- 1Department of Agricultural, Food and Environmental Sciences, Università Politecnica delle Marche, Ancona, Italy
- 2J. T. Environmental Consultants Ltd, Cambridge, United Kingdom
The improvement of woody fruit species by traditional plant breeding techniques has several limitations mainly caused by their high degree of heterozygosity, the length of their juvenile phase and auto-incompatibility. The development of new biotechnological tools (NBTs), such as RNA interference (RNAi), trans-grafting, cisgenesis/intragenesis, and genome editing tools, like zinc-finger and CRISPR/Cas9, has introduced the possibility of more precise and faster genetic modifications of plants. This aspect is of particular importance for the introduction or modification of specific traits in woody fruit species while maintaining unchanged general characteristics of a selected cultivar. Moreover, some of these new tools give the possibility to obtain transgene-free modified fruit tree genomes, which should increase consumer's acceptance. Over the decades biotechnological tools have undergone rapid development and there is a continuous addition of new and valuable techniques for plant breeders. This makes it possible to create desirable woody fruit varieties in a fast and more efficient way to meet the demand for sustainable agricultural productivity. Although, NBTs have a common goal i.e., precise, fast, and efficient crop improvement, individually they are markedly different in approach and characteristics from each other. In this review we describe in detail their mechanisms and applications for the improvement of fruit trees and consider the relationship between these biotechnological tools and the EU biosafety regulations applied to the plants and products obtained through these techniques.
Introduction
Conventional breeding for genetic improvement of woody fruit crops is a slow and difficult process, with drawbacks caused by high heterozygosity, extended juvenile periods, and auto-incompatibility (Petri and Burgos, 2005; Rai and Shekhawat, 2014). Furthermore, improvement of woody fruit species using conventional breeding methods is a long-term process because of their long generation time. New biotechnological tools (NBTs) including genetic engineering methods can promote the prompt insertion of important genes into the genome of commercial woody fruit cultivars, thus resulting in more efficient and reliable genetic improvement (Lusser et al., 2012) of clonal propagated plants, maintaining high stability of the major traits of the clone. The introduction of recombinant DNA technology paved the way for an immense potential in the field of plant biotechnology. In order to attain food security and to guarantee nutritional quality, NBTs for generating genetically modified (GM) plants with useful agronomic and quality traits are already of high significance for many crops (Datta, 2013; Qaim and Kouser, 2013).
Genetic engineering in plants has been in practice for more than three decades. Direct transformation methods (Biolistic) and indirect methods (Agrobacterium tumefaciens-mediated transformation), developed decades ago, have been the primary strategies of heterologous DNA introduction into plants (Chilton et al., 1977; Gelvin, 2003; Altpeter et al., 2005). All genetically modified crops commercially grown, including woody fruit species, were produced using one of these methods (Parisi et al., 2016). Often the ability to obtain fruit tree plants with new traits or mutations by genetic engineering or by NBTs depends on the existence of a well-established in vitro regeneration protocol, which depends on the genotype and the type of starting plant tissue used (Wang et al., 2011; Rai and Shekhawat, 2014; Saporta et al., 2017). Furthermore, it is more advisable from an agronomic point of view to in vitro regenerate a new fruit tree plant from mature tissues, due to the high degree of heterozygosity, which characterize the majority of these species (Cervera et al., 1998; Pérez-Jiménez et al., 2012). In this sense relevant progress have been made during the last two decades for some difficult-to-transform woody species, such as peach or grapevine genotypes, in which efficient protocols for the regeneration of adventitious shoots have been developed starting from adult tissues (Mezzetti et al., 2002; Pérez-Jiménez et al., 2012; Sabbadini et al., 2015). Introduction of one or more new genes or regulatory elements using genetic engineering techniques, directly manipulates the genome of an organism in order to express or silence specific traits (Tzfira and Citovsky, 2006; Mittler and Blumwald, 2010; Rai and Shekhawat, 2014). Transgenic approaches having global impact are aimed mainly at the production of crops with new resistance genes against pests and diseases, or herbicide tolerance, such as Monsanto's roundup ready crops (soya, maize, and cotton; Funke et al., 2006; Lombardo et al., 2016; Parisi et al., 2016), and plants with enhanced desirable qualities and nutritional levels, such as the golden rice with an increased vitamin A content (Paine et al., 2005; Bhullar and Gruissem, 2013; Pérez-Massot et al., 2013; Zhu et al., 2013; Giuliano, 2017).
In woody fruit species, the use of conventional plant breeding techniques such as traditional mutation, translocation breeding, and intergeneric crosses, is very limiting due to the non-specific approaches often leading to mutation of thousands of untargeted nucleotides instead of the single desired one or the transfer of a large part of the genome instead of a single gene (Hartung and Schiemann, 2014). It is for this reason that gene transfer, site-specific integration, and specific regulation of gene expression are crucial advancements in plant biotechnology (Datta, 2013). In this review we describe the mechanisms of the more advanced biotechnological techniques and their application in woody fruit species improvement.
NBTs used for modifying an existing DNA sequence in a plant, comprise of insertion/deletion and gene replacement, or stable silencing of a gene or promoter sequence. In this category we consider techniques such as RNA interference (RNAi), cisgenesis/intragenesis, trans-grafting, and gene editing techniques including zinc finger nucleases (ZFNs) as well as clustered regularly interspaced short palindromic repeats/CRISPR-associated protein 9 (CRISPR/Cas9 system), to introduce new traits into a host plant genome. All these technologies have been successfully applied in different crops, but there are still limited applications in woody fruit species.
Cisgenesis and Intragenesis
The term cisgenesis was introduced by Schouten et al. (2006a), defining it as the genetic modification of plants using genes that originate only from the species itself or from a species that can be crossed conventionally with this species. The added gene is an extra copy to the existing genome and is a natural variant, which includes its introns, flanking native promoter and terminator in normal sense orientation (Lusser and Davies, 2013). In intragenesis, the introduced genetic element (intragene) originates from the same species or a species from a sexually compatible gene pool. The intragenes are considered hybrid genes since they can be driven by different promoter or terminator regions of different genes and loci (Rommens, 2007). The inserted DNA sequence will form a new arrangement of genetic elements leading to a modified functional version compared to the starting genome (Conner et al., 2007). Furthermore, in intragenic plants, when using Agrobacterium-mediated transformation as strategy to insert the new trait, plant-derived transfer DNA (P-DNA) borders sequences from the sexually compatible DNA pool are used in order to avoid accidental insertion of vector sequences (Rommens, 2004). Thus, it is possible to obtain transformed plants which do not contain any foreign DNA. These approaches avoid the potential for “linkage drag” (the transfer of other undesirable genes along with the gene of interest), associated with classical introgression in conventional breeding (Jacobsen and Schouten, 2007). Whole genomic sequencing studies are providing information on the cisgenes that can be used for genetic improvement of specific crops, but in many cases the availability of cisgenic promoters and efficient marker genes are limited. An illustration of the two techniques is shown in Figure 1.
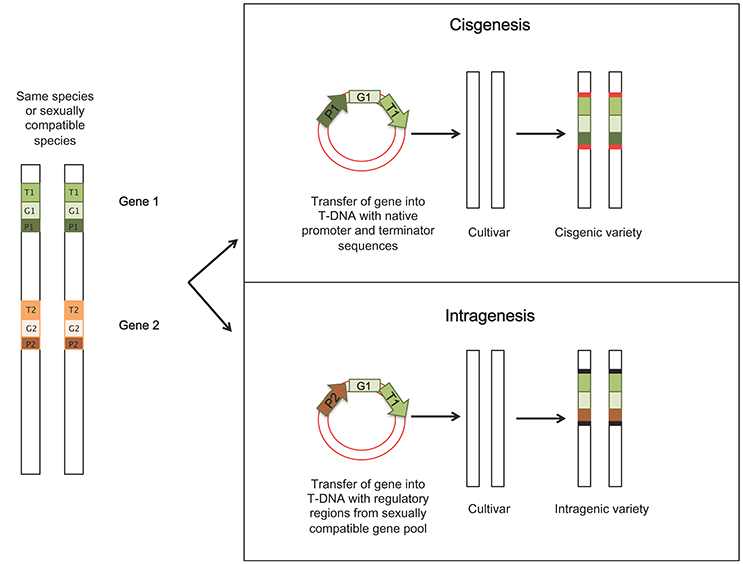
Figure 1. Illustration of principles and procedures of obtaining cisgenic and intragenic crops. In cisgenesis, the new trait is derived from a sexually compatible species and it is transferred to the recipient as it is, including the Agrobacterium-derived T-DNA borders; in intragenesis, the gene construct is a hybrid of different components from different genes within the same species or sexually compatible species. Red boxes: Agrobacterium-derived T-DNA borders; black boxes: borders belonging to sexually compatible DNA pool (P-DNA borders), when using Agrobacterium-mediated transformation. P, promoter; G, engineered gene; T, terminator.
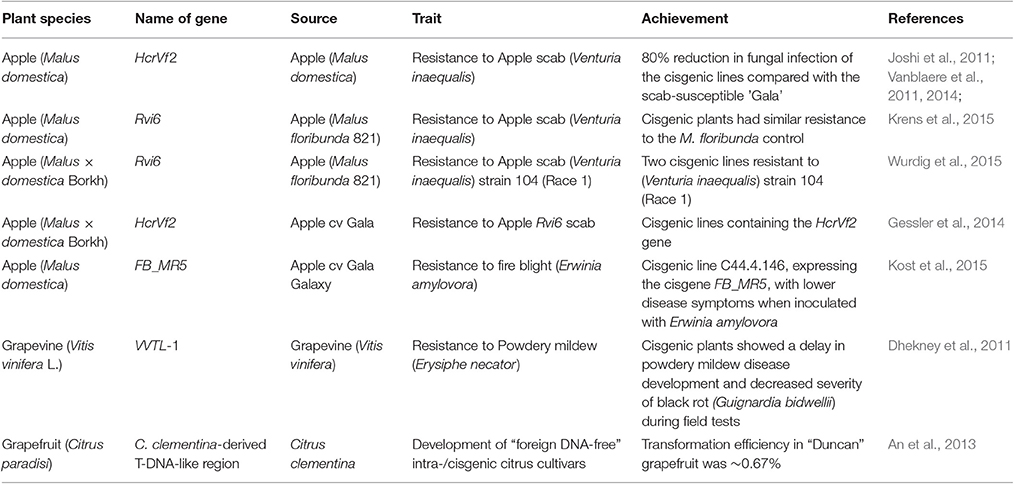
Cisgenesis/intragenesis has been applied in different woody fruit species including apples. Fruit breeders are developing solutions for the various diseases affecting apples, including fire blight disease caused by Erwinia amylovora. Kost et al. (2015) recently developed a cisgenic apple line C44.4.146 from a fire blight susceptible cultivar “Gala Galaxy” using the cisgene FB_MR5 from wild apple Malus × robusta 5 (Mr5), which confers resistance to fire blight (Peil et al., 2007). After elimination of the selectable markers through heat-induced recombinase, both PCR and Southern blot analysis did not detect any transgenes. The transformed line C44.4.146 carried just the cisgene FB_MR5 and its native regulatory sequences (Kost et al., 2015). Cisgenesis and intragenesis have also been successfully applied to induce resistance to other diseases in both apple and other woody fruit tree and vines, as shown in Table 1.
RNA Interference (RNAi)
The first discovery of the silencing phenomenon in plants was made in 1990, by scientists trying to deepen the purple color of petunias through the overexpression of Chalcone synthase gene. Contrary to their expectations, the flowers became white indicating that the gene had been turned off (Napoli et al., 1990; Metzlaff et al., 1997). The suppression of endogenous gene expression through the introduction of a homologous sequence into the genome was referred to as “co-suppression” in petunia (Campbell and Choy, 2005), later correlated to the phenomenon of post-transcriptional gene silencing (PTGS).
RNAi is an endogenous cellular process that occurs naturally to “turn off” unwanted or harmful specific nucleic sequences, or to regulate gene expression before translation (Baum et al., 2007; De Alba et al., 2013). RNAi has been discovered and studied in many organisms such as fungi, animals, and ciliates (Romano and Macino, 1992; Fire et al., 1998; Billmyre et al., 2013; Scott et al., 2013), and has more recently been studied in plants (Matzke et al., 2001; Baulcombe, 2004; Ipsaro and Joshua-Tor, 2015).
