- The Institute of Vegetables and Flowers, Chinese Academy of Agricultural Sciences, Beijing, China
Cucumber seeds with shallow dormancy start to germinate in fruit that are harvested late. ABSCISIC ACID INSENSITIVE3 (ABI3), a transcription factor in the abscisic acid (ABA) signaling pathway, is one of the most important regulators in the transition from late embryogenesis to germination. Our analysis found a candidate cis-regulatory motif for cucumber BASIC PENTACYSTEINE (CsBPC) in the promoter of CsABI3. Yeast one-hybrid and chromatin immunoprecipitation (ChIP) assays showed that CsBPCs bound to the promoter of CsABI3. Examination of β-glucuronidase (GUS) activity driven by the CsABI3 promoter in transgenic Arabidopsis thaliana plants overexpressing CsBPCs and a Nicotiana benthamiana (tobacco) luciferase assay indicated that CsBPCs inhibited the expression of CsABI3. Transgenic plants overexpressing CsBPCs were constructed to confirm that CsBPCs participates in the control of seed germination. This study of the cucumber BPC-ABI3 pathway will help to explore and characterize the molecular mechanisms underlying seed germination and will provide necessary information for seed conservation in agriculture and forestry.
Introduction
Cucumber (Cucumis sativus L.), grown worldwide, is a model species of the Cucurbitaceae family. Cucumber is a typical example of fruit with shallow dormancy. During ripening, cucumber seeds start to germinate inside the fruit, also known as pre-harvest sprouting or vivipary, severely damaging the seed quantity and quality. In agriculture, vivipary is a phenomenon that results in worldwide losses in cereal crops, especially in humid regions (Farnsworth, 2000). Seeds undergo a period of dormancy until the proper environment for survival is present. Primary dormancy is generally used to describe an intact freshly-harvested viable seed that cannot germinate in favorable conditions (Bewley, 1997). Warm stratification, chilling (cold stratification), light, or hormones (including gibberellins; Kucera et al., 2005; Miransari and Smith, 2014) can release seeds from primary dormancy. Non-dormant seeds can also re-enter dormancy, known as secondary dormancy, to avoid unfavorable conditions such as season changes (Finkelstein et al., 2008). The breaking of seed dormancy to establish seedling growth is a critical step in the life of seed plants. The regulation of this developmental transition is not only necessary for plant survival but also important for agriculture and forestry; however, the regulatory mechanism underlying remains elusive.
Plant hormones such as Abscisic acid (ABA), gibberellins, brassinosteroids, ethylene, and cytokinins have long been known to participate in the control of seed dormancy and germination (Kucera et al., 2005; Finkelstein et al., 2008; Wang et al., 2011). ABA is a hormone that plays a vital role during the dormancy-to-germination transition (Finkelstein et al., 2002; Nambara and Marion-Poll, 2003; Chen et al., 2008; Kang et al., 2015). ABSCISIC ACID INSENSITIVE3 (ABI3) is an important transcription factor in the ABA signaling pathway (Finkelstein et al., 2008), participating in many different development stages including plastid development (Rohde et al., 2000), bud dormancy and flowering time (Rohde et al., 2002), lateral root development (Brady et al., 2003), desiccation tolerance (Khandelwal et al., 2010), and leaf development (Rohde et al., 1999). In particular, ABI3 is one of the most important regulators in the transition from late embryogenesis to germination (Parcy et al., 1994; Jones et al., 1997; Li and Foley, 1997; Zeng et al., 2003). The Arabidopsis thaliana mutant abi3 had decreased seed storage and a reduction in the level of the late embryogenesis abundant proteins Early methionine-labeled 1 (Em1) and Em6. It was reported that, via ABI3, ABA induced seed maturation behavior including the accumulation of storage substances, the acquisition of desiccation tolerance, and the imposition of dormancy (McCarty et al., 1989; Li and Foley, 1997; Nambara et al., 2000; Raz et al., 2001). Although the transcription level of ABI3 drops quickly after the breaking of seed dormancy (Park et al., 2011), ABI3 participates in the regulation of seed germination (Parcy et al., 1994; Nambara et al., 2000; Lopezmolina et al., 2002; Bassel et al., 2006; Feng et al., 2014). Given the range of functions that ABI3 is involved in, its expression is strictly regulated. LEAFY COTYLEDON1 (LEC1), LEC2, FUSCA 3 (FUS3), and ABI3 itself are the four important partially redundant transcription factors that regulate expression of ABI3 in seed development (To et al., 2006). ABI3 is also regulated by ABA during germination (Lopezmolina et al., 2002). During germination and early seedling development, the ABA pathway factor RELATED TO ABI3/VP1 (RAV1) represses the expression of ABI3, ABI4, and ABI5 (Feng et al., 2014). WRKY41 controls seed dormancy via ABI3 activation (Ding et al., 2014). During seed germination, ABI3 is repressed by the chromatin remodeling factor PICKLE (PKL; Perruc et al., 2007). The BES1-TPL-HDA19 repressor complex also participates in the ABA signaling pathway by controlling the epigenetic silencing of ABI3 during early seedling development (Ryu et al., 2014). At the protein level, ABI3 is targeted to 26S proteasome degradation by ABI3-INTERACTING PROTEIN 2 (AIP2; Zhang et al., 2005; Zeng et al., 2013), while protein-protein interactions between ABI3 and PIL5 are also responsible for the regulation of ABI3 (Park et al., 2011). Additionally, ABI3 is regulated by alternative splicing (Sugliani et al., 2010; Gao et al., 2013).
The BASIC PENTACYSTEINE/BARLEY B RECOMBINANT (BPC/BBR) family of transcription factors exists only in the plant kingdom (Sangwan and O'Brian, 2002; Santi et al., 2003; Kooiker et al., 2005; Berger et al., 2011; Simonini and Kater, 2014). While BPCs are known to participate in a wide range of developmental process (Monfared et al., 2011), the mechanism that determines how BPCs are specialized to specific developmental processes remains largely unknown. The barley BBR protein is involved in leaf and flower development, probably by regulating the expression of the homeotic gene Barley Knox3 (BKn3; Santi et al., 2003). The BPC1 protein participated in ovule and embryo development by regulating transcription factors LEC2, INNER NO OUTER (INO), and SEEDSTICK (STK; Meister et al., 2004; Kooiker et al., 2005; Berger et al., 2011). Class I BPCs regulate the inflorescence meristem via the regulation of SHOOT MERISTEMLESS (STM) and BREVIPEDICELLUS/KNAT1 (BP; Simonini and Kater, 2014).
While the mechanisms underlying seed dormancy have been studied extensively, less is known about the transition from dormancy to germination. ABI3 is one of the most important factors controlling seed germination. In this study, we used yeast one-hybrid and ChIP assays to show that CsBPCs act upstream of CsABI3. We further determined the negative relationship between the CsBPCs and CsABI3 using GUS staining and a luciferase assay. Moreover, the role of cucumber CsBPCs in seed germination was investigated. Analysis of the transcription factors controlling ABI3 during seed germination is useful for understanding the molecular mechanisms underlying seed germination and will provide information for seed conservation in agriculture and forestry.
Material and Methods
Plant Materials and Growth Conditions
Cucumber (C. sativus L.) was used in this study; line 9930 donated by Huang et al. (2009) was used for gene cloning to obtain the target construct and “Xintai Mici” was used for the construction of transgenic plants. Arabidopsis (ecotype Columbia) was used as the wild-type and the bpc1-1 bpc2 bpc4 bpc6 mutant was generously provided by Professor Charles S. Gasser (Monfared et al., 2011).
Cucumber seeds were germinated in darkness at 28°C and then seedlings with two true leaves were transplanted to a growth chamber under a 14-h light (350 μmol m−2 s−1) at 28°C/10-h dark at 18°C cycle. Arabidopsis seeds were surface-sterilized, sown on Murashige and Skoog (MS) medium plus 1.5% sucrose, and then 7-day-old-seedlings were transplanted to a growth chamber at 22°C under a 14-h light (60 μmol m−2 s−1) /10-h dark cycle.
