- 1Independent Researcher, Hyderabad, India
- 2School of Biological Sciences, Institute of Agriculture, University of Western Australia, Perth, WA, Australia
- 3Plant and AgriBiosciences Research Centre, Ryan Institute, National University of Ireland Galway, Galway, Ireland
- 4Department of Plant Breeding, Swedish University of Agricultural Sciences, Alnarp, Sweden
There is a need to accelerate crop improvement by introducing alleles conferring host plant resistance, abiotic stress adaptation, and high yield potential. Elite cultivars, landraces and wild relatives harbor useful genetic variation that needs to be more easily utilized in plant breeding. We review genome-wide approaches for assessing and identifying alleles associated with desirable agronomic traits in diverse germplasm pools of cereals and legumes. Major quantitative trait loci and single nucleotide polymorphisms (SNPs) associated with desirable agronomic traits have been deployed to enhance crop productivity and resilience. These include alleles associated with variation conferring enhanced photoperiod and flowering traits. Genetic variants in the florigen pathway can provide both environmental flexibility and improved yields. SNPs associated with length of growing season and tolerance to abiotic stresses (precipitation, high temperature) are valuable resources for accelerating breeding for drought-prone environments. Both genomic selection and genome editing can also harness allelic diversity and increase productivity by improving multiple traits, including phenology, plant architecture, yield potential and adaptation to abiotic stresses. Discovering rare alleles and useful haplotypes also provides opportunities to enhance abiotic stress adaptation, while epigenetic variation has potential to enhance abiotic stress adaptation and productivity in crops. By reviewing current knowledge on specific traits and their genetic basis, we highlight recent developments in the understanding of crop functional diversity and identify potential candidate genes for future use. The storage and integration of genetic, genomic and phenotypic information will play an important role in ensuring broad and rapid application of novel genetic discoveries by the plant breeding community. Exploiting alleles for yield-related traits would allow improvement of selection efficiency and overall genetic gain of multigenic traits. An integrated approach involving multiple stakeholders specializing in management and utilization of genetic resources, crop breeding, molecular biology and genomics, agronomy, stress tolerance, and reproductive/seed biology will help to address the global challenge of ensuring food security in the face of growing resource demands and climate change induced stresses.
Assessing Crop Functional Diversity
Producing sufficient food for the growing population is a major challenge, with climate change emerging as an additional threat to the food security and livelihood of millions of people (Abberton et al., 2016). Achieving significant yield gains in staple crops is essential because rising demand requires a twofold increase in crop production by 2050 (Tilman et al., 2011). The increasing frequency of droughts and heat stress is impacting crop productivity (Deryng et al., 2014; Lesk et al., 2016), and the increased frequency and severity of flooding events may cause yield loss in regions such as Asia, where prolonged flooding of rice fields already substantially reduces yields (Mackill et al., 2012). To meet the challenges of increasing demand in a changing climate, there is a need to more rapidly generate new and improved crop cultivars.
Cereals and grain legumes constitute the major components of the human diet and of livestock feed. Grain legumes also enrich soil with nitrogen and improve soil texture for other crops (Graham and Vance, 2003). The discovery of semi-dwarfing genes fuelled the stark increase in yields (known as the ‘Green Revolution’) in rice and wheat production globally (Trethowan et al., 2007). However, the reliance on a narrow range of elite cultivars has likely led to some negative effects on agroecosystems productivity (Dwivedi et al., 2017), though this assumption remains controversial and empirical research provides contradictory evidence (Fu, 2015). More recent evidence also suggests that productivity of major food crops is either stagnating or not increasing at the rate needed to ensure food security (Ortiz, 2015). Accelerated progress in plant breeding is required to better harness crop genetic resources and produce higher-yielding, climate-resilient cultivars.
As the methods to assess functional diversity in crops have become more sophisticated during the last 100 years, our understanding of the mechanisms underlying this diversity has grown. Functional diversity refers to a component of biodiversity related to what organisms do in communities and ecosystems (Petchey and Gaston, 2006). The decreasing cost of high-throughput DNA sequencing has facilitated the recent rise of genome-wide methods such as genotyping by sequencing (Scheben et al., 2017a) for assessing functional diversity of crops using single nucleotide polymorphisms (SNPs) (Kilian and Graner, 2012; Huang and Han, 2014). Common targets of breeding are yield-related traits such as abiotic stress tolerance, pest resistance and flowering time. The potential yield gains are substantial, considering that abiotic stress can reduce average yields of major crops by 50% (Bray et al., 2000) and pests can cause 26–40% yield losses (Oerke, 2006). Assessing and using functional diversity in pathways controlling flowering time is also important for yield, particularly as control of crop development can enhance adaptation to the predicted impact of climate change. The genomics era has led to a rapid increase in sequence data capturing the genetic diversity underlying heritable target traits in elite cultivars, landraces and crop wild relatives. However, while there were already over 100 plant genomes available in 2015 (Michael and VanBuren, 2015), over half of which were crops, the functions of the vast majority of plant genes remain unknown (Rhee and Mutwil, 2014).
Powerful and high-throughput forward and reverse genetic techniques are required to help elucidate these unknown gene functions to assist targeted breeding. Genetic mapping approaches also play an important role in associating genomic regions with phenotypic traits. Vast improvements in our understanding of the functional knowledge of crop genomes is an important prerequisite for targeted genome editing based approaches to access novel diversity for breeding programs (Scheben et al., 2017b), which often remain limited by the natural diversity found in germplasm resources. Both understanding and shaping of crop functional diversity using genomic technologies will be necessary to ensure continuing yield increases to keep pace with growing global food demand. In this review article, we focus on the latest developments in assessing and exploiting functional diversity associated with abiotic stress adaptation, phenology, plant architecture, and yield attributing traits in cereals and food legumes germplasm pools using genomics-led methods for crop genetic enhancement. We focus on three questions: (1) How do we characterize functional diversity? (2) What are the key breeding targets? (3) How can we apply knowledge of functional diversity to improve crop traits using genomic prediction and genome editing?
Approaches for Uncovering Functional Diversity
Analysis of DNA variation regulating phenotypes (traits) in crops can facilitate the identification of causal genes associated with desired agronomic traits. Advances in genome sequencing have dramatically reduced costs of measuring DNA variation, facilitating the identification of candidate genes for complex traits. To date, many crop genomes are sequenced, yielding millions of SNPs, while resequencing of diverse germplasm (including wild species) across crop genepools further provides a wealth of genomic information (in some instances related with discrete phenotypes). Single nucleotide polymporphisms (SNPs) are most abundant genetic markers that are amenable to automation and cost-effective for use and integration with crop breeding research. In particular, SNPs which are robustly associated with desirable agronomic phenotypes can provide a better understanding of gene function while also providing markers that can be used for more-efficient plant breeding schemes (Huq et al., 2016).
Genome-Wide SNP Polymorphism
Legumes
Soybean (Glycine max) has been extensively investigated for SNP variation using diverse genepools (Table 1). Valliyodan et al. (2016) reported over 10 million high quality SNPs and 0.75 m InDels, mostly (82.6%) in intergenic regions. Wild soybeans had 15% more SNPs than landraces and elite lines. Soybean cultivars also showed high SNP polymorphism (dos Santos et al., 2016). SNP-based arrays in soybean include the SoySNP1.5K (Shen et al., 2005), SoySNP6K (Akond et al., 2013), SoySNP50K (Song et al., 2013), and 180 K AXIOM® SoyaSNP (Lee et al., 2015) arrays. The SoySNP355K array, which covers the whole genome, is also available (Wang J. et al., 2016).
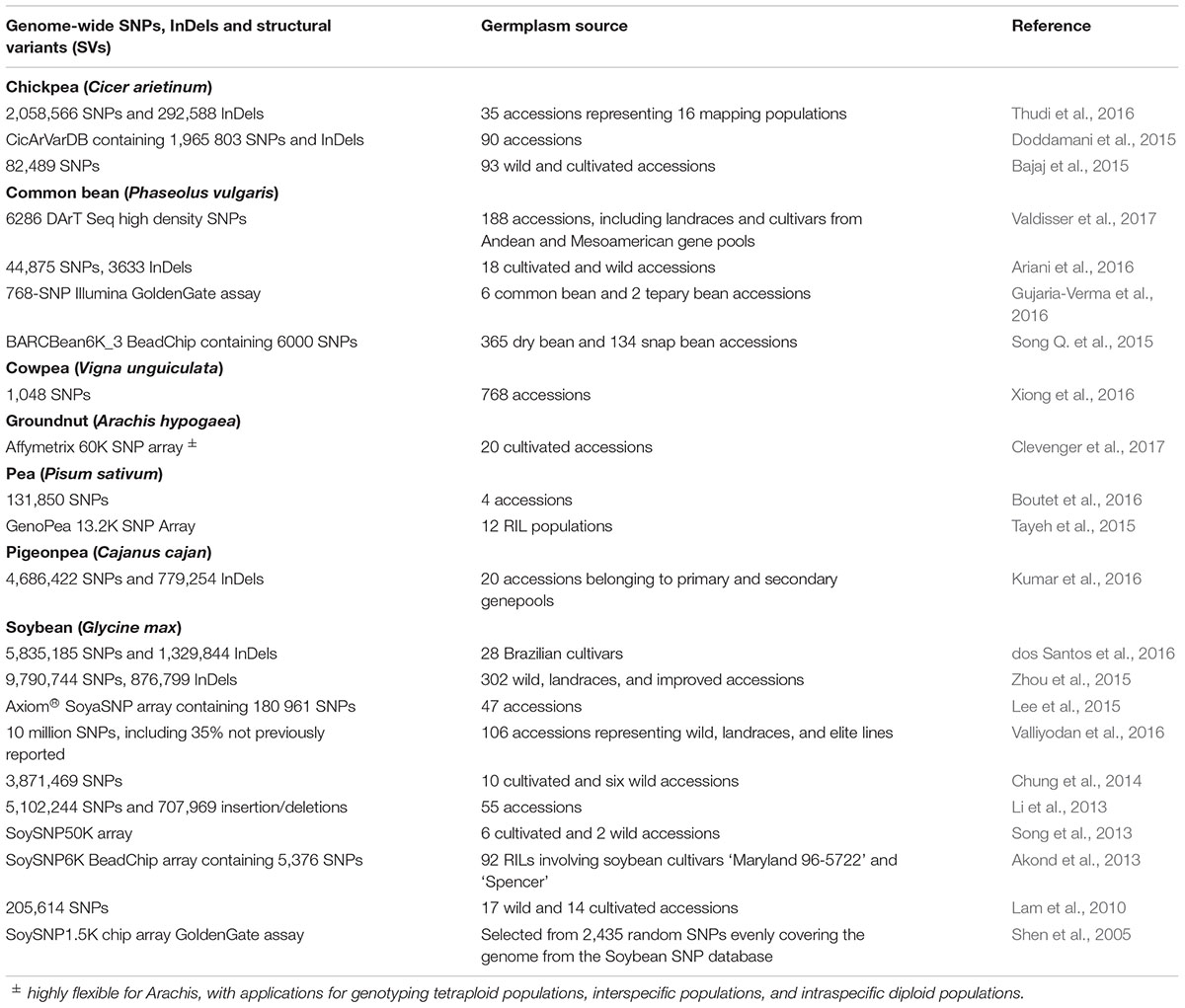
TABLE 1. Genome wide SNPs discovered in chickpea, common bean, cowpea, groundnut, pea, pigeonpea, and soybean.
Clevenger et al. (2017) re-sequenced 20 diverse groundnut (Arachis hypogaea) accessions to identify SNP variations and constructed a large-scale genotyping array, which contains 58,233 putative SNPs, including those from groundnut ancestors A. duranensis (21,547 SNPs) and A. ipaensis (22,933 SNPs). The array is designed to be highly flexible for Arachis, with applications for genotyping A. hypogaea populations, interspecific populations, and intraspecific diploid populations. A unique feature of this array is its set of 1,193 SNPs indicative of tetrasomic recombination (i.e., tetrasomic inheritance) events. Thus, this newly developed SNP array will be very useful for further genetic and breeding applications in Arachis.
The RCBean6K_3BeadChip array containing 6,000 SNPs is widely used in beans (Song Q. et al., 2015). A gene-based SNP array in tepary bean (Phaseolus acutifolius) provided greater insight of this species’ population structure and its relationship with common bean (Phaseolus vulgaris), facilitating the introgression of agriculturally important traits (Gujaria-Verma et al., 2016). Development of high throughput genotyping arrays, GenoPea 13.2KSNP in pea (Pisum sativum) (Tayeh et al., 2015) and Axiom®Cicer50.6SNP array in chickpea (Cicer arietinum) (Roorkiwal et al., 2016), are expected to accelerate genetic research. The chickpea database repository CicArVarDB contains 1.9 million SNPs and InDels anchored on eight pseudomolecules, allowing to select for variation associated with quantitative trait loci (QTL) (Doddamani et al., 2015).
The International Cowpea Consortium and Illumina have developed a new SNP genotyping array for cowpea (Vigna unguiculata). This 60,000-marker iSelect array provides a 40-fold increase in marker density compared to an older, 1,536-marker GoldenGate Illumina panel (Close, 2015). Pigeonpea (Cajanus cajan) has lagged behind in array technology, though abundant SNPs have been identified (Kumar et al., 2016).
Cereals
Both maize (Zea mays) and rice (Oryza sativa) have been extensively studied for SNP variation using diverse germplasm (Table 2). A publicly available high-density SNP array (609,442 SNPs and 6,759 InDels) optimized for European and American temperate maize, the Affymetrix® Axiom® Maize Genotyping Array, was recently developed (Unterseer et al., 2014). MaizeSNP3072 array containing 3,072 SNPs is more efficient than MaizeSNP50 array in fingerprinting Chinese cultivars (Tian et al., 2015). A maize 55 K SNP array with improved genome coverage was developed on an Affymetrix® Axiom® platform with 55,229 SNPs evenly distributed across the genome, which contains 451 markers associated with 368 known genes including those for drought tolerance and kernel oil biosynthesis, 4067 markers not assigned to any chromosome or position in the current reference genome, 734 markers differentiating heterotic groups, and 132 markers tagged for important transgenic events. This array improves MaizeSNP50 (Ganal et al., 2011), and is a powerful tool for germplasm evaluation, marker-assisted breeding, QTL mapping and association studies for both tropical and temperate maize (Xu C. et al., 2017).
Rice SNP50 array contains 51,478 evenly distributed markers (Chen et al., 2014). This array incorporates 50,051 SNPs from 18,980 single copy genes (3,710 conserved between wheat and rice, 14,959 unique to rice, 194 agronomically important cloned rice genes) and 117 multicopy rice genes, mapped on 12 rice chromosomes. The utility of this assay was demonstrated for genetic diversity and phylogenetic research, using a panel of diverse genepools, and in breeding (Singh et al., 2015).
A high-density array in wheat (Triticum aestivum) contains about 90,000 gene-associated SNPs from populations of diverse geographical origins. This array consists of 46,977 SNPs that were mapped using eight segregating populations (Wang S. et al., 2014).
Whole-genome resequencing (16–45× genome coverage) of 44 accessions of the diverse origins, end-use and taxonomic groups unravels 8 million high-quality SNPs and 1.9 million InDels in sorghum (Sorghum bicolor) (Mace et al., 2013), while resequencing three sorghum inbred lines uncovered 1 million SNPs, 0.099 million InDels, 0.106 million presence/absence variations, and 0.017 million copy number variations (Zheng et al., 2011). Sequencing of 500 pearl millet (Pennisetum glaucum) accessions identified 83,875 SNPs (Hu et al., 2015), while targeted resequencing of 433 diverse accessions generated a genome-wide panel of 544,318 high quality SNP in barley (Hordeum vulgare) (Pankin et al., unpublished).
Clearly, technological innovations in genomics have already led to discovery of abundant polymorphic SNPs in most cereal and legume crops, thus facilitating trait discovery and introgression. As the pace of technological advances and cost-reductions in next-generation sequencing technologies is extremely rapid, it is possible that SNP-array based platforms may be superseded by or become largely integrated with high-throughput sequencing approaches to genotyping (Pérez-Enciso et al., 2015).
QTL and Candidate Genes for Complex Traits
Genome-wide association studies (GWAS) carried out on diversity panels can provide higher mapping resolution than linkage mapping based on biparental crosses, thus allowing better detection of candidate causal genes (Ingvarsson and Street, 2011; Huang and Han, 2014). GWAS success depends, however, on data quality, population size, and the degree of linkage disequilibrium (LD) (Flint-Garcia et al., 2005; Mackay and Powell, 2007). Mutation, population structure, epistasis, and population perturbations such as migration, inbreeding, and selection all affects LD (Jannink and Walsh, 2002). LD decay varies between species, among different populations within species, and among different loci within a given genome (Tenaillon et al., 2001; Gupta et al., 2005; Caldwell et al., 2006).
Multigenic complex traits such as plant architecture, yield and related traits, and stress adaptation are typically affected by many genes and are also influenced by genotype × environment interactions. GWAS has been successful for detecting natural variation underlying some complex traits which has enabled researchers to identify several associated SNPs, some of which were co-located with previously reported QTL or candidate genes.
Phenology and Pod/Seed Traits in Legumes
Table 3 lists selected candidate genes for crop phenology in soybean and common bean or those associated with pod or seed characteristics in chickpea and cowpea. Zhang J. et al. (2015) reported new loci and refined genomic regions of known loci associated with crop duration (i.e., number of days from sowing to harvesting) and plant height in soybean. Candidate genes homologous to flowering genes in Arabidopsis thaliana were located near the peak SNP associated with flowering in soybean (Zhang J. et al., 2015). An allelic variant of the CesA-type cellulose synthase gene, Ca_Kabuli_CesA3, was found to regulate pod and seed numbers plant-1 in chickpea (Kujur et al., 2015). Phvul.001G221100 was associated with days to flower and maturity in common bean (Kamfwa et al., 2015). SNPs were also identified which are associated with pod length in cowpea (Xu P. et al., 2017).
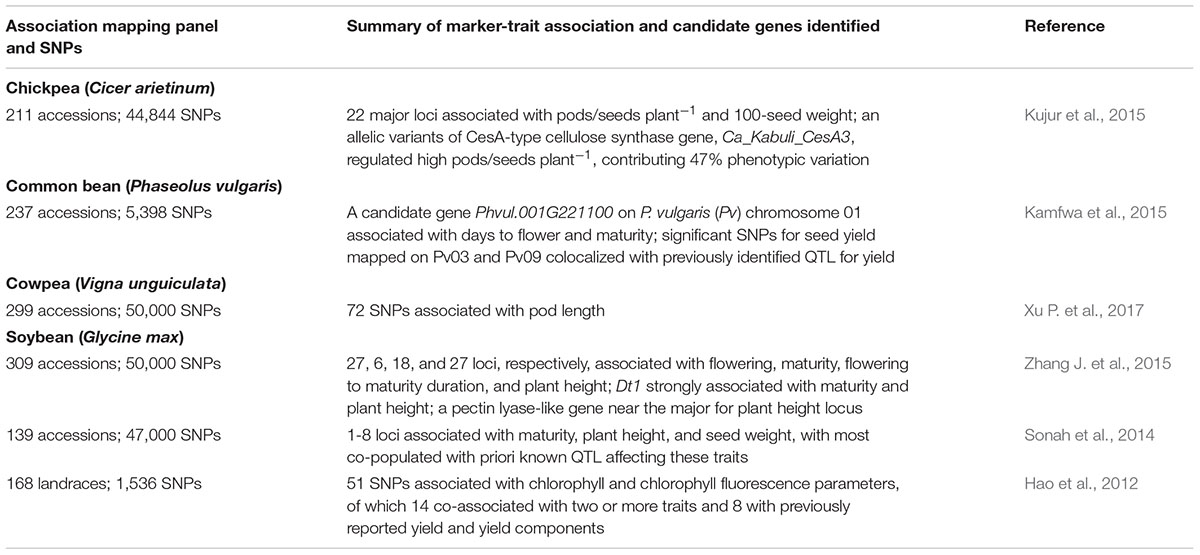

TABLE 3. SNPs and germplasm-based genome-wide association studies (GWAS) for phenology and yield in chickpea, common bean, cowpea and soybean.
Plant Architecture and Edible Yield in Cereals
Domestication and subsequent artificial selection by humans has dramatically changed plant architecture, phenology and components of grain yield in many cereals, largely to address agronomic needs and to adapt the crops to various stress-prone environments. Candidate genes and SNPs associated with crop phenology, plant architecture, and yield-attributing traits are known in cereals (Table 4). Several unique candidate gene regions related to plant growth and development and grain yield have been identified in maize (Farfan et al., 2015; Li X. et al., 2016). Bouchet et al. (2017) found 34 and 6 QTL for individual or combinatorial trait combinations in maize, respectively. They identified a QTL cluster in a 5 Mb region around Tb1 associated with tiller number and ear row number. The latter was positively correlated with flowering (days to anthesis for male and female flowering and anthesis to silking interval measured in days) and negatively correlated to grain yield. Kn1 and ZmNIP1 have been identified as candidate genes for tillering, along with ZCN8 for leaf number and Rubisco Activase 1 for kernel weight. A more upright leaf in maize has been shown to be influenced by variation in liguleless genes (Tian et al., 2011).
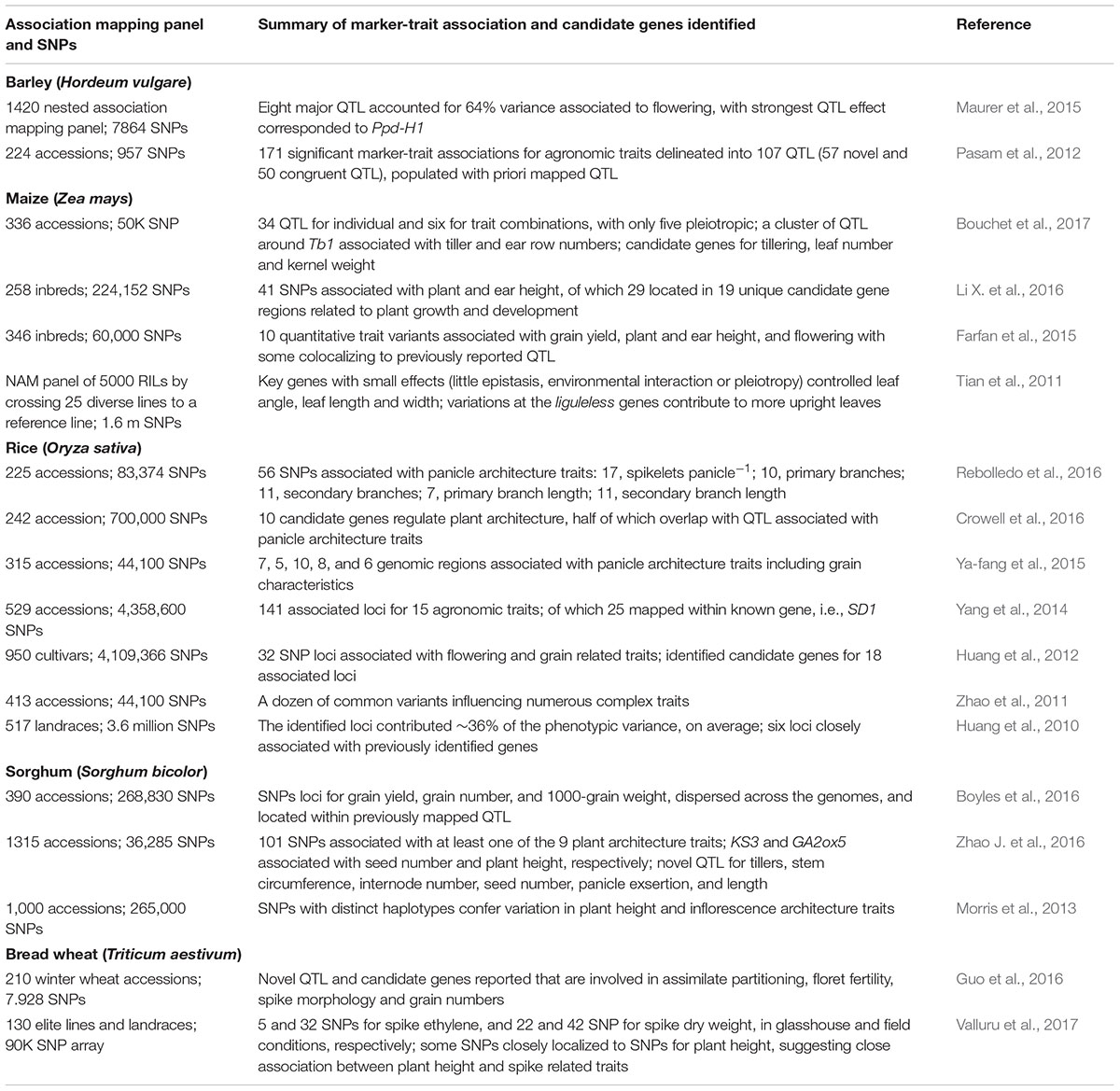
TABLE 4. SNPs and germplasm-based genome-wide association studies for plant architecture traits in barley, maize, rice, sorghum, and wheat.
