- 1Key Laboratory for Biological Control of the Ministry of Agriculture, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing, China
- 2State Key Laboratory of Agrobiotechnology, China Agricultural University, Beijing, China
Abiotic stresses, especially drought, seriously threaten cereal crops yields and quality. In this study, we observed that the rice plants of overexpression the Alternariatenuissima PeaT1 gene showed enhanced drought stress tolerance and increased the survival rate following a drought treatment. In PeaT1-overexpressing (PeaT1OE) plants, abscisic acid and chlorophyll content significantly increased, while the malondialdehyde (MDA) content decreased compared with the wild-type plants. Additionally, we confirmed that the transcript levels of drought-responsive genes, including OsAM1, OsLP2, and OsDST, were prominently lower in the PeaT1OE plants. In contrast, expression levels of genes encoding positive drought stress regulators including OsSKIPa, OsCPK9, OsNAC9, OSEREBP1, and OsTPKb were upregulated in PeaT1OE plants. Furthermore, combing the yeast two-hybrid assay, we found that PeaT1 could interact with amyo-inositol oxygenase (OsMIOX), which was verified by pull-down assay. Interestingly, OsMIOX was highly expressed in PeaT1OE plants during the drought treatment. Additionally, the OsMIOX-GFP fusion protein co-localized with the endoplasmic reticulum (ER) marker in tobacco protoplasts, suggesting OsMIOX performs its function in ER. Therefore, our results are useful for elucidating the molecular mechanism underlying the improvement of drought tolerance by PeaT1.
Introduction
Drought is one of main abiotic stresses that negatively influences plant development, growth, and seed production (Huang et al., 2009). Plants have evolved diverse mechanisms to respond to environmental stresses. Therefore, details regarding to the molecular mechanism regulating plant responses to stresses may be useful for improving plant resistance to abiotic stresses, especially water deficiency (Huang et al., 2009). Previous studies on Arabidopsis thaliana and rice revealed that complex plant responses to drought stress are mediated by several factors, including abscisic acid (ABA) (Ma et al., 2009; Park et al., 2009), reactive oxygen species (Pei et al., 2000; Kwak et al., 2003), and transcription factors (Huang et al., 2009; Fujita et al., 2011). However, biological functions of well known stress-responsive genes remain uncharacterized. Additionally, there are likely many plant genes associated with drought tolerance yet to be identified.
Abscisic acid, which is an important plant hormone, mediates some aspects of the plant life cycle such as seed dormancy, flowering, and fruit ripening. It is also crucial for plant responses to diverse abiotic stresses such as drought, cold, and salinity. Furthermore, ABA regulates stomatal closure to help plants adapt to water deficiency (Zhu, 2002; Fan et al., 2004). In response to abiotic stresses, ABA content immediately increases to protect plants from damages. When plants face with drought stress, ABA signal pathway becomes active and involves in ABA receptors PYR/PYL/RCAR (Park et al., 2009) as well as two phosphatase/kinase enzyme pairs, PP2Cs (Nishimura et al., 2009) and SnRK2s (Gonzalez-Guzman et al., 2012), with opposite functions. In the end, a number of transcription factors under the control of SnRK2s activate ABA dependent genes expression such as ABI5 and ABRE binding factors/ABRE binding proteins (ABF/AREBs), which specifically bind to promoters containing ABA-responsive elements (ABREs) (Antoni et al., 2011), and regulate guard cell channel activities (Lee et al., 2013), resulting in drought-resistant responses (Kim, 2014).
Elicitors isolated from several fungal and bacterial pathogens include proteins, peptides, glycoproteins, lipids, and oligosaccharides (Nürnberger, 1999). Elicitors, which function as signaling molecules, were originally identified in studies that focused on host specificity and disease development (Paré et al., 2005). Harpin, the first reported elicitor from the bacterium Erwinia amylovora (Wei et al., 1992; He et al., 1993), could activate the hypersensitive response and growth systems in several crops. The PeaT1 elicitor isolated from Alternaria tenuissima induces the systemic accumulation of the pathogenesis-related proteins PR-1a and PR-1b, which are markers for systemic acquired resistance (Zhang W. et al., 2011). Elicitors may also influence plant resistance to environmental stresses, especially drought. For example, PebC1, which was isolated from Botrytis cinerea, induces plant drought tolerance (Zhang et al., 2010). Other studies have revealed that PeaT1 could promote wheat and cotton growth and development (Wang et al., 2011; Tang et al., 2012). Additionally, rice and wheat leaves sprayed with exogenous PeaT1 were observed to exhibit increased drought resistance (Liu et al., 2009; Wang et al., 2011). Furthermore, overexpression Harpin-encoding hrf1 gene in rice enhances drought tolerance (Zhang L. et al., 2011).
Myo-inositol oxygenase (MIOX) is considered as a unique monooxygenase that catalyzes the transfer of myo-inositol to D-glucuronic acid (D-GlcUA), which is an important sugar precursor for plant cell walls (Kanter et al., 2005). Myo-inositol oxygenase balances the myo-inositol content and is essential for the synthesis of some low molecular weight compounds in plants (Smirnoff et al., 2001). Recently, OsMIOX expression was observed to be induced by drought, H2O2, salt, cold, and ABA. Additionally, overexpression the OsMIOX considerably improves plant growth under mannitol stress conditions (Duan et al., 2012).
Our lab previously generated PeaT1-overexpressing rice plants (PeaT1OE) (Sheng et al., 2011). In the current study, we analyzed the resistance of PeaT1OE plants to drought stress and using yeast two-hybrid and pull-down assays, found that PeaT1surely interacts with OsMIOX. Thus, our resultss might clarify PeaT1effects on drought tolerance regarding to survival rate, ABA content, and phenotype.
Materials and Methods
Drought Tolerance Analysis
In our previous study, an elicitor peaT1 gene was introduced into pCAMBIA2300 vector under the control of CaMV 35S promoter and then transformed into the Nipponbare rice (Sheng et al., 2011). Drought tolerance of transgenic plants was examined by germinating T2 transgenic plants. Seeds were submerged in water at room temperature for 48 h, and then allowed to germinate for 48 h at 37°C in a growth chamber. After 4 days, seedlings were transplanted to containers filled with soil, and then incubated in a greenhouse at 24–30°C and 50–60% relative humidity. Wild-type (WT) Oryza sativa L. japonica cv Nipponbare seedlings and the transgenic seedlings were on soil with three control lines and the PeaT1OE1, PeaT1OE40, and PeaT1OE43 lines, which contained the Nipponbare background. Each line consisted of 10 seedlings. Approximately 21 days later, watering of the seedlings was stopped for 12 days to simulate drought conditions. Seedling survival rates were recorded 15 days after watering was resumed. The drought stress experiment was conducted at least three times.
Chlorophyll, ABA, Malondialdehyde and Proline Measurements
The chlorophyll extracted from 50 mg (fresh weight) leaf tissue was quantified using the absorbance values at 652 nm according to the Infinite M200 multimode reader (Tecan Group Ltd, Switzerland) as previously described by Kong et al. (2006). ABA was extracted from seedling leaves and quantified according to a previously described method (Fu et al., 2012). Chlorophyll and ABA contents were measured before and during the simulated drought treatment. For every line (10 seedlings), a total of 50 mg tissue was collected from three leaves. Malondialdehyde (MDA) content was measured using a published procedure (Heath and Packer, 1968). Proline concentration was determined as described (Bates et al., 1973). All measurements contained three independent biological replicates. Significant differences are determined with Student’s t-test (∗0.01 < P < 0.05, ∗∗P < 0.01).
