- 1Department of Engineering Technology, College of Technology, University of Houston, Houston, TX, United States
- 2College of Agriculture and Technology, Arkansas State University, Jonesboro, AR, United States
Colchicine is one of the oldest plant-based medicines used to treat gout and one of the most important alkaloid-based antimitotic drugs with anticancer potential, which is commercially extracted from Gloriosa superba. Clinical trials suggest that colchicine medication could prevent atrial fibrillation recurrence after cardiac surgery. In addition, therapeutic colchicine is undergoing clinical trials to treat non-diabetic metabolic syndrome and diabetic nephropathy. However, the industrial-scale biomanufacturing of colchicine have not yet been established. Clearly, further studies on detailed biorhizome-specific transcriptome analysis, gene expression, and candidate gene validation are required before uncover the mechanism of colchicine biosynthesis and biorhizome-based colchicine biomanufacturing. Annotation of 32312 assembled multiple-tissues transcripts of G. superba represented 15088 unigenes in known plant specific gene ontology. This could help understanding colchicine biosynthesis in G. superba. This review highlights the biorhizomes, rhizome specific genes or gene what expressed with high level in rhizomes, and deep fluid dynamics in a bioreactor specifically for the biomanufacture of colchicine.
Introduction
Alkaloids are one of the most chemically diverse nitrogenous small molecules which are synthesized from amino acids. Many bioactive alkaloids are extracted from plants which have been used for human medicine (Schläger and Dräger, 2016). The Colchicaceae family has a unique colchicine alkaloid biosynthetic mechanism (Chacón et al., 2014). Gloriosa superba L. is a member of Colchicaceae, and is a very successful commercial source of pharmaceutical colchicine (Sivakumar, 2013). Colchicine has several molecular functions (Kwon et al., 2017; Prins et al., 2017). First, colchicine has very strong binding affinity for tubulin that prevents the microtubule assembly and thereby inhibits cell division (Herdman et al., 2016). This antimitotic mechanism has been used in chemotherapy to prevent cancer cell growth (Johnson et al., 2017). In addition, colchicine enhances the interleukin-8 production which could inhibit the human pancreatic cancer (Yokoyama et al., 2017). However, the anticancer applications of colchicine have been limited due to high clinically acceptable concentrations (Lin et al., 2016). Colchicine has been successfully used in plant cytogenetics to double chromosome numbers. For instance, colchicine inhibits the formation of spindle fibers at anaphase, resulting in replicated homozygous chromosomes as in cabbage and broccoli (Yuan et al., 2015). Second, colchicine has been widely used for centuries to treat gout (Wilson and Saseen, 2016; Abhishek et al., 2017). Colchicine treatment could decrease systemic inflammation (Akodad et al., 2017). Indeed, colchicine had antifibrotic effects in diabetic nephropathy (Solak et al., 2017). Finally, clinical data suggested that colchicine treatment could inhibit cardiovascular diseases, among others (Frommeyer et al., 2017).
Medical studies indicated that patients administered with the dose of 0.6 mg colchicine per day would show plasma concentration after single dosing of approximately 2 ng/ml, which has been shown to promote gout inhibition, while 6 ng/ml is required to observe gastric cancer inhibition (Terkeltaub et al., 2010; Lin et al., 2016). Overdoses can have devastating consequences or toxicity (Medani and Wall, 2016). Notably, appropriate G. superba crude extract doses could prevent unintended contraindications which have been reported in traditional treatments (Capistrano et al., 2016; Kande Vidanalage et al., 2016). The pharmaceutical quality control NMR analysis of enantiomer and synthetic racemic mixture of colchicine has been recently reported (Menéndez-López et al., 2017). G. superba seed and field grown rhizomes contain a unique colchicine scaffold with a high concentration of colchicine, approximately 0.9 and 0.3%, respectively (Sivakumar, 2013). Therefore, public biosafety is important in field cultivation, handling, and processing to prevent accidental poisoning of workers. Despite colchicine being highly studied in the medical sector, little is known about the biosynthesis in plants and biosynthetic genes have not yet been identified. Due to lack of this knowledge, there has been limited success in increasing the yield of G. superba rhizomes. Nevertheless, stable high colchicine accumulation is challenging and the cultivation is labor-intensive, time consuming, and expensive (Vanitha and Manimalathi, 2013). Use of natural colchicine has been increasing substantially in the pharmaceutical industry, thus, alternative biomanufacturing platforms must be developed (Sivakumar, 2017).
Plant cell and root culture systems have been typically used in biotech industry to biomanufacture therapeutic molecules (Sivakumar et al., 2011; Tekoah et al., 2015). Despite considerable metabolic engineering or synthetic biotechnology efforts, the yield of bioactive alkaloid molecules are still very low in these systems because, in part, the lack of knowledge of the biosynthetic mechanism, pathways, and gene expression (Li and Smolke, 2016). G. superba and colchicum species root, callus and cell cultures have been conducted in vitro, but these cultures have yielded insignificant concentrations of colchicine (Daradkeh et al., 2012; Ghosh et al., 2015; Nikhila et al., 2017). Clearly, further advancement is needed to effectively overcome these barriers. Notably, in vitro bulbs are capable of producing montanine and hemanthamine alkaloids (Zayed et al., 2011). Since, rhizomatousness is one of the key lifecycle features in the perenniality of G. superba, the biorhizome can be used as an alternative colchicine production system. For instance, rhizomes are the predominant field propagation system for commercially grown G. superba (Phatak and Hegde, 2014; Padmapriya et al., 2015). Each G. superba daughter rhizome arises from a bifurcated mother rhizome, and each rhizome fork possesses one apical vegetative meristem (Mallya Suma et al., 2014). The apical rhizome buds are dynamic asexual organs which involve complex cross-talk between different regulatory levels, and grow into a complete plant which eventually becomes self-supporting (Salvato et al., 2015). There is very little gene expression information regarding rhizome development and cascade mechanisms involving biosynthesis of small molecules (Li et al., 2014). However, G. superba in vitro tuber cultures accumulate 0.01–0.1% DW of colchicine (Selvarasu and Kandhasamy, 2012; Kumar et al., 2015). Dormancy mechanisms may counteract biosynthesis of colchicine in field grown rhizomes, but this impediment has been overcome in the G. superba biorhizome. This review highlights new biotechnological biorhizome-based biomanufacturing to improve the therapeutic colchicine production in G. superba (Figure 1).

FIGURE 1. Illustration of workflow for Gloriosa superba biorhizome biomanufacturing. (A) Biorhizome induction from callus (50 ml flask) – the chemical structure is colchicine; (B) Biorhizome scaled-up in a 5 L airlift bioreactor (height: 16 inches; diameter 8 inches); (C) Harvested biorhizome from bioreactors.