A large GWAS study in rice detected 42 significant genotype–phenotype associations for plant morphology, grain quality, and root architecture traits, which in most cases were co-localized with QTL and candidate genes controlling the phenotypic variation of single or multiple traits (Biscarini et al., 2016). Several SNPs in rice were associated with plant and panicle architecture, biomass and yield (Zhao et al., 2011; Yang et al., 2014; Ya-fang et al., 2015; Rebolledo et al., 2016), while candidate genes in pathways regulating plant architecture overlap with QTL associated with panicle architecture traits (Crowell et al., 2016; Rebolledo et al., 2016).
In wheat, candidate genes associated with SNPs were involved in carbohydrate metabolism, floral fertility, spike morphology and grain number, providing valuable targets for selection (Guo et al., 2016). Significant marker-trait associations also provided insight into genetic architecture of flowering, plant height and grain weight in barley (Pasam et al., 2012). Individual QTL accounted, however, only for a small portion of phenotypic variation.
In sorghum, several SNPs were associated with plant and inflorescence architecture traits, with many located within previously mapped QTL (Morris et al., 2013; Maurer et al., 2015; Boyles et al., 2016; Zhao J. et al., 2016). Candidate genes KS3 (associated with seed number) and GA2ox5 (associated with plant height) were also reported (Zhao J. et al., 2016). A QTL with a major effect corresponded to the priori known photoperiod response gene Ppd-H1 (Maurer et al., 2015).
Abiotic Stress Adaptation in Soybean
Multiple SNPs are reported to be associated with tolerance to drought and heat stress in soybean (Table 5). Dhanapal et al. (2015) reported 39 SNPs associated with carbon isotope ratio (δ13C), which is a surrogate trait to measure water use efficiency. The genomic distribution of these SNPs revealed that several are co-located and likely tag the same locus, suggesting that markers for δ13C can be identified in soybean using GWAS. Dhanapal et al. (2016) reported 52 unique SNPs for total chlorophyll content tagged on 27 loci across 16 chromosomes. While many of these putative loci were near genes previously identified or annotated as related to chlorophyll traits (Hao et al., 2012), numerous SNPs marked chromosomal regions with unknown-function genes.
Non-photochemical quenching (NPQ) under abiotic stress conditions protects plants from heat when more light is absorbed than can be used for photosynthesis (Li et al., 2009). Canopy reflectance measured as photochemical reflectance index (PRI), amenable for high throughput field phenotyping, is a surrogate to measure NPQ (Gamon et al., 1992). Thirty-one PRI-specific SNPs, tagged in 15 loci on 11 chromosome harboring candidate genes associated with NPQ, photosynthesis, and sugar transport, may provide an opportunity to improve photosynthesis in soybean (Herritt et al., 2016).
Abiotic Stress Adaptation in Cereals
Cereal crops have been extensively investigated for SNPs and candidate genes associated with abiotic stress adaptation (Table 6). Ethylene levels have been linked to yield penalty under heat stress in wheat, largely due to reduction in spike fertility and grain weight (Hays et al., 2007). Valluru et al. (2017) reported 5 and 32 significant SNPs associated with spike ethylene, and 22 and 142 significant SNPs associated with spike dry weight, in greenhouse and field studies, respectively. Some of these SNPs are close to SNPs associated with plant height, suggesting associations between plant height and spike-related traits. This opens the possibility of gene discovery and breeding of wheat cultivars with reduced ethylene effects on yield under heat. The D genome progenitor of bread wheat Aegilops tauschii has potential as an excellent source of abiotic stress tolerance. Qin et al. (2016) reported 25 SNPs and several putative candidate genes (enzyme, storage protein, and drought-induced protein) associated with drought adaptation, while Liu Y. et al. (2015) found 13 SNPs and putative candidate genes related to P-deficiency tolerance.
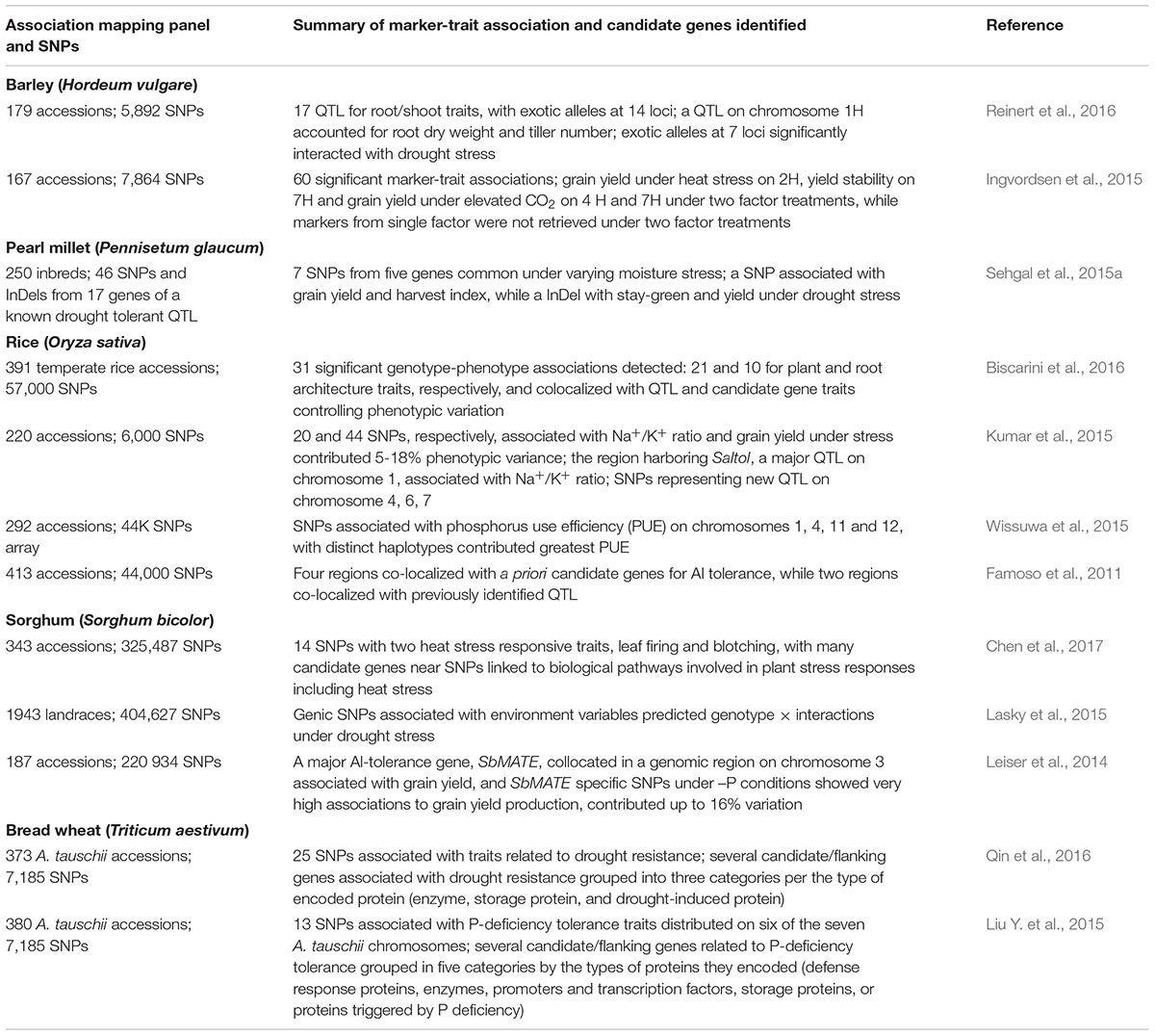
TABLE 6. SNPs and germplasm-based GWAS for abiotic stress tolerance in barley, pearl millet, rice, and sorghum.
A major Al-tolerance gene SbMATE on chromosome 3 has been shown to be associated with grain yield in sorghum, where SbMATE specific SNPs under –P conditions contributed up to 16% genotypic variance (Leiser et al., 2014). Forty-eight genomic regions associated with Al tolerance were reported in rice, four of which co-localized with a priori known candidate genes, and two co-located with previously identified QTL (Famoso et al., 2011).
In barley, a genomic region on chromosome 2H was associated with grain yield under heat stress, a region on chr 7H with grain yield, and a region on chr 4H and chr 7H with elevated CO2 under two factor treatments (high temperature and elevated CO2). None of the SNPs associated with single factor treatments were retrieved under two factor treatments, thus emphasizing the importance of multifactor treatments (Ingvordsen et al., 2015).
Genic SNPs associated with environmental variations (but independent of geographical location) predicted genotype × environment interactions for drought stress and aluminum toxicity in sorghum (Lasky et al., 2015). Wissuwa et al. (2015) reported several SNP loci associated with phosphorus use efficiency (PUE) in rice on chromosomes 1, 4, 11, and 12. A minor indica-specific haplotype on chromosome 1 and a rare aus-specific haplotype on chromosome 11 displayed the highest PUE, and could have potential for targeted introgression while breeding for rice under P-limited cropping systems.
Emerging evidence suggests that responses to stress combinations cannot be reliably predicted from the responses to individual stresses (Makumburage et al., 2013). An integrated approach is therefore needed to model the genetics of responses to a range of single and combined stresses. For example, association analysis report QTL with contrasting and with similar responses to biotic versus abiotic stresses, and below-ground versus above-ground stresses. There is a need to conduct multi-trait GWAS to identify robust candidate genes for multiple stresses (Thoen et al., 2016).
The proliferation of genome wide association analyses has led to identification of candidate loci (often co-located with major QTLs or candidate genes) associated with abiotic stress adaptation, phenology and plant architecture, and edible yield. The identification of such loci can facilitate genomics-assisted breeding in cereal and legumes.
TILLING: Mutagenesis and Reverse Genetics for Elucidating Gene Function
Chemical mutagenesis and subsequent screening for mutations linked to altered agronomic phenotypes is a reverse genetic technique to identify candidate genes for crop improvement. Targeting Induced Local Lesions IN Genomics (TILLING) is the commonly used approach, employing a mismatch-specific endonuclease to detect single base pair (bp) allelic variation in a target gene (Gilchrist and Haughn, 2005). TILLING by sequencing (Tsai et al., 2011) can greatly increase throughput and novel allele discovery by applying second-generation sequencing approaches rather than endonucleases to facilitate variant discovery across the genome rather than in individual genes (Henry et al., 2014; Kumar et al., 2017). TILLING has been successfully used to detect both induced and natural variations in a wide range of plant species, including: novel allelic variation in the barley genes HvCO1, Rpg1, elF4E, HvHox1, BMY1, GBSS1, LDA1, SSI, SSlla, mlo and Mla (Mejlhede et al., 2006; Talamé et al., 2008; Gottwald et al., 2009; Sparla et al., 2014); the maize genes DMT101, DMT102, DMT103, DMT 106, HAC110, HDA105 (Till et al., 2004); and the wheat genes PpD-1, Rubisco activase A and Rubisco activase B (Chen et al., 2012). In sorghum, TILLING generated a functional-effect point mutation in the CYP79A1 gene, generating sorghum lines with reduced levels of the cyanogenic glycoside dhurrin, which has potential to enhance the use of this widely grown crop as forage for livestock (Blomstedt et al., 2012). A TILLING-induced mutation in a TI1 protease inhibitor increased the digestibility and thus nutritional value of pea. Although mutagenesis in TILLING approaches is untargeted and does not provide the versatility of genome editing, crops improved using chemical or radiation mutagenesis via TILLING are not regulated as GMOs in most jurisdictions, increasing their commercial competitiveness with more precise genome editing approaches (Kumar et al., 2017).
Using Haplotypes to Identify Alleles in Cultigen Pools
A haplotype is a combination of DNA polymorphisms (markers, alleles) that are tightly linked to each other on a chromosome and hence tend to be inherited together from parent to offspring. Maize was among the first crops for which a comprehensive haplotype map was generated, which showed highly divergent haplotypes and recombination rates based on several million sequence polymorphisms in 27 diverse inbred lines (Gore et al., 2009). This research also identified hundreds of selective sweeps and highly distinct chromosome regions likely bearing loci related to domestication and geographic adaptation. Genetic structure and subpopulation structure are also associated with origin of germplasm and post-domestication selection, as revealed by comparative haplotype analysis in tropical and temperate maize germplasm (Lu et al., 2011). Moreover, Thirunavukkarasu et al. (2017) were able to identify 252 haplotype blocks in subtropical elite inbred maize lines, which varied in size from 1 to 15.8 Mb, with slow LD decay (200–300 Kb) across all chromosomes, suggesting selection of favorable traits around low LD regions in breeding programs. Due to strong population substructure, this subtropical maize germplasm grouped into three distinct clusters, which provides means for exploiting heterotic potential among them. The use of haplotypes improved mapping efficiency to detect QTL related to drought adaptation in maize (Lu et al., 2009). Furthermore, integrated mapping (based on independent linkage and LD analysis) along with haplotypes led to identification of significant QTL explaining up to ca. 35% of phenotypic variation. Two significant haplotypes were involved in the control of flowering time, and encoding aldo-keto reductases associated with detoxification pathways contributing to cellular damage due to stress.
There is a continual need to identify allelic variants conferring desirable agronomic traits. For example, recent haplotype analysis in Indian wild rice identified the variants H5 and H1 of HKT1;5 and HKT2;3 as associated with high salinity tolerance (Mishra et al., 2016b). Haplotype variation of major and few minor alleles seems to be distributed over distant geographic regions (Mishra et al., 2016a). Such alleles may be useful for broadening the range of cultivars to enhance rice productivity in salt-prone areas. The rice DNA markers RM 464A and RM 219 at the Sub-1 locus of chromosome 9 (which accounts for 70% of phenotypic variation for submergence tolerance) have assisted in breeding cultivars that are tolerant to submergence for up to 2 weeks during the vegetative growth stage (Rathnayake et al., 2012). The Sub-1 locus encodes the ethylene-responsive factor (ERF) genes sub1B (from the submergence tolerant FR13A landrace) and Sub1C in all Oryza sativa cultivars, while the ERF paralog Sub1A is found in a subset of O. sativa ssp. indica accessions, and seems to arise from duplication of Sub1B (Fukao et al., 2009). Some submergence tolerant rice accessions lack Sub1A (Tamang et al., 2011), which appears to suppress leaf elongation under submergence (Singh et al., 2010). This suggests that Sub1A may not be the only contributing factor to submergence tolerance in rice (Samal et al., 2014).
The haplotype map of disomic hexaploid bread wheat, based on resequencing 62 wheat lines using exome capturing and genotype-by-sequencing, has exposed distinct patterns of directional selection in homeologous genomes (Jordan et al., 2015). This finding suggests that the likelihood of beneficial allele recovery was increased in bread wheat by broadening the set of selection targets. Haplotype analysis of stem rust resistance genes revealed that most breeding lines (83 out of 115) released by CIMMYT until the 2000s carry Sr2. Five were found to carry the Sr25 haplotype, while a small number of (5 out of 22) cultivars bred by the United States Department of Agriculture haboured the Sr2, Sr24, Sr36 haplotypes. Sr2 was also found in two out of 43 wheat breeding lines from China (Yu et al., 2010). Diverse bread wheat lines bred in different Africa countries have been found to harbor the Sr2, Sr36, Sr24, Sr31and Lr34/Yr18/Sr57 haplotypes (Prins et al., 2016). Tetraploid Ethiopian durum wheat landraces and bred cultivars carry the Sr2 and Sr22 haplotypes, with only a few bearing Sr13 (Haile et al., 2013). Haplotype analysis also located the origin of Sr33, an ortholog of a barley mildew resistance Mla gene that was introgressed to bread wheat from the wild relative Aegilops tauschii (Periyannan et al., 2013). Such analyses can identify sources of novel alleles for use in improving host plant resistance through breeding.
Landraces and Wild Relatives
Landraces are a repository of crop genetic diversity that have evolved through natural and artificial selection over millennia, and represent valuable resources for crop adaptation to stresses. For example, the allelic variation amongst rice and wheat landraces has provided agronomically beneficial traits for abiotic stress tolerance (Dwivedi et al., 2016). Pasam et al. (2014) noted that widely adapted (5°–62.5° N, 16°–71° E) spring barley landraces (LRC1485), which showed abundant genetic diversity, clustered into six major germplasm groups, differentiated by geographical origin and latitude, ear row type, caryopsis types, and climate zones. Creole wheat landraces introduced into Mexico from Europe are adapted to a wide range of climatic regimes and represent a useful genetic resource. Vikram et al. (2016) characterized 9,416 landrace accessions using genotyping-by-sequencing and identified 15 genetic groups that are likely adapted to specific environments of Mexico, with some groups adapted to extreme environments. For example, landraces from Michoacán (high temperature and rainfall) and Durango (high annual average temperature and low precipitation) had an exceptionally high frequency of rare alleles, which may be a contributing factor of landrace adaptation to these climates. A similar study on local adaptation of barley landraces in Ethiopia revealed that environmental differences (temperature and precipitation) and geographic effects contributed 40 and 29% of the explained genetic variation, respectively (Abebe et al., 2015). Pearl millet landraces (249 accessions) from Senegal were genetically distinct from many global accessions1, 262 accessions from Africa, Asia, and the American contents, with the greatest representation from India, Kenya, South Africa, Yemen, and Zimbabwe, showed little population structure, and higher-levels of linkage disequilibrium decay, providing a valuable resource for use in breeding (Hu et al., 2015). Population structure analysis involving cowpea landraces and wild relatives delineated most African landraces into two major genepools, with most landraces from West Africa forming genepool 1, while the majority of the landraces in genepool 2 were from East Africa. Furthermore, the authors noted that each genepool was closely related to wild cowpea in the same geographic region, suggesting divergent domestication leading to the formation of two genepools in cowpea (Huynh et al., 2013). Lentil (Lens culinaris) landraces (predominantly from Greece and Turkey) also have revealed high levels of genetic diversity (Lombardi et al., 2014).
Wild and weedy relatives of crops are an important source of adaptation and stress tolerance genes. Wild species often grow in harsh environments and therefore could be the source of genes conferring abiotic stress adaptation. The greatest impact of wild relatives in crop improvement to date have been in increasing host plant resistance to pathogens and pests in several crops. Wild species have also been the source of genes for edible yield and quality traits in some crops (Dwivedi et al., 2008). Poets et al. (2015) compared SNP polymorphism between landraces and wild barley accessions and noted that landraces comprised multiple source populations with unequivocal contributions from wild barley populations across the genome. Furthermore, two genomic regions on the 2H and 5H chromosomes contributed to geographic differentiation in allele frequencies (Fang et al., 2014). Wild barley accessions collected at ‘Evolution Canyon’ at Nahal Oren, Israel were more genetically diverse than those from other regions in northern Israel, while those from the hot and dry south-facing slope were genetically more distinct from north-facing slope accessions (Bedada et al., 2014).
A study on genetic basis of phenotypic variations among wild pearl millet populations from two north–south aridity gradients in West Africa revealed that the size of the inflorescence, the number of flowers and above-ground dry mass co-varied positively with rainfall decrease. Moreover, two SNPs located in the Myosin XI gene were significantly associated with variation in the average flower number. Both the allele frequency of the two SNPs and the average flower number co-varied with the rainfall gradient on the two gradients. Myosin XI is a good candidate for fitness-related adaptation in wild populations (Ousseini et al., 2017). Structure analysis of 99 ecotypes of wild soybean, sampled across their native geographic range and genotyped by SoySNP50K array, identified four genetic groups that largely corresponded to geographic regions of central China, northern China, Korea, and Japan, with high levels of admixture between genetic groups. Moreover, the environmental factors contributed 23.6% to population differentiation, while geographical factors accounted for 6.6%. Precipitation variables explained divergence of the groups along longitudinal axes, whereas temperature variables contributed more to latitudinal divergence (Leamy et al., 2016).
Such delineation of landraces and wild relatives into groups of genetic relatedness associated with geographic or environmental differences, and identification of accessions harboring higher numbers of rare alleles (with functional effects) will be valuable genetic resources in breeding and for improving the management and utilization of germplasm in crop improvement. There are significant barriers (both pre-fertilization and post-fertilization) to inter-specific hybridization. Technology for circumventing these barriers are required for increased introgression of allelic variation from wild relatives to primary crop genepools (Dwivedi et al., 2008).
Key Breeding Targets to Enhance Adaptation and Productivity
Optimizing Crop Productivity Using Mutagenesis in the Florigen Pathway
Flowering time, the transition from vegetative to reproductive growth, is a major determinant of crop yield (Cockram et al., 2007; Jung and Muller, 2009). The universal flowering activator florigen has several genetic components, including the key FLOWERING LOCUS T (FT) gene (Koornneef et al., 1991; Samach et al., 2000; Turck et al., 2008). While FT and FT-like genes generally activate flowering, another group of genes including TERMINAL FLOWER 1 (TFL1) and TFL1-like genes act as flowering repressors (Karlgren et al., 2011). These genes belong to the CENTRORADIALIS/TERMINAL FLOWER 1/SELF-PRUNING (CETS) gene family, which display sequence similarity to the phosphatidylethanolamine binding protein (PEBP) genes. The balance and interplay between flowering activators and repressors determines flowering response. Selection of variant genes in the florigen pathway to increase environmental flexibility and yield has played in important role in the domestication and improvement of many crops, including barley (Comadran et al., 2012) and rice (Ogiso-Tanaka et al., 2013).
Recently, more targeted breeding approaches have used florigen pathway genes to control flowering in crops. A study in tomato showed that reproductive growth could be influenced by combining mutations in SINGLE FLOWER TRUSS (SFT; an FT homolog) and a bZIP transcription factor within the florigen pathway, to produce plant architecture increasing yields (Park et al., 2014). Similarly, SFT heterozygosity in tomato was also observed to alter plant architecture (Jiang K. et al., 2013). Using the CRISPR/Cas9 system in tomato, it has been demonstrated that mutation of the floral repressor gene SP5 can produce early yielding plants (Soyk et al., 2017).
Rational engineering of flowering time remains constrained by a lack of knowledge on the functions of many components of the florigen pathway in different plant species, which contains many closely related genes with diverse functions, including unknown interaction and regulation networks. Because of the diversification of the florigen pathway genes in flowering plants, an improved understanding of species-specific florigen pathways will be important for crop breeding (Zhang et al., 2010). Recent studies investigating genetic control of flowering have identified an important regulator of florigen transport in rice (Song et al., 2017). Other studies have identified a loss of vernalization requirement in narrow-leafed lupin (Lupinus albus), caused by a deletion in the FT promoter (Nelson et al., 2017). In Arabidopsis thaliana, mutagenesis of codons within the FT gene identified differences critical for the related antagonist TFL1 and indicated potential candidate transcription factors interacting with the protein. As knowledge of species-specific flowering mechanisms develops, fine-tuning of the florigen pathway should allow yield increases through better control of flowering and growth in crops.
Enhancing Abiotic Stress Adaptation
Breeding for adaptation to abiotic-stress remains a challenging task (Dwivedi et al., 2010, 2017; Kole et al., 2015). Conventional crossing and selection for abiotic stress adaptation had had limited success. However, when supported by applied genomics tools and genetic engineering, accelerated introgression of beneficial alleles has enhanced yield and abiotic stress adaptation in cereals and legumes.
Six large-effect QTL related to drought adaptation have been shown to be effective in multiple genetic backgrounds and production environments in rice. Pyramiding of these large-effect QTL has improved drought adaptation of widely grown Asian cultivars (Kumar et al., 2014). Submergence tolerant rice cultivars bearing SUB1A-1 have had significant impacts in Asia (Bailey-Serres et al., 2010; Dar et al., 2013). Furthermore, pyramiding SUB1A-1 and drought-tolerant QTL (Kumar et al., 2014) or salt (Saltol1) and flood (Sub1) tolerance QTL (Mackill et al., 2012) is expected to lead to improved cultivars adapted to stress-prone lands in Asia. The Pup1 allele increases P uptake and confers significant grain yield advantage in rice grown on P-deficient soils (Wissuwa et al., 2002). Introgressed lines containing Pup1 significantly increased grain yield on P-deficient soils (Chin et al., 2011). Also, overexpression of a Pup1-specific protein kinase gene (PSTOL1) significantly enhances grain yield in P-deficient soils. POSTL1 promotes early root growth, thereby enabling plants to acquire more P and other nutrients (Gamuyao et al., 2012).
