- 1Central Horticultural Experiment Station, ICAR-Indian Institute of Horticultural Research, Bhubaneswar, India
- 2World Vegetable Center (WorldVeg), Tainan, Taiwan
- 3Diversity Array Technologies, Canberra, ACT, Australia
Male sterility is of high importance in hybrid seed production of hot and sweet peppers. Genic (or nuclear) male sterility (GMS) is a simply inherited (usually monogenic recessive) and highly stable trait. However, one major disadvantage of using GMS is 1:1 segregation of male sterile to male fertile plants in every subsequent generation. Molecular markers tightly linked to genic male sterility (ms) genes would facilitate an efficient and rapid transfer of ms genes into different genetic backgrounds through marker-assisted backcrossing. The two non-allelic genic male sterility genes ms3 and msw in hot and sweet pepper backgrounds, respectively, are monogenic recessive. Genotyping by sequencing (GBS) in an F2 population segregating for ms3 gene in hot pepper and in an F6 inbred near-isogenic line (NIL) population segregating for msw gene in sweet pepper yielded 9,713 and 7,453 single nucleotide polymorphism markers, respectively. Four candidate SNPs co-segregating with ms3 gene and one co-segregating with msw gene were identified by bulk segregant analysis and physically mapped to chromosomes 1 and 5, respectively. In hot pepper, two markers [HPGMS2 (CAPS) and HPGMS3 (dCAPS)] located 3.83 cM away from the ms3 gene and in sweet pepper the dCAPS marker SPGMS1 co-segregated (completely linked) with the msw gene were developed. These markers will increase the efficacy of the male sterility genes for pepper breeding, as they can be useful in developing the genic male sterile lines in parental inbred lines of commercial hybrids through marker-assisted backcrossing, hybrid seed production, and genetic purity testing of hybrid seeds.
Introduction
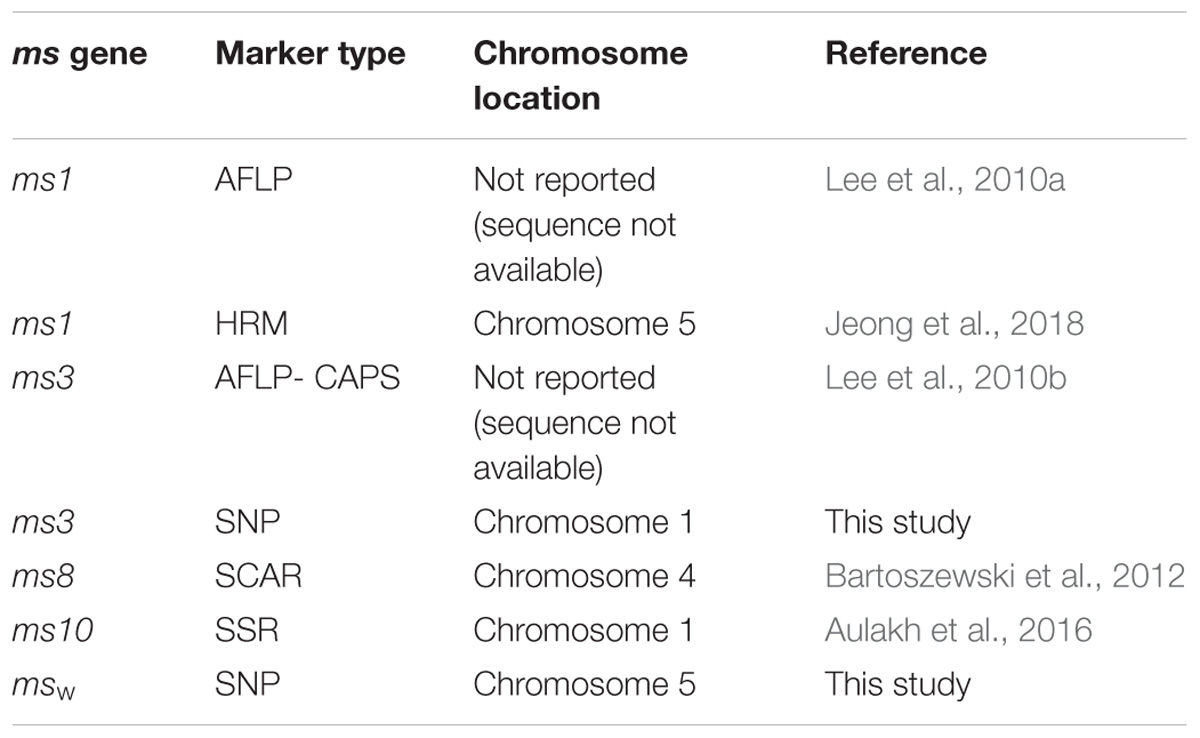
The mechanism of male sterility has long been of interest among plant scientists and breeders. This interest stems from the use of male sterility systems to produce hybrids and their seeds in a cost-effective manner, as well as the differential/spatial expression of male sterility genes, the interaction between the nuclear and cytoplasmic genome, and serves as a model to better understand the endosymbiosis. Peppers (Capsicum spp.) are important commercial cash crops valued globally for their pungent (hot pepper) and non-pungent (sweet pepper) fruits that are used as a spice and vegetable, respectively. Both nuclear male sterility (genic male sterility, GMS) and cytoplasmic male sterility (CMS) systems are used to produce the cost-effective commercial pepper hybrid seed (Dhaliwal and Jindal, 2014; Lin et al., 2016). The CMS hot pepper lines have been used for commercial hybrid seed production, especially in India (Reddy et al., 2015; Lin et al., 2016; Schreinemachers et al., 2016). However, the use of the CMS system in sweet pepper has been limited because of high instability of male sterility expression at low temperatures and poor fertility restoration (Rf) ability in most of the sweet pepper genotypes (Lin et al., 2015). Since two independently isolated and commercially utilized S-cytoplasms of pepper are genetically similar (Kumar et al., 2009), the diversification of S (CMS)-cytoplasm is advisable to avoid risks associated with vulnerability of commercial hot pepper hybrids derived from single S-cytoplasm (Kumar et al., 2009; Yeh et al., 2016). To avoid such risks due to the monopolistic use of the same S-cytoplasm in peppers, GMS lines with highly stable and absolute expression of male sterility (Dhaliwal and Jindal, 2014) should also be increasingly used. More than 20 monogenic recessive nuclear male sterile (ms) genes in pepper are known (Dhaliwal and Jindal, 2014). The ethyl methanesulphonate (EMS)-induced ms10 allele (originally named ms509) (Daskalov and Poulos, 1994) was found to be allelic with the natural msk and ms2 alleles but was non-allelic to the ms1 allele (Shifriss, 1997). Molecular markers linked to ms1 (Lee et al., 2010a), ms3 (Lee et al., 2010b), ms8 (Bartoszewski et al., 2012), ms10 (Aulakh et al., 2016), and to one ms with undisclosed or unknown origin (Lee et al., 2012) have been reported. The ms8 and ms10 genes were mapped to chromosome 4 and chromosome 1, respectively. The ms3-linked AFLP converted CAPS marker was reported, but the primer sequence and chromosome location were not provided (Lee et al., 2010b).
