- 1Crop Research Institute, Shandong Academy of Agricultural Sciences/Key Laboratory of Wheat Biology and Genetic Improvement in the Northern Yellow-Huai Rivers Valley of Ministry of Agriculture/National Engineering Laboratory for Wheat and Maize, Jinan, China
- 2College of Plant Protection, Agricultural University of Hebei, Baoding, China
- 3College of Life Science, Shandong Normal University, Jinan, China
Aegilops caudata is an important gene source for wheat breeding. Intensive evaluation of its utilization value is an essential first step prior to its application in breeding. In this research, the agronomical and quality traits of Triticum aestivum-Ae. caudata additions B–G (homoeologous groups not identified) were analyzed and evaluated. Disease resistance tests showed that chromosome D of Ae. caudata might possess leaf rust resistance, and chromosome E might carry stem rust and powdery mildew resistance genes. Investigations into agronomical traits suggested that the introduction of the Ae. caudata chromosome in addition line F could reduce plant height. Grain quality tests showed that the introduction of chromosomes E or F into wheat could increase its protein and wet gluten content. Therefore, wheat-Ae. caudata additions D–F are all potentially useful candidates for chromosome engineering activities to create useful wheat-alien chromosome introgressions. A total of 55 EST-based molecular markers were developed and then used to identify the chromosome homoeologous group of each of the Ae. caudata B–G chromosomes. Marker analysis indicated that the Ae. caudata chromosomes in addition lines B to G were structurally altered, therefore, a large population combined with intensive screening pressure should be taken into consideration when inducing and screening for wheat-Ae. caudata compensating translocations. Marker data also indicated that the Ae. caudata chromosomes in addition lines C–F were 5C, 6C, 7C, and 3C, respectively, while the homoeologous group of chromosomes B and G of Ae. caudata are as yet undetermined and need further research.
Introduction
Aegilops caudata L. [syn. Ae. markgrafii (Greuter) Hammer] is an annual diploid species (2n = 2x = 14, genome CC), naturally occurring mainly in the Aegean Region and in western Turkey, less commonly and more sporadically in inland Turkey and through the Fertile Crescent (Kilian et al., 2011). Ae. caudata can form dense stands, often together with other Aegilops species. Collected annual rainfall data indicate a range of 300–700 mm. From sea level up to 1,850 m (Kilian et al., 2011). Ae. caudata has been found to be resistant to wheat stripe rust (Puccinia striiformis Westend) (Valkoun et al., 1985; Baldauf et al., 1992; Toor et al., 2016), leaf rust (P. recondita Roberge ex Desmaz. f. sp. tritici) (Gill et al., 1985; Valkoun et al., 1985; Iqbal et al., 2007; Riar et al., 2012), stem rust (Puccinia graminis f. sp. tritici) (Valkoun et al., 1985; Dyck et al., 1990), powdery mildew (Blumeria graminis f. sp. tritici) (Gill et al., 1985; Valkoun et al., 1985; Baldauf et al., 1992), barley yellow dwarf luteovirus (Makkouk et al., 1994), snow mold (Typhula ishikariensis S. Imai) (Iriki et al., 2001), greenbug [Schizaphis graminum (Rondani)] (Baldauf et al., 1992) and hessian fly [Mayetiola destructor (Say)] (Gill et al., 1985). Moreover, some species of Ae. caudata has freezing tolerance (Barashkova and Migushova, 1984; Iriki et al., 2001), salt tolerance (Gorham, 1990), and could be used for iron and zinc fortification (Wang et al., 2011). Therefore, Ae. caudata is an excellent gene source for wheat improvement.
Wheat-Ae. caudata amphiploids, addition, substitution and translocation lines are bridging materials for transferring desirable genes from Ae. caudata to wheat. The creation and identification of these bridge materials is the first step in the gene transfer procedure. Muramatsu (1973) produced and identified a bread wheat-Ae. caudata 5C (5A,5D) substitution. Biithner et al. (1988) created a set of bread wheat (cv. Alcedo)-Ae. caudata addition lines, temporarily named as additions A to G (Schubert and Bluthner, 1995). Later, Friebe et al. (1992) described the C-banded karyotype of this set of addition lines except addition A. Latter, Molnár et al. (2016) studied the homoeologous relationships of flow sorted wheat and Ae. caudata chromosomes using COS markers using this set of additions. More recently, Danilova et al. (2017) used single gene FISH and exome capture sequencing approaches and revised the nomenclature of Ae. caudata chromosomes A, B, C, D, E, F and G to 1C, 2C, 5C, 6C, 7C, 3C, and 4C, respectively. Moreover, Kong et al. (1999a) also synthesized a Triticum durum-Ae. caudata amphiploid, and identified chromosome translocations within the backcrossed progenies between wheat and the T. durum-Ae. caudata amphiploid. However, less useful markers for C chromosome were available in transferring genes from Ae. caudata into wheat.
Comprehensive evaluation of the disease resistance status, agronomical characters and quality traits of wheat-Ae. caudata addition lines will provide useful background information for future research to create useful wheat-Ae. caudata chromosome translocations for wheat breeding programs. Although the wheat-Ae. caudata addition lines A–G have been produced and identified (Schubert and Bluthner, 1995), the breeding value of this set of material has not yet been evaluated which stymied the creation and utilization of compensating wheat-Ae. caudata translocations. In this research, the level of disease resistance, agronomical characteristics and quality traits of wheat-Ae. caudata addition lines B–G (addition A was not available) were investigated or measured. Moreover, EST-based molecular markers specific for Ae. caudata chromosomes were developed to identify the homoeologous group of Ae. caudata chromosomes.
Materials and Methods
Plant Material
Triticum aestivum cv. Alcedo (ALCD), ALCD-Ae. caudata additions B–E (TA3558-TA3561), and G (TA3563) (Schubert and Bluthner, 1995) were provided by Prof. WX Liu, College of Life Science, Henan Agricultural University. Ae. caudata (TA1908), ALCD-Ae. caudata additions F (TA3562) (Schubert and Bluthner, 1995) and T. turgidum (TA10543) were provided by Prof. BS Gill, Wheat Genetic and Genomic Resource Center, Kansas State University. T. aestivum cv. Chinese Spring (CS), Mianyang11 (MY11) and Mianyang15 (MY15) were provided by Prof. ZJ Yang, School of Life Science and Technology, University of Electronic Science and Technology of China.
Disease Resistance Testing
Stripe rust, leaf rust, stem rust and powdery mildew resistances of 20 individual plants of each of ALCD-Ae. caudata additions B–G, ALCD, TA1908, CS, MY11, and MY15 were tested. Among these lines, CS, MY11, and MY15 are highly susceptible to all four diseases, hence the disease response scoring did not begin until these three control genotypes were fully infected. The disease responses were scored on a 0–4 rating scale according to Wang et al. (2014), whereas 0 indicates immune, 0; means nearly immune but showing a small fleck on the leaf, 1 indicates highly resistant, 2 means moderately resistant, 3 indicates moderately susceptible, and 4 means highly susceptible. Record disease resistance levels (DRL) of 20 individual plants of each material truthfully, if there are resistance segregations, for example, the DRL of some plants are 1, some are 3, record as 1, 3. If the DRL of all 20 individual plants are completely same, just record only one DRL value.