Quantitative Real-Time PCR Analysis
Total RNA was extracted from WT and PeaT1OE transgenic rice plants using Trizol reagent (TaKaRa, Japan) according to the procedure recommended by the manufacturer. The first-strand cDNA was synthesized in a 20-μl solution using the QuantiTect Reverse Transcription Kit (Qiagen, Germany). The 20 μl qRT-PCR solutions included of 0.5 μl cDNA, 0.2 μM primer mix, and reagents from the SYBR Premix Ex Taq Kit (TaKaRa, Japan). The qRT-PCR was conducted using the ABI PRISM 7900HT system (Applied Biosystems, United States) (Livak and Schmittgen, 2001), with the endogenous rice gene Ubiquitin (LOC_Os03g13170) serving as the control. Semi quantitative RT-PCR was determined according to method (Marone et al., 2001). qRT-PCR was performed with three technical and three biological replicates. All qRT-PCR primers are listed in Supplementary Table S1. Significant differences are determined with Student’s t-test (∗0.01 < P < 0.05, ∗∗P < 0.01).
Drought-Responsive Genes Expression
Drought-responsive genes were detected by quantitative real-time polymerase chain reaction (qRT-PCR) analysis. Water was withheld from 4-week-old rice seedlings for 12 days, after which transcript levels of drought-responsive genes in leaves were analyzed (OsAM1, OsLP2, OsDST, OsCPK9, OsSKIPa OsNCED2, OsNAC9, OsTPKb, and OsEREBP1). Additionally, OsMIOX expression was detected by exposing 4-week-old transgenic seedlings to air (without water), and then collecting leaves 0.5, 1, 2, 3, 5, 7, and 9 h later for a subsequent qRT-PCR, qRT-PCR was performed with three technical and three biological replicates. All primers are listed in Supplementary Table S1.
Yeast Two-Hybrid Assay
The PeaT1 and OsMIOX coding sequences were amplified using gene-specific primer sets (Supplementary Table S1). The full-length PeaT1 sequence was ligated into the pDBleu vector to form the BD-PeaT1 plasmid. Meanwhile, the full-length OsMIOX sequence was cloned into the pPC86 vector to generate the AD-OsMIOX plasmid. Yeast cells were transformed and the subsequent reporter gene assays were completed according to the manufacturer’s instructions (Invitrogen, United States).
Pull-Down Assay
For the in vitro pull-down assay, the OsMIOX and PeaT1 coding regions were inserted into pMAL-c2X and pGEX6p-1 vector, respectively, to generate the OsMIOX–maltose-binding protein (MBP) and PeaT1-glutathione S-transferase (GST) plasmids (Supplementary Table S1 for primer sets). The amylose resin for purifying MBP (New England Biolabs) and the GST-binding resin for purifying GST (Merck, Germany) were used to purify the fusion proteins and empty tags. The pull-down assay was conducted as previously described (Zhou et al., 2013). Protein products were detected using an enhanced chemiluminescence reagent (GE Healthcare, China).
Subcellular Localization of OsMIOX
To prepare the pCAMBIA1305-d35S-MIOX-green fluorescent protein (GFP) plasmid, the MIOX coding sequence was cloned into the Bgl II site of the pCAMBIA1305-GFP vector. The construct was transiently expressed in tobacco (Nicotiana benthamiana) epidermal cells as previously described (Batoko et al., 2000).
Results
Enhanced Drought Tolerance of PeaT1OE Rice Plants
To investigate PeaT1OE activities in drought-stressed plants, 3-week-old WT and PeaT1OE seedlings (Figure 1A) were subjected to simulated drought conditions for 12 days (Figure 1B). Control leaves demonstrated curled 2 days earlier than PeaT1OE leaves. Additionally, three PeaT1OE lines exhibited leaf wilting and rolling symptoms after the 12-day drought treatment. Fifteen days after watering was resumed (Figure1C), 60% of PeaT1OE plants were still alive, whereas the survival rate of the control plants was only 40% (Figure 1D).
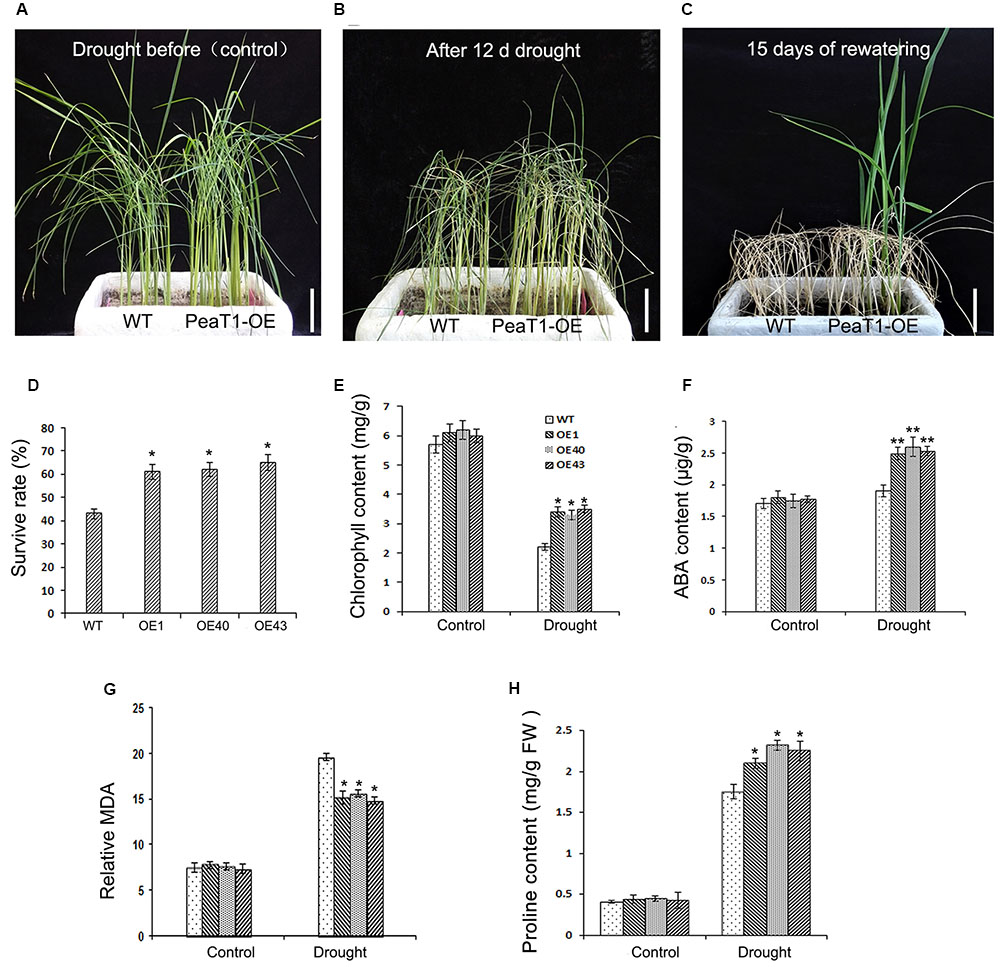
FIGURE 1. Drought stress tolerance in wild-type (WT) and PeaT1OE lines. (A) Three-week-old seedlings. (B) Phenotypes of plants deprived of water for 12 days. (C) Phenotypes of plants 15 days after watering was resumed. (D) Chlorophyll contents of control and PeaT1OE plants, with or without the drought treatment. (E) Survival rates were calculated 15 days after watering was resumed. (F) Abscisic acid (ABA) contents of the control and PeaT1OE plants, with or without the drought treatment. (G) Malondialdehyde (MDA) contents of the control and PeaT1OE plants, with or without the drought treatment. (H) Proline contents of control and PeaT1OE plants, with or without the drought treatment. ∗0.01 < P < 0.05, ∗∗P < 0.01; Bar in (A–C) is 5 cm.