Biorhizome
Biotechnological biorhizomes are asexually produced rootstocks grown in vitro, whose buds develop new shoots, adventitious roots, and daughter biorhizomes to serve as reproductive as well as storage organs (Figure 1C). They may be used to biosynthesize high-value pharmaceutical molecules. Biorhizomes are unique and efficient biosynthetic mechanisms in rhizomatous plants, and an advanced biotechnological platform compared to root and cell cultures (Sivakumar, 2017). Notably, the size of the shoot is directly related to the age and size of the biorhizome, perhaps because the rhizome is not only energy source but hormones source for the developing shoot (Winkel et al., 2011). The coordinating mechanism of the shoot and rhizome could balance the inorganic and organic carbon via photosynthesis and respiration, respectively (Sakamaki and Ino, 2006; Srinivasan et al., 2016). Biorhizomes continuously synthesize colchicine. This functional characteristic of continuous colchicine production is a decided advantage for biomanufacturing compared to root culture, in which colchicine production is quite low (Sivakumar, 2013). The biosynthesis of colchicine exploits the immobilization of the biosynthetic machinery within a differentiated specialized biorhizome.
At the molecular level, regulation of biorhizome formation is very complex but genes controlling shoot production might be involved (Balbuena et al., 2012; Kim et al., 2013). There is evidence that rhizome morphogenesis in Lotus is regulated by photoperiod (Hu et al., 2011; Cheng et al., 2013b). Hormone auxin are involved in the initiation and development of rhizomes in Lotus. Many genes exhibit significant changes in their expression during development, however, genes associated with auxin hormone signaling appear to trigger rhizome induction (Masuda et al., 2007; Cheng et al., 2013a; Novak and Whitehouse, 2013). In bamboo, about 26 genes are highly expressed in the rhizome buds, which are related to auxin biosynthesis and signaling. The transcriptional factor REVOLUTA was highly expressed in rhizome buds of bamboo, which plays an important role in meristem initiation (Wang et al., 2010). In potato, calmodulin-binding protein plays a regulatory role in signal transduction for tuber formation (Reddy et al., 2002). For instance, FT, Lov Kelch protein 2, CONSTAN, and GIGANTEA genes have been involved in the transduction of photoperiodic signals which might be promoting the rhizome budding in potato (Navarro et al., 2011; Yang et al., 2015). There were 14 other important rhizome formation-related genes, including a MADS-box that could be involved in rhizome enlargement (Cheng et al., 2013b). Genes encoding phytochrome B, CO, GI, and FT were identified in Lotus rhizomes, with differing gene expression and regulation in the shoot and rhizome (Yang et al., 2015). The transcription factor families such as AP2-EREBP, bHLH, MYB, NAC, and WRKY play an important role in regulating secondary metabolic pathways in rhizomes (Yang et al., 2012). In addition, miRNAs were differentially expressed in aerial shoots and rhizomes (Zonga et al., 2014). Thus, at the transcriptional level, shoots and biorhizomes are sharing the functional coordination.
Genomic and transcriptomic data generally suggest that gene transcripts involved in translation, transcription regulation, and metabolism were abundant in the rhizome, while in the leaf the gene transcripts for photosynthesis, stress response, and translation were the most dominant (Huang et al., 2016). Hence, the biorhizome is a unique system for identifying rhizome-specific genes for elucidating the colchicine pathway, and the biorhizome can be used as a biofactory to produce pharmaceutical colchicine. Interestingly, colchicine biosynthesis appears to be upregulated in the biorhizome relative to that in adventitious root culture. Gene expression patterns in the rhizome were quite diverse, while the primary and secondary metabolisms were upregulated (Chen and Li, 2016; Gurung et al., 2016). Apparently, the biorhizome biomass and the colchicine biosynthesis are interconnected with shoot production, but more colchicine was produced in the biorhizome than the shoot. For instance, the leaves and stems accumulate less than 0.1% colchicine whereas the biorhizome accumulate over 0.5% (DW) colchicine (Sivakumar, 2017). Indeed, the sprouts upregulate the colchicine production in the biorhizome. In bioreactor culture, the roots-detached biorhizome continuously grows and synthesizes colchicine, whereas shoots-detached biorhizome loses its function to synthesize biomass or colchicine. Despite this, metabolic adaptation or a gene network could enhance the translocation of colchicine from the shoots to the biorhizome, which is important for the plant’s survival.
Indeed, the shoots-detached biorhizome induces the new daughter biorhizome in bioreactor culture. This phenomenon suggests that shoots play a key molecular mechanisms in biorhizome and colchicine biosynthesis. This characteristic could be associated with changes in the fundamental expression pattern of genes, and alterations in various biochemical and physiological processes that would be crucial for growth and survival of biorhizomes. Genes involved in stress response were greatly upregulated in the rhizome (Yang et al., 2016). For instance, the rhizome encodes a mobile signaling protein, which could control the biorhizome formation (Lee et al., 2013). This suggests that biorhizome might have a complete set of the stress response pathway enzymes. In addition, increased levels of dissolved nutrients, oxygen and hormone in bioreactor culture could stimulate daughter biorhizome development. However, G. superba biorhizome transcriptome analysis and gene expression patterns need to be understood to ascertain and unravel the underlying biorhizome regulatory network.
