- Guangdong Provincial Key Laboratory of Biotechnology for Plant Development, College of Life Sciences, South China Normal University, Guangzhou, China
Abscisic acid (ABA) transporters are essential for the transport of ABA from its sites of synthesis to its multiple sites of action within plants and are key players in plant stress responses. Despite their importance, there is limited information on ABA transporters in crop plants. In this study, we isolated and characterized an ABA transporter-like 1 (AhATL1) gene from peanut (Arachis hypogaea L.) whose cognate protein, AhATL1, is a member of the ATP-binding cassette transporter G subfamily and localizes to the plasma membrane. The expression of both the AhATL1 transcript and the corresponding protein were upregulated by water stress and treatment with exogenous ABA. Overexpression of AhATL1 in ecotype Columbia (Col) Arabidopsis (AhATL1-OX) plants reduced ABA sensitivity. When AhATL1-OX and Arabidopsis Col plants were subjected to dehydration stress, the expression of 9-cis-epoxycarotenoid dioxygenase 3 (AtNCED3) and responsive to desiccation 29 A (AtRD29A) accumulated rapidly in rosette leaves of both lines. In contrast, while expression of ATP-binding cassette G 40 (AtABCG40) was increased in Col rosette leaves, there was no change in expression of AtABCG40 in AhATL1-OX leaves. Similarly, water loss from detached leaves of AhATL1-OX plants was more rapid than from Col leaves. Therefore, we suggest that the function of AhATL1 is probably to modulate ABA sensitivity by specifically influencing ABA import into cells.
Introduction
Abscisic acid (ABA) is a key phytohormone involved in a host of biological processes, including embryo and seed maturation, postgerminative growth, and abiotic stress responses (Tuteja, 2007). Drought is the most damaging abiotic stress affecting plant productivity (Govind et al., 2009). When higher plants are exposed to drought, the levels of endogenous ABA increase and downstream signaling pathways are activated (Hwang et al., 2016). Indeed, many drought-responsive genes are induced by exogenous application of ABA (Li T. et al., 2016; Zhao et al., 2016; Li et al., 2017). Local active ABA levels reflect a balance of ABA biosynthesis and inactivation by turnover or conjugation, and levels are further modified by compartmentation and transport (Nambara and Marion-Poll, 2005; Finkelstein, 2013). The translocation of ABA between cells, tissues and organs also plays an important role in the physiological response of plants to stress conditions. For example, ABA is rapidly synthesized in root vascular parenchyma under root stress, and is then rapidly transported to leaf tissues, where it induces stomatal closure in peanut and Arabidopsis (Boursiac et al., 2013; Puértolas et al., 2015; Hu et al., 2016).
ATP-binding cassette transporters are a highly conserved family of ATP-driven pump proteins consisting of two hydrophobic transmembrane domains (TMDs), which constitute the membrane-spanning pore, and two cytosolic domains, known as the nucleotide-binding domains (NBDs) or nucleotide-binding folds (NBFs), which contain the ATP-binding Walker A and B motifs (Kerr et al., 2011). In Arabidopsis, four ABA transporters have been identified (AtABCG25, AtABCG40, AtABCG30, and AtABCG31), all four of which are ATP-binding cassette transporter G subfamily members (Kang et al., 2010, 2015; Kuromori et al., 2010; Kanno et al., 2012). AtABCG25 is involved in exporting ABA from the vasculature (Kuromori et al., 2010), while AtABCG40 is a plasma-membrane ABA-uptake transporter in guard cells, and is necessary for timely closure of stomata in response to drought stress and seed germination (Kang et al., 2010, 2015). AtABCG30 mediates ABA-uptake into the embryo, while AtABCG31 brings about ABA secretion from the endosperm (Kang et al., 2015). These data indicate that the ABA transport system plays a significant role in water deficit tolerance and growth regulation. Therefore, investigation of the function of ABA transporters in ABA signaling is essential for an understanding of plant resistance to water stress. However, ABA transporters have not yet been characterized in crop plants of economic importance.
Like other crops, the peanut plant (Arachis hypogaea L.), which is a major oil and protein seed crop worldwide, is subjected to periodic soil moisture-deficits of varying degree and duration that can result in substantial loss of yield (Nautiyal et al., 2002). Therefore, how water stress affects growth sensitivity and stomatal movement in peanut, and the role of ABA biosynthesis and transport in this stress response, is of considerable importance. In our previous study in peanut, we found that ABA was rapidly synthesized in roots treated with polyethylene glycol 6000 (PEG 6000), mainly accumulating in root vascular parenchyma, and was then quickly transported to the leaf tissues where it induced stomatal closure (Hu et al., 2016). However, it is currently unclear how ABA transport to leaves from biosynthesis sites affects stomatal movement during water stress in peanut.
Here we report an ABA transporter-like1 (AhATL1) gene from A. hypogaea that encodes a plasma membrane-localized protein containing ABC family domains. AhATL1 gene transcription and protein expression are enhanced when peanut roots are treated with PEG 6000 or exogenous ABA. Furthermore, no AhATL1 protein is detected when peanut root is incubated with ABA biosynthesis inhibitor fluridone (Flu) following PEG 6000 treatment. On the other hand, AhATL1 protein expression is similar to that with PEG 6000 treatment alone if plants are pretreated with Flu and then Flu is replaced with ABA. This suggests that AhATL1 expression specifically responds to drought stress via ABA signals in peanut. Overexpression of AhATL1 in Arabidopsis Col lines enhances seed germination, and also increases the axial root length compared with Col plants after exposure to various concentrations of ABA, indicating that AhATL1 decreases ABA sensitivity when expressed heterogenously in Arabidopsis. At the same time, while drought increased AtABCG40 expression in Col plants, there was no such change in AhATL1-OX plants. Moreover, stomatal movement in AhATL1-OX plants in response to drought stress was lower than that in Col plants, while water loss rates were higher. In summary, AhATL1 appears to reduce ABA sensitivity by specifically influencing ABA import.
Materials and Methods
Peanut Plant Material and Growth Conditions
Peanut (Arachis hypogaea L.) cultivar Yueyou 7, a line resistant to water stress, was provided by the Crop Research Institute, Guangdong Academy of Agricultural Sciences, China. Seeds were soaked in water for 12 h, and then placed on moist filter paper in a growth chamber with a cycle of 16 h light from fluorescent and incandescent lamps (200 μmol m-2 s-1) at 26°C followed by 8 h darkness at 22°C for 48 h until the cusp was exposed. Then germinated seeds were sown in saturated peat-containing soil in a growth chamber. Plants were well watered as previously (Su et al., 2015).
Isolation and Sequence Analysis of AhATL1 from Arachis hypogaea L.
First-strand cDNA was synthesized by reverse transcription (RT) of 1 μg total RNA from peanut leaves, using 200 units Superscript III Reverse Transcriptase (Invitrogen, catalog no. 18080) and 500 ng oligo-dT primer under the following conditions: 70°C for 10 min, 42°C for 1 h, and 15 min at 70°C (Su et al., 2015). The AhATL1 open reading frame was amplified by PCR using the following primers: AhATL1 forward primer 5′-ATGGCATCTGAAACCATGGTGGT-3′, AhATL1 reverse primer 5′-TTATGCTTGATGACTTAGTGCAG-3′. PCR amplification was performed as follows: 94°C for 5 min, then 35 cycles of 94°C for 30 s, 55°C for 3 min and 72°C for 1 min, with a final extension step at 72°C for 10 min. PCR products were ligated into the pMD19 T-vector (TaKaRa, catalog no. 6013) and confirmed by sequencing. The AhATL1 amino acid sequence and transmembrane structure was analyzed, respectively, by BLAST1 and TMHMM 2.02.