Chlorophyll and ABA contents are used to determine whether plants have been exposed to drought stress (Seki et al., 2007; Li et al., 2013). In the current study, we determined that there was no difference in the chlorophyll content between WT and PeaT1OE plants grown under normal conditions. In contrast, among plants exposed to simulated drought conditions, the chlorophyll content of PeaT1OE lines was 50% higher than that of WT plants (Figure 1E). Leaf ABA content is correlated with adaptations to environmental stresses via a feedback loop. Thus, we quantified the ABA contents of PeaT1OE and WT leaves under normal and drought conditions. Results showed that there were no significant differences between the two lines before the drought treatment (Figure 1F). However, under drought stress, ABA content increased by 38 and 11% in the PeaT1OE and WT plants, respectively.
Malondialdehyde is the end-product of the peroxidation of membrane lipids. It is a consequence of oxidative damage derived from the decomposition of polyunsaturated fatty acid hydroperoxides. A comparison of the MDA contents of WT and PeaT1OE plants revealed that no obvious differences were detected before the drought treatment. However, in response to drought conditions, MDA content decreased significantly (0.01 < P < 0.05) in PeaT1OE plants. The result indicated that minor lipid peroxidation occurring in PeaT1OE plants contributed to drought tolerance.
Proline is associated with adaptation to stress abiotic stress in plant (Jiang et al., 2016). We also checked the content of proline in WT and PeaT1OE plant before and during the drought treatment. Nonetheless, no differences were observed under normal conditions (Figure 1H). Proline content improved to higher level in PeaT1OE plants, whereas WT plants showed a low increase in proline. These changes demonstrate that proline accumulation are corresponded to the increased drought tolerance of PeaT1OE plants.
Analysis of Drought-Responsive Gene Expression Levels
For a more thorough characterization of the mechanism regulating drought tolerance in PeaT1OE plants, expression levels of several stress-responsive genes were investigated. OsAM1 gene encodes a K+ efflux antiporter that is mainly present in plastid-containing organisms, from lower green algae to higher flowering plants. The expression of OsAM1 is reportedly induced by polyethylene glycol and salt. Additionally, an am1 mutant was observed to be more drought tolerant than WT plants (Sheng et al., 2014). In this study, the OsAM1 transcript level decreased significantly in PeaT1OE plants (Figure 2A), suggesting decreased abundance of K+ efflux antiporters may be associated with increased drought tolerance of PeaT1OE plants. Furthermore, OsLP2 (Leaf Panicle 2) (Sheng et al., 2014; Wu et al., 2014) and OsDST (Huang et al., 2009) expression levels were much lower in PeaT1OE lines than in WT (Figures 2B,C). Additionally, OsCPK9 encodes a calcium-dependent protein kinase that affects abiotic stress tolerance (Wei et al., 2014). This gene was highly expressed in PeaT1OE plants during the exposure to drought conditions (Figure 2D). The transcript level of OsSKIPa, which encodes a rice homolog of the human Ski-interacting protein, was higher in PeaT1OE plants than in WT (Figure 2E), indicating the expression of drought-responsive genes changed much a lot in PeaT1OE plants.
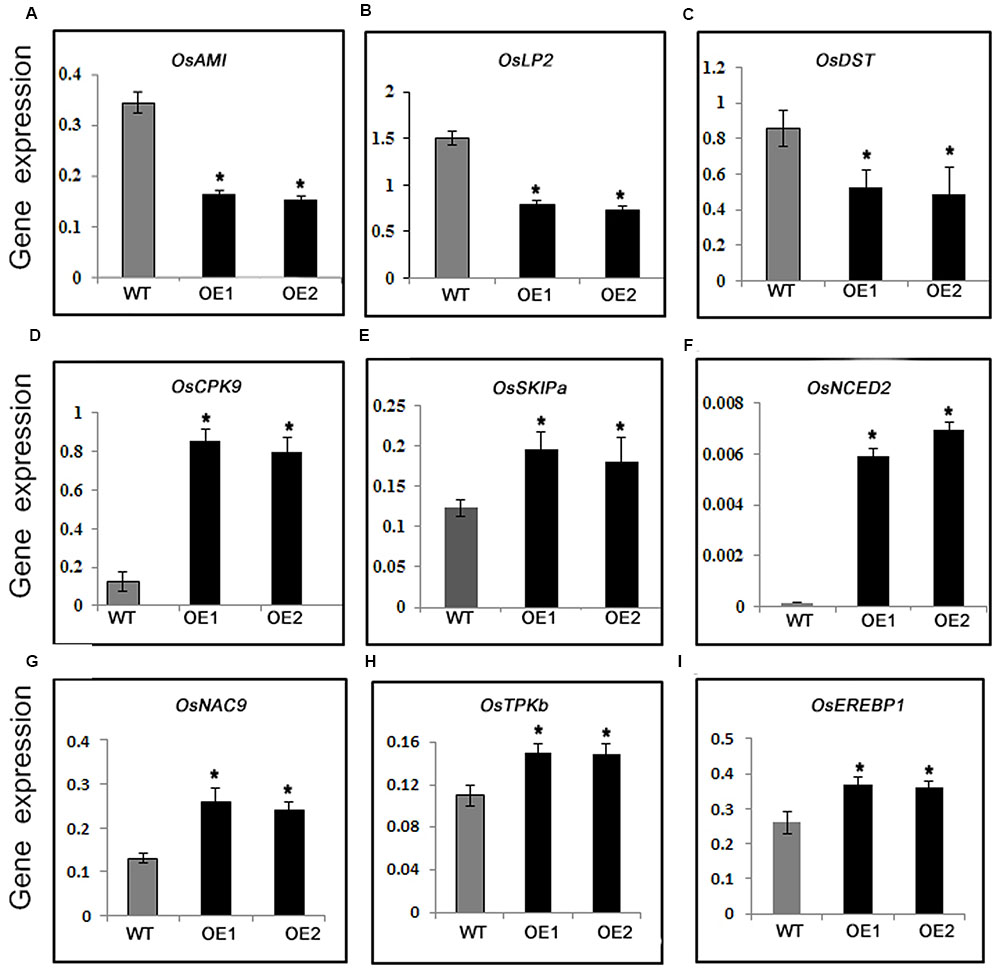
FIGURE 2. Relative expression levels of drought-responsive genes in WT and PeaT1OE plants. The analyzed genes were OsAM1 (A), OsLP2 (B), OsDST (C), OsCPK9 (D), OsSKIPa (E), and OsNCED2 (F), OsNAC9 (G), OSTPKb (H), OsEREBP1 (I). Data is presented as the mean ± standard deviation (n = 9). Significant differences are determined with Student’s t-test (∗0.01 < P < 0.05, ∗∗P < 0.01).