The pathogen inoculation methods of stripe rust, leaf rust and powdery mildew were according to Liu et al. (2013), while stem rust inoculation was according to Wu et al. (2014). Stripe rust resistance was determined on both seedlings and adult plants using isolates of races CY32, CY33, and Su-4 in the experimental farmland of School of Life Science and Technology, University of Electronic Science and Technology of China. Stem rust resistance was determined on seedlings using mixed isolates of 34MKGQM and 21C3CTHSM in the greenhouse of College of Plant Protection, Shenyang Agricultural University. Leaf rust resistance was determined on seedlings using mixed leaf rust isolates of THTT, PHTT, THKS, THTS, and THKT in the greenhouse of College of Plant Protection, Agricultural University of Hebei. Powdery mildew resistance was determined on both seedlings (in greenhouse) and adult plants (field) following inoculation with mixed powdery mildew races collected from four different cities including Jinan, Linyi, Dezhou and Heze of Shandong Province.
Agronomical Trait Investigation and Quality Measurement
ALCD and ALCD-Ae. caudata addition lines B–G were planted in the farmland at four different cities including Jinan, Dezhou, Heze and Linyi of Shandong Province on October 25, 2015. The experimental design consisted of three biological replications arranged in a randomized block, and all plots and sites followed the same standard cultivation practices and were grown under irrigated conditions. Spacing between plants in a row was 20 cm and the between row spacing was 33 cm. Each experimental plot contains seven rows. A border buffer consisting of 18 rows of wheat variety Jimai22 surrounded the experimental plots so as to eliminate the margin effect. The four cities were in a temperate continental monsoon climate, characterized by dry, cold winters and rainy, hot summers. During wheat growing season, total precipitation in 2016 was 264.7 mm in Jinan, 176.3 mm in Dezhou, 279.4 mm in Heze, and 339.6 mm in Linyi, respectively. Averaged temperatures in 2016 growing season were 11.5°C in Jinan, 9.7°C in Dezhou, 10.5°C in Heze and 11.0°C in Linyi, respectively. Soil types of all four stations were fluvo-aquic soil, same amount of compound fertilizer was used, weeds and diseases were controlled. The climate information was obtained from the official website of Shandong Meteorological Bureau. The soil type information of four cities was obtained from the official website of Soil and Fertilizer Station, Shandong Provincial Department of Agriculture.
Randomly select 10 plants of each material for the measurement of plant height, spike length (the selected spikes were painted by red lacquer using a manual spray painting pot), flag leaf length and width, tiller number, spikelet number before leaf rolling or shrinking on May, 2016. Randomly harvest one spike of each individual plants (the 10 spikes painted by red lacquer were included) after they were fully mature in June, 2016. Spikes were threshed manually to prevent seed loss so as to determine grain number of the 30 spikes and thousand-kernel weight. Data on the number of tillers, grain number of 30 spikes and thousand-kernel weight from Jinan was not obtainable.
Grain samples for quality tests were collected from four cities of Shandong Province as mentioned above. The grains were milled using wheat grinding machine 3100 (Perten, Sweden), the protein content was measured with a near-infrared (NIR) spectrometer DA7200 (Perten, Sweden) according to the approved method 46-12 (AACC, 2000), three replications for each sites. The wet gluten content was measured with a gluten tester 2200 (Perten, Sweden), three replications for each sites. Data processing and t-test was performed by Microsoft Excel 2010 and the statistical software SPSS v. 13.0. The data from four sites were completely consistent with each other (tiller number, grain number of 30 spikes and thousand-kernel weight, across the three cities), the trait variation when compared to the background genotype ALCD will be regarded as attributable to the presence of the alien chromatin. Alternatively, it might be considered as a result of interaction of genotype and environments. In this research, only the former will be discussed.
DNA Isolation, Primer Design and PCR
Total genomic DNA was prepared from young leaves using the SDS protocol (Liu et al., 2006). A total of 410 bin mapped Expressed Sequence Tags (ESTs) were selected from the wheat EST mapping project (http://wheat.pw.usda.gov/NSF/data.html) for EST-Sequence Tagged Site (EST-STS) primer design using the software Primer 3 (http://frodo.wi.mit.edu). EST-STS PCR amplifications were performed as described by Gong et al. (2014). To obtain higher levels of polymorphism, the PCR products were digested with the 4-base cutter enzymes AluI, HaeIII, MspI or RsaI. The PCR products were separated on a 2% agarose gel.
A total of 258, 107 and 185 EST-Simple Sequence Repeat (EST-SSR), Conserved Orthologous Sequence (COS) and PCR-based Landmark Unique Gene (PLUG) primers were selected and synthesized, and PCR protocol were followed that according to Xue et al. (2008), Quraishi et al. (2009), and Ishikawa et al. (2007), respectively. To obtain high levels of polymorphism, the PLUG PCR products were digested with the four-base cutter enzymes HaeIII or TaqI according to Ishikawa et al. (2007), whereas the COS and EST-SSR PCR products were separated on a native polyacrylamide gel electrophoresis and stained in a silver solution according to Xue et al. (2008) and Quraishi et al. (2009). The molecular markers specific for Ae. caudata chomosomes were determined using Ae. caudata accession TA1908, ALCD-Ae. caudata addition lines TA3598-TA3563 as positive control and wheat genotypes Alcedo, CS, T. turgidum accession TA10543, MY11 and MY15 as negative control.
Results
Disease Resistance Tests of ALCD-Ae. caudata Additions
In this research, wheat stripe rust, leaf rust, stem rust and powdery mildew resistance of Ae. caudata, the ALCD- Ae. caudata B–G addition lines, and wheat controls ALCD, CS, MY11, and MY15 were tested (Table 1). The results showed that CS, MY11 and MY15 were highly susceptible to all four diseases, indicating that the infection races were fully inoculated. Ae. caudata, ALCD and ALCD-Ae. caudata B–G additions were nearly immune or highly resistant to stripe rust at the seedling and adult plant stages, suggesting that there is at least one stripe rust resistant gene in the six additions which was derived from the wheat line ALCD. Ae. caudata and the ALCD-Ae. caudata D addition line were highly resistant to leaf rust, while ALCD and other five additions were susceptible, indicating that D chromosome of Ae. caudata might carry a leaf rust resistance gene. Ae. caudata and the ALCD-Ae. caudata E addition line were highly resistant to stem rust and powdery mildew, while ALCD and other five additions were susceptible, indicating that E chromosome of Ae. caudata might possess stem rust and powdery mildew resistant gene (s).
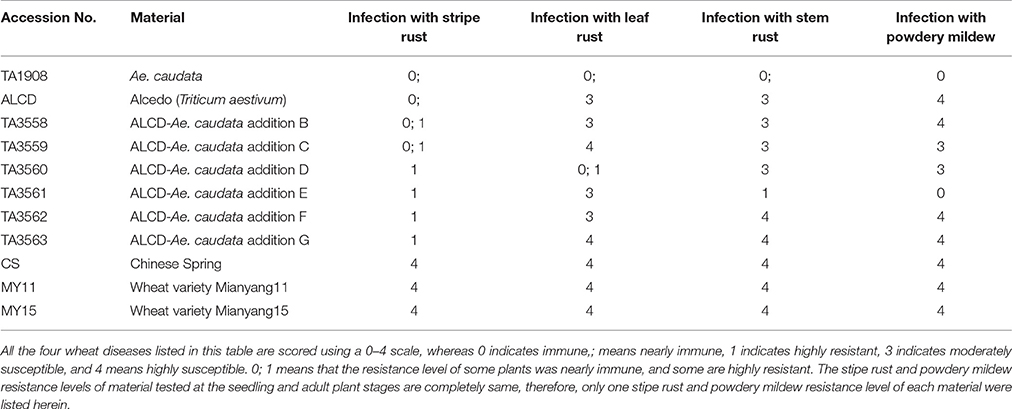
Table 1. Stripe rust, leaf rust, stem rust and powdery mildew resistances of ALCD-Ae. caudata additions.
