × 0.7 cm in dimension); (B) Longitudinal section (L.S.) of vegetative bud. The shoot apical meristem (SAM) is dome shaped (marked with arrow); (C) External morphology of an early stage inflorescence bud (~0.3 × 0.3 cm in dimension); (D) LS of the early stage inflorescence bud having triangular inflorescence meristem (marked with arrow); (E) External morphology of middle stage inflorescence bud (~0.8 × 0.5 cm in dimension); (F) LS of middle stage inflorescence bud showing differentiated floral primordia (marked with arrow); (G) External morphology of late stage inflorescence bud (~1.2 × 0.6 cm in dimension); (H) LS of late stage inflorescence bud having differentiated anther primordia (marked with arrow).
Gene Family Expansion, High Sequence Homology and Associated Challenges
Bamboos are highly polyploid plants with big genomes (2075 Mb for P. heterocycla compared to 125 Mb for A. thaliana). Consequently, the majority of genes are present in multiple copies. It would be important to dissect their evolutionary origin (orthologs-functional, paralogs-old/recent vs. tandem duplicates) and deduce their functional conservation or divergence by studying detailed transcriptional expression patterns (Das et al., 2016). However, the majority of these genes are very similar in sequence, which creates challenges in maintaining specificity in gene expression analyses. Example of this are FT and TFL1 genes, which are members of the Phosphatidylethanolamine-binding protein (PEBP) family and share high sequence similarity (>60%). However, they are functionally antagonistic to each other. There are diagnostic amino acids, which are crucial to maintain either FT (Tyr-85) or TFL1 (His-88) function (Hanzawa et al., 2005). Our BLAST analyses identified five P. heterocycla homologs each for FT and TFL1 and they are completely overlapping with each other (Table 1). Follow-up analysis indicated PH01002288G0050 as the predicted FT gene, while the other four, PH01001134G0390, PH01003363G0220, PH01002570G0010, PH01007086G0020 are TFL1. Therefore, in addition to large-scale sequence analyses such as BLAST, individual gene sequences should be checked for correct gene function annotation.
Genetic Tools for functional Validation
With the completion of gene sequencing and expression pattern characterization, the next challenge would be to confirm gene functions using loss- or gain-of-function mutants. This is especially important for multi copy genes for which expression data is not indicative of functional differentiation among copies. Therefore, a model plant is needed in which tissue culture and genetic transformation are easy to perform. Woody bamboos are generally recalcitrant and present several challenges (Das and Pal, 2005a). Since loss-of-function mutation analyses would be challenging, other model plants could be exploited to perform genetic complementation analyses by ectopically expressing bamboo flowering genes. Rice could be useful for such purposes due to its close evolutionary relationship, related floral biology and availability of mutant lines for several genes. However, many rice genes and associated mutant phenotypes have yet to be characterized.
Development of a New Model System for Tropical Bamboo
The majority of available research reports are on the tetraploid bamboo Phyllostachys, predominantly found in the temperate regions of China and Japan. However, enormous biodiversity is found in the tropical regions and dominated by members of the genus Bambusa. Therefore, the genome/transcriptomes of a tropical bamboo should be characterized. These have enormous economic importance, a large population size, wide genetic diversity (Das et al., 2008), molecular methods for species level identification (Das et al., 2005), a standardized micropropagation protocol (Das and Pal, 2005b), incidents of both gregarious (Mohan Ram and Harigopal, 1981) and sporadic flowering (Bhattacharya et al., 2006), which taken together makes B. tulda a good choice as a model species of tropical bamboos.
Author Contributions
MD and AP collaborated in this study. PB, SC, and SD had done the bioinformatics and histological analyses. MD wrote the paper with input from all co-authors.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The research results reported in this paper are funded by Council of Scientific and Industrial Research, India [38(1386)/14/EMR-II], Department of Biotechnology, India (BT/PR10778/PBD/16/1070/2014) and Faculty Research and Professional Development Fund (FRPDF) grant, Presidency University. PB acknowledges a JRF fellowship from University Grant Commission, India. We thank Prof. James Westwood, Virginia Tech for carefully editing the manuscript and the three reviewers and the editor for their critical comments to substantially improve the quality of the manuscript.