Abscisic acid is crucial for plant drought stress responses, especially the closure of stomata (Lim et al., 2015). Therefore, we analyzed the expression of OsNCED2 (9-cis-epoxycarotenoid dioxygenase), which is important for ABA biosynthesis. The OsNCED2expression level was higher in the PeaT1OE plants than in the WT plants (Figure 2F), suggesting that increased ABA biosynthesis may regulate drought resistance in rice plants.
OsNAC9, a member of rice NAC domain family, had been reported to improve drought tolerance in rice (Redillas et al., 2012). In our study, OsNAC9 was up-regulated in PeaT1OE plants (Figure 2G). OsTPKb, which encodes a two pore potassium channel protein (Ahmad et al., 2016), decreased the transpiration in its overexpression rice plants under drought stress. We found that the transcription level of OsTPKb was higher in PeaTOE plants (Figure 2H). Overexpression of an AP2/ERF transcription factor OsEREBP1 confers drought tolerance in rice (Jisha et al., 2015). Our detection showed that OsEREBP1 expression level was higher in PeaT1OE plants than WT (Figure 2I). Above all, the regulation of drought inducible genes was contributed in PeaT1OE plants to drought tolerance.
PeaT1 Interacts with OsMIOX
In order to excavate the molecular mechnism for PeaT1’s role in drought, a yeast two-hybrid screen using the Nipponbare cDNA was conducted to determine if there are any rice proteins which interact with PeaT1. We were also interested in characterizing the functions of any interacting protein. The BD-PeaT1 plasmid was used as the bait. Because PeaT1 contains a self-activated domain, we added 50 mM 3-AT to SD/-Leu/-Trp/-His medium. Dozens of potential interacting proteins were detected, including chloroplast precursor protein S1 and OsMIOX. To verify the accuracy of interaction, we made PeaT1 as the bait, and OsMIOX were as preys, respectively, in a yeast two-hybrid analysis. Finally, we found that there really existed interaction between PeaT1 and OsMIOX based on the results of X-gal (5-Bromo- 4-chloro-3-indolyl β-D-galactopyranoside) assays involving SD/-Leu/-Trp/-His medium supplemented with 3-AT (Figure 3A).
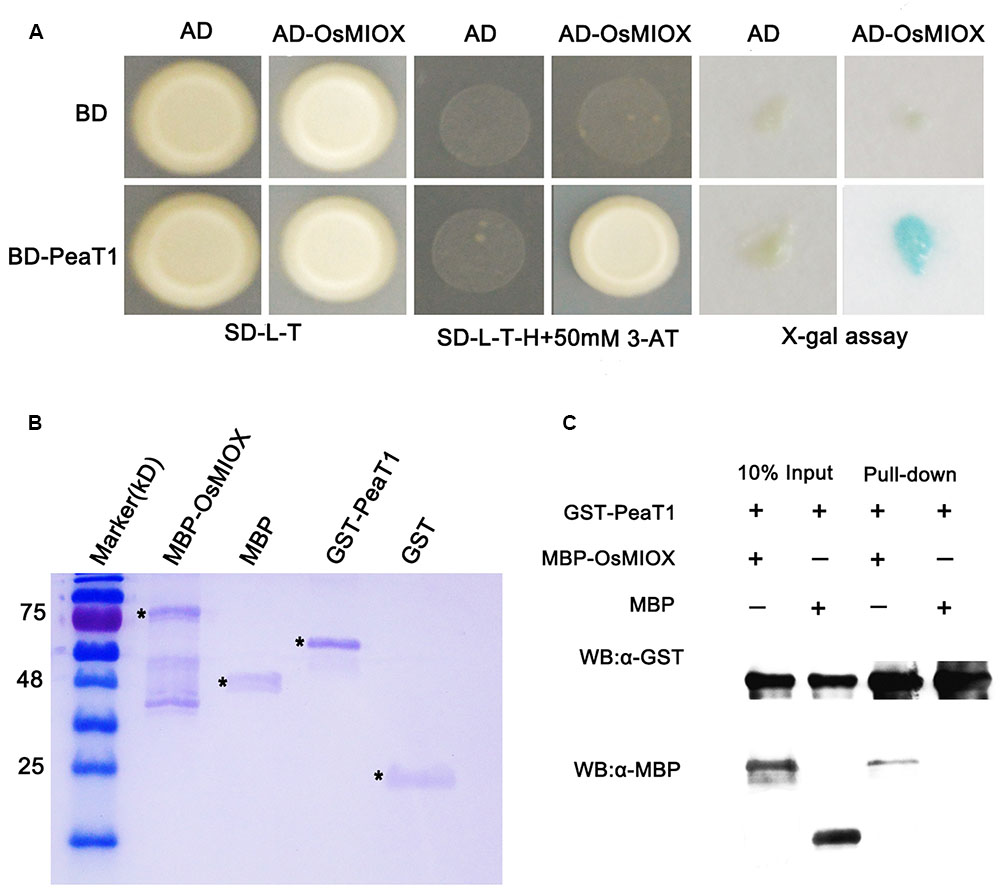
FIGURE 3. PeaT1 interacts with OsMIOX. (A) Results of a yeast two-hybrid assay involving BD-PeaT1 and AD-OsMIOX. The transformed yeast cells were plated on SD/-Leu/-Trp medium (left) and SD/-Leu/-Trp/-His medium supplemented with 50 mM 3-AT (middle). The results of the X-gal assay are also provided (right). (B) Purified proteins stained with Coomassie Brilliant Blue. (C) A solution consisting of PeaT1-GST bound to the GST-binding resin was incubated with an equal volume of cell lysate containing OsMIOX-MBP or MBP. The binding of OsMIOX-MBP or MBP to the GST-binding resin before (Input) and after (Pull-down) washes was examined in a western blot involving the anti-MBP antibody. The same protein gel was used for a western blot involving the anti-GST antibody to verify that equal amounts of GST-PeaT1 were present in all protein mixtures. ∗ Represents the target protein band.
We also confirmed the interaction using a pull-down assay involving the PeaT1-MBP and OsMIOX1-GST fusion proteins (Figure 3B), which were produced in Escherichia coli cells. The OsMIOX1-GST fusion protein was able to pull down PeaT1-MBP, but GST alone could not (Figure 3C). These results verified the direct interaction between PeaT1 and OsMIOX.
OsMIOX Expression Was Upregulated during the Drought Treatment
To assess whether OsMIOX expression in PeaT1OE and WT plants was affected by drought stress, water-cultivated seedlings were exposed to air to simulate drought conditions. Leaves were collected at specific time points to measure the OsMIOX expression levels by qRT-PCR and semi-quantitative RT-PCR (Figure 4). The upregulated expression of OsMIOX followed a two-phase curve reminiscent of the interaction process. The first significant increase in PeaT1OE plants occurred at 0.5 and 2 h after the initiation of the drought treatment. A second peak was observed when PeaT1 interacted with OsMIOX at 5, 7, and 9 h after starting the drought treatment. There were no differences between the PeaT1OE and WT plants at the 1- and 3-h time points. These results suggested that PeaT1 might function cooperatively with OsMIOX during stress responses in PeaT1OE plants.