Peanut Plant Treatments
To simulate the effects of different plant hormone treatments, peanut plants were gently removed from soil at the four-leaf seedling stage, maintained in 1/8 Murashige and Skoog medium (MS) for up to 2 h, and then replaced with 100 μM ABA, 100 μM gibberellin3 (GA3), 50 nM 1-naphthylacetic acid (NAA) or 25 μM 6-benzylaminopurine (6-BA) for 1, 3, 5 or 8 h. For pretreatment with the ABA biosynthesis inhibitor fluridone (Flu), the peanut taproot was first pretreated with 100 μM Flu for 2 h followed by treatment with 20% (w/v) PEG 6000 solution or 100 μM ABA for 1, 3, 5, 8, 12 or 24 h. To examine the effects of water stress on peanut plants, taproots were transferred to 1/8 MS for 2 h and then replaced with 20% (w/v) PEG 6000 solution for 1, 3, 5, 8, 12 or 24 h under standard growth conditions.
Protein Extraction
Peanut leaf samples (first functional leaves; 100 mg) were frozen in liquid nitrogen immediately after treatment and stored at -70°C until further use. Total leaf protein samples were extracted by grinding leaves in liquid nitrogen and 1 mL lysis buffer [50 mM Tris-HCl pH 7.2, 10% glycerol, 2% SDS, 1% β-mercaptoethanol, protease inhibitors cocktail (Roche), 100 mM PMSF (Sigma)].
Antibody Preparation and Immunoblotting
To analyze the production of AhATL1 protein in peanut leaf samples by immunoblotting, three synthetic peptides (each of 12–14 amino acid residues) corresponding to residues 69–82 (QKPSDETRSTEERT), 132–143 (GKITKQTLKRTG), and 328–340 (GVTEREKPNVRQT) of the AhATL1 protein were used to prepare three polyclonal anti-AhATL1 antibodies in rabbits. Total protein from peanut leaves was used to test antibody specificity.
The concentration of protein samples was determined using a TaKaRa BCA Protein Assay Kit (TaKaRa, Code No. T9300A). Total leaf protein was suspended in 5 × SDS–PAGE loading buffer (0.25 M Tris-HCl, pH 6.8, 10% SDS, 50% glycerol, 5% 2-mercaptoethanol), then loaded and run on 15% polyacrylamide gels, which were subsequently stained with Coomassie Brilliant Blue or blotted onto a 0.22 μm PVDF membrane. The membrane was blocked overnight in Tris-buffered saline with 0.1% Tween 20 (TBST, pH 7.6) containing 5% dry milk and then incubated with 0.01–0.05 mg/mL of anti-AhATL1 antibody for 14 h at 4°C. After washing with TBST three times, the primary antibody was detected with secondary HRP-labeled goat anti-rabbit IgG (H + L) antibody (Dingguo, catalog no. HD001-1) at room temperature for 45 min. Visualization was achieved using the ECL system (Millipore, catalog no. 345818).
Expression Analysis
Total RNA was extracted from peanut leaf or Arabidopsis thaliana rosette leaf tissue after various treatments using a TRIzol Kit (TaKaRa), and a Prime Script TM RT Reagent Kit (Perfect Real Time, TaKaRa) was used for RT. Real-time quantitative PCR (qRT-PCR) was performed with the ChamQ SYBR qPCR Master Mix (Low ROX Premixed, Vazyme Biotech Co., Ltd) in an ABI 7500 system to quantify AhATL1 transcript levels. Each reaction consisted of 10 ng cDNA, 0.05 mM primers and 5 μL 2 × ChamQ SYBR qPCR Master Mix in a final volume of 10 μL. The reactive cycle consisted of 95°C for 30 s, then 40 cycles of 95°C for 5 s and 60°C for 34 s. Gene expression level was calculated using the relative 2-ΔΔCT method (Li et al., 2013). Expression data for A. hypogaea and A. thaliana were normalized using the geometric mean (geomean) of the validated housekeeping genes AhACTIN and AtACTIN2, respectively, as described (Su et al., 2015; Li X. et al., 2016). Primers are listed in Supplementary Table S1.
Subcellular Localization
Full-length AhATL1 cDNA was amplified using KOD FX polymerase (Toyobo) with forward primer 5′-CGGGTTC GAAATCGATGGATCCATGGCATCTGAAACCATGGTGGT-3′ and reverse primer 5′-GTCCTAGGCTACGTAGGATCCTTATGCTTGATGACTTAGTGCAG-3′. PCR products were first cloned into pMD19 and confirmed by sequencing. Then, the AhATL1 open reading frame was transferred into the plasmid p35S::eGFP, which contains the 35S promoter, to obtain p35S::AhATL1-eGFP. Constructs were then transformed into Arabidopsis protoplasts, with p35S::eGFP as a positive control. After growing in darkness for 16 h, GFP fluorescence was determined by confocal microscopy (Carl Zeiss LSM 710).
Overexpressing Arabidopsis Plants
The p35S::AhATL1-eGFP construct was transformed into Arabidopsis Col plants by the floral dip method using Agrobacterium tumefaciens strain EHA105. From T2 plants, transgenic lines were identified by PCR. T3 seeds were used for subsequent experiments. Plants were grown on peat-containing soil with a daily cycle of 16 h light and 8 h dark in well-watered conditions at 22 ± 2°C and 60–70% relative humidity.
Arabidopsis Seed Germination, Cotyledon Greening and Root Growth
AhATL1-OX and Col seeds were used to assess the effects of exogenous ABA at different concentrations on seed germination, cotyledon greening and root growth. Seeds were surface sterilized in 70% ethanol containing 0.5% Tween-100 and were air-dried on filter paper. To examine the effects of ABA on germination and the development of green cotyledons, seeds were sown on 1/2 MS medium with 0.8% agar containing 0, 0.1, 0.5 or 2 μM ABA. After 2 days stratification at 4°C, germinating seeds were transferred to a growth chamber with a daily cycle of 16 h light and 8 h dark at 20 ± 2°C. Germination was observed and recorded every 6 h until 96 h. Three days after germination on 1/2 MS medium, seedlings were transferred to new 1/2 MS medium with 0.8% agar and 2% sucrose containing 0, 10 or 50 μM ABA for root growth experiments. Root growth was recorded by photography 7 days after transfer to the new plates and measured using Digimizer software (Li X. et al., 2016).