Transcriptome Analysis
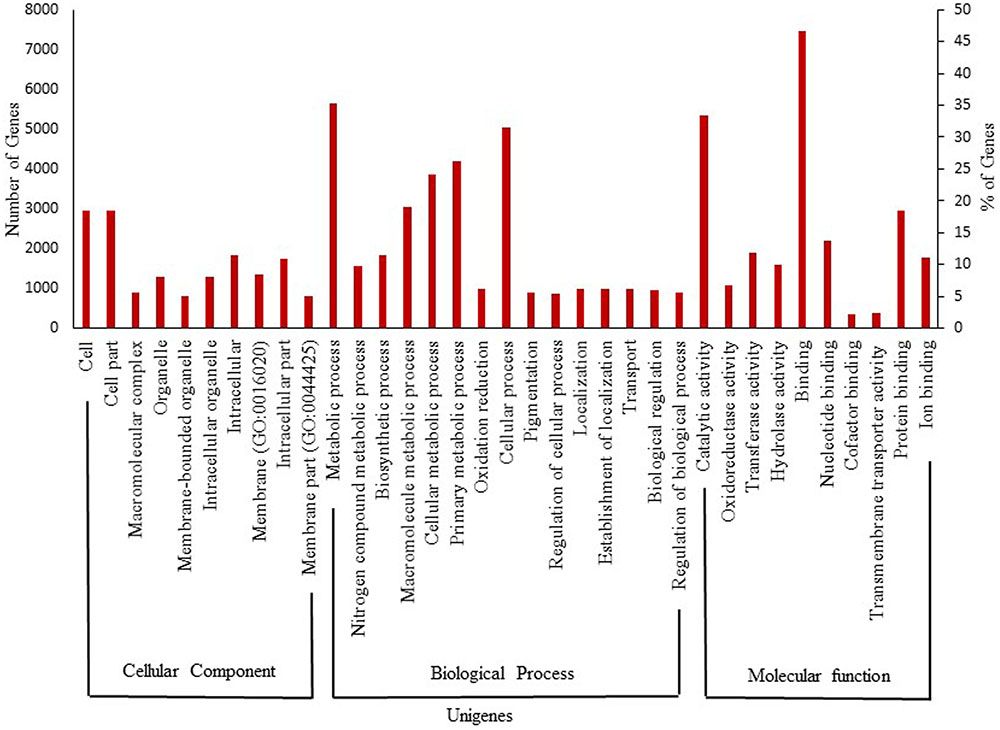
The turmeric and ginger ESTs revealed that over 770 gene transcripts expressed in rhizomes, which are absent in other tissues. These transcripts were enriched for genes associated with rhizome development and regulation. The bioactive small molecules such as curcuminoids and gingerols synthesizing candidate genes were highly expressed in the rhizomes (Koo et al., 2013). Recently, deep sequencing transcriptome data was used to identify various unigenes involved in genome cellular component, biological process, molecular function, and proanthocyanidin biosynthesis in rhizome (Chen and Li, 2016). Notably, the benzylisoquinoline alkaloids biosynthetic genes were highly upregulated during bulb development in Corydalis yanhusuo (Liao et al., 2016). This suggests that rhizome has unique small molecule biosynthetic mechanism. However, there is no molecular information revealing the colchicine biosynthetic pathway in biorhizome. Advanced genomic, proteomic, metabolic, and bioprocess engineering efforts are required to overcome this barrier. Annotation of 32312 assembled transcript sequences, for multi-tissues including dormant rhizomes of G. superba, from the medicinal plant database1 represents 15088 unique genes having homology to known plant specific protein GO terms. For instance, in the cellular component domain, the terms cell (2795 genes, 18.5%, GO:0005623) and cell part (2795, 18.5%, GO:0044464) were mostly assigned. Within the biological function domain, the assignments were mostly enriched in the terms metabolic process (5306, 35.2%, GO:0008152) and cellular process (4746, 31.5%, GO:0009987). For the molecular function domain, the most evident matches were to the terms binding (7026, 46.6%, GO:0005488) and catalytic activity (5038, 33.4%, GO:0003824) (Figure 2). In addition, the G. superba transcriptome contains desired colchicine pathway candidate genes such as of N-methyltransferase, O-methyltransferases, P450s, and N-acetyltransferase (Sivakumar, 2017). Further studies on detailed biorhizome transcriptome analysis, gene expression, and candidate gene validation could uncover the mechanism of colchicine biosynthesis and development in G. superba biorhizomes, and facilitate metabolic engineering and industrial-scale biomanufacturing of colchicine.
Biomanufacturing
Many human medicines are now biomanufactured by genetic engineering or recombinant DNA technology (Tekoah et al., 2015; Roh et al., 2016). Therapeutic small molecules with bioactive natural isomers are derived from biomanufacturing as part of a living system or cells (Sivakumar et al., 2006; Neville et al., 2017). The pharmaceutical quality control colchicine profile is important in raw plant tissue, necessitating that the colchicine molecule drug should not be altered. Therefore, the biomanufacturing is not only to transform a biorhizome system to produce therapeutic colchicine, but also to develop a safer production and quality control as mandated by regulatory agencies. Biomanufacturing colchicine from biorhizomes could lower upstream bioprocessing costs, incorporate economy of scale, speed production, reduce pesticide contamination of drugs.
Ginseng adventitious root culture has been successfully scaled-up in a BTBR (Sivakumar et al., 2005, 2011). Therefore, to scale-up Gloriosa biorhizome a BTBR has been used (Figures 1B, 3). Successful biorhizome scale-up in BTBR require a deep fluid dynamics understanding, because the biorhizomes are completely immersed in the media. For instance, many engineering parameters are involved in the design of a BCR such as; gas density, 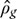 , liquid density,
, liquid density, 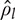 , viscosity,
, viscosity, 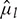 , volumetric gas flow rate,
, volumetric gas flow rate, 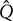 , interfacial tension between gas and liquid phases,
, interfacial tension between gas and liquid phases, 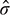 , sparger pore size,
, sparger pore size, 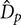 , column diameter,
, column diameter, 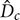 , and length,
, and length, 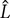 . Such parameters will define mean diameter of the bubbles,
. Such parameters will define mean diameter of the bubbles, 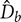 , gas holdup (ratio of the gas phase to the total volume), ε, and superficial velocity defined as
, gas holdup (ratio of the gas phase to the total volume), ε, and superficial velocity defined as 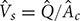 (Kantarci et al., 2005). Here,
(Kantarci et al., 2005). Here, 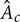 is the cross-sectional area of the column. The flow regimes in BCR are mainly classified according to the column diameter,
is the cross-sectional area of the column. The flow regimes in BCR are mainly classified according to the column diameter, 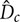 , and the superficial gas velocity,
, and the superficial gas velocity, 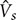 .
.
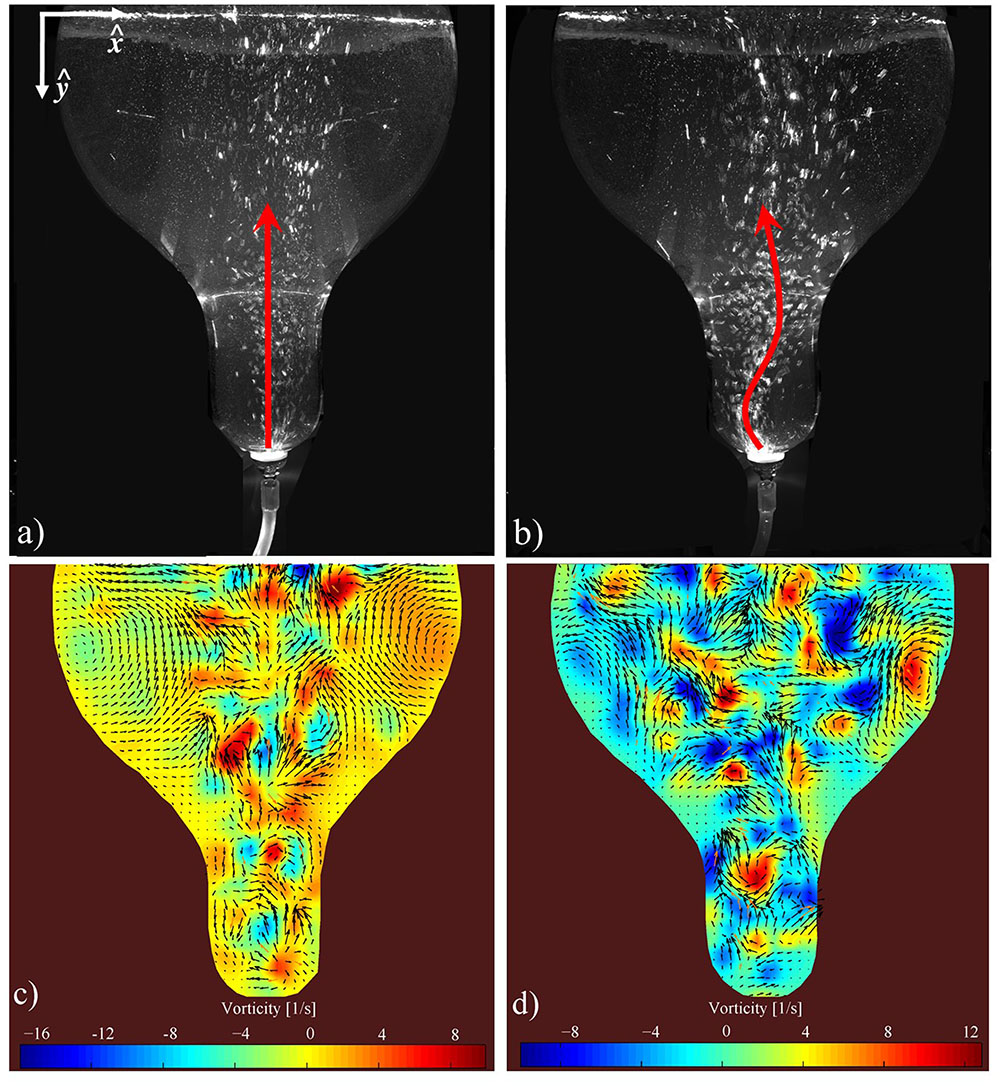
FIGURE 3. Multiphase flow mixing pattern in BTBR: (a,b) snapshots of air injection rates of 50 and 550 cc/min respectively within a 4-L working volume (water seeded with Polyamide Seeding Particles). The arrows indicate the direction of the ascending air bubbles (a,b). (c,d) Corresponding Particle Image Velocimetry to (a,b), respectively. The arrows represent the velocity field while the contours show the local vorticity intensity (c,d).