Genotyping by sequencing (GBS) is an efficient technology for the simultaneous discovery and genotyping of SNPs (Elshire et al., 2011; Taranto et al., 2016). Because of its cost effectiveness, GBS has been widely adopted for high density genotyping for linkage analyses, diversity studies, and genome wide association studies (He et al., 2014). The SNP markers have become a broadly used marker system due to the relative abundance of SNPs in most genomes and the co-dominant and bi-allelic nature of these markers (Mammadov et al., 2012). Once the SNP markers associated with a trait of interest are identified, for genotyping large numbers of individuals, suitable SNP assays can be developed (Semagn et al., 2014) or the SNPs can be converted to PCR-based CAPs or dCAPs markers (Lee et al., 2010b; Schafleitner et al., 2016).
The molecular markers linked to the recessive male sterility genes provide an unprecedented increase in efficacy for developing male sterile female inbred lines for hybrid seed production by identifying the male sterile and male fertile plants at the seedling stage. Hence, this study was conducted to perform an allelic test between two monogenic recessive genic male sterility genes and develop and validate molecular markers linked to male sterility genes using GBS. The results are presented and discussed in the regards to currently available molecular markers linked to ms genes in peppers for their wider application in pepper breeding.
Materials and Methods
Plant Materials
Male Sterile Lines and Genes
Two GMS lines, one each in the genetic background of hot and sweet peppers, were used. The hot pepper GMS line possessed the monogenic recessive ms3 allele for male sterility. The ms3 allele was originally derived from the hybrid “Novator F1,” which was provided to the World Vegetable Center (WorldVeg) by the Vegetable Crops Research Institute (VCRI), Hungary in 1987. For mapping the ms3 gene and for marker development, 71 F2 plants derived from a cross between Novator (ms3ms3) and PBC315 (Ms3Ms3) were phenotyped and 47 of these were used for GBS. For ms3-associated molecular marker validation, a separate F2 population of 183 lines was used. The sweet pepper male sterile line possessed an unknown ms allele (designated temporarily as msw) derived through continuous selfing and simple selection of a GMS-based sweet pepper hybrid “Forever” developed by Syngenta Private Limited. A stable F6 inbred population of 91 plants segregating into 50% male sterile (mswmsw) and 50% male fertile (Mswmsw) progenies was created and used for mapping and marker development.
Crosses for Testing Allelism of ms Genes
Since the sweet pepper male sterile line segregated into 50% male sterile (mswmsw) and 50% male fertile (Mswmsw) and the exact genotype of hot pepper male fertile plants was unknown at male sterility locus (Ms3Ms3 or Ms3ms3), we crossed one hot pepper male sterile plant (ms3ms3) with one sweet pepper male fertile plant (Mswmsw) for testing allelism between the ms3 and the msw alleles. A total of 60 F1 plants were phenotyped for male sterility expression with the hypothesis that both the alleles would be allelic if F1 plants segregate at a 1:1 ratio of male sterility and male fertility.
Phenotyping for Male Sterility
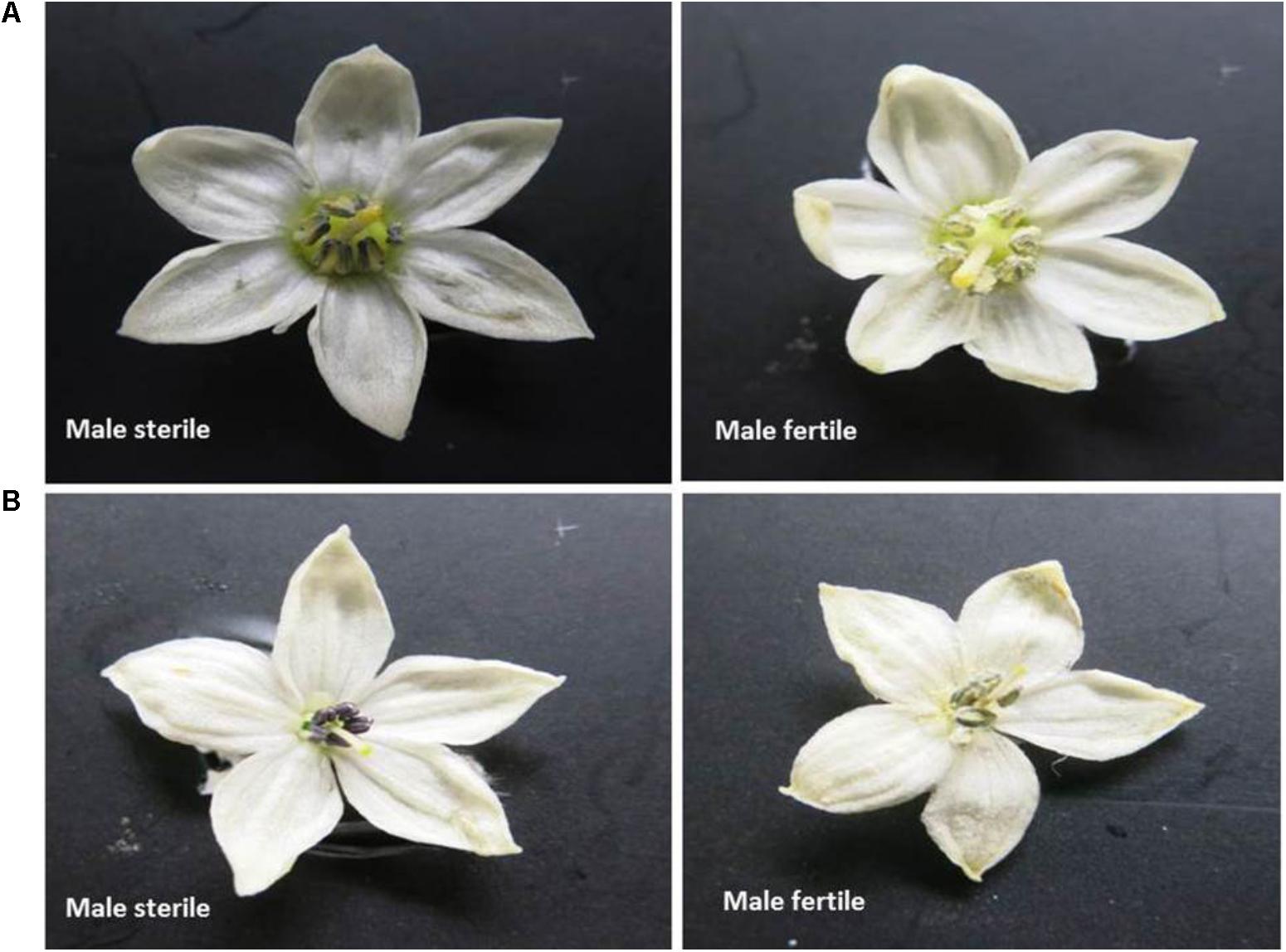
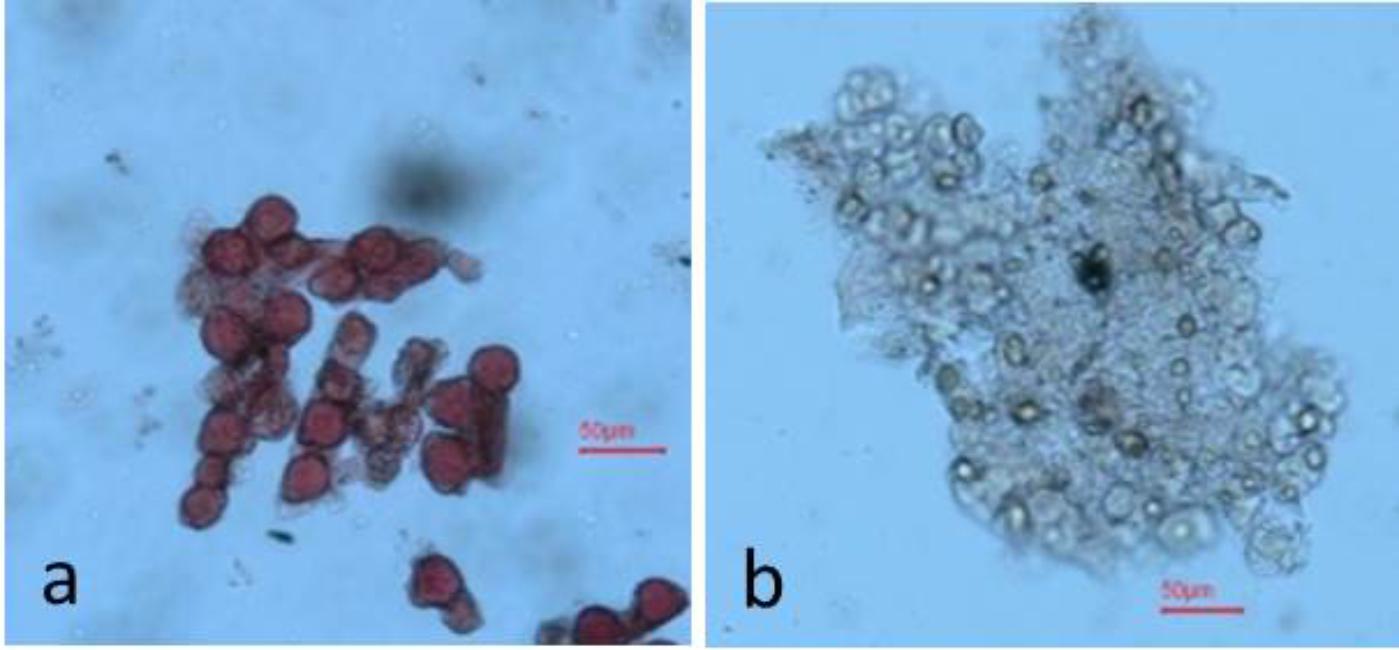
Seeds were sown in small pots (6 inches diameter) in an insect-proof nethouse at WorldVeg, Tainan, Taiwan. All the seedlings were allowed to flower and set fruits. Ten freshly opened flowers of each plant were examined visually by physically tapping on the anthers of the flowers and observing the presence of abundant pollen as a powdery mass (male fertile) or absence of a powdery mass (male sterile) (Figure 1). Ambiguous flowers were tested for pollen viability by 1% acetocarmine staining (Marks, 1954; Figure 2). The final phenotype of each plant was assessed based on their ability (male fertile) or inability (male sterile) to produce fruits with seeds after self-pollination.