Agronomic Trait Investigation
Plant height, spike length, spikelet number and five other agronomic traits of ALCD and ALCD-Ae. caudata B–G additions were studied. The results showed that there were no significant agronomic trait influences with the addition of chromosome B of Ae. caudata into ALCD (Figures 1A–H). Chromosome C of Ae. caudata introduced into ALCD showed an increase in the thousand-kernel weight (Figure 1H), however, that chromosome also seemed to produce a negative impact on grain number per spike (Figure 1G). Wheat plants carrying chromosome D of Ae. caudata showed reduced flag leaf width and decreased grain number per spike (Figures 1E,G) compared with ALCD. Chromosome E of Ae. caudata reduced wheat flag leaf width (Figure 1E), and had a negative influence on spikelet number and grain number per spike (Figures 1C,G). Chromosome F appeared to reduce plant height (Figure 1A), but had a negative influence on nearly all of the other agronomic traits (Figures 1B–H). The introduction of chromosome G into wheat had no significant influence on wheat agronomical traits due to the fact that data from four or three locations were not consistent with each other (Figure 1G).
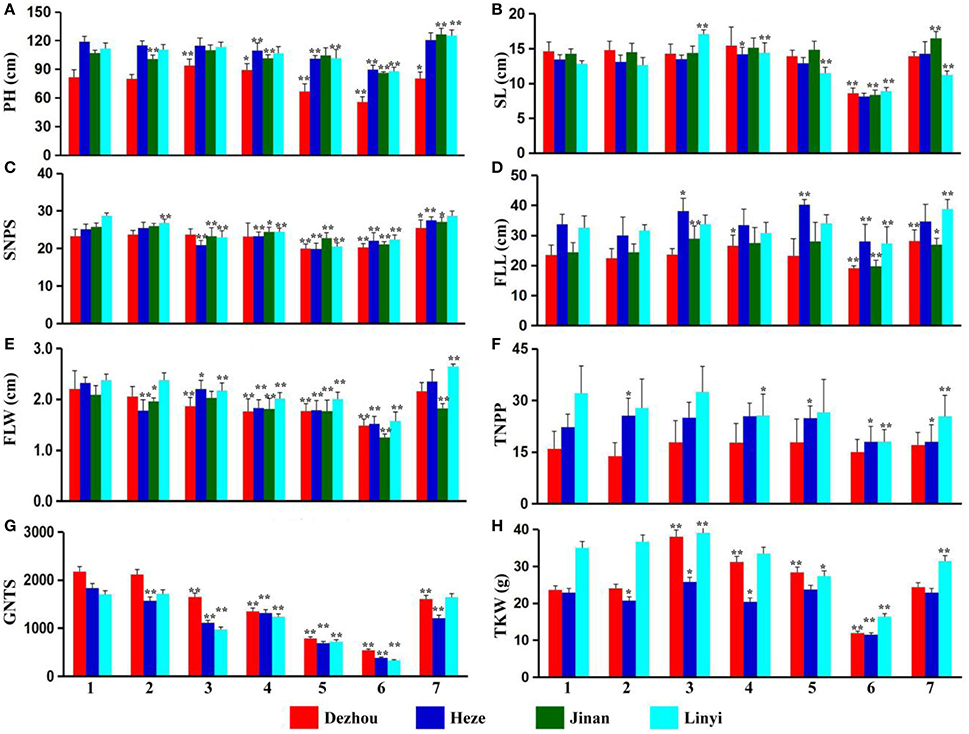
Figure 1. Agronomical traits investigation result of the material tested. Spikelet number, grain number per 30 spikes and thousand-kernel weight data of Ji'nan are not obtained due to crop rotation.1–7 represent T. aestivum cv. Alcedo, Alcedo-Ae. caudata B to G addition lines. PH (A), SL (B), SNPS (C), FLL (D), FLW (E), TNPP (F), GNTS (G), and TKW (H) are the abbreviations of Plant Height, Spike Length, Spikelet Number Per Spike, Flag Leaf Length, Flag Leaf Width, Tiller Number Per Plant, Grain Number of 30 Spikes and Thousand Kernel Weight, respectively. *significant at P < 0.05 by t-test as compared to relative data of ALCD; **significant at P < 0.01 by t-test as compared to relative data of ALCD. Bar represents standard deviation.
Quality Trait Measurements
Protein content and wet gluten content of ALCD and ALCD-Ae. caudata B–G addition lines were measured, and the results showed that data from Dezhou, Heze and Linyi were similar across all sites (Figures 2A,B). Protein content and wet gluten content of ALCD were 16.1–16.3% (Figure 2A) and 33.3–33.8% (Figure 2B), respectively, while measurements of 15.4–20.4% (Figure 2A) (protein content) and 30.8–42.5% (Figure 2B) (wet gluten content) were recorded for the for B–G additions. There were no significant quality differences with the introduction of chromosomes B, D, and G of Ae. caudata into ALCD (Figures 2A,B). However, the presence of chromosome C significantly reduced both wheat protein and wet gluten contents, while chromosomes E and F significantly increased wheat protein and wet gluten contents.
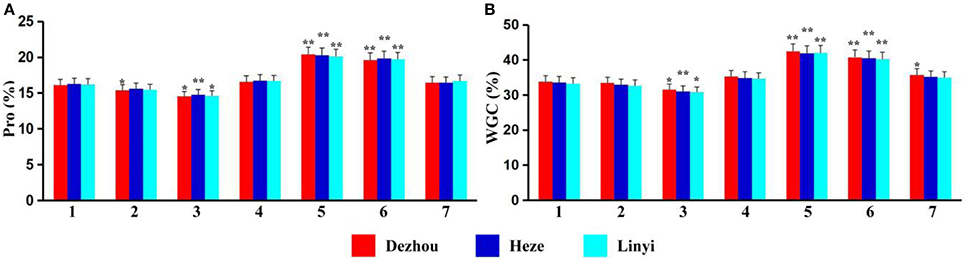
Figure 2. Protein and wet gluten contents of the material tested. Pro (A), Protein; WGC (B), Wet Gluten Content; 1–7 represent T. aestivum cv. Alcedo, Alcedo-Ae. caudata B to G addition lines. *significant at P < 0.05 by t-test as compared to relative data of ALCD; **significant at P < 0.01 by t-test as compared to relative data of ALCD. Bar represents standard deviation.