RNAi refers to a complex of molecular mechanisms, which have the main function of gene expression inhibition or suppression, activated by the presence of double-stranded RNA molecules (dsRNAs; Voinnet, 2008; Parent and Vaucheret, 2012). The discovery of this process led to the possibility of creating custom “knock-downs” of gene activity. In both plants and animals it has been shown that RNAi utilizes the dsRNAs as trigger molecules that detect homologous mRNAs, whose transcription is negatively regulated (Almeida and Allshire, 2005; Ketting, 2011; Ipsaro and Joshua-Tor, 2015). Consequently, RNA silencing has emerged as a preferred method for gene targeting in fungi (Nakayashiki, 2005; Salame et al., 2011), insects (Scott et al., 2013), bacteria (Escobar et al., 2001; Navarro et al., 2006), viruses (Baulcombe, 2004; Ding, 2010), and plants (Brodersen and Voinnet, 2006; Frizzi and Huang, 2010). Presently, there are several routes of gene silencing identified in plants, these include: PTGS (Vaucheret et al., 2001; Borges and Martienssen, 2015), transcriptional gene silencing (TGS; Vaucheret and Fagard, 2001; De Alba et al., 2013), and microRNA silencing (miRNA; Bartel, 2004; Jonas and Izaurralde, 2015). All these pathways rely on the presence of dsRNA molecules of different sizes, which are processed into the plant cell by specific protein families, i.e., Dicer or Dicer-like (DCL), Argonaute (AGO), and RNA-dependent RNA polymerases (RDRs; Molnar et al., 2011).
Long dsRNAs constitute the precursors for siRNA molecules (siRNAs) production, thanks to specific Dicer enzymes, whose action determines their final length. DCL1 is responsible for the creation of miRNAs, which originate in the plant cell's nucleus from endogenous precursors characterized by a stem-loop with imperfect double-strand structure (Voinnet, 2009). Twenty-two to twenty-four nucleotide long siRNAs are produced into the cell's cytoplasm by the action of DCL2, which cleaves exogenous long dsRNAs originated from viral intermediates or transgenes (Meister and Tuschl, 2004; Carthew and Sontheimer, 2009). DCL3 cleaves long dsRNAs transcribed in the nucleus by the plant RNA polymerase Pol IV, to produce 24-nt long siRNAs with the main function of heterochromatin modifications (Qi and Hannon, 2005; Brosnan and Voinnet, 2011; Matzke and Mosher, 2014). Finally, DCL4 is responsible for the production of 21-nt secondary siRNAs involved in cell-to-cell silencing signaling (Dunoyer et al., 2005). The siRNAs produced through the various RNAi pathways, are unwound into the passenger and the guide strand; the latter is bound to Argonaute (AGO) proteins to form the nucleus of the RNA-Induced Silencing Complex (RISC). It has been observed that AGO 1, AGO2, AGO 7, and AGO10 bind to siRNAs to induce degradation of the complementary mRNAs or to inhibit translation (Brodersen et al., 2008). The association of siRNAs with other types of AGO proteins (4, 6, or 9) can activate the TGS mechanism, which has the main role of inducing epigenetic changes by chromatin modifications and histone methylation through the RNA-directed DNA methylation (RdDM) pathway (Xie et al., 2004; Brosnan and Voinnet, 2011; Matzke and Mosher, 2014). The silencing signal can be amplified through the action of an RNA-dependent RNA polymerase (RdRP), which helps the perpetuation of the silencing response by the synthesis of secondary siRNAs (Figure 2). This pool of secondary molecules can induce systemic silencing in the plant (Meister and Tuschl, 2004; Carthew and Sontheimer, 2009). The silencing signal, constituted mainly by 21 and 24 nt long siRNAs, has the ability to move from cell to cell and systematically over long distances inside the plant, with the principal role of defense from invasive nucleic acids or to induce epigenetic modification (Molnar et al., 2011). In particular, the local cell to cell movement of siRNAs takes place through the plasmodesmata or by apoplastic transfer; while, the systemic movement occurs through the vascular system, generally starting from a photosynthetic source to end in a sucrose sink (Melnyk et al., 2011).
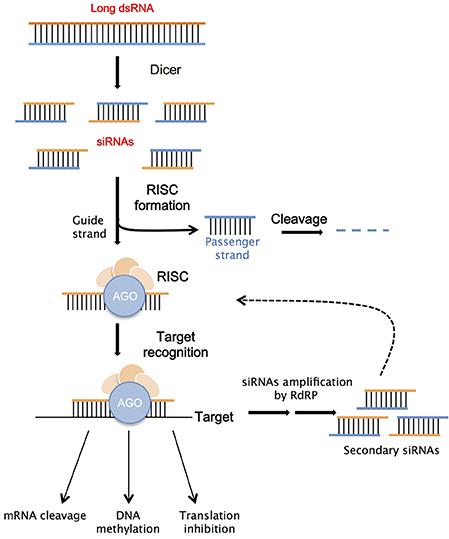
Figure 2. Schematic illustration of RNAi mechanism. Double stranded RNA (dsRNA) molecule binds to a Dicer protein, which cleaves it into small interfering RNAs (siRNAs); these siRNAs bind to an Argonaute (AGO) protein, part of the RNA-Induced Silencing Complex (RISC). The RISC separates the siRNAs into two strands: the passenger strand (blue) is degraded while the guide strand (orange) serves as a search probe, which links RISC to complementary RNA targets. After this recognition target's expression can be regulated through several different mechanisms. In plants, the silencing signal can be perpetuated by the action of the RNA-dependent RNA polymerase (RdRP).
The most recent discovery in RNA silencing is the cross—talk occurring between kingdoms (Knip et al., 2014). Studies carried out on plants and their fungal pathogens in the laboratory indicate that both parties can move RNAs back and forth into each other's cells. Fungal microbes utilize RNAi to enhance their spread whereas, plants seem to use this mechanism to encounter infection by these pathogens. In both cases desired outcome is achieved through the same molecular process of RNA interference, which interrupts gene expression through target messenger RNA degradation (Cheng et al., 2015; Grens, 2017).
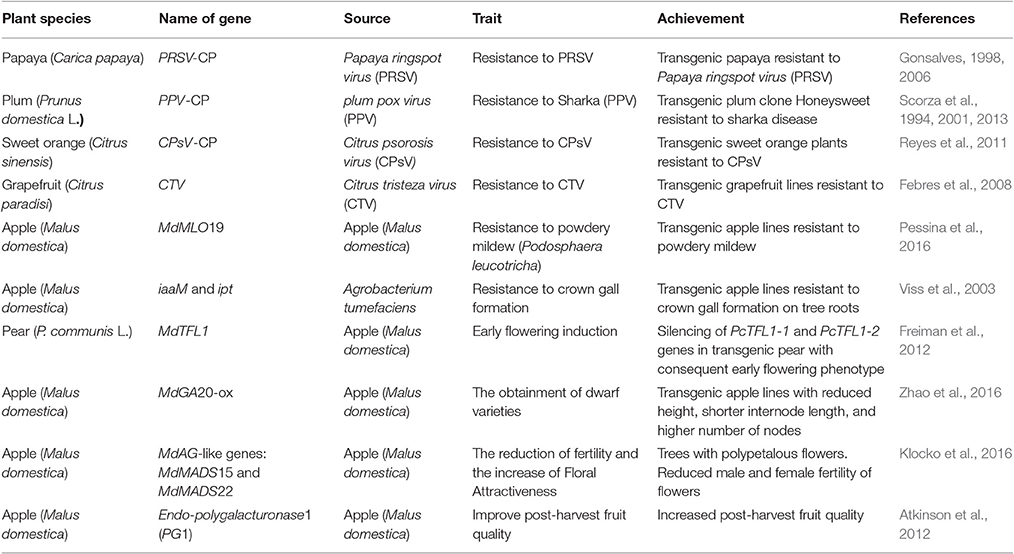
In plants, RNA silencing affects the regulation of endogenous gene expression, and it is also an evolutionary conserved mechanism that serves as host defense against viruses (Ding and Voinnet, 2007; Carbonell and Carrington, 2015). RNAi has been mainly applied in woody fruit species to induce pathogen resistance. Pathogen derived resistance (PDR) is based on the expression of pathogen genetic elements (Sanford and Johnston, 1985; Baulcombe, 1996) which has led to various forms of plant virus resistance (Simón-Mateo and García, 2011). One of the first applications of this approach was the induction of virus resistance through the introduction of gene constructs expressing viral sequences, such as coat protein (CP), movement protein, and replicase (Abel et al., 1986; Baulcombe, 1996; Gottula and Fuchs, 2009). Subsequently several studies showed that the virus resistant phenotypes were often based on the induction of an RNA-mediated mechanism and not on a protein-mediated resistance (English et al., 1996; Hannon, 2002). In early 1990s' a regeneration and transformation protocol to genetically engineer papaya for PRSV resistance, the most widespread and damaging virus disease of papaya, was developed. The objective was to introduce a gene construct that codes for a chimeric coat protein (CP) containing 17 amino acids of the Cucumber mosaic virus and the N terminus of the CP gene of PRSV HA 5-1. The inhibition of PRSV obtained in one of the transgenic papaya lines showed an RNA-mediated resistance (Gonsalves, 1998, 2006). Another case of virus resistance in fruit trees, based on RNA-mediated mechanism of PDR, is represented by the transgenic plum clone Honeysweet resistant to sharka or plum pox disease (Scorza et al., 2013). Sharka is considered one of the most devastating diseases in stone fruits and is caused by the plum pox virus (PPV; Cambra et al., 2006). PPV-resistant plum was obtained using hypocotyl slices as starting explants, which were transformed with the coat protein gene of plum pox virus (PPV-CP). The integration of the engineered CP gene was confirmed in five transgenic lines (Scorza et al., 1994). After years of testing, it was shown that the high resistance to PPV of the transgenic clone C5 displayed the typical characteristics of PTGS mechanism (Scorza et al., 2001); 24-nt long siRNAs were detected in the resistant clone when infected by PPV, which were considered responsible for the Honeysweet PPV resistance. dsRNA was demonstrated to be the triggering molecule of RNA silencing, and virus-resistant plants were obtained through a second generation technology based on the introduction of inverted repeat RNA (hairpin RNA) or intron—hairpin-RNA (ihpRNA) constructs into cells, which are able to provide efficient virus resistance by eliciting PTGS in the host plant (Smith et al., 2000; Collinge et al., 2010; Khalid et al., 2017). Hily et al. (2007) demonstrated for the first time that ihpRNA technology could be exploited to obtain virus resistance in Prunus domestica. Table 2 shows applications of RNAi technique in other woody fruit species performed over the years with the purpose of inducing disease resistance, post-harvest quality improvement as well as gene functional studies.
Trans-grafting Technique
This technique focuses mainly on grafting, a horticultural technique that has been practized for centuries to improve the quality and yield of fruit crops (Melnyk and Meyerowitz, 2015). The method pairs two autonomous genotypes selected individually for their rooting ability and fruiting characteristics. They are grafted together in order to combine their superior traits in the scion and the rootstock. It has been extensively used to improve crop quality and productivity and also to propagate woody perennial crops like fruits, and ornamental plants (Mudge et al., 2009). The rootstock can alter the phenotype of the scion, for example by reducing its vigor and encouraging more fruit set, but the rootstock and scion retain their genetic integrity, in that the grafted tissues are joined but their genetic materials do not mix. Other tissue grafting techniques include applications ranging from plant breeding to animal organ transplants.
Traditionally grafting is used to improve disease resistance, in particular against soil-borne fungi and bacteria, and growth characteristics such as rooting ability, nutrient, and water acquisition (Haroldsen et al., 2012b). Trans grafting is a method which combines traditional grafting practices with genetic modification of plants. The technique involves grafting of a non-genetically modified scion onto a genetically modified rootstock. The scion acquires benefits and traits conferred by transgenes in the rootstock, but the end products, such as fruits, do not contain the transgene and hence are not genetically modified (Schaart and Visser, 2009; Haroldsen et al., 2012a; Lemgo et al., 2013).