Plant Transformation
To generate overexpression lines, the coding sequences of CsBPC1 and CsBPC3 were amplified and cloned into the BamHI/SpeI sites of the pCAMBIA-2300 vector to obtain the Pro35S:CsBPC1-GFP and Pro35S:CsBPC3-GFP constructs.
For Arabidopsis plant transformation, constructs were introduced into Agrobacterium tumefaciens GV3101. The floral dip method was then used to transform Arabidopsis plants with these constructs (Clough and Bent, 1998). The transgenic plants used in this study were selected on plates containing hygromycin or kanamycin and were confirmed by PCR amplification.
For cucumber plant transformation, constructs were transformed into the cucumber pure line Xintai Mici as previously described (Cheng et al., 2015). Agrobacterium LB4404 harboring the constructs were used for cucumber transformation. Briefly, cucumber seeds were treated with 70% alcohol for 20 s followed by 3% sodium hypochlorite solution for 7 min before being rinsed five times in sterile deionized water and placed on MS0 medium (MS plus 3% sucrose) for 2–3 days. The basal half of the cotyledon was then harvested and incubated with Agrobacterium harboring the target constructs for 15 min. The inoculated explants were transferred onto sterile filter paper and cultured on MS1 medium (MS0 medium plus 0.5 mg L−1 6-Benzylaminopurine and 1 mg L−1 ABA) for 2 d in the dark. The explants were incubated on MS1 medium for 15–20 d until the shoot was 1–1.5 cm long. The shoot was then transferred to MS2 (MS plus 200 mg L−1 cefotaxime) to develop the root. Integration of the construct in the regenerated plants was confirmed by PCR.
GFP Assays
The coding sequences of the CsBPCs were amplified by RT-PCR and cloned into the XhoI/EcoRI sites of the transient expression vector PBSK+-35s-EGFP, which contains the eGFP coding sequence under the control of the constitutive cauliflower mosaic virus 35S promoter, to generate a CsBPCs fusion protein with GFP at the C-terminus. The fibrillarin gene (Sun et al., 2011) was used as a nuclear target control. The resulting constructs were transfected into Arabidopsis mesophyll protoplasts according to the method of Yoo et al. (2007). Fluorescence analysis was performed using an OLYMPUS DP72 digital camera.
GUS Activity Assays
To generate ProCsABI3:GUS transgenic plants, the CsABI3 promoter sequence was amplified and cloned into the HindIII/NcoI sites upstream of the GUS coding sequence in the pCAMBIA1305 plasmid. The resulting construct was transformed into Arabidopsis via the Agrobacterium-mediated floral dip method (Clough and Bent, 1998). Plants were incubated in GUS staining solution containing 100 mM sodium phosphate buffer (pH 7.2), 0.1% Triton X-100, 10 mM EDTA, 0.5 mM K4Fe(CN)6·3H2O, 0.5 mM K3Fe(CN)6, and 1 mg L−1 X-Gluc at 37°C overnight. Images were taken on a dissecting microscope (OLYMPUS SZX2-ILLB) using a digital camera (Olympus). To quantify the GUS level, Arabidopsis seedlings were collected and ground in 100 μL of 1 × Cell Culture Lysis Reagent buffer (Promega), after which the extracts (5 μL) were mixed with 50 μL of luciferase assay substrate (Promega). Then, the extracts (5 μL) were incubated with 50 μL of 4-methylumbelliferyl β-D-glucuronide (MUG) substrate mix (10 mM Tris–HCl, pH 8.0, containing 1 mM MUG and 2 mM MgCl2) at 37°C for 30 min, after which the reaction was stopped by adding 945 μL of 0.2 M Na2CO3 and the fluorescence intensity was measured using a Modulus Luminometer/Fluorometer and an ultraviolet fluorescence optical kit (Turner Biosystems). GUS activity in each sample is normalized per unit tissue weight.
Yeast One-Hybrid
Yeast one-hybrid assays were performed according to the manufacturer's instructions (Clontech). The CsABI3 promoter sequence was fused to the KpnI/SalI sites in the pLacZi plasmid (Clontech) that was then linearized by digestion with NcoI and integrated into the genome of yeast strain YM4271 to obtain the integrated yeast clone harboring the ProCsABI3-pLacZi construct. To generate CsBPCs-PGAD, the CsBPCs coding sequences were amplified and cloned into EcoRI/XhoI sites downstream of the GAL4 activation domain coding sequence in PGADT7. CsBPCs-PGAD plasmids or the pGADT7 plasmid alone (negative control) were transformed into the yeast clone harboring the ProCsABI3-pLacZi construct. Transformants were grown on SD/-Ura-Leu dropout plates containing X-gal (5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside) for blue/white colony screening.
LUC Activity Assay for DNA-Protein Interactions in N. benthamiana Leaves
The reporter construct ProCsABI3:LUC was prepared using the primers listed in Table S1 and the effector constructs Pro35S:CsBPC1-GFP and 35S:CsBPC3-GFP as described above. To generate the ProCsABI3:LUC construct, the promoter sequence of CsABI3 was PCR-amplified and cloned into the HindIII/BamHI sites of the pCAMBIA1305 plasmid.
The reporter and effector constructs were transformed into Agrobacterium strain GV3101 to infiltrate N. benthamiana leaves as previously reported (Voinnet et al., 2003). The fluorescence intensity was measured under a Modulus Luminometer/Fluorometer using an ultraviolet fluorescence optical kit (Turner Biosystems). The relative LUC levels were expressed as the ratio of LUC to GUS.
qRT-PCR
Total RNA was isolated using a Qiagen RNeasy Plant Mini Kit and first-strand cDNA was synthesized using the SuperScript® III First-Strand Synthesis System (Invitrogen). PCR was then carried out using the gene-specific primers listed in Table S1 and SYBR PrimeScript Ready Mix (Takara) with an Mx3000p Real-time PCR System (Agilent, Stratagene) according to the manufacturer's instructions. Three biological replicates were included for each sample, and the expression levels were normalized to Tubulin.
ChIP
Cucumber seedlings containing the constructs Pro35S:CsBPC1-GFP and 35S:CsBPC3-GFP were generated as described above and were used to perform the ChIP assay as described by Wang et al. (2002). Transgenic cucumber chromatin was extracted from the GFP-tagged seedlings and then incubated with anti-GFP antibody. The immunoprecipitated DNA was analyzed by qRT-PCR with specific primers listed in Table S1.
Seed Germination Rate
To determine the germination rate of cucumber seeds, cucumber fruit were collected 35 d after pollination and the seeds were removed after 5 d of ripening. The seeds were put on Petri dishes (9.0 cm) containing a double layer of filter paper soaked with 5 mL sterilized distilled water and cultured at 28°C for 2 d in the dark. Seeds were considered to have germinated when 3 mm of the radicle had emerged.
For Arabidopsis, seeds collected at the same times were stored at room temperature for 40 d before being used. Seeds were sown on MS agar plates to observe germination. The plates were incubated at 23°C under continuous light for 8 d. Germination was scored based on radicle emergence (Liu et al., 2013).
Approximately 50 seeds were used for each replicate for both cucumber and Arabidopsis.
Results
Cucumber CsBPCs Influence Seed Germination
In this study, we found that seeds of the Arabidopsis quadruple mutant bpc1-1 bpc2 bpc4 bpc6 exhibited decreased germination rates (Figure 1A). This led us to question whether BPC homologs in cucumber could also affect germination. There are seven BPC proteins (BPC1-BPC7) encoded by the Arabidopsis genome. These have been categorized into three classes (Class I-III) based on their sequence similarity (Meister et al., 2004; Monfared et al., 2011). Database (http://www.icugi.org/cgi-bin/ICuGI/index.cgi) searches revealed that four BPC homologs exist in cucumber with highly conserved C-terminal regions. The cucumber BPC homologs were divided into two classes based on amino acid sequence alignments (Meister et al., 2004; Figure 2A) and phylogenic methods (Figure 2B). Class I included CsBPC1 (Csa5M092910) and CsBPC2 (Csa5M092920), while Class II included CsBPC3 (Csa7M007860) and CsBPC4 (Csa2M365700). Like their homologs in other plant species, nuclear fluorescence was observed in the nucleus of Arabidopsis protoplast cells expressing the CsBPCs-GFP fusion proteins (Figure S1). For the present study, two members were selected for further analysis, CsBPC1 and CsBPC3, representing Class I and Class II, respectively.