Identifying SNPs that are robustly associated with environmental adaptation can be useful for crop improvement. Environment-associated SNPs independent of geographic origins predicted genotype × environment interactions under drought stress and aluminum toxicity in sorghum (Lasky et al., 2015). Climatic variables accounted for most of the genetic variation in barley (Abebe et al., 2015). Many SNPs were associated with putative adaptive loci and candidate genes conferring enhanced adaptation. In barley, five and two SNPs were correlated with length of growth season and precipitation, respectively (Selçuk et al., 2015). However, none were correlated with both. More recently, Russell et al. (2016) noted extensive sequence variations amongst known flowering associated barley genes HvCEN, HvPPD, and HvFT1 (Turner et al., 2005; Casas et al., 2011; Comadran et al., 2012), with haplotypes exhibiting strong geographical structuring, likely contributed to range-wide ecogeographical adaptation. In maize, Westengen et al. (2012) noted that 79 and 22 SNPs associated with maximum temperature and mean precipitation, respectively, with many located in genes functioning in abiotic stress adaptation. Non-synonymous SNPs clustered in the region harboring six known QTL associated with relatively high phenotypic variation for drought adaptation in maize (Xu et al., 2014). Pyhäjärvi et al. (2013) observed that SNPs in wild teosinte ancestor were associated with altitude. Millet et al. (2016) noted 8 and 12 QTL associated with heat and drought stress adaptation in maize, respectively, with low or negative effects in favorable environments. Another 24 QTL improved yield in favorable environments but without any effects under stress, thereby indicating that QTL effects were expressed as functions of environmental variables and scenarios. Such genomics-assisted knowledge can potentially be used to accelerate breeding for drought-prone environments (Millet et al., 2016).
African pearl millet landraces collected in 2003 displayed a short life cycle, reduced plant and spike size, and increased frequency of early flowering alleles (from 9.9% in 1976 to 18.3% in 2003) at the flowering locus PHYC (Saïdou et al., 2009), which suggests that recurrent drought may promote shortening of growth duration in pearl millet (Vigouroux et al., 2011). Similar observations were noted after monitoring changes in functional diversity due to possible climate effects in wild emmer wheat and wild barley populations. Populations collected in 2008 flowered earlier than those collected in 1980, with greater shortening of flowering time after 28 years for wild barley than wild emmer wheat. However, the study indicated that emmer wheat lost more alleles than wild barley. The allelic reduction in emmer wheat was negatively correlated with altitude (-0.854∗) and humidity (-0.673∗), while in barley the difference between the sampling years was positively correlated with rainfall (0.790∗) but negatively with evaporation (-0.692∗) (Nevo et al., 2012).
Villordo-Pineda et al. (2015) have identified 37 SNPs with a potential drought adaptation function in common bean. A ‘QTL-hotspot’ region harboring 12 QTL associated with drought adaptation traits contributed up to 58% of the phenotypic variation in chickpea (Varshney et al., 2014). Indeed, introgressions containing this region in JG 11, a widely grown cultivar in India, have improved root traits and drought tolerance (Varshney et al., 2013). Subsequently, Kale et al. (2015) fine-mapped this ‘QTL-hotspot’ and identified four candidate genes from this region that are associated with drought tolerance. Anderson et al. (2016) identified several candidate loci that putatively contributed to adaptation to abiotic stresses, which may permit targeted use of Glycine soja germplasm for enhancing the genetic potential of cultivated soybeans. Qi et al. (2014) noted that sequence variations in GmCHX1 were associated with salt tolerance in a wild soybean, W05. Likewise, 20 loci associated with P efficiency-related traits have been identified in soybean, some coinciding with known P efficiency-related genes GmACP1 and GmPT1, while Glyma.04G214000 and Glyma.13G161900 displayed differential expression in low-P soils (Zhang et al., 2016c).
The evidence to date suggests that understanding (and deployment) of major QTL or candidate genes associated with abiotic stress adaptation has led to the development and release of several maize and rice cultivars adapted to different abiotic stresses (Ortiz, 2013). A large effort is underway to introgress major QTL associated with drought and heat stress adaptation in common bean and chickpea (Dwivedi et al., 2017). The discovery of several SNPs associated with variation in both temperature and precipitation responses in barley, maize and sorghum provide a further opportunity to develop cultivars with enhanced fitness in the context of a changing climate.
Phenology, Yield and Adaptation
Crop Duration and Yield
Understanding the nucleotide variation and mechanism of molecular evolution of flowering, maturity and plant height genes could accelerate the development of cultivars of specific duration to better adapt them to growing seasons. Knowledge on sequence variation in genes related to plant or panicle architecture (and yield) may provide opportunities to genetically enhance crop productivity per se.
Rice adaptation to climate is influenced by days to flowering and its sensitivity to photoperiod variation. OsPRR37 (PRR37) is within the Early heading 7-2 (EH7-2)/Heading date 2 (Hd2) QTL in rice. The japonica cultivars having Ghd7/Hd4 and PRR37/Hd2 non-functional alleles flower early under extended photoperiod, and are adapted to the northernmost region of cultivation, up to 53°N latitude. Genetic analysis reveals that the effects PRR37 and Ghd7 effects on heading date are additive (Koo et al., 2013). PRR37 down-regulates Hd3a expression to suppress flowering under extended photoperiods, thus suggesting that PRR37/Hd2 and Ghd7/Hd4 contributed to adaptation of rice in temperate and cool regions. Further investigation using accessions from the O. japonica core collection have revealed that RICE FLOWERING LOCUS T1 (RFT1) is the major contributor to flowering among japonica cultivars adapted to northern areas (Naranjo et al., 2014). Ghd7 is a gene with pleotropic effects that controls plant height, heading date and yield in rice. Lu et al. (2012) noted 76 SNPs and six indels within a 3932bp DNA fragment of Ghd7 derived from two distinct ancestral genepools (indica and japonica), of which SNP S_55 was associated with plant height while another seven SNPs were in complete linkage with spikelets per panicle, regardless of photoperiod. Their finding suggests major flexibility of Ghd7 to improving phenology, panicle architecture, and yield in rice.
The genes IDEAL PLANT ARCHITECTURE (IPA), LONG PANICLE1 (LP1), SPIKELET NUMBER (SPIKE), Gna1 (grain number), Ghd7 (grain number, plant height, and flowering), GS3 (grain weight and length), GW5 (grain weight), and DEP1 (DENSE AND ERECT PANICLE1) greatly influence panicle architecture and seed yield in rice (Ashikari et al., 2005; Fan et al., 2006; Weng et al., 2008; Xue et al., 2008; Huang et al., 2009; Jiao et al., 2010; Miura et al., 2010; Fujita et al., 2013; Liu E. et al., 2016). DEP1 locus has been widely used for developing high yield rice cultivars with erect panicle architecture (Yan et al., 2013). Mining allelic variations for panicle traits unravels 45 SNPs and 26 InDels within the DNA fragment of DEP1 and replacement of 637 bp by 12 bp fragment explain most of the phenotypic variations for panicle architecture, and SNP(G/C) largely affects branches and grains panicle-1 (Zhao M. et al., 2016).
Ghd7 (Ma6) and pseudoresponse regulator protein 37 (PRR37) alleles in sorghum confer differences in photoperiod sensitivity and flowering times that are critical for production of high-biomass energy or grain sorghum (Murphy et al., 2011, 2014). Furthermore, Wang Y. et al. (2014) investigated nucleotide diversity of Ma3, another maturity gene in sorghum, and identified three and 17 SNPs that affected flowering at high-latitude and at low-latitude environments, respectively. Indeed, a major QTL on chromosome 6, FlrAvgD1, which contributed 85.7% of variation in flowering under LD, was narrowed to a 10 kb interval containing the only one annotated protein-coding gene (Sb06g012260) with potential to accelerate cross-utilization of temperate and tropical germplasm for production of grain or bioenergy sorghum types (Cuevas et al., 2016).
The discovery and deployment of alleles of semi-dwarfing genes have contributed to enhanced lodging resistance and increased productivity in both rice and wheat (Peng et al., 1999). For example, the use of semi-dwarf1 (sd1) in rice and reduced height (Rht) alleles in wheat, which encode a GA biosynthesis enzyme and a dominant suppressor protein of GA signal transduction, respectively, have been widely used to confer lodging resistance in these crops (Peng et al., 1999; Sasaki et al., 2002). A semi-dwarfing gene (sdw1) locus has been widely introgressed into barley cultivars grown worldwide. At least four alleles (sdw1.a, sdw1.c, sdw1.d, and sdw1.e) have been reported (Franckowiak and Lundqvist, 2012). The gibberellin 20-oxidase gene (HvGA20ox2) is the functional gene of sdw1 mutants, and deletions resulted in different functional alleles for breeding purposes. Diagnostic markers can differentiate the wild type allele from the sdw1.d, sdw1.a, and sdw1.c alleles (Xu Y. et al., 2017). In sorghum, four unlinked dwarfing genes (Dw1–Dw4) were combined to reduce plant height to increase lodging resistance and improve mechanized harvesting (Quinby and Karper, 1954). Of these, only Dw3 has been cloned (Multani et al., 2003). Yamaguchi et al. (2016) isolated the Dw1 gene, which encodes a novel uncharacterized protein. A histological analysis comparing the Near Isogenic Line (NIL)-dw1 with that of wild type showed similar longitudinal parenchymal cell lengths of the internode, but significantly reduced number of cells per internode in NIL-dw1. NILs containing dw1 and dw3 displayed a synergistic phenotype, which contributes to improved lodging resistance and mechanical harvesting in sorghum (Yamaguchi et al., 2016).
To date, 10 major genes (E1 to E9 and J) and several QTL have been shown to be involved in control of flowering in soybean. Different allele combinations at E1-E4 and E9 loci produce diverse flowering habits in soybean cultivars (Xu et al., 2013; Kong et al., 2014; Tsubokura et al., 2014; Zhao C. et al., 2016). Multi-locus genotypes involving E1 to E4 account for 62–66% of natural variation in the flowering time and identified a new allele in E1 locus, e1-re, for flowering in soybean (Tsubokura et al., 2014). FT2a and FT5a, the orthologs of FLOWERING LOCUS T (FT) (Kong et al., 2010), play a major role in initiation of flowering. Indeed, their expression in response to photoperiod is controlled by different allelic combinations involving E1 to E4. More recently, Takeshima et al. (2016) identified a QTL in LG J which was localized to a genomic region of 107 kb (harboring FT5a). The study detected SNP polymorphisms between the parents involving early (ef) and late (lf) flowering alleles, and also detected ef, a rare haplotype distinct from others including lf. A higher transcript abundance of FT5a in NILs containing ef allele suggests that differential transcriptional activities or mRNA stability may cause differences in flowering (Takeshima et al., 2016).
An investigation of flowering time variation and SNP polymorphisms in key regulatory genes in common bean revealed that PvVRN1 and Pv PHYB are associated with days to flowering, PvMYB29 with number of flower buds per inflorescence, and PvTFL1z and PvFCA with inflorescence length (Raggi et al., 2014). More recently, a QTL on linkage group Pv01, harboring the Phvul.001G189200 gene (with sequence similarity to the TERMINALFLOWER1 (TFL1) gene in Arabidopsis thaliana), explained up to 32% of phenotypic variation for time to flowering, 66% for vegetative growth, and 19% for rate of plant production, supporting Phvul.001G189200 (referred as PvTFL1y) as a candidate gene for determinacy locus in common bean (González et al., 2016). CcTFL1, a candidate gene for determinacy in pigeonpea, contributed substantial phenotypic variations for determinacy (45–96%), flowering (45%) and plant height (77%) (Mir et al., 2014). Foucher et al. (2003) isolated three TFL1 homologs, PsTFL1a, PsTFL1b, and PsTFL1c in pea. PsTFL1a controls indeterminacy of the apical meristem during flowering, while PsTFL1c delays the induction of flowering by lengthening the vegetative phase in pea. The development of genetic markers has potential to allow manipulation of the determinacy trait in these and other legume species.
Understanding the molecular basis of allelic variation associated with flowering, plant height, maturity, plant architecture, and yield provides opportunities to tailor crop ideotypes that are better adapted to specific agro-ecosystems or meeting end-use preferences.
Tropical vs. Temperate Adaptation
Although maize, rice and sorghum were domesticated in tropical regions, they are all commercially grown both in tropical and temperate climates. Understanding the molecular basis of such adaptation differences for these important cereal crops is critical for targeted introgression of beneficial alleles from one genepool to another, or for developing grain or bioenergy sorghum types. Sorghum is a short-day plant requiring a daylength below 12 h 20 min to induce flowering. Hence, most of the tropical sorghum germplasm flowers too late or is too tall to be exploited for seed production in temperate environments. In the 1970s a large-scale sorghum conversion program was initiated by USDA-ARS to convert tropical accessions to plants adapted to temperate zones by introgressing recessive day-neutral flowering alleles and dwarf-height genes into the exotic backgrounds via a backcross scheme to recover the exotic genome in early flowering, combine-height inbred lines (Stephens et al., 1967). This effort resulted in the release of 40 such converted lines for use in temperate zones worldwide (Klein et al., 2016). Three genomic regions, each with multiple linked loci for phenology (plant height and flowering), have been found to control adaptation of grain type sorghum in temperate zones (Thurber et al., 2013).
Although maize is highly sensitive to low temperature, there is natural variation in freezing and chilling tolerance. The mechanisms responsible for chilling tolerance include modification of photosynthetic apparatus modification, cell wall properties, and developmental processes (Sobkowiak et al., 2016). Differential gene expression in response to freezing identified nine candidate genes with higher expression levels and eight candidate genes with lower expression levels in the tolerant compared to the intolerant lines (Li Z. et al., 2016; di Fenza et al., 2017). Dent and flint maize, which differ in their kernel phenotypes, represent two major temperate gene pools in maize. The flint contributes to early vigor and cold tolerance, while dent increases productivity in hybrids. Unterseer et al. (2016) identified candidate genes under differential selection pressure in these two genepools. Most flint-specific candidate genes were associated with endogenous pathways, whereas dent candidate genes were mainly involved in response to environmental factors such as light and photoperiod.
Low temperature is one of the major constraints limiting rice productivity and cultivation in high-altitude regions. The major rice QTL COLD1, which functions as a regulator of G-protein signaling, confers chilling tolerance. The allele SNP2 in COLD1jap/ind enhances the ability to activate G-protein α GTPase, as COLD1 interacts with G protein to activate the Ca2+ channel for temperature sensing (Ma et al., 2015). A novel gene CTB4a controlling cold tolerance at booting stage in rice enhances seed setting and grain yield under cold stress conditions (Zhang et al., 2017). Oryza glaberrima has contributed the major QTL OgTT1, which confers adaptation to heat stress. OgTT1 protects cells from heat stress through more efficient elimination of cytotoxic denatured proteins and more effective maintenance of heat-response processes. Overexpression of this gene was associated with markedly enhanced thermotolerance in rice, Arabidopsis and Festuca elata (Li et al., 2015). Thus, deployment of QTL conferring chilling- and thermo-tolerance is expected to aid development of rice cultivars with enhanced adaptation to these climatic variables.
Soybean was domesticated in temperate regions and is highly sensitive to photoperiod. However, temperate soybean cultivars are not adapted to tropical and sub-tropical climates. The discovery of the long juvenile (LJ) trait in tropical soybean germplasm and its deployment has extended cultivation of temperate soybean to low altitude tropical and sub-tropical climates (Hartwig and Kiihl, 1979; Neumaier and James, 1993). A major locus J identified as the ortholog of Arabidopsis ELF3 confers the LJ trait: J promotes flowering, while j delays flowering, providing new insight into soybean adaptation to tropical climates (Lu et al., 2017).
The discovery of novel allelic variation and investigation of its genetic and molecular basis has facilitated the successful conversion and adaption of tropical genepools to temperate climates (or temperate genepools to tropical climates) as noted in maize, rice, sorghum, and soybean. This enables exploration of new adaptation niches in agro-ecosystems where farmers are currently growing crops that may become unsuitable due to future climate change.
Rare Alleles to Benefit Future Genetic Improvement
While common alleles in crops are more likely to be involved in beneficial traits, useful variation may persist as rare alleles that have not undergone strong natural or human selection. The potential agronomic benefits of rare alleles are evident in many traits associated with domestication that are rare in natural populations. For instance, dwarf height, reduced tillering, non-shattering seeds and male sterility alleles will be rare in natural populations. In humans, the search for the ‘missing heritability’ of diseases has led to the understanding that rare alleles (at frequencies as low as <0.1%) can have major phenotypic effects (Fritsche et al., 2016). However, identifying rare alleles in wild or crop populations is challenging because extremely large populations need to be phenotyped and genotyped to detect rare alleles. There is also considerable ascertainment bias against rare alleles because these alleles can be confounded with sequencing errors (Heslot et al., 2013) and routine filtering often excludes alleles with a frequency below 5% to facilitate detection of common variants linked to traits.
Despite the challenges involved in detecting trait-linked rare alleles to use as candidates for crop improvement, substantial progress has been made as the cost of sequencing has decreased and the power of association mapping has risen. A remarkable success was achieved in maize, where the discovery of beneficial rare alleles LcyE and crtRB1 using association mapping (Harjes et al., 2008; Yan et al., 2010) later allowed introgression of the high-provitamin A trait into cultivars consumed in developing countries where vitamin A deficiency in children is an important public health concern (Azmach et al., 2013). In rice, a rare allelic variant of an upstream promoter of OsglHAT1 was shown to enhance grain weight and yield (Song X.J. et al., 2015), and a rare allele of the grain length QTL, qGL3, increases grain length, filling, and weight (Zhang et al., 2012).
Harnessing Epigenetic Variation for Crop Improvement
Epigenetics is broadly defined as “the study of mitotically and/or meiotically heritable changes in gene function that cannot be explained by changes in DNA sequence” (Russo et al., 1996). In the context of nuclear genes, epialleles are epigenetically modified alleles whose function is altered as a result of the particular epigenetic modification(s) (Finnegan, 2002). The range of reversible molecular mechanisms that can generate epialleles includes DNA methylation and a range of possible modifications (methylation, acetylation, phosphorylation, ubiquitination) to histones that can change chromatin states (Pikaard and Scheid, 2014). Additional molecular mechanisms that can generate or alter epialleles involve structural proteins and enzymes involved in chromatin assembly and remodeling (Pikaard and Scheid, 2014).
While transposable elements are key drivers of genetic variation in crop genepools (Bennetzen and Wang, 2014), they are also major drivers of epigenetic variation (Seymour and Becker, 2017; Song and Cao, 2017). In addition, there is a strong interplay between genetic variation (e.g., SNPs) and epiallelic variation (as measured by DNA methylation variation) (Eichten et al., 2013; Meng et al., 2016). Transposon-associated epigenetic variation has been shown to have functional effects in a range of crops, including rice (Zhang X. et al., 2015), melon (Citrullus lanatus) (Martin et al., 2009) and oil palm (Elaeis guineensis) (Ong-Abdullah et al., 2015).
Unlike mutational changes, epigenetic changes are potentially reversible. For instance, research on paramutation (since Brink’s pioneering studies of the R locus in maize) showed that at some loci, one allele can induce a heritable epigenetic change in the other allele (Brink, 1958; Hollick, 2016). Similar reversible epigenetic states are evident for loci regulated by genomic imprinting (Garnier et al., 2008), nucleolar dominance (Tucker et al., 2010) and gene silencing (Pikaard and Scheid, 2014). Indeed, epigenetic studies in plants (e.g., for the Lcyc and FWA loci) demonstrated that epialleles associated with biological functions can be heritable over multiple generations (Cubas et al., 1999; Soppe et al., 2000). Fundamental research using genetically identical yet epigenetically diverse recombinant inbred lines (epiRILs) has shown that epigenetic variation is associated with phenotypic variation and that such epiRILs can remain stable over generations (Johannes et al., 2009), at least in inbreeding plant species. Epigenetic quantitative trait loci (epiQTL) have been identified associated with traits such as root length and flowering time (Cortijo et al., 2014; Kooke et al., 2015). It has been also demonstrated that RNA interference (RNAi) pathways are important to maintain DNA methylation pattern fidelity over generations (Teixeira et al., 2009).
While it is clear that epialleles contribute to functional effects in plants, that can be trans-generationally inherited and be reversible (e.g., in response to abiotic or biotic environmental stimuli) (Becker et al., 2011; Schmitz et al., 2011), it is also emerging that epigenetic variation (epialleles) and mechanisms can potentially make contributions to functional traits in crop genepools (Eichten et al., 2011; Li Q. et al., 2014). Plant epigenome diversity research revealed that while different geographic origins display different genome-wide DNA methylation levels and epiallelic gene expression (Kawakatsu et al., 2016), there is no detectable signal of DNA methylome adaptation to the environment (Hagmann et al., 2015). In addition, the contribution of the DNA methylome to gene expression regulation has been demonstrated to be much less than the contribution from SNPs (Meng et al., 2016). While over a 1000 expression traits displayed significant SNP associations, less than 60 of these displayed an association with DNA methylation polymorphisms (Meng et al., 2016). Such findings have implications for crop improvement, particularly if epiallelic variation is contributing to specific adaptation of crops or their wild relatives to agro-environments.
Heterosis refers to the superior performance of F1 progeny compared to their parents and is extensively harnessed for crop improvement (Duvick, 2001). While heterosis in plants may be due to genetic dominance (complementation), overdominance and pseudo-overdominance effects (Birchler et al., 2010; McKeown et al., 2013; Schnable and Springer, 2013), there is emerging evidence that epigenetic variation and mechanisms may contribute to heterosis effects in plants (Shivaprasad et al., 2012; Groszmann et al., 2013; Offermann and Peterhansel, 2014). For instance, “heterosis without hybridization” has been demonstrated in plants using epiRILs and triploid lines that are genetically identical but epigenetically different (Duszynska et al., 2013; Dapp et al., 2015; Fort et al., 2016).
Metastable epialleles are alleles that display variable expressivity despite being in an identical genetic background. Research using metastable epialleles of the red color r1 locus in maize did not, however, support a metastable epigenetic contribution to heterosis or inbreeding depression (Auger et al., 2004). While non-additive DNA methylation effects have been observed in F1 hybrids that display heterosis (Greaves et al., 2014), the functional significance of such DNA methylation changes is unclear, as genetic ablation of such RNA-mediated DNA methylation interactions did not affect heterosis biomass, while the chromatin remodeller DDM1 has been identified as a modifier of heterosis (Groszmann et al., 2011; Shen et al., 2012; Kawanabe et al., 2016; Zhang et al., 2016a,b).
Apomixis refers to asexual reproduction via plant seeds. Despite being a naturally occurring phenomenon, the fixation of heterosis via apomixis to generate true-breeding lines in crop improvement programs has been extensively proposed but has not been realized to date (Spillane et al., 2004). Indeed, while it was expected that apomixis could fix F1 heterosis effects that have a genetic basis, this had not been demonstrated until recently where it has been shown that apomixis could fix 90% of traits generated in Pilosella F1 hybrids over two successive generations (Sailer et al., 2016).
Research in tomato (Solanum lycopersicum) and emerging studies on other fleshy fruit crops are revealing a role for epigenetic control of fruit ripening (Gallusci et al., 2016), e.g., DEMETER-like DNA demethylases and CHROMOMETHYLASE3 (SICMT3) genes regulate fruit-ripening associated transcription factors (e.g., RIN) and epi-alleles (Cnr) in tomato (Manning et al., 2006; Zhong et al., 2013; Chen et al., 2015; Liu R. et al., 2015). Epigenetic variation in crop genepools at loci that are sensitive to environmental signals (e.g., temperature, vernalization, photoperiod, flowering time, and ripening) has potential for more effective harnessing through breeding (King, 2015), involving selection of functional epiallelic variants, and potentially future epi-genome editing (Klann et al., 2017).