Two types of flow regimes are commonly observed in BCR, namely homogenous (bubbly) and heterogeneous (churn-turbulent). A heterogeneous slug flow regime could also appear with small diameters at high gas flow rates. The bubbly flows, which can be either perfect or imperfect depending on the degree of the non-uniformity in bubble sizes that are usually obtained at low superficial gas velocities (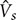 < 5 cm/s) (Bouaifi et al., 2001). The bubbles’ rising velocity and distribution in this regime is relatively steady, the mixing is gentle over the entire reactor and there is no bubble coalescence and/or break-up (Hua and Lou, 2007). Therefore, the bubble size is almost fully dictated by the sparger design and system properties (Ruzicka et al., 2001; Dhotre et al., 2004; Tang and Heindel, 2004; Thorat and Joshi, 2004). The gas holdup, ε, is found to increase linearly with superficial gas velocity,
< 5 cm/s) (Bouaifi et al., 2001). The bubbles’ rising velocity and distribution in this regime is relatively steady, the mixing is gentle over the entire reactor and there is no bubble coalescence and/or break-up (Hua and Lou, 2007). Therefore, the bubble size is almost fully dictated by the sparger design and system properties (Ruzicka et al., 2001; Dhotre et al., 2004; Tang and Heindel, 2004; Thorat and Joshi, 2004). The gas holdup, ε, is found to increase linearly with superficial gas velocity, 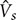 . For higher gas injection rates (
. For higher gas injection rates (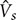 < 5 cm/s), churn-turbulent regimes are found, characterized by the coalescence/break-up of bubbles and increased turbulence and circulation (Hibiki and Ishii, 2000; Olmos et al., 2001; Buwa and Ranade, 2002; Michele and Hempel, 2002). This results in unsteady patterns and various bubble sizes ranging from a few millimeters to a few centimeters. Heat and mass transfer as well as liquid foaming may also introduce additional complexities (Lin and Wang, 2001; Cho et al., 2002; Li and Prakash, 2002; Chen et al., 2003; Krishna and Van Baten, 2003; Ruzicka and Thomas, 2003; Veera et al., 2004). Although several studies have identified the boundaries of possible BCR flow regimes, flow regimes in dimensionless maps have not been reported which is important for industrial design and scale-up. To generate dimensionless maps, the following Buckingham-π theorem analysis was used (Sopan Rahtika et al., 2017).
< 5 cm/s), churn-turbulent regimes are found, characterized by the coalescence/break-up of bubbles and increased turbulence and circulation (Hibiki and Ishii, 2000; Olmos et al., 2001; Buwa and Ranade, 2002; Michele and Hempel, 2002). This results in unsteady patterns and various bubble sizes ranging from a few millimeters to a few centimeters. Heat and mass transfer as well as liquid foaming may also introduce additional complexities (Lin and Wang, 2001; Cho et al., 2002; Li and Prakash, 2002; Chen et al., 2003; Krishna and Van Baten, 2003; Ruzicka and Thomas, 2003; Veera et al., 2004). Although several studies have identified the boundaries of possible BCR flow regimes, flow regimes in dimensionless maps have not been reported which is important for industrial design and scale-up. To generate dimensionless maps, the following Buckingham-π theorem analysis was used (Sopan Rahtika et al., 2017).
Following standard visualization techniques, the dynamics of the flow was characterized in a BTBR in the absence of nutrients and biorhizome to identify homogenous and heterogeneous regimes (Chen and Fan, 1992). The 5 L BTBR was used with 2 and 4 L working volume of polyamide seeding particles (PSP)-water solution at two different air injection rates (low injection rate 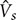 = 0.25 mm/s (
= 0.25 mm/s (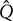 = 50 cm3/min) and higher injection rate 2.76 mm/s (
= 50 cm3/min) and higher injection rate 2.76 mm/s (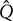 = 550 cm3/min) (Figure 3). Figure 3a suggests that in 4 L the air bubbles at low injection rate ascend up a fairly straight vertical path, concentrating mostly toward the center of the BTBR. However, at higher injection rates, a more chaotic flow forms (Figure 3b). In fact, upon leaving the sparger, the air bubbles oscillate in various directions over time. It is suggested that larger bubbles form at higher injection rate. Figures 3c,d show the 4 L velocity field corresponding to the experiments shown in Figures 3a,b, respectively. The formation of two major vortices are evident of the BTBR at low injection rate (Figure 3c). These major circulatory zones are disturbed (and thus shrunk) at higher flow rate (Figure 3d). The generated fluid mixing and circulation in a bioreactor can significantly affect the quality/quantity of the biorhizome biomass. In order to quantify the strength of the circulatory zones within the flow may calculate the vorticity,
= 550 cm3/min) (Figure 3). Figure 3a suggests that in 4 L the air bubbles at low injection rate ascend up a fairly straight vertical path, concentrating mostly toward the center of the BTBR. However, at higher injection rates, a more chaotic flow forms (Figure 3b). In fact, upon leaving the sparger, the air bubbles oscillate in various directions over time. It is suggested that larger bubbles form at higher injection rate. Figures 3c,d show the 4 L velocity field corresponding to the experiments shown in Figures 3a,b, respectively. The formation of two major vortices are evident of the BTBR at low injection rate (Figure 3c). These major circulatory zones are disturbed (and thus shrunk) at higher flow rate (Figure 3d). The generated fluid mixing and circulation in a bioreactor can significantly affect the quality/quantity of the biorhizome biomass. In order to quantify the strength of the circulatory zones within the flow may calculate the vorticity, 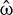 , as
, as 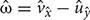 , where
, where 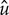 and
and 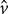 are the velocity components in
are the velocity components in 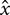 and
and 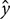 directions, respectively (Figure 3a). Here,
directions, respectively (Figure 3a). Here, 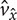 and
and 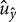 are simply the amount of flow shearing in
are simply the amount of flow shearing in 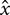 and
and 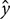 directions (Alba et al., 2014). The vorticity contours (in unit 1/s) have also been added to the velocity vectors shown in Figures 3c,d for comparison. The positive/negative values of the vorticity,
directions (Alba et al., 2014). The vorticity contours (in unit 1/s) have also been added to the velocity vectors shown in Figures 3c,d for comparison. The positive/negative values of the vorticity, 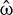 , correspond to clockwise/counter-clockwise directions (Figures 3c,d). The positive and negative vorticity zones are propagated throughout a much larger BTBR domain at higher injection rate suggesting a more uniform mixing (Figure 3d). Both the strongest clockwise (positive
, correspond to clockwise/counter-clockwise directions (Figures 3c,d). The positive and negative vorticity zones are propagated throughout a much larger BTBR domain at higher injection rate suggesting a more uniform mixing (Figure 3d). Both the strongest clockwise (positive 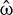 ) and counter-clockwise (negative
) and counter-clockwise (negative 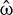 ) rotations were at higher injection rates. The 2 L flow pattern and dimensionless mapping are similar to 4 L. Further analysis is required to understand the counter-intuitive dynamics and flow regimes of such a complex system with biorhizome. Such flow analysis will not only be able to address the geometric patterns of mixing but extend to the nature of liquids, solutions, and injection gasses with various combinations of density, viscosity and surface tension that eventually will improve the biomanufacturing process design.