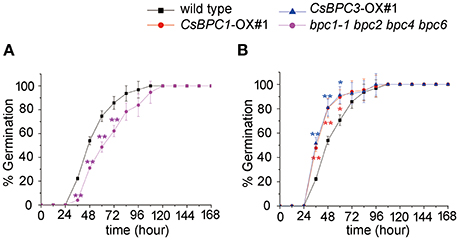
Figure 1. Germination rate of Arabidopsis seeds. (A) Seed germination rate of the bpc1-1 bpc2 bpc4 bpc6 mutant and wild-type Arabidopsis. (B) Seed germination rate of CsBPC1-OX#1, CsBPC3-OX#1, and wild-type Arabidopsis. Germination rates (%) of the seeds were analyzed at the indicated time points. The data represent means ± SD of three independent replicates with at least 50 seeds counted per replicate. Values that differed significantly from wild type are indicated. *p < 0.05 and **p < 0.01 by Bonferroni post-hoc test.
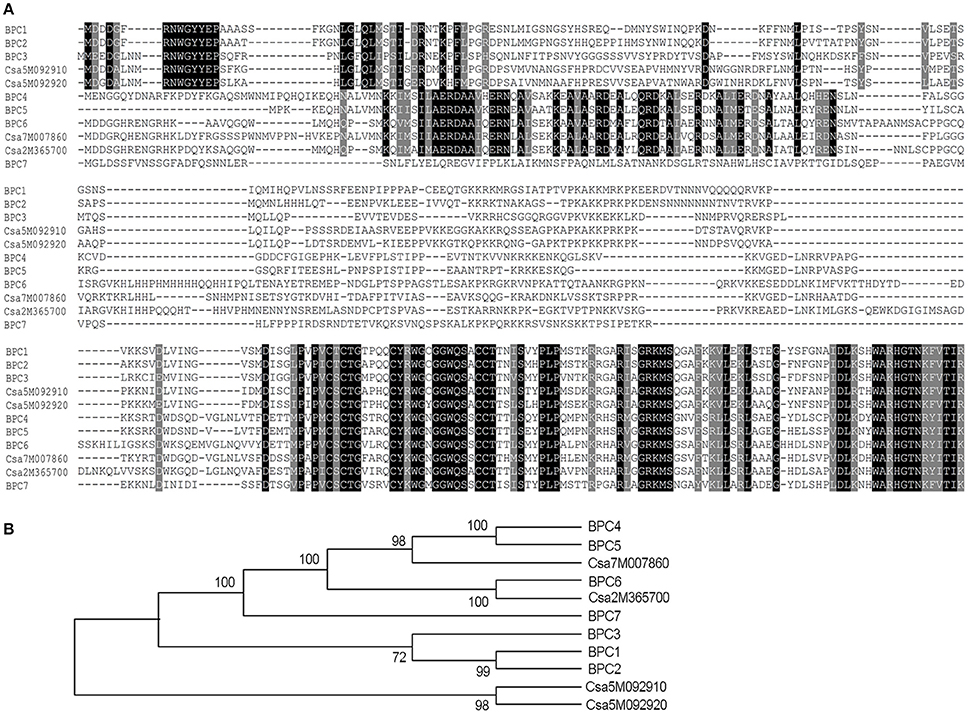
Figure 2. Arabidopsis and cucumber BPC homologs. (A) Arabidopsis and cucumber BPC homologs were aligned using CLUSTAL X. Conserved regions between single classes or the whole family are shown in white text on black, with conserved substitutions shaded in gray. Conserved cysteines are marked by an asterisk. (B) Phylogeny of the Arabidopsis and Cucumber BPC family. The 11 BPC proteins were classified into three groups using the Neighbor-Joining method (Saitou and Nei, 1987). The optimal tree with the sum of branch length = 2.95032244 is shown. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) are shown next to the branches. The evolutionary distances were computed using the Poisson correction method and are in the units of the number of amino acid substitutions per site. The analysis involved 11 amino acid sequences. All positions containing gaps and missing data were eliminated; there were a total of 199 positions in the final dataset. Evolutionary analyses were conducted in MEGA7 (Kumar et al., 2016).
To investigate whether the CsBPCs function in germination, Arabidopsis lines CsBPC1-OX and CsBPC3-OX overexpressing CsBPC1 and CsBPC3 (Figure S2), respectively, were constructed. The seed germination rate increased in CsBPC1-OX and CsBPC3-OX lines (CsBPC1-OX#1, CsBPC1-OX#4, CsBPC3-OX#1, and CsBPC3-OX#3) compared with wild-type Arabidopsis (Figure 1B and Figure S3). We next investigated if cucumber CsBPCs can rescue the late germination phenotype of the Arabidopsis quadruple mutant bpc1-1 bpc2 bpc4 bpc6. CsBPC1 and CsBPC3, driven by the cauliflower mosaic virus 35S promoter, was transferred into Arabidopsis bpc1-1 bpc2 bpc4 bpc6. The resulting transgenic lines CsBPC1 com#1, CsBPC1 com#2, CsBPC3 com#1, and CsBPC3 com#2 in which the CsBPCs were detected by RT-PCR (Figure 3A) were selected for the germination experiments. The results showed the germination rate of the complement lines was similar to that of wild type (Figure 3B).
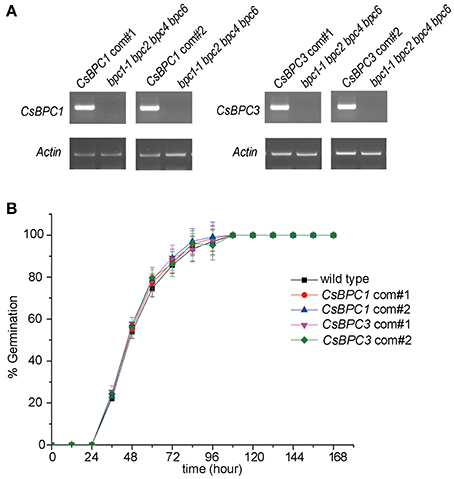
Figure 3. CsBPCs could complement the germination phenotype of the Arabidopsis bpc1-1 bpc2 bpc4 bpc6 mutant. (A) Expression of the CsBPCs were confirmed by RT-PCR. Genomic DNA of the bpc1-1 bpc2 bpc4 bpc6 mutant was used as negative control. The PCR primers used for detecting CsBPC1, CsBPC3, and Actin (control) genes are listed in Table S1. (B) Seed germination rates of CsBPC1 com#1, CsBPC1 com#2, CsBPC3 com#1, CsBPC3 com#3, and wild-type Arabidopsis. Germination rates (%) of the seeds were analyzed at the indicated time points. The data represent means ± SD of three independent replicates with at least 50 seeds counted per replicate. Values that differed significantly from wild type are indicated. Bonferroni post hoc test detected no significant difference in germination rates between the wild type and complementation of the bpc1-1 bpc2 bpc4 bpc6 mutant.
To determine whether the CsBPCs function similarly in cucumber, overexpression lines of the CsBPC1 and CsBPC3 in cucumber were constructed. We obtained 9 independent lines for CsBPC1-cuOX, and 9 lines for CsBPC3-cuOX. Quantitative RT-PCR (qRT-PCR) was used to measure the expression level of CsBPC1 and CsBPC3, respectively, in CsBPC1-cuOX and CsBPC3-cuOX lines. Two lines for each with the highest expression level were selected (two- to three- fold higher compared with wild-type cucumber; Figure S4). CsBPC1-cuOX#1, CsBPC1-cuOX#3, CsBPC3-cuOX#2, and CsBPC3-cuOX#8 were therefore used to assess germination rates. This experiment showed that the increase in germination rates of these overexpression lines was not significant (Figure 4 and Figure S5).