Epigenetic variation that contributes to adaptive phenotypic variation may be particularly important in fluctuating environments, and could play an important phenotypic plasticity role as a buffer to environmental stimuli, and both abiotic and biotic stresses. For instance, tight epigenetic regulation of the antagonistic NLR receptors PigmR and PigMS is necessary to confer rice blast resistance with minimal yield penalty (Deng et al., 2017). The identification of such epialleles and epigenetic regulatory systems conferring functional impacts on agronomic traits can feed into a range of different approaches emerging for epigenetic breeding of crop plants, including use of mutant lines (Yang et al., 2015), recurrent epi-selection (Hauben et al., 2009; Greaves et al., 2015), hybrid mimics (Wang L. et al., 2015), epigenomic selection (Jonas and de Koning, 2013; Oakey et al., 2016) and epigenome editing (Park et al., 2016).
Many studies on the relationship between epigenetic variation and epialleles and phenotypic variation (including expression variation) have been conducted under controlled conditions. As a result, little is known of the extent of epigenotype × genotype × environment (epiG × G × E) interactions of crops under field conditions. For crop improvement, harnessing both multilocus epiallele interactions (with associated epistasis effects) and single locus “major effect” epialleles offer opportunities for developing novel approaches for increased epigenetic gain in crop breeding programs.
Harnessing Functional Diversity with New Trait Improvement Technologies
Genomic-Estimated Breeding Values to Predict the Utility of Germplasm Accessions
While there are 100s or 1000s of germplasm accessions conserved in ex situ genebanks globally, the lack of phenotyping and genotyping data limits their use. Advances in genomics, phenomics and bioinformatics are increasing the availability and quality of data to better leverage this germplasm for breeding (McCouch, 2013). Association genetics along with genomic prediction further allows expansion of use of genetic variation, with the aim of increasing yield-related genetic gains in cereals (Spindel et al., 2016). Ayling (2016) provides an overview of promising methods for increasing the knowledge on (and utility of) genebank accessions using next generation sequencing (NGS). Moreover, emerging cross-disciplinary “genoplasmics” has been proposed as new term to refer to genomics-assisted plant germplasm research (Jia et al., 2017). Such a research methodology involves defining core collections or core subsets (that capture maximum evolutionary history in a limited number of accessions) are promoted for genetic enhancement or gene discovery (van Hintum et al., 2000). For example, investigation of spring bread wheat diversity (in a genebank from mega environments) by high quality genotyping-by-sequencing (GBS) loci and gene-based markers permitted selection of novel variation for further use in breeding crops with traits such as adaptation to drought or heat stress (Sehgal et al., 2015b).
Longin and Reif (2014) proposed a stepwise strategy for better use of wheat genetic resources that are available in genebanks. They propose using representative core subsets of accessions that are defined after genotyping and assessing their genetic relationships, including consideration of whether different accessions harbor major adaptation genes to stressful environments. Specific accessions are selected according to phenotyping and genome-wide data; i.e., genotypic and phenotypic data are used to estimate effects for all genomic regions and to develop models for predicting genomic estimated breeding values (GEBV) of genebank accessions that may be candidate parents of elite wheat breeding lines. This approach targets the entire genome rather than focusing on major genes with large effects related to traits of interest. Genomic prediction models including the genotype × environment interaction have already been shown to be promising for introgressing highly heritable traits from exotic wheat landrace germplasm stored in genebanks into elite breeding lines (Crossa et al., 2016). These results validate the direct use in crop breeding of the substantial landrace genetic diversity that is conserved in genebanks. This introgression breeding approach also requires GEBV to predict the value of the resulting offspring. In the last step of the proposed strategy, genotypic and phenotypic data along with passport and pedigree information are shared through a database platform to facilitate breeding.
As noted by Brown (2016), the major interest of using GEBV for predicting traits lies on replacing expensive phenotyping with inexpensive genotyping. In this regard, Yu et al. (2016) provided a proof-of-concept study that integrated genomic prediction into the evaluation of germplasm with a broad genetic base. They first characterized a sorghum core subset (962 accessions) with GBS. Next, 299 accessions representing the overall diversity of the core subset were selected as a training set for biomass yield and other related phenotypic traits such as plant height, stalk number and root lodging, amongst others. Cross-validation demonstrated a high prediction accuracy for stalk number and biomass yield. Similarly, Gorjanc et al. (2016) used GEBV to harness multigenic variation from maize landraces. Their results suggest that genetic enhancement using high levels of genetic diversity can begin directly with landraces. They also indicated that early introgression into elite germplasm seems to be feasible for loci with large effects, but not for landrace haplotypes harboring multi-genic variation because further improvement will favor the elite haplotypes and limit the distinctness of resulting germplasm. Similarly, Burstin et al. (2015) were able to predict flowering, seeds per plant, and seed weight using diverse pea accessions after characterizing the accessions with DNA markers. Genomic prediction, as shown by Jarquin et al. (2016), depends on having both the target population and environment in the training set, and on including data from diverse geographical locations and genetic clusters. Their research highlights the value of historical germplasm data to develop predictive models that assist in selecting genebank accessions for introgressing useful genetic variation into breeding populations and programs.
CRISPR/Cas9 to Release Novel Variation
Genome editing is a rapidly emerging targeted mutagenesis approach that offers unique opportunities to elucidate gene functions and introduce novel beneficial alleles into crop germplasm (Voytas, 2013; Scheben and Edwards, 2017). Although engineered nucleases such as transcription activator-like effector nucleases (TALENs) and zinc finger nucleases (ZFNs) can be used for genome editing, the dominant tool at present is the type II clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein (Cas) system using the Cas9 nuclease (Jinek et al., 2012). The CRISPR/Cas system targets specific genomic regions via a guide RNA which hybridizes to a G(N)19-22NGG target DNA sequence downstream of an NGG protospacer adjacent motif (Gasiunas et al., 2012; Jinek et al., 2012). The guide RNA forms a complex with the Cas9 nuclease which then cleaves double stranded DNA at the target site. The error-prone non-homologous end joining (NHEJ) DNA repair pathway repairs the double-strand break, typically introducing a deletion. By delivering a DNA repair template together with the CRISPR/Cas system, in principle precise insertions or deletions can be achieved via the error-free homology directed repair (HDR) pathway.
CRISPR/Cas has now been demonstrated to improve agronomic traits in numerous crops. For example, genome edited lines of the rice genes gn1a, dep1, gs3, dep1 and gs3 showed enhanced grain number, dense erect panicle, larger grain size, semi-dwarf stature and long-awned grains, respectively (Li M. et al., 2016). Disrupting pest-susceptibility genes increased resistance to fungal blast in rice (Wang F. et al., 2016), resistance to powdery mildew in bread wheat (Wang Y. et al., 2014) and broad-spectrum disease resistance in tomato (de Toledo Thomazella et al., unpublished). The CRISPR/Cas system has also unraveled several biallelic mutations of Glyma06g14180 and Glmya08g02290 with varying gene expression during hairy root development in soybean (Sun et al., 2015). While these studies have relied on disruption of target genes via error-prone NHEJ to change plant phenotypes, progress has also been made in the more difficult to achieve precise gene targeting using HDR. In a recent study in maize, a promoter of the drought-tolerance associated ARGOS8 gene was swapped using CRISPR/Cas and HDR with the U3 maize promoter to increase expression of the ARGOS8 gene (Shi et al., 2016). Field trials showed that genome edited maize plants had significantly higher yield under drought stress and no yield loss under normal conditions.
The major current limitations in the application of CRISPR/Cas genome editing for crop improvement relate to the inefficient precise gene targeting via HDR (Steinert et al., 2016). However, targeted base editing has now been demonstrated in rice, wheat, maize and tomato using Cas9-cytidine deaminase fusions (Zong et al., 2017; Shimatani et al., 2017). In addition, given that most cereal and legume crops are auto- or allo-polyploids, genome editing now offers the exciting opportunity for targeted mutagenesis in polyploid plant genomes (Ryder et al., 2017), which has not been possible with conventional mutagenesis techniques (Kumar et al., 2017). While there has been some focus on potential lack of target site specificity (Fu et al., 2013; Jiang W.Z. et al., 2013; Pattanayak et al., 2013) from early generation CRISPR/Cas systems, improved systems are under development to minimize any off-target edits. For instance, innovative techniques such as paired Cas9 nickases (Ran et al., 2013) and highly specific Cas9 variants (Kleinstiver et al., 2016) are increasing the DNA target specificity of genome editing. In any event, weighing up the benefits of gene targeting against any (hypothetical) costs associated with off-target editing in crop genomes, very low levels of off-target editing in crop genomes is unlikely to be of any major concern, given that (chemical and radiation) mutagenized lines have been used for decades for crop improvement. Genome editing has major potential not only for crop improvement but also for rapid domestication of novel crops from wild species or minor crops by simultaneously editing genes related to domestication such as grain size, shattering, plant stature, and flowering time. As wild plants may harbor greater diversity in climate-related traits such as stress tolerance and pest resistance, CRISPR/Cas-assisted breeding approach may play an important role in increasing global food production in a changing climate.
Storage and Integration of Genetic and Phenotypic Information
Although vast amounts of sequence and expression data are hosted by the European Molecular Biology Laboratory (EMBL) (Kanz et al., 2005), GenBank (Benson et al., 2012), and the DNA Data Bank of Japan (DDBJ) (Mashima et al., 2016), crop improvement relies on the integration of such data with more widely dispersed data on sequence variation and phenotypes. Storing and managing the increasing amounts of public and private genotypic and phenotypic data on crops, however, is challenging (Batley and Edwards, 2009; Lee et al., 2012). In the past decade or so, crop-specific databases and bioinformatics services have been developed for many crops (Table 7). These public databases provide access to genomes and the corresponding annotation data, together with data on phenotypes and genotypes. Tools such as ngs.plot (Shen et al., 2014) and QTLNetMiner2 enable integrated analysis of genotype and phenotype information contained in databases. Mining of genomic databases in this way or using more advanced machine learning approaches can facilitate discovery of genes related to specific target functions. A range of databases for the management of crop germplasm have also been developed by the European Cooperative Programme for Plant Genetic Resources Networks (ECPGR) and the United States. National Plant Germplasm System (NPGS). Linking germplasm information to broader genetic and phenotypic resources would allow easier accessibility of germplasm for experimentation.
The large scale of genotypic and phenotypic data requires a powerful computational platform for data management and parallel processing. The open-source Apache Hadoop framework suits the demands of large-scale processing of genomic data (Niemenmaa et al., 2012; Nordberg et al., 2013; O’Driscoll et al., 2013). Cloud computing services such as those provided by Amazon (Madduri et al., 2014), or institutional dedicated computing clusters, allow researchers cost-effective access to the computational power required for integrated analysis of large biological datasets. While the infrastructure for developing databases to host and help analyze big biological data is available, long-term funding of database projects is rare, despite being essential infrastructure to maintain and curate the continuously growing databases. Moreover, the usefulness of existing databases is often severely limited by a lack of phenotype data because the generation of genotypic data has fast outstripped that of other data types. Without phenotype data, it is not possible to carry out association genetics or identifying candidate genes linked to agronomic traits (Cobb et al., 2013). Advances in phenomics such as remote sensing, robotics and automated environmental data collection may help overcome the bottleneck in phenotyping data (Furbank and Tester, 2011; Araus and Cairns, 2014).
Another step required to facilitate the integration of large-scale crop data is the use of shared vocabularies for genetic and phenotypic information. The gene ontology (GO) project has made major strides in the use of universal vocabularies for the annotation of genes, gene products and sequences. Nevertheless, the vocabularies used for describing sequence variation and phenotypic traits remain inconsistent and hamper data integration. Researchers must collaborate closely in developing consistent vocabularies to accelerate the development of broadly informative crop databases that are useful for plant breeders.
Finally, an important step for crop improvement would be an increase in data sharing between public and private sector institutions (Spindel and McCouch, 2016). Political disagreements on access to crop genetic resources and the distribution of benefits have led to protectionist attitudes regarding plant genetic resources, indicating the need for an acceptable data sharing framework. The 2001 International Treaty on Plant Genetic Resources for Food and Agriculture (IT PGRFA) provides a multilateral framework (agreed between the world’s governments) for exchange of plant genetic resources (using a common material transfer agreement (MTA) for each accession) between countries (and their institutions). The IT PGRFA covers access to the vast majority of the world major crop and forage species, which are listed in Annex 1 of the Treaty. Future challenges will undoubtedly emerge regarding access and benefit sharing relating to genomic or phenomic data derived from crop genetic resources, which fall under the auspices of the International Treaty on Plant Genetic Resources.
Current Knowledge and Future Challenges in Functional Diversity
Advances in genomic technologies have led to an unprecedented availability of crop sequences and sequence variation data. As crop genomes are re-sequenced to better represent the genetic diversity in the gene pool, pangenomes capturing core and variable genes in crop species are becoming available, e.g., in maize (Hirsch et al., 2014), rice (Schatz et al., 2014), wheat (Montenegro et al., 2017), soybean (Li Y.H. et al., 2014), Brassica rapa (Lin et al., 2014), and Brassica oleracea (Golicz et al., 2016). Despite this wealth of genomic data, gene functions and networks remain very poorly characterized, even in crops such as rice (Rhee and Mutwil, 2014). The genetic mechanisms controlling important agronomic traits are only slowly being elucidated, revealing complex interaction networks and considerable diversity between crops, as in the case of the universal florigen flowering pathway (Turck et al., 2008). The genes underlying other complex traits such as abiotic stress adaptation are still only partly known and often confounded by gene by environment interactions (Fleury et al., 2010). Association mapping of these complex traits, combined with reverse genetic screening to elucidate trait-gene associations, will be crucial to uncover both common and rare alleles of yield-related traits.
While trait-gene association is currently hampered by a lack of extensive phenotype data from well characterized environments, recent advances in high-throughput phenotyping platforms that can be used in the field may soon help overcome this challenge (Furbank and Tester, 2011; Araus and Cairns, 2014). By utilizing the diverse germplasm resources available from crops and their wild relatives to uncover genes that can be introgressed into elite breeding germplasm, crops can continue to be improved for potential yield and yield stability. An important step to accelerate such breeding efforts worldwide is the integration of this information in openly accessible databases, which is currently lagging behind the rapid generation of data. As climate change and a growing population put increasing pressure on plant breeders in the public and private sectors to produce high-yielding, climate resilient cultivars, a consensus needs to also be reached on the merits of genome editing to produce novel and useful diversity in crop germplasm, to rapidly improve agronomic traits associated with known genes. It is increasingly clear that crop improvement must draw on diverse germplasm pools and leverage advances in biotechnology to ensure future global food security.
Conclusion
Plant genetic resources provide raw materials for mining allelic variations associated with target traits (Figure 1). Crop improvement continues to rely on combining diversity in crop populations and their wild relatives via genetic recombination. Sequencing technology advances and bioinformatic tools used for assessing diversity in germplasm panels have identified millions of polymorphic SNPs in cereals and legumes, as noted by the examples included in this article. Unlocking functional diversity for key agronomic traits such as crop phenology, plant architecture, yield and stress tolerance is facilitating greater use of germplasm in crop breeding. Major QTL and candidate SNPs associated with such agronomi traits, identified through genome wide association research (and ideally confirmed by functional studies), have been deployed in crop breeding to enhance adaptation and productivity of staple food crops. The discovery and deployment of alleles associated with variation in response to photoperiod and flowering has allowed the cultivation of tropical crops such as maize, rice and sorghum in temperate climates (CTB4a, Ghd7, PRR37, RFT1, Sb06g012260) or temperate crops such as soybean in tropical regions (J locus).
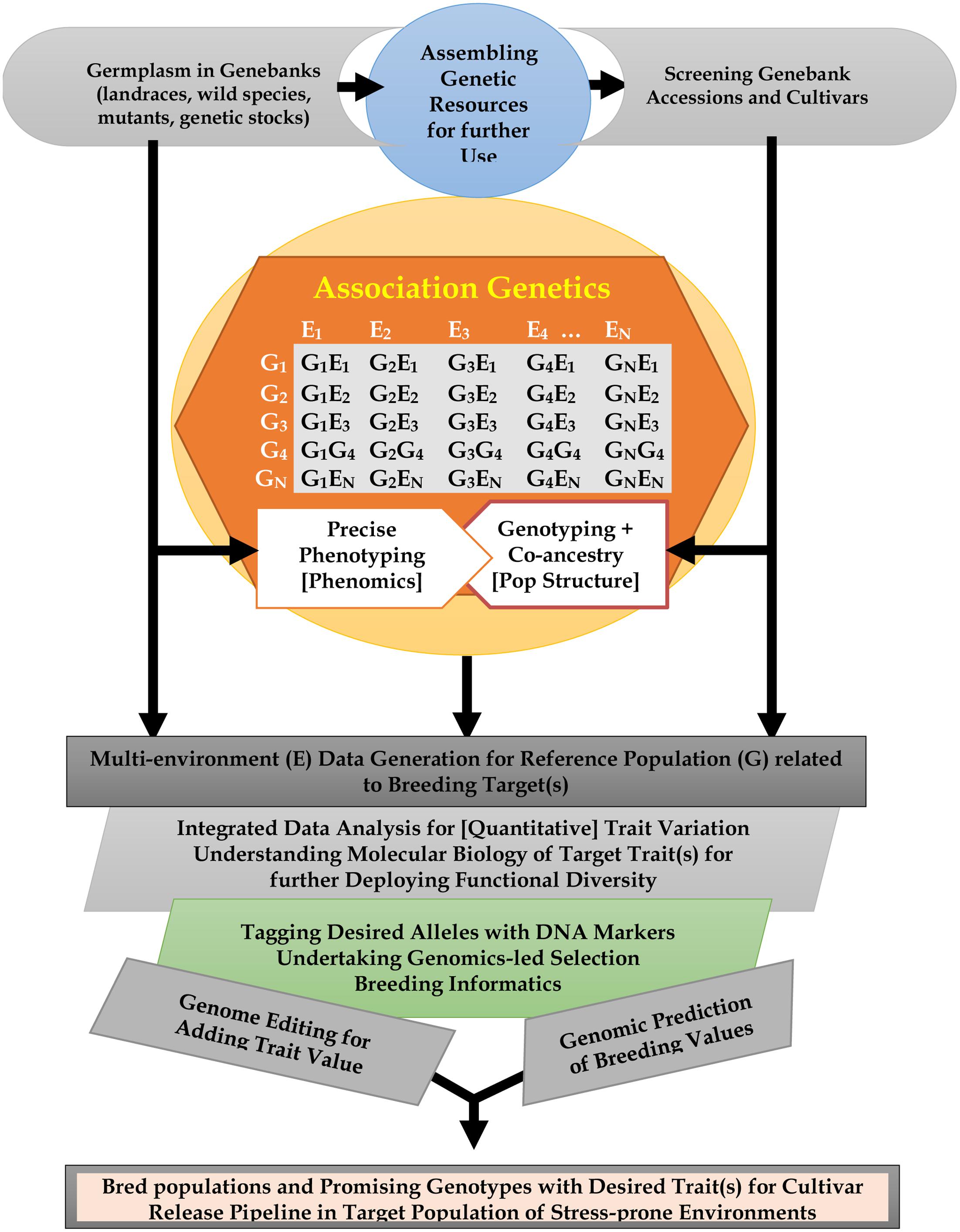
FIGURE 1. Assessing and exploiting functional diversity in germplasm pools in the omics era for plant breeding under a changing climate.
Flowering time is a major determinant of crop yield, and selection of variants in florigen pathway and their deployment in crop breeding has increased yield in several crops. In particular. fine-tuning of the florigen pathway has allowed yield increases through better control of flowering and growth in crops. Grain legumes have indeterminate flowering, leading to low productivity compared to cereals. The discovery of candidate genes, Phvul.001G189200 in common bean and CcTFL1 in pigeonpea associated with determinate flowering may allow manipulation of determinacy trait in these and other legumes.
Functional allelic diversity has been successfully harnessed through breeding to enhance abiotic stress (drought, salinity, low soil P, and submergence tolerance) adaptation and productivity by manipulating panicle architecture in rice. Likewise, QTL hotspots associated with drought adaptation in chickpea have been introgressed in several leading cultivars in Asia and Africa. Glycine soja is an interesting source of variation for abiotic stress adaptation in soybean. Candidate SNPs associated with drought adaptation are known in common bean. Changes in functional diversity due to global warming were noted for flowering in wild barley and emmer wheat from Israel or among pearl millet landraces from Africa, thus providing valuable resource for enhancing crop adaptation to variable climates leading to shortening of growing season. Genomics have unraveled SNPs associated with precipitation and length of growing season in barley or with precipitation and high temperature in maize and sorghum, with many located in genes known for abiotic stress adaptation, thus providing valuable resource to accelerate breeding for drought-prone environments.
Maize has comprehensive haplotype maps that has enabled researchers to identify selective sweeps and chromosome regions harboring loci related to domestication and geographic adaptation. Identification of distinct haplotypes in subtropical maize germplasm provided opportunity to exploit heterotic potential among them. Likewise, distinct haplotypes discovered amongst Indian wild rice accessions associated with high salinity tolerance from distant geographic regions may be useful for broadening the available cultigen pool to enhance rice productivity in salt-prone areas.
Landraces and wild relatives are proven genetic resource to identify genetic variants associated with environmental adaptation, particularly temperature and precipitation. In addition, such genetic resources are also the source of discovering rare alleles; however, identifying such alleles is challenging because of their low presence in populations, necessitating phenotyping and genotyping of extremely large populations. Evidence suggests that epigenetic variation (i.e., epialleles) can also be successfully exploited to enhance abiotic stress adaptation and productivity in crops. Genome editing in maize, rice and soybean or estimating genomic-estimated breeding values of genebank accessions in maize, pea, sorghum, and wheat provide means to access and generate additional variability for agronomic and stress tolerance traits.
Phenomics and genomics are enabling generation of vast data sets in crop breeding. However, archival and easy retrieval of these data set is a challenge. Crop-specific databases along with bioinformatics services provide access to genomes and the corresponding annotation data, together with data on phenotypes and genotypes for many crops. An integrated analysis of genotype and phenotype information contained in databases facilitates the discovery of genes related to specific target functions. Such insights on crop biodiversity and trait inheritance along with mapping of genetic variation controlling key traits, and using them for developing breeding germplasm will accelerate crop improvement, increase genetic gains and allowing improved crop yields and yield stability under a changing climate and in stress-prone environments.
Author Contributions
All authors participated in outlining the manuscript contents, searching the literature, writing and editing the text.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
SLD acknowledges the contribution of Ramesh Kotnana of Knowledge Sharing and Innovation Program of ICRISAT for arranging reprints on functional diversity as valuable literature resources for drafting this manuscript. CS acknowledges funding support from Science Foundation Ireland, and the Irish Department of Food, Agriculture and the Marine, while grants provided during the writing of this manuscript from PlantePigment and Annato –projects led by Chr. Hansen A/S with grants from Green Development and Demonstration Program, (GUDP, Denmark) and Innovationsfonden (Denmark), respectively– and Utveckling av ett molekylärt förädlingsprogram för durum-vete i Senegal: Kapacitetsuppbyggnad för att möta den globala uppvärmningen –a project with funding from Vetenskapsrådet (VR, Sweden) Development Research, are acknowledged by RO. The sketch used for drawing Figure 1 was kindly shared by Dr. Sandeep Kumar Kushwaha (SLU, VF, Alnarp. Sweden). We are grateful to the three reviewers for helpful suggestions on improving the manuscript.
Footnotes
References
Abberton, M., Batley, J., Bentley, A., Bryant, J., Cai, H., Cockram, J., et al. (2016). Global agricultural intensification during climate change: a role for genomics. Plant Biotechnol. J. 14, 1095–1098. doi: 10.1111/pbi.12467
Abebe, T. D., Naz, A., and Léon, J. (2015). Landscape genomics reveal signatures of local adaptation in barley (Hordeum vulgare L.). Front. Plant Sci. 6:813. doi: 10.3389/fpls.2015.00813
Akond, M., Liu, S., Schoener, L., Anderson, J. A., Kantartzi, S. K., Meksem, K., et al. (2013). A SNP-based genetic linkage map of soybean using the SoySNP6K Illumina Infinium BeadChip genotyping array. J. Plant Genome Sci. 1, 80–89. doi: 10.5147/jpgs.2013.0090
Anderson, J. E., Kono, T. J. Y., Stupar, R. M., Kanta, M. B., and Morrell, P. L. (2016). Environmental association analyses identify candidates for abiotic stress tolerance in Glycine soja, the wild progenitor of cultivated soybean. G3 6, 835–843. doi: 10.1534/g3.116.026914
Araus, J. L., and Cairns, J. E. (2014). Field high-throughput phenotyping: the new crop breeding frontier. Trends Plant Sci. 19, 52–61. doi: 10.1016/j.tplants.2013.09.008
Ariani, A., Teran, J. C. B. M., and Gepts, P. (2016). Genome-wide identification of SNPs and copy number variation in common bean (Phaseolus vulgaris L.) using genotyping-by-sequencing. Mol. Breed. 36:87. doi: 10.1007/s11032-016-0512-9
Ashikari, M., Sakakibara, H., Lin, S., Yamamoto, T., Takashi, T., Nishimura, A., et al. (2005). Cytokinin oxidase regulates rice grain production. Science 309, 533–538. doi: 10.1126/science.1113373
Auger, D., Ream, T., and Birchler, J. (2004). A test for a metastable epigenetic component of heterosis using haploid induction in maize. Theor. Appl. Genet. 108, 1017–1023. doi: 10.1007/s00122-003-1521-8
Ayling, S. C. (2016). Technical Appraisal of Strategic Approaches to Large-Scale Germplasm Evaluation. Norwich: The Genome Analysis Center.