Identification of ALCD-Ae. caudata Additions Using Molecular Markers
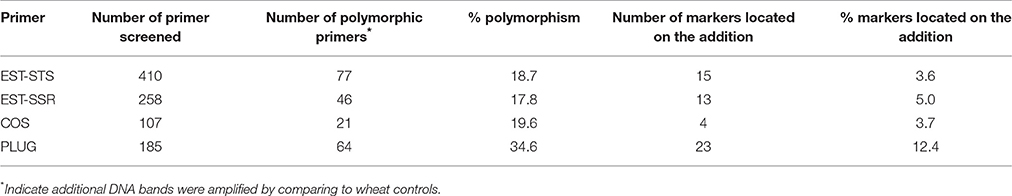
Ae. caudata, CS, T. turgidum, MY11 and MY15 were used to screen 410 pairs of EST-STS primers, 258 pairs of EST-SSR primers, 107 pairs of COS primers and 185 pairs of PLUG primers. The results showed that 77 of EST-STS primer pairs (18.7% of the total primer pairs tested), 46 of EST-SSR primer pairs (17.8%), 21 of the COS primer pairs (19.6%) and 64 of the PLUG primer pairs (34.6%) could generate additional DNA band(s) from Ae. caudata compared to wheat controls as listed in Table 2. The PCR patterns of primer pairs TNAC1497 and TNAC1605 are shown in Figures 3A,C.
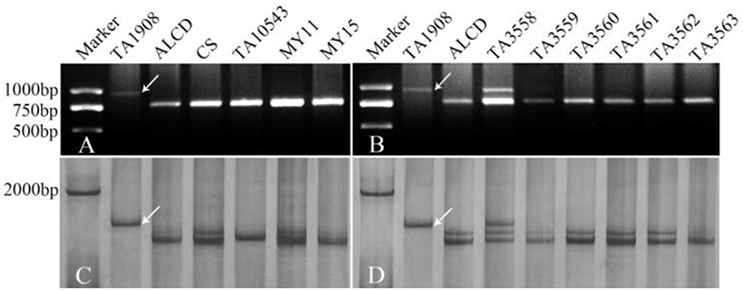
Figure 3. PCR patterns of primer TNAC1497 (A,B) and TNAC1605 (C,D). Arrows indicate polymorphic bands; Panels A,B are from agarose gel, while C,D are from polyacrylamide gel. TA1908 represents Ae. caudata, while ALCD, CS, MY11, and MY15 mean T. aestivum cv. Alcedo, cv. Chinese Spring, cv. Mianyang11 and cv. Miyang15, respectively. TA10543 means T. turgidum, TA3598-TA3563 mean ALCD-Ae. caudata additions B–G, respectively.
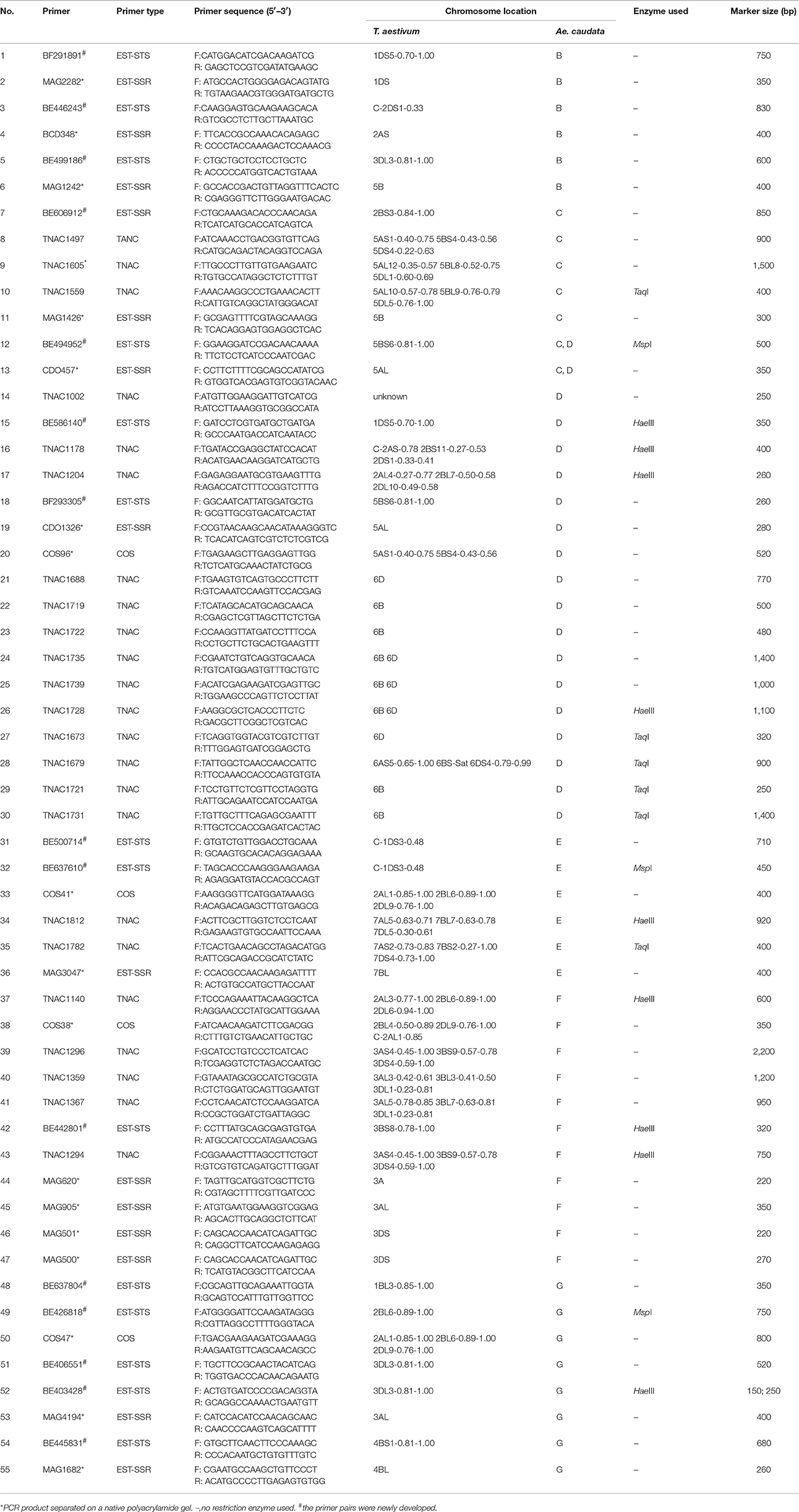
PCR using the polymorphic primer pairs was performed on ALCD and ALCD-Ae. caudata B–G addition lines to locate the polymorphic bands to the Ae. caudata chromosomes. The results suggested that there were 15 (3.6% of the polymorphic primer pairs number), 13 (5.0%), 4 (3.7%), and 23 (12.4%) EST-STS, EST-SSR, COS, and PLUG polymorphic markers, respectively, that could be located to Ae. caudata chromosomes. The PCR patterns of primer pairs TNAC1497 and TNAC1605 as applied to DNA of the addition lines are shown in Figures 3B,D. The detailed information concerning the 55 markers developed by the current research is listed in Table 3.
Primer pairs belong to homoeologous groups 1, 2, 3, and 5 could amplify polymorphic bands from ALCD-Ae. caudata B addition compared to wheat controls (Table 3; Figure 4), implying that a complicated rearrangement involving 1C, 2C, 3C, and 5C might have occurred to chromosome B of Ae. caudata. Primer pairs belong to homoeologous groups 2 and 5 could amplify polymorphic bands from ALCD-Ae. caudata C addition compared to wheat controls (Table 3; Figure 4), implying that a rearrangement involving 2C and 5C might have occurred to chromosome C of Ae. caudata. Molecular marker data physically mapped in wheat also showed that a rearrangement involving 2C, 5C, and 6C might have occurred to chromosome D of Ae. caudata (Table 3; Figure 4). Similarly, a 1C, 2C, and 7C rearrangement might have occurred to chromosome E of Ae. caudata (Table 3; Figure 4), a 2C and 3C rearrangement might have occurred to chromosome F of Ae. caudata (Table 3; Figure 4), a 1C, 2C, 3C, and 4C rearrangement might have occurred to chromosome G of Ae. caudata (Table 3; Figure 4).