References
Abou-Elwafa, S. F., Büttner, B., Chia, T., Schulze-Buxloh, G., Hohmann, U., Mutasa-Gottgens, E., et al. (2011). Conservation and divergence of autonomous pathway genes in the flowering regulatory network of Beta vulgaris. J. Exp. Bot. 62, 3359–3374. doi: 10.1093/jxb/erq321
Anderson, J. V., Horvath, D. P., Chao, W. S., Foley, M. E., Hernandez, A. G., Thimmapuram, J., et al. (2007). Characterization of an EST database for the perennial weed leafy spurge: an important resource for weed biology research. Weed Sci. 55, 193–203. doi: 10.1614/WS-06-138.1
Austin, A. T., and Marchesini, V. A. (2012). Gregarious flowering and death of understorey bamboo slow litter decomposition and nitrogen turnover in a southern temperate forest in Patagonia, Argentina. Funct. Ecol. 26, 265–273. doi: 10.1111/j.1365-2435.2011.01910.x
Azad-Thakur, N. S., and Firake, D. M. (2014). Population dynamics of rodents during bamboo flowering in north east India. Indian J. Agric. Sci. 84, 6.
Ben-Naim, O., Eshed, R., Parnis, A., Teper-Bamnolker, P., Shalit, A., Coupland, G., et al. (2006). The CCAAT binding factor can mediate interactions between CONSTANS-like proteins and DNA. Plant J. 46, 462–476. doi: 10.1111/j.1365-313X.2006.02706.x
Bhattacharya, S., Das, M., Bar, R., and Pal, A. (2006). Morphological and molecular characterization of Bambusa tulda with a note on flowering. Ann. Bot. 98, 529–535. doi: 10.1093/aob/mcl143
Bhattacharya, S., Ghosh, J. S., Das, M., and Pal, A. (2009). Morphological and molecular characterization of Thamnocalamus spathiflorus subsp. spathiflorus at population level. Plant Syst. Evol. 282, 13–20. doi: 10.1007/s00606-008-0092-1
Bo, T., Yongyan, C., Yuanxin, Y., and Dezhu, L. (2005). Isolation and ectopic expression of a bamboo MADS-box gene. Chin. Sci. Bull. 50, 217–224. doi: 10.1007/BF02897530
Chaubey, O. P., Sharma, A., and Prakash, R. (2013). Impact of fires and grazing closure on rehabilitation of gregariously flowered bamboo (Dendrocalamus strictus. Roxb. Nees) forests. Int. J. Bio-Sci. Bio-Tech. 5, 207–220. doi: 10.14257/ijbsbt.2013.5.6.22
Choi, K., Kim, J., Hwang, H. J., Kim, S., Park, C., Kim, S. Y., et al. (2011). The FRIGIDA complex activates transcription of FLC, a strong flowering repressor in Arabidopsis, by recruiting chromatin modification factors. Plant Cell 23, 289–303. doi: 10.1105/tpc.110.075911
Colasanti, J., and Coneva, V. (2009). Mechanisms of floral induction in grasses: something borrowed something new. Plant Physiol. 149, 56–62. doi: 10.1104/pp.108.130500
Das, M., Bhattacharya, S., and Pal, A. (2005). Generation and characterization of SCARs by cloning and sequencing of RAPD products: a strategy for species-specific marker development in bamboo. Ann. Bot. 95, 835–841. doi: 10.1093/aob/mci088
Das, M., Bhattacharya, S., Singh, P., Filgueiras, T., and Pal, A. (2008). Bamboo taxonomy and diversity in the era of molecular markers. Adv. Bot. Res. 47, 225–268. doi: 10.1016/S0065-2296(08)00005-0
Das, M., Haberer, G., Panda, A., Das Laha, S., Ghosh, T. C., and Schäffner, A. R. (2016). Expression pattern similarities support the prediction of orthologs retaining common functions after gene duplication events. Plant Physiol. 171, 2343–2357. doi: 10.1104/pp.15.01207
Das, M., and Pal, A. (2005a). In vitro regeneration of Bambusa balcooa Roxb: factors affecting changes of morphogenetic competence in the axillary buds. Plant Cell Tiss. Organ Cult. 81, 109–112. doi: 10.1007/s11240-004-3017-x
Das, M., and Pal, A. (2005b). Clonal propagation and production of genetically uniform regenerants from axillary meristems of adult bamboo. J. Plant Biochem. Biotech. 14, 185–188. doi: 10.1007/BF03355956
Dubcovsky, J., Loukoianov, A., Fu, D., Valarik, M., Sanchez, A., and Yan, L. (2006). Effect of photoperiod on the regulation of wheat vernalization genes VRN1 and VRN2. Plant Mol. Biol. 60, 469–480. doi: 10.1007/s11103-005-4814-2
Gao, J., Ge, W., Zhang, Y., Cheng, Z., Li, L., Hou, D., et al. (2015). Identification and characterization of microRNAs at different flowering developmental stages in moso bamboo (Phyllostachys edulis) by high throughput sequencing. Mol. Genet. Genomics 290, 2335–2353. doi: 10.1007/s00438-015-1069-8
Gao, J., Zhang, Y., Zhang, C., Qi, F., Li, X., Mu, S., et al. (2014). Characterization of the floral transcriptome of moso bamboo (Phyllostachys edulis) at different flowering developmental stages by transcriptome sequencing and RNA-seq analysis. PLoS ONE 9:e98910. doi: 10.1371/journal.pone.0098910
Ge, W., Zhang, Y., Cheng, Z., Hou, D., Li, X., and Gao, J. (2016). Main regulatory pathways, key genes, and microRNAs involved in flower formation and development of moso bamboo (Phyllostachys edulis). Plant Biotechnol. J. doi: 10.1111/pbi.12593. [Epub ahead of print].