The movement of RNA molecules through the vascular system from the rootstock to the scion is at the basis of trans-grafting technique (Mallory et al., 2003; Lucas and Lee, 2004; Stegemann and Bock, 2009; Haroldsen et al., 2012a). Higher plants function as integrated organisms due to long-distance transport of signaling molecules through phloem, which has emerged as a major communication mechanism that ensures synchronized differentiation and supply of nutrients (McGarry and Kragler, 2013). For example, Lang et al. (1977) demonstrated the mobility of florigen in tobacco plants for the promotion and inhibition of flower formation in a neutral-day plant by grafting with a short-day plant and a long-day plant. This discovery helped in understanding the regulation and coordination of tissue formation by plants, making it possible to manipulate flowering time and meiosis thereby controlling crop breeding processes. Recent studies have shown that phloem transports some specific RNA molecules to coordinate organ development (Palauqui et al., 1997; Melnyk et al., 2011; Nazim and Kim, 2013). Research of functional analyses of phloem shows that over 15% of the transcripts are signal transduction related (Omid et al., 2007). If RNAi-based rootstocks can efficiently transfer the silencing molecules to non-transformed scions, then RNAi can be applied to obtain virus resistant transgenic plants (Schaart and Visser, 2009; Lemgo et al., 2013). Recent researches show that siRNA molecules derived from hairpin gene constructs can spread between cells and systemically over long distances (i.e., 1.2 m above the graft union) in woody plants (Haroldsen et al., 2012a; Zhao and Song, 2014), and can induce direct epigenetic modifications at the DNA level of the recipient cells in Arabodopsis thaliana (Molnar et al., 2010). In addition, microRNAs and trans-acting siRNAs have been associated with the transmission of silencing signals systemically via phloem and from cell to cell through the plasmodesmata (Nazim and Kim, 2013; Zhao and Song, 2014). Compatibility is important for scion-rootstock interactions for the downward flow of photosynthesis products and upward movement of water and mineral nutrients (Aloni et al., 2010), as well as for the transmission of the RNAi silencing signal into the scion, and initiation of systemic silencing.
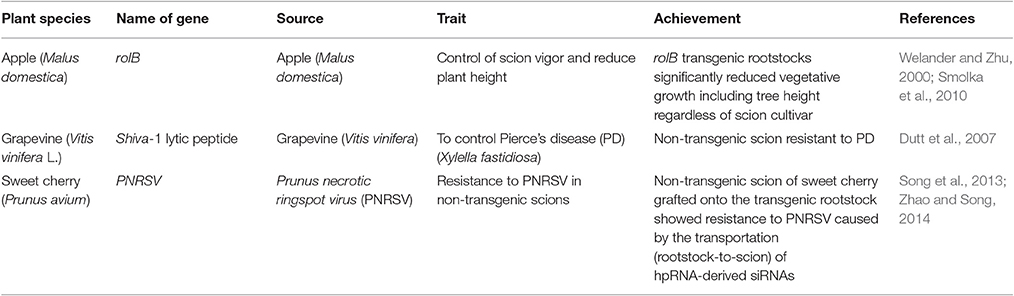
Genetically modified rootstocks have the potential to boost production of standard, non-genetically modified fruit varieties while avoiding concerns about transgene flow and exogenous protein production that occur in other types of transformed fruits (Haroldsen et al., 2012a; Song et al., 2013). Considering the applications of this technique (Table 3), it is evident that the use of genetically modified rootstocks for grafting might be the answer to disease control in many woody fruit species through the production of healthy non-genetically modified fruits. These fruits should not need the level of biosafety scrutiny normally required for traditional genetically modified plants.
Gene Editing Techniques
A decade ago, a new approach emerged that makes it possible for researchers to manipulate almost any gene in different cell types and organisms. This fundamental technique, commonly referred to as “genome editing” integrates, deletes, and/or mutates genes of interest. Engineered nucleases composed of sequence-specific DNA-binding domains attached to non-specific DNA cleavage modules are at the heart of genome editing techniques (Urnov et al., 2010). The potential to manipulate genetic information in a precise manner and obtain improved plants not only provides the opportunity to create novel phenotypes but also enables biological mechanism and gene function studies. The ability to cleave specific DNA sequences and to induce different DNA repair mechanisms allows for a range of genomic modifications from single-nucleotide mutations to large sequence deletions, rearrangements and/or integrations (Figure 3; Curtin et al., 2012).
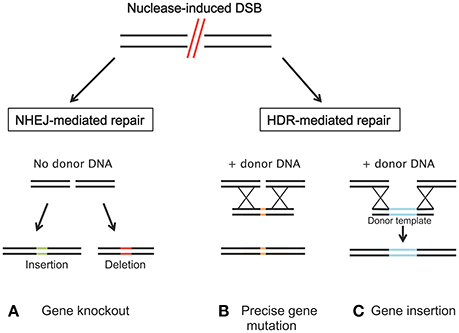
Figure 3. Induced double strand breaks (DSBs) of the target DNA by nucleases can be repaired by either non-homologous end-joining (NHEJ) or homology directed repair (HDR). (A) NHEJ usually leads to gene knockout by the insertion (green) or deletion (red) of random base pairs. (B) If a donor template, that shares regions of homology to the sequence next to the DSB is available, HDR can introduce precise gene modification or (C) specific nucleotide/gene insertion.
Zinc Finger Nuclease (ZFN) and TALE Nucleases (TALENs)
After the discovery of the functional principles of the Cys2-His2 zinc finger (ZF) motifs, and of truncated transcription activator-like effector (TALE) domains, a first generation of engineered endonucleases (EENs), zinc-finger nucleases (ZFNs), and TALE nucleases (TALENs), were developed (Pabo et al., 2001; Wood et al., 2011). Both ZFNs and TALENs can easily target DNA cleavage and they have been adopted as tools for making directed genetic changes, as they also facilitate the rearrangement of their DNA-binding domain. They are designed and utilized for generating double-strand breaks (DSBs) at almost any specific genomic position to enable genome editing (Urnov et al., 2010; Gaj et al., 2013). DSBs are subsequently exposed to cellular DNA repair mechanisms, which include error-prone non-homologous end joining (NHEJ) and homology-directed repair (HDR), that lead to high frequencies of both targeted mutagenesis, genome editing and targeted gene replacement/integration (Carroll, 2011; Petolino, 2015; Figure 3). ZFNs and TALENs are composed of two proteins, one that is necessary for DNA targeting and binding, which can be engineered to target specific DNA sequences, and a fused nuclease, usually FokI, which cuts the target DNA in a non-specific manner; the two proteins are artificially connected by a peptide linker (Hartung and Schiemann, 2014). Zinc finger proteins (ZFPs) bind to their DNA targets as monomers (fingers), each of which recognizes 3 bp of DNA. ZFNs of 3–6 monomers can be used to target specific DNA sequences of about 9–18-bases long as shown in Figure 4. In the case where a longer target sequence needs to be edited, longer ZFN recognition sequences (24–36 bp) are required for binding to achieve a higher specificity level and reduction of off-site cleavage (Miller et al., 2007; Petolino, 2015). In contrast with ZFNs, TALENs are characterized by DNA binding domains composed by repeats of 33–35 amino acids, each of which is able to recognize a single DNA base pair. This represents an advantage in terms of design flexibility (Gaj et al., 2013). The target specifity of TALENs relies on the presence of two amino acids called repeat-variable di-residues (RVDs; Deng et al., 2012). As with ZFNs, TALENs can also act as modular repeats to target adjoining DNA sequences.
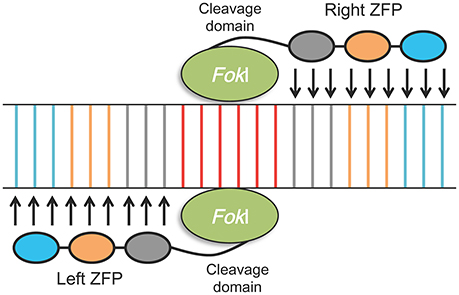
Figure 4. Schematic illustration of zinc-finger nuclease (ZFN) structure and mechanism of inducing double strand breaks (DBSs) on its target. The target site of the ZFN is recognized by the “left” and “right” monomers consisting of a tandem array of three to six engineered zinc finger proteins (ZFPs) (three are shown here); single engineered ZFP can recognize a nucleotide triplet (shown in different colors). Each ZNF is linked to a nuclease domain from the FokI restriction enzyme. Recognition of the target sequence by the left and right ZFPs results in dimerization of the FokI nuclease; DNA cleavage takes place along the spacer sequence (usually 6 bp long, shown in red) between the two ZFP recognition sites.
To our knowledge, only ZFNs have been applied in woody fruit species, in particular in apple and fig in a targeted mutagenesis experiment. Protocol optimisation for the use of ZFNs in apple and fig trees was developed by Peer et al. (2015). In this study, the ability of QQR-ZFN to repair a mutated uidA gene, which encodes for a non-functional GUS reporter protein, was explored. Both transient and stable transformation studies were carried out in fig and apple in vitro tissues. Whole plants with repaired uidA gene were regenerated; GUS assay results showed an overall gene editing efficiency of 80–100% per leaf explant in fig and 10–40% per leaf explant in apple.
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-Associated Protein (Cas9)
ZFNs (Townsend et al., 2009; Carroll, 2011) and transcription activator-like effector nucleases (TALENs; Boch et al., 2009; Moscou and Bogdanove, 2009) were the main genome editing tools until recently. Due to the difficulties related to the creation of flexible DNA-binding proteins, new methods of targeting such as CRISPR/Cas9 significantly simplified the creation of custom nucleases. Engineered nucleases have been designed as tools for genome editing as efficient genetic engineering methods to target and cleave DNA sequences at specific locations in the genome of both plants and animals (Bortesi and Fischer, 2015; Osakabe and Osakabe, 2015). CRISPR/Cas9 systems are an integrated part of the adaptive immune system of bacteria (Streptococcus pyogenes) and archaea (Bhaya et al., 2011; Jinek et al., 2012; Barrangou, 2015; Bortesi and Fischer, 2015), which protects them from invading nucleic acids such as viruses. This adaptive immunity is provided through silencing of the invading nucleic acids using CRISPR RNAs (crRNAs) and the Cas9 nucleases (Horvath and Barrangou, 2010). Cas9 gene in bacterial genomes was found to be closely linked with short, highly homologous sequences arranged in tandem repeats with a varying size between 21 and 37 bp interspaced with non-homologous spacer sequences (Jansen et al., 2002; Bhaya et al., 2011). Immunity is acquired by integrating into the genome short fragments of DNA from the invading organism (spacers), between two adjacent repeats at the proximal end of a CRISPR locus (Bortesi and Fischer, 2015). The spacer sequences determine the target to be cleaved by the endonuclease. The CRISPR arrays, which include the spacers, are transcribed during every encounter with invading DNA and are processed into 40 bp-long small interfering CRISPR RNAs (crRNAs), which combine with the trans-activating CRISPR RNA (tracrRNA) to activate and guide the Cas9 nuclease in cleaving the invading nucleic acid (Bortesi and Fischer, 2015). Target recognition is dependent on the “protospacer adjacent motif” (PAM) which is downstream of the target sequence and usually has the sequence 5′-NGG-3′ adjacent to the 3′ end of the 20 bp target (Jinek et al., 2012; Bortesi and Fischer, 2015). The application of this natural immune system to plant genome editing needs the creation of a single guide RNA molecule (sgRNA), obtained by fusing the 3′ end of the crRNA to the 5′ end of the tracrRNA. In this way Cas9 is reprogrammed to induce the cleavage of specific DNA sequences. A schematic illustration of the CRISPR/Cas9 mechanism is shown in Figure 5.
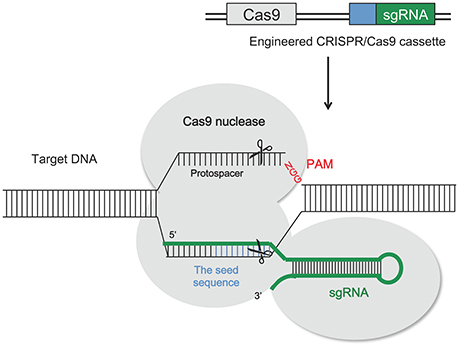
Figure 5. Schematic illustration of the CRISPR/Cas9 system structure and the principle of CRISPR/Cas9-mediated genomic modifications. Cas9 can be reprogrammed to induce the cleavage of specific DNA sequences by the production of a synthetic guide RNA (sgRNA). It contains a region (seed sequence, usually 8–12 bp long, shown in blue) complementary to the target DNA on the genomic loci that mediates the binding of the Cas9 protein. The cleavage site stays 3 bp upstream of the protospacer adjacent motif (PAM, NGG; shown in red), which is required for the cleavage of the target DNA sequence. Induced DSBs of the target DNA are repaired by either NHEJ or HDR, producing gene mutations that include nucleotide insertion, deletion or substitution around the cleavage sites (see Figure 3).