Chi Square Test
The probability values between the expected and observed segregation ratios of male sterile and male fertile plants were tested through Chi square for goodness of fit in the Microsoft excel sheet. The calculated chi square value was compared with the t-value. If the calculated value was found to be lower than the t-value, it was considered non-significant, indicating the best fit for the observed and expected ratios.
Genotyping by Sequencing
From 71 F2 hot pepper plants, 47 plants consisting of 28 male fertile (Ms3Ms3 or Ms3ms3) and 19 male sterile (ms3ms3) were selected for GBS. In the case of sweet pepper, from 91 NILs, 47 plants consisting of 23 male fertile (Mswmsw) and 24 male sterile (mswmsw) were selected for GBS. The DNA was isolated using the DNEasy Mini Kit (Qiagen) according to the instructions of the manufacturer. Fluorometric quantification of the DNA was done on a Qubit device (Invitrogen). Sequencing of both the sweet and hot peppers was done at Diversity Array technologies, Canberra, Australia using the DArTSeqTM technology (Sansaloni et al., 2011). Restriction enzymes PstI and MseI were used for complexity reduction, and digestion/ligation reactions were performed as described by Kilian et al. (2012), replacing a single PstI compatible adapter with a PstI barcode adapter, also including the Illumina flow cell attachment and sequencing primer sequence, and a MseI common adapter. Barcodes were used as described (Elshire et al., 2011). For PCR amplification, primer pairs that only bound to fragments containing the PstI adapter on one side and the MseI adapter on the other side were used, thereby leading to selective amplification of fragments having both restriction enzyme cutting ends. The PCR conditions were 1 min at 94°C for initial denaturation; 30 cycles each consisting of 20 s at 94°C for denaturation, 30 s at 58°C for annealing, and 45 s at 72°C for extension; and a final extension step for 7 min at 72°C. After PCR, equimolar amounts of the amplification products from each sample were bulked and applied to c-Bot (Illumina) bridge PCR followed by sequencing on Illumina Hiseq 2500 (Ren et al., 2015). Sequences generated from each lane were processed using the proprietary DArT analytical pipelines, as described (Ren et al., 2015).
Bulk Segregant Analysis
In total, 9713 and 7453 SNP markers were used for bulk segregant analysis for the hot and sweet pepper populations, respectively. Individual male fertile and male sterile plants used for genotyping were grouped into male fertile and male sterile bulks, respectively. In hot pepper, the F2 population segregation did not deviate from a 1:2:1 ratio for the ms3 gene (Ms3Ms3: Ms3ms3: ms3ms3), therefore heterozygous or homozygous SNPs (N/N or N/n) in male fertile plants (28 plants) and homozygous SNPS (n/n) in male sterile plants (19 plants) were selected as candidates associated with the ms3 gene. For the sweet pepper NIL population, segregation did not deviate from a 1:1 ratio into heterozygote male fertile (Mswmsw) and homozygote recessive male sterile (mswmsw) plants, and hence a heterozygous SNP (N/n) in male fertile plants (23 plants) and homozygous (n/n) in male sterile plants (23 out of 24 male sterile plants) were selected as candidate SNPs for marker development.
Marker Development
The sequence context of the candidate SNPs was searched in the Capsicum annuum cv CM334 genome sequence (Kim et al., 2014) by BLAST alignment1 to acquire elongated sequences for the marker development. The SNPs were converted to cleaved amplified polymorphic sequences (CAPS) using the CAPS designer tool2. In cases where conversion of SNPs to CAPS was not possible, dCAPS markers were developed using the dCAPS finder3 (Neff et al., 1998). Primer 3 input (version 0.4.0) program4 was used for designing primers.