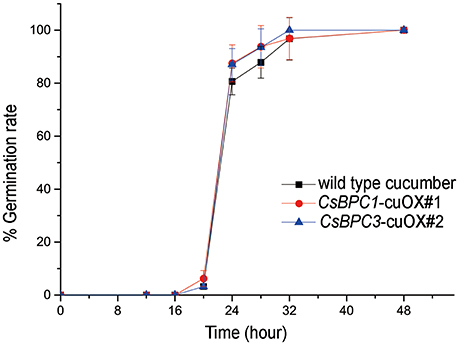
Figure 4. Germination rate of cucumber. The seed germination rates of wild-type cucumber and overexpression lines CsBPC1-cuOX#1 and CsBPC3-cuOX#2. Germination rates (%) were analyzed at the indicated time points. The data represent means ± SD of three independent replicates with at least 50 seeds counted for each replicate. Bonferroni post hoc test detected no difference between the wild-type and the overexpression lines.
CsBPCs Act Upstream of CsABI3
To explore the mechanism by which CsBPCs regulate germination rates, the gene sequence of the Arabidopsis germination-related gene ABI3 was used to search the cucumber genome. We found that the promoter sequence of a cucumber ABI3 homolog Csa7M336510.1, named CsABI3, showed high similarity (54.6%) to that of Arabidopsis ABI3. The promoter of CsABI3 contains three typical 9 bp DNA consensus sequences (RGARAGRRA) that comprise the BPC protein binding motif (Kooiker et al., 2005; Figure S6). The first cis-regulatory element is composed of 9 nucleotides (5′-AAAGAGAGA-3′) and is located 303 bp upstream of the translational start codon; the second cis-regulatory element, which consists of 16 nucleotides (5′-CTTCTTTCTCTTTCCT-3′), is located 279 bp upstream of the translational start point (inverted 5′-AGGAAAGAGAAAGAAG-3′); the third element, consisting of 9 nucleotides (5′-AAAGAAAGA-3′), is found 13 bp upstream of the translational start point. To determine whether CsABI3 is targeted by the CsBPCs, a yeast one-hybrid assay was performed which showed that all four CsBPCs interacted with the CsABI3 promoter (Figure 5A). Furthermore, a ChIP assay was performed to confirm the binding ability of the CsBPC1 and CsBPC3 to the CsABI3 promoter. Results showed that CsABI3 promoter fragments were enriched in anti-GFP samples compared with the ChIP samples prepared using Tubulin controls (Figure 5B).
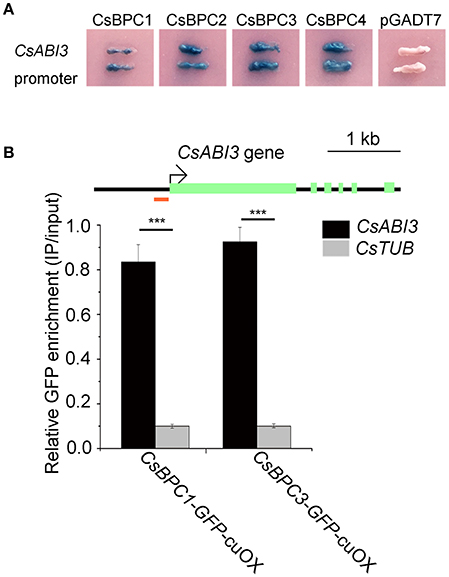
Figure 5. CsBPC proteins bind to the CsABI3 promoter. (A) Yeast one-hybrid assay of interactions between CsBPCs and the CsABI3 promoter. The CsABI3 promoter was fused to the pLacZi plasmid that was then linearized and integrated into the genome of the YM4271 yeast strain. Four CsBPC genes were cloned into pGADT7 to obtain the CsBPC1/2/3/4-pGAD plasmids. The CsBPCs-pGAD plasmids were transformed into a YM4271 clone harboring the CsABI3 promoter. The transformants were grown on SD/-Ura-Leu dropout plates containing X-gal (5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside) for white/blue screening. Unmodified PGADT7 was used as a negative control. (B) A ChIP assay showed that the CsBPCs bound to the CsABI3 promoter in vivo. GFP enrichment is expressed relative to the chromatin input. One pool comprising three independent transgenic lines was analyzed for each construct. Genomic DNA was obtained via ChIP from cucumber transgenic lines over-expressing CsBPC-GFP fusion proteins (CsBPC1/3-GFP-cuOX) using an anti-GFP antibody and was then subjected to qPCR analysis. The fragment used for qPCR is indicated by a red bar in the schematic diagram of the CsABI3 promoter. A control experiment with the Tubulin gene was used to establish the ChIP specificity. The mean values ± SD are representative of three independent experiments. ***p < 0.001 by Bonferroni post-hoc test.
CsBPCs Act as Negative Regulators of CsABI3
The interactions between the CsBPCs and the CsABI3 promoter were further investigated by evaluating the capacity of the CsBPCs to regulate the CsABI3 promoter. We developed a transient transcription assay system in which tobacco leaves were cotransfected with a 35S:CsBPC1/3 effector plasmid and a reporter construct carrying a fusion between the CsABI3 promoter and firefly luciferase (fLUC) that acted as a reporter for transcriptional activation (Figure 6A). The results showed that CsBPC1 and CsBPC3 resulted in a reduction in fLUC levels to about 10% of the controls, suggesting a negative relationship between the CsBPCs and CsABI3. Consistent with this negative relationship, CsABI3 mRNA levels were significantly lower in the cucumber CsBPCs overexpression lines (CsBPC1-cuOX#1 and CsBPC3-cuOX#2) compared with wild-type cucumber (Figure 6B). A similar result was found in the CsBPC1-cuOX#3 and CsBPC3-cuOX#8 lines (data not shown). Furthermore, overexpression of both CsBPC1 and CsBPC3 decreased the level of GUSexpression driven by the 1.0-kb fragment of the CsABI3 promoter (ProCsABI3:GUS) (Figures 6C,D).
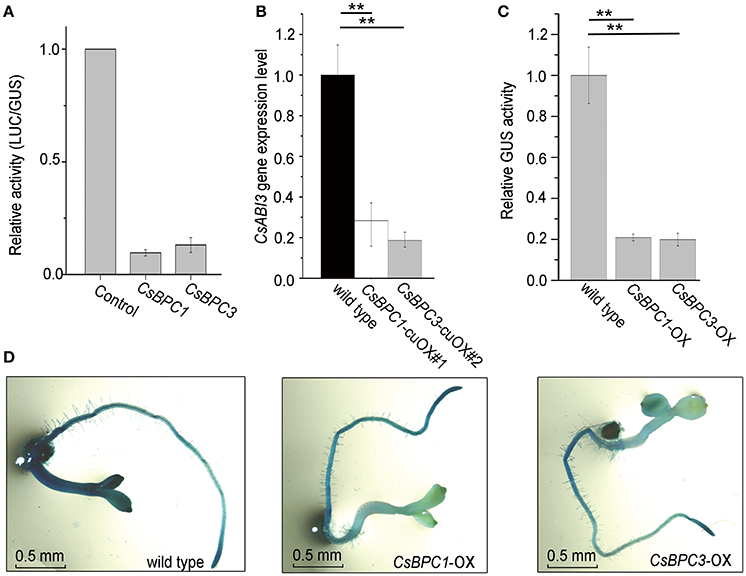
Figure 6. CsBPCs act as negative regulators of CsABI3. (A) LUC activity in a tobacco leaves transiently expressing the CsABI3 promoter co-infiltrated with empty control plasmid (CK) or plasmids containing the CsBPCs genes fused to the 35S promoter. Data shown are averages ± SD (n = 3). (B) CsABI3 mRNA levels in CsBPC1-cuOX#1, CsBPC3-cuOX#2, and wild-type cucumber seedlings grown for 4 d. Data shown are averages ± SD (n = 3). Three biological replicates were included for each experiment and 10 seedlings were included for each line. **p < 0.01 by Bonferroni post hoc test. (C) Quantitation of GUS activity of ProABI3:GUS in wild-type, CsBPC1-OX, and CsBPC3-OX Arabidopsis seedlings grown for 4 d. The mean values ± SD are representative of three independent experiments. **p < 0.01 by Bonferroni post hoc test. (D) Histochemical GUS staining for GUS activity shown in (C). A similar trend was observed in each of the three biological repeats, with representative images shown here.