Azmach, G., Gedil, M., Menkir, A., and Spillane, C. (2013). Marker-trait association analysis of functional gene markers for provitamin A levels across diverse tropical yellow maize inbred lines. BMC Plant Biol. 13:227. doi: 10.1186/1471-2229-13-227
Bailey-Serres, J., Fukao, T., Ronald, P., Ismail, A., Heuer, S., and Mackill, D. (2010). Submergence tolerant rice: SUB1’s journey from landrace to modern cultivar. Rice 3, 138–147. doi: 10.1007/s12284-010-9048-5
Bajaj, D., Das, S., Badoni, S., Kumar, V., Singh, M., Bansal, K. C., et al. (2015). Genome-wide high throughput SNP discovery and genotyping for understanding natural (functional) allelic diversity and domestication patterns in wild chickpea. Sci. Rep. 5:12468. doi: 10.1038/srep12468
Batley, J., and Edwards, D. (2009). Genome sequence data: management, storage, and visualization. Biotechniques 46, 333–334. doi: 10.2144/000113134
Becker, C., Hagmann, J., Müller, J., Koenig, D., Stegle, O., Borgwardt, K., et al. (2011). Spontaneous epigenetic variation in the Arabidopsis thaliana methylome. Nature 480, 245–249. doi: 10.1038/nature10555
Bedada, G., Westerbergh, A., Nevo, E., Korol, A., and Schmid, K. J. (2014). DNA sequence variation of wild barley Hordeum spontaneum (L.) across environmental gradients in Israel. Heredity 112, 646–655. doi: 10.1038/hdy.2014.2
Bennetzen, J. L., and Wang, H. (2014). The contributions of transposable elements to the structure, function, and evolution of plant genomes. Annu. Rev. Plant Biol. 65, 505–530. doi: 10.1146/annurev-arplant-050213-035811
Benson, D. A., Karsch-Mizrachi, I., Clark, K., Lipman, D. J., Ostell, J., and Sayers, E. W. (2012). GenBank. Nucleic Acids Res. 40, D48–D53. doi: 10.1093/nar/gkr1202
Birchler, J. A., Yao, H., Chudalayandi, S., Vaiman, D., and Veitia, R. A. (2010). Heterosis. Plant Cell 22, 2105–2112. doi: 10.1105/tpc.110.076133
Biscarini, F., Cozzi, P., Casella, L., Riccardi, P., Vattari, A., Orasen, G., et al. (2016). Genome-wide association study for traits related to plant and grain morphology, and root architecture in temperate rice accessions. PLoS ONE 11:e0155425. doi: 10.1371/journal.pone.0155425
Blake, V. C., Birkett, C., Matthews, D. E., Hane, D. L., Bradbury, P., and Jannink, J. L. (2016). The triticeae toolbox: combining phenotype and genotype data to advance small-grains breeding. Plant Genome 9. doi: 10.3835/plantgenome2014.12.0099
Blomstedt, C. K., Gleadow, R. M., O’Donnell, N., Naur, P., Jensen, K., Laursen, T., et al. (2012). A combined biochemical screen and TILLING approach identifies mutations in Sorghum bicolor L. Moench resulting in acyanogenic forage production. Plant Biotechnol. J. 10, 54–66. doi: 10.1111/j.1467-7652.2011.00646.x
Bouchet, S., Bertin, P., Presterl, T., Jamin, P., Coubriche, D., Gouesnard, B., et al. (2017). Association mapping for phenology and plant architecture in maize shows higher power for developmental traits compared with growth influenced traits. Heredity 118, 249–259. doi: 10.1038/hdy.2016.88
Boutet, G., Carvalho, S. A., Falque, M., Peterlongo, P., Lhuillier, E., Bouchez, O., et al. (2016). SNP discovery and genetic mapping using genotyping by sequencing of whole genome genomic DNA from a pea RIL population. BMC Genomics 17:121. doi: 10.1186/s12864-016-2447-2
Boyles, R. E., Cooper, E. A., Myers, M. T., Brenton, Z., Rauh, B. L., Morris, G. P., et al. (2016). Genome-wide association studies of grain yield components in diverse sorghum germplasm. Plant Genome 9. doi: 10.3835/plantgenome2015.09.0091
Bray, E. A., Bailey-Serres, J., and Weretilnyk, E. (2000). Responses to Abiotic Stresses. Rockville, MD: American Society of Plant Physiologists.
Brink, R. A. (1958). Paramutation at the R locus in maize. Cold Spring Harbor. Symp. Quant. Biol. 23, 379–391. doi: 10.1101/SQB.1958.023.01.036
Brown, P. J. (2016). Effective use of genetic diversity. Nature Plants 2:16154. doi: 10.1038/NPLANTS.2016.154
Burstin, J., Salloignon, P., Chabert-Martinello, M., Magnin-Robert, J.-B., Siol, M., Jacquin, F., et al. (2015). Genetic diversity and trait genomic prediction in a pea diversity panel. BMC Genomics 16:105. doi: 10.1186/s12864-015-1266-1
Caldwell, K. S., Russell, J., Langridge, P., and Powell, W. (2006). Extreme population dependent linkage disequilibrium detected in an inbreeding plant species. Genetics 172, 557–567. doi: 10.1534/genetics.104.038489
Casas, A. M., Djemel, A., Ciudad, F. J., Yahiaoui, S., Ponce, L. J., Contreras-Moreira, B., et al. (2011). HvFT1 (VrnH3) drives latitudinal adaptation in Spanish barleys. Theor. Appl. Genet. 122, 1293–1304. doi: 10.1007/s00122-011-1531-x
Cavanagh, C. R., Chao, S., Wang, S., Huang, B. E., Stephen, S., Kiani, S., et al. (2013). Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploidy wheat landraces and cultivars. Proc. Natl. Acad. Sci. U.S.A. 110, 8057–8062. doi: 10.1073/pnas.1217133110
Chen, H., Xie, W., He, H., Yu, H., Chen, W., Li, J., et al. (2014). A high density SNP genotyping array for rice biology and molecular breeding. Mol. Plant 7, 541–553. doi: 10.1093/mp/sst135
Chen, J., Chopra, R., Hayes, C., Morris, G., Marla, S., Burke, J., et al. (2017). Genome-wide association study of developing leaves’ heat tolerance during vegetative growth stages in a sorghum association panel. Plant Genome 10. doi: 10.3835/plantgenome2016.09.0091
Chen, L., Huang, L. Z., Min, D. H., Phillips, A., Wang, S. Q., Madgwick, P. J., et al. (2012). Development and characterization of a new TILLING population of common bread wheat (Triticum aestivum L.). PLoS ONE 7:e41570. doi: 10.1371/journal.pone.0041570
Chen, W., Kong, J., Qin, C., Yu, S., Tan, J., Chen, Y.-R., et al. (2015). Requirement of CHROMOMETHYLASE3 for somatic inheritance of the spontaneous tomato epimutation colourless non-ripening. Sci. Rep. 5:9192. doi: 10.1038/srep09192
Chin, J. H., Gamuyao, R., Dalid, C., Bustamam, M., Prasetiyono, J., Moeljopawiro, S., et al. (2011). Developing rice with high yield under phosphorus deficiency: Pup1 sequence to application. Plant Physiol. 156, 1202–1216. doi: 10.1104/pp.111.175471
Chung, W. H., Jeong, N., Kim, J., Lee, W. K., Lee, Y. G., Lee, S. H., et al. (2014). Population structure and domestication revealed by high-depth resequencing of Korean cultivated and wild soybean genomes. DNA Res. 21, 153–167. doi: 10.1093/dnares/dst047
Clevenger, J., Chu, Y., Chavarro, C., Agrawal, G., Bertioli, D. J., Leal-Bertioli, S. C. M., et al. (2017). Genome-wide SNP genotyping resolves signatures of selection and tetrasomic recombination in peanut. Mol. Plant 10, 309–322. doi: 10.1016/j.molp.2016.11.015
Close, T. J. (2015). Development of a 60k iSelect to Facilitate Cowpea Breeding. Riverside, CA: University of California - Riverside.
Cobb, J. N., DeClerck, G., Greenberg, A., Clark, R., and McCouch, S. (2013). Next-generation phenotyping: requirements and strategies for enhancing our understanding of genotype-phenotype relationships and its relevance to crop improvement. Theor. Appl. Genet. 126, 867–887. doi: 10.1007/s00122-013-2066-0
Cockram, J., Jones, H., Leigh, F. J., O’Sullivan, D., Powell, W., Laurie, D. A., et al. (2007). Control of flowering time in temperate cereals: genes, domestication, and sustainable productivity. J. Exp. Bot. 58, 1231–1244. doi: 10.1093/jxb/erm042
Comadran, J., Kilian, B., Russell, J., Ramsay, L., Stein, N., Ganal, M., et al. (2012). Natural variation in a homolog of Antirrhinum CENTRORADIALIS contributed to spring growth habit and environmental adaptation in cultivated barley. Nat. Genet. 44, 1388–1392. doi: 10.1038/ng.2447
Cortijo, S., Wardenaar, R., Colomé-Tatché, M., Gilly, A., Etcheverry, M., Labadie, K., et al. (2014). Mapping the epigenetic basis of complex traits. Science 343, 1145–1148. doi: 10.1126/science.1248127
Crossa, J., Jarquín, D., Franco, J., Pérez-Rodríguez, P., Burgueño, J., Saint-Pierre, C., et al. (2016). Genomic prediction of gene bank wheat landraces. G3 6, 1819–1834. doi: 10.1534/g3.116.029637
Crowell, S., Korniliev, P., Falcão, A., Ismail, A., Gregorio, G., and Mezey, J. (2016). Genome-wide association and high-resolution phenotyping link Oryza sativa panicle traits to numerous trait-specific QTL clusters. Nat. Commun. 7:10527. doi: 10.1038/ncomms10527
Cubas, P., Vincent, C., and Coen, E. (1999). An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401, 157–161. doi: 10.1038/43657
Cuevas, H. E., Zhou, C., Tang, H., Khadke, P. P., Das, S., Lin, Y. R., et al. (2016). The evolution of photoperiod insensitive flowering in sorghum, a genomic model for panicoid grasses. Mol. Biol. Evol. 33, 2417–2428. doi: 10.1093/molbev/msw120
Dapp, M., Reinders, J., Bédiée, A., Balsera, C., Bucher, E., Theiler, G., et al. (2015). Heterosis and inbreeding depression of epigenetic Arabidopsis hybrids. Nat. Plants 1:5092. doi: 10.1038/nplants.2015.92
Dar, M. H., de Janvry, A., Emerick, K., Raitzer, D., and Sadoulet, E. (2013). Flood tolerant rice reduces yield variability and raises expected yield, differentially benefitting socially disadvantaged groups. Sci. Rep. 3:3315. doi: 10.1038/srep03315
Deng, Y., Zhai, K., Xie, Z., Yang, D., Zhu, X., Liu, J., et al. (2017). Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science 355, 962–965. doi: 10.1126/science.aai8898
Deryng, D., Conway, D., Ramankutty, N., Price, J., and Warren, R. (2014). Global crop yield response to extreme heat stress under multiple climate change futures. Environ. Res. Lett. 9:034011. doi: 10.1088/1748-9326/9/3/034011
Dhanapal, A. P., Ray, J. D., Singh, S. K., Hoyos-Villegas, V., Smith, J. R., Purcell, L. C., et al. (2015). Genome-wide association study (GWAS) of carbon isotope ratio (δ13C) in diverse soybean [Glycine max (L.) Merr.] genotypes. Theor. Appl. Gent. 128, 73–91. doi: 10.1007/s00122-014-2419-9
Dhanapal, A. P., Ray, J. D., Singh, S. K., Hoyos-Villegas, V., Smith, J. R., Purcell, L. C., et al. (2016). Genome-wide association mapping of soybean chlorophyll traits based on canopy spectral reflectance and leaf extracts. BMC Plant Biol. 16:174. doi: 10.1186/s12870-016-0861-x
di Fenza, M., Hogg, B., Grant, J., and Barth, S. (2017). Transcriptomic response of maize primary roots to low temperatures at seedling emergence. Peer J. 5, e2839. doi: 10.7717/peerj.2839
Doddamani, D., Khan, A. W., Katta, M. A. V. S. K., Agrawal, G., Thudi, M., Ruperao, P., et al. (2015). CicArVarDB: SNP and InDel database for advancing genetics research and breeding applications in chickpea. Database (Oxford) 2015:bav078. doi: 10.1093/database/bav078
dos Santos, J. V. M., Valliyodan, B., Joshi, T., Khan, S. M., Liu, Y., and Wang, J. (2016). Evaluation of genetic variation among Brazilian soybean cultivars through genome resequencing. BMC Genomics 17:110. doi: 10.1186/s12864-016-2431-x
Duszynska, D., McKeown, P. C., Juenger, T. E., Pietraszewska-Bogiel, A., Geelen, D., and Spillane, C. (2013). Gamete fertility and ovule number variation in selfed reciprocal F1 hybrid triploid plants are heritable and display epigenetic parent-of-origin effects. New Phytol. 198, 71–81. doi: 10.1111/nph.12147
Duvick, D. N. (2001). Biotechnology in the 1930s: the development of hybrid maize. Nat. Rev. Genet. 2, 69–74. doi: 10.1038/35047587
Dwivedi, S. L., Ceccarelli, S., Grando, S., Lammerts van Bueren, E. T., Upadhyaya, H. D., and Ortiz, R. (2017). Diversifying food systems in the pursuit of sustainable food production and healthy diets. Trends Plant Sci. doi: 10.1016/j.tplants.2017.06.011 [Epub ahead of print].
Dwivedi, S. L., Salvatore, C., Blair, M. W., Upadhyaya, H. D., Are, A. K., and Ortiz, R. (2016). Landrace germplasm for improving yield and abiotic stress adaptation. Trends Plant Sci. 21, 31–41. doi: 10.1016/j.tplants.2015.10.012
Dwivedi, S. L., Stalker, H. T., Blair, M. W., Bertioli, D., Upadhyaya, H. D., Nielen, S., et al. (2008). Enhancing crop gene pool with beneficial traits using wild relatives. Plant Breed. Rev. 30, 179–230. doi: 10.1002/9780470380130.ch3
Dwivedi, S. L., Upadhyaya, H. D., Subudhi, P., Gehring, C., Bajic, V., and Ortiz, R. (2010). Abiotic stress tolerance in cereals through breeding and transgenic interventions. Plant Breed. Rev. 33, 31–114. doi: 10.1002/9780470535486.ch2
Eichten, S. R., Briskine, R., Song, J., Li, Q., Swanson-Wagner, R., Hermanson, P. J., et al. (2013). Epigenetic and genetic influences on DNA methylation variation in maize populations. Plant Cell 25, 2783–2797. doi: 10.1105/tpc.113.114793
Eichten, S. R., Swanson-Wagner, R. A., Schnable, J. C., Waters, A. J., Hermanson, P. J., Liu, S., et al. (2011). Heritable epigenetic variation among maize inbreds. PLoS Genet. 7:e1002372. doi: 10.1371/journal.pgen.1002372
Famoso, A. N., Zhao, K., Clark, R. T., Tung, C.-W., Wright, M. H., Bustamante, C., et al. (2011). Genetic architecture of aluminum tolerance in rice (Oryza sativa) determined through genome-wide association analysis and QTL mapping. PLoS Genet. 7:e1002221. doi: 10.1371/journal.pgen.1002221
Fan, C., Xing, Y., Mao, H., Lu, T., Han, B., Xu, C., et al. (2006). GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor. Appl. Genet. 112, 1164–1171. doi: 10.1007/s00122-006-0218-1
Fang, Z., Gonzales, A. M., Clegg, M. T., Smith, K. V., Muehlbauer, G. J., Steffenson, B. J., et al. (2014). Two genomic regions contribute disproportionately to geographic differentiation in wild barley. G3 4, 1193–1203. doi: 10.1534/g3.114.010561/-/DC1
Farfan, I. D. B., De La Fuente, G. N., Murray, S. C., Isakeit, T., Huang, P. C., Warburton, M., et al. (2015). Genome-wide association study for drought, aflatoxin resistance, and important agronomic traits of maize hybrids in the subtropics. PLoS ONE 10:e0117737. doi: 10.1371/journal.pone.0117737
Finnegan, E. J. (2002). Epialleles—a source of random variation in times of stress. Curr. Opin. Plant Biol. 5, 101–106. doi: 10.1016/S1369-5266(02)00233-9
Fleury, D., Jefferies, S., Kuchel, H., and Langridge, P. (2010). Genetic and genomic tools to improve drought tolerance in wheat. J. Exp. Bot. 61, 3211–3222. doi: 10.1093/jxb/erq152
Flint-Garcia, S. A., Thuillet, A. C., Yu, J., Pressoir, G., Romero, S. M., Mitchell, S. E., et al. (2005). Maize association population: a high-resolution platform for quantitative trait locus determination. Plant J. 44, 1054–1064. doi: 10.1111/j.1365-313X.2005.02591.x
Fort, A., Ryder, P., McKeown, P. C., Wijnen, C., Aarts, M. G., Sulpice, R., et al. (2016). Disaggregating polyploidy, parental genome dosage and hybridity contributions to heterosis in Arabidopsis thaliana. New Phytol. 209, 590–599. doi: 10.1111/nph.13650
Foucher, F., Martin, J., Courtiade, J., Cadioux, S., Ellis, N., Banfield, M. J., et al. (2003). Determinate and late flowering are two TERMINAL FLOWER1/CENTRORADIALIS homologs that control two distinct phases of flowering initiation and development in pea. Plant Cell 15, 2742–2754. doi: 10.1105/tpc.015701
Franckowiak, J. D., and Lundqvist, U. (2012). Descriptions of barley genetic stocks for 2012. Barely Genet. Newsl. 42, 36–173.