) rotations were at higher injection rates. The 2 L flow pattern and dimensionless mapping are similar to 4 L. Further analysis is required to understand the counter-intuitive dynamics and flow regimes of such a complex system with biorhizome. Such flow analysis will not only be able to address the geometric patterns of mixing but extend to the nature of liquids, solutions, and injection gasses with various combinations of density, viscosity and surface tension that eventually will improve the biomanufacturing process design.
Critical culture conditions optimized in lab-scale (5–20 L) bioreactor for nutrients, temperature, and culture density may be emulated, at least in part, by that of colchicine biomanufacturing from biorhizomes. Workflow for G. superba upstream biomanufacturing has recently been reported for colchicine (Sivakumar, 2017). However, large-scale data and process validation are required for biorhizomes because during scale-up many working parameters inevitably differ from lab-scale to industrial-scale biomanufacturing. For instance, the nutrient utilization, oxygen level, convective media mixing, and growth factors become more challenging and airflow rate, shear stress profile, and mass transfer are significantly different from small- to large-scale (Roh et al., 2016). Moreover, maintaining reproducibility of biorhizome biomass and colchicine concentration requires homogenous microenvironmental parameters such as nutrients, oxygen, pH, and continuous removal of undesired molecules. These parameters should ideally be monitored online by automated computerized sensors, thereby standardizing the process control during the biomanufacturing processes, as has been done in industrial-scale bioreactors.
Conclusion
Biomanufacturing utilizes the molecular mechanism of living systems and modifies their genome with upstream and downstream processes to develop efficient therapeutic products that help improve human health. Indeed, large-scale biomanufacturing of biopharmaceuticals is a rapidly growing sector of the bioeconomy. Biomanufacturing has utilized regulatory guidance to advance biopharmaceuticals for developing safe and effective medicine. The biorhizome has unique biosynthetic mechanism over plant cell or root cultures which could overcome small molecules production barriers in biomanufacturing. Moreover, biorhizome platforms could revolutionize colchicine upstream biomanufacturing, but first must resolve colchicine pathway elucidation challenges and biomass scale-up for the pharmaceutical industry. For cost-effective robust colchicine biomanufacturing, overproduction via metabolic engineering becomes an important upstream manufacturing step. Reprograming of colchicine biosynthetic pathway in biorhizome or synthetic biotechnology requires detailed pathway elucidation. While studies with large-scale airlift bioreactors for biorhizome manufacturing have not been conducted, a suitable model for colchicine biomanufacturing might be the industrial-scale process for ginsenosides biomanufacturing. More insight into the molecular mechanism of the biorhizome, its interactions with the shoot, as well as mass transfer are needed to fully understand and optimize the biosynthetic pathway for biomanufacturing of colchicine.
Author Contributions
GS lead and designed the experiments and performed the biorhizome biomanufacturing, bioprocess engineering and analytical studies. GP helped bioreactor maintenance. KA performed the fluid mechanics.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
This research has been supported with National Research University (# 110661) and Global Faculty Development Fund from the University of Houston, TX.
Abbreviation
BCR, bubble column reactor; BTBR, balloon type bubble reactor; CO, CONSTAN; DW, dry weight; ESTs, expressed sequence tags; FDA, food and drug administration; FT, Flowering Locus T; GI, GIGANTEA; GO, gene ontology; KEGG, Kyoto encyclopedia of genes and genomes; NAT, N-acetyltransferase; NMR, nuclear magnetic resonance; NMT, N-methyltransferase; OMT, O-methyltransferases.
Footnotes
References
Abhishek, A., Roddy, E., and Doherty, M. (2017). Gout - a guide for the general and acute physicians. Clin. Med. 17, 54–59. doi: 10.7861/clinmedicine.17-1-54
Akodad, M., Lattuca, B., Nagot, N., Georgescu, V., Buisson, M., Cristol, J. P., et al. (2017). COLIN trial: Value of colchicine in the treatment of patients with acute myocardial infarction and inflammatory response. Arch. Cardiovasc. Dis. doi: 10.1016/j.acvd.2016.10.004 [Epub ahead of print].
Alba, K., Taghavi, S., and Frigaard, I. (2014). Miscible heavy-light displacement flows in an inclined two-dimensional channel: a numerical approach. Phys. Fluids 26:122104. doi: 10.1063/1.4903822
Balbuena, T. S., He, R., Salvato, F., Gang, D. R., and Thelen, J. J. (2012). Large-scale proteome comparative analysis of developing rhizomes of the ancient vascular plant Equisetum Hyemale. Front Plant. Sci. 3:131. doi: 10.3389/fpls.2012.00131
Bouaifi, M., Hebrard, G., Bastoul, D., and Roustan, M. (2001). A comparative study of gas hold-up, bubble size, interfacial area and mass transfer coefficients in stirred gas–liquid reactors and bubble columns. Chem. Eng. Process 40, 97–111. doi: 10.1016/S0255-2701(00)00129-X
Buwa, V., and Ranade, V. (2002). Dynamics of gas–liquid flow in a rectangular bubble column: experiments and single/multi-group CFD simulations. Chem. Eng. Sci. 57, 4715–4736. doi: 10.1016/S0009-2509(02)00274-9
Capistrano, R., Vangestel, C., Wouters, A., Dockx, Y., Pauwels, P., Stroobants, S., et al. (2016). Efficacy screening of Gloriosa superba extracts in a murine pancreatic cancer model using (18)F-FDG PET/CT for monitoring treatment response. Cancer Biother. Radiopharm. 31, 99–109. doi: 10.1089/cbr.2015.1954
Chacón, J., Cusimano, N., and Renner, S. S. (2014). The evolution of Colchicaceae, with a focus on chromosome numbers. Syst. Bot. 39, 415–427. doi: 10.1600/036364414X680852
Chen, C., and Li, A. (2016). Transcriptome analysis of differentially expressed genes involved in proanthocyanidin accumulation in the rhizomes of Fagopyrum dibotrys and an irradiation-induced mutant. Front. Physiol. 7:100. doi: 10.3389/fphys.2016.00100
Chen, R. C., and Fan, L. S. (1992). Particle image velocimetry for characterizing the flow structure in three-dimensional gas-liquid-solid fluidized beds. Chem. Eng. Sci. 47, 3615–3622. doi: 10.1016/0009-2509(92)85077-O
Chen, W., Hasegawa, T., Tsutsumi, A., Otawara, K., and Shigaki, Y. (2003). Generalized dynamic modeling of local heat transfer in bubble columns. Chem. Eng. J. 96, 37–44. doi: 10.1016/j.cej.2003.08.016
Cheng, L., Li, S., Xu, X., Hussain, J., Yin, J., Zhang, Y., et al. (2013a). Identification of differentially expressed genes relevant to corm formation in Sagittaria trifolia. PLoS ONE 8:e54573.