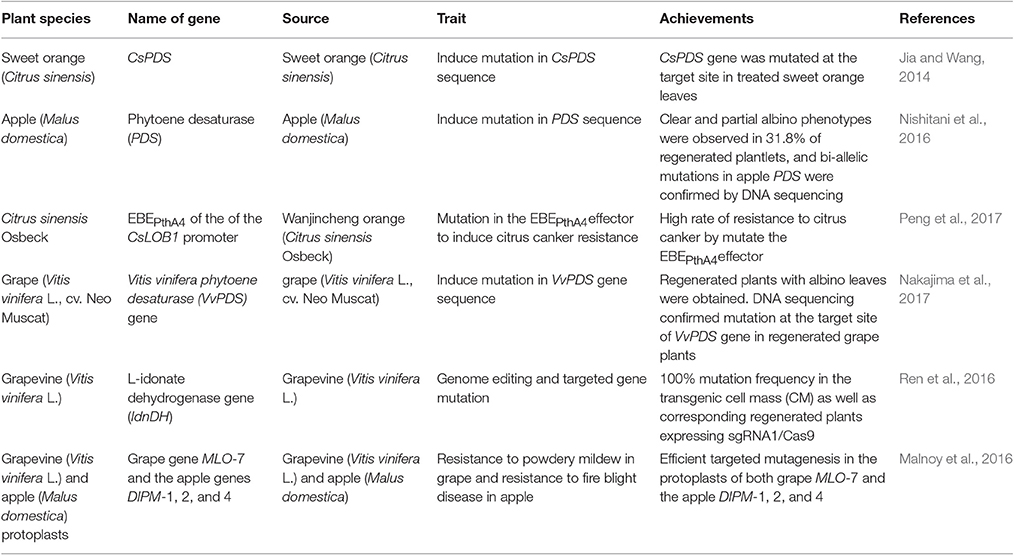
In comparison to ZFNs and TALENs which are larger in size and require a pair of proteins to recognize target DNA strands for DSBs induction, CRISPR/Cas9 is smaller in size making it easier to co-deliver multiple sgRNAs with Cas9 to the cell, so that the simultaneous editing of more than one target sequence is achievable in a process called “multiplex gene editing” (Cong et al., 2013). CRISPR/Cas9 system specificity is determined by the guide sequence of sgRNA complementing that of the target DNA. For efficient target cleavage by Cas9 to occur, there must be perfect base pairing between the last 8–12 bases of the guide sequence, called the “seed sequence,” and the complementary region of the target DNA (i.e., the region proximal to the 5′ end of the PAM; Jiang et al., 2013). Cas9 acts alone to bind and cleave the DNA target in a sequence-dependent manner (Anders et al., 2014; Nishimasu et al., 2014). The CRISPR/Cas9 system is used widely for genome editing because of its simplicity, design flexibility and high efficiency, which is easily applicable in laboratories. It is also the least expensive and most user-friendly of the three genome-editing tools (Nagamangala et al., 2015). CRISPR/Cas9 system has been applied in a number of woody fruit species to determine the possibility of precise gene mutations induction (Table 4). The most recent application of CRISPR/Cas9 system in inducing disease resistance in woody fruit species has been done in citrus by Peng et al. (2017). This genome editing technique was applied to increase resistance against Citrus canker, caused by Xanthomonas citri subsp. citri (Xcc), a deadly disease threatening the citrus industry worldwide (Stover et al., 2014). Peng et al. (2017) targeted the modification of the EBEPthA4 (effector binding element) of the susceptibility gene LATERAL ORGAN BOUNDARIES 1 (CsLOB1) promoter (Hu et al., 2014) in Wanjincheng orange. During infection, the main Xcc transcription activator-like (TAL) effector, PthA4, connects with the EBEPthA4 effector present on the promoter of CsLOB1 susceptibility gene, activating its expression and inducing Citrus canker development (Hu et al., 2014). Results showed that editing of CsLOB1 gene promoter was sufficient to increase the resistance of Wanjincheng orange against Citrus canker. In addition high levels of resistance to this disease were also induced by deletion of the entire EBEPthA4 sequence from both CsLOB1 alleles (Peng et al., 2017).
Oligonucleotide-Directed Mutagenesis (ODM)
Oligonucleotide-directed mutagenesis (ODM) is a gene-editing technique aimed to introduce a new mutation in the plant genome by replacing one or few base pairs (Lusser et al., 2011). This site-specific mutation occurs by the introduction of chemically synthesized DNA oligonucleotides or also chimeric DNA-RNA fragments of 20–100 nucleotides, which are delivered into the plant cells mainly by biolistic methods or electroporation of protoplasts (Breyer et al., 2009; Sauer et al., 2016). The introduced oligonucleotide hybridizes with a complementary predetermined DNA sequence in the plant genome, leading to the creation of a mismatch of one or two base pairs corresponding to the non-homologous nucleotides. This mechanism induces the cell's natural repair machinery to recognize the single base mismatch and to correct it. As a result, the desired specific change in the plant's genome is produced and the oligonucleotide is subsequently degraded by the cell (Schaart et al., 2016). ODM represents theoretically an improved technique over conventional breeding and traditional mutagenesis techniques, due to its controlled and accurate manner of action, through which random mutations are prevented and no recombinant DNA introduction is involved. Thus the final product produced is often similar to conventionally bred or traditional mutagenesis products (Breyer et al., 2009). To our knowledge there are no examples of ODM applications to woody fruit species and few data exist for other plant species (maize, rice, tobacco, and wheat; Zhu et al., 2000; Iida and Terada, 2005; Dong et al., 2006), probably due to some drawbacks, such as the low mutation efficiency and frequency, difficulties in the regeneration of mutated plants and the introduction of spontaneous somatic mutations which hide the changes introduced by ODM (Ruiter et al., 2003; Sauer et al., 2016).
Biosafety Considerations for the Application of NBTs in Fruit Trees
Advancements in agricultural technology offer new products and new solutions toward a sustainable future. However, these come with new concerns and new issues to address. Biosafety risk assessment principles, procedures, and policies have been adopted for ensuring the environmental and personal safety of genetically modified organisms (GMOs). Genetically modified crops present numerous biosafety issues and plant breeders are required to demonstrate the safety of their product before releasing GM crops into the environment for commercial purposes. NBTs have been developed to enable more precise genetic modifications of plants compared to conventional and some early genetic modification methods. However, it is still not clear whether crops obtained using some of these techniques should be classified and regulated as GMOs or the same as a product from traditional breeding or mutagenesis. In particular at EU level there is a lack of clear regulation concerning the use of these new techniques, while the world wide scientific community suggests that the evaluation of plants obtained by NBTs should focus on the changes made to the plant itself and on the final products obtained (Hartung and Schiemann, 2014; Sprink et al., 2016). Therefore, the products from the application of these technologies should be assessed with a simplified procedure mostly addressed to consider whether the induced genomic modifications are within the normal genetic variability of the species.
In terms of risk assessment, one of the main concerns related to GM plants is linked to the production of a new protein in the modified plant and their possible off-target effects. This concern should be avoided by the applications of the cisgenic approach since all components are obtained from the same species or from a sexually compatible species, and thus do not produce a novel protein in the recipient genetically modified plant, and therefore do not provide different outcomes compared to traditional breeding (Podevin et al., 2012).
The EFSA GMO Panel considers that “the Guidance for risk assessment of food and feed from genetically modified plants and the Guidance on the environmental risk assessment of genetically modified plants are applicable for the evaluation of food and feed products derived from cisgenic and intragenic plants/crops. It can be envisaged that on a case-by-case basis lesser amounts of event specific data are needed for the risk assessment” [European Food Safety Authority (EFSA), 2012].
From these considerations, the assessment of a new cisgenic plant in the EU could be reduced to the genomic characterization of the product so that molecular studies can confirm the absence of heterologous DNA sequences and products (proteins and enzymes), and no additional environmental and food safety risk assessment might be needed (Schouten et al., 2006b). Regulatory authorities in some North and South American countries consider that cisgenic plants and intragenic plants not containing antibiotic resistance markers do not need to be considered as other types of GM plants and so would not require regulation (Waltz, 2011).
The full genome sequences now available for many crops offer enormous possibilities to identify useful genes/promoters directly from the same species to be transferred to improve the commercial cultivars. However, the transformation approach can be limited by the availability of efficient selectable markers suitable to replace the commonly used antibiotic or herbicide resistance markers. To solve this problem, progress is being made in developing reporter genes derived from the large class of myeloblastosis (MYB) transcription factors involved in anthocyanin pigment activation in plant species (Elomaa et al., 2003). This approach has been applied in grapevines by Kandel et al. (2016), who compared the grapevine-derived VvMybA1 transcription factor with existing reporter genes gfp and gus. The MybA1 reporter gene was found to be suitable for identification of gene expression events at the cell culture level (Kandel et al., 2016). MYB markers can be identified for each plant species, but it is not always easy to develop efficient regeneration systems that allow the application of only a reporter gene without the application of selectable markers.
RNAi-based GM plants regulate the expression of specific genes, determined by the production of dsRNA molecules, without the production of new homologous and heterologous proteins/enzymes. However, impacts on the RNAi gene target and possible off-target effects also need to be considered. Several approaches have been developed to silence plant endogenous genes (e.g., silencing genes for fruit ripening; Atkinson et al., 2012) or the MLO gene for mildew resistance (Pessina et al., 2016). In these cases, if there are no other transgenic sequences inserted, the plant could be considered a cisgenic plant and the risk assessment could be reduced to molecular characterization of the event and of the target gene silenced. In case of RNAi systems to induce resistance to other organisms interacting with the plant (e.g., virus, fungi, bacteria, and insects) a study of possible off-target gene silencing through the dsRNAs produced by the plant, in both target and non-target organisms, should be investigated.
The application of GM techniques and, in particular, RNAi technology on rootstocks for producing non-GM grafted scions, are offering new important opportunities for fruit and other vegetatively propagated plants. In the scion, including the fruit, there is no presence of homologous and heterologous proteins but only of small fragments of RNA. The scion does not contain transgenic DNA or novel proteins therefore; the main concerns should be related only to the off-target effects on non-target organisms and sequences by RNAi. An additional benefit of the trans-grafting approach is the absence of pollen or seed dispersal of transgenic material from the non-genetically modified scion.
Biosafety concerns affecting genetically engineered fruit trees also include off-target mutations. Taking into consideration the gene editing approaches (CRISPR/Cas9 system, ZFN, TALEN's, and ODM) it has been demonstrated that their application can result in mutations similar to those that occur in nature or by use of traditional breeding techniques, with the important difference that these new technologies act in a much more specific way (Curtin et al., 2011; Tzfira et al., 2012; Hartung and Schiemann, 2014; Ren et al., 2016). In the CRISPR/Cas9 technique, bioinformatics tools are used in the designing of sgRNAs to identify both target and off-target sites. These tools choose the gene regions for sgRNA creation on the basis of specific sequence characteristics such as size (usually 21–23 bases) and specific nucleotide constitution, thus minimizing the possibility of off-target mutations (Brazelton et al., 2016). One of the final products of this novel gene editing approach can be considered as a point mutation as the double strand break is made in a very precise manner in that it is un-differentiable from natural mutation. For this reason the scientific community has supported the view that CRISPR/Cas9-edited plants should not be classified as GMOs unless they contain transgenic elements. However, the method commonly used to introduce this genome editing system is with Agrobacterium-mediated transformation. In this case, the first product obtained is a GMO, for this reason some authorities suggest for the product's regulation. The transgenic complex can be eliminated only by an F1 segregation from mutant F0 after selfing or back-crossing with the wild type. Segregants in the offspring no longer contain the transgenes, or foreign DNA in their genome. These F1 mutants should only differ from the wild type by a small deletion in the target gene, and so they are usually indistinguishable from those arising spontaneously or through mutation breeding (Jones, 2015). In homozygous (seed propagated plants), where selfing- can be used, the F1 mutant maintains all the traits of the original cultivar. In heterozygous plants, fruit trees and many other vegetatively propagated plants, back-crossing is required and the gene-edited offspring show a larger variability in comparison with the original clone. This aspect remains a major limiting factor in the application of CRISPR/Cas9 via stable transformation for improving woody fruit crop species. The most common alternative now proposed is the transient cell transformation by the insertion of the CRISPR/Cas9 Ribonucleoproteins (RNPs) complex in protoplast cells (Malnoy et al., 2016). In this case, no stable genetic modification occurs, while the inserted protein complex induces the mutation, the CRISPR/Cas9 RNPs are quickly cleared from the cell via protein degradation pathways resulting in a GM free gene-edited plant. This could be considered the most appropriate approach for applying CRISPR/Cas9 to induce point mutations, or for gene insertion/knockdown in woody fruit species. However, a major limitation is the inability to regenerate plants from modified protoplasts of many important woody fruit species (Mezzetti et al., 2001), indeed few examples exist of optimal regeneration protocols for woody fruit species protoplasts (Patat-Ochatt et al., 1988; Vardi et al., 1990).