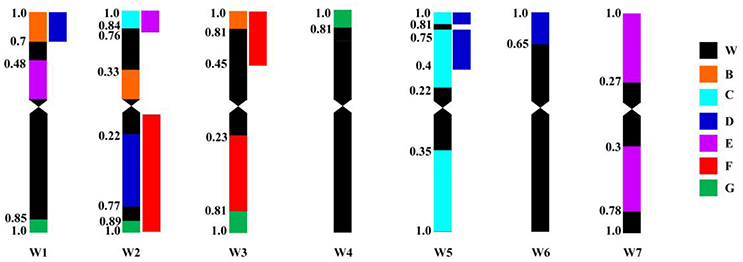
Figure 4. Schematic patterns of Ae. caudata chromosome rearrangement compared to wheat chromosomes as a reference revealed by PLUG markers physically mapped. Ae. caudata chromosomes were marked as B, C, D, E, F and G. W means wheat, W1–W7 represents wheat chromosome group 1–7, respectively. Define the lengths from centromere to both chromosome ends are 1, the values shown on the left of the chromosomes are fragment length (FL).
Discussion
The C genome of Ae. caudata is known to carry many useful genes that can be used for wheat breeding. Whilst investigating the potential of exploiting useful genes from the C genome of Ae. caudata, Riar et al. (2012) mapped a leaf rust resistance gene LrAC originating from a wheat-Ae. caudata introgression line onto the short arm of chromosome 5D of wheat. The LrAc gene is a homoeoallele of an ortholog Lr57. Iqbal et al. (2007) mapped an Ae. caudata-derived major leaf rust resistant quantitative trait locus (Qlr.ipk-2A) on chromosome 2AS of wheat. Weidner et al. (2012) mapped two Ae. caudata-derived powdery mildew resistance loci, QPm.ipk-1A and QPm.ipk-7A, on wheat chromosome arms 1AS and 7AL, respectively. Toor et al. (2016) mapped an Ae. caudata-derived stripe rust resistance gene on wheat arm 5DS. Dyck et al. (1990) selected stem rust resistant germplasm from cross combinations of Ae. caudata and the 5B monosomics of wheat. None of the afore-mentioned studies indicated which specific chromosome of Ae. caudata was carrying the resistance gene(s). The C genome of Ae. caudata has been found to be the source of the C genome of Ae. triuncialis L. (2n = 4x = 28, CCUU) and Ae. cylindrica Host (2n = 4x = 28,CCDD) (Kong et al., 1999a,b). However, there have been earlier reports of disease or pest resistance found in these latter species which clearly has originated from the U or D genomes (not the C genome) (Martin-Sanchez et al., 2003), or alternatively, the resistance was derived from the C genome but the specific identity of that C chromosome based on its conformity to a Triticeae homoeologous group remained unknown (Romero et al., 1998; Galaev et al., 2006; Kuraparthy et al., 2007; Ghazvini et al., 2012). In this current research, we found that the D chromosome of Ae. caudata might possess a leaf rust resistant gene, and the E chromosome might possess stem rust and powdery mildew resistant gene (s). Therefore, these two C-genome chromosome addition lines deserve further investigations involving chromosome engineering activities to produce agronomically useful translocations.
Apart from evaluation of wheat-Ae. caudata germplasm for disease and pest resistance (Romero et al., 1998; Galaev et al., 2006; Kuraparthy et al., 2007; Ghazvini et al., 2012), reports of the agronomical and quality traits are rather rare. In this present research, both agronomical and quality traits of wheat-Ae. caudata additions B–G were investigated or measured, and the introduced Ae. caudata chromosomes into wheat appeared to bring negative influences to several agronomical traits (Figure 1). However, the introduced E or F chromosomes appeared to significantly increase seed protein and gluten contents. Therefore, the induction of wheat-Ae. caudata translocations involving chromosomes E or F could find application in breeding programs which targeted high-yielding or superior-quality wheat. Especially, the superior-quality wheat cultivars with high protein and wet gluten contents might be a candidate food that can provide nutrients for people with malnutrition.
In the aspect of marker development for Ae. caudata chromosomes, Peil et al. (1997, 1998) developed RAPD and SSR markers which could be used to detect Ae. caudata chromatin in a wheat background. Friebe et al. (1992) established the cytogenetic markers, namely the standard C-banding pattern of Ae. caudata chromosomes which could be used for Ae. caudata chromosome identification in wheat. Kong et al. (1999a,b) cloned specific repetitive DNA from the Ae. caudata genome and used it as a marker for the detection of Ae. caudata chromatin in wheat. Badaeva et al. (1996) described the pSc119 and pAs1 FISH karyotype together with the C-banding karyotype of the diploid Aegilops species including Ae. caudata. In this research, a total of 55 EST-based molecular markers which are specific for Ae. caudata chromosomes were developed (Table 3), providing new detection approaches for the quick selection and identification of wheat-Ae. caudata introgressions.
In the aspect of primer pair selection for suitable markers, the data of the current research showed that the rate of successful allocation of polymorphic EST-STS, EST-SSR, COS, and PLUG primers to Ae. caudata chromosomes was 3.6, 5.0, 3.7, and 12.4%, respectively. By comparison, the percentage for marker development of Ae. markgrafii and Ae. cylindrica using COS primers was 80.0% (Molnár et al., 2015), that for Ae. umbellulata, Ae. comosa, Ae. speltoides, and Ae. markgrafii using COS primers range from 46.49 to 53.38% (Molnár et al., 2016), that for Ae. searsii chromosomes by using EST-STS, EST-SSR, COS and PLUG primer was 1.0, 8.6, 5.7, and 16.7%, respectively (Gong et al., 2016), that for Ae. mutica chromosomes by using EST-STS, EST-SSR and PLUG primers was 2.0, 0, and 6.9%, respectively (Liu et al., 2015), that for Ae. uniaristata chromosomes by using EST-STS, EST-SSR, and PLUG primers was 3.5, 11.0, and 11.8%, respectively (Gong et al., 2014), that for Ae. biuncialis, Ae. umbellulata, Ae. comosa, Ae. Biuncialis, and Ae. geniculata, chromosomes rang) range from 54.1 to 80.3% by using COS primers (Molnár et al., 2013), that for Ae. peregrina with US chromosomes and the synthetic KU37 with USsh chromosomes by using COS marker was 43.09%, respectively (Howard et al., 2011), and that for Ae. ventricosa chromsome was 27% by using COS primers (Burt and Nicholson, 2011). The percentage for marker development of Lophopyrum elongatum chromosomes by using EST-SSR and PLUG primers was 6.6 and 11.0%, respectively (Hu et al., 2012), and that for Dasypyrum breviaristatum chromosomes by using EST-STS and PLUG primer was 4.8 and 10.7%, respectively (Liu et al., 2011). The percentage for marker development by using different primer pairs varies. In this research, COS marker development rate using agarose gel electrophoresis is too much lower (3.7%) than that of other reports (27–80.3%) (Burt and Nicholson, 2011; Molnár et al., 2013, 2015, 2016), this might due to that the detection sensitiveness of capillary sequencer or silver staining is too much higher than agarose gel electrophoresis. In this research, PLUG primers appear to generate a higher percentage than other primer pairs, therefore, it should be the system of first choice for marker development of chromosomes belonging to Triticeae species when agarose gel electrophoresis was used. However, COS primer should also be a good choice for marker development if capillary sequencer or silver staining was used.