Fritsche, L. G., Igl, W., Bailey, J. N., Grassmann, F., Sengupta, S., Bragg-Gresham, J. L., et al. (2016). A large genome-wide association study of age-related macular degeneration highlights contributions of rare and common variants. Nat. Genet. 48, 134–143. doi: 10.1038/ng.3448
Fu, C.-Y., Liu, W.-G., Liu, D.-N., Li, J.-H., Zhu, M.-S., Liao, Y.-L., et al. (2016). Genome-wide DNA polymorphism in the indica rice varieties RGD-7S and Taifeng B as revealed by whole genome resequencing. Genome 59, 197–207. doi: 10.1139/gen-2015-0101
Fu, Y., Foden, J. A., Khayter, C., Maeder, M. L., Reyon, D., Joung, J. K., et al. (2013). High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826. doi: 10.1038/nbt.2623
Fu, Y. B. (2015). Understanding crop genetic diversity under modern plant breeding. Theor. Appl. Genet. 128, 2131–2142. doi: 10.1007/s00122-015-2585-y
Fujita, D., Trjatmiko, K. R., Tagle, A. G., Sapasap, M. V., Koide, Y., Sasaki, K., et al. (2013). NAL1 allele from a rice landrace greatly increases yield in modern indica cultivars. Proc. Natl. Acad. Sci. U.S.A. 110, 20431–20436. doi: 10.1073/pnas.1310790110
Fukao, T., Harris, T., and Bailey-Serres, J. (2009). Evolutionary analysis of the Sub1 gene cluster that confers submergence tolerance to domesticated rice. Ann. Bot. 103, 143–150. doi: 10.1093/aob/mcn172
Furbank, R. T., and Tester, M. (2011). Phenomics - technologies to relieve the phenotyping bottleneck. Trends Plant Sci. 16, 635–644. doi: 10.1016/j.tplants.2011.09.005
Gallusci, P., Hodgman, C., Teyssier, E., and Seymour, G. B. (2016). DNA methylation and chromatin regulation during fleshy fruit development and ripening. Front. Plant Sci. 7:807. doi: 10.3389/fpls.2016.00807
Gamon, J., Penuelas, J., and Field, C. (1992). A narrow-waveband spectral index that tracks diurnal changes in photosynthetic efficiency. Remote Sens. Environ. 41, 35–44. doi: 10.1016/0034-4257(92)90059-s
Gamuyao, R., Chin, J. H., Pariasca-Tanaka, J., Pesaresi, P., Catausan, S., Dalid, C., et al. (2012). The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488, 535–541. doi: 10.1038/nature11346
Ganal, M. W., Durstewitz, G., Polley, A., Bérard, A., Buckler, E. S., Charcosset, A., et al. (2011). A large maize (Zea mays L.) SNP genotyping array: Development and germplasm genotyping, and genetic mapping to compare with B73 reference genome. PLoS ONE 6:e28334. doi: 10.1371/journal.pone.0028334
Garnier, O., Laoueillé-Duprat, S., and Spillane, C. (2008). Genomic imprinting in plants. Epigenetics 3, 14–20. doi: 10.4161/epi.3.1.5554
Gasiunas, G., Barrangou, R., Horvath, P., and Siksnys, V. (2012). Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. U.S.A. 109, E2579–E2586. doi: 10.1073/pnas.1208507109
Gilchrist, E. J., and Haughn, G. W. (2005). TILLING without a plough: a new method with applications for reverse genetics. Curr. Opin. Plant Biol. 8, 211–215. doi: 10.1016/j.pbi.2005.01.004
Golicz, A. A., Bayer, P. E., Barker, G. C., Edger, P. P., Kim, H., Martinez, P. A., et al. (2016). The pangenome of an agronomically important crop plant Brassica oleracea. Nat. Commun. 7:13390. doi: 10.1038/ncomms13390
González, A. M., Yuste-Libona, F. J., Saburido, S., Bretons, S., DeRon, A. M., Lozano, R., et al. (2016). Major contribution of flowering time and vegetative growth to plant production in common bean as deduced from a comparative genetic mapping. Front. Plant Sci. 7:1940. doi: 10.3389/fpls.2016.01940
Gore, M. A., Chia, J.-M., Elshire, R. J., Sun, Q., Ersoz, E. S., Hurtwitz, B. L., et al. (2009). A first-generation haplotype map of maize. Science 326, 1115–1117. doi: 10.1126/science.1177837
Gorjanc, G., Jenko, J., Hearne, S. J., and Hickey, J. (2016). Initiating maize pre-breeding programs using genomic selection to harness polygenic variation from landrace populations. BMC Genomics 17:30. doi: 10.1186/s12864-015-2345-z
Gottwald, S., Bauer, P., Komatsuda, T., Lundqvist, U., and Stein, N. (2009). TILLING in the two-rowed barley cultivar ’Barke’ reveals preferred sites of functional diversity in the gene HvHox1. BMC Res. Notes 2:258. doi: 10.1186/1756-0500-2-258
Graham, P. H., and Vance, C. P. (2003). Legumes: importance and constraints to greater use. Plant Physiol. 131, 872–877. doi: 10.1104/pp.017004
Greaves, I. K., Gonzalez-Bayon, R., Wang, L., Zhu, A., Liu, P.-C., Groszmann, M., et al. (2015). Epigenetic changes in hybrids. Plant Physiol. 168, 1197–1205. doi: 10.1104/pp.15.00231
Greaves, I. K., Groszmann, M., Wang, A., Peacock, W. J., and Dennis, E. S. (2014). Inheritance of trans chromosomal methylation patterns from Arabidopsis F1 hybrids. Proc. Natl. Acad. Sci. U.S.A. 111, 2017–2022. doi: 10.1073/pnas.1323656111
Groszmann, M., Greaves, I. K., Albertyn, Z. I., Scofield, G. N., Peacock, W. J., and Dennis, E. S. (2011). Changes in 24-nt siRNA levels in Arabidopsis hybrids suggest an epigenetic contribution to hybrid vigor. Proc. Natl. Acad. Sci. U.S.A. 108, 2617–2622. doi: 10.1073/pnas.1019217108
Groszmann, M., Greaves, I. K., Fujimoto, R., Peacock, W. J., and Dennis, E. S. (2013). The role of epigenetics in hybrid vigour. Trends Genet. 29, 684–690. doi: 10.1016/j.tig.2013.07.004
Gujaria-Verma, N., Ramsay, L., Sharpe, A. G., Sanderson, L. A., Debouck, D. G., Tar’an, B., et al. (2016). Gene-based SNP discovery in tepary bean (Phaseolus acutifolius) and common bean (P. vulgaris) for diversity analysis and comparative mapping. BMC Genomics 17:239. doi: 10.1186/s12864-016-2499-3
Guo, Z., Chen, D., Alqudah, A. M., Röder, M. S., Ganal, M. W., and Schnurbusch, T. (2016). Genome-wide association analyses of 54 traits identified multiple loci for the determination of floral fertility in wheat. New Phytol. 214, 257–270. doi: 10.1111/nph.14342
Gupta, P. K., Rustgi, S., and Kulwal, P. L. (2005). Linkage disequilibrium and association studies in higher plants: present status and future prospects. Plant Mol. Biol. 57, 461–485. doi: 10.1007/s11103-005-0257-z
Hagmann, J., Becker, C., Müller, J., Stegle, O., Meyer, R. C., Wang, G., et al. (2015). Century-scale methylome stability in a recently diverged Arabidopsis thaliana lineage. PLoS Genet. 11:e1004920. doi: 10.1371/journal.pgen.1004920
Haile, J. K., Hammer, K., Badebo, A., Singh, R. P., and Röder, M. S. (2013). Haplotype analysis of molecular markers linked to stem rust resistance genes in Ethiopian improved durum wheat varieties and tetraploid wheat landraces. Genet. Resour. Crop Evol. 60, 853–864. doi: 10.1007/s10722-012-9880-0
Hao, D., Cheng, H., Yin, Z., Cui, S., Zhang, D., Wang, H., et al. (2012). Identification of single nucleotide polymorphisms and haplotypes associated with yield and yield components in soybean (Glycne max) landraces across multiple environments. Theor. Appl. Genet. 124, 447–458. doi: 10.1007/s00122-011-1719-0
Harjes, C. E., Rocheford, T. R., Bai, L., Brutnell, T. P., Kandianis, C. B., Sowinski, S. G., et al. (2008). Natural genetic variation in lycopene epsilon cyclase tapped for maize biofortification. Science 319, 330–333. doi: 10.1126/science.1150255
Hartwig, E. E., and Kiihl, R. A. S. (1979). Identification and utilization of delayed flowering character in soybean for short day conditions. Field Crops Res. 2, 145–151. doi: 10.1016/0378-4290(79)90017-0
Hauben, M., Haesendonckx, B., Standaert, E., van der Kelen, K., Azmi, A., Akpo, H., et al. (2009). Energy use efficiency is characterized by an epigenetic component that can be directed through artificial selection to increase yield. Proc. Natl. Acad. Sci. U.S.A. 106, 20109–20114. doi: 10.1073/pnas.0908755106
Hays, D. B., Do, J. H., Mason, R. E., Morgan, G., and Finlayson, S. A. (2007). Heat stress induced ethylene production in developing wheat grains induces kernel abortion and increased maturation in a susceptible cultivar. Plant Sci. 172, 1113–1123. doi: 10.1016/j.plantsci.2007.03.004
Henry, I. M., Nagalakshmi, U., Lieberman, M. C., Ngo, K. J., Krasileva, K. V., Vasquez-Gross, H., et al. (2014). Efficient genome-wide detection and cataloging of EMS-induced mutations using exome capture and next-generation sequencing. Plant Cell 26, 1382–1397. doi: 10.1105/tpc.113.121590
Herritt, M., Dhanapal, A. P., and Fritschi, F. B. (2016). Identification of genomic loci associated with the photochemical reflectance index by genome-wide association study in soybean. Plant Genome 9. doi: 10.3835/plantgenome2015.08.0072
Heslot, N., Rutkoski, J., Poland, J., Jannink, J. L., and Sorrells, M. E. (2013). Impact of marker ascertainment bias on genomic selection accuracy and estimates of genetic diversity. PLoS ONE 8:e74612. doi: 10.1371/journal.pone.0074612
Hirsch, C. D., Hamilton, J. P., Childs, K. L., Cepela, J., Crisovan, E., Vaillancourt, B., et al. (2014). Spud DB: a resource for mining sequences, genotypes, and phenotypes to accelerate potato breeding. Plant Genome 7. doi: 10.3835/plantgenome2013.12.0042
Hollick, J. B. (2016). Paramutation and related phenomena in diverse species. Nat. Rev. Genet. 18, 5–23. doi: 10.1038/nrg.2016.115
Hu, Z., Mbacke, B., Perumal, R., Guéye, M. C., Sy, O., Bouchet, S., et al. (2015). Population genomics of pearl millet (Pennisetum glaucum (L.) R. Br.): Comparative analysis of global accessions and Senegalese landraces. BMC Genomic 16:1048. doi: 10.1186/s12864-015-2255-0
Huang, X., and Han, B. (2014). Natural variations and genome-wide association studies in crop plants. Annu. Rev. Plant Biol. 65, 531–551. doi: 10.1146/annurev-arplant-050213-035715
Huang, X., Qian, Q., Liu, Z., Sun, H., He, S., Luo, D., et al. (2009). Natural variation at the DEEP1 locus enhances grain yield in rice. Nat. Genet. 41, 494–497. doi: 10.1038/ng.352
Huang, X., Wei, X., Sang, T., Zhao, Q., Feng, Q., Zhao, Y., et al. (2010). Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967. doi: 10.1038/ng.695
Huang, X., Zhao, Y., Wei, X., Li, C., Wang, A., Zhao, Q., et al. (2012). Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat. Genet. 44, 32–41. doi: 10.1038/ng.1018
Huq, M. A., Akter, S., Nou, I. S., Kim, H. T., Jung, Y. J., and Kang, K. K. (2016). Identification of functional SNPs in genes and their effects on plant phenotypes. J. Plant Biotechnol. 43, 1–11. doi: 10.5010/JPB.2016.43.1.1
Huynh, B. L., Close, T. J., Roberts, P. A., Hu, Z., Wanamaker, S., Lucas, M. R., et al. (2013). Gene pools and the genetic architecture of domesticated cowpea. Plant Genome 6. doi: 10.3835/plantgenome2013.03.0005
Ingvarsson, P. K., and Street, N. R. (2011). Association genetics of complex traits in plants. New Phytol. 189, 909–922. doi: 10.1111/j.1469-8137.2010.03593.x
Ingvordsen, C. H., Backes, G., Lyngkjaer, M. F., Peltonen-Sainio, P., Jahoor, A., Mikkelsen, T. N., et al. (2015). Genome-wide association study of production and stability traits in barley cultivated under future climate scenarios. Mol. Breed. 35:84. doi: 10.1007/s11032-015-0283-8
Jannink, J. L., and Walsh, B. (2002). “Association mapping in plant populations,” in Quantitative Genetics, Genomics and Plant Breeding, ed. M. S. Kong (Wallingford: CABI), 59–68.
Jarquin, D., Specht, J., and Lorenz, A. (2016). Prospects of genomic prediction in the USDA soybean germplasm collection: historical data creates robust models for enhancing selection of accessions. G3 (Bethesda) 6, 2329–2341. doi: 10.1534/g3.116.031443
Jia, J., Li, H., Zhang, X., Li, Z., and Qiu, L. (2017). Genomics-based plant germplasm research (GPGR). Crop J. 5, 166–174. doi: 10.1016/j.cj.2016.10.006
Jiang, K., Liberatore, K. L., Park, S. J., Alvarez, J. P., and Lippman, Z. B. (2013). Tomato yield heterosis is triggered by a dosage sensitivity of the florigen pathway that fine-tunes shoot architecture. PLoS Genet. 9:e1004043. doi: 10.1371/journal.pgen.1004043
Jiang, W. Z., Zhou, H. B., Bi, H. H., Fromm, M., Yang, B., and Weeks, D. P. (2013). Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res. 41, e188. doi: 10.1093/nar/gkt780
Jiao, Y., Wang, Y., Xue, D., Wang, J., Yan, M., Liu, J., et al. (2010). Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 42, 541–544. doi: 10.1038/ng.591
Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., and Charpentier, E. (2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821. doi: 10.1126/science.1225829
Johannes, F., Porcher, E., Teixeira, F. K., Saliba-Colombani, V., Simon, M., Agier, N., et al. (2009). Assessing the impact of transgenerational epigenetic variation on complex traits. PLoS Genet. 5:e1000530. doi: 10.1371/journal.pgen.1000530
Jonas, E., and de Koning, D. J. (2013). Does genomic selection have a future in plant breeding? Trends Biotechnol. 31, 497–504. doi: 10.1016/j.tibtech.2013.06.003
Jordan, K. W., Wang, S., Lun, Y., Gardiner, L.-J., MacLachlan, R., Hucl, P., et al. (2015). A haplotype map of allohexaploid wheat reveals distinct patterns of selection on homoeologous genomes. Genome Biol. 16, 48. doi: 10.1186/s13059-015-0606-4
Joshi, T., Fitzpatrick, M. R., Chen, S. Y., Liu, Y., Zhang, H. X., Endacott, R. Z., et al. (2014). Soybean knowledge base (SoyKB): a web resource for integration of soybean translational genomics and molecular breeding. Nucleic Acids Res. 42, D1245–D1252. doi: 10.1093/nar/gkt905
Jung, C., and Muller, A. E. (2009). Flowering time control and applications in plant breeding. Trends Plant Sci. 14, 563–573. doi: 10.1016/j.tplants.2009.07.005
Kale, S. M., Jaganathan, D., Ruperao, P., Chen, C., Punna, R., Kudapa, H., et al. (2015). Prioritization of candidate genes in ‘QTL-hotspot’ region for drought tolerance in chickpea (Cicer arietinum L.). Sci. Rep. 5:15296. doi: 10.1038/srep15296
Kamfwa, K., Cichy, K. A., and Kelly, J. D. (2015). Genome-wide association study of agronomic traits in common bean. Plant Genome 8. doi: 10.3835/plantgenome2014.09.0059
Kanz, C., Aldebert, P., Althorpe, N., Baker, W., Baldwin, A., Bates, K., et al. (2005). The EMBL nucleotide sequence database. Nucleic Acids Res. 33, D29–D33.
Karlgren, A., Gyllenstrand, N., Kallman, T., Sundstrom, J. F., Moore, D., Lascoux, M., et al. (2011). Evolution of the PEBP gene family in plants: functional diversification in seed plant evolution. Plant Physiol. 156, 1967–1977. doi: 10.1104/pp.111.176206
Kawakatsu, T., Huang, S.-C. C., Jupe, F., Sasaki, E., Schmitz, R. J., Urich, M. A., et al. (2016). Epigenomic diversity in a global collection of Arabidopsis thaliana accessions. Cell 166, 492–505. doi: 10.1016/j.cell.2016.06.044
Kawanabe, T., Ishikura, S., Miyaji, N., Sasaki, T., Wu, L. M., Itabashi, E., et al. (2016). Role of DNA methylation in hybrid vigor in Arabidopsis thaliana. Proc. Natl. Acad. Sci. U.S.A. 113, E6704–E6711. doi: 10.1073/pnas.1613372113
Kilian, B., and Graner, A. (2012). NGS technologies for analyzing germplasm diversity in genebanks. Brief. Funct. Genomics 11, 38–50. doi: 10.1093/bfgp/elr046
King, G. J. (2015). Crop epigenetics and the molecular hardware of genotype × environment interactions. Front. Plant Sci. 6:968. doi: 10.3389/fpls.2015.00968
Klann, T. S., Black, J. B., Chellappan, M., Safi, A., Song, L., Hilton, I. B., et al. (2017). CRISPR-Cas9 epigenome editing enables high-throughput screening for functional regulatory elements in the human genome. Nat. Biotechnol. 35, 561–568. doi: 10.1038/nbt.3853
Klein, R. R., Miller, F. R., Bean, S., and Klein, P. E. (2016). Registration of 40 converted germplasm sources from the reinstated sorghum conversion program. J. Plant Reg. 45, 57–61. doi: 10.3198/jpr2015.05.0034crg
Kleinstiver, B. P., Pattanayak, V., Prew, M. S., Tsai, S. Q., Nguyen, N. T., Zheng, Z. L., et al. (2016). High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495. doi: 10.1038/nature16526
Kole, C., Muthamilarasan, M., Henry, R., Edwards, D., Sharma, R., Abberton, M., et al. (2015). Application of genomics for generation of climate resilient crops: progress and prospects. Front. Plant Sci. 6:563. doi: 10.3389/fpls.2015.00563
Kong, F., Nan, H., Cao, D., Li, Y., Wu, F., Wang, J., et al. (2014). A new dominant gene E9 conditions early flowering and maturity in soybean. Crop Sci. 54, 1220–1231. doi: 10.2135/cropsci2014.03.0228
Kong, F. J., Liu, B., Xia, Z., Sato, S., Kim, B. M., Watanabe, S., et al. (2010). Two coordinately regulated homologs of FLOWERING LOCUS T are involved in the control of photoperiodic flowering in soybean. Plant Physiol. 154, 1220–1231. doi: 10.1104/pp.110.160796
Koo, B.-H., Yoo, S.-C., Park, J.-W., Kwon, C.-T., Lee, B.-D., An, G., et al. (2013). Natural variation in OsPRR37 regulates heading date and contributes to rice cultivation at a wide range of latitudes. Mol. Plants 6, 1877–1888. doi: 10.1093/mp/sst088
Kooke, R., Johannes, F., Wardenaar, R., Becker, F., Etcheverry, M., Colot, V., et al. (2015). Epigenetic basis of morphological variation and phenotypic plasticity in Arabidopsis thaliana. Plant Cell 27, 337–348. doi: 10.1105/tpc.114.133025
Koornneef, M., Hanhart, C. J., and Vanderveen, J. H. (1991). A genetic and physiological analysis of late flowering mutants in Arabidopsis thaliana. Mol. Gen. Genet. 229, 57–66. doi: 10.1007/BF00264213
Kujur, A., Bajaj, D., Upadhyaya, H. D., Das, S., Ranjan, R., Shree, T., et al. (2015). A genome-wide SNP scan accelerates trait-regulatory genomic loci identification in chickpea. Sci. Rep. 5:11166. doi: 10.1038/srep11166
Kumar, A., Dixit, S., Ram, T., Yadav, R. B., Mishra, K. K., and Mandal, N. P. (2014). Breeding high-yielding drought-tolerant rice: genetic variations and conventional and molecular approaches. J. Exp. Bot. 65, 6265–6278. doi: 10.1093/jxb/eru363
Kumar, A. P., McKeown, P. C., Boualem, A., Ryder, P., Brychkova, G., Bendahmane, A., et al. (2017). TILLING by Sequencing (TbyS) for targeted genome mutagenesis in crops. Mol. Breed. 37, 14. doi: 10.1007/s11032-017-0620-1
Kumar, V., Khan, A. W., Saxena, R. K., Garg, V., and Varshney, R. K. (2016). First-generation hapmap in Cajanus spp. reveals untapped variations in parental lines of mapping populations. Plant Biotechnol. J. 14, 1673–1681. doi: 10.1111/pbi.12528
Kumar, V., Singh, A., Amitha Mitra, S. V., Krishnamurthy, S. L., Parida, S. K., Jain, S., et al. (2015). Genome-wide association mapping of salinity tolerance in rice (Oryza sativa). DNA Res. 22, 133–145. doi: 10.1093/dnares/dsu046
Lai, J., Li, R., Xu, X., Jin, W., Xu, M., Zhao, H., et al. (2010). Genome-wide patterns of genetic variation among elite maize inbred lines. Nat. Genet. 42, 1027–1031. doi: 10.1038/ng.684
Lai, K., Lorenc, M. T., Lee, H. C., Berkman, P. J., Bayer, P. E., Visendi, P., et al. (2015). Identification and characterization of more than 4 million intervarietal SNPs across the group 7 chromosomes of bread wheat. Plant Biotechnol. J. 13, 97–104. doi: 10.1111/pbi.12240
Lam, H.-M., Xu, X., Liu, X., Chen, W., Yang, G., Wong, F.-L., et al. (2010). Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat. Gent. 42, 1053–1059. doi: 10.1038/ng.715
Lasky, J. R., Upadhyaya, H. D., Ramu, P., Deshpande, S., Hash, C. T., Bonnette, J., et al. (2015). Genome-environment associations in sorghum landraces predict adaptive traits. Sci. Adv. 1:e1400218. doi: 10.1126/sciadv.1400218
Lawrence, C. J., Dong, O. F., Polacco, M. L., Seigfried, T. E., and Brendel, V. (2004). MaizeGDB, the community database for maize genetics and genomics. Nucleic Acids Res. 32, D393–D397. doi: 10.1093/nar/gkh011
Leamy, L. J., Lee, C. R., Song, Q., Mujacic, I., Luo, Y., Chen, C. Y., et al. (2016). Environmental versus geographical effects on genomic variation in wild soybean (Glycine soja) across its native range in northeast Asia. Ecol. Evol. 6, 6332–6344. doi: 10.1022/ece3.2351
Lee, H. C., Lai, K. T., Lorenc, M. T., Imelfort, M., Duran, C., and Edwards, D. (2012). Bioinformatics tools and databases for analysis of next-generation sequence data. Brief. Funct. Genomics 11, 12–24. doi: 10.1093/bfgp/elr037
Lee, Y. G., Jeong, N., Kim, J. H., Lee, K., Kim, K. H., and Pirani, A. (2015). Development, validation and genetic analysis of a large soybean SNP genotyping array. Plant J. 81, 625–636. doi: 10.1111/tpj.12755
Leiser, W. L., Rattunde, H. F. W., Weltzein, E., Cisse, N., Abdou, M., Diallo, A., et al. (2014). Two in one sweep: aluminum tolerance and grain yield in P-limited soils are associated to the same genomic region in West African Sorghum. BMC Plant Biol. 14:206. doi: 10.1186/s12870-014-0206-6
Lesk, C., Rowhami, P., and Ramankutty, N. (2016). Influence of extreme weather disasters on global crop production. Nature 529, 84–87. doi: 10.1038/nature16467
Li, M., Li, X., Zhou, Z., Wu, P., Fang, M., Pan, X., et al. (2016). Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front. Plant Sci. 7:377. doi: 10.3389/fpls.2016.00377
Li, Q., Eichten, S. R., Hermanson, P. J., and Springer, N. M. (2014). Inheritance patterns and stability of DNA methylation variation in maize near-isogenic lines. Genetics 196, 667–676. doi: 10.1534/genetics.113.158980
Li, X., Zhou, Z., Ding, J., Wu, Y., Zhou, B., Wang, R., et al. (2016). Combined linkage and association mapping reveals QTL and candidate genes for plant and ear height in maize. Front. Plant Sci. 7:833. doi: 10.3389/fpls.2016.00833
Li, X. M., Chao, D. Y., Wu, Y., Huang, X., Chen, K., Cui, L. G., et al. (2015). Natural alleles of a proteasome α2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat. Genet. 47, 827–833. doi: 10.1038/ng.3305
Li, Y.-H., Zhao, S.-C., Ma, J.-X., Li, D., Yan, L., Li, J., et al. (2013). Molecular footprints of domestication and improvement in soybean revealed by whole genome re-sequencing. BMC Genomics 14:579. doi: 10.1186/1471-2164-14-579
Li, Y. H., Zhou, G., Ma, J., Jiang, W., Jin, L. G., Zhang, Z., et al. (2014). De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat. Biotechnol. 32, 1045–1052. doi: 10.1038/nbt.2979
Li, Z., Hu, G., Liu, X., Zhou, Y., Li, Y., Zhang, X., et al. (2016). Transcriptome sequencing identified genes and gene ontologies associated with early freezing tolerance in maize. Front. Plant Sci. 7:1477. doi: 10.3389/fpls.2016.01477
Li, Z., Wakao, S., Fischer, B. B., and Niyogi, K. K. (2009). Sensing and responding to excess light. Annu. Rev. Plant Biol. 60, 391–395. doi: 10.1146/annurev.arplant.58.032806.103844
Lin, K., Zhang, N., Severing, E. I., Nijveen, H., Cheng, F., Visser, R. G., et al. (2014). Beyond genomic variation–comparison and functional annotation of three Brassica rapa genomes: a turnip, a rapid cycling and a Chinese cabbage. BMC Genomics 15:250. doi: 10.1186/1471-2164-15-250
Liu, E., Liu, Y., Wu, G., Zeng, S., Thi, T. G. T., Liang, L., et al. (2016). Identification of a candidate gene for panicle length in rice (Oryza sativa L.) via association and linkage analysis. Front. Plant Sci. 7:596. doi: 10.3389/fpls.2016.00596
Liu, H., Wang, X., Wei, B., Wang, Y., Liu, Y., and Zhang, J. (2016). Characterization of genome-wide variation in four-row wax, a waxy landrace with a reduced kernel row phenotype. Front. Plant Sci. 7:667. doi: 10.3389/fpls.2016.00667
Liu, Y., Wang, L., Deng, M., Li, Z., Lu, Y., Wang, J., et al. (2015). Genome-wide association study of phosphorus-deficiency-tolerance traits in Aegilops tauschii. Theor. Appl. Genet. 128, 2203–2212. doi: 10.1007/s00122-015-2578-x
Liu, R., How-Kit, A., Stammitti, L., Teyssier, E., Rolin, D., Mortain-Bertrand, A., et al. (2015). A DEMETER-like DNA demethylase governs tomato fruit ripening. Proc. Natl. Acad. Sci. U.S.A. 112, 10804–10809. doi: 10.1073/pnas.1503362112
Lombardi, M., Materne, M., Cogan, N. O. I., Rodda, M., Daetwyler, H. D., Slater, A. T., et al. (2014). Assessment of genetic variation within a global collection of lentil (Lens culinaris Medik.) cultivars and landraces using SNP markers. BMC Genet. 15:150. doi: 10.1186/s12863-014-0150-3
Longin, C. F. H., and Reif, J. C. (2014). Redesigning the exploitation of wheat genetic resources. Trends Plant Sci. 19, 631–636. doi: 10.1016/j.tplants.2014.06.012
Lu, L., Yan, W., Xue, W., Shao, D., and Xing, Y. (2012). Evolution and association analysis of Ghd7 in rice. PLoS ONE 7:e34021. doi: 10.1371/journal.pone.0034021
Lu, S., Zhao, X., Hu, Y., Liu, S., Nan, H., Li, X., et al. (2017). Natural variation at the soybean J locus improves adaptation to the tropics and enhances yield. Nat. Genet. 49, 773–779. doi: 10.1038/ng.3819
Lu, Y., Shah, T., Hao, Z., Taba, S., Zhang, S., Gao, S., et al. (2011). Comparative SNP and haplotype analysis reveals a higher genetic diversity and rapider LD decay in tropical than temperate germplasm in maize. PLoS ONE 6:e24861. doi: 10.1371/journal.pone.0024861
Lu, Y., Zhang, S., Shah, T., Xie, C., Hao, Z., Li, X., et al. (2009). Joint linkage–linkage disequilibrium mapping is a powerful approach to detecting quantitative trait loci underlying drought tolerance in maize. Proc. Natl. Acad. Sci. U.S.A. 107, 19585–19590. doi: 10.1073/pnas.1006105107
Ma, Y., Dai, X., Xu, Y., Luo, W., Zheng, X., Zeng, D., et al. (2015). COLD1 confers chilling tolerance in rice. Cell 160, 1209–1221. doi: 10.1016/j.cell.2015.01.046
Mace, E. S., Tai, S., Gilding, E. K., Li, Y., Prentis, P. J., Bian, L., et al. (2013). Whole-genome sequencing reveals untapped genetic potential in Africa’s indigenous cereal crop sorghum. Nat. Commun. 4, 2320. doi: 10.1038/ncomms3320
Mackay, I., and Powell, W. (2007). Methods of linkage disequilibrium mapping in crops. Trends Plant Sci. 44, 1054–1064. doi: 10.1016/j.tplants.2006.12.001
Mackill, D. J., Ismail, A. M., Singh, U. S., Labios, R. V., and Paris, T. R. (2012). Development and rapid adoption of sub-mergence-tolerant (Sub1) rice varieties. Adv. Agron. 155, 299–352. doi: 10.1016/B978-0-12-394276-0.00006-8
Madduri, R. K., Sulakhe, D., Lacinski, L., Liu, B., Rodriguez, A., Chard, K., et al. (2014). Experiences building globus genomics: a next-generation sequencing analysis service using Galaxy, Globus, and Amazon Web Services. Concurr. Comput. 26, 2266–2279. doi: 10.1002/cpe.3274
Makumburage, G. B., Richbourg, H. L., La Torre, K. D., Capps, A., Chen, C. X., and Stapleton, A. E. (2013). Genotype to phenotype maps: multiple input abiotic signals combine to produce growth effects via attenuating signaling interactions in maize. G3 (Bethesda) 3, 2195–2204. doi: 10.1534/g3.113.008573
Manning, K., Tör, M., Poole, M., Hong, Y., Thompson, A. J., Graham, J. K., et al. (2006). A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat. Genet. 38, 948–952. doi: 10.1038/ng1841
Martin, A., Troadec, C., Boualem, A., Rajab, M., Fernandez, R., Morin, H., et al. (2009). A transposon-induced epigenetic change leads to sex determination in melon. Nature 461, 1135–1138. doi: 10.1038/nature08498
Mashima, J., Kodama, Y., Kosuge, T., Fujisawa, T., Katayama, T., Nagasaki, H., et al. (2016). DNA data bank of Japan (DDBJ) progress report. Nucleic Acids Res. 44, D51–D57. doi: 10.1093/nar/gkv1105
Maurer, A., Draba, V., Jiang, Y., Schnaithmann, F., Sharma, R., Schumann, E., et al. (2015). Modelling the genetic architecture of flowering time in barley through nested association mapping. BMC Genomics 16:290. doi: 10.1186/s12864-015-1459-7
McKeown, P. C., Fort, A., Duszynska, D., Sulpice, R., and Spillane, C. (2013). Emerging molecular mechanisms for biotechnological harnessing of heterosis in crops. Trends Biotechnol. 31, 549–551. doi: 10.1016/j.tibtech.2013.06.008
Mejlhede, N., Kyjovska, Z., Backes, G., Burhenne, K., Rasmussen, S. K., and Jahoor, A. (2006). EcoTILLING for the identification of allelic variation in the powdery mildew resistance genes mlo and Mla of barley. Plant Breed. 125, 461–467. doi: 10.1111/j.1439-0523.2006.01226.x
Meng, D., Dubin, M., Zhang, P., Osborne, E. J., Stegle, O., Clark, R. M., et al. (2016). Limited contribution of DNA methylation variation to expression regulation in Arabidopsis thaliana. PLoS Genet. 12:e1006141. doi: 10.1371/journal.pgen.1006141
Michael, T. P., and VanBuren, R. (2015). Progress, challenges and the future of crop genomes. Curr. Opin. Plant Biol. 24, 71–81. doi: 10.1016/j.pbi.2015.02.002
Millet, E. J., Welcker, C., Kruijer, W., Negro, S., Coupel-Ledru, A., Nicolas, S. D., et al. (2016). Genome-wide analysis of yield in Europe: allelic effects vary with drought and heat scenarios. Plant Physiol. 172, 749–764. doi: 10.1104/pp.16.00621
Mir, R. R., Kudapa, H., Srikanth, S., Saxena, R. K., Sharma, A., Azam, S., et al. (2014). Candidate gene analysis for determinacy in pigeonpea (Cajanus spp.). Theor. Appl. Genet. 127, 2663–2678. doi: 10.1007/s00122-014-2406-8
Mishra, S., Singh, B., Misra, P., Rai, V., and Singh, N. K. (2016a). Haplotype distribution and association of candidate genes with salt tolerance in Indian wild rice germplasm. Plant Cell Rep. 35, 2295–2308. doi: 10.1007/s00299-016-2035-6
Mishra, S., Singh, B., Panda, K., Singh, B. P., Singh, N., Misra, P., et al. (2016b). Association of SNP haplotypes of HKT family genes with salt tolerance in Indian wild rice germplasm. Rice 9, 15. doi: 10.1186/s12284-016-0083-8
Miura, K., Ikeda, M., Matsubara, A., Song, X., Ito, M., Asano, K., et al. (2010). OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 42, 545–549. doi: 10.1038/ng.592
Montenegro, J. D., Golicz, A. A., Bayer, P. E., Hurgobin, B., Lee, H., Chan, C. K. K., et al. (2017). The pangenome of hexaploid bread wheat. Plant J. 90, 1007–1013. doi: 10.1111/tpj.13515
Morris, G. P., Ramu, P., Deshpande, S. P., Hash, C. T., Shah, T., Upadhyaya, H. D., et al. (2013). Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc. Natl. Acad. Sci. .U.S.A. 110, 453–458. doi: 10.1073/pnas.1215985110
Multani, D. S., Briggs, S. P., Chamberlin, M. A., Blakeslee, J. J., Murphy, A. S., and Johal, G. S. (2003). Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 302, 81–84. doi: 10.1126/science.1086072
Muraya, M. M., Schmutzer, T., Ulipinnis, C., Scholz, U., and Altmann, T. (2015). Targeted sequencing reveals large-scale sequence polymorphism in maize candidate genes for biomass production and composition. PLoS ONE 10:e0132120. doi: 10.1371/journal.pone.0132120
Murphy, R. L., Klein, R. R., Morishige, D. T., Brady, J. A., Rooney, W. L., Miller, F. R., et al. (2011). Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum. Proc. Natl. Acad. Sci. U.S.A. 108, 16469–16474. doi: 10.1073/pnas.1106212108
Murphy, R. L., Morishige, D. T., Brady, J. A., Rooney, W. L., Yang, S., Klein, P. E., et al. (2014). Ghd7 (Ma6) represses sorghum flowering in long days: Ghd7 alleles enhance biomass accumulation and grain production. Plant Genome 7, 1–10. doi: 10.3835/plantgenome2013.11.0040
Naranjo, L., Talón, M., and Domingo, C. (2014). Diversity of floral regulatory genes of japonica rice cultivated at northern latitudes. BMC Genomics 15:101. doi: 10.1186/1471-2164-15-101
Nelson, M. N., Ksiazkiewicz, M., Rychel, S., Besharat, N., Taylor, C. M., Wyrwa, K., et al. (2017). The loss of vernalization requirement in narrow-leafed lupin is associated with a deletion in the promoter and de-repressed expression of a Flowering Locus T (FT) homologue. New Phytol. 213, 220–232. doi: 10.1111/nph.14094
Neumaier, N., and James, A. T. (1993). Exploiting the long-juvenile trait to improve adaptation of soybean to the tropics. Food Legume Newsl. 8, 12–14.