Conclusions
Biotechnological techniques have undergone rapid development adding novel and valuable tools for plant breeders. These techniques make it possible to create desirable crop cultivars in fast and more efficient ways to meet the demand for improved crops to support sustainable agricultural productivity and in order to cater for the ever-increasing world population.
Although the new biotechnological techniques have one common goal i.e., precise, fast, and efficient crop improvement, individually they are different in approach and characteristics from each other. Some of these techniques, such as RNAi and trans-grafting, can be combined to achieve the desired results. Commercial applications of genetically modified fruit trees are so far limited. The only fruit plants available on the market are the “Rainbow” virus resistant papaya since September 1997, when all the necessary procedures for approval had been completed successfully (Gonsalves, 2004), and the arctic apple approved by the US Department of Agriculture (USDA) in February 2015, making it the first genetically modified apple resistant to browning (Waltz, 2015). The virus resistant Honey Sweet plum cultivar attained approval for commercialization in USA but has not reached the market yet (Scorza et al., 2013). The limited application of GM technology in fruit trees can be explained by (1) the difficulties in developing efficient regeneration and transformation protocols for many cultivars of the different species as many fruit tree species are recalcitrant, (2) the regulatory requirements. These reasons lead to the limited commercial exploitation of GM fruit trees by the fruit industry hence limited investment in fruit tree biotechnologies by plant breeders and the biotech industry. In this connection therefore, it is mainly public research institutions, with limited budgets, that are developing biotechnology research on these crops.
The NBTs such as cisgenesis and intragenesis could raise less biosafety concerns and should be considered more similar to conventional breeding techniques; RNAi introduces no new proteins in the plant, which means no novel allergenicity issues and that a lightened risk assessment process should be required. Furthermore, gene editing techniques, especially CRISPR/Cas9 combined with RNPs delivered directly to the protoplast, are more precise and targeted techniques and less likely to create unintended off-target mutations as RNPs are quickly cleared from the cell via protein degradation pathways resulting in a modified plant free from any foreign materials from the CRISPR/Cas9 RNPs complex.
Author Contributions
CL designed and contributed to the writing of the main body of article; SS contributed by writing specific parts of the article and revising it critically; JS contributed to the development of specific parts of the article and to its critical revision; BM conceived and contributed toward the writing of specific parts of the article and in its critical revision.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The reviewer UM and handling Editor declared their shared affiliation, and the handling Editor states that the process met the standards of a fair and objective review.
Acknowledgments
Our studies described here were supported by COST action CA15223 iPlanta project.
References
Abel, P. P., Nelson, R. S., De, B., Hoffmann, N., Rogers, S. G., Fraley, R. T., et al. (1986). Delay of disease development in transgenic plants that express the tobacco mosaic virus coat protein gene. Science 232, 738–743. doi: 10.1126/science.3457472
Almeida, R., and Allshire, R. C. (2005). RNA silencing and genome Regulation. Trends Cell Biol. 15, 251–258. doi: 10.1016/j.tcb.2005.03.006
Aloni, B., Cohen, R., Karni, L., Aktas, H., and Edelstein, M. (2010). Hormonal signaling in rootstock-scion interactions. Sci. Hortic. 127, 119–126. doi: 10.1016/j.scienta.2010.09.003
Altpeter, F., Baisakh, N., Beachy, R., Bock, R., Capell, T., Christou, P., et al. (2005). Particle bombardment and the genetic enhancement of crops: myths and realities. Mol. Breed. 15, 305–327. doi: 10.1007/s11032-004-8001-y
An, C., Orbović, V., and Mou, Z. (2013). An efficient intragenic vector for generating intragenic and cisgenic plants in citrus. Am. J. Plant Sci. 4, 2131–2137. doi: 10.4236/ajps.2013.411265
Anders, C., Niewoehner, O., Duerst, A., and Jinek, M. (2014). Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573. doi: 10.1038/nature13579
Atkinson, R. G., Sutherland, P. W., Johnston, S. L., Gunaseelan, K., Hallett, I. C., Mitra, D., et al. (2012). Down-regulation of POLYGALACTURONASE1 alters firmness, tensile strength and water loss in apple (Malus x domestica) fruit. BMC Plant Biol. 12:129. doi: 10.1186/1471-2229-12-129
Barrangou, R. (2015). The roles of CRISPR–Cas systems in adaptive immunity and beyond. Curr. Opin. Immunol. 32, 36–41. doi: 10.1016/j.coi.2014.12.008
Bartel, D. P. (2004). MicroRNAs: genomics, biogenesis, mechanism and function. Cell 116, 281–297. doi: 10.1016/S0092-8674(04)00045-5
Baulcombe, D. C. (1996). Mechanisms of pathogen-derived resistance to viruses in transgenic plants. Plant Cell 8, 1833–1844. doi: 10.1105/tpc.8.10.1833
Baum, J. A., Bogaert, T., Clinton, W., Heck, G. R., Feldmann, P., Ilagan, O., et al. (2007). Control of coleopteran insect pests through RNA interference. Nat. Biotechnol. 25, 1322–1326. doi: 10.1038/nbt1359
Bhaya, D., Davison, M., and Barrangou, R. (2011). CRISPR-Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annu. Rev. Genet. 45, 273–297. doi: 10.1146/annurev-genet-110410-132430
Bhullar, N. K., and Gruissem, W. (2013). Nutritional enhancement of rice for human health: the contribution of biotechnology. Biotechnol. Adv. 31, 50–57. doi: 10.1016/j.biotechadv.2012.02.001
Billmyre, R. B., Calo, S., Feretzaki, M., Wang, X., and Heitman, J. (2013). RNAi function, diversity, and loss in the fungal kingdom. Chromosome Res. 21, 561–572. doi: 10.1007/s10577-013-9388-2
Boch, J., Scholze, H., Schornack, S., Landgraf, A., Hahn, S., Kay, S., et al. (2009). Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512. doi: 10.1126/science.1178811
Borges, F., and Martienssen, R. A. (2015). The expanding world of small RNAs in plants. Nat. Rev. Mol. Cell Biol. 16, 727–741. doi: 10.1038/nrm4085
Bortesi, L., and Fischer, R. (2015). The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 33, 41–52. doi: 10.1016/j.biotechadv.2014.12.006
Brazelton, V. A. Jr., Zarecor, S., Wright, D. A., Wang, Y., Liu, J., Chen, K., et al. (2016). A quick guide to CRISPR sgRNA design tools. GM Crops Food 6, 266–276. doi: 10.1080/21645698.2015.1137690
Breyer, D., Herman, P., Brandenburger, A., Gheysen, G., Remaut, E., Soumillion, P., et al. (2009). Commentary: genetic modification through oligonucleotide-mediated mutagenesis. A GMO regulatory challenge? Environ. Biosaf. Res. 8, 57–64. doi: 10.1051/ebr/2009007
Brodersen, P., and Voinnet, O. (2006). The diversity of RNA silencing pathways in plants. Trends Genet. 22, 268–280. doi: 10.1016/j.tig.2006.03.003
Brodersen, P., Sakvarelidze-Achard, L., Bruun-Rasmussen, M., Dunoyer, P., Yamamoto, Y. Y., Sieburth, L., et al. (2008). Widespread translational inhibition by plant miRNAs and siRNAs. Science 320, 1185–1190. doi: 10.1126/science.1159151
Brosnan, C. A., and Voinnet, O. (2011). Cell-to-cell and long-distance siRNA movement in plants: mechanisms and biological implications. Curr. Opin. Plant Biol. 14, 580–587. doi: 10.1016/j.pbi.2011.07.011
Cambra, M., Capote, N., Myrta, A., and Llácer, G. (2006). Plum pox virus and estimated costs associated to sharka disease. Bull. OEPP/EPPO Bull. 36, 202–204. doi: 10.1111/j.1365-2338.2006.01027.x
Campbell, T. N., and Choy, F. Y. (2005). RNA interference: past, present and future. Curr. Issues Mol. Biol. 7, 1–6.