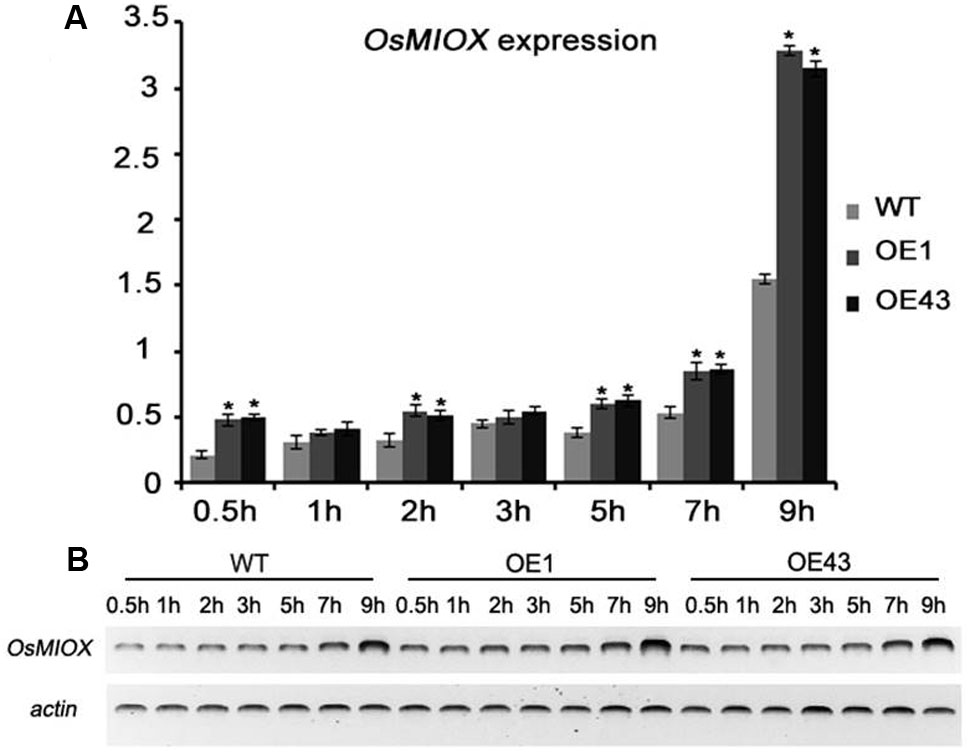
FIGURE 4. Expression of OsMIOX in WT and PeaT1OE plants exposed to air. (A) qRT-PCR analysis of OsMIOX expression. (B) OsMIOX expression was detected by semi-quantitative RT-PCR. Data is presented as the mean ± standard deviation (n = 9). Significant differences are determined with Student’s t-test (∗0.01 < P < 0.05, ∗∗P < 0.01).
Subcellular Localization of OsMIOX
To investigate the subcellular localization of OsMIOX, the OsMIOX cDNA was fused in frame to the GFP marker gene under the control of the cauliflower mosaic virus 35S promoter (35S::MIOX::GFP). The vector with the GFP gene was used as the control (Figure 5A). Mcherry fluorescent protein (mRFP)-tagged organelle markers (Figures 5B–E) were incorporated into the epidermal cells of N. benthamiana leaves. Protoplasts were prepared from the tobacco leaves to enhance the fluorescent signals detected by confocal microscopy. The MIOX-GFP fluorescence did not completely merge with the plasma membrane (PM: SCAMP1 [Lam et al., 2007]) (Figure 5B), which was consistent with previous results of predicted localization in cytoplasm. Furthermore, four organelle markers tagged with mRFP (Gomord et al., 1997; Nebenführ et al., 1999; Miao et al., 2006; Lam et al., 2007) were co-expressed with MIOX-GFP in tobacco protoplasts. We observed that the MIOX-GFP fluorescence co-localized only with the endoplasmic reticulum (ER) marker HDEL (Figure 5C).
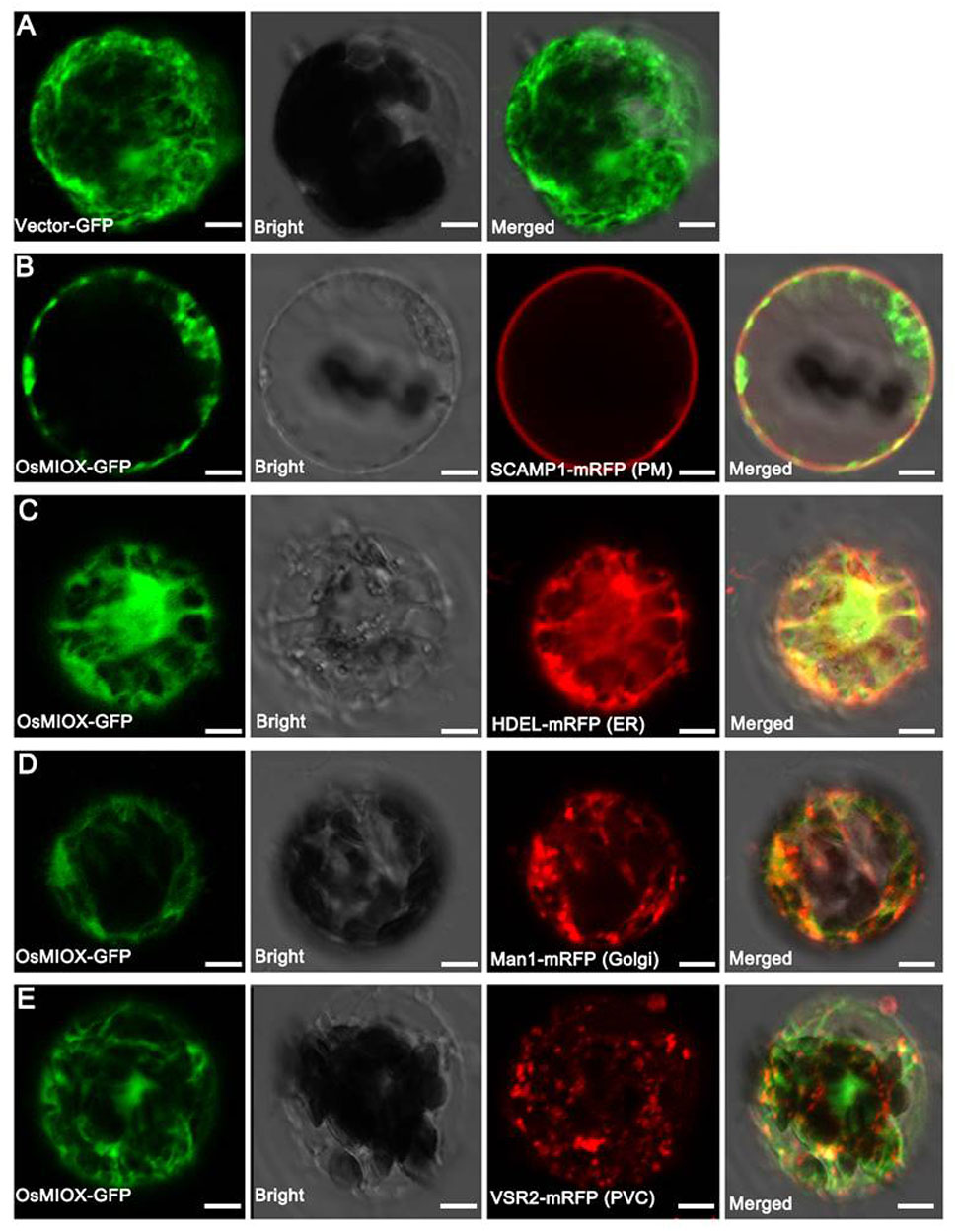
FIGURE 5. Subcellular localization of OsMIOX in protoplasts of N. benthamiana leaves. (A) Fluorescence signal from the expression of the GFP tag. Fluorescence signal from the co-expression of the OsMIOX-GFP fusion protein and (B) SCAMP1-mRFP (PM marker), (C) HDEL-mRFP (ER marker), (D) Man1-mRFP (Golgi marker), (E) VSR2-mRFP (PVC marker). The scale bar corresponds to (A) 9 μm, (B) 5 μm, (C) 8 μm, and (D,E) 10 μm. PM, plasma membrane; ER, endoplasmic reticulum; PVC, prevacuolar compartment.