The Interaction between CsBPCs and CsABI3 during Germination
To validate the regulatory role of CsBPCs on CsABI3 in seed germination, the level of GUS expression driven by the CsABI3 promoter was observed. CsABI3 promoter activity was high for 2 d after germination before dropping quickly over the following days to a barely detectable level 6 d after germination (Figure 7A). Conversely, the expression levels of the CsBPCs increased gradually in cucumber from 2–6 d after germination (Figure 7B). These results are consistent with CsABI3 expression being negatively regulated by the four CsBPC transcription factors in the germination and early development period.
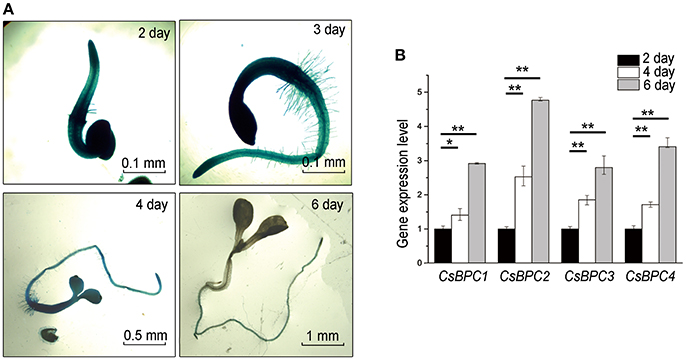
Figure 7. Expression of CsABI3 and the CsBPCs. (A) Expression of ProABI3:GUS in wild-type Arabidopsis seedlings grown for different periods of time. A similar trend was observed in each of three biological repeats, with representative images shown here. (B) Expression of the CsBPCs in cucumber seedlings grown for different periods of time. RNA was extracted and the expression levels of the CsBPCs were analyzed by qRT-PCR. Data shown are averages ± SD (n = 3). Three biological replicates were included for each experiment and 10 seedlings were included for each line. *p < 0.05 and **p < 0.01 by Bonferroni post hoc test.
ABI3 is an essential embryogenesis factor (Giraudat et al., 1992; Finkelstein et al., 1998) and is particularly important for late embryo development that takes place after embryonic cell division and morphogenesis are complete (Mccourt, 2003). The Arabidopsis seed maturation genes Em1, Em6, and the 2S ALBUMIN 3 (2S3) are all regulated by ABI3 (Parcy et al., 1994; Carles et al., 2002). Considering the interactions between the CsBPCs and CsABI3, it was reasonable for us to speculate that the CsBPCs might be involved in ABI3-mediated regulation during seed germination. To validate this possibility, we conducted a database search and identified a number of cucumber homologs of these Arabidopsis seed maturation genes including Csa3M078820.1, named cu2S3-1; Csa3M080340.1, named cu2S3-2; Csa4M311730.1, named CuEm1-1; Csa4M311230.1, named CuEm1-2; Csa4M311230.1, named cuEm6-1; and Csa4M311730.1, named cuEm6-2. qRT-PCR results showed that the expression of each of these genes was reduced in the cucumber CuBPC1/3 overexpression lines, strongly suggesting a role for the CsBPCs in the ABI3-mediated seed germination (Figure 8). Similar expression patterns were observed in the CsBPC1-cuOX#3 and CsBPC3-cuOX#8 lines (data not shown).
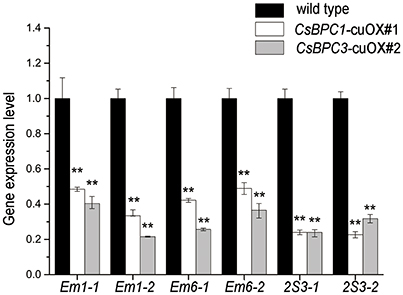
Figure 8. Overexpression of the CsBPCs reduces the expression of cucumber homologs of seed maturation genes. The expression levels of cucumber homologs of Em1, Em6, and 2S3 were examined in CsBPC1-cuOX#1, CsBPC3-cuOX#2, and wild-type cucumber plants. RNA was extracted from seedlings grown for 3 d and the expression levels were analyzed by qRT–PCR. Data shown are averages ± SD (n = 3). Three biological replicates were included for each experiment and 10 seedlings were included for each line. Values that differed significantly from wild type are indicated. ** p < 0.01 by Bonferroni post-hoc test.
Discussion
The breaking of seed dormancy and the beginning of autotrophic growth is a critical transition for seed plants. During this process, plants must monitor the environment and respond properly. The mechanism underlying seed dormancy and the transition to autotrophic growth requires further characterization. In this study, the role of cucumber CsBPCs during germination was investigated.
Cucumber, a well-known vegetable that is grown worldwide, displays shallow seed dormancy that often results in vivipary and causes serious damage to cucumber seed production and quality (Cao et al., 2016). Unfortunately, few published studies have focused on vivipary in cucumber. Previously, we found that the seeds of an Arabidopsis quadruple mutant with mutations in four BPC genes had significantly reduced germination rates (Figure 1A). To investigate whether the homologous cucumber CsBPCs have a similar function, CsBPC-overexpressing lines were generated showing that increased CsBPC expression resulted in greater seed germination rates in Arabidopsis (Figure 1B and Figure S3). Furthermore, cucumber CsBPCs functionally complement the germination phenotype of Arabidopsis bpc1-1 bpc2 bpc4 bpc6 mutants (Figure 3).
However, the effect of CsBPC overexpression on seed germination was not significant in cucumber (Figure 4 and Figure S5). While the enhancement of gene expression driven by the CaMV 35S promoter is efficient in Arabidopsis (Shin et al., 2007), the expression levels of the CsBPCs in transgenic cucumber lines increased by only two- to three- fold compared with those of wild-type cucumber (Figure S1) which is likely to result in the unaffected germination rate of these cucumber seeds. Thus, a more effective mrthod such as the CRISPR/Cas9 system must be established in the future to enhance gene expression levels in cucumber. These results show that cucumber CsBPCs are vital for the regulation of seed germination. Additionally, previous studies have shown that the ovules of the Arabidopsis BPC quadruple mutant bpc1-1 bpc2 bpc4 bpc6 had excess cell layers in the inner integument, suggesting that BPCs were also involved in seed development (Monfared et al., 2011).
Given that CsBPCs affect seed germination, we aimed to examine the molecular mechanism underlying their regulation of this process. Consistent with previous reports that Arabidopsis BPCs bind specifically to GA-rich domains (RGARAGRRA; Kooiker et al., 2005), three GA-rich domains were found in the CsABI3 promoter that may represent the BPC binding motif (Figure S6). In animals, such GAGA-motif containing DNA is bound by GAGA factors (GAFs) that play a role in both the repression and activation of homeobox genes through their interactions with Polycomb (PcG) and trithorax group (trxG) proteins, and thus contributes to a wide range of development events (Strutt et al., 1997; Francis and Kingston, 2001; Lehmann, 2004; Muller and Kassis, 2006; Schuettengruber et al., 2009; Berger and Dubreucq, 2012). In contrast to chromatin factors PcG and TrxG that are well-conserved between animals and plants, there are no homologs of the animal GAFs in plants (Strutt et al., 1997; Francis and Kingston, 2001; Lehmann, 2004; Muller and Kassis, 2006; Schuettengruber et al., 2009; Berger and Dubreucq, 2012). Given the functional similarities between BPCs and GAFs, BPCs were suggested to be GAF substitutes in plants (Berger et al., 2011; Berger and Dubreucq, 2012; Simonini et al., 2012). Nevertheless, a GAGA element in the LEC2 gene promoter bound by BPC1 was not required for the activity of PcG or TrxG, suggesting that BPCs might have other unknown functions (Berger et al., 2011).