Carbonell, A., and Carrington, J. C. (2015). Antiviral roles of plant ARGONAUTES. Curr. Opin. Plant Biol. 27, 111–117. doi: 10.1016/j.pbi.2015.06.013
Carroll, D. (2011). Genome engineering with zinc-finger nucleases. Genetics 188, 773–782. doi: 10.1534/genetics.111.131433
Carthew, R. W., and Sontheimer, E. J. (2009). Origins and Mechanisms of miRNAs and siRNAs. Cell 136, 642–655. doi: 10.1016/j.cell.2009.01.035
Cervera, M., Juarez, J., Navarro, A., Pina, J. A., Duran-Vila, N., Navarro, L., et al. (1998). Genetic transformation and regeneration of mature tissues of woody fruit plants bypassing the juvenile stage. Transgenic Res. 7, 51–59. doi: 10.1023/A:1008855922283
Cheng, W., Song, X. S., Li, H. P., Cao, L. H., Sun, K., Qiu, X. L., et al. (2015). Host induced gene silencing of an essential chitin synthase gene confers durable resistance to Fusarium head blight and seedling blight in wheat. Plant Biotechnol. J. 13, 1335–1345. doi: 10.1111/pbi.12352
Chilton, M. D., Drummond, M. H., Merio, D. J., Sciaky, D., Montoya, A. L., Gordon, M. P., et al. (1977). Stable incorporation of plasmid DNA into higher plant cells: the molecular basis of crown gall tumorigenesis. Cell 11, 263–271. doi: 10.1016/0092-8674(77)90043-5
Collinge, D. B., Jørgensen, H. J., Lund, O. S., and Lyngkjær, M. F. (2010). Engineering pathogen resistance in crop plants: current trends and future prospects. Annu. Rev. Phytopathol. 48, 269–291. doi: 10.1146/annurev-phyto-073009-114430
Cong, L., Ran, F. A., Cox, D., Lin, S., Barretto, R., Habib, N., et al. (2013). Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823. doi: 10.1126/science.1231143
Conner, A. J., Barrell, P. J., Baldwin, S. J., Lokerse, A. S., Cooper, P. A., Erasmuson, A. K., et al. (2007). Intragenic vectors for gene transfer without foreign DNA. Euphytica 154, 341–353. doi: 10.1007/s10681-006-9316-z
Curtin, S. J., Voytas, D. F., and Stupar, R. M. (2012). Genome engineering of crops with designer nucleases. Plant Genome 5, 42–50. doi: 10.3835/plantgenome2012.06.0008
Curtin, S. J., Zhang, F., Sander, J. D., Haun, W. J., Starker, C., Baltes, N. J., et al. (2011). Targeted mutagenesis of duplicated genes in soybean with zinc-finger nucleases. Plant Physiol. 156, 466–473. doi: 10.1104/pp.111.172981
Datta, A. (2013). Genetic engineering for improving quality and productivity of crops. Agric. Food Security 2:15. doi: 10.1186/2048-7010-2-15
De Alba, A. E. M., Elvira-Matelot, E., and Vaucheret, H. (2013). Gene silencing in plants: a diversity of pathways. Biochim. Biophys. Acta Gene Regul. Mech. 1829, 1300–1308. doi: 10.1016/j.bbagrm.2013.10.005
Deng, D., Yan, C., Pan, X., Mahfouz, M., Wang, J., Zhu, J. K., et al. (2012). Structural basis for sequence-specific recognition of DNA by TAL effectors. Science 335, 720–723. doi: 10.1126/science.1215670
Dhekney, S. A., Li, Z. T., and Gray, D. J. (2011). Grapevines engineered to express cisgenic Vitis vinifera thaumatin-like protein exhibit fungal disease resistance in vitro cell. Dev. Biol. Plant 47, 458–466. doi: 10.1007/s11627-011-9358-3
Ding, S. W. (2010). RNA-based antiviral immunity. Nat. Rev. Immunol. 10, 632–644. doi: 10.1038/nri2824
Ding, S. W., and Voinnet, O. (2007). Antiviral immunity directed by small RNAs. Cell 130, 413–426. doi: 10.1016/j.cell.2007.07.039
Dong, C., Beetham, P., Vincent, K., and Sharp, P. (2006). Oligonucleotide-directed gene repair in wheat using a transient plasmid gene repair assay system. Plant Cell Rep. 25, 457–465. doi: 10.1007/s00299-005-0098-x
Dunoyer, P., Himber, C., and Voinnet, O. (2005). DICER-LIKE 4 is required for RNA interference and produces the 21-nucleotide small interfering RNA component of the plant cell-to-cell silencing signal. Nat. Genet. 37, 1356–1360. doi: 10.1038/ng1675
Dutt, M., Li, Z. T., Kelley, K. T., Dhekney, S. A., Van Aman, M., Tattersall, J., et al. (2007). Transgenic rootstock protein transmission in grapevines. Acta Hortic. 738, 749–754. doi: 10.17660/ActaHortic.2007.738.99
Elomaa, P., Uimari, A., Mehto, M., Albert, V. A., Laitinen, R. A., and Teeri, T. H. (2003). Activation of anthocyanin biosynthesis in Gerbera hybrida (Asteraceae) suggests conserved protein-protein and protein-promoter interactions between the anciently diverged monocots and eudicots. Plant Physiol. 133, 1831–1842. doi: 10.1104/pp.103.026039
English, J. J., Mueller, E., and Baulcombe, D. C. (1996). Suppression of virus accumulation in transgenic plants exhibiting silencing of nuclear genes. Plant Cell 8, 179–188. doi: 10.1105/tpc.8.2.179
Escobar, M. A., Civerolo, E. L., Summerfelt, K. R., and Dandekar, A. M. (2001). RNAi-mediated oncogene silencing confers resistance to crown gall tumorigenesis. Proc. Nat. Acad. Sci. U.S.A. 98, 13437–13442. doi: 10.1073/pnas.241276898
European Food Safety Authority (EFSA) (2012). Scientific opinion addressing the safety assessment of plants developed through cisgenesis and intragenesis. EFSA J. 10:2561. doi: 10.2903/j.efsa.2012.2561
Febres, V. J., Lee, R. F., and Moore, G. A. (2008). Transgenic resistance to Citrus tristeza virus in grapefruit. Plant Cell Rep. 27, 93–104. doi: 10.1007/s00299-007-0445-1
Fire, A., Xu, S. Q., Montgomery, M. K., Kostas, S. A., Driver, S. E., and Mello, C. C. (1998). Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811. doi: 10.1038/35888
Freiman, A., Shlizerman, L., Golobovitch, S., Yablovitz, Z., Korchinsky, R., Cohen, Y., et al. (2012). Development of a transgenic early flowering pear (Pyrus communis L.) genotype by RNAi silencing of PcTFL1-1 and PcTFL1-2. Planta 235, 1239–1251. doi: 10.1007/s00425-011-1571-0
Frizzi, A., and Huang, S. (2010). Tapping RNA silencing pathways for plant biotechnology. Plant Biotechnol. J. 8, 655–677. doi: 10.1111/j.1467-7652.2010.00505.x
Funke, T., Han, H., Healy-Fried, M. L., Fischer, M., and Schönbrunn, E. (2006). Molecular basis for the herbicide resistance of Roundup Ready crops. Proc. Natl. Acad. Sci. U.S.A. 103, 13010–13015. doi: 10.1073/pnas.0603638103
Gaj, T., Gersbach, C. A., and Barbas, C. F. (2013). ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405. doi: 10.1016/j.tibtech.2013.04.004
Gelvin, S. B. (2003). Agrobacterium-mediated plant transformation: the biology behind the “gene-jockeying” tool. Microbiol. Mol. Biol. Rev. 67, 16–37. doi: 10.1128/MMBR.67.1.16-37.2003
Gessler, C., Vanblaere, T., Parravicini, G., and Broggini, G. A. L. (2014). Cisgenic ‘Gala’ containing the scab resistance gene from Malus floribunda 821 and the fire blight resistance genes from M. ‘Evereste’. Acta Horticult. 1048, 43–50. doi: 10.17660/ActaHortic.2014.1048.4
Giuliano, G. (2017). Provitamin A biofortification of crop plants: a gold rush with many miners. Curr. Opin. Biotechnol. 44, 169–180. doi: 10.1016/j.copbio.2017.02.001
Gonsalves, D. (1998). Control of papaya ringspot virus in papaya: a case study. Annu. Rev. Phytopathol. 36, 415–437. doi: 10.1146/annurev.phyto.36.1.415
Gonsalves, D. (2006). Transgenic papaya: development, release, impact and challenges. Adv. Virus Res. 67, 317–354. doi: 10.1016/S0065-3527(06)67009-7
Gottula, J., and Fuchs, M. (2009). Toward a quarter century of pathogen derived resistance and practical approaches to plant virus disease control. Adv. Virus Res. 75, 161–183. doi: 10.1016/S0065-3527(09)07505-8
Grens, K. (2017). RNA Interference Between Kingdoms. Available online at: http://www.the-scientist.com/?articles.view/articleNo/48073/title/RNAInterferenceBetweenKingdoms/&_hsenc=p2ANqtz-_gT8pO7h21CVKZggjSAJSIGfBMHMB7hsrba2Ggg82zZCKW6skKkR8qLPrY_M7AxCl0394O9EFjs14yUYwSAKKv1zQ&_hsmi=41791015/
Haroldsen, V. M., Chi-Ham, C. L., and Bennett, A. B. (2012a). Transgene mobilization and regulatory uncertainty for non-GE fruit products of transgenic rootstocks. J. Biotechnol. 161, 349–353. doi: 10.1016/j.jbiotec.2012.06.017
Haroldsen, V. M., Szczerba, M. W., Aktas, H., Lopez-baltazar, J., Odias, M. J., Chi-ham, C. L., et al. (2012b). Mobility of transgenic nucleic acids and proteins within grafted rootstocks for agricultural improvement. Front. Plant Sci. 3:39. doi: 10.3389/fpls.2012.00039
Hartung, F., and Schiemann, J. (2014). Precise plant breeding using new genome editing techniques: opportunities, safety and regulation in the EU. Plant J. 78, 742–752. doi: 10.1111/tpj.12413
Hily, J. M., Ravelonandro, M., Damsteegt, V., Bassett, C., Petri, C., Liu, Z., et al. (2007). Plum pox virus coat protein gene Intron-hairpin-RNA (ihpRNA) constructs provide resistance to plum pox virus in Nicotiana benthamiana and Prunus domestica. J. Am. Soc. Hortic. Sci. 132, 850–858.
Horvath, P., and Barrangou, R. (2010). CRISPR/Cas, the immune system of Bacteria and Archaea. Science 327, 167. doi: 10.1126/science.1179555
Hu, Y., Zhang, J. L., Jia, H. G., Sosso, D., Li, T., Frommer, W. B., Yang, B., et al. (2014). Lateral organ boundaries 1 is a disease susceptibility gene for citrus bacterial canker disease. Proc. Natl. Acad. Sci. U.S.A. 111, E521–E529. doi: 10.1073/pnas.1313271111
Iida, S., and Terada, R. (2005). Modification of endogenous natural genes by gene targeting in rice and other higher plants. Plant Mol. Biol. 59, 205–219. doi: 10.1007/s11103-005-2162-x
Ipsaro, J. J., and Joshua-Tor, L. (2015). From guide to target: molecular insights into eukaryotic RNA-interference machinery. Nat. Struct. Mol. Biol. 22, 20–28. doi: 10.1038/nsmb.2931
Jacobsen, E., and Schouten, H. J. (2007). Cisgenesis strongly improves introgression breeding and induced translocation breeding of plants. Trends Biotechnol. 25, 219–223. doi: 10.1016/j.tibtech.2007.03.008
Jansen, R., Embden, J. D., Gaastra, W., and Schouls, L. M. (2002). Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 43, 1565–1575. doi: 10.1046/j.1365-2958.2002.02839.x
Jia, H., and Wang, N. (2014). Targeted genome editing of sweet orange using Cas9/sgRNA. PLoS ONE 9:e93806. doi: 10.1371/journal.pone.0093806
Jiang, W., Bikard, D., Cox, D., Zhang, F., and Marraffini, L. A. (2013). RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 31, 233–239. doi: 10.1038/nbt.2508
Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., and Charpentier, E. (2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821. doi: 10.1126/science.1225829
Jonas, S., and Izaurralde, E. (2015). Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 16, 421–433. doi: 10.1038/nrg3965
Jones, H. D. (2015). Regulatory uncertainty over genome editing. Nat. Plants 1, 14011. doi: 10.1038/nplants.2014.11
Joshi, S. G., Schaart, J. G., Groenwold, R., Jacobsen, E., Schouten, H. J., and Krens, F. A. (2011). Functional analysis and expression profiling of HcrVf1 and HcrVf2 for development of scab resistant cisgenic and intragenic apples. Plant Mol. Biol. 75, 579–591. doi: 10.1007/s11103-011-9749-1
Kandel, R., Bergey, D. R., Dutt, M., Sitther, V., Li, Z. T., Gray, D. J., et al. (2016). Evaluation of a grapevine-derived reporter gene system for precision breeding of Vitis. Plant Cell Tiss. Organ Cult. 124, 599–609. doi: 10.1007/s11240-015-0918-9
Ketting, R. F. (2011). The many faces of RNAi. Dev. Cell. 20, 148–161. doi: 10.1016/j.devcel.2011.01.012
Khalid, A., Zhang, Q., Yasir, M., and Li, F. (2017). Small RNA based genetic engineering for plant viral resistance: application in crop protection. Front. Microbiol. 8:43. doi: 10.3389/fmicb.2017.00043
Klocko, A. L., Borejsza-Wysocka, E., Brunner, A. M., Shevchenko, O., Aldwinckle, H., and Strauss, S. H. (2016). Transgenic suppression of AGAMOUS genes in apple reduces fertility and increases floral attractiveness. PLoS ONE 11:e0159421. doi: 10.1371/journal.pone.0159421
Knip, M., Constantin, M. E., and Thordal-Christensen, H. (2014). Trans-kingdom cross-talk: small RNAs on the move. PLoS Genet. 10:e1004602. doi: 10.1371/journal.pgen.1004602
Kost, T. D., Gessler, C., Jänsch, M., Flachowsky, H., Patocchi, A., and Broggini, G. A. L. (2015). Development of the first cisgenic apple with increased resistance to fire blight. PLoS ONE 10:e0143980. doi: 10.1371/journal.pone.0143980
Krens, F. A., Schaart, J. G., Van der Burgh, A. M., Tinnenbroek-Capel, I. E., Groenwold, M. R., Kodde, L. P., et al. (2015). Cisgenic apple trees; development, characterization, and performance. Front. Plant Sci. 6:286. doi: 10.3389/fpls.2015.00286
Lang, A., Chailakhyan, M. K., and Frolova, I. A. (1977). Promotion and inhibition of flower formation in a day neutral plant in grafts with a short-day plant and a long-day plant. Proc. Natl. Acad. Sci. U.S.A. 74, 2412–2416. doi: 10.1073/pnas.74.6.2412
Lemgo, G., Sabbadini, S., Pandolfini, T., and Mezzetti, B. (2013). Biosafety considerations of RNAi-mediated virus resistance in fruit-tree cultivars and in rootstock. Transgenic Res. 22, 1073–1088. doi: 10.1007/s11248-013-9728-1
Lombardo, L., Coppola, G., and Zelasco, S. (2016). New technologies for insect-resistant and herbicide-tolerant plants. Trends Biotechnol. 34, 49–57. doi: 10.1016/j.tibtech.2015.10.006
Lucas, W. J., and Lee, J.-Y. (2004). Plasmodesmata as a supracellular control network in plants. Nat. Rev. Mol. Cell Biol. 5, 712–726. doi: 10.1038/nrm1470
Lusser, M., and Davies, H. V. (2013). Comparative regulatory approaches for groups of new plant breeding techniques. Nat. Biotechnol. 30, 437–446. doi: 10.1016/j.nbt.2013.02.004
Lusser, M., Parisi, C., Plan, D., and Rodríguez-Cerezo, E. (2011). New Plant Breeding Techniques. State of-the-Art And Prospects for Commercial Development. JRC Scientific and Technical Reports/EUR 24760 EN. European Commission.