Stomatal Movement Assay
Detached rosette leaves from 3-week-old AhATL1-OX and Col plants were used to determine stomatal apertures. For assays of stomatal movement under normal conditions or following dehydration treatment, detached whole leaves were floated on “open stomatal buffer” containing 10 mM MES, 5 mM KCl, 50 mM CaCl2, pH 6.15, in a growth chamber under incandescent lamps (200 μmol m-2 s-1) at 20°C for 3 h to allow stomata to open, and were then transferred onto filter paper in a growth chamber under incandescent lamps (200 μmol m-2 s-1) at 20°C for 2 h for drought treatment. Stomatal apertures were measured as described previously (Kang et al., 2010). Digimizer software was used to measure stomatal apertures.
Statistical Analysis
Quantitative data were expressed as mean ± SD. The statistical significance of experimental data was assessed by Student t-test or ANOVA (one-way analysis of variance with a LSD post hoc test), as appropriate, using the SPSS17.0 statistical package.
Accession Numbers
Sequence data from this article can be found in the Arabidopsis genome initiative database or National Center for Biotechnology Information under the following accession numbers: AtNCED3 (AT3G14440), AtABCG40 (AT1G15520), AtABCG25 (AT1G71960), ATACT2 (AT3G18780), AtRD29A (AT5G52310), AhATL1 (KY621345), GsABCG11 (KHN26506), MnABCG11 (XP_010094142), AtABCG11 (NC_00370), and MtWBC (XP_010094142).
Results
AhATL1 Belongs to the ABCG Subfamily
We conducted a BLAST alignment to find conserved protein domains in AhATL1 (Gen Bank accession no. KY621345) after translation of the AhATL1 open reading frame. Two domains were found that characterize ABC transporters: an ABC2 membrane domain (residues 366–572) and an ABC transporter domain (residues 56–207). Sequences signifying Walker A, Walker B, Q-loop, D-loop, and ATP-binding sites were also found in the AhATL1 protein sequence (Figure 1A). A prediction of secondary structure suggested the protein has six transmembrane structures (Figure 1B), consistent with AhATL1 being a member of the ABCG family (Kerr et al., 2011). To determine the subcellular localization of AhATL1, we engineered a plasmid construct comprising a green fluorescent protein (GFP) gene, eGFP, fused to AhATL1 under the control of the cauliflower mosaic virus (CaMV) 35S promoter. The p35S::AhATL1-eGFP recombinant gene was transiently expressed in Arabidopsis protoplast cells, resulting in the green fluorescent AhATL-eGFP protein being present at the cell surface, as shown by confocal microscopy. This expression pattern differed from that of GFP alone, which was distributed throughout the cell (Figure 1C). These results suggest that AhATL1 encodes a plasma membrane protein containing conserved ABCG subfamily domains.
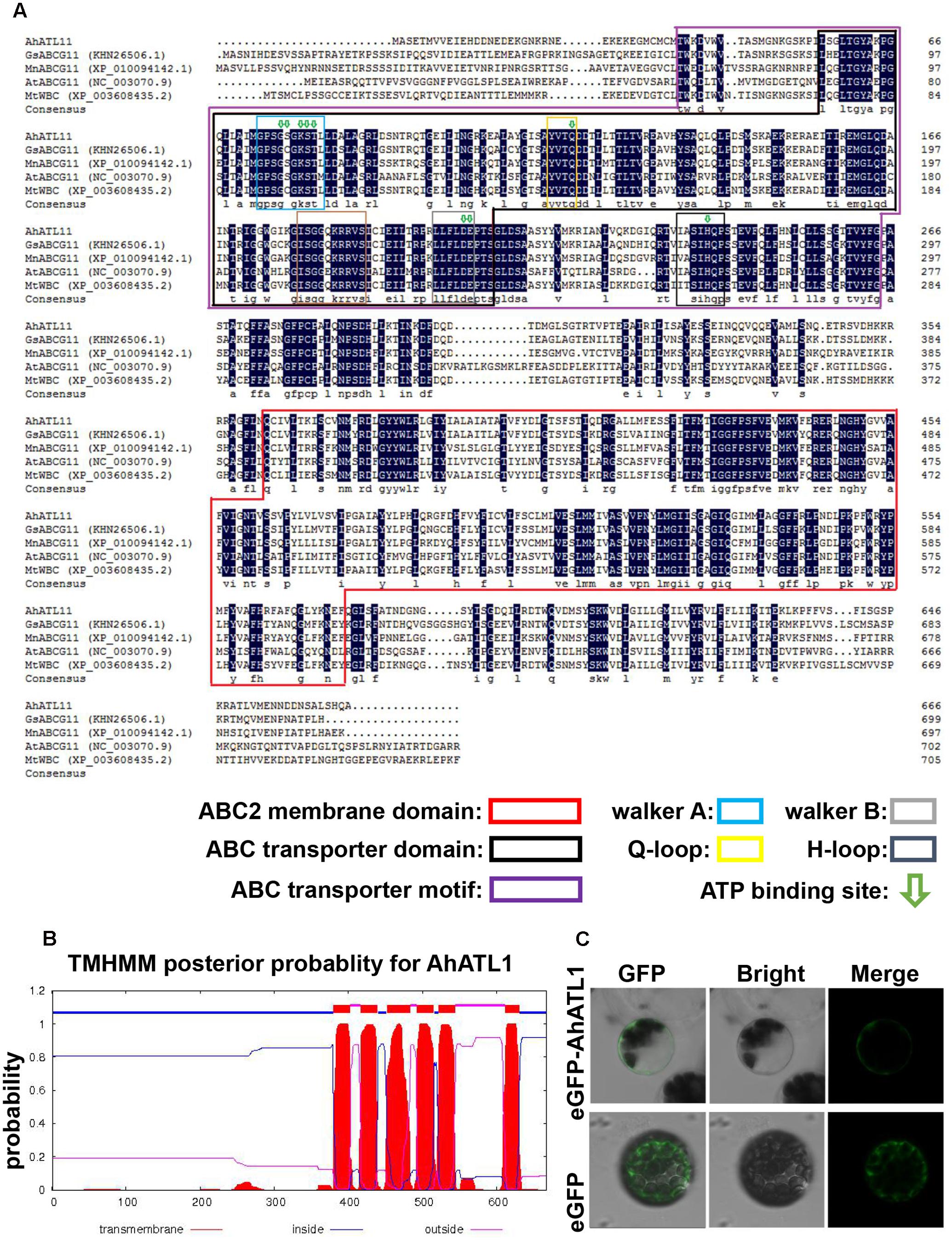
FIGURE 1. AhATL1 is a member of the ABC transporter family. (A) Alignment of amino acid sequences of AhATL1 with four ABCG subfamily proteins. AhATL1: GenBank number KY621345; GsABCG11: GenBank number KHN26506.1; MnABCG11: GenBank number XP_010094142.1; AtABCG11: GenBank number NC_00370.9; MtWBC: GenBank number XP_010094142.1. Identical amino acid residues are shaded in dark blue. Conserved functional domains are marked with rectangular boxes of different colors, as indicated in the key. ATP-binding sites are labeled with green hollow arrows. (B) AhATL1 protein contains a total of six transmembrane structures. Transmembrane helical structures are represented by amino acid residues 380–402, 417–439, 452–483, 493–515, 522–544, and 612–630 of the AhATL1 protein. Consequently, residues 1–379, 440–451, 516–521, and 631–667 are located inside the bilayer (i.e., are intracellular). Amino acid residues 403–416, 484–492, and 545–611 are thus outside the bilayer. AhATL1 transmembrane structure was analyzed using TMHMM 2.0 (online at http://www.cbs.dtu.dk/services/). (C) The intracellular localization of AhATL1 in Arabidopsis protoplasts. Expression of the fusion protein AhATL-eGFP or eGFP alone after transient expression in protoplasts, determined by confocal and bright-field microscopy.