Cheng, L., Li, S., Yin, J., Li, L., and Chen, X. (2013b). Genome-wide analysis of differentially expressed genes relevant to rhizome formation in Lotus root (Nelumbo nucifera Gaertn). PLoS ONE 8:e67116. doi: 10.1371/journal.pone.0067116
Cho, Y., Woo, K., Kang, Y., and Kim, S. (2002). Dynamic characteristics of heat transfer coefficient in pressurized bubble columns with viscous liquid medium. Chem. Eng. Process. 41, 699–706. doi: 10.1016/S0255-2701(02)00002-8
Daradkeh, N. Q., Shibli, R. A., Makhadmeh, I. M., Alali, F., and Al-Qudah, T. S. (2012). Cell suspension and in vitro production of colchicine in wild colchicum hierosolymitanum Feib. TOPROCJ. 3, 52–59. doi: 10.2174/1876326X01203020052
Dhotre, M., Ekambara, K., and Joshi, J. (2004). CFD simulation of sparger design and height to diameter ratio on gas hold-up profiles in bubble column reactors. Exp. Therm. Fluid Sci. 28, 407–421. doi: 10.1016/j.expthermflusci.2003.06.001
Frommeyer, G., Krawczyk, J., Dechering, D. G., Kochhäuser, S., Leitz, P., Fehr, M., et al. (2017). Colchicine increases ventricular vulnerability in an experimental whole-heart model. Basic Clin. Pharmacol. Toxicol. 120, 505–508. doi: 10.1111/bcpt.12702
Ghosh, S., Ghosh, B., and Jha, S. (2015). Role of exogenous carbohydrate and amino acid sources on biomass and colchicine production in non- transformed root cultures of Gloriosa superba. Plant Tissue Cult. Biotechnol. 25, 247–256. doi: 10.3329/ptcb.v25i2.26258
Gurung, B., Bhardwaj, P. K., and Talukdar, N. C. (2016). Subtractive transcriptome analysis of leaf and rhizome reveals differentially expressed transcripts in Panax sokpayensis. Funct. Integr. Genomics 16, 619–639. doi: 10.1007/s10142-016-0517-9
Herdman, C. A., Strecker, T. E., Tanpure, R. P., Chen, Z., Winters, A., Gerberich, J., et al. (2016). Synthesis and biological evaluation of benzocyclooctene-based and indene-based anticancer agents that function as inhibitors of tubulin polymerization. Medchemcomm 7, 2418–2427. doi: 10.1039/C6MD00459H
Hibiki, T., and Ishii, M. (2000). Two-group interfacial area transport equations at bubbly-to-slug flow transition. Nucl. Eng. Des. 202, 39–76. doi: 10.1016/S0029-5493(00)00286-7
Hu, F., Wang, D., Zhao, X., Zhang, T., Sun, H., Zhu, L., et al. (2011). Identification of rhizome-specific genes by genome-wide differential expression analysis in Oryza longistaminata. BMC Plant Biol. 11:18. doi: 10.1186/1471-2229-11-18
Hua, J., and Lou, J. (2007). Numerical simulation of bubble rising in viscous liquid. J. Comput. Phys. 222, 769–795. doi: 10.1016/j.jcp.2006.08.008
Huang, Q., Huang, X., Deng, J., Liu, H., Liu, Y., Yu, K., et al. (2016). Differential gene expression between leaf and rhizome in Atractylodes lancea: a comparative transcriptome analysis. Front. Plant Sci. 7:348. doi: 10.3389/fpls.2016.00348
Johnson, L., Goping, I. S., Rieger, A., Mane, J. Y., Huzil, T., Banerjee, A., et al. (2017). Novel colchicine derivatives and their anti-cancer activity. Curr. Top. Med. Chem. doi: 10.2174/1568026617666170104143618 [Epub ahead of print].
Kande Vidanalage, C. J., Ekanayeka, R., and Wijewardane, D. K. (2016). Case report: a rare case of attempted homicide with Gloriosa superba seeds. BMC Pharmacol. Toxicol. 17:26. doi: 10.1186/s40360-016-0069-6
Kantarci, N., Borak, F., and Ulgen, K. (2005). Bubble column reactors. Process Biochem. 40, 2263–2283. doi: 10.1016/j.procbio.2004.10.004
Kim, M. J., Nelson, W., Soderlund, C., and Gang, D. R. (2013). Next-generation sequencing-based transcriptional profiling of Sacred Lotus “China Antique”. Trop Plant Biol. 6, 161–179.