Nevo, E., Fu, Y.-B., Pavlicek, T., Khalifa, S., Tavasi, M., and Beiles, A. (2012). Evaluation of wild cereals during 28 years of global warming in Israel. Proc. Natl. Acad. Sci. .U.S.A. 109, 3412–3415. doi: 10.1073/pnas.1121411109
Niemenmaa, M., Kallio, A., Schumacher, A., Klemela, P., Korpelainen, E., and Heljanko, K. (2012). Hadoop-BAM: directly manipulating next generation sequencing data in the cloud. Bioinformatics 28, 876–877. doi: 10.1093/bioinformatics/bts054
Ning, L., Kan, G., Du, W., Guo, S., Wang, Q., Zhang, G., et al. (2016). Association analysis for detecting significant single nucleotide polymorphisms for phosphorus-deficiency tolerance at the seedling stage in soybean [Glycine max (L) Merr.]. Breed. Sci. 66, 191–203. doi: 10.1270/jsbbs.66.191
Nordberg, H., Bhatia, K., Wang, K., and Wang, Z. (2013). BioPig: a Hadoop-based analytic toolkit for large-scale sequence data. Bioinformatics 29, 3014–3019. doi: 10.1093/bioinformatics/btt528
Oakey, H., Cullis, B., Thompson, R., Comadran, J., Halpin, C., and Waugh, R. (2016). Genomic selection in multi-environment crop trials. G3 (Bethesda) 6, 1313–1326. doi: 10.1534/g3.116.027524/-/DC1
O’Driscoll, A., Daugelaite, J., and Sleator, R. D. (2013). ’Big data’, Hadoop and cloud computing in genomics. J. Biomed. Inform. 46, 774–781. doi: 10.1016/j.jbi.2013.07.001
Oerke, E. C. (2006). Crop losses to pests. J. Agric. Sci. 144, 31–43. doi: 10.1017/S0021859605005708
Offermann, S., and Peterhansel, C. (2014). Can we learn from heterosis and epigenetics to improve photosynthesis? Curr. Opin. Plant Biol. 19, 105–110. doi: 10.1016/j.pbi.2014.05.010
Ogiso-Tanaka, E., Matsubara, K., Yamamoto, S., Nonoue, Y., Wu, J. Z., Fujisawa, H., et al. (2013). Natural variation of the RICE FLOWERING LOCUS T 1 contributes to flowering time divergence in rice. PLoS ONE 8:e75959. doi: 10.1371/journal.pone.0075959
Ong-Abdullah, M., Ordway, J. M., Jiang, N., Ooi, S.-E., Kok, S.-Y., Sarpan, N., et al. (2015). Loss of Karma transposon methylation underlies the mantled somaclonal variant of oil palm. Nature 525, 533–537. doi: 10.1038/nature15365
Ortiz, R. (2013). “Drought tolerance,” in Genomics and Breeding for Climate-Resilient Crops, Target Traits, Vol. 2, ed. C. Kole (Dordrecht: Springer), 203–223. doi: 10.1007/978-3-642-37048-9_5
Ortiz, R. (2015). Plant Breeding in the Omics Era. New York, NY: Springer, 249. doi: 10.1007/978-3-319-20532-8
Ousseini, I. S., Bakasso, Y., Kane, N. A., Couderc, M., Zekraoui, L., Mariac, C., et al. (2017). Myosin XI is associated with fitness and adaptation to aridity in wild pearl millet. Heredity 119, 88–94. doi: 10.1038/hdy.2017.13
Park, M., Keung, A. J., and Khalil, A. S. (2016). The epigenome: the next substrate for engineering. Genome Biol. 17, 183. doi: 10.1186/s13059-016-1046-5
Park, S. J., Jiang, K., Tal, L., Yichie, Y., Gar, O., Zamir, D., et al. (2014). Optimization of crop productivity in tomato using induced mutations in the florigen pathway. Nat. Genet. 46, 1337–1342. doi: 10.1038/ng.3131
Pasam, R. K., Sharma, R., Malosetti, M., van Eeuwijk, F. A., Haseneyer, G., Kilian, B., et al. (2012). Genome-wide association studies for agronomical traits in a worldwide spring barley collection. BMC Plant Biol. 12:16. doi: 10.1186/1471-2229-12-16
Pasam, R. K., Sharma, R., Walther, A., Özkan, H., Graner, A., and Kilian, B. (2014). Genetic diversity and population structure in a legacy collection of spring barley landraces adapted to a wide range of climates. PLoS ONE 9:e116164. doi: 10.1371/journal.pone.0116164
Pattanayak, V., Lin, S., Guilinger, J. P., Ma, E. B., Doudna, J. A., and Liu, D. R. (2013). High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 31, 839–843. doi: 10.1038/nbt.2673
Peng, J., Richards, D. E., Hartley, N. M., Murphy, G. P., Devos, K. M., Flintham, J. E., et al. (1999). Green revolution genes encode mutant gibberellin response modulators. Nature 400, 256–261. doi: 10.1038/22307
Pérez-Enciso, M., Rincón, J. C., and Legarra, A. (2015). Sequence-vs. chip-assisted genomic selection: accurate biological information is advised. Genet. Sel. Evol. 47, 43. doi: 10.1186/s12711-015-0117-5
Periyannan, S., Moore, J., Ayliffe, M., Bansal, U., Wang, X., Huang, L., et al. (2013). The gene Sr33, an ortholog of barley Mla genes, encodes resistance to wheat stem rust race Ug99. Science 341, 786–788. doi: 10.1126/science.1239028
Petchey, O. L., and Gaston, K. J. (2006). Functional diversity: back to basics and looking forward. Ecol. Lett. 9, 741–758. doi: 10.1111/j.1461-0248.2006.00924.x
Pikaard, C. S., and Scheid, O. M. (2014). Epigenetic regulation in plants. Cold Spring Harb. Perspect. Biol. 6:a019315. doi: 10.1101/cshperspect.a019315
Poets, A. M., Fang, Z., Clegg, M. T., and Morrell, P. L. (2015). Barley landraces are characterized by geographically heterogeneous genomic origins. Genome Biol. 16, 173. doi: 10.1186/s13059-015-071-3
Prins, R., Dreisigacker, S., Pretorius, Z., Schalkwyk, H., Wessels, E., Smit, C., et al. (2016). Stem rust resistance in a geographically diverse collection of spring wheat lines collected from across Africa. Front. Plant Sci. 7:973. doi: 10.3389/fpls.2016.00973
Pyhäjärvi, T., Hufford, M. B., Mezmouk, S., and Ross-Ilbarra, J. (2013). Complex patterns of local adaptation in teosinte. Genome Biol. Evol. 5, 1594–1609. doi: 10.1093/gbe.evt109
Qi, X., Li, M. W., Xie, M., Liu, X., Ni, M., Shao, G., et al. (2014). Identification of a novel salt tolerant gene in wild soybean by whole-genome sequencing. Nat. Commun. 5, 4340. doi: 10.1038/ncomm5340
Qin, P., Lin, Y., Hu, Y., Liu, K., Mao, S., Li, Z., et al. (2016). Genome-wide association study of drought-related resistance traits in Aegilops tauschii. Genet. Mol. Biol. 39, 398–407. doi: 10.1590/1678-4685-GMB-2015-0232
Quinby, J. R., and Karper, R. E. (1954). Inheritance of height in sorghum. Agron. J. 46, 211–216. doi: 10.2134/agronj1954.00021962004600050007x
Raggi, L., Tissi, C., Mazzucato, A., and Negri, V. (2014). Molecular polymorphism related flowering trait variation in a Phaseolus vulgaris L. collection. Plant Sci. 216, 180–189. doi: 10.1016/j.plantsci.2013.11.001
Ran, F. A., Hsu, P. D., Lin, C.-Y., Gootenberg, J. S., Konermann, S., Trevino, A. E., et al. (2013). Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389. doi: 10.1016/j.cell.2013.08.021
Rathnayake, N., Bentota, A. P., Dissanayake, D. M. N., Perera, K., Sooriyapathirana, S. D. S. S., and Jayasekera, G. A. U. (2012). DNA markers RM 464A and RM 219 haplotypes are effective in selecting Sub-1 locus for the introgression of submergence tolerance into new rice varieties. Ceylon J. Sci. 41, 125–136.
Rebolledo, M. G., Peña, A. L., Duitama, J., Cruz, D. F., Dingkuhn, M., Grenier, C., et al. (2016). Combining image analysis, genome wide association studies and different field trials to reveal stable genetic regions related to panicle architecture and the number of spikelets per panicle in rice. Front. Plant Sci. 7:1384. doi: 10.3389/fpls.2016.01384
Reinert, S., Kortz, A., Léon, J., and Naz, A. A. (2016). Genome-wide association mapping in the global diversity set reveals new QTL controlling root system and related shoot variation in barley. Front. Plant Sci. 7:1061. doi: 10.3389/fpls.2016.01061
Rhee, S. Y., and Mutwil, M. (2014). Towards revealing the functions of all genes in plants. Trends Plant Sci. 19, 212–221. doi: 10.1016/j.tplants.2013.10.006
Romay, M. C., Millard, M. J., Glaubitz, J. C., Peiffer, J. A., Swarts, K. L., Castevens, T. M., et al. (2013). Comprehensive genotyping of the United States national maize inbred seed bank. Genome Biol. 14:R55. doi: 10.1186/gb-2013-14-6-r55
Roorkiwal, M., Jain, A., Kale, S. M., Chitikineni, A., Thudi, M., Gaur, P. M., et al. (2016). “AxiomCicerSNP Array”: A High-Throughput SNP Genotyping Platform for High Resolution Genetic Analysis and Breeding Applications in Chickpea. Available at: https://pag.confex.com/pag/asia2016/webprogram/Paper22628.html
Russell, J., Mascher, M., Dawson, I. K., Kyriakidis, S., Calixto, C., Freund, F., et al. (2016). Exome sequencing of geographically diverse landraces and wild relatives gives insights into environmental adaptation. Nat. Genet. 48, 1024–1030. doi: 10.1038/ng.3612
Russo, V. E., Martienssen, R. A., and Riggs, A. D. (1996). Epigenetic Mechanisms of Gene Regulation. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Ryder, P., McHale, M., Fort, A., and Spillane, C. (2017). Generation of stable nulliplex autopolyploid lines of Arabidopsis thaliana using CRISPR/Cas9 genome editing. Plant Cell 36, 1005–1008. doi: 10.1007/s00299-017-2125-0
Saïdou, A. A., Mariac, C., Luong, V., Pham, J. L., Bezançon, G., and Vigouroux, Y. (2009). Association studies identify natural variation at the PHYC linked to flowering time and morphological variation in pearl millet. Genetics 182, 899–910. doi: 10.1534/genetics.109.102756
Sailer, C., Schmid, B., and Grossniklaus, U. (2016). Apomixis allows the transgenerational fixation of phenotypes in hybrid plants. Curr. Biol. 26, 331–337. doi: 10.1016/j.cub.2015.12.045
Samach, A., Onouchi, H., Gold, S. E., Ditta, G. S., Schwarz-Sommer, Z., Yanofsky, M. F., et al. (2000). Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288, 1613–1616. doi: 10.1126/science.288.5471.1613
Samal, R., Reddy, J. N., Rao, G. J. N., Roy, P. S., Subudhi, H. N., and Pani, D. R. (2014). Haplotype diversity for Sub1QTL associated with submergence tolerance in rice landraces of Sundarban region (West Bengal) of India. J. Exp. Biol. Agric. Sci. 2, 315–322.
Sasaki, A., Ashikari, M., Ueguchi-Tanaka, M., Itoh, H., Nishimura, A., Swapan, D., et al. (2002). Green revolution: a mutant gibberellin synthesis gene in rice. Nature 416, 701–702. doi: 10.1038/416701a
Schatz, M. C., Maron, L. G., Stein, J. C., Hernandez, W. A., Gurtowski, J., Biggers, E., et al. (2014). Whole genome de novo assemblies of three divergent strains of rice, Oryza sativa, document novel gene space of aus and indica. Genome Biol. 15, 506. doi: 10.1186/s13059-014-0506-z
Scheben, A., Batley, J., and Edwards, D. (2017a). Genotyping-by-sequencing approaches to characterize crop genomes: choosing the right tool for the right application. Plant Biotechnol. J. 15, 149–161. doi: 10.1111/pbi.12645
Scheben, A., Wolter, F., Batley, J., Puchta, H., and Edwards, D. (2017b). Towards CRISPR/Cas crops – bringing together genomics and genome editing. New Phytol. doi: 10.1111/nph.14702 [Epub ahead of print].