AhATL1 Responds to Drought Stress via ABA-Specific Signals in Peanut
We investigated AhATL1 protein expression in response to treatment of peanut seedlings with the plant hormones ABA, GA3, NAA, and 6-BA over a period of 8 h. Figures 2A,D shows that AhATL1 protein levels begin to increase after 1 h treatment with 100 μM ABA, reaching a maximum after 5–8 h. Treatment with 100 μM GA3 resulted in a gradual decrease in AhATL1 expression, while 50 nM NAA and 25 μM 6-BA also caused a decrease, although with 6-BA this was reversed after 5 h. When we carried out qRT-PCR on total RNA extracted from the same batch of treated seedlings to quantitate levels of AhATL1 transcripts, we found that gene expression levels were broadly consistent with the immunoblotting results, although GA3 treatment did not show a decline in AhATL1 RNA abundance (Figure 2B).
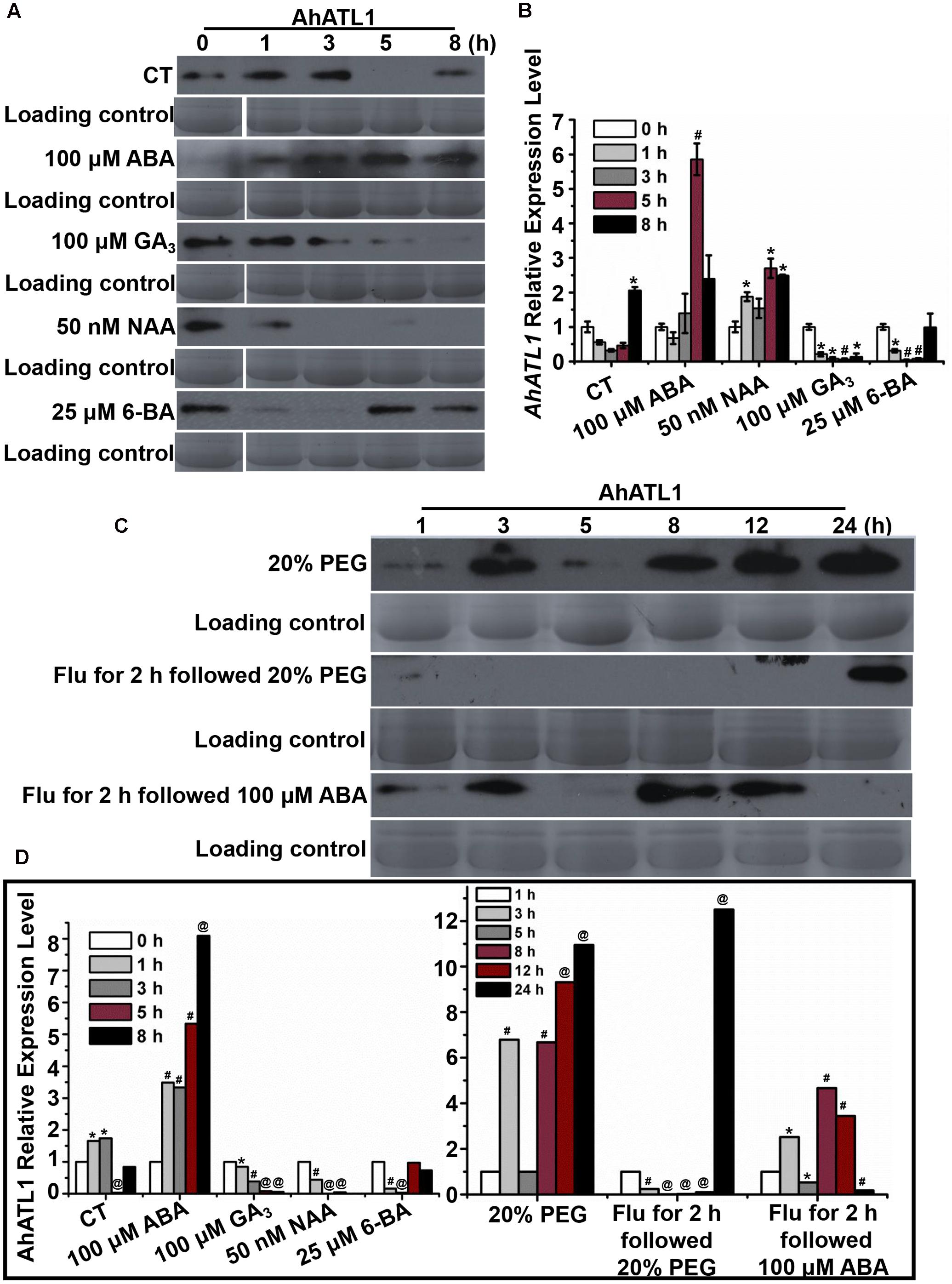
FIGURE 2. AhATL1 is up-regulated by abscisic acid (ABA) and responds to drought stress in peanut. (A) AhATL1 protein accumulation detected by immunoblotting in the first functional leaves of four-leaf peanut plants subjected to 1/8 MS medium (control: CT), 100 μM ABA, 100 μM GA3, 50 nM NAA, and 25 μM 6-BA plant hormone treatments and sampled after 0, 1, 3, 5, and 8 h. (B) Quantitative PCR of AhATL1 transcripts in the first functional leaves of four-leaf peanut plants under CT, 100 μM ABA, 100 μM GA3, 50 nM NAA, and 25 μM 6-BA plant hormone treatments, sampled after 0, 1, 3, 5, and 8 h. Bars indicate standard deviation (n = 3). ‘∗’ or ‘#’ indicate a significant difference at the level of P < 0.05 or P < 0.01, respectively, compared with 0 h under the same treatment. (C) AhATL1 protein accumulation detected by immunoblotting in the first functional leaves of four-leaf peanut plants subjected to treatments with 20% polyethylene glycol (PEG), 20% PEG with 100 μM Flu pretreatment or 100 μM ABA with 100 μM Flu pretreatment and sampled after 1, 3, 5, 8, 12, and 24 h. (D) Statistical analysis of AhATL1 protein accumulation in (A,C) by ANOVA. ‘∗’, ‘#’ or ‘@’ indicate a significant difference in protein accumulation at the level of P < 0.05, P < 0.01 or P < 0.001, respectively, compared with 0 or 1 h of the same treatment.