Koo, H. J., McDowell, E. T., Ma, X., Greer, K. A., Kapteyn, J., Xie, Z., et al. (2013). Ginger and turmeric expressed sequence tags identify signatures for rhizome identity and development and biosynthesis of curcuminoids, gingerols and terpenoid. BMC Plant Biol. 13:27. doi: 10.1186/1471-2229-13-27
Krishna, R., and Van Baten, J. (2003). Mass transfer in bubble columns. Catal Today 79, 67–75. doi: 10.1016/S0920-5861(03)00046-4
Kumar, C. N., Jadhav, S. K., Tiwari, K. L., and Afaque, Q. (2015). In vitro tuberization and colchicine content analysis of Gloriosa superba L. Biotechnology 14, 142–147. doi: 10.3923/biotech.2015.142.147
Kwon, O. C., Hong, S., Ghang, B., Kim, Y. G., Lee, C. K., and Yoo, B. (2017). Risk of colchicine-associated myopathy in gout: influence of concomitant use of statin. Am. J. Med. 130, 583–587. doi: 10.1016/j.amjmed.2016.12.006
Lee, R., Baldwin, S., Kenel, F., McCallum, J., and Macknight, R. (2013). Flowering locus T genes control onion bulb formation and flowering. Nat. Commun. 4:2884. doi: 10.1038/ncomms3884
Li, B., Bhandari, D. R., Janfelt, C., Rompp, A., and Spengler, B. (2014). Natural products in Glycyrrhiza glabra (licorice) rhizome imaged at the cellular level by atmospheric pressure matrixassisted laser desorption/ionization tandem mass spectrometry imaging. Plant J. 80, 161–171. doi: 10.1111/tpj.12608
Li, H., and Prakash, A. (2002). Analysis of flow patterns in bubble and slurry bubble columns based on local heat transfer measurements. Chem. Eng. J. 86, 269–276. doi: 10.1016/S1385-8947(01)00186-3
Li, Y., and Smolke, C. D. (2016). Engineering biosynthesis of the anticancer alkaloid noscapine in yeast. Nat. Commun. 7:12137. doi: 10.1058/ncomms12137
Liao, D., Wang, P., Jia, C., Sun, P., Qi, J., Zhou, L., et al. (2016). Identification and developmental expression profiling of putative alkaloid biosynthetic genes in Corydalis yanhusuo bulbs. Sci. Rep. 6:19460. doi: 10.1038/srep19460
Lin, T., and Wang, S. (2001). Effects of macroscopic hydrodynamics on heat transfer in bubble columns. Chem. Eng. Sci. 56, 1143–1149. doi: 10.1016/S0009-2509(00)00333-X
Lin, Z. Y., Kuo, C. H., Wu, D. C., and Chuang, W. L. (2016). Anticancer effects of clinically acceptable colchicine concentrations on human gastric cancer cell lines. Kaohsiung J. Med. Sci. 32, 68–73. doi: 10.1016/j.kjms.2015.12.006
Mallya Suma, V., Sunil Kumar, K. N., and Kamat Shrilatha, K. (2014). Macro-microscopic standards of an abortifacient drug-langali (tubers of Gloriosa superba Linn.). J. Phytopharmacol. 3, 242–247.
Masuda, J., Ozaki, Y., and Okubo, H. (2007). Rhizome transition to storage organ is under phytochrome control in lotus (Nelumbo nucifera). Planta 226, 909–915. doi: 10.1007/s00425-007-0536-9
Medani, S., and Wall, C. (2016). Colchicine toxicity in renal patients - Are we paying attention? Clin. Nephrol. 86, 100–105. doi: 10.5414/CN108343
Menéndez-López, N., Valimaña-Traverso, J., Castro-Puyana, M., Salgado, A., García, M. Á, and Marina, M. L. (2017). Enantiomeric separation of the antiuremic drug colchicine by electrokinetic chromatography. Method development and quantitative analysis. J. Pharm. Biomed. Anal. 138, 189–196. doi: 10.1016/j.jpba.2017.02.001
Michele, V., and Hempel, D. (2002). Liquid flow and phase holdup-measurement and CFD modeling for two-and three-phase bubble columns. Chem. Eng. Sci. 57, 1899–1908. doi: 10.1016/S0009-2509(02)00051-9
Navarro, C., Abelenda, J. A., Cruz-Oró, E., Cuéllar, C. A., Tamaki, S., Silva, J., et al. (2011). Control of flowering and storage organ formation in potato by Flowering Locus T. Nature 478, 119–122. doi: 10.1038/nature10431
Neville, J. J., Orlando, J., Mann, K., McCloskey, B., and Antoniou, M. N. (2017). Ubiquitous chromatin-opening elements (UCOEs): applications in biomanufacturing and gene therapy. Biotechnol. Adv. doi: 10.1016/j.biotechadv.2017.05.004 [Epub ahead of print].
Nikhila, G. S., Sangeetha, G., Chinmayee, D. M., Preetha, T. S., and Swapna, T. S. (2017). Cell suspension culture – An improved system for production of colchicine from Gloriosa superba L. Int. J. Adv. Res. 5, 1184–1190. doi: 10.21474/IJAR01/2869
Novak, S. D., and Whitehouse, G. A. (2013). Auxin regulates first leaf development and promotes the formation of protocorm trichomes and rhizome-like structures in developing seedlings of Spathoglottis plicata (Orchidaceae). AoB Plants 5:pls053. doi: 10.1093/aobpla/pls053
Olmos, E., Gentric, C., Vial, C., Wild, G., and Midoux, N. (2001). Numerical simulation of multiphase flow in bubble column reactors. Influence of bubble coalescence and break-up. Chem. Eng. Sci. 56, 6359–6365. doi: 10.1016/S0009-2509(01)00204-4
Padmapriya, S., Rajamani, K., and Sathiyamurthy, V. A. (2015). Glory lily (Gloriosa superba L.) - A review. Int. J. Curr. Pharmaceut. Rev. Res. 7, 43–49.
Phatak, R. S., and Hegde, L. N. (2014). Glory lily (Gloriosa superba L.): an important medicinal crop - A review. HortFlora Res. Spectrum 3, 282–287.
Prins, K. W., Tian, L., Wu, D., Thenappan, T., Metzger, J. M., and Archer, S. L. (2017). Colchicine depolymerizes microtubules, increases junctophilin-2, and improves right ventricular function in experimental pulmonary arterial hypertension. J. Am. Heart Assoc. 2017:e006195. doi: 10.1161/JAHA.117.006195
Reddy, A. S., Day, I. S., Narasimhulu, S. B., Safadi, F., Reddy, V. S., Golovkin, M., et al. (2002). Isolation and characterization of a novel calmodulin-binding protein from potato. J. Biol. Chem. 277, 4206–4214. doi: 10.1074/jbc.M104595200
Roh, K. H., Nerem, R. M., and Roy, K. (2016). Biomanufacturing of therapeutic cells: state of the art, current challenges, and future perspectives. Annu. Rev. Chem. Biomol. Eng. 7, 455–478. doi: 10.1146/annurev-chembioeng-080615-033559
Ruzicka, M., and Thomas, N. (2003). Buoyancy-driven instability of bubbly layers: analogy with thermal convection. Int. J. Multiphase Flow 29, 249–270. doi: 10.1016/S0301-9322(02)00150-7
Ruzicka, M., Zahradnık, J., Drahos, J., and Thomas, N. (2001). Homogeneous-heterogeneous regime transition in bubble columns. Chem. Eng. Sci. 56, 4609–4626. doi: 10.1016/S0009-2509(01)00116-6
Sakamaki, Y., and Ino, Y. (2006). Tubers and rhizome fragments as propagules: competence for vegetative reproduction in Equisetum arvense. J. Plant Res. 119, 677–683. doi: 10.1007/s10265-006-0026-3
Salvato, F., Balbuena, T. S., Nelson, W., Rao, S. H., He, R., Soderlund, C. A., et al. (2015). Comparative proteomic analysis of developing rhizomes of the ancient vascular plant Equisetum hyemale and different monocot species. J. Proteome Res. 14, 1779–1791. doi: 10.1021/pr501157w
Schläger, S., and Dräger, B. (2016). Exploiting plant alkaloids. Curr. Opin. Biotechnol. 37, 155–164. doi: 10.1016/j.copbio.2015.12.003
Selvarasu, A., and Kandhasamy, R. (2012). In vitro tuberization of glory lily (Gloriosa superba L.). J. Hortic. For. 4, 81–84. doi: 10.5897/JHF11.067
Sivakumar, G. (2013). Colchicine semisynthetics: chemotherapeutics for cancer? Curr. Med. Chem. 20, 892–898. doi: 10.2174/0929867311320070005
Sivakumar, G. (2017). Upstream biomanufacturing of pharmaceutical colchicine. Crit. Rev. Biotechnol. doi: 10.1080/07388551.2017.1312269 [Epub ahead of print].