Marker Validation
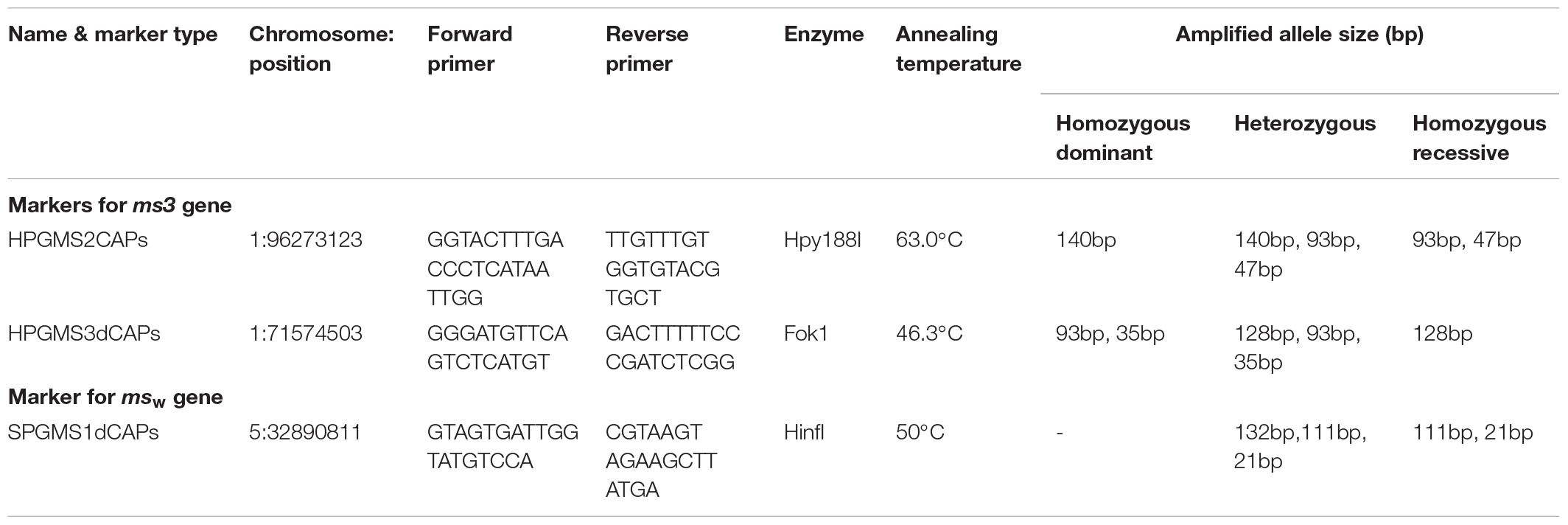
Genomic DNA was extracted from actively growing young leaves (Fulton et al., 1995). Polymerase chain reaction was performed using specific primers listed in Table 2. Each PCR contained 30 ng of DNA, 0.2 mM of each primer, of deoxy ribonucleotides, 50 mM KCl, 10 mM TrisHCl (pH 8.3), 1.5 mM MgCl2, and 0.5 unit of Taq DNA polymerase. The PCR conditions include initial denaturation at 94°C for 5 min, followed by 30 cycles of 94°C for 30 s, annealing temperature for 45 s, 72°C for 45 s, and final extension at 72°C for 7 min. The annealing temperature was optimized for each primer combination by gradient PCR.
Results
Phenotypic Segregation
Seventy-one and 183 F2 plants were, respectively, used for molecular marker development and validation of hot pepper segregating for ms3 gene. Ninety-one plants of the sweet pepper NIL population segregating for the unknown ms gene (Mswmsw/mswmsw) were phenotyped for the expression of male sterility.
In the first set of the F2 population of hot pepper, 52 out of 71 plants were male fertile and 19 were male sterile, which were in agreement with the single recessive gene ratio of 3:1 (χ2 = 0.089 at P = 0.76). Likewise, in the second set of 183 F2 plants, the segregation ratio was also found to be a good fit to a single recessive gene segregation (χ2 = 0.117 at P = 0.73) (Table 1). The segregation of the NIL population of sweet pepper did not deviate from a 1:1 ratio for male fertile: male sterile, which fit well in the expected single gene segregation (χ2 = 0.01 at P = 0.92) (Table 1). Segregation ratios revealed that the male sterility genes in both populations of sweet pepper (msw) and hot pepper (ms3) are monogenic recessive.
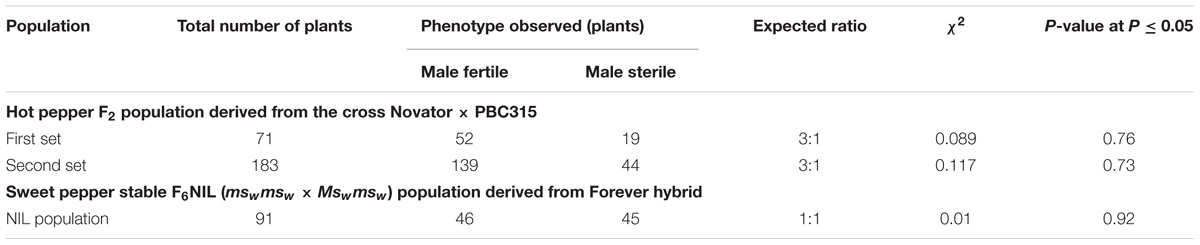

TABLE 1. Segregation of male fertile and male sterile phenotypes in F2 population of hot pepper and near isogenic lines population of sweet pepper.
Allelism Test
Sixty F1 plants produced from a cross between male sterile hot pepper (ms3ms3) and the sweet pepper plants with the unknown heterozygous male sterility allele (Mswmsw) were found to be 100% male fertile, indicating that these two genes (ms3 and msw) are non-allelic. Alternatively, had these been different alleles of the same gene there would have been 1:1 segregation of male fertile and male sterile plants.
Genotyping by Sequencing
From the 9,713 SNPs identified in the hot pepper library, 6909 (71%) could be mapped to the reference genome. Likewise, in the sweet pepper library, 7453 SNPs were identified and the proportion of mapped markers was 68.4%, which is similar to the hot pepper library. The SNPs covered all the 12 chromosomes of pepper, with 458 to 707 SNPs per chromosome for hot pepper and 376–533 SNPs per chromosome for sweet pepper. From all the SNPs, 2,804 and 2,354 SNPs could not be attributed to a chromosome in hot and sweet peppers, respectively. The average distance between two markers in hot pepper amounted to 497 kb (maximum distance 15 Mb) while it was 656 kb (maximum distance 16 Mb) in sweet pepper.
Bulk Segregant Analysis
Since the expression of male sterile genes ms3 and msw was strictly monogenic recessive, we conducted a bulk segregant analysis (BSA) comparing the marker genotypes between bulks. For hot pepper, 4 SNPs were selected as candidates for molecular markers from 47 genotyped plants because they were homozygous recessive (n/n) in 19 male sterile (ms3ms3) plants, heterozygous (N/n) in 20 male fertile (Ms3ms3) plants, and homozygous dominant (N/N) in 8 male fertile (Ms3Ms3) plants. All these four candidate SNPs were physically mapped on chromosome 1 at positions 70,730,824–96,273,123 bp and co-segregated with male sterility.
In sweet pepper, only one SNP marker was heterozygous in all 23 male fertile (Mswmsw) plants and homozygous recessive in 23 out of 24 male sterile plants (mswmsw). This candidate SNP was physically mapped to chromosome 5 at the physical position 32,890,811.
Marker Development
The candidate SNPs were converted to PCR-based CAPS and dCAPS markers. In hot pepper, for candidate SNPs 1 and 3 CAPS markers were developed, whereas for candidate SNPs 2 and 4, dCAPS markers were developed because these SNPs could not be converted to CAPS markers (SNP accession # PRJEB28302). In sweet pepper, the candidate marker was physically converted to a dCAPS marker. The details of the (d)CAPS markers such as primer sequence, restriction enzyme, and PCR product sizes are presented in Table 2.
Validation of Markers and Mapping
Molecular markers were validated in sets of phenotyped plants.