Together, Figures 2A,B show that AhATL1 and its corresponding protein are positively regulated by ABA. Therefore, because ABA is a key phytohormone involved in drought stress (Hwang et al., 2016), we speculated that AhATL1 might be associated with the drought response in peanut. To test this, we simulated water stress by treating peanut roots with 20% PEG 6000 over a period of 24 h, and observed a complicated, non-linear effect on AhATL1 expression, although the overall trend was an increase in protein levels. Thus, at first, the immunoblotting signal increased from 1 to 2 h, decreased between 3 and 5 h, and finally increased continuously from 8 to 24 h (Figures 2C,D). To confirm a role for ABA in this induction pattern, peanut roots were pretreated with the ABA synthesis inhibitor Flu (100 μM for 2 h), followed by 20% PEG 6000 treatment, or exogenous ABA (100 μM) was applied alone. Interestingly, no AhATL1 protein signal was detected until 8 h during the 20% PEG 6000 treatment; in contrast, AhATL1 was observed immediately when roots were subject to 100 μM ABA following Flu pretreatment (Figures 2C,D). These results demonstrate that AhATL1 is upregulated by ABA and responds to drought stress via ABA-specific signals in peanut.
AhATL1 Reduces Drought Resistance and ABA Sensitivity by Mediating ABA Transport in AhATL1-OX Arabidopsis Plants
To understand AhATL1 function in the context of ABA signaling, we generated transgenic AhATL1-OX Arabidopsis lines (i.e., which overexpress AhATL1). T3 seedlings from the resultant transgenic lines were tested for ABA sensitivity by measuring germination rate, cotyledon greening and axial root growth. The germination rates of AhATL1-OX plants were significantly faster than Col plants when treated with 0.5 or 2 μM ABA, but a difference was even observed without ABA treatment (Figure 3A). With respect to cotyledon greening, there was no significant difference between AhATL1-OX and Col plants without ABA treatment but, remarkably, AhATL1-OX seedlings showed ABA insensitivity at all ABA concentrations tested (0.1, 0.5 or 2 μM; Figures 3B,D). Similarly, the axial root growth of AhATL1-OX and Col seedlings was identical without ABA, but with ABA treatment root growth of the AhATL1-OX seedlings was suppressed to a lesser extent than that of Col seedlings (Figures 3C,E). These results indicate that heterologous expression of AhATL1 reduces ABA sensitivity in Arabidopsis.
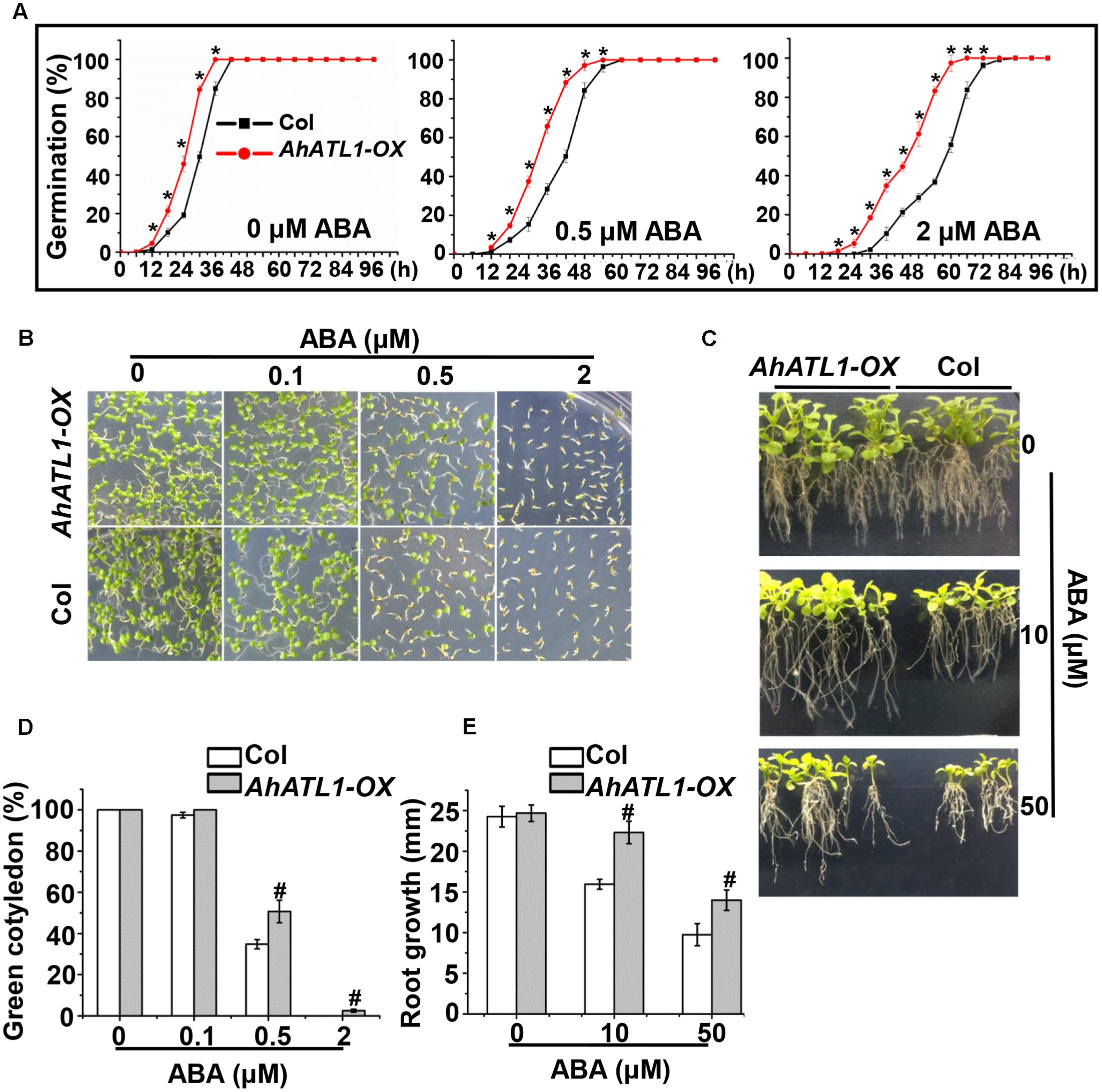
FIGURE 3. AhATL1 decreases ABA sensitivity in Arabidopsis. (A) Seed germination rate of AhATL1-OX and Col lines in response to different concentrations of ABA. Numbers of germinated seedlings were recorded from 0 to 96 h after stratification on 1/2 MS agar plates containing 0, 0.5 or 2 μM ABA. ‘∗’ indicates a significant difference at the level of P < 0.05. (B) Photographs of seedlings grown for 7 days after stratification on agar plates containing 0, 0.1 or 0.5 μM ABA. (C) Photographs of seedlings grown for 20 days after transfer to plates containing 0, 10 or 50 μM ABA. (D) Proportion of green cotyledons in (B). Bars indicate standard deviation (n = 100). ‘#’ indicates a significant difference between AhATL1-OX and Col plants under the same ABA treatment conditions (P < 0.01). (E) Measurement of axial root growth in (C). Bars indicate standard deviation (n = 15). ‘#’ indicates a significant difference between AhATL1-OX and Col plants under the same ABA treatment conditions (P < 0.01).