Gui, Y., Wang, S., Quan, L., Zhou, C., Long, S., Zheng, H., et al. (2007). Genome size and sequence composition of moso bamboo: a comparative study. Sci. China C Life Sci. 50, 700–705. doi: 10.1007/s11427-007-0081-6
Guo, X., Wang, Y., Wang, Q., Xu, Z., and Lin, X. (2015). Molecular characterization of FLOWERING LOCUS T (FT) genes from bamboo (Phyllostachys violascens). J. Plant Biochem. Biotechnol. 25, 168–178. doi: 10.1007/s13562-015-0322-x
Hanzawa, H., Money, T., and Bradely, D. (2005). A single amino acid converts a repressor to an activator of flowering. Proc. Natl. Acad. Sci. U.S.A. 102, 7748–7753. doi: 10.1073/pnas.0500932102
Hisamoto, Y., Kashiwagi, H., and Kobayashi, M. (2008). Use of flowering gene FLOWERING LOCUS T (FT) homologs in the phylogenetic analysis of bambusoid and early diverging grasses. J. Plant Res. 121, 451–461. doi: 10.1007/s10265-008-0181-9
Hisamoto, Y., and Kobayashi, M. (2007). Comparison of nucleotide sequences of fragments from rice FLOWERING LOCUS T (RFT1) homologues in Phyllostachys (Bambusoideae, Poaceae) with particular reference to flowering behaviour. Kew Bull. 62, 463–473.
Hisamoto, Y., and Kobayashi, M. (2013). Flowering habit of two two bamboo species, Phyllostachys meyeri and Shibataea chinensis, analyzed with flowering gene expression. Plant Species Biol. 28, 109–117. doi: 10.1111/j.1442-1984.2012.00369.x
Huijser, P., and Schmid, M. (2011). The control of developmental phase transitions in plants. Development 138, 4117–4129. doi: 10.1242/dev.063511
Imaizumi, T., Schultz, T. F., Harmon, F. G., Ho, L. A., and Kay, S. A. (2005). FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science 309, 293–297. doi: 10.1126/science.1110586
Jansson, S., and Douglas, C. J. (2007). Populus: a model system for plant biology. Annu. Rev. Plant Biol. 58, 435–458. doi: 10.1146/annurev.arplant.58.032806.103956
Janzen, D. H. (1976). Why bamboos wait so long to flower. Annu. Rev. Ecol. Syst. 7, 347–391. doi: 10.1146/annurev.es.07.110176.002023
Kwon, C. T., and Paek, N. C. (2016). Gibberellic acid: a key phytohormone for spikelet fertility in rice grain production. Int. J. Mol. Sci. 17:794. doi: 10.3390/ijms17050794
Lee, J. H., Cho, Y. S., Yoon, H. S., Suh, M. C., Moon, J., Lee, I., et al. (2005). Conservation and divergence of FCA function between Arabidopsis and rice. Plant Mol. Biol. 58, 823–838. doi: 10.1007/s11103-005-8105-8
Lee, Y. S., Lee, D. Y., Cho, L. H., and An, G. (2014). Rice miR172 induces flowering by suppressing OsIDS1 and SNB, two two AP2 genes that negatively regulate expression of Ehd1 and florigens. Rice J. 7, 31. doi: 10.1186/s12284-014-0031-4
Levy, Y. Y., Mesnage, S., Mylne, J. S., Gendall, A. R., and Dean, C. (2002). Multiple roles of Arabidopsis VRN1 in vernalization and flowering time control. Science 297, 243–246. doi: 10.1126/science.1072147
Lin, E. P., Peng, H. Z., Jin, Q. Y., Deng, M. J., Li, T., Xiao, X. C., et al. (2009). Identification and characterization of two two bamboo (Phyllostachys praecox) AP1/SQUA-like MADS-box genes during floral transition. Planta 231, 109–120. doi: 10.1007/s00425-009-1033-0
Lin, X. C., Chow, T. Y., Chen, H. H., Liu, C. C., Chou, S. J., Huang, B. L., et al. (2010). Understanding bamboo flowering based on large-scale analysis of expressed sequence tags. Genet. Mol. Res. 9, 1085–1093. doi: 10.4238/vol9-2gmr804