Four SNP markers physically mapped on chromosome 1 at positions 71,091,336 (SNP1), 96,273,123 (SNP 2), 71,574,503 (SNP 3), and 70,730,824 (SNP 4) co-segregated with the male sterile/fertile phenotype. In the initial validation of the GBS data in the 47 genotyped plants of the BSA, HPGMS 1 primers designed for SNP 1 was not polymorphic, indicating that the GBS gave a false positive SNP at this position. In contrast, HPGMS2 and HPGMS3 were polymorphic among male fertile and sterile plants and co-segregated perfectly with the male sterile/fertile phenotype in the 47 genotypes. The HPGMS4 marker showed partial digestion interfering with scoring and was discarded. Therefore, the HPGMS2 (CAPS) and HPGMS3 (dCAPS) markers were validated in a test set of 183 F2 plants with known male fertility phenotype. Both markers differentiated heterozygous (Ms3ms3- male fertile), homozygous dominant (Ms3Ms3–male fertile), and homozygous recessive (ms3ms3 – male sterile) genotypes without any ambiguity (Table 3 and Figures 3, 4). Seven recombinants were found in 183 test plants between HPGMS2 and HPGMS3, and the sterility locus suggested a genetic distance between both markers and the ms3 gene to be 3.83 cM.
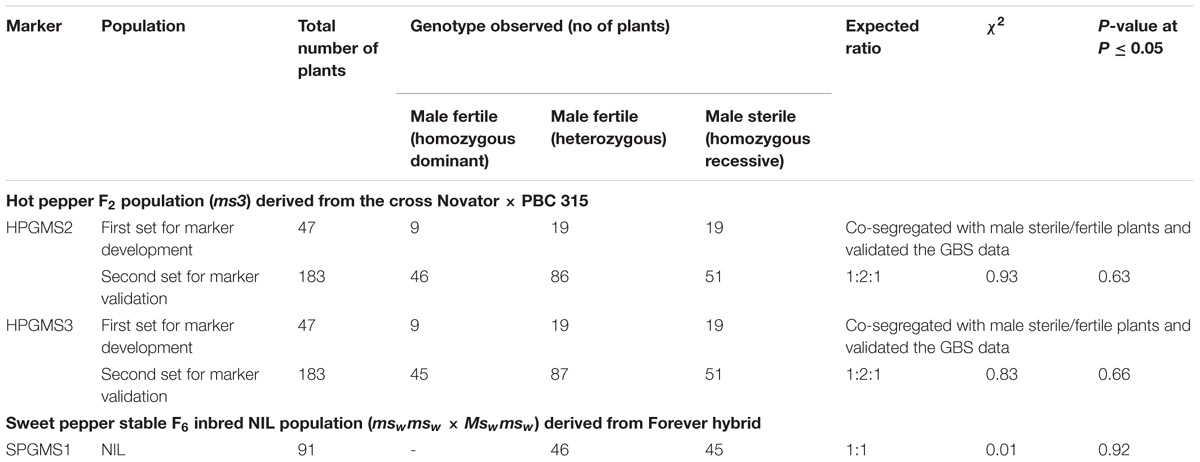

TABLE 3. Segregation of SNP-derived markers linked to ms3 and msw genes in hot pepper (F2) and sweet pepper (near isogenic; NIL) populations.
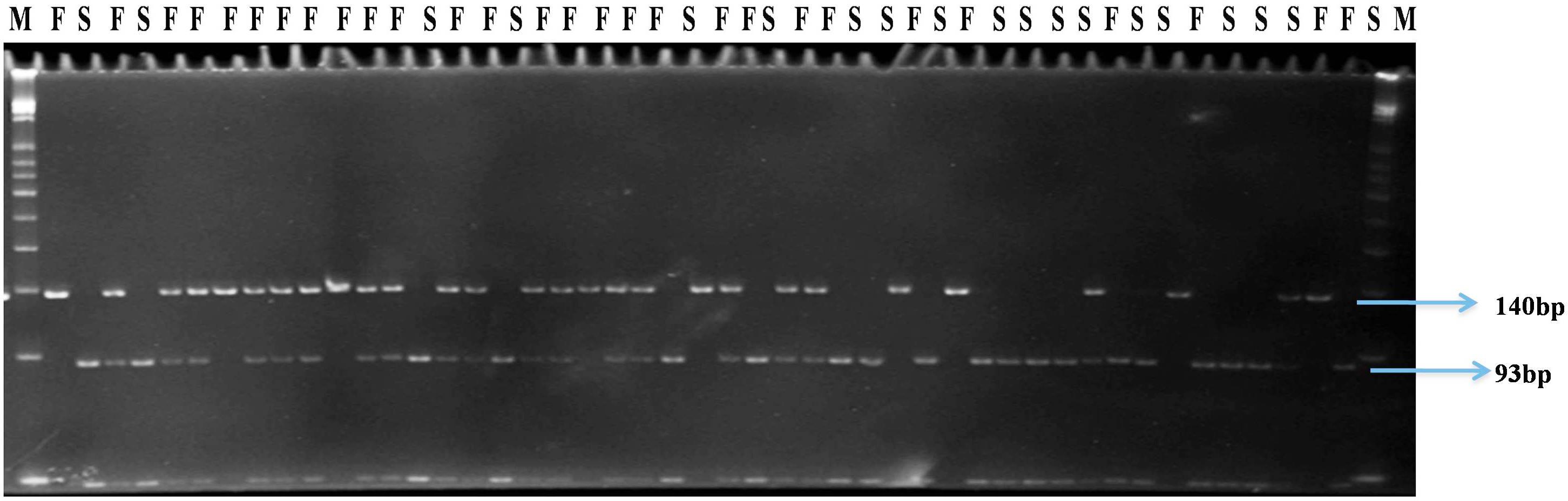
FIGURE 3. Gel profile of HPGMS2-CAPS marker linked to ms3 gene in an F2 segregating population; M, 50 bp DNA ladder; F, male fertile (Ms3Ms3/Ms3ms3); S, male sterile (ms3ms3).
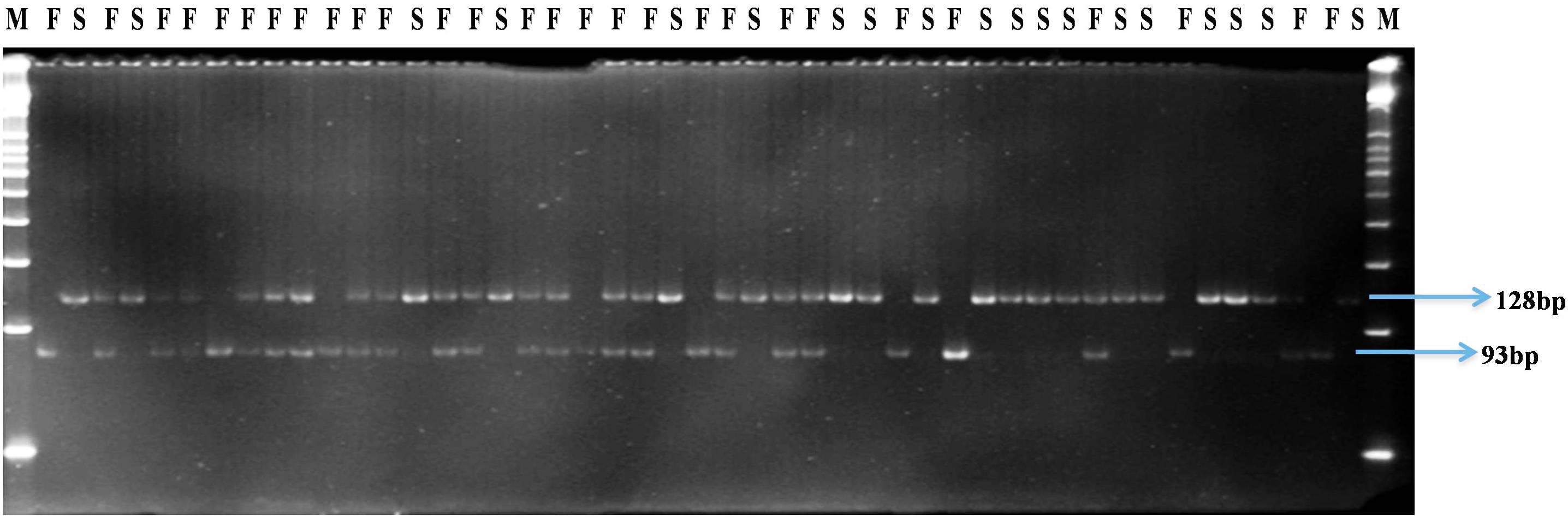
FIGURE 4. Gel profile of HPGMS3-dCAPS marker linked to ms3 gene in an F2 segregating population; M, 50 bp DNA ladder; F, male fertile (Ms3Ms3/Ms3ms3); S, male sterile (ms3ms3).