Sivakumar, G., Medina-Bolivar, F., Lay, J. O., Dolan, M. C., Condori, J., Grubbs, S. K., et al. (2011). Bioprocess and bioreactor: next generation technology for production of potential plant-based antidiabetic and antioxidant molecules. Curr. Med. Chem. 18, 79–90. doi: 10.2174/092986711793979724
Sivakumar, G., Yu, K. W., Lee, J. S., Kang, J. K., Lee, H. L., Kim, W. J., et al. (2006). Tissue cultured mountain ginseng adventitious roots: safety and toxicity evaluation. Eng. Life Sci. 6, 372–383. doi: 10.1002/elsc.200520139
Sivakumar, G., Yu, K. W., and Paek, K. Y. (2005). Production of biomass and ginsenosides from adventitious roots of Panax ginseng in bioreactor cultures. Eng. Life Sci. 5, 333–342. doi: 10.1002/elsc.200520085
Solak, Y., Siriopol, D., Yildiz, A., Yilmaz, M. I., Ortiz, A., Covic, A., et al. (2017). Colchicine in renal medicine: new virtues of an ancient friend. Blood Purif. 43, 125–135. doi: 10.1159/000454669
Sopan Rahtika, I. P. G., Wardana, I. N. G., Sonief, A. A., and Siswanto, E. (2017). Experimental investigation on flutter similitude of thin-flat plates. Adv. Acoust. Vib. 2017:7091425. doi: 10.1155/2017/7091425
Srinivasan, V., Thankamani, C. K., Dinesh, R., Kandiannan, K., Zachariah, T. J., Leela, N. K., et al. (2016). Nutrient management systems in turmeric: effects on soil quality, rhizome yield and quality. Ind. Crops Prod. 85, 241–250. doi: 10.1016/j.indcrop.2016.03.027
Tang, C., and Heindel, T. (2004). Time-dependent gas holdup variation in an air-water bubble column. Chem. Eng. Sci. 59, 623–632. doi: 10.1016/j.ces.2003.10.016
Tekoah, Y., Shulman, A., Kizhner, T., Ruderfer, I., Fux, L., Nataf, Y., et al. (2015). Large-scale production of pharmaceutical proteins in plant cell culture - the protalix experience. Plant Biotechnol. J. 13, 1199–1208. doi: 10.1111/pbi.12428
Terkeltaub, R. A., Furst, D. E., Bennett, K., Kook, K. A., Crockett, R. S., and Davis, M. W. (2010). High versus low dosing of oral colchicine for early acute gout flare: twenty-four-hour outcome of the first multicenter, randomized, double-blind, placebo-controlled, parallel-group, dose-comparison colchicine study. Arthritis Rheum. 62, 1060–1068. doi: 10.1002/art.27327
Thorat, B., and Joshi, J. (2004). Regime transition in bubble columns: experimental and predictions. Exp. Therm. Fluid Sci. 28, 423–430. doi: 10.1016/j.expthermflusci.2003.06.002
Vanitha, B., and Manimalathi, P. (2013). Cost and returns from the cultivation of Gloriosa superba. Int. J. Sci. Res. 2, 127–128.
Veera, U., Kataria, K., and Joshi, J. (2004). Effect of superficial gas velocity on gas hold-up profiles in foaming liquids in bubble column reactors. Chem. Eng. J. 99, 53–58. doi: 10.1016/j.cej.2003.09.003
Wang, K., Peng, H., Lin, E., Jin, Q., Hua, X., Yao, S., et al. (2010). Identification of genes related to the development of bamboo rhizome bud. J. Exp. Bot. 61, 551–561. doi: 10.1093/jxb/erp334
Wilson, L., and Saseen, J. J. (2016). Gouty arthritis: a review of acute management and prevention. Pharmacotherapy 36, 906–922. doi: 10.1002/phar.1788
Winkel, A., Colmer, T. D., and Pedersen, O. (2011). Leaf gas films of Spartina anglica enhance rhizome and root oxygen during tidal submergence. Plant Cell Environ. 34, 2083–2092. doi: 10.1111/j.1365-3040.2011.02405.x
Yang, C. Q., Fang, X., Wu, X. M., Mao, Y. B., Wang, L. J., and Chen, X. Y. (2012). Transcriptional regulation of plant secondary metabolism. J. Integr. Plant Biol. 54, 703–712. doi: 10.1111/j.1744-7909.2012.01161.x
Yang, D. L., Sun, P., and Li, M. F. (2016). Chilling temperature stimulates growth, gene over-expression and podophyllotoxin biosynthesis in Podophyllum hexandrum royle. Plant Physiol. Biochem. 107, 197–203. doi: 10.1016/j.plaphy.2016.06.010
Yang, M., Zhu, L., Pan, C., Xu, L., Liu, Y., Ke, W., et al. (2015). Transcriptomic analysis of the regulation of rhizome formation in temperate and Tropical Lotus (Nelumbo nucifera). Sci. Rep. 5:13059. doi: 10.1038/srep13059
Yokoyama, C., Yajima, C., Machida, T., Kawahito, Y., Uchida, M., and Hisatomi, H. (2017). Interleukin-8 enhances the effect of colchicine on cell death. Biochem. Biophys. Res. Commun. 485, 89–94. doi: 10.1016/j.bbrc.2017.02.025
Yuan, S., Su, Y., Liu, Y., Li, Z., Fang, Z., Yang, L., et al. (2015). Chromosome doubling of microspore-derived plants from cabbage (Brassica oleracea var. capitata L.) and broccoli (Brassica oleracea var. italica L.). Front. Plant Sci. 22:1118. doi: 10.3389/fpls.2015.01118
Zayed, R., El-Shamy, H., Berkov, S., and Codina, C. (2011). In vitro micropropagation and alkaloids of Hippeastrum vittatum. In vitro Cell Dev. Biol. Plant 47, 695–701. doi: 10.1007/s11627-011-9368-1
Keywords: anticancer, antigout, bioprocess, Gloriosa superba, transcriptome
Citation: Sivakumar G, Alba K and Phillips GC (2017) Biorhizome: A Biosynthetic Platform for Colchicine Biomanufacturing. Front. Plant Sci. 8:1137. doi: 10.3389/fpls.2017.01137
Received: 11 April 2017; Accepted: 13 June 2017;
Published: 30 June 2017.
Edited by:
Agnieszka Ludwików, Adam Mickiewicz University in Poznań, PolandReviewed by:
Javier Palazon, University of Barcelona, SpainTaras P. Pasternak, Albert Ludwig University of Freiburg, Germany
Copyright © 2017 Sivakumar, Alba and Phillips. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ganapathy Sivakumar, c2dhbmFwYTNAY2VudHJhbC51aC5lZHU=
 Ganapathy Sivakumar
Ganapathy Sivakumar Kamran Alba1
Kamran Alba1