We further investigated the effect of AhATL1 on specific members of the ABA pathway by measuring transcription levels of AtNCED3, AtABCG25, AtABCG40, and AtRD29A in AhATL1-OX and Col plants subjected to dehydration stress. AtNCED3 and AtRD29A expression was enhanced in both AhATL1-OX and Col plants under dehydration conditions, while AtABCG25 transcription was not affected in either line (Figure 4A). A different profile was noted, however, for AtABCG40, whose expression was increased in Col plants, but was unchanged in AhATL1-OX plants under drought conditions (Figure 4A). AtABCG40 is a plasma-membrane ABA-uptake transporter in guard cells, and is necessary for timely closure of stomata in response to drought stress and seed germination (Kang et al., 2010). Therefore, we examined stomatal movement in AhATL1-OX and Col plants under water stress. The stomatal aperture in leaves of AhATL1-OX plants was larger than Col plants under both normal and dehydration conditions (Figures 4B,C). In line with these results, the water loss from detached leaves of AhATL1-OX plants was higher than from Col leaves under dehydration conditions (Figure 4D). In summary, the above results suggest that AhATL1 may reduce ABA transport into guard cells, thereby leading to ABA desensitization in Arabidopsis.
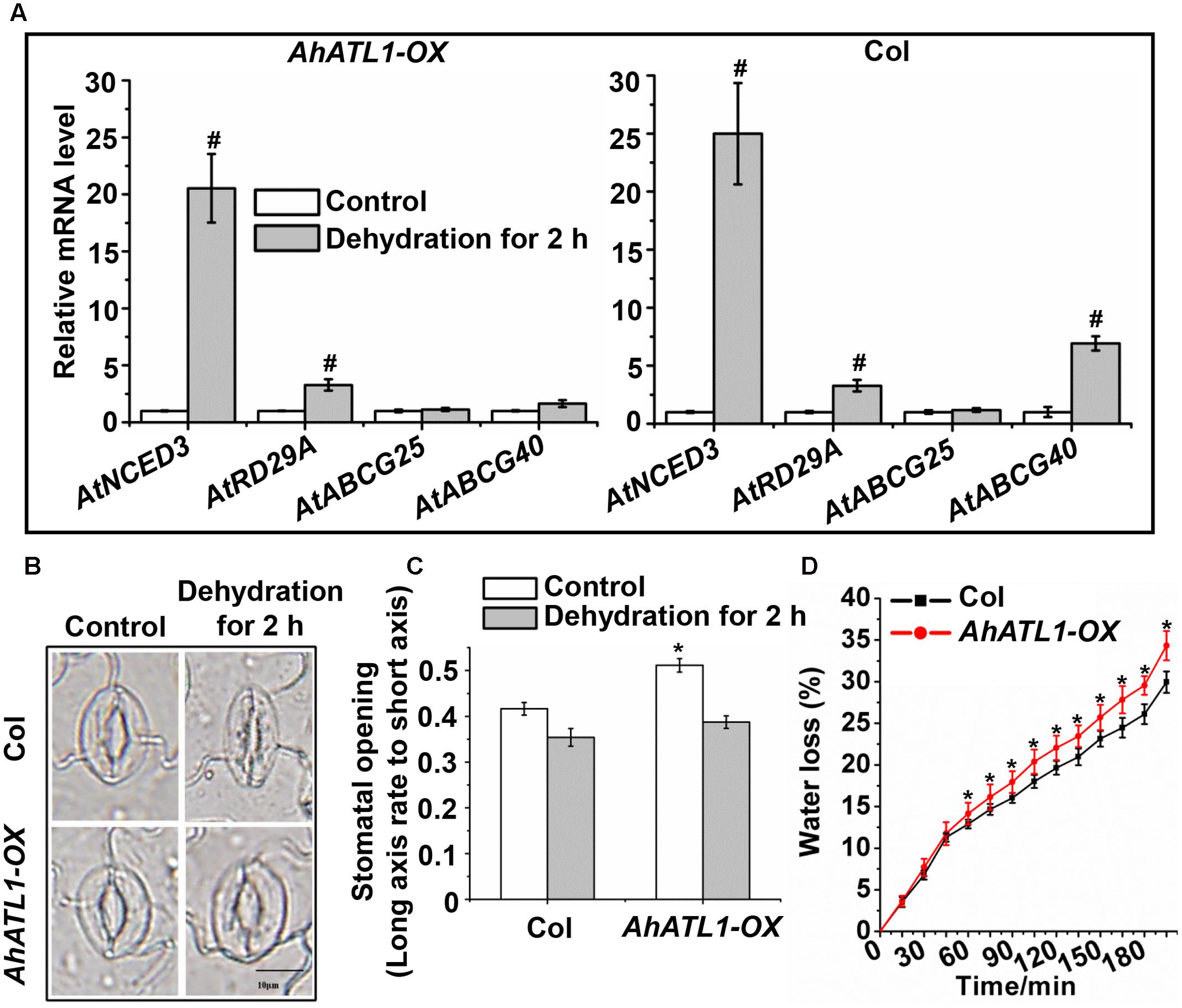
FIGURE 4. AhATL1 mediates stomatal movement by reducing expression of ABA import transporter AtABCG40 in Arabidopsis. (A) Expression of AtNCED3, AtABCG40, AtABCG25, and AtRD29A in AhATL1-OX and Col plants under normal or dehydration conditions. Three-week-old AhATL1-OX and Col plants were grown normally or under dehydration conditions for 2 h. Bars indicate standard deviation. ‘#’ indicates a significant difference in AhATL1-OX or Col plants at the level of P < 0.01 between drought stress and control treatment conditions. (B) Images (bright-field microscopy) of stomatal apertures in the leaves of 3-week-old AhATL1-OX and Col plants under control or dehydration conditions. Bar = 10 μm. (C) Stomatal opening in the leaves of 3-week-old AhATL1-OX and Col plants under control conditions or after 2 h dehydration; n = 150. ‘∗’ indicates a significant difference at the level of P < 0.05 between AhATL1-OX and Col plants under control treatment or drought stress conditions. (D) Water loss in the detached leaves of 3-week-old AhATL1-OX and Col plants under drought conditions. ‘∗’ indicates a significant difference at the level of P < 0.05.
Discussion
The plant hormone ABA regulates various physiological processes, including the responses to stress caused by heavy metals, drought, heat, high levels of salinity, low temperature and radiation (Finkelstein et al., 2002). However, its main function is to regulate plant water balance and osmotic stress tolerance (Finkelstein et al., 2002; Zhu, 2002). The endogenous ABA level is modulated by a precise balance between biosynthesis and catabolism of this hormone (Nambara and Marion-Poll, 2005). The prevailing view has been that, when a plant experiences water stress, the ABA concentration increases in the root due to a local increase in its biosynthesis, and then ABA moves via phloem transport from the roots to the leaves, where it regulates stomatal aperture (Nambara and Marion-Poll, 2005; Rodrigues et al., 2009). More recently, studies in A. thaliana report that ABA can also be transported from the leaves to the roots via the phloem when plants are subjected to water limitation or well-watered conditions (Ikegami et al., 2009; McAdam et al., 2016).