ABI3, the candidate target gene of the CsBPC proteins, has been known as an essential regulator of seed dormancy and germination (Parcy et al., 1994; Nambara et al., 2000; Lopezmolina et al., 2002; Bassel et al., 2006; Tamura et al., 2006; Park et al., 2011; Feng et al., 2014). ABI3 regulates a substantial number of seed development-related genes and, therefore, functions primarily in the establishment of seed dormancy (Parcy et al., 1994; Nambara et al., 2000; Lopezmolina et al., 2002; Nakashima et al., 2006). During seed germination, ABI3 expression levels remain relatively high (Figure 7A; Park et al., 2011). Despite the fact that ABI3 gene expression is regulated by transcription factors like FUS2, LEC2, LEC3 (To et al., 2006), and RAV1 (Feng et al., 2014), WRKY41(Ding et al., 2014) is the only known upstream regulator that continues to regulate ABI3 expression at seed germination and over the following days (Ding et al., 2014).
The identification of unknown upstream regulators of ABI3 in the seed germination pathway is vital for us to understand the molecular mechanisms underlying this process. Here, we used yeast one-hybrid assays to demonstrate that the CsABI3 promoter was bound by the four cucumber CsBPCs, CsBPC1–4 (Figure 5A). Additionally, ChIP assays confirmed that CsBPC1 and CsBPC3 (representing Class I and Class II BPCs, respectively) could bind directly to the CsABI3 promoter (Figure 5B). Having shown that the cucumber CsBPCs act upstream of CsABI3, we explored the relationship between the CsABI3 and CsBPCs further. Analyses of the CsABI3 promoter and CsBPC proteins in vivo showed that the overexpression of either CsBPC1 or CsBPC3 resulted in decrease of ~90%in the activity of the CsABI3 promoter (Figure 6A). The regulatory functions of the CsBPCs were further confirmed in transgenic cucumber plants overexpressing CsBPC1 or CsBPC3 fused to GFP. The qRT–PCR analysis showed that expression of CsABI3 was reduced in these lines compared with wild-type cucumber (Figure 6B). Consistent with these results, GUS activity driven by the CsABI3 promoter was significantly reduced in Arabidopsis lines overexpressing CsBPC1 or CsBPC3 (Figures 6C,D). The negative relationship between the CsBPCs and the CsABI3 promoter, together with the ability of the BPCs to bind to the CsABI3 promoter, suggests that CsBPCs might be involved in the negative regulation of CsABI3.
Having shown that cucumber CsBPCs are involved in seed germination and that CsBPCs act upstream of CsABI3, it is reasonable for us to speculate that the CsBPCs affect seed germination via the CsABI3 pathway.
Previous studies have shown that Em1, Em6, and 2S3 are representative seed maturation genes that are controlled by the ABA pathway through ABI3 (Parcy et al., 1994). The qRT-PCR assays showed that, in 3-day-old transgenic cucumber lines, the overexpression of either CsBPC1 or CsBPC3 significantly reduced the expression levels of Em1, Em6, and 2S3 homologs in cucumber (Figure 8), supporting a role for CsBPCs upstream of CsABI3 in seed germination. It has also been suggested that there are more putative BPC target genes in the plant kingdom. Sequence analysis identified more than 12,000 Arabidopsis genes containing more than one GA-rich stretch within their regulatory regions (Santi et al., 2003; Berendzen et al., 2006; Deng et al., 2013; Hecker et al., 2015), suggesting that BPCs might act on a variety of target genes in diverse developmental processes. The significance of the relatively high number of GAGA-binding factors in the plant kingdom indicates the importance of the BPC family in growth and development processes.
Moreover, a previous study showed that LEC2, a positive regulator of ABI3, controlled most aspects of seed maturation (To et al., 2006). Arabidopsis BPCs were reported to act redundantly to regulate LEC2 expression through its specific GAGA cis-regulatory motif (Berger et al., 2011). This implies that the BPCs might also be involved in the ABI3 pathway indirectly via LEC2 regulation. In the present study, we describe four cucumber CsBPC transcription factors, CsBPC1, CsBPC2, CsBPC3, and CsBPC4, that are direct upstream regulators of CsABI3. Furthermore, we suggest that by regulating CsABI3, CsBPCs control seed germination. Further, investigation is needed to characterize the precise mechanism regulating the CsBPC-CsABI3 pathway controlling seed germination.
Author Contributions
Substantial contributions to the conception or design of the work; or the acquisition, analysis, or interpretation of data for the work (YM, YML, LB, SL, CH, YY, XY, and YSL); Drafting the work or revising it critically for important intellectual content (YM, YML, LB, SL, CH, YY, XY, and YSL); Final approval of the version to be published (YM, YML, LB, SL, CH, YY, XY, and YSL); Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved (YM, YML, LB, SL, CH, YY, XY, and YSL).
Funding
Sincere thanks for the funding support provided by the Earmarked fund for Modern Agro-industry Technology Research System (CARS-25-C-01), Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences (CAAS-ASTIP-IVFCAAS) and the support by the Key Laboratory of Horticultural Crop Biology and Germplasm Innovation, Ministry of Agriculture, China. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We would like to thank Professor Charles S. Gasser for providing the bpc1-1 bpc2 bpc4 bpc6 mutant and Professor Huang for providing cucumber line 9930.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2017.00459/full#supplementary-material
References
Bassel, G. W., Mullen, R. T., and Bewley, J. D. (2006). ABI3 expression ceases following, but not during, germination of tomato and Arabidopsis seeds. J. Exp. Bot. 57, 1291–1297. doi: 10.1093/jxb/erj101
Berendzen, K. W., Stuber, K., Harter, K., and Wanke, D. (2006). Cis-motifs upstream of the transcription and translation initiation sites are effectively revealed by their positional disequilibrium in eukaryote genomes using frequency distribution curves. BMC Bioinform. 7:522. doi: 10.1186/1471-2105-7-522
Berger, N., and Dubreucq, B. (2012). Evolution goes GAGA: GAGA binding proteins across kingdoms. Biochim. Biophys. Acta 1819, 863–868. doi: 10.1016/j.bbagrm.2012.02.022
Berger, N., Dubreucq, B., Roudier, F., Dubos, C., and Lepiniec, L. (2011). Transcriptional regulation of Arabidopsis LEAFY COTYLEDON2 involves RLE, a cis-element that regulates trimethylation of histone H3 at lysine-27. Plant Cell 23, 4065–4078. doi: 10.1105/tpc.111.087866
Bewley, J. D. (1997). Seed germination and dormancy. Plant Cell 9, 1055–1066. doi: 10.1105/tpc.9.7.1055
Brady, S. M., Sarkar, S. F., Bonetta, D., and McCourt, P. (2003). The ABSCISIC ACID INSENSITIVE 3 (ABI3) gene is modulated by farnesylation and is involved in auxin signaling and lateral root development in Arabidopsis. Plant J. 34, 67–75. doi: 10.1046/j.1365-313X.2003.01707.x
Cao, M., Guan, W., Wang, H., Deng, Q., Li, S., and Yang, R. (2016). Research progress of cucumber seeds pre-harvest sprouting. Tianjin Agric. Sci. 22, 127–130.