Discussion
Drought results in considerable adverse consequences for crop yield and quality. Previous studies have concluded that the PeaT1 elicitor is critical for the drought resistance and development in rice, wheat, and cotton plants (Liu et al., 2009; Wang et al., 2011; Tang et al., 2012). In the study, we observed that PeaT1OE plants showed increased drought-tolerant compared with WT, with higher survival rate in OE (Figure 1). Thus, PeaT1’s function in drought resistance is similar to another elicitor Harpin, which could enhances drought tolerance in rice (Zhang W. et al., 2011). These results suggested that PeaT1 played crucial roles in plant improvement tolerance of drought.
Expression of several genes related to drought resistance also changed differently between PeaT1OE plants and WT. For example, OsDST transduction significantly decreased in OE plants compared with WT (Figure 2C). As OsDST is a negative regulator in drought (Huang et al., 2009), lower expression of the gene might partially increase stomatal closure and reduce stomatal density, thus causing enhanced drought in OE plants. A study reported that the expression of OsLP2 is down-regulated by drought and the gene’s overexpression plants are drought sensitivity (Wu et al., 2014). Our data detected that lower OsLP2 expression existed in PeaT1 OE plants than in the WT (Figure 2B), suggesting lower expression of OsLP2 contributed to the plant drought resistance in OE. Taking together, these results implied that the overexpression of PeaT1 in plants might impact the transduction of other drought-responsive genes, thereby inducing OE plants drought tolerance.
Abscisic acid exerts key roles in affecting stomatal defense against drought stresses via activating diverse plant physiological and developmental processes (Finkelstein et al., 2002; Robertseilaniantz et al., 2007; Ton et al., 2009). Under drought conditions, ABA is produced or accumulated to induce stomatal closure, ultimately helping plant conserve water (Bauer et al., 2013; Lee and Luan, 2012). ABA-controlled processes are necessary for plant survival and deficient mutants generally are susceptible to water stress (Finkelstein et al., 2002; Kang et al., 2002). The upregulated expression of NCED3 (an ABA biosynthesis gene) is induced by drought stress, in turn decreasing transpiration rates and enhancing plant water stress (Schwartz et al., 1997, 2003; Tan et al., 1997; Thompson et al., 2000). Here, we analyzed the expression of OsNCED2 (another ABA biosynthesis gene) and found that it was significantly increased in PeaT1OE plants (Figure 2F), similar to NCED3’s upregulation, which both improve plant drought resistance. These results indicated that overexpressing PeaT1 affected some ABA biosynthesis genes expression, which increased ABA content, finally improved plant drought tolerance. However, several ABA signaling genes expression should be detected in future in order to know whether upregulation of PeaT1 has effects on ABA signal tranduction.
Myo-inositol oxygenase (MIOX) could regulate the abundance of myo-inositol, thus affecting the ascorbic acid biosynthesis (Kanter et al., 2005). Ascorbic acid, as a major plant antioxidant, plays roles in countering balances of any oxidative damage (Shao et al., 2008). However, little knowledge about the sub-location of MIOX, except rice OsMIOX was predicted to be localized in cytoplasm (Duan et al., 2012), it was unclear whether it is associated with specific subcellular structures. In the study, we co-expressed OsMIOX with crucial organelle markers in tobacco protoplasts and observed that OsMIOX just co-localized with ER marker, suggesting OsMIOX functions in ER.
A recent study showed that overexpression of OsMIOX in rice could improve drought tolerance (Duan et al., 2012). Our results showed that OsMIOX expression was significantly upregulated in PeaT1OE plants under drought conditions, suggesting that PeaT1 might promote the transcription of OsMIOX. However, the molecuar function of PeaT1 or OsMIOX on drought tolerance remians unclear. By yeast screening system, we found that PeaT1 could interact with OsMIOX in vivo and in vitro, which offered evidence of PeaT1’s roles in drought reistance on molecular level. And it is the first report that elicitor PeaT1 could physicall interact with Myo-inositol oxygenase together to carry out their function. Further studies on whether PeaT1 interacting with other protein will be required to deeply understand PeaT1’s molecular function.
Author Contributions
XY and DQ designed experiments. FS carried out experiments and analyzed experimental results. FS wrote the manuscript. YZ and YD assistant the experiment and wrote the MS.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31672097), The Special Fund for Agro-scientific Research in the Public Interest in China (201303021).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2017.00970/full#supplementary-material
References
Ahmad, I., Devonshire, J., Mohamed, R., Chultze, M., and Maathuis, F. J. (2016). Overexpression of the potassium channel TPKb in small vacuoles confers osmotic and drought tolerance to rice. New Phytol. 209, 1040–1048. doi: 10.1111/nph.13708
Antoni, R., Rodriguez, L., Gonzalez-Guzman, M., Pizzio, G. A., and Rodriguez, P. L. (2011). News on ABA transport, protein degradation, and ABFs/WRKYs in ABA signaling. Curr. Opin. Plant Biol. 14, 547–553. doi: 10.1016/j.pbi.2011.06.004
Bates, L. S., Waldren, R. P., and Teare, I. D. (1973). Rapid determination of free proline for water-stress studies. Plant Soil 39, 205–207. doi: 10.1016/j.dental.2010.07.006
Batoko, H., Zheng, H. Q., Hawes, C., and Moore, I. (2000). A Rab1 GTPase is required for transport between the endoplasmic reticulum and Golgi apparatus and for normal Golgi movement in plants. Plant Cell 12, 2201–2218. doi: 10.1105/tpc.12.11.2201
Bauer, H., Ache, P., Lautner, S., Fromm, J., Hartung, W., Alrasheid, K. A. S., et al. (2013). The stomatal response to reduced relative humidity requires guard cell-autonomous ABA synthesis. Curr. Biol. 23, 53–57. doi: 10.1016/j.cub.2012.11.022
Duan, J., Zhang, M., Zhang, H., Xiong, H., Liu, P., Ali, J., et al. (2012). OsMIOX, a myo-inositol oxygenase gene, improves drought tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L.). Plant Sci. 196, 143–151. doi: 10.1016/j.plantsci.2012.08.003
Fan, L. M., Zhao, Z., and Assmann, S. M. (2004). Guard cells: a dynamic signaling model. Curr. Opin. Plant Biol. 7, 537–546. doi: 10.1016/j.pbi.2004.07.009
Finkelstein, R. R., Gampala, S. S. L., and Rock, C. D. (2002). Abscisic acid signaling in seeds and seedlings. Plant Cell 14, S15–S45.