Currently, four ABA transporters have been identified in Arabidopsis; these have various transport functions depending on physiological state and tissue (Kang et al., 2010, 2015; Kuromori et al., 2010; Kanno et al., 2012). An understanding of such ABA transporters is essential if we wish to predict the patterns of ABA accumulation at different stress sites and how stomatal apertures respond to water stress. In a previous study in peanut, we found that, when roots are treated with PEG 6000, ABA is synthesized rapidly and accumulates mainly in root vascular parenchyma, after which it is quickly transported to the leaf tissues where it induces stomatal closure. When peanut leaves are treated with PEG 6000, ABA biosynthesis initially increases in the leaf, then rapidly accumulates in the vascular cambium of leaves and induces stomatal closure; ABA produced in root tissues is also transported to the leaf tissues to maintain stomatal closure (Hu et al., 2016). In the current paper, we characterized a novel peanut ABA transporter-like1 gene, AhATL1, which responds to drought stress or exogenous ABA. We found that expression of the AhATL1 gene and the corresponding protein increased rapidly after treatment with ABA or PEG 6000 (Figures 2A,B). However, when roots were pretreated with the ABA synthesis inhibitor Flu, and then subjected to water stress, AhATL1 protein was not detectable (Figure 2C). On the other hand, when pretreated with Flu and followed up with ABA treatment, the expression of AhATL1 protein was similar to that observed with PEG 6000 treatment alone (Figure 2C). These results prompted us to further explore the role of AhATL1 in ABA signaling.
ABC transporters contain four core structural domains: two of these are TMDs that are located toward the C-terminus of the protein and contain multiple (usually 4-6) membrane-spanning α-helices, which together allow transport of substrates across the lipid bilayer; there are also two NBDs, which are responsible for ATP-binding and hydrolysis (Higgins, 1992; Kerr et al., 2011). In plants, functional ABC transporters consist of a combination of two NBDs and two TMDs organized into a single transporter (full-molecule category), although the core domains may be expressed as separate polypeptides or as multidomain proteins (half-molecule category) (Higgins, 1992). The predicted amino acid sequence shows that AhATL1 is an ABC transporter with TMDs comprising a total of six transmembrane structures, as well as an ABC transporter domain (containing Walker A and Walker B motifs) and ATP-binding sites, consistent with AhATL1 belonging to the family of half-molecule ABC transporters (Figures 1A,B). In addition, protoplast transient expression experiments showed that AhATL1 is a plasma membrane protein (Figure 1C).
ABA plays a central role in the induction and maintenance of seed dormancy. It also inhibits the transition from embryonic to germination growth (Sah et al., 2016). AtABCG30 and AtABCG40 are ABA-uptake transporters and Atabcg30 and Atabcg40 mutant plants are less inhibited by 0.1–2.0 μM ABA than the corresponding Col (wild-type) seeds. Atabcg40 plants are also defective in stomatal closure, resulting in enhanced water loss and main root growth (Kang et al., 2015). Prompted by these reports, we examined the effects of ABA on the germination and seedling processes on AhATL1-OX and Col plants and found that the germination rate of AhATL1 overexpressing (AhATL1-OX) plants is higher than that of Col plants when treated with 0–2.0 μM ABA (Figure 3A). There was also more axial root growth and cotyledon greening in AhATL1-OX plants, consistent with the enhanced germination rate (Figures 3B–E). Hence, these data suggest that overexpression of AhATL1 decreases ABA sensitivity in Arabidopsis. However, Atabcg30 and Atabcg40 mutant plants were inhibited to a lesser degree by ABA than Col seeds, because there is less exogenous ABA transport into plant tissues (Kang et al., 2010, 2015). The key question, then, is why do AhATL1-OX plants show a decreased ABA sensitivity phenotype?
Endogenous ABA is rapidly produced during drought, triggering a cascade of physiological responses, including stomatal closure, which is regulated by a signal transduction network. NCED3 in Arabidopsis catalyzes a key step in ABA biosynthesis, and NCED3 expression is rapidly induced by drought stress. RD29A is known as an ABA signaling marker gene due to its rapid response to ABA treatment and drought stress (Shinozaki et al., 2003; Osakabe et al., 2014). Because AhATL1 likely functions as an ABA transporter-like protein in the ABA signaling pathway, we examined the AtNCED3, AtABCG40, AtABCG25, and AtRD29A gene transcription levels in AhATL1-OX and Col plants under normal and dehydration conditions. The expression of AtNCED3 and AtRD29A was enhanced in both AhATL1-OX and Col plants during water stress, but while AtABCG40 transcription was increased in Col plants, there was no change in expression in AhATL1-OX plants. Stomatal activity, which is affected by environmental stress, can influence CO2 absorption and water loss, and thus impact photosynthesis and plant growth (Osakabe et al., 2014). When plants experience drought stress, ABA synthesized in roots or leaves is transported into guard cells by specific transporters, such as AtABCG40, which trigger stomatal closure (Hartung et al., 2002; Bauer et al., 2013). Under dehydration stress conditions, we observed reduced stomatal closure in AhATL1-OX plants, and, consistent with this, a higher rate of water loss compared to Col plants. This effect is compounded in AhATL1-OX plants by the lower levels of AtABCG40 transcription relative to controls, and the consequently reduced amount of ABA transport into guard cells, causing a further reduction in stomatal movement and higher water loss. Taken together, these observations support the notion that AhATL1 is involved in ABA transport from synthesis sites to guard cells under dehydration conditions.
In summary, our study reveals for the first time a potential ABA transporter-like gene in peanut, which reduces ABA sensitivity by specifically influencing ABA import. This provides insight into the spatial regulation of the ABA transport system and its regulatory networks in A. hypogaea.
Author Contributions
KG, BH, and LL designed the research. KG and XL performed the research. KG, XL, and XyL analyzed the data and prepared figures. KG wrote the manuscript in consulation with BH, XyL, and LL.
Funding
This work was supported by Natural Science Foundation of Guangdong Province (8151063101000011 to LL), Innovation Project of Graduate School of South China Normal University (2016lkxm39 to KG).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
We are grateful to Prof. Alan Tunnacliffe (Chief Editor) at Cambridge Academic Manuscripts, for technical support and critical reading of the manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2017.01150/full#supplementary-material
Footnotes
References
Bauer, H., Ache, P., Lautner, S., Fromm, J., Hartung, W., Al-Rasheid, K. A., et al. (2013). The stomatal response to reduced relative humidity requires guard cell-autonomous ABA synthesis. Curr. Biol. 23, 53–57. doi: 10.1016/j.cub.2012.11.022
Boursiac, Y., Léran, S., Corratgé-Faillie, C., Gojon, A., Krouk, G., and Lacombe, B. (2013). ABA transport and transporters. Trends Plant Sci. 18, 325–333.
Finkelstein, R. (2013). Abscisic acid synthesis and response. Arabidopsis Book 11:e0166. doi: 10.1199/tab.0166
Finkelstein, R. R., Gampala, S. S., and Rock, C. D. (2002). Abscisic acid signaling in seeds and seedlings. Plant Cell 14(Suppl. 1), S15–S45.
Govind, G., ThammeGowda, H. V., Kalaiarasi, P. J., Iyer, D. R., Muthappa, S. K., Nese, S., et al. (2009). Identification and functional validation of a unique set of drought induced genes preferentially expressed in response to gradual water stress in peanut. Mol. Genet. Genomics 281, 591–605. doi: 10.1007/s00438-009-0432-z
Hartung, W., Sauter, A., and Hose, E. (2002). Abscisic acid in the xylem: where does it come from, where does it go to? J. Exp. Bot. 53, 27–32.