Carles, C., Bies-Etheve, N., Aspart, L., Leon-Kloosterziel, K. M., Koornneef, M., Echeverria, M., et al. (2002). Regulation of Arabidopsis thaliana Em genes: role of ABI5. Plant J. 30, 373–383. doi: 10.1046/j.1365-313X.2002.01295.x
Chen, S., Kuo, S., and Chien, C. (2008). Roles of gibberellins and abscisic acid in dormancy and germination of red bayberry (Myrica rubra) seeds. Tree Physiol. 28, 1431–1439. doi: 10.1093/treephys/28.9.1431
Cheng, J., Wang, Z., Yao, F., Gao, L., Ma, S., Sui, X., et al. (2015). Down-Regulating CsHT1, a cucumber pollen-specific hexose transporter, inhibits pollen germination, tube growth, and seed development. Plant Physiol. 168, 635–647. doi: 10.1104/pp.15.00290
Clough, S. J., and Bent, A. F. (1998). Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743. doi: 10.1046/j.1365-313x.1998.00343.x
Deng, W., Buzas, D. M., Ying, H., Robertson, M., Taylor, J., Peacock, W. J., et al. (2013). Arabidopsis polycomb repressive complex 2 binding sites contain putative GAGA factor binding motifs within coding regions of genes. BMC Genomics 14:593. doi: 10.1186/1471-2164-14-593
Ding, Z., Yan, J., Li, G., Wu, Z., Zhang, S., and Zheng, S. (2014). WRKY41 controls Arabidopsis seed dormancy via direct regulation of ABI3 transcript levels not downstream of ABA. Plant J. 79, 810–823. doi: 10.1111/tpj.12597
Farnsworth, E. (2000). The ecology and physiology of viviparous and recalcitrant seeds. Annu. Rev. Ecol. Syst. 31, 107–138. doi: 10.1146/annurev.ecolsys.31.1.107
Feng, C. Z., Chen, Y., Wang, C., Kong, Y. H., Wu, W. H., and Chen, Y. F. (2014). Arabidopsis RAV1 transcription factor, phosphorylated by SnRK2 kinases, regulates the expression of ABI3, ABI4, and ABI5 during seed germination and early seedling development. Plant J. 80, 654–668. doi: 10.1111/tpj.12670
Finkelstein, R. R., Gampala, S. S., and Rock, C. D. (2002). Abscisic acid signaling in seeds and seedlings. Plant Cell 14(Suppl), S15–S45. doi: 10.1105/tpc.010441
Finkelstein, R. R., Reeves, W., Ariizumi, T., and Steber, C. M. (2008). Molecular aspects of seed dormancy. Annu. Rev. Plant Biol. 59, 387–415. doi: 10.1146/annurev.arplant.59.032607.092740
Finkelstein, R. R., Wang, M. L., Lynch, T. J., Rao, S., and Goodman, H. M. (1998). The arabidopsis abscisic acid response locus ABI4 encodes an APETALA2 domain protein. Plant Cell 10, 1043–1054. doi: 10.2307/3870689
Francis, N. J., and Kingston, R. E. (2001). Mechanisms of transcriptional memory. Nat. Rev. Mol. Cell Biol. 2, 409–421. doi: 10.1038/35073039
Gao, Y., Liu, J., Zhang, Z., Sun, X., Zhang, N., Fan, J., et al. (2013). Functional characterization of two alternatively spliced transcripts of tomato ABSCISIC ACID INSENSITIVE3 (ABI3) gene. Plant Mol. Biol. 82, 131–145. doi: 10.1007/s11103-013-0044-1
Giraudat, J., Hauge, B. M., Valon, C., Smalle, J., Parcy, F., and Goodman, H. M. (1992). Isolation of the Arabidopsis ABI3 gene by positional cloning. Plant Cell 4, 1251–1261. doi: 10.1105/tpc.4.10.1251
Hecker, A., Brand, L. H., Peter, S., Simoncello, N., Kilian, J., Harter, K., et al. (2015). The Arabidopsis GAGA-Binding Factor BASIC PENTACYSTEINE6 Recruits the POLYCOMB-REPRESSIVE COMPLEX1 Component LIKE HETEROCHROMATIN PROTEIN1 to GAGA DNA Motifs. Plant Physiol. 168, 1013–1024. doi: 10.1104/pp.15.00409
Huang, S., Li, R., Zhang, Z., Li, L., Gu, X., Fan, W., et al. (2009). The genome of the cucumber, Cucumis sativus L. Nat. Genet. 41, 1275–1281. doi: 10.1038/ng.475
Jones, H. D., Peters, N. C., and Holdsworth, M. J. (1997). Genotype and environment interact to control dormancy and differential expression of the VIVIPAROUS 1 homologue in embryos of Avena fatua. Plant J. 12, 911–920. doi: 10.1046/j.1365-313X.1997.12040911.x
Kang, J., Yim, S., Choi, H., Kim, A., Lee, K. P., Lopezmolina, L., et al. (2015). Abscisic acid transporters cooperate to control seed germination. Nat. Commun. 6, 8113–8113. doi: 10.1038/ncomms9113
Khandelwal, A., Cho, S. H., Marella, H., Sakata, Y., Perroud, P. F., Pan, A., et al. (2010). Role of ABA and ABI3 in desiccation tolerance. Science 327:546. doi: 10.1126/science.1183672
Kooiker, M., Airoldi, C. A., Losa, A., Manzotti, P. S., Finzi, L., Kater, M. M., et al. (2005). BASIC PENTACYSTEINE1, a GA binding protein that induces conformational changes in the regulatory region of the homeotic Arabidopsis gene SEEDSTICK. Plant Cell 17, 722–729. doi: 10.1105/tpc.104.030130
Kucera, B., Cohn, M. A., and Leubnermetzger, G. (2005). Plant hormone interactions during seed dormancy release and germination. Seed Sci. Res. 15, 281–307. doi: 10.1079/SSR2005218
Kumar, S., Stecher, G., and Tamura, K. (2016). MEGA7: molecular Evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874. doi: 10.1093/molbev/msw054
Lehmann, M. (2004). Anything else but GAGA: a nonhistone protein complex reshapes chromatin structure. Trends Genet. 20, 15–22. doi: 10.1016/j.tig.2003.11.005
Li, B., and Foley, M. E. (1997). Genetic and molecular control of seed dormancy. Trends Plant Sci. 2, 384–389. doi: 10.1016/S1360-1385(97)90053-4
Liu, X., Zhang, H., Zhao, Y., Feng, Z., Li, Q., Yang, H. Q., et al. (2013). Auxin controls seed dormancy through stimulation of abscisic acid signaling by inducing ARF-mediated ABI3 activation in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 110, 15485–15490. doi: 10.1073/pnas.1304651110
Lopezmolina, L., Mongrand, S., Mclachlin, D., Chait, B., and Chua, N. (2002). ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J. 32, 317–328. doi: 10.1046/j.1365-313X.2002.01430.x
McCarty, D. R., Carson, C. B., Stinard, P. S., and Robertson, D. S. (1989). Molecular analysis of viviparous-1: an abscisic acid-insensitive mutant of maize. Plant Cell 1, 523–532. doi: 10.1105/tpc.1.5.523
Mccourt, P. (2003). Genetic analysis of hormone signaling. Annu. Rev. Plant Biol. 50, 219–243. doi: 10.1146/annurev.arplant.50.1.219
Meister, R. J., Williams, L. A., Monfared, M. M., Gallagher, T. L., Kraft, E. A., Nelson, C. G., et al. (2004). Definition and interactions of a positive regulatory element of the Arabidopsis INNER NO OUTER promoter. Plant J. 37, 426–438. doi: 10.1046/j.1365-313X.2003.01971.x
Miransari, M., and Smith, D. (2014). Plant hormones and seed germination. Environ. Exp. Bot. 99, 110–121. doi: 10.1016/j.envexpbot.2013.11.005
Monfared, M. M., Simon, M. K., Meister, R. J., Roig-Villanova, I., Kooiker, M., Colombo, L., et al. (2011). Overlapping and antagonistic activities of BASIC PENTACYSTEINE genes affect a range of developmental processes in Arabidopsis. Plant J. 66, 1020–1031. doi: 10.1111/j.1365-313X.2011.04562.x
Muller, J., and Kassis, J. A. (2006). Polycomb response elements and targeting of Polycomb group proteins in Drosophila. Curr. Opin. Genet. Dev. 16, 476–484. doi: 10.1016/j.gde.2006.08.005
Nakashima, K., Fujita, Y., Katsura, K., Maruyama, K., Narusaka, Y., Seki, M., et al. (2006). Transcriptional regulation of ABI3- and ABA-responsive genes including RD29B and RD29A in seeds, germinating embryos, and seedlings of Arabidopsis. Plant Mol. Biol. 60, 51–68. doi: 10.1007/s11103-005-2418-5
Nambara, E., Hayama, R., Tsuchiya, Y., Nishimura, M., Kawaide, H., Kamiya, Y., et al. (2000). The role of ABI3 and FUS3 loci in Arabidopsis thaliana on phase transition from late embryo development to germination. Dev. Biol. 220, 412–423. doi: 10.1006/dbio.2000.9632
Nambara, E., and Marion-Poll, A. (2003). ABA action and interactions in seeds. Trends Plant Sci. 8, 213–217. doi: 10.1016/S1360-1385(03)00060-8
Parcy, F., Valon, C., Raynal, M., Gaubier-Comella, P., Delseny, M., and Giraudat, J. (1994). Regulation of gene expression programs during Arabidopsis seed development: roles of the ABI3 locus and of endogenous abscisic acid. Plant Cell 6, 1567–1582. doi: 10.1105/tpc.6.11.1567
Park, J., Lee, N., Kim, W., Lim, S., and Choi, G. (2011). ABI3 and PIL5 collaboratively activate the expression of SOMNUS by directly binding to its promoter in imbibed Arabidopsis seeds. Plant Cell 23, 1404–1415. doi: 10.1105/tpc.110.080721
Perruc, E., Kinoshita, N., and Lopez-Molina, L. (2007). The role of chromatin-remodeling factor PKL in balancing osmotic stress responses during Arabidopsis seed germination. Plant, J. 52, 927–936. doi: 10.1111/j.1365-313X.2007.03288.x
Raz, V., Bergervoet, J. H., and Koornneef, M. (2001). Sequential steps for developmental arrest in Arabidopsis seeds. Development 128, 243–252.