In sweet pepper, the dCAPS marker SPGMS1 (Table 3) co-segregated with sterility and differentiated the heterozygous (Mswmsw) male fertile plants from the homozygous recessive (mswmsw) male sterile plants (χ2 = 0.01 at P = 0.92) (Table 3 and Figure 5).
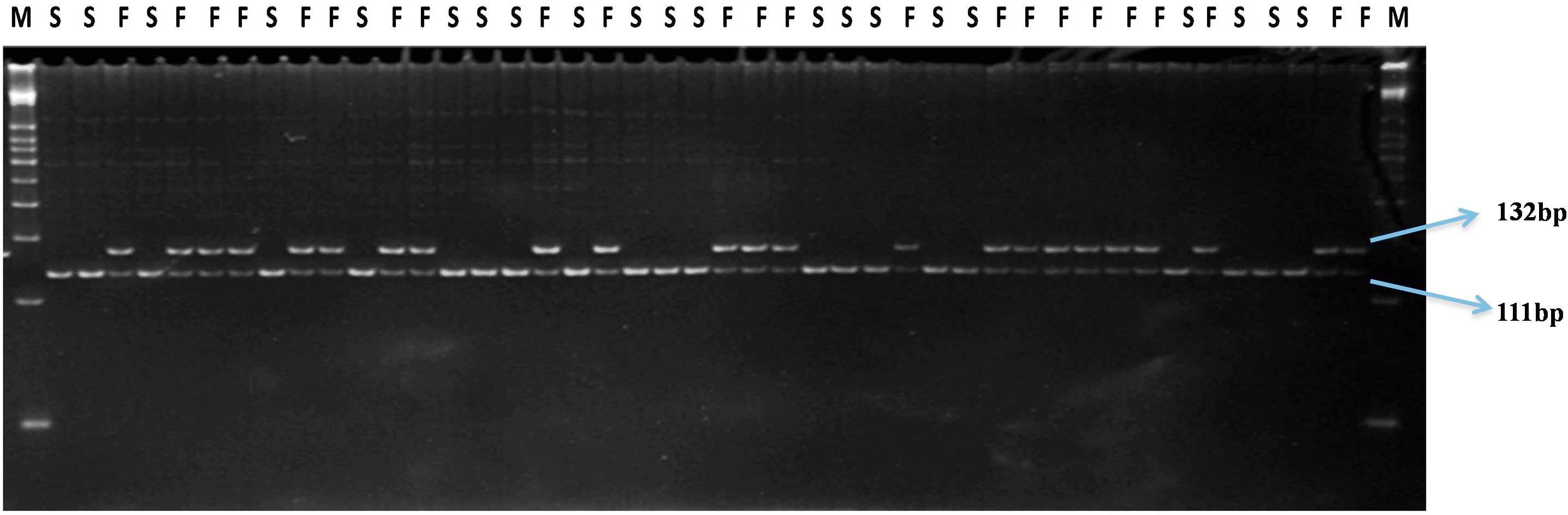
FIGURE 5. Gel profile of SPGMS1-dCAPS marker linked to msw gene in an F6 NIL population; M, 50 bp DNA ladder; F, male fertile (Mswmsw); S, male sterile (mswmsw).
Discussion
Hybrid seed production through manual emasculation and pollination in pepper is labor demanding and not cost-effective. The introduction of male sterility systems can reduce up to 40% of the labor costs by eliminating the manual emasculation process (Lin et al., 2015). Both GMS and CMS systems are available and well documented in peppers. For commercial hybrid development and seed production, the CMS is currently used only in hot pepper, while the GMS is used in both hot and sweet peppers (Dhaliwal and Jindal, 2014). This is because fertility restoration in sweet pepper is weak (Lin et al., 2015) and the expression of male sterility is conditioned by the presence of several restorer-of-fertility (Rf) loci including partial restoration loci in pepper (Lee et al., 2008). However, the worldwide commercially used male sterile cytoplasm is genetically similar (Kumar et al., 2009). This monopolistic use of a single source of a male sterile cytoplasm is not desirable and needs either diversification of CMS source in the long term, or promotion of GMS-based hybrid crops in the short term. Hence, to increase the efficacy of GMS in hybrid seed production in hot and sweet peppers, we mapped the ms3 and msw genes and developed molecular markers to distinguish the alleles for male fertility and sterility. Both genes (ms3 and msw) caused complete male sterility and their inheritance was monogenic recessive. Among the approximately 20 known spontaneously occurring or artificially induced nuclear male sterile genes in peppers (Wang and Bosland, 2006; Dhaliwal and Jindal, 2014), all are monogenic recessive (Shifriss, 1997; Lee et al., 2012; Aulakh et al., 2016) with the single exception of the dominant mutant male sterility gene Dms (Daskalov and Poulos, 1994). Very few allelic tests have been performed among these male sterile genes (Dhaliwal and Jindal, 2014). However, ms10 was found to be non-allelic to ms3 (Yu, 1990) but allelic to the msk gene (Yu, 1985). We found that ms3 and msw are non-allelic as all the F1 plants from the hybridization ms3ms3 ×Mswmsw were male fertile. This non-allelic relationship established through a genetic test was also confirmed through genotypic sequence data, as ms3 and msw genes were physically mapped on different chromosomes, chromosomes 1 and 5, respectively (Table 4). The ms10 (originally ms509) gene has also been recently mapped on chromosome 1 and the associated Simple Sequence Repeat (SSR) marker (AVRDC-PP12) was developed (Aulakh et al., 2016). However, as expected, the AVRDC-PP12 marker for ms10 did not produce polymorphism in our ms3 population (data not shown), ascertaining that both are non-allelic. GBS revealed to be highly efficient in obtaining markers tightly linked to monogenic male sterility (ms) genes using a small population. Hence, the method is rapid and involves relatively low cost (He et al., 2014). We employed GBS to identify the SNP markers for ms3 and msw genes and successfully converted the SNP markers into PCR-based CAPS and dCAPS markers and validated these markers in segregating populations. Since both the male sterile genes were monogenic recessive, 47 plants were sufficient to identify the markers associated with msw in the NIL population and ms3 in the F2 population. For the ms3 gene, one CAPS (HPGMS2) and one dCAPS (HPGMS3) marker were developed. Both of these markers co-segregated with the ms3 gene and the phenotypes in 176 out of 183 plants (Table 3), and therefore are suggested to be located at 3.82 cM from the ms3 gene. In spite of the distance between the markers and the ms3 gene, these markers could be used in marker-assisted selection (MAS) and hybrid seed production by facilitating identification of most of the male sterile and male fertile plants at the seedling stage. As expected, six successive generations of backcrossing introgression carrying the male sterility gene in the NIL population was small, and therefore, only one candidate marker was obtained for this population, while four candidates were obtained for the F2 population. The dCAPS marker SPGMS1 was developed for the msw gene for sweet pepper, which showed complete co-segregation with the msw gene differentiating the heterozygous male fertile plants from the homozygous recessive sterile plants. MAS is important for the GMS system because (i) transferring the ms gene into different genetic backgrounds through conventional backcrossing is tedious and time-consuming, as selfing after each successive backcross is required to select the male fertile heterogonous plant and (ii) for hybrid seed production, 50% male fertile plants need to be identified and removed in the hybrid seed production plot, which is also a time- and resource-consuming process. Both these obstacles are overcome by the use of molecular markers linked to the male sterility locus. Hence, the co-dominant molecular markers linked to the ms3 (HPGMS2 and HPGMS3) and msw (SPGMS1) genes developed in this study will greatly reduce the amount of time and labor resources required for early (seedling stage) identification of the heterozygous male fertile plants (Msms) in backcross breeding (Dhaliwal and Jindal, 2014). These molecular markers will be equally useful in selecting the male sterile seedlings for hybrid seed production (Lee et al., 2010b) and also in hybrid purity testing (Jang et al., 2004). In addition, the molecular markers identified in this study may be useful in the cloning of ms3 and msw genes for various uses in strategic and applied research including map-based cloning of these two male sterility genes.