Schubert and Bluthner (1995) developed the set of wheat-Ae. caudata chromosome additions A–G. Among them, addition A was identified as chromosome 1C by use of isozymes, and also by chromosome characteristics such as the presence of a satellite and C-banding pattern. Friebe et al. (1992) identified additions B–G using the established standard C-banding pattern of the Ae. caudata chromosomes, assuming that additions C, D, and F might be 5C, 6C, and 3C. However, no molecular marker data at the time existed to support this conjecture. The marker results of our current research showed that 7, 19, and 11 markers could be used to identify additions C, D, and F, and among these markers, 6 (85.7%), 10 (52.6%), and 9 (81.8%) belong to homoeologous groups 5, 6, and 3 (Table 3). Therefore, additions C, D, and F should be chromosomes 5C, 6C, and 3C, which confirms Friebe's conjecture (Friebe et al., 1992). Furthermore, the recent single gene FISH mapping data for identifying homoeologous relationships of Ae. caudata chromosomes (Danilova et al., 2016, 2017) also supports this conclusion. Only six markers in this present study could be used to identify addition E, among them, three, two, and one marker(s) belong to homoeologous groups 7, 1, and 2. Addition line A was earlier shown to be the chromosome 1C addition (Schubert and Bluthner, 1995), therefore, addition E could not be addition 1C. The solitary homoeologous group 2 marker associated with addition line E was located on a chromosome terminal region (Table 3), while the two homoeologous group 7 markers were located on the subtelomeric regions of both chromosome long and short arm. Therefore, it is more likely that addition E might be the 7C addition, supporting the results of Danilova et al. (2016, 2017). Cytogenetic evidence has shown that the chromosome B of Ae. caudata might have relationship to both 4C and 5C chromosomes, while chromosome G of Ae. caudata might have a relationship to both 4C and 3C chromosomes (Friebe et al., 1992). Molecular data from this current research suggests that chromosome B of Ae. caudata not only has a relationship to homoeologous group 5, but also to groups 1, 2, and 3 (Table 3; Figure 4), however, we have not found a homoeologous group 4 marker herein. Meanwhile, chromosome G of Ae. caudata not only has a relationship to Triticeae groups 4 and 3, but also to groups 1 and 2 (Table 3; Figure 4). Seed high molecular weight (HWM) protein subunit evidence also supports the conclusion that addition G has a relationship to homoeologous group 1 (Han et al., 2015). Hence, complex chromosomal structural rearrangements might have occurred on chromosomes B and G of Ae. caudata. The molecular data of this research, the recent molecular evidence (Molnár et al., 2016) and cytogenetic evidence (Danilova et al., 2016, 2017) all support that chromosomal structural rearrangements have occurred on chromosomes of Ae. caudata, therefore, further studies into the detailed structures of each of these Ae. caudata chromosomes are warranted.
Wheat-alien introgressions play an important role in wheat resistance breeding. The most notable examples are the wheat-rye 1BL.1RS translocation (Rabinovich, 1998) which for many years was part of most wheat cultivars grown around the world, and also wheat-Dasypyrum villosum 6VS/6AL (Cao et al., 2011) translocation carrying the powdery mildew resistance gene Pm21. Therefore, production of novel wheat-alien species translocations, particularly for disease resistance, has always been the research hot topic. Non-compensating translocations are rarely used in wheat breeding due to the genetic drag or bad agronomical traits (Sears, 1993; Friebe et al., 1996). Therefore, identification of whether the target wheat and alien species chromosomes have been structurally rearranged or not is essential before embarking on the exhaustive and time consuming task of trying to produce compensating translocations for commercial agriculture. Based on the molecular data of this research (Table 3), we found evidence that chromosomes of Ae. caudata have been structurally rearranged. However, evidence from standard C-banding patterns of Ae. caudata chromosomes suggested that no chromosomal arrangements had occurred (Friebe et al., 1992), but in contrast, single gene FISH data support the conclusion that Ae. caudata chromosomes had undergone extensive structural rearrangments (Danilova et al., 2016). Moreover, our molecular data indicate that the rearrangements of B–G chromosomes of Ae. caudata involved at least two homoeologous groups, therefore, a large population and intensive screening pressure needs to be taken into consideration when inducing and searching for wheat-Ae. caudata compensating translocations.
Author Contributions
CL and JL conceived and designed the experiments. RH, HL, JS, and GL performed the experiments. AL and XC contributed reagents/materials/analysis tools. HY, CL, and JG performed disease resistance testing. SZ, DC, and ZZ analyzed the data. WG and RH wrote the paper.
Funding
This research was funded by the National Natural Science Foundation of China (31201203), National key research and development program (2017YFD0100606), a Seed Industry Foundation grant to Taishan Scholar (2014-2019), Natural Science Foundation of Shandong Province (ZR201702070094), the Modern Agricultural Industry Technology System, and Chuang Xin Gong Cheng sponsored by Shandong Academy of Agricultural Sciences (CXGC2016B01).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We would like to express our appreciations to associate Professors Peng Liu, Fengzhi Guo, and Baoqiang Li from Dezhou Academy of Agricultural Sciences, Heze Academy of Agricultural Sciences, Linyi Academy of Agricultural Sciences, respectively, for field management and agricultural traits investigation. We are thankful to associate Professor Tianya Li, College of Plant Protection, Shenyang Agricultural University, for performing stem rust resistance testing. We particularly thank Dr. I. Dundas, School of Agriculture, Food and Wine, The University of Adelaide, Australia, for reviewing and editing of the manuscript.
References
AACC (2000). Approved Methods of the American Association of Cereal Chemists, 10th Edn. St Paul, MN: American Association of Cereal Chemists.
Badaeva, E. D., Friebe, B., and Gill, B. S. (1996). Genome differentiation in Aegilops. 1. Distribution of highly repetitive DNA sequences on chromosomes of diploid species. Genome 39, 293–306.
Baldauf, F., Schubert, V., and Metzlaff, M. (1992). Repeated DNA sequences of Aegilops markgrafii (Greuter) Hammer var. markgrafii: cloning, sequencing and analysis of distribution in Poaceae species. Hereditas 116, 71–78. doi: 10.1111/j.1601-5223.1992.tb00802.x
Barashkova, E. A., and Migushova, E. F. (1984). Genotypic differences between Aegilops in resistance to low temperature. Sel' skokhozyaistevennaya Biol. 997, 41–46.
Biithner, W. D., Schubert, V., and Mettin, D. (1988). “Instability in amphiploids and backcross derivativesofa Triticum aestivum × Ae. caudata cross,” in Proceeding 7th International Wheat Genet Syrup, eds T. E. Miller, R. M. D. Koebner (Cambridge), 209–213.