Scheben, A., and Edwards, D. (2017). Genome editors take on crops. Science 355, 1122–1123. doi: 10.1126/science.aal4680
Schmitz, R. J., Schultz, M. D., Lewsey, M. G., O’Malley, R. C., Urich, M. A., Libiger, O., et al. (2011). Transgenerational epigenetic instability is a source of novel methylation variants. Science 334, 369–373. doi: 10.1126/science.1212959
Schnable, P. S., and Springer, N. M. (2013). Progress toward understanding heterosis in crop plants. Annu. Rev. Plant Biol. 64, 71–88. doi: 10.1146/annurev-arplant-042110-103827
Sehgal, D., Skot, L., Singh, R., Srivastava, R. K., Das, S. P., Taunk, J., et al. (2015a). Exploring potential of pearl millet germplasm association panel for association mapping of drought tolerance traits. PLoS ONE 10:e0122165. doi: 10.1371/journal.pone.0122165
Sehgal, D., Vikram, P., Sansaloni, C. P., Ortiz, C., Pierre, C. S., Payne, T., et al. (2015b). Exploring and mobilizing the gene bank biodiversity for wheat improvement. PLoS ONE 10:e0132112. doi: 10.1371/journal.pone.013211
Selçuk, A., Forsberg, N., Hagenblad, J., and Leino, M. W. (2015). Molecular genotyping of historical barley landraces reveals novel candidate regions for local adaption. Crop Sci. 55, 2766–2776. doi: 10.2135/cropsci2015.02.0119
Seymour, D. K., and Becker, C. (2017). The causes and consequences of DNA methylome variation in plants. Curr. Opin. Plant Biol. 36, 56–63. doi: 10.1016/j.pbi.2017.01.005
Shen, H., He, H., Li, J., Chen, W., Wang, X., Guo, L., et al. (2012). Genome-wide analysis of DNA methylation and gene expression changes in two Arabidopsis ecotypes and their reciprocal hybrids. Plant Cell 24, 875–892. doi: 10.1105/tpc.111.094870
Shen, L., Shao, N. Y., Liu, X. C., and Nestler, E. (2014). ngs.plot: quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15:284. doi: 10.1186/1471-2164-15-284
Shen, R., Fan, J., Campbell, D., Chang, W., Chen, J., Doucet, D., et al. (2005). High throughput SNP genotyping on universal bead arrays. Mutat. Res. 573, 70–82. doi: 10.1016/j.mrfmmm.2004.07.022
Shi, J., Gao, H., Wang, H., Lafitte, H. R., Archibald, R. L., Yang, M., et al. (2016). ARGOS8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol. J. 15, 207–216. doi: 10.1111/pbi.12603
Shimatani, Z., Kashojiya, S., Takayama, M., Terada, R., Arazoe, T., Ishii, H., et al. (2017). Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 35, 441–443. doi: 10.1038/nbt.3833
Shivaprasad, P. V., Dunn, R. M., Santos, B. A., Bassett, A., and Baulcombe, D. C. (2012). Extraordinary transgressive phenotypes of hybrid tomato are influenced by epigenetics and small silencing RNAs. EMBO J. 31, 257–266. doi: 10.1038/emboj.2011.458
Singh, N., Jayaswal, P. K., Panda, K., Mandal, P., Kumar, V., Singh, B., et al. (2015). Single-copy gene based 50K SNP chip for genetic studies and molecular breeding in rice. Sci. Rep. 5:11600. doi: 10.1038/srep11600
Singh, N., Trang, T. M., Vergara, G. V., Pandey, D. M., Sanchez, D., Neeraja, C. N., et al. (2010). Molecular marker survey and expression analyses of the rice submergence-tolerance gene SUB1A. Theor. Appl. Genet. 121, 1441–1453. doi: 10.1007/s00122-010-1400-z
Sobkowiak, A., Joñzyk, M., Adamczyk, J., Szczpanik, J., Soleck, D., Kuciara, I., et al. (2016). Molecular foundation of chilling tolerance of modern maize. BMC Genomics 17:125. doi: 10.1186/s12864-016-2453-4
Sonah, H., O’Donaughue, L., Cober, E., Rajcan, I., and Belzile, F. (2014). Identification of loci governing eight agronomic traits using a GBS-GWAS approach and validation by QTL mapping in soybean. Plant Biotechnol. J. 13, 211–221. doi: 10.1111/pbi.12249
Song, Q., Hyten, D. L., Jia, G., Quigley, C. V., Fickus, E. W., and Nelson, R. L. (2013). Development and evaluation of SoySNP50K, a high-density genotyping array for soybean. PLoS ONE 8:e54985. doi: 10.1371/journal.pone.0054985
Song, Q., Jia, G., Hyten, D. L., Jenkins, J., Hwang, E. Y., Schroeder, S. G., et al. (2015). SNP assay development for linkage map construction, anchoring whole genome sequence and other genetic and genomic applications in common bean. G3 5, 2285–2290. doi: 10.1534/g3.115.020594
Song, X. J., Kuroha, T., Ayano, M., Furuta, T., Nagai, K., Komeda, N., et al. (2015). Rare allele of a previously unidentified histone H4 acetyltransferase enhances grain weight, yield, and plant biomass in rice. Proc. Natl. Acad. Sci. U.S.A. 112, 76–81. doi: 10.1073/pnas.1421127112
Song, S., Chen, Y., Liu, L., Wang, Y., Bao, S., Zhou, X., et al. (2017). OsFTIP1-mediated regulation of Florigen Transport in rice is negatively regulated by a ubiquitin-like domain kinase OsUbDKgamma4. Plant Cell 29, 491–507. doi: 10.1105/tpc.16.00728
Song, X., and Cao, X. (2017). Transposon-mediated epigenetic regulation contributes to phenotypic diversity and environmental adaptation in rice. Curr. Opin. Plant Biol. 36, 111–118. doi: 10.1016/j.pbi.2017.02.004
Soppe, W. J., Jacobsen, S. E., Alonso-Blanco, C., Jackson, J. P., Kakutani, T., Koornneef, M., et al. (2000). The late flowering phenotype of fwa mutants is caused by gain-of-function epigenetic alleles of a homeodomain gene. Mol. Cell 6, 791–802. doi: 10.1016/S1097-2765(05)00090-0
Soyk, S., Muller, N. A., Park, S. J., Schmalenbach, I., Jiang, K., Hayama, R., et al. (2017). Variation in the flowering gene SELF PRUNING 5G promotes day-neutrality and early yield in tomato. Nat. Genet. 49, 162–168. doi: 10.1038/ng.3733
Sparla, F., Falini, G., Botticella, E., Pirone, C., Talame, V., Bovina, R., et al. (2014). New starch phenotypes produced by TILLING in barley. PLoS ONE 9:e107779. doi: 10.1371/journal.pone.0107779
Spillane, C., Curtis, M. D., and Grossniklaus, U. (2004). Apomixis technology development—virgin births in farmers’ fields? Nat. Biotechnol. 22, 687–691. doi: 10.1038/nbt976
Spindel, J. E., Begum, H., Akdemir, D., Collard, B., Redoña, E., Jannink, J.-L., et al. (2016). Genome-wide prediction models that incorporate de novo GWAS are a powerful new tool for tropical rice improvement. Heredity 116, 395–408. doi: 10.1038/hdy.2015.113
Spindel, J. E., and McCouch, S. R. (2016). When more is better: how data sharing would accelerate genomic selection of crop plants. New Phytol. 212, 814–826. doi: 10.1111/nph.14174
Steinert, J., Schiml, S., and Puchta, H. (2016). Homology-based double-strand break-induced genome engineering in plants. Plant Cell Rep. 35, 1429–1438. doi: 10.1007/s00299-016-1981-3
Stephens, J. C., Miller, F. R., and Rosenow, D. T. (1967). Conversion of alien sorghums to early maturing genotypes. Crop Sci. 7, 396. doi: 10.2135/cropsci1967.0011183x000700040036x
Sun, X. J., Hu, Z., Chen, R., Jiang, Q. Y., Song, G. H., Zhang, H., et al. (2015). Targeted mutagenesis in soybean using the CRISPR-Cas9 system. Sci. Rep. 5, 103–142. doi: 10.1038/srep10342
Takeshima, R., Hayashi, T., Zhu, J., Zhao, C., Xu, M., Yamaguchi, N., et al. (2016). A soybean quantitative trait locus that promotes flowering under long days is identified as FT5a, a FLOWERING LOCUS T ortholog. J. Exp. Bot. 67, 5247–5258. doi: 10.1093/jxb/erw283
Talamé, V., Bovina, R., Sanguineti, M. C., Tuberosa, R., Lundqvist, U., and Salvi, S. (2008). TILLMore, a resource for the discovery of chemically induced mutants in barley. Plant Biotechnol. J. 6, 477–485. doi: 10.1111/j.1467-7652.2008.00341.x
Tamang, B. G., Niroula, R. K., Amgai, R. B., Sad, B. P., and Ghimire, S. K. (2011). Determination of adaptive mechanisms for flash flooding tolerance in Nepalese cultivated rice genepool based on morpho-physiological and molecular analysis. Asian J. Plant Sci. 10, 347–356. doi: 10.3923/ajps.2011.347.356
Tayeh, N., Aluome, C., Falque, M., Jackuin, F., Klein, A., Chauveau, A., et al. (2015). Development of two major resources for pea genomics: the GenoPea 13.2K SNP Array and a high-density, high-resolution consensus genetic map. Plant J. 84, 1257–73. doi: 10.1111/tpj.13070
Teixeira, F. K., Heredia, F., Sarazin, A., Roudier, F., Boccara, M., Ciaudo, C., et al. (2009). A role for RNAi in the selective correction of DNA methylation defects. Science 323, 1600–1604. doi: 10.1126/science.1165313
Tello-Ruiz, M. K., Stein, J., Wei, S., Preece, J., Olson, A., Naithani, S., et al. (2016). Gramene 2016: comparative plant genomics and pathway resources. Nucleic Acids Res. 44, D1133–D1140. doi: 10.1093/nar/gkv1179
Tenaillon, M. I., Sawkins, M. C., Long, A. D., Gaut, R. L., Doebley, J. F., and Gaut, B. S. (2001). Patterns of DNA sequence polymorphism along chromosome 1 of maize (Zea mays ssp. mays L.). Proc. Natl. Acad. Sci. U.S.A. 98, 9161–9166. doi: 10.1073/pnas.151244298
Thirunavukkarasu, N., Hossain, F., Shiriga, K., Mittal, S., Arora, K., Rathore, A., et al. (2017). Unraveling the genetic architecture of subtropical maize (Zea mays L.) lines to assess their utility in breeding programs. BMC Genomics 14:877. doi: 10.1186/1471-2164-14-877
Thoen, M. P. M., Olivas, N. H. D., Kloth, K. J., Coolen, S., Huang, P.-P., Aarts, M. G., et al. (2016). Genetic architecture of plant stress resistance: multi-trait genome-wide association mapping. New Phytol. 213, 1346–1362. doi: 10.1111/nph.14220
Thudi, M., Khan, A. W., Kumar, V., Gaur, P. M., Katta, K., Garg, V., et al. (2016). Whole genome re-sequencing reveals genome-wide variations among parental lines of 16 mapping populations in chickpea (Cicer arietinum L.). BMC Plant Biol. 16(Suppl. 1):10. doi: 10.1186/s12870-015-0690-3
Thurber, C. S., Ma, J. M., Higgins, R. H., and Brown, P. J. (2013). Retrospective genomic analysis of sorghum adaptation to temperate-zone grain production. Genome Biol. 14:R68. doi: 10.1186/gb-2013-14-6-r68
Tian, F., Bradbury, P. J., Brown, P. J., Hung, H., Sun, Q., Flint-Garcia, S., et al. (2011). Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat. Genet. 43, 159–162. doi: 10.1038/ng.746
Tian, H.-L., Wang, F.-G., Zhao, J.-R., Yi, H.-M., Wang, L., Wang, R., et al. (2015). Development of maizeSNP3072, a high-throughput compatible SNP array, for DNA fingerprinting identification of Chinese maize varieties. Mol. Breed. 35, 136. doi: 10.1007/s11032-015-0335-0
Till, B. J., Reynolds, S. H., Weil, C., Springer, N., Burtner, C., Young, K., et al. (2004). Discovery of induced point mutations in maize genes by TILLING. BMC Plant Biol. 4:12. doi: 10.1186/1471-2229-4-12
Tilman, D., Balzer, C., Hill, J., and Befort, B. L. (2011). Global food demand and the sustainable intensification of agriculture. Proc. Natl. Acad. Sci. U.S.A. 108, 20260–20264. doi: 10.1073/pnas.1116437108
Trethowan, R. M., Reynolds, M. P., Ortiz-Monasterio, I., and Ortiz, R. (2007). The genetic basis of the Green Revolution in wheat production. Plant Breed. Rev. 28, 39–58. doi: 10.1002/9780470168028.ch2
Tsai, H., Howell, T., Nitcher, R., Missirian, V., Watson, B., Ngo, K. J., et al. (2011). Discovery of rare mutations in populations: TILLING by sequencing. Plant Physiol. 156, 1257–1268. doi: 10.1104/pp.110.169748
Tsubokura, Y., Watanabe, S., Xia, Z., Kanamori, H., Yamagata, H., Kaga, A., et al. (2014). Natural variation in genes responsible for maturity loci E1, E2, E3, and E4 in soybean. Ann. Bot. 113, 429–441. doi: 10.1093/aob/mct269
Tucker, S., Vitins, A., and Pikaard, C. S. (2010). Nucleolar dominance and ribosomal RNA gene silencing. Curr. Opin. Cell Biol. 22, 351–356. doi: 10.1016/j.ceb.2010.03.009
Turck, F., Fornara, F., and Coupland, G. (2008). Regulation and identity of florigen: FLOWERING LOCUS T moves center stage. Annu. Rev. Plant Biol. 59, 573–594. doi: 10.1146/annurev.arplant.59.032607.092755
Turner, A., Beales, J., Faure, S., Dunford, R. P., and Laurie, D. A. (2005). The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310, 1031–1034. doi: 10.1126/science.1117619
Unterseer, S., Bauer, E., Haberer, G., Seidel, M., Knaak, C., Ouzonova, M., et al. (2014). A powerful tool for genome analysis in maize: development and evaluation of the high density 600 K SNP genotyping array. BMC Genomics 15:823. doi: 10.1186/1471-2164-15-823
Unterseer, S., Pophaly, S. D., Peis, R., Westermeier, P., Mayer, M., Seidel, M. A., et al. (2016). A comprehensive study of the genomic differentiation between temperate dent and flint maize. Genome Biol. 17, 137. doi: 10.1186/s13059-016-1009-x
Valdisser, P. A. M. R., Pereira, W. J., Filho, J. E., Müller, B. S. F., Coelho, G. R. C., de Menezes, I. P. P., et al. (2017). In-depth genome characterization of a Brazilian common bean core collection using DArTseq high-density SNP genotyping. BMC Genomics 18:423. doi: 10.1186/s12864-017-3805-4
Valliyodan, B. V., Qiu, D., Patil, G., Zeng, P., Huang, J., Dai, L., et al. (2016). Landscape of genomic diversity and trait discovery in soybean. Sci. Rep. 6:23598. doi: 10.1038/srep23598
Valluru, R., Reynolds, M. P., Davies, W. J., and Sukumaran, S. (2017). Phenotypic and genome-wide association analysis of spike ethylene in diverse wheat genotypes under heat stress. New Phytol. 214, 271–283. doi: 10.1111/nph.14367
van Hintum, T. J. L., Brown, A. H. D., Spillane, C., and Hodgkin, T. (2000). Core Collections of Plant Genetic Resources. IPGRI Technical Bulletin No. 3. Rome: IPGRI, 48.
Varshney, R. K., Gaur, P. M., Chamarthi, S. K., Krishnamurthy, L., Tripathi, S., Kashiwagi, J., et al. (2013). Fast-track introgression of ‘QTL-hotspot’ for root traits and other drought tolerance traits in J11, an elite and leading variety of chickpea. Plant Genome 6, 1–9. doi: 10.3835/plantgenome2013.07.0022
Varshney, R. K., Thudi, M., Nayak, S. N., Gaur, P. M., Kashiwagi, J., Krishnamurthy, L., et al. (2014). Genetic dissection of drought tolerance in chickpea (Cicer arietinum L.). Theor. Appl. Genet. 127, 445–462. doi: 10.1007/s00122-013-2230-6
Vigouroux, Y., Mariac, C., De Mita, S., Pham, J. L., Gérard, B., Kapran, I., et al. (2011). Selection for earlier flowering crop associated with climatic variations in Sahel. PLoS ONE 6:e19563. doi: 10.1371/journal.pone.0019563
Vikram, P., Franco, J., Burgueño-Ferreira, J., Li, H., Sehgal, D., Pierre, C. S., et al. (2016). Unlocking the genetic diversity of Creole wheats. Sci. Rep. 6:23092. doi: 10.1038/srep23092
Villordo-Pineda, E., González-Chavira, M. M., Giraldo-Carbajo, P., Acosta-Gallegos, J. A., and Caballero-Pérez, J. (2015). Identification of novel drought-tolerant-associated SNPs in common bean (Phaseolus vulgaris). Front. Plant Sci. 6:546. doi: 10.3389/fpls.2015.00546
Voytas, D. F. (2013). Plant genome engineering with sequence-specific nucleases. Annu. Rev. Plant Biol. 64, 327–350. doi: 10.1146/annurev-arplant-042811-105552
Wang, J., Chu, S., Zhang, H., Zhu, Y., Cheng, H., Yu, D., et al. (2016). Development and application of a novel genome-wide SNP array reveals domestication history in soybean. Sci. Rep. 6:20728. doi: 10.1038/srep20728
Wang, F., Wang, C. L., Liu, P. Q., Lei, C. L., Hao, W., Gao, Y., et al. (2016). Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS ONE 11:e0154027. doi: 10.1371/journal.pone.0154027
Wang, L., Greaves, I. K., Groszmann, M., Wu, L. M., Dennis, E. S., and Peacock, W. J. (2015). Hybrid mimics and hybrid vigor in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 112, E4959–E4967. doi: 10.1073/pnas.1514190112
Wang, Y., Tan, L., Fu, Y., Zhu, Z., Liu, F., Sun, C., et al. (2015). Molecular evolution of the sorghum maturity gene Ma3. PLoS ONE 10:e0124435. doi: 10.1371/journal.pone.0124435
Wang, S., Wong, D., Forrest, K., Allen, A., Chao, S., Huang, B. E., et al. (2014). Characterization of polyploid wheat genomic diversity using a high-density 90,000 single nucleotide polymorphism array. Plant Biotechnol. J. 12, 787–796. doi: 10.1111/pbi.12183
Wang, Y., Cheng, X., Shan, Q. W., Zhang, Y., Liu, J. X., Gao, C. X., et al. (2014). Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951. doi: 10.1038/nbt.2969
Weng, J., Gu, S., Wan, X., Gao, H., Guo, T., Su, N., et al. (2008). Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res. 18, 1199–1209. doi: 10.1038/cr.2008.307
Westengen, O. T., Berg, P. R., Kent, M. P., and Brysting, A. K. (2012). Spatial structure and climatic adaptation in African maize revealed by surveying SNP diversity in relation to global breeding and landrace panels. PLoS ONE 7:e47832. doi: 10.1371/journal.pone.0047832
Wissuwa, M., Kondo, K., Fukuda, T., Mori, A., Rose, M. T., Pariasca-Tanaka, J., et al. (2015). Unmasking novel loci for internal phosphorus utilization efficiency in rice germplasm through genome-wide association analysis. PLoS ONE 10:e0124215. doi: 10.1371/journal.pone.0124215
Wissuwa, M., Wegner, J., Ae, N., and Yano, M. (2002). Substitution mapping of the Pup1: a major QTL increasing phosphorus uptake of rice from a phosphorus deficient soil. Theor. Appl. Genet. 105, 890–897. doi: 10.1007/s00122-002-1051-9
Xiong, H., Shi, A., Mou, B., Qin, J., Motes, D., Lu, W., et al. (2016). Genetic diversity and population structure of cowpea (Vigna unguiculata L. Walp). PLoS ONE 11:e0160941. doi: 10.1371/journal.pone.0160941
Xu, C., Ren, Y., Jian, Y., Guo, Z., Zhang, Y., Xie, C., et al. (2017). Development of a maize 55 K SNP array with improved genome coverage for molecular breeding. Mol. Breed. 37, 20. doi: 10.1007/s11032-017-0622-z
Xu, J., Yuan, Y., Xu, Y., Zhang, G., Guo, X., Wu, F., et al. (2014). Identification of candidate genes for drought tolerance by whole-genome resequencing in maize. BMC Plant Biol. 14:83. doi: 10.1186/1471-2229-14-83
Xu, M., Xu, Z., Liu, B., Kong, F., Tsubokura, Y., Watanabe, S., et al. (2013). Genetic variation in four maturity genes affects photoperiod sensitivity and PHYA-regulated post-flowering responses of soybean. BMC Plant Biol. 13:91. doi: 10.1186/1471-2229-13-91
Xu, P., Wu, X., Muñoz-Amatriaín, M., Wang, B., Wu, X., Hu, Y., et al. (2017). Genomic regions, cellular components and gene regulatory basis underlying pod length variations in cowpea (V. unguiculata L. Walp). Plant Biotechnol. J. 15, 547–557. doi: 10.1111/pbi.12639
Xu, X., Liu, X., Ge, S., Jensen, J. D., Hu, F., Li, X., et al. (2012). Resequencing 50 accessions of cultivated and wild rice yield markers for identifying agronomically important genes. Nat. Biotechnol. 30, 105–111. doi: 10.1038/nbt.2050
Xu, Y., Jia, Q., Zhou, G., Zhang, X. Q., Angessa, T., Broughton, S., et al. (2017). Characterization of the sdw1 semi-dwarf gene in barley. BMC Plant Biol. 7:11. doi: 10.1186/s12870-016-0964-4
Xue, W., Xing, Y., Weng, X., Zhao, Y., Tang, W., Wang, L., et al. (2008). Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 40, 761–767. doi: 10.1038/ng.143
Ya-fang, Z., Yu-yin, M. A., Zong-xiang, C., Jie, Z., Tian-xiao, C., Qian-qian, L., et al. (2015). Genome-wide association studies reveal new genetic targets for five panicle traits of international rice varieties. Rice Sci. 22, 217–226. doi: 10.1016/j.rsci.2015.07.001
Yamaguchi, M., Fujimoto, H., Hirano, K., Araki-Nakamura, S., Ohmae-Shinohara, K., Fujii, A., et al. (2016). Sorghum Dw1, an agronomically important gene for lodging resistance, encodes a novel protein involved in cell proliferation. Sci. Rep. 6:28366. doi: 10.1038/srep28366
Yan, J. B., Kandianis, C. B., Harjes, C. E., Bai, L., Kim, E. H., Yang, X. H., et al. (2010). Rare genetic variation at Zea mays crtRB1 increases beta-carotene in maize grain. Nat. Genet. 42, 322–327. doi: 10.1038/ng.551
Yan, W., Liu, H., Zhou, X., Li, Q., Zhang, J., Liu, T., et al. (2013). Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res. 23, 969–971. doi: 10.1038/cr.2013.43
Yang, W., Guo, Z., Huang, C., Duan, L., Chen, G., Jiang, N., et al. (2014). Combining high throughput phenotyping and genome-wide association studies to reveal natural genetic variation in rice. Nat. Commun. 5:5087. doi: 10.1038/ncomm6087s
Yang, X., Kundariya, H., Xu, Y.-Z., Sandhu, A., Yu, J., Hutton, S. F., et al. (2015). MutS HOMOLOG1-derived epigenetic breeding potential in tomato. Plant Physiol. 168, 222–232. doi: 10.1104/pp.15.00075
Yu, H., Xie, W., Li, J., Zhou, F., and Zhang, Q. (2014). A whole-genome SNP array (RICE6K) for genomic breeding in rice. Plant Biotechnol. J. 12, 28–37. doi: 10.1111/pbi.12113
Yu, L. X., Liu, S., Anderson, J. A., Singh, R. P., Jin, Y., Dubcovsky, J., et al. (2010). Haplotype diversity of stem rust resistance loci in uncharacterized wheat lines. Mol. Breed. 26, 667–680. doi: 10.1007/s11032-010-9403-7
Yu, X., Li, X., Guo, T., Zhu, C., Wu, Y., Mitchell, S. E., et al. (2016). Genomic prediction contributing to a promising global strategy to turbocharge gene banks. Nat. Plants 2:16150. doi: 10.1038/NPLANTS.2016.150
Zhang, H. L., Harry, D. E., Ma, C., Yuceer, C., Hsu, C. Y., Vikram, V., et al. (2010). Precocious flowering in trees: the FLOWERING LOCUS T gene as a research and breeding tool in Populus. J. Exp. Bot. 61, 2549–2560. doi: 10.1093/jxb/erq092
Zhang, J., Song, Q., Cregan, P. B., Nelson, R. L., Wang, X., Wu, J., et al. (2015). Genome-wide association study for flowering time, maturity dates and plant height in early maturing soybean (Glycine max) germplasm. BMC Genomics 16:217. doi: 10.1186/s12864-015-1441-4
Zhang, X., Sun, J., Cao, X., and Song, X. (2015). Epigenetic mutation of RAV6 affects leaf angle and seed size in rice. Plant Physiol. 169, 2118–2128. doi: 10.1104/pp.15.00836
Zhang, Q., Li, Y., Xu, T., Srivastava, A. K., Wang, D., Zeng, L., et al. (2016a). The chromatin remodeler DDM1 promotes hybrid vigor by regulating salicylic acid metabolism. Cell Discov. 2:16027. doi: 10.1038/celldisc.2016.27
Zhang, Q., Wang, D., Lang, Z., He, L., Yang, L., Zeng, L., et al. (2016b). Methylation interactions in Arabidopsis hybrids require RNA-directed DNA methylation and are influenced by genetic variation. Proc. Natl. Acad. Sci. U.S.A. 113, E4248–E4256. doi: 10.1073/pnas.1607851113
Zhang, D., Li, H., Wang, J., Zhang, H., Hu, Z., Chu, S., et al. (2016c). High-density genetic mapping identifies new major loci for tolerance to low-phosphorus stress in soybean. Front. Plant Sci. 7:372. doi: 10.3389/fpls.2016.00372
Zhang, X. J., Wang, J. F., Huan, G. J., Lan, H. X., Wang, C. L., Yin, C. F., et al. (2012). Rare allele of OsPPKL1 associated with grain length causes extra-large grain and a significant yield increase in rice. Proc. Natl. Acad. Sci. U.S.A. 109, 21534–21539. doi: 10.1073/pnas.1219776110
Zhang, Z., Li, J., Pan, Y., Li, J., Zhou, L., Shi, H., et al. (2017). Natural variation in CTB4a enhances rice adaptation to cold habitats. Nat. Commun. 8:14788. doi: 10.1038/ncomms14788
Zhao, J., Montilla Perez, M. B., Hu, J., and Salas Fernandez, M. G. (2016). Genome-wide association study for nine plant architecture traits. Plant Genome 9. doi: 10.3835/plantgenome2015.06.0044
Zhao, M., Sun, J., Xiao, Z., Cheng, F., Xu, H., Tang, L., et al. (2016). Variations in DENSE AND ERECT PANICLE1 (DEP1) contribute to the diversity of the panicle trait in high yielding japonica rice varieties in northern China. Breed. Sci. 66, 599–605. doi: 10.1270/jsbbs.160.58
Zhao, C., Takeshima, R., Zhu, J., Xu, M., Sato, M., Watanabe, S., et al. (2016). A recessive allele for delayed flowering at the soybean maturity locus E9 is a leaky allele of F2a, a FLOWERING LOCUS T ortholog. BMC Plant Biol. 16:20. doi: 10.1186/s12870-016-0704-9
Zhao, K., Tung, C.-W., Eizenga, G. C., Wright, M.-H., Ali, M. L., Price, A. H., et al. (2011). Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. 2:467. doi: 10.1038/ncomms1467
Zheng, L. Y., Guo, X.-S., He, B., Sun, L.-J., Peng, Y., and Domg, S.-S. (2011). Genome-wide pattern of variation in sweet and grain sorghum (Sorghum bicolor). Genome Biol. 12:R114. doi: 10.1186/gb-2011-12-11-r114
Zhong, S., Fei, Z., Chen, Y.-R., Zheng, Y., Huang, M., Vrebalov, J., et al. (2013). Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening. Nat. Biotechnol. 31, 154–159. doi: 10.1038/nbt.2462
Zhou, Z., Jiang, Y., Wang, Z., Gou, Z., Lyu, J., Li, W., et al. (2015). Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat. Biotechnol. 33, 408–414. doi: 10.1038/nbt.3096
Keywords: crop improvement, epigenetic variation, florigen pathways, functional diversity, genome editing, genomic estimated breeding value, haplotypes, TILLING
Citation: Dwivedi SL, Scheben A, Edwards D, Spillane C and Ortiz R (2017) Assessing and Exploiting Functional Diversity in Germplasm Pools to Enhance Abiotic Stress Adaptation and Yield in Cereals and Food Legumes. Front. Plant Sci. 8:1461. doi: 10.3389/fpls.2017.01461
Received: 21 April 2017; Accepted: 07 August 2017;
Published: 29 August 2017.
Edited by:
Jaime Prohens, Universitat Politècnica de València, SpainReviewed by:
Karl Kunert, University of Pretoria, South AfricaUday Chand Jha, Indian Institute of Pulses Research, India
Filippo Biscarini, Consiglio Nazionale Delle Ricerche (CNR), Italy
Copyright © 2017 Dwivedi, Scheben, Edwards, Spillane and Ortiz. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Rodomiro Ortiz, cm9kb21pcm8ub3J0aXpAc2x1LnNl
 Sangam L. Dwivedi
Sangam L. Dwivedi Armin Scheben
Armin Scheben David Edwards2
David Edwards2 Charles Spillane
Charles Spillane Rodomiro Ortiz
Rodomiro Ortiz