In peppers, although more than 20 ms genes have been reported, molecular markers linked to male sterility loci are known for only a few of them (Table 4). AFLP markers associated with the ms1 gene (Lee et al., 2010a), CAPS markers associated with the unknown ms gene (Lee et al., 2010c), SCAR markers associated with the ms8 gene (Bartoszewski et al., 2012), and a distantly linked SSR marker for the ms10 gene (Aulakh et al., 2016) have been developed. However, for the ms3 gene, the marker information (primer sequence) has not previously been revealed (Lee et al., 2010b; Table 4) and was not available for public use. For the first time we are reporting markers for the ms3 gene with complete and detailed information for public use. During the preparation of this manuscript, Jeong et al. (2018) reported that ms1 in Magnipio hybrid of Syngenta is located on chromosome 5, similar to msw derived from Forever hybrid of Syngenta mapped in this study (Table 4). Moreover, the location of the marker for msw (32187928) is within the fine mapped 869.9 kb ms1 gene (31516016–32385930) reported by Jeong et al. (2018). Hence, it is highly likely that msw and ms1 are the same gene (allelic); however, a genetic test between msw and ms1 would be desirable to confirm their functional allelism.
Nuclear male sterile genes in peppers are usually highly stable and possess a monogenic simply inherited trait; therefore, they could be relatively easy to use in breeding programs if molecular markers are available. We employed GBS technology to develop molecular markers linked to two non-allelic male sterility genes (ms3 and msw) in peppers. These markers will facilitate rapid and cost-effective development of GMS lines, useful in hybrid seed production and genetic purity testing of hybrid seeds in peppers.
Author Contributions
PN executed the research work and data analysis and led the manuscript preparation. SWL helped in greenhouse experiments. CYL helped in marker development. YWW helped in markers validation. RS supervised the genotyping by sequencing progress and revised the manuscript. AK helped in GBS data analysis. SK planned the entire work and helped substantially to develop the draft and revisions of the manuscript.
Funding
The operation costs of this research were provided by WorldVeg through its innovation fund project. PN was provided post-doc fellowship by Department of Science and Technology (DST), India to conduct this research at WorldVeg.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
PN thanks the Science and Engineering Research Board (SERB, DST), the Government of India for fellowship and Indian Council of Agricultural Research (ICAR), New Delhi for deputing for the project.
Footnotes
- ^ https://www.solgenomics.net/tools
- ^ https://www.solgenomics.net/tools/caps_designer
- ^ http://helix.wustl.edu/dcaps/dcaps.html
- ^ http://bioinfo.ut.ee/primer3-0.4.0
References
Aulakh, P. K., Dhaliwal, M. S., Jindal, S. K., Schafleitner, R., and Singh, K. (2016). Mapping of male sterility gene ms10 in chilli pepper (Capsicum annuum L.). Plant Breed. 135, 531–535. doi: 10.1111/pbr.12389
Bartoszewski, G., Waszczak, C., Gawronski, P., Stepien, I., Bragoszewska, H. B., Palloix, A., et al. (2012). Mapping of the ms8 male sterility gene in sweet pepper (Capsicum annuum L.) on the chromosome P4 using PCR-based markers useful for breeding programmes. Euphytica 186, 453–461. doi: 10.1007/s10681-012-0637-9
Daskalov, S., and Poulos, J. M. (1994). Updated Capsicum gene list. Capsicum and Eggplant. News Lett. 13, 16–26.
Dhaliwal, M. S., and Jindal, S. K. (2014). Induction and exploitation of nuclear and cytoplasmic male sterility in pepper (Capsicum spp.): a review. J. Hortic. Sci. Biotechnol. 89, 471–479. doi: 10.1080/14620316.2014.11513108
Elshire, R. J., Glaubitz, J. C., Sun, Q., Poland, J. A., Kawamoto, K., Buckler, E. S., et al. (2011). A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6:e19379. doi: 10.1371/journal.pone.0019379
Fulton, T. M., Chunwongse, J., and Tanksley, S. D. (1995). Microprep protocol for extraction of DNA from tomato and other herbaceous plants. Plant Mol. Biol. Rep. 13, 207–209. doi: 10.1007/BF02670897
He, J., Zhao, X., Laroche, A., Lu, Z. X., Liu, H. K., and Li, Z. (2014). Genotyping-by-sequencing (GBS), an ultimate marker-assisted selection (MAS) tool to accelerate plant breeding. Front. Plant Sci. 5:484. doi: 10.3389/fpls.2014.00484
Jang, I., Moon, J. H., Yoon, J. B., Yoo, J. H., Yang, T. J., Kim, Y. J., et al. (2004). Application of RAPD and SCAR markers for purity testing of F1 hybrid seed in chilli pepper (Capsicum annuum L.). Mol. Cells 18, 295–299.
Jeong, K., Choi, D., and Lee, J. (2018). Fine mapping of the genic male sterile ms1 gene in Capsicum annuum L. Theor. Appl. Genet. 131, 183–191. doi: 10.1007/s00122-017-2995-0
Kilian, A., Wenzl, P., Huttner, E., Carling, J., Xia, L., Blois, H., et al. (2012). Diversity arrays technology: a generic genome profiling technology on open platforms. data production and analysis in population genomics. Methods Mol. Biol. 888, 67–89. doi: 10.1007/978-1-61779-870-2_5
Kim, S., Park, M., Yeom, S. I., Kim, Y. M., Lee, J. M., Lee, H. A., et al. (2014). Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat. Genet. 46, 270–278. doi: 10.1038/ng.2877
Kumar, R., Kumar, S., Dwivedi, N., Kumar, S., Rai, A., Singh, M., et al. (2009). Validation of SCAR markers, diversity analysis of male sterile (S-) cytoplasms and isolation of an alloplasmic S-cytoplasm in Capsicum. Sci. Hortic. 120, 167–172. doi: 10.1016/j.scienta.2008.10.012
Lee, H. R., An, H. J., Yang, D. C., Choi, S. H., Kim, H. J., Rhee, H. G., et al. (2012). Development of a high resolution melting (HRM) marker linked to genic male sterility in Capsicum annuum L. Plant Breed. 131, 444–448.