Burt, C., and Nicholson, P. (2011). Exploiting co-linearity among grass species to map the Aegilops ventricosa-derived Pch1 eyespot resistance in wheat and establish its relationship to Pch2. Theor. Appl. Genet. 123, 1387–1400. doi: 10.1007/s00122-011-1674-9
Cao, A., Xing, L., Wang, X., Yang, X., Wang, W., Sun, Y., et al. (2011). Serine/threonine kinase gene Stpk-V, a key member of powdery mildew resistance gene Pm21, confers powdery mildew resistance in wheat. Proc. Natl. Acad. Sci. U.S.A. 108, 7727–7732. doi: 10.1073/pnas.1016981108
Danilova, T. V., Akhunova, A. R., Akhunov, E. D., Friebe, B., and Gill, B. S. (2017). Major structural genomic alterations can be associated with hybrid speciation in Aegilops markgrafii (Triticeae). Plant J. doi: 10.1111/tpj.13657. [Epub ahead of print].
Danilova, T. V., Friebe, B., and Gill, B. S. (2016). “Homoeologous relationships of Aegilops caudata and wheat chromosomes as determined by single gene FISH mapping,” in Plant and Animal Genome Conference XXIV (San Diego, CA).
Dyck, P. L., Kerber, E. R., and Martens, J. W. (1990). Transfer of a gene for stem rust resistance from Aegilops caudata to common wheat. Can. J. Plant Sci. 70l, 931–934. doi: 10.4141/cjps90-114
Friebe, B., Jiang, J., Raupp, W. J., McIntosh, R. A., and Gill, B. S. (1996). Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91, 59–87. doi: 10.1007/BF00035277
Friebe, B., Schubert, V., Blüthner, W. D., and Hammer, K. (1992). C-banding pattern and polymorphism of Aegilops caudata, and chromosomal constitutions of the amphiploid T. aestivum-Ae. caudata, and six derived chromosome addition lines. Theor. Appl. Genet. 83, 589–596. doi: 10.1007/BF00226902
Galaev, A. V., Babaiants, L. T., and Ium, S. (2006). Molecular marker mapping of the gene resistant to common bunt transferred from Aegilops cylindrica into bread wheat. Tsitol. Genet. 40, 3–11.
Ghazvini, H., Hiebertcolin, W., Zegeye, T., and Tom, F. (2012). Inheritance of stem rust resistance derived from Aegilops triuncialis. Can. J. Plant Sci. 9, 1037–1041. doi: 10.4141/cjps2011-248
Gill, B. S., Sharma, H. C., and Raupp, W. J. (1985). Evaluation of Aegilops species for resistance to wheat powdery mildew, wheat leaf rust, hessian fly, and greenbug. Plant Dis. 69, 314–316.
Gong, W., Li, G., Zhou, J., Li, G., Liu, C., Huang, C., et al. (2014). Cytogenetic and molecular markers for detecting Aegilops uniaristata chromosomes in a wheat background. Genome 57, 489–497. doi: 10.1139/gen-2014-0111
Gong, W. Y., Gong, W. P., Han, R., Li, G. Y., Sehgal, S. K., Li, H. S., et al. (2016). Development of chromosome-arm-specific markers from Aegilops searsii permits targeted introgression. Biologia 71, 87–92. doi: 10.1515/biolog-2016-0003
Gorham, J. (1990). Salt tolerance in the Triticeae: K+/Na+ discrimination in Aegilops species. J. Exp. Bot. 41, 615–621. doi: 10.1093/jxb/41.5.615
Han, R., Sui, X. X., Yang, H. M., Gong, W. Y., Li, G. Y., Chu, X. S., et al. (2015). High molecular weight glutenin composition and powdery mildew resistance of wheat-alien chromosome lines. J. Triticeae Crops 35, 1494–1501.
Howard, T., Rejab, N. A., Griffiths, S., Leigh, F., Leverington-Waite, M., Simmonds, J., et al. (2011). Identification of a major QTL controlling the content of B-type starch granules in Aegilops. J. Exp. Bot. 62, 2217–2228. doi: 10.1093/jxb/erq423
Hu, L. J., Liu, C., Zeng, Z. X., Li, G. R., Song, X. J., and Yang, Z. J. (2012). Genomic rearrangement between wheat and Lophopyrum elongatum revealed by mapped functional molecular markers. Genes Genom. 34, 67–75. doi: 10.1007/s13258-011-0153-7
Iqbal, N., Eticha, F., Khlestkina, E. K., Weidner, A., Röder, S. M., and Borner, A. (2007). The use of simple sequence repeat (SSR) markers to identify and map alien segments carrying genes for effective resistance to leaf rust in bread wheat. Plant Genet. Resour. 5, 100–103. doi: 10.1017/S1479262107672311
Iriki, N., Kawakami, A., Takata, K., Kuwabara, T., and Ban, T. (2001). Screening relatives of wheat for snow mold resistance and freezing tolerance. Euphytica 122, 335–341.
Ishikawa, G., Yonemaru, J., Saito, M., and Nakamura, T. (2007). PCR-based landmark unique gene (PLUG) markers effectively assign homoeologous wheat genes to A, B and D genomes. BMC Genomics 8:135. doi: 10.1186/1471-2164-8-135
Kilian, B., Mammen, K., Millet, E., Sharma, R., Graner, A., Salamini, F., et al. (2011). Wild Crop Relatives: Genomic and Breeding Resources. Berlin; Heidelberg: Springer.
Kong, X. Y., Zhou, R. H., Dong, Y. C., and Jia, J. Z. (1999a). Hybridization of Aegilops caudata with Triticum durum and the detection of alien chromatins. Acta Bot. Sin. 41, 1164–1168.
Kong, X. Y., Zhou, R. H., Dong, Y. C., and Jia, J. Z. (1999b). Cloning of C-genome specific repetitive DNA sequence of the Aegilops caudata. Chin. Sci. Bull. 44, 828–832.
Kuraparthy, V., Sood, S., and Chhuneja, P. (2007). A cryptic wheat-Aegilops triuncialis translocation with leaf rust resistance gene Lr58. Crop Sci. 47, 1995–2003. doi: 10.2135/cropsci2007.01.0038
Liu, C., Li, G. R., Gong, W. P., Li, G. Y., Han, R., Li, H. S., et al. (2015). Molecular and cytogenetic characterization of powdery mildew resistant wheat-Aegilops mutica partial amphiploid and addition line. Cytogenet. Genome Res. 147, 186–194. doi: 10.1159/000443625
Liu, C., Li, G. R., Yang, Z. J., Feng, J., Zhou, J. P., and Ren, Z. L. (2006). Isolation and application of specific DNA segment of rye genome. Acta Bot. Boreal. Occident. Sin. 26, 2434–2438.
Liu, C., Li, G. R., Yan, H. F., Zhou, J. P., Hu, L. J., Lei, M. P., et al. (2011). Molecular and cytogenetic identification of new wheat-D. breviaristatum additions conferring resistance to stem rust and powdery mildew. Breed. Sci. 61, 366–372. doi: 10.1270/jsbbs.61.366
Liu, C., Yan, H. F., Gong, W. P., Li, G. R., Liu, D. Q., and Yang, Z. J. (2013). Screening of new resistance sources of wheat leaf rust. J. Plant Genet. Resour. 14, 936–944.