Lusser, M., Parisi, C., Plan, D., and Rodríguez-Cerezo, E. (2012). Deployment of new biotechnologies in plant breeding. Nat. Biotechnol. 30, 231–239. doi: 10.1038/nbt.2142
Mallory, A. C., Mlotshwa, S., Bowman, L. H., and Vance, V. B. (2003). The capacity of transgenic tobacco to send a systemic RNA silencing signal depends on the nature of the inducing transgene locus. Plant J. 35, 82–92. doi: 10.1046/j.1365-313X.2003.01785.x
Malnoy, M., Viola, R., Jung, M.-H., Koo, O.-J., Kim, S., Kim, J.-S., et al. (2016). DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front. Plant Sci. 7:1904. doi: 10.3389/fpls.2016.01904
Matzke, M. A., and Mosher, R. A. (2014). RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat. Rev. Genet. 15, 394–408. doi: 10.1038/nrg3683
Matzke, M. A., Matzke, A. J., Pruss, G. J., and Vance, V. B. (2001). RNA-based silencing strategies in plants. Curr. Opin. Genet. Dev. 11, 221–227. doi: 10.1016/S0959-437X(00)00183-0
McGarry, R. C., and Kragler, F. (2013). Phloem-mobile signals affecting flowers: applications for crop breeding. Trends Plant Sci. 18, 198–206. doi: 10.1016/j.tplants.2013.01.004
Meister, G., and Tuschl, T. (2004). Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343–349. doi: 10.1038/nature02873
Melnyk, C. W., and Meyerowitz, E. M. (2015). Plant grafting. Curr. Biol. 25, R183–R188. doi: 10.1016/j.cub.2015.01.029
Melnyk, C. W., Molnar, A., and Baulcombe, D. C. (2011). Intercellular and systemic movement of RNA silencing signals. EMBO J. 30, 3553–3563. doi: 10.1038/emboj.2011.274
Metzlaff, M., O'dell, M., Cluster, P. D., and Flavell, R. B. (1997). RNA-mediated RNA degradation and chalcone synthase A silencing in petunia. Cell 88, 845–854.
Mezzetti, B., Landi, L., Phan, B. H., Taruschio, L., and Lim, Y. K. (2001). PEG-mediated fusion of Rubus idaeus (raspberry) and R. fructicosus (blackberry) and protoplast, selection and characterization of callus lines. Plant Biosyst. 135, 63–70. doi: 10.1080/11263500112331350660
Mezzetti, B., Pandolfini, T., Navacchi, O., and Landi, L. (2002). Genetic transformation of Vitis vinifera via organogenesis. BMC Biotechnol. 2:18. doi: 10.1186/1472-6750-2-18
Miller, J. C., Holmes, M. C., Wang, J., Guschin, D. Y., Lee, Y. L., Rupniewski, I., et al. (2007). An improved zinc-finger nuclease architecture for highly specific genome editing. Nat. Biotechnol. 25, 778–785. doi: 10.1038/nbt1319
Mittler, R., and Blumwald, E. (2010). Genetic engineering for modern agriculture: challenges and perspectives. Annu. Rev. Plant Biol. 61, 443–462. doi: 10.1146/annurev-arplant-042809-112116
Molnar, A., Melnyk, C. W., Bassett, A., Hardcastle, T. J., Dunn, R., and Baulcombe, D. C. (2010). Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Science 328, 872–875. doi: 10.1126/science.1187959
Molnar, A., Melnyk, C., and Baulcombe, D. C. (2011). Silencing signals in plants: a long journey for small RNAs. Genome Biol. 12:215. doi: 10.1186/gb-2010-11-12-219
Moscou, M. J., and Bogdanove, A. J. (2009). A simple cipher governs DNA recognition by TAL effectors. Science 326:1501. doi: 10.1126/science.1178817
Mudge, K., Janick, J., Scofield, S., and Goldschmidt, E. E. (2009). A history of grafting. Hortic. Rev. 35, 437–493. doi: 10.1002/9780470593776.ch9
Nagamangala, K. C., Sargent, D. J., Velasco, R., Maffei, M. E., and Malnoy, M. (2015). Looking forward to genetically edited fruit crops. Trends Biotechnol. 33, 62–64. doi: 10.1016/j.tibtech.2014.07.003
Nakajima, I., Ban, Y., Azuma, A., Onoue, N., Moriguchi, T., Yamamoto, T., et al. (2017). CRISPR/Cas9-mediated targeted mutagenesis in grape. PLoS ONE 12:e0177966. doi: 10.1371/journal.pone.0177966
Nakayashiki, H. (2005). RNA silencing in fungi: mechanisms and applications. FEBS Lett. 579, 5950–5970. doi: 10.1016/j.febslet.2005.08.016
Napoli, C., Lemieux, C., and Jorgensen, R. (1990). Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2, 279–289. doi: 10.1105/tpc.2.4.279
Navarro, L., Dunoyer, P., Jay, F., Arnold, B., Dharmasiri, N., Estelle, M., et al. (2006). A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312, 436–439. doi: 10.1126/science.1126088
Nazim, U. M., and Kim, J. Y. (2013). Intercellular and systemic spread of RNA and RNAi in plants. Wiley Interdiscip. Rev. RNA. 4, 279–293. doi: 10.1002/wrna.1160
Nishimasu, H., Ran, F. A., Hsu, P. D., Konermann, S., Shehata, S. I., Dohmae, N., et al. (2014). Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949. doi: 10.1016/j.cell.2014.02.001
Nishitani, C., Hirai, S. K., Masato, W., Kazuma, O., Keishi, O., Toshiya, Y., et al. (2016). Efficient genome editing in apple using a CRISPR/Cas9 system. Sci. Rep. 6:31481. doi: 10.1038/srep31481
Omid, A., Keilin, T., Glass, A., Leshkowitz, D., and Wolf, S. (2007). Characterization of phloem-sap transcription profile in melon plants. J. Exp. Bot. 58, 3645–3656. doi: 10.1093/jxb/erm214
Osakabe, Y., and Osakabe, K. (2015). Genome editing with engineered nucleases in plants. Plant Cell Physiol. 5, 389–400. doi: 10.1093/pcp/pcu170
Pabo, C. O., Peisach, E., and Grant, R. A. (2001). Design and selection of novel Cys2His2 zinc finger proteins. Annu. Rev. Biochem. 70, 313–340. doi: 10.1146/annurev.biochem.70.1.313
Paine, J., Shipton, C., Chaggar, S., Howells, R., Kennedy, M., Vernon, G., et al. (2005). Improving the nutritional value of Golden Rice through increased pro-vitamin A content. Nat. Biotechnol. 23, 482–487. doi: 10.1038/nbt1082
Palauqui, J. C., Elmayan, T., Pollien, J. M., and Vaucheret, H. (1997). Systemic acquired silencing: transgene-specific post-transcriptional silencing is transmitted by grafting from silenced stocks to non-silenced scions. EMBO J. 16, 4738–4745. doi: 10.1093/emboj/16.15.4738
Parent, J. S., and Vaucheret, H. (2012). The origin and effect of small RNA signaling in plants. Front. Plant Sci. 3:179. doi: 10.3389/fpls.2012.00179
Parisi, C., Tillie, P., and Rodríguez-Cerezo, E. (2016). The global pipeline of GM crops out to 2020. Nat. Biotechnol. 34, 32–36. doi: 10.1038/nbt.3449
Patat-Ochatt, E. M., Ochatt, S. J., and Power, J. B. (1988). Plant regeneration from protoplasts of apple rootstocks and scion varieties (Malus x domestica Borkh.). J. Plant Physiol. 133, 460–465. doi: 10.1016/S0176-1617(88)80037-3
Peer, R., Rivlin, G., Golobovitch, S., Lapidot, M., Gal-On, A., Vainstein, A., et al. (2015). Targeted mutagenesis using zinc-finger nucleases in perennial fruit trees. Planta 241, 941–951. doi: 10.1007/s00425-014-2224-x
Peil, A., Garcia-Libreros, T., Richter, K., Trognitz, F. C., Trognitz, B., Hanke, M. V., et al. (2007). Strong evidence for a fire blight resistance gene of Malus robusta located on linkage group 3. Plant Breed. 126, 470–475. doi: 10.1111/j.1439-0523.2007.01408.x
Peng, A., Chen, S., Lei, T., Xu, L., He, Y., Wu, L., et al. (2017). Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotech. J. doi: 10.1111/pbi.12733. [Epub ahead of print].
Pérez-Jiménez, M., Carrillo-Navarro, A., and Cos-Terrer, J. (2012). Regeneration of peach (Prunus persica L. Batsch) cultivars and Prunus persica× Prunus dulcis rootstocks via organogenesis. Plant Cell Tissue Organ Cult. 108, 55–62. doi: 10.1007/s11240-011-0011-y
Pérez-Massot, E., Banakar, R., Gómez-Galera, S., Zorrilla-López, U., Sanahuja, G., Arjó, G., et al. (2013). The contribution of transgenic plants to better health through improved nutrition: opportunities and constraints. Genes Nutr. 8, 29–41. doi: 10.1007/s12263-012-0315-5
Pessina, S., Angeli, D., Martens, S., Visser, R., Bai, Y., Salamini, F., et al. (2016). The knock-down of the expression of MdMLO19 reduces susceptibility to powdery mildew (Podosphaera leucotricha) in apple (Malus domestica). Plant Biotechnol. J. 14, 2033–2044. doi: 10.1111/pbi.12562
Petolino, J. F. (2015). Genome editing in plants via designed zinc finger nucleases. In Vitro Cell. Dev. Biol. 51:1. doi: 10.1007/s11627-015-9663-3
Petri, C., and Burgos, L. (2005). Transformation of fruit trees. Useful breeding tool or continued future prospect? Transgenic Res. 14, 15–26. doi: 10.1007/s11248-004-2770-2
Podevin, N., Devos, Y., Davies, H. V., and Nielsen, K. M. (2012). Transgenic or not? No simple answer! New biotechnology-based plant breeding techniques and the regulatory landscape. EMBO Rep. 13, 1057–1061. doi: 10.1038/embor.2012.168
Qaim, M., and Kouser, S. (2013). Genetically modified crops and food security. PLoS ONE 8:e64879. doi: 10.1371/journal.pone.0064879
Qi, Y., and Hannon, G. J. (2005). Uncovering RNAi mechanisms in plants: biochemistry enters the foray. FEBS Lett. 579, 5899–5903. doi: 10.1016/j.febslet.2005.08.035
Rai, M. K., and Shekhawat, N. S. (2014). Recent advances in genetic engineering for improvement of fruit crops. Plant Cell Tissue Organ Cult 116, 1–15. doi: 10.1007/s11240-013-0389-9
Ren, C., Liu, X., Zhang, Z., Wang, Y., Duan, W., Li, S., et al. (2016). CRISPR/Cas9-mediated efficient targeted mutagenesis in Chardonnay (Vitis vinifera L.) Sci. Rep. 6:32289. doi: 10.1038/srep32289
Reyes, C. A., Francesco, A. D., Pe na, E. J., Costa, N., Plata, M. I., Sendin, L., et al. (2011). Resistance to Citrus psorosis virus in transgenic sweet orange plants is triggered by coat protein–RNA silencing. J. Biotechnol. 151, 151–158. doi: 10.1016/j.jbiotec.2010.11.007
Romano, N., and Macino, G. (1992). Quelling: transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol. Microbiol. 6, 3343–3353. doi: 10.1111/j.1365-2958.1992.tb02202.x
Rommens, C. M. (2004). All-native DNA transformation: a new approach to plant genetic engineering. Trends Plant Sci. 9, 457–464. doi: 10.1016/j.tplants.2004.07.001
Rommens, C. M. (2007). Intragenic crop improvement: combining the benefits of traditional breeding and genetic engineering. J. Agric. Food Chem. 55, 4281–4288. doi: 10.1021/jf0706631
Ruiter, R., Van Den Brande, I., Stals, E., Delauré, S., Cornelissen, M., and D'halluin, K. (2003). Spontaneous mutation frequency in plants obscures the effect of chimeraplasty. Plant Mol. Biol. 53, 715–729. doi: 10.1023/B:PLAN.0000019111.96107.01
Sabbadini, S., Pandolfini, T., Girolomini, L., Molesini, B., and Navacchi, O. (2015). “Peach (Prunus persica L.)” in Agrobacterium Protocols, ed K. Wang (New York, NY: Springer), 205–215.