Higgins, C. F. (1992). ABC transporters: from microorganisms to man. Annu. Rev. Cell Biol. 8, 67–113.
Hu, B., Cao, J., Ge, K., and Li, L. (2016). The site of water stress governs the pattern of ABA synthesis and transport in peanut. Sci. Rep. 6:32143. doi: 10.1038/srep32143
Hwang, J. U., Song, W. Y., Hong, D., Ko, D., Yamaoka, Y., Jang, S., et al. (2016). Plant ABC transporters enable many unique aspects of a terrestrial plant’s lifestyle. Mol. Plant 9, 338–355. doi: 10.1016/j.molp.2016.02.003
Ikegami, K., Okamoto, M., Seo, M., and Koshiba, T. (2009). Activation of abscisic acid biosynthesis in the leaves of Arabidopsis thaliana in response to water deficit. J. Plant Res. 122, 235–243. doi: 10.1007/s10265-008-0201-9
Kang, J., Hwang, J. U., Lee, M., Kim, Y. Y., Assmann, S. M., Martinoia, E., et al. (2010). PDR-type ABC transporter mediates cellular uptake of the phytohormone abscisic acid. Proc. Natl. Acad. Sci. U.S.A. 107, 2355–2360. doi: 10.1073/pnas.0909222107
Kang, J., Yim, S., Choi, H., Kim, A., Lee, K. P., Lopez-Molina, L., et al. (2015). Abscisic acid transporters cooperate to control seed germination. Nat. Commun. 6:8113. doi: 10.1038/ncomms9113
Kanno, Y., Hanada, A., Chiba, Y., Ichikawa, T., Nakazawa, M., Matsui, M., et al. (2012). Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc. Natl. Acad. Sci. U.S.A. 109, 9653–9658. doi: 10.1073/pnas.1203567109
Kerr, I. D., Haider, A. J., and Gelissen, I. C. (2011). The ABCG family of membrane-associated transporters: you don’t have to be big to be mighty. Br. J. Pharmacol. 164, 1767–1779. doi: 10.1111/j.1476-5381.2010.01177.x
Kuromori, T., Miyaji, T., Yabuuchi, H., Shimizu, H., Sugimoto, E., Kamiya, A., et al. (2010). ABC transporter AtABCG25 is involved in abscisic acid transport and responses. Proc. Natl. Acad. Sci. U.S.A. 107, 2361–2366. doi: 10.1073/pnas.0912516107
Li, J., Li, Y., Yin, Z., Jiang, J., Zhang, M., Guo, X., et al. (2017). OsASR5 enhances drought tolerance through a stomatal closure pathway associated with ABA and H2O2 signalling in rice. Plant Biotechnol. J. 15, 183–196. doi: 10.1111/pbi.12601
Li, T., Wu, X. Y., Li, H., Song, J. H., and Liu, J. Y. (2016). A dual-function transcription factor, AtYY1, is a novel negative regulator of the Arabidopsis ABA response network. Mol. Plant 9, 650–661. doi: 10.1016/j.molp.2016.02.010
Li, X., Li, X., Li, M., Yan, Y., Liu, X., and Li, L. (2016). Dual function of NAC072 in ABF3-mediated ABA-responsive gene regulation in Arabidopsis. Front. Plant Sci. 7:1075. doi: 10.3389/fpls.2016.01075
Li, X. Y., Liu, X., Yao, Y., Li, Y. H., Liu, S., He, C. Y., et al. (2013). Overexpression of Arachis hypogaea AREB1 gene enhances drought tolerance by modulating ROS scavenging and maintaining endogenous ABA content. Int. J. Mol. Sci. 14, 12827–12842. doi: 10.3390/ijms140612827
McAdam, S. A., Brodribb, T. J., and Ross, J. J. (2016). Shoot-derived abscisic acid promotes root growth. Plant Cell Environ. 39, 652–659. doi: 10.1111/pce.12669
Nambara, E., and Marion-Poll, A. (2005). Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 56, 165–185.
Nautiyal, P. C., Rachaputi, N. R., and Joshi, Y. C. (2002). Moisture-deficit-induced changes in leaf-water content, leaf carbon exchange rate and biomass production in groundnut cultivars differing in specific leaf area. Field Crops Res. 74, 67–79.
Osakabe, Y., Osakabe, K., Shinozaki, K., and Tran, L. S. P. (2014). Response of plants to water stress. Front. Plant Sci. 5:86. doi: 10.3389/fpls.2014.00086
Puértolas, J., Conesa, M. R., Ballester, C., and Dodd, I. C. (2015). Local root abscisic acid (ABA) accumulation depends on the spatial distribution of soil moisture in potato: implications for ABA signalling under heterogeneous soil drying. J. Exp. Bot. 66, 2325–2334. doi: 10.1093/jxb/eru501
Rodrigues, A., Santiago, J., Rubio, S., Saez, A., Osmont, K. S., Gadea, J., et al. (2009). The short-rooted phenotype of the brevis radix mutant partly reflects root abscisic acid hypersensitivity. Plant Physiol. 149, 1917–1928. doi: 10.1104/pp.108.133819
Sah, S. K., Reddy, K. R., and Li, J. (2016). Abscisic acid and abiotic stress tolerance in crop plants. Front. Plant Sci. 7:571. doi: 10.3389/fpls.2016.00571
Shinozaki, K., Yamaguchi-Shinozaki, K., and Seki, M. (2003). Regulatory network of gene expression in the drought and cold stress responses. Curr. Opin. Plant Biol. 6, 410–417.
Su, L. C., Deng, B., Liu, S., Li, L. M., Hu, B., Zhong, Y. T., et al. (2015). Isolation and characterization of an osmotic stress and ABA induced histone deacetylase in Arachis hypogaea. Front. Plant Sci. 6:512. doi: 10.3389/fpls.2015.00512
Zhao, Y., Chan, Z., Gao, J., Xing, L., Cao, M., Yu, C., et al. (2016). ABA receptor PYL9 promotes drought resistance and leaf senescence. Proc. Natl. Acad. Sci. U.S.A. 113, 1949–1954. doi: 10.1073/pnas.1522840113
Keywords: peanut, AhATL1, ABA transporter, AtABCG40, stomatal movement, drought stress
Citation: Ge K, Liu X, Li X, Hu B and Li L (2017) Isolation of an ABA Transporter-Like 1 Gene from Arachis hypogaea That Affects ABA Import and Reduces ABA Sensitivity in Arabidopsis. Front. Plant Sci. 8:1150. doi: 10.3389/fpls.2017.01150
Received: 08 April 2017; Accepted: 15 June 2017;
Published: 30 June 2017.
Edited by:
Junhua Peng, Center for Life Sci&Tech of China National Seed Group Co., Ltd., ChinaReviewed by:
Liang Chen, University of Chinese Academy of Sciences, ChinaHong Luo, Clemson University, United States
Copyright © 2017 Ge, Liu, Li, Hu and Li. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Bo Hu, aHVibzA2MTBAMTI2LmNvbQ== Ling Li, bGlsaW5nNTAyQDEyNi5jb20=
 Kui Ge
Kui Ge Xing Liu
Xing Liu Xiaoyun Li
Xiaoyun Li