Makkouk, K. M., Comeau, A., and Ghulam, W. (1994). Resistance to barley yellow dwarf luteovirus in Aegilops species. Can. J. Plant Sci. 74, 631–634. doi: 10.4141/cjps94-113
Martín-Sánchez, J. A., Gómez-Colmenajero, M., Del Moral, J., SinE Montes, M. J., González-Belinchón, C., et al. (2003). A new Hessian fly resistance gene (H30) transferred from the wild grass Aegilops triuncialis to hexaploid wheat. Theor. Appl. Genet. 106, 1248–1255. doi: 10.1007/s00122-002-1182-z
Molnár, I., Šimková, H., Leverington-Waite, M., Goram, R., Cseh, A., Vrána, J., et al. (2013). Syntenic relationships between the U and M genomes of Aegilops, wheat and the model species Brachypodium and rice as revealed by COS markers. PLoS ONE 8:e70844. doi: 10.1371/journal.pone.0070844
Molnár, I., Vrána, J., Burešová, V., Cápal, P., Farkas, A., Darkó, É., et al. (2016). Dissecting the U, M, S and C genomes of wild relatives of bread wheat (Aegilops spp.) into chromosomes and exploring their synteny with wheat. Plant J. 88, 452–467. doi: 10.1111/tpj.13266
Molnár, I., Vrána, J., Farkas, A., Kubaláková, M., Cseh, A., Molnár-Láng, M., et al. (2015). Flow sorting of C-genome chromosomes from wild relatives of wheat Aegilops markgrafii, Ae. triuncialis and Ae. cylindrica, and their molecular organization. Ann. Bot. 116, 189–200. doi: 10.1093/aob/mcv073
Muramatsu, M. (1973). “Genic homology and cytological differentiation of the homeologous-group-5 chromosomes of wheat and related species,” in Proceedings of the Fourth International Wheat Genetics Symposium (Columbia: University of Missouri), 719–724.
Peil, A., Korzun, V., Schubert, V., Schumann, E., Weber, W. E., and Röder, S. M. (1998). The application of wheat microsatellites to identify disomic Triticum aestivum-Aegilops markgrafii addition lines. Theor. Appl. Genet. 96, 138–146. doi: 10.1007/s001220050720
Peil, A., Schubert, V., Schumann, E., and Weber, W. E. (1997). RAPDs as molecular markers for the detection of Aegilops markgrafii chromatin in addition and euploid introgression lines of hexaploid wheat. Theor. Appl. Genet. 94, 934–940. doi: 10.1007/s001220050498
Quraishi, U. M., Abrouk, M., Bolot, S., Pont, C., Throude, M., Guilhot, N., et al. (2009). Genomics in cereals: from genome-wide conserved orthologous (COS) sequences to candidate genes for trait dissection. Funct. Integr. Genomics 9, 473–484. doi: 10.1007/s10142-009-0129-8
Rabinovich, S. V. (1998). Importance of wheat-rye translocations for breeding modern cultivar of Triticum aestivum. Euphytica 100, 323–340. doi: 10.1023/A:1018361819215
Riar, A. K., Kaur, S., Dhaliwal, H. S., Singh, K., and Chhuneja, P. (2012). Introgression of a leaf rust resistance gene from Aegilops caudata to bread wheat. J. Genet. 91, 155–161. doi: 10.1007/s12041-012-0161-7
Romero, M. D., Montes, M. J., Sin, E., Lopez-Brana, L., Duce, A., Martin-sanchez, J. A., et al. (1998). A cereal cyst nematode (Heterodera avenae Woll.) resistance gene transferred from Aegilops triuncialis to hexaploid wheat. Theor. Appl. Genet. 96, 1135–1140. doi: 10.1007/s001220050849
Schubert, V., and Bluthner, W. D. (1995). “Triticum aestivum-Aegilops markgrafii addition lines: production and morphology,” in Proceedings of 8th Wheat International Genet Symposium (Beijing: China Agricultural Scientech Press), 421–425.
Sears, E. R. (1993). Use of radiation to transfer alien segments to wheat. Crop Sci. 33, 897–901. doi: 10.2135/cropsci1993.0011183X003300050004x
Toor, P. I., Kaur, S., Bansal, M., Yadav, B., and Chhuneja, P. (2016). Mapping of stripe rust resistance gene in an Aegilops caudata introgression line in wheat and its genetic association with leaf rust resistance. J. Genetics 95, 1–6. doi: 10.1007/s12041-016-0718-y
Valkoun, J., Hammer, K., Kucerova, D., and Bartos, P. (1985). Disease resistance in the genus Aegilops L.-stem rust, leaf rust, stripe rust, powdery mildew. Kulturpflanze 33, 133–153. doi: 10.1007/BF01997267
Wang, S. W., Yin, L. N., Tanaka, H., Tanaka, K., and Tsujimoto, H. (2011). Wheat-Aegilops chromosome addition lines showing high iron and zinc contents in grains. Breed. Sci. 61, 189–195. doi: 10.1270/jsbbs.61.189
Wang, Z., Cui, Y., Chen, Y., Zhang, D., Liang, Y., Zhang, D., et al. (2014). Comparative genetic mapping and genomic region collinearity analysis of the powdery mildew resistance gene Pm41. Theor. Appl. Genet. 127, 1741–1751. doi: 10.1007/s00122-014-2336-5
Weidner, A., Röder, M. S., and Börner, A. (2012). Mapping wheat powdery mildew resistance derived from Aegilops markgrafii. Plant Genet. Res. 10, 137–140. doi: 10.1017/S1479262112000123
Wu, X. X., Li, T. Y., Chen, S., Wang, G. Q., Cao, Y. Y., Ma, S. L., et al. (2014). Stem rust resistance evaluation and Ug99-resistance gene detection of 139 wheat cultivars. Scientia Agricultura Sinica. 47, 4618–4626.
Keywords: Aegilops caudata, agronomic traits, disease resistance, molecular marker, chromosome rearrangement
Citation: Gong W, Han R, Li H, Song J, Yan H, Li G, Liu A, Cao X, Guo J, Zhai S, Cheng D, Zhao Z, Liu C and Liu J (2017) Agronomic Traits and Molecular Marker Identification of Wheat–Aegilops caudata Addition Lines. Front. Plant Sci. 8:1743. doi: 10.3389/fpls.2017.01743
Received: 15 July 2017; Accepted: 25 September 2017;
Published: 12 October 2017.
Edited by:
Luigi Cattivelli, Consiglio per la Ricerca in Agricoltura e L'analisi Dell'economia Agraria (CREA), ItalyReviewed by:
Annaliese Mason, Justus Liebig Universität Gießen, GermanyIstván Molnár, Centre for Agricultural Research (MTA), Hungary
Ghader Mirzaghaderi, University of Kurdistan, Iran
Marta Molnar-Lang, Hungarian Academy of Sciences (MTA), Hungary
Copyright © 2017 Gong, Han, Li, Song, Yan, Li, Liu, Cao, Guo, Zhai, Cheng, Zhao, Liu and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Cheng Liu, bGNoNjY4ODQwN0AxNjMuY29t
Jianjun Liu, bGpqc2Fhc0AxNjMuY29t
†These authors have contributed equally to this work.
 Wenping Gong
Wenping Gong Ran Han
Ran Han Haosheng Li1
Haosheng Li1 Genying Li
Genying Li Aifeng Liu
Aifeng Liu Cheng Liu
Cheng Liu