Salame, T. M., Ziv, C., Hadar, Y., and Yarden, O. (2011). RNAi as a potential tool for biotechnological applications in fungi. Appl. Microbiol. Biotechnol. 89, 501–512. doi: 10.1007/s00253-010-2928-1
Sanford, J. C., and Johnston, S. A. (1985). The concept of pathogen-derived resistance: deriving resistance genes from the parasite's own genome. J. Theor. Biol. 113, 395–405. doi: 10.1016/S0022-5193(85)80234-4
Saporta, R., San Pedro, T., and Gisbert, C. (2017). Attempts at grapevine (Vitis vinifera L.) breeding through genetic transformation: the main limiting factors. VITIS J. Grape. Res. 55, 173–186. doi: 10.5073/vitis.2016.55.173-186
Sauer, N. J., Mozoruk, J., Miller, R. B., Warburg, Z. J., Walker, K. A., Beetham, P. R., et al. (2016). Oligonucleotide-directed mutagenesis for precision gene editing. Plant Biotechnol. J. 14, 496–502. doi: 10.1111/pbi.12496
Schaart, J. G., and Visser, R. G. F. (2009). Novel Plant Breeding Techniques - Consequences of New Genetic Modification-Based Techniques in Comparison to Conventional Plant Breeding. The Netherlands Commission on Genetic Modification. COGEM Research Report number 2009–02.
Schaart, J. G., van de Wiel, C. C., Lotz, L. A., and Smulders, M. J. (2016). Opportunities for products of new plant breeding techniques. Trends Plant Sci. 21, 438–449. doi: 10.1016/j.tplants.2015.11.006
Schouten, H. J., Krens, F. A., and Jacobsen, E. (2006a). Cisgenic plants are similar to traditionally bred plants: international regulations for genetically modified organisms should be altered to exempt cisgenesis. EMBO Rep. 7, 750–753. doi: 10.1038/sj.embor.7400769
Schouten, H. J., Krens, F. A., and Jacobsen, E. (2006b). Do cisgenic plants warrant less stringent oversight? Nat. Biotechnol. 24:753. doi: 10.1038/nbt0706-753
Scorza, R., Callahan, A., Dardick, C., Ravelonandro, M., Polak, J., Malinowski, T., et al. (2013). Genetic engineering of Plum pox virus resistance:‘HoneySweet’plum—from concept to product. Plant Cell Tissue Organ Cult. 115, 1–12. doi: 10.1007/s11240-013-0339-6
Scorza, R., Callahan, A., Levy, L., Damsteegt, V., Webb, K., and Ravelonandro, M. (2001). Post-transcriptional gene silencing in plum pox virus resistant transgenic European plum containing the plum pox poty virus coat protein gene. Transgenic Res. 10, 201–209. doi: 10.1023/A:1016644823203
Scorza, R., Ravelonandro, M., Callahan, A. M., Cordts, J. M., Fuchs, M., and Dunez, J. (1994). Transgenic plums (Prunus domestica L.) express the plum pox virus coat protein gene. Plant Cell Rep. 14, 18–22. doi: 10.1007/BF00233291
Scott, J. G., Michel, K., Bartholomay, L. C., Siegfried, B. D., Hunter, W. B., Smagghe, G., et al. (2013). Towards the elements of successful insect RNAi. J. Insect Physiol. 59, 1212–1221. doi: 10.1016/j.jinsphys.2013.08.014
Simón-Mateo, C., and García, J. A. (2011). Antiviral strategies in plants based on RNA silencing. BBA Gene Regul. Mech. 1809, 722–731. doi: 10.1016/j.bbagrm.2011.05.011
Smith, N. A., Singh, S. P., Wang, M. B., Stoutjesdijk, P. A., Green, A. G., and Waterhouse, P. M. (2000). Gene expression: total silencing by intron-spliced hairpin RNAs. Nature 407, 319–320. doi: 10.1038/35030305
Smolka, A., Li, X. Y., Heikelt, C., Welander, M., and Zhu, L. H. (2010). Effects of transgenic rootstocks on growth and development of non-transgenic scion cultivars in apple. Transgenic Res. 19:933. doi: 10.1007/s11248-010-9370-0
Song, G. Q., Sink, K. C., Walworth, A. E., Cook, M. A., Allison, R. F., and Lang, G. A. (2013). Engineering cherry rootstocks with resistance to Prunus necrotic ring spot virus through RNAi-mediated silencing. Plant Biotechnol. J. 11, 702–708. doi: 10.1111/pbi.12060
Sprink, T., Eriksson, D., Schiemann, J., and Hartung, F. (2016). Regulatory hurdles for genome editing: process-vs. product-based approaches in different regulatory contexts. Plant Cell Rep. 35, 1493–1506. doi: 10.1007/s00299-016-1990-2
Stegemann, S., and Bock, R. (2009). Exchange of genetic material between cells in plant tissue grafts. Science 324, 649–651. doi: 10.1126/science.1170397
Stover, E., Driggers, R., Richardson, M. L., Hall, D. G., Duan, Y. P., and Lee, R. F. (2014). Incidence and severity of asiatic citrus canker on diverse citrus and citrus-related germplasm in a FLORIDA field planting. HortScience 49, 4–9.
Townsend, J. A., Wright, D. A., Winfrey, R. J., Fu, F., Maeder, M. L., Joung, J. K., et al. (2009). High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 459, 442–445. doi: 10.1038/nature07845
Tzfira, T., and Citovsky, V. (2006). Agrobacterium-mediated genetic transformation of plants: biology and biotechnology. Curr. Opin. Biotechnol. 17, 147–154. doi: 10.1016/j.copbio.2006.01.009
Tzfira, T., Weinthal, D., Marton, I., Zeevi, V., Zuker, A., and Vainstein, A. (2012). Genome modifications in plant cells by custom-made restriction enzymes. Plant Biotechnol. J. 10, 373–389. doi: 10.1111/j.1467-7652.2011.00672.x
Urnov, F. D., Rebar, E. J., Holmes, M. C., Zhang, H. S., and Gregory, P. D. (2010). Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 11, 636–646. doi: 10.1038/nrg2842
Vanblaere, T., Flachowsky, H., Gessler, C., and Broggini, G. A. (2014). Molecular characterization of cisgenic lines of apple ‘Gala’ carrying the Rvi6 scab resistance gene. Plant Biotechnol. J. 12, 2–9. doi: 10.1111/pbi.12110
Vanblaere, T., Szankowski, I., Schaart, J., Schouten, H., Flachowsky, H., Broggini, G. A. L., et al. (2011). The development of a cisgenic apple plant. J. Biotechnol. 154, 304–311. doi: 10.1016/j.jbiotec.2011.05.013
Vardi, A., Bleichman, S., and Aviv, D. (1990). Genetic transformation of Citrus protoplasts and regeneration of transgenic plants. Plant Sci. 69, 199–206. doi: 10.1016/0168-9452(90)90118-8
Vaucheret, H., and Fagard, M. (2001). Transcriptional gene silencing in plants: targets, inducers and regulators. Trends Genet. 17, 29–35. doi: 10.1016/S0168-9525(00)02166-1
Vaucheret, H., Béclin, C., and Fagard, M. (2001). Post-transcriptional gene silencing in plants. J. Cell Sci. 114, 3083–3091.
Viss, W. J., Pitrak, J., Humann, J., Cook, M., Driver, J., and Ream, W. (2003). Crown-gall-resistant transgenic apple trees that silence Agrobacterium tumefaciens oncogenes. Mol. Breed. 12, 283–295. doi: 10.1023/B:MOLB.0000006805.76717.08
Voinnet, O. (2008). Post-transcriptional RNA silencing in plant-microbe interactions: a touch of robustness and versatility. Curr. Opin. Plant Biol. 11, 464–470. doi: 10.1016/j.pbi.2008.04.006
Voinnet, O. (2009). Origin, biogenesis, and activity of plant microRNAs. Cell 136, 669–687. doi: 10.1016/j.cell.2009.01.046
Waltz, E. (2015). Non-browning GM apple cleared for market. Nat. Biotechnol. 33, 326–327. doi: 10.1038/nbt0415-326c
Wang, H., Alburquerque, N., Burgos, L., and Petri, C. (2011). Adventitious shoot regeneration from hypocotyl slices of mature apricot (Prunus armeniaca L.) seeds: a feasible alternative for apricot genetic engineering. Sci. Hortic. 128, 457–464. doi: 10.1016/j.scienta.2011.02.020
Welander, M., and Zhu, L. H. (2000). The rooting ability of rolB transformed clones of the apple rootstock M26 and its relation to gene expression. Acta Hortic. 521, 133–138. doi: 10.17660/ActaHortic.2000.521.14
Wood, A. J., Lo, T. W., Zeitler, B., Pickle, C. S., Ralston, E. J., Lee, A. H., et al. (2011). Targeted genome editing across species using ZFNs and TALENs. Science 333, 307–307. doi: 10.1126/science.1207773
Wurdig, J., Flachowsky, H., Sab, A., Peil, A., and Hanke, M. (2015). Improving resistance of different apple cultivars using the Rvi6 scab resistance gene in a cisgenic approach based on the FLP/FRT recombinase system. Mol. Breed. 35:95. doi: 10.1007/s11032-015-0291-8
Xie, Z., Johansen, L. K., Gustafson, A. M., Kasschau, K. D., Lellis, A. D., Zilberman, D., et al. (2004). Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2:e104. doi: 10.1371/journal.pbio.0020104
Zhao, D., and Song, G. Q. (2014). Rootstock-to-scion transfer of transgene-derived small interfering RNAs and their effect on virus resistance in non-transgenic sweet cherry. Plant Biotechnol. J. 12, 1319–1328. doi: 10.1111/pbi.12243
Zhao, K., Zhang, F., Yang, Y., Ma, Y., Liu, Y., Li, H., et al. (2016). Modification of plant height via RNAi suppression of MdGA20-ox gene expression in apple. J. Am. Soc. Hortic. Sci. 141, 242–248.
Zhu, C., Sanahuja, G., Yuan, D., Farré, G., Arjó, G., Berman, J., et al. (2013). Biofortification of plants with altered antioxidant content and composition: genetic engineering strategies. Plant Biotechnol. J. 11, 129–141. doi: 10.1111/j.1467-7652.2012.00740.x
Keywords: plant breeding, RNA interference (RNAi), trans-grafting, cisgenesis/intragenesis, Crispr/Cas9, genome editing, EU biosafety regulations
Citation: Limera C, Sabbadini S, Sweet JB and Mezzetti B (2017) New Biotechnological Tools for the Genetic Improvement of Major Woody Fruit Species. Front. Plant Sci. 8:1418. doi: 10.3389/fpls.2017.01418
Received: 17 May 2017; Accepted: 31 July 2017;
Published: 15 August 2017.
Edited by:
Joachim Hermann Schiemann, Julius Kühn-Institut, GermanyReviewed by:
Elena Corredoira, Consejo Superior de Investigaciones Científicas (CSIC), SpainJeff Wolt, Iowa State University, United States
Ulrike Manske, Julius Kühn-Institut, Germany
Copyright © 2017 Limera, Sabbadini, Sweet and Mezzetti. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Bruno Mezzetti, Yi5tZXp6ZXR0aUB1bml2cG0uaXQ=
 Cecilia Limera
Cecilia Limera Silvia Sabbadini
Silvia Sabbadini Jeremy B. Sweet
Jeremy B. Sweet Bruno Mezzetti1*
Bruno Mezzetti1*