Fu, J., Chu, J., Sun, X., Wang, J., and Yan, C. (2012). Simple, rapid, and simultaneous assay of multiple carboxyl containing phytohormones in wounded tomatoes by UPLC-MS/MS using single SPE purification and isotope dilution. Anal. Sci. 28, 1081–1087. doi: 10.2116/analsci.28.1081
Fujita, Y., Fujita, M., Shinozaki, K., and Yamaguchishinozaki, K. (2011). ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 124, 509–525. doi: 10.1007/s10265-011-0412-3
Gomord, V., Denmat, L. A., Fitchette-Lainé, A. C., Satiat-Jeunemaitre, B., Hawes, C., and Faye, L. (1997). The C-terminal HDEL sequence is sufficient for retention of secretory proteins in the endoplasmic reticulum (ER) but promotes vacuolar targeting of proteins that escape the ER. Plant J. 11, 313–325. doi: 10.1046/j.1365-313X.1997.11020313.x
Gonzalez-Guzman, M., Pizzio, G. A., Antoni, R., Vera-Sirera, F., Merilo, E., Bassel, G. W., et al. (2012). Arabidopsis PYR/PYL/RCAR receptors play a major role in quantitative regulation of stomatal aperture and transcriptional response to abscisic acid. Plant Cell 24, 2483–2496. doi: 10.1105/tpc.112.098574
He, S. Y., Huang, H. C., and Collmer, A. (1993). Pseudomonas syringae pv. syringae harpinPss: a protein that is secreted via the Hrp pathway and elicits the hypersensitive response in plants. Cell 73, 1255–1266. doi: 10.1016/0092-8674(93)90354-S
Heath, R. L., and Packer, L. (1968). Photoperoxidation in isolated chloroplasts: I. Kinetics and stoichiometry of fatty acid peroxidation. Arch. Biochem. Biophys. 125, 189–198. doi: 10.1016/0003-9861(68)90654-1
Huang, X. Y., Chao, D. Y., Gao, J. P., Zhu, M. Z., Shi, M., and Lin, H. X. (2009). A previously unknown zinc finger protein, DST, regulates drought and salt tolerance in rice via stomatal aperture control. Genes Dev. 23, 1805–1817. doi: 10.1101/gad.1812409
Jiang, Y., Qiu, Y., Hu, Y., and Yu, D. (2016). Heterologous expression of AtWRKY57 confers drought tolerance in Oryza sativa. Front. Plant Sci. 7:145. doi: 10.3389/fpls.2016.00145
Jisha, V., Dampanaboina, L., Vadassery, J., Mithöfer, A., Kappara, S., and Ramanan, R. (2015). Overexpression of an AP2/ERF type transcription factor OsEREBP1 confers biotic and abiotic stress tolerance in rice. PLoS ONE 10:e0127831. doi: 10.1371/journal.pone.0127831
Kang, J. Y., Choi, H. I., Im, M. Y., and Kim, S. Y. (2002). Arabidopsis basic leucine zipper proteins that mediate stress-responsive abscisic acid signaling. Plant Cell 14, 343–357. doi: 10.1105/tpc.010362
Kanter, U., Usadel, B., Guerineau, F., Li, Y., and Pauly, M. (2005). The inositol oxygenase gene family of Arabidopsis is involved in the biosynthesis of nucleotide sugar precursors for cell-wall matrix polysaccharides. Planta 221, 243–254. doi: 10.1007/s00425-004-1441-0
Kim, T. H. (2014). Mechanism of ABA signal transduction: agricultural highlights for improving drought tolerance. J. Plant Biol. 57, 1–8. doi: 10.1007/s12374-014-0901-8
Kong, Z., Li, M., Yang, W., Xu, W., and Xue, Y. (2006). A novel nuclear-localized CCCH-type zinc finger protein, OsDOS, is involved in delaying leaf senescence in rice. Plant Physiol. 141, 1376–1388. doi: 10.1104/pp.106.082941
Kwak, J. M., Mori, I. C., Pei, Z. M., Leonhardt, N., Torres, M. A., Dangl, J. L., et al. (2003). NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J. 22, 2623–2633. doi: 10.1093/emboj/cdg277
Lam, S. K., Siu, C. L., Hillmer, S., Jang, S., An, G., Robinson, D. G., et al. (2007). Rice SCAMP1 defines clathrin-coated, trans-golgi–located tubular-vesicular structures as an early endosome in tobacco BY-2 cells. Plant Cell 19, 296–319. doi: 10.1105/tpc.106.045708
Lee, S. C., Lim, C. W., Lan, W., He, K., and Luan, S. (2013). ABA signaling in guard cells entails a dynamic protein–protein interaction relay from the PYL-RCAR family receptors to ion channels. Mol. Plant 6, 528–538. doi: 10.1093/mp/sss078
Lee, S. C., and Luan, S. (2012). ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ. 35, 53–60. doi: 10.1111/j.1365-3040.2011.02426.x
Li, G. L., Wu, H. X., Sun, Y. Q., and Zhang, S. Y. (2013). Response of chlorophyll fluorescence parameters to drought stress in sugar beet seedlings. Russ. J. Plant Physiol. 60, 337–342. doi: 10.1134/S1021443713020155
Lim, C. W., Baek, W., Jung, J., Kim, J. H., and Lee, S. C. (2015). Function of ABA in stomatal defense against biotic and drought stresses. Int. J. Mol. Sci. 16, 15251–15270. doi: 10.3390/ijms160715251
Liu, Q., Guang-Yue, L. I., Zeng, H. M., Yang, X. F., and Qiu, D. W. (2009). Acquisition of microbial protein elicitor PeaT1 and preliminary research on inducing drought resistance of rice. J. Agric. Sci. Technol. 11, 51–55.
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative pcr and the 2-ΔΔCT method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Ma, Y., Szostkiewicz, I., Korte, A., Moes, D., Yang, Y., Christmann, A., et al. (2009). Regulators of PP2C phosphatase activity function as abscisic acid sensors. Sci. 324, 1064–1068. doi: 10.1126/science.1172408
Marone, M., Mozzetti, S., De, R. D., Pierelli, L., and Scambi, G. (2001). Semiquantitative RT-PCR analysis to assess the expression levels of multiple transcripts from the same sample. Biol. Proced. Online 3, 19–25. doi: 10.1251/bpo20
Miao, Y., Yan, P. K., Kim, H., Hwang, I., and Jiang, L. (2006). Localization of green fluorescent protein fusions with the seven Arabidopsis vacuolar sorting receptors to prevacuolar compartments in tobacco BY-2 cells. Plant Physiol. 142, 945–962. doi: 10.1104/pp.106.083618
Nebenführ, A., Gallagher, L. A., Dunahay, T. G., Frohlick, J. A., Mazurkiewicz, A. M., Meehl, J. B., et al. (1999). Stop-and-go movements of plant Golgi stacks are mediated by the acto-myosin system. Plant Physiol. 121, 1127–1141. doi: 10.1104/pp.121.4.1127
Nishimura, N., Sarkeshik, A., Nito, K., Park, S. Y., Wang, A., Carvalho, P. C., et al. (2009). PYR/PYL/RCAR family members are major in-vivo ABI1 protein phosphatase 2C-interacting proteins in Arabidopsis. Plant J. 61, 290–299. doi: 10.1111/j.1365-313X.2009.04054.x
Nürnberger, T. (1999). Signal perception in plant pathogen defense. Cell. Mol. Life Sci. CMLS 55, 167–182. doi: 10.1007/s000180050283
Paré, P. W., Farag, M. A., Krishnamachari, V., Zhang, H., Ryu, C. M., and Kloepper, J. W. (2005). Elicitors and priming agents initiate plant defense responses. Photosynth. Res. 85, 149–159. doi: 10.1007/s11120-005-1001-x
Park, S. Y., Fung, P., Nishimura, N., Jensen, D. R., Fujii, H., Zhao, Y., et al. (2009). Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324, 1068–1071. doi: 10.1126/science.1173041
Pei, Z. M., Murata, Y., Benning, G., Thomine, S., Klüsener, B., Allen, G. J., et al. (2000). Calcium channels activated by hydrogen peroxide mediate abscisic acid signalling in guard cells. Nature 406, 731–734. doi: 10.1038/35021067
Redillas, M. C., Jeong, J. S., Kim, Y. S., Jung, H., Bang, S. W., Choi, Y. D., et al. (2012). The overexpression of OsNAC9 alters the root architecture of rice plants enhancing drought resistance and grain yield under field conditions. Plant Biotechnol. J. 10, 792–805. doi: 10.1111/j.1467-7652.2012.00697.x
Robertseilaniantz, A., Navarro, L., Bari, R., and Jones, J. D. (2007). Pathological hormone imbalances. Curr. Opin. Plant Biol. 10, 372–379. doi: 10.1016/j.pbi.2007.06.003
Schwartz, S. H., Qin, X., and Zeevaart, J. A. (2003). Elucidation of the indirect pathway of abscisic acid biosynthesis by mutants, genes, and enzymes. Plant Physiol. 131, 1591–1601. doi: 10.1104/pp.102.017921
Schwartz, S. H., Tan, B. C., Gage, D. A., Zeevaart, J. A. D., and Mccarty, D. R. (1997). Specific oxidative cleavage of carotenoids by VP14 of maize. Science 276, 1872–1874. doi: 10.1126/science.276.5320.1872
Seki, M., Umezawa, T., Urano, K., and Shinozaki, K. (2007). Regulatory metabolic networks in drought stress responses. Curr. Opin. Plant Biol. 10, 296–302. doi: 10.1016/j.pbi.2007.04.014
Shao, H. B., Chu, L. Y., Shao, M. A., Jaleel, C. A., and Mi, H. M. (2008). Higher plant antioxidants and redox signaling under environmental stresses. C. R. Biol. 331, 433–441. doi: 10.1016/j.crvi.2008.03.011
Sheng, D., Yushanjiang, M., Guo, L., Zeng, H., Yang, X., Yuan, J., et al. (2011). Transfer of the protein elicitor gene peaT1 into duoxi 1. Mol. Plant Breed. 9, 1450–1456.