Rohde, A., De Rycke, R., Beeckman, T., Engler, G., Van Montagu, M., and Boerjan, W. (2000). ABI3 affects plastid differentiation in dark-grown Arabidopsis seedlings. Plant Cell 12, 35–52. doi: 10.1105/tpc.12.1.35
Rohde, A., Prinsen, E., De Rycke, R., Engler, G., Van Montagu, M., and Boerjan, W. (2002). PtABI3 impinges on the growth and differentiation of embryonic leaves during bud set in poplar. Plant Cell 14, 1885–1901. doi: 10.1105/tpc.003186
Rohde, A., Van Montagu, M., and Boerjan, W. (1999). The ABSCISIC ACID-INSENSITIVE 3 (ABI3) gene is expressed during vegetative quiescence processes in Arabidopsis. Plant Cell Environ. 22, 261–270. doi: 10.1046/j.1365-3040.1999.00428.x
Ryu, H., Cho, H., Bae, W., and Hwang, I. (2014). Control of early seedling development by BES1/TPL/HDA19-mediated epigenetic regulation of ABI3. Nat. Commun. 5:4138. doi: 10.1038/ncomms5138
Saitou, N., and Nei, M. (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425.
Sangwan, I., and O'Brian, M. R. (2002). Identification of a soybean protein that interacts with GAGA element dinucleotide repeat DNA. Plant Physiol. 129, 1788–1794. doi: 10.1104/pp.002618
Santi, L., Wang, Y., Stile, M. R., Berendzen, K., Wanke, D., Roig, C., et al. (2003). The GA octodinucleotide repeat binding factor BBR participates in the transcriptional regulation of the homeobox gene Bkn3. Plant J. 34, 813–826. doi: 10.1046/j.1365-313X.2003.01767.x
Schuettengruber, B., Ganapathi, M., Leblanc, B., Portoso, M., Jaschek, R., Tolhuis, B., et al. (2009). Functional anatomy of polycomb and trithorax chromatin landscapes in Drosophila embryos. PLoS Biol. 7:e13. doi: 10.1371/journal.pbio.1000013
Shin, R., Burch, A. Y., Huppert, K. A., Tiwari, S. B., Murphy, A. S., Guilfoyle, T. J., et al. (2007). The Arabidopsis transcription factor MYB77 modulates auxin signal transduction. Plant Cell 19, 2440–2453. doi: 10.1105/tpc.107.050963
Simonini, S., and Kater, M. M. (2014). Class I BASIC PENTACYSTEINE factors regulate HOMEOBOX genes involved in meristem size maintenance. J. Exp. Bot. 65, 1455–1465. doi: 10.1093/jxb/eru003
Simonini, S., Roig-Villanova, I., Gregis, V., Colombo, B., Colombo, L., and Kater, M. M. (2012). Basic pentacysteine proteins mediate MADS domain complex binding to the DNA for tissue-specific expression of target genes in Arabidopsis. Plant Cell 24, 4163–4172. doi: 10.1105/tpc.112.103952
Strutt, H., Cavalli, G., and Paro, R. (1997). Co-localization of Polycomb protein and GAGA factor on regulatory elements responsible for the maintenance of homeotic gene expression. EMBO J. 16, 3621–3632. doi: 10.1093/emboj/16.12.3621
Sugliani, M., Brambilla, V., Clerkx, E. J., Koornneef, M., and Soppe, W. J. (2010). The conserved splicing factor SUA controls alternative splicing of the developmental regulator ABI3 in Arabidopsis. Plant Cell 22, 1936–1946. doi: 10.1105/tpc.110.074674
Sun, X., Feng, P., Xu, X., Guo, H., Ma, J., Chi, W., et al. (2011). A chloroplast envelope-bound PHD transcription factor mediates chloroplast signals to the nucleus. Nat. Commun. 2:477. doi: 10.1038/ncomms1486
Tamura, N., Yoshida, T., Tanaka, A., Sasaki, R., Bando, A., Toh, S., et al. (2006). Isolation and characterization of high temperature-resistant germination mutants of Arabidopsis thaliana. Plant Cell Physiol. 47, 1081–1094. doi: 10.1093/pcp/pcj078
To, A., Valon, C., Savino, G., Guilleminot, J., Devic, M., Giraudat, J., et al. (2006). A network of local and redundant gene regulation governs Arabidopsis seed maturation. Plant Cell 18, 1642–1651. doi: 10.1105/tpc.105.039925
Voinnet, O., Rivas, S., Mestre, P., and Baulcombe, D. (2003). An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J. 33, 949–956. doi: 10.1046/j.1365-313X.2003.01676.x
Wang, H., Tang, W. N., Zhu, C., and Perry, S. E. (2002). A chromatin immunoprecipitation (ChIP) approach to isolate genes regulated by AGL15, a MADS domain protein that preferentially accumulates in embryos. Plant J. 32, 831–843. doi: 10.1046/j.1365-313X.2002.01455.x
Wang, Y., Li, L., Ye, T., Zhao, S., Liu, Z., Feng, Y. Q., et al. (2011). Cytokinin antagonizes ABA suppression to seed germination of Arabidopsis by downregulating ABI5 expression. Plant J. 68, 249–261. doi: 10.1111/j.1365-313X.2011.04683.x
Yoo, S. D., Cho, Y. H., and Sheen, J. (2007). Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572. doi: 10.1038/nprot.2007.199
Zeng, Y., Raimondi, N., and Kermode, A. R. (2003). Role of an ABI3 homologue in dormancy maintenance of yellow-cedar seeds and in the activation of storage protein and Em gene promoters. Plant Mol. Biol. 51, 39–49. doi: 10.1023/A:1020762304937
Zeng, Y., Zhao, T., and Kermode, A. (2013). An ABI3-interactor of conifers responds to multiple hormones. Plant Signal Behav. 8:e26225. doi: 10.4161/psb.26225
Keywords: CsBPC, CsABI3, germination, cucumber, transcription factors
Citation: Mu Y, Liu Y, Bai L, Li S, He C, Yan Y, Yu X and Li Y (2017) Cucumber CsBPCs Regulate the Expression of CsABI3 during Seed Germination. Front. Plant Sci. 8:459. doi: 10.3389/fpls.2017.00459
Received: 23 December 2016; Accepted: 16 March 2017;
Published: 03 April 2017.
Edited by:
Keqiang Wu, National Taiwan University, TaiwanReviewed by:
Chenlong Li, Agriculture and Agriculture-Food Canada, CanadaWim Soppe, Max Planck Institute for Plant Breeding Research (MPG), Germany
Copyright © 2017 Mu, Liu, Bai, Li, He, Yan, Yu and Li. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xianchang Yu, eGN5dTE5NjJAMTYzLmNvbQ==
Yansu Li, bGl5YW5zdUBjYWFzLmNu
 Ying Mu
Ying Mu Shuzhen Li
Shuzhen Li Yansu Li
Yansu Li