Lee, J., Han, J. H., An, C. G., Lee, W. P., and Yoon, J. B. (2010c). A CAPS marker linked to a genic male-sterile gene in the colored sweet pepper, ‘Paprika’ (Capsicum annuum L). Breed. Sci. 60, 93–98. doi: 10.1270/jsbbs.60.93
Lee, J., Yoon, J. B., Han, J. H., Lee, W. P., Do, J. W., Ryu, H., et al. (2010a). A co-dominant SCAR marker linked to the genic male sterility gene (ms1) in chili pepper (Capsicum annuum). Plant Breed. 129, 35–38. doi: 10.1111/j.1439-0523.2009.01643.x
Lee, J., Yoon, J. B., Han, J. H., Lee, W. P., Kim, S. H., and Par, H. G. (2010b). Three AFLP markers tightly linked to the genic male sterility ms3 gene in chili pepper (Capsicum annuum L.) and conversion to a CAPS marker. Euphytica 173, 55–61. doi: 10.1007/s10681-009-0107-1
Lee, J. D., Yoon, J. B., and Park, H. G. (2008). Linkage analysis between the partial restoration (pr) and the restorer-of-fertility (Rf) loci in pepper cytoplasmic male sterility. Theor. Appl. Genet. 117, 383–389. doi: 10.1007/s00122-008-0782-7
Lin, S. W., Shieh, H. C., and Kumar, S. (2016). “Male sterility research in peppers at World Vegetable Center,” in Proceedings of the XVI EUCARPIA Meeting on Genetics and Breeding of Capsicum and Eggplant, Kecskemét, HU, 46–50.
Lin, S. W., Shieh, H. C., Wang, Y. W., Gniffke, P. A., Tan, C. W., Schafleitner, R., et al. (2015). Restorer breeding in sweet pepper: introgressing Rf allele from hot pepper through marker-assisted backcrossing. Sci. Hortic. 197, 170–175. doi: 10.1016/j.scienta.2015.09.036
Mammadov, J., Aggarwal, R., Buyyarapu, R., and Kumpatla, S. (2012). SNP markers and their impact on plant breeding. Int. J. Plant Genomics 2012:728398. doi: 10.1155/2012/728398
Marks, G. E. (1954). An acetocarmine glycerol jelly for use in pollen fertility counts. Biotech. Histochem. 29, 277–278.
Neff, M. M., Neff, J. D., Chory, J., and Pepper, A. E. (1998). dCAPS, a simple technique for the genetic analysis of single nucleotide polymorphisms: experimental applications in Arabidopsis thaliana genetics. Plant J. 14, 387–392. doi: 10.1046/j.1365-313X.1998.00124.x
Reddy, M. K., Srivastava, A., Lin, S. W., Kumar, R., Shieh, H. C., Ebert, A. W., et al. (2015). Exploitation of AVRDC’s chili pepper (Capsicum spp.) germplasm in India. J. Taiwan Soc. Hortic. Sci. 61, 1–9.
Ren, R., Ray, R., Li, P., Xu, J., Zhang, M., Liu, G., et al. (2015). Construction of a high-density DArTseq SNP-based genetic map and identification of genomic regions with segregation distortion in a genetic population derived from a cross between feral and cultivated-type watermelon. Mol. Genet. Genomics 290, 1457–1470. doi: 10.1007/s00438-015-0997-7
Sansaloni, C., Petroli, C., Jaccoud, D., Carling, J., Detering, F., Grattapaglia, D., et al. (2011). Diversity arrays technology (DArT) and next-generation sequencing combined: genome-wide, high throughput, highly informative genotyping for molecular breeding of Eucalyptus. BMC Proc. 5(Suppl.7):54. doi: 10.1186/1753-6561-5-S7-P54
Schafleitner, R., Huang, S. M., Chu, S. H., Yen, J. Y., Lin, C. Y., Yan, M. R., et al. (2016). Identification of single nucleotide polymorphism markers associated with resistance to bruchids (Callosobruchus spp.) in wild mungbean (Vigna radiata var. sublobata) and cultivated V. radiata through genotyping by sequencing and quantitative trait locus analysis. BMC Plant Biol. 16:159. doi: 10.1186/s12870-016-0847-8
Schreinemachers, P., Rao, K. P. C., Easdown, W., Hanson, P., and Kumar, S. (2016). The contribution of international vegetable breeding to private seed companies in India. Genet. Resour. Crop Evol. 64, 1037–1049. doi: 10.1007/s10722-016-0423-y
Semagn, K., Babu, R., Hearne, S., and Olsen, M. (2014). Single nucleotide polymorphism genotyping using kompetitive allele specific PCR (KASP): overview of the technology and its application in crop improvement. Mol. Breed. 33, 1–14. doi: 10.1007/s11032-013-9917-x
Shifriss, C. (1997). Male sterility in pepper (Capsicum annuum L.). Euphytica 93, 83–88. doi: 10.1023/A:1002947907046
Taranto, F., D’Agostino, N., Greco, B., Cardi, T., and Tripodi, P. (2016). Genome-wide SNP discovery and population structure analysis in pepper (Capsicum annuum) using genotyping by sequencing. BMC Genomics 17:943. doi: 10.1186/s12864-016-3297-7
Yeh, T., Lin, S., Teoh, Y., and Kumar, S. (2016). Markers for cytoplasmic male sterility (CMS) traits in chili peppers (Capsicum annuum L.): multiplex PCR and validation. SABRAO J. Breed. Genet. 48, 465–473.
Yu, I. W. (1985). Inheritance of Cytoplasmic Male Sterility in Pepper (Capsicum annuum L.). M.Sc. Thesis, Kyung Hee University, Seoul.
Keywords: genic male sterility, genotyping by sequencing, hot pepper, hybrid seed production, SNP, sweet pepper
Citation: Naresh P, Lin S-w, Lin C-y, Wang Y-w, Schafleitner R, Kilian A and Kumar S (2018) Molecular Markers Associated to Two Non-allelic Genic Male Sterility Genes in Peppers (Capsicum annuum L.). Front. Plant Sci. 9:1343. doi: 10.3389/fpls.2018.01343
Received: 24 January 2018; Accepted: 24 August 2018;
Published: 16 October 2018.
Edited by:
Jianjun Chen, University of Florida, United StatesReviewed by:
Anil Khar, Indian Agricultural Research Institute (ICAR), IndiaPatricia Corral-Martínez, Instituto Universitario de Conservación y Mejora de la Agrodiversidad Valenciana, Spain
Francesca Taranto, Research Centre for Industrial Crops (CRA), Italy
Copyright © 2018 Naresh, Lin, Lin, Wang, Schafleitner, Kilian and Kumar. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Sanjeet Kumar, c2FuamVldC5rdW1hckB3b3JsZHZlZy5vcmc=
 Ponnam Naresh
Ponnam Naresh Shih-wen Lin2
Shih-wen Lin2 Yen-wei Wang
Yen-wei Wang Sanjeet Kumar
Sanjeet Kumar