Sheng, P., Tan, J., Jin, M., Wu, F., Zhou, K., Ma, W., et al. (2014). Albino midrib 1, encoding a putative potassium efflux antiporter, affects chloroplast development and drought tolerance in rice. Plant Cell Rep. 33, 1581–1594. doi: 10.1007/s00299-014-1639-y
Smirnoff, N., Conklin, P. L., and Loewus, F. A. (2001). BIOSYNTHESIS OF ASCORBIC ACID IN PLANTS: a renaissance. Plant Biol. 52, 437–467. doi: 10.1146/annurev.arplant.52.1.437
Tan, B. C., Schwartz, S. H., Zeevaart, J. A. D., and Mccarty, D. R. (1997). Genetic control of abscisic acid biosynthesis in maize. Proc. Natl. Acad. Sci. U.S.A. 94, 12235–12240. doi: 10.1073/pnas.94.22.12235
Tang, H., Zeng, H., Yang, X., Li, G., Sheng, D., Yuan, J., et al. (2012). Identification of transgenic cotton with a protein elicitor-encoding gene peaT1. Biotechnol. Bull. 1, 60–64.
Thompson, A. J., Jackson, A. C., Symonds, R. C., Mulholland, B. J., Dadswell, A. R., Blake, P. S., et al. (2000). Ectopic expression of a tomato 9-cis-epoxycarotenoid dioxygenase gene causes over-production of abscisic acid. Plant J. 23, 363–374. doi: 10.1046/j.1365-313x.2000.00789.x
Ton, J., Flors, V., and Mauch-Mani, B. (2009). The multifaceted role of ABA in disease resistance. Trends Plant Sci. 14, 310–317. doi: 10.1016/j.tplants.2009.03.006
Wang, L., Yang, X., Zeng, H., Qiu, D., Guo, L., and Liu, Z. (2011). Extracellular expression of protein elicitor PeaT1 in Bacillus subtilis to enhance drought tolerance and growth in wheat. Chin. J. Biotechnol. 27, 1355–1362.
Wei, S., Hu, W., Deng, X., Zhang, Y., Liu, X., Zhao, X., et al. (2014). A rice calcium-dependent protein kinase OsCPK9 positively regulates drought stress tolerance and spikelet fertility. BMC Plant Biol. 14:133. doi: 10.1186/1471-2229-14-133
Wei, Z. M., Laby, R. J., Zumoff, C. H., Bauer, D. W., He, S. Y., Collmer, A., et al. (1992). Harpin, elicitor of the hypersensitive response produced by the plant pathogen Erwinia amylovora. Science 257, 85–88. doi: 10.1126/science.1621099
Wu, F., Sheng, P., Tan, J., Chen, X., Lu, G., Ma, W., et al. (2014). Plasma membrane receptor-like kinase leaf panicle 2 acts downstream of the DROUGHT AND SALT TOLERANCE transcription factor to regulate drought sensitivity in rice. J. Exp. Bot. 66, 271–281. doi: 10.1093/jxb/eru417
Zhang, L., Xiao, S., Li, W., Feng, W., Li, J., Wu, Z., et al. (2011). Overexpression of a Harpin-encoding gene hrf1 in rice enhances drought tolerance. J. Exp. Bot. 62, 4229–4238. doi: 10.1093/jxb/err131
Zhang, W., Yang, X., Qiu, D., Guo, L., and Zeng, H. (2011). PeaT1-induced systemic acquired resistance in tobacco follows salicylic acid-dependent pathway. Mol. Biol. Rep. 38, 2549–2556. doi: 10.1007/s11033-010-0393-7
Zhang, Y., Yang, X., Liu, Q., Qiu, D., Zhang, Y., Zeng, H., et al. (2010). Purification of novel protein elicitor from Botrytis cinerea that induces disease resistance and drought tolerance in plants. Microbiol. Res. 165, 142–151. doi: 10.1016/j.micres.2009.03.004
Zhou, F., Lin, Q., Zhu, L., Ren, Y., Zhou, K., Shabek, N., et al. (2013). D14-SCFD3-dependent degradation of D53 regulates strigolactone signalling. Nature 504, 406–410. doi: 10.1038/nature12878
Keywords: elicitor, PeaT1, interaction, OsMIOX, overexpression
Citation: Shi F, Dong Y, Zhang Y, Yang X and Qiu D (2017) Overexpression of the PeaT1 Elicitor Gene from Alternaria tenuissima Improves Drought Tolerance in Rice Plants via Interaction with a Myo-Inositol Oxygenase. Front. Plant Sci. 8:970. doi: 10.3389/fpls.2017.00970
Received: 07 March 2017; Accepted: 23 May 2017;
Published: 09 June 2017.
Edited by:
Sagadevan G. Mundree, Queensland University of Technology, AustraliaReviewed by:
Sailendra Nath Sarkar, University of Calcutta, IndiaDanuta Babula Skowrońska, Institute of Plant Genetics (PAN), Poland
Copyright © 2017 Shi, Dong, Zhang, Yang and Qiu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Dewen Qiu, cWl1ZGV3ZW5AY2Fhcy5jbg==
 Fachao Shi
Fachao Shi Yijie Dong
Yijie Dong Yi Zhang
Yi Zhang Xiufeng Yang
Xiufeng Yang Dewen Qiu
Dewen Qiu