Liu, M., Qiao, G., Jiang, J., Yang, H., Xie, L., Xie, J., et al. (2012). Transcriptome sequencing and de novo analysis for mabamboo (Dendrocalamus latiflorus Munro) using the illumina platform. PLoS ONE 7:e46766. doi: 10.1371/journal.pone.0046766
Liu, S. N., Zhu, L. F., Lin, X. C., and Ma, L. Y. (2015). Overexpression of the repressor gene PvFRI-L from Phyllostachys violascens delays flowering time in transgenic Arabidopsis thaliana. Biol. Plant. 60, 401–409. doi: 10.1007/s10535-016-0614-6
Marchesini, V. A., Sala, O. E., and Austin, A. T. (2009). Ecological consequences of a massive flowering event of bamboo (Chusquea culeou) in a temperate forest of Patagonia, Argentina. J. Veg. Sci. 20, 424–432. doi: 10.1111/j.1654-1103.2009.05768.x
Michaels, S. D., and Amasino, R. M. (1999). FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11, 949–956. doi: 10.1105/tpc.11.5.949
Mohan Ram, H. Y., and Harigopal, B. (1981). Some observations on the flowering of bamboos in Mizoram. Curr. Sci. 50, 708–710.
Nadgauda, R. S., John, C. K., Parasharami, V. A., Joshi, M. S., and Mascarenhas, A. F. (1997). A comparison of in vitro and in vivo flowering in bamboo: Bambusa arundinacea. Plant Cell Tiss. Org. Cult. 48, 181–188. doi: 10.1023/A:1005800700024
Peng, Z., Lu, Y., Li, L., Zhao, Q., Feng, Q., Gao, Z., et al. (2013). The draft genome of the fast-growing non-timber forest species moso bamboo (Phyllostachys heterocycla). Nat. Genet. 45, 456–461. doi: 10.1038/ng.2569
Putterill, J., Laurie, R., and Macknight, R. (2004). It's time to flower: the genetic control of flowering time. Bioessays 26, 363–373. doi: 10.1002/bies.20021
Quesada, V., Dean, C., and Simpson, G. G. (2005). Regulated RNA processing in the control of Arabidopsis flowering. Int. J. Dev. Biol. 49, 773–780. doi: 10.1387/ijdb.051995vq
Rai, V., and Dey, N. (2012). Identification of programmed cell death related genes in bamboo. Gene 497, 243–248. doi: 10.1016/j.gene.2012.01.018
Ramanayake, S. M. S. D., and Yakandawala, K. (1998). Incidence of flowering, death and the phenology of the giant bamboo (Dendrocalamus giganteus Wall. Ex Munro). Ann. Bot. 82, 779–785. doi: 10.1006/anbo.1998.0754
Ream, T. S., Woods, D. P., Schwartz, C. J., Sanabria, C. P., Mahoy, J. A., Walters, E. M., et al. (2014). Interaction of photoperiod and vernalization determines flowering time of Brachypodium distachyon. Plant Physiol. 164, 694–709. doi: 10.1104/pp.113.232678
Reid, D. G., Alan, H., Taylor, A. H., Jinchui, H., and Zisheng, Q. (1991). Environmental influences on bamboo Bashania fangiana growth and implications for giant panda conservation. J. Appl. Ecol. 28, 855–868. doi: 10.2307/2404212
Ruelens, P., Maagd, R., Proost, S., Theiben, G., Geuten, K., and Kaufmann, K. (2013). FLOWERING LOCUS C in monocots and the tandem origin of angiosperm-specific MADS-box genes. Nat. Commun. 4, 2280. doi: 10.1038/ncomms3280
Sakamoto, T., Miura, K., Itoh, H., Tatsumi, T., Ueguchi-Tanaka, M., Ishiyama, K., et al. (2004). An overview of gibberellin metabolism enzyme genes andtheir related mutants in rice. Plant Physiol. 134, 1642–1653. doi: 10.1104/pp.103.033696
Sertse, D., Disasa, T., Bekele, K., Alebachew, M., Kebede, Y., Eshete, N., et al. (2011). Mass flowering and death of bamboo: a potential threat to biodiversity and livelihoods in Ethiopia. J. Biol. Environ. Sci. 1, 16–25.
Shih, M. C., Chou, M. L., Yue, J. J., Hsu, C. T., Chang, W. J., Ko, S. S., et al. (2014). BeMADS1 is a key to delivery MADSs into nucleus in reproductive tissues-De novo characterization of Bambusa edulis transcriptome and study of MADS genes in bamboo floral development. BMC Plant Biol. 14:179. doi: 10.1186/1471-2229-14-179
Simpson, G. G. (2004). The autonomous pathway: epigenetic and posttranscriptional gene regulation in the control of Arabidopsis flowering time. Curr. Opin. Plant Biol. 7, 570–574. doi: 10.1016/j.pbi.2004.07.002
Sung, S., and Amasino, R. (2004). Vernalization and epigenetics: how plants remember winter. Curr. Opin. Plant Biol. 7, 4–10. doi: 10.1016/j.pbi.2003.11.010
Wang, X., Zhang, X., Zhao, L., and Guo, Z. (2014). Morphology and quantitative monitoring of gene expression patterns during floral induction and early flower development in Dendrocalamus latiflorus. Int. J. Mol. Sci. 15, 12074–12093. doi: 10.3390/ijms150712074
Wysocki, W. P., Ruiz-Sanchez, E., Yin, Y., and Duvall, M. R. (2016). The floral transcriptomes of four four bamboo species (Bambusoideae; Poaceae): support for common ancestry among woody bamboos. BMC Genomics 17:384. doi: 10.1186/s12864-016-2707-1
Xie, N., Chen, L. N., Wong, K. M., Cui, Y. Z., and Yang, H. Q. (2016). Seed set and natural regeneration of Dendrocalamus membranaceus munro after mass and sporadic flowering in Yunnan, China. one PLoS ONE 11:e0153845. doi: 10.1371/journal.pone.0153845
Xiong, L., Wu, C., and Xie, K. (2006). Genomic organization, differential expression, and interaction of SQUAMOSA promoter-Binding-Like transcription factors and microRNA156 in rice. Plant Physiol. 142, 280–293. doi: 10.1104/pp.106.084475
Zeng, H. Y., Lu, Y. T., Yang, X. M., Xu, Y. H., and Lin, X. C. (2015). Ectopic expression of the BoTFL1-like gene of Bambusa oldhamii delays blossoming in Arabidopsis thaliana and rescues the tfl1 mutant phenotype. Genet. Mol. Res. 14, 9306–9317. doi: 10.4238/2015.August.10.11
Zhang, X. M., Zhao, L., Larson-Rabin, Z., Li, D. Z., and Guo, Z. H. (2012). De Novo sequencing and characterization of the floral transcriptome of Dendrocalamus latiflorus (Poaceae: Bambusoideae). PLoS ONE 7:e42082. doi: 10.1371/journal.pone.0042082
Keywords: bamboo, flowering pathways, genes, drought, plant age, future research
Citation: Biswas P, Chakraborty S, Dutta S, Pal A and Das M (2016) Bamboo Flowering from the Perspective of Comparative Genomics and Transcriptomics. Front. Plant Sci. 7:1900. doi: 10.3389/fpls.2016.01900
Received: 28 July 2016; Accepted: 01 December 2016;
Published: 15 December 2016.
Edited by:
Stefan A. Rensing, University of Marburg, GermanyReviewed by:
Annette Becker, University of Giessen, GermanyMarcelo Carnier Dornelas, State University of Campinas, Brazil
Amanda E. Fisher, California State University, Long Beach, USA
Copyright © 2016 Biswas, Chakraborty, Dutta, Pal and Das. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Malay Das, bWFsYXkuZGJzQHByZXNpdW5pdi5hYy5pbg==
†These authors have contributed equally to this work.
 Prasun Biswas
Prasun Biswas Sukanya Chakraborty
Sukanya Chakraborty Smritikana Dutta
Smritikana Dutta Amita Pal
Amita Pal Malay Das
Malay Das