- 1The State Key Laboratory of Plant Cell and Chromosome Engineering, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing, China
- 2College of Life Sciences, University of Chinese Academy of Sciences, Beijing, China
Iron, as an essential micronutrient, is required by all living organisms. In plants, the deficiency and excess of iron will impair their growth and development. For maintaining a proper intracellular iron concentration, plants evolved different regulation mechanisms to tightly control iron uptake, translocation and storage. FIT, a bHLH transcription factor, is the master regulator of the iron deficiency responses and homeostasis in Arabidopsis. It interacts with different proteins, functioning in controlling the expression of various genes involved in iron uptake and homeostasis. In this review, we summarize the recent progress in the studies of FIT and FIT-binding proteins, and give an overview of FIT-regulated network in iron deficiency response and homeostasis.
Introduction
Iron plays an essential role in a variety of biologic processes in plants, such as photosynthesis, respiration, nitrogen fixation and vitamin synthesis (Schmidt, 2003; Hansch and Mendel, 2009; Jeong and Guerinot, 2009; Zhao et al., 2011). Although iron is the fourth most abundant element in the earth’s crust, the deficiency of iron is one of the factors mostly limiting plant growth because it is mainly present as insoluble form Fe3+ and not available for plants in soil, especially in the neutral and basic soil. In order to overcome iron deficiency, plant developed the reduction-based (strategy I) and chelation-based (strategy II) mechanisms to mobilize and enhance iron uptake from soil (Marschner et al., 1986; Römheld and Marschner, 1986). All dicots and non-graminaceous plants use the strategy I mechanism whereas grasses apply the strategy II. Under iron limitation condition, the strategy I plants secrete proton and phenolic compounds from roots to rhizosphere for increasing the solubility of ferric iron (Santi and Schmidt, 2009; Fourcroy et al., 2014); the mobilized ferric iron will then be reduced to ferrous iron on root surface by the ferric chelate reductase, such as Ferric Reduction Oxidase2 (FRO2) (Robinson et al., 1999); finally, the reduced ferrous iron will be transported into root epidermal cells by the metal transporter Iron-Regulated Transporter1 (IRT1) (Vert et al., 2002). The strategy II plants (grasses) secrete phytosiderophores (PS) from roots into soil to chelate Fe(III), and the PS-Fe(III) complex will be transported into root cells by YS1 under the stress of iron deficiency (Takagi et al., 1984; Mori and Nishizawa, 1987; Curie et al., 2001). On the other hand, excessive iron can generate oxidative stress in cells and become toxic for plants. Therefore, the uptake, translocation and storage of iron in plants should be carefully regulated.
In strategy I plants, the expression of many genes is induced or upregulated at transcriptional level under iron limitation. In the processes, transcription factors are required and play important roles. FER, the first isolated regulatory gene of iron homeostasis from tomato, encodes a typical basic helix-loop-helix protein (Ling et al., 2002). The FER-null mutant T3238fer, which was caused by the insertion of a Ty1-copia-like element in the first exon of FER (Cheng et al., 2009), lost the whole iron deficiency responses. The expression of ferric-chelate reductase LeFRO1 and ferrous transporter LeIRT1 was not able to be upregulated in the mutant at the stress of iron deficiency (Ling et al., 2002; Li et al., 2004). The mutant showed severe chlorosis and died at young seedling stage under normal culture condition. FIT (AtbHLH29/FRU), a FER ortholog in Arabidopsis, was confirmed to enable complementation of the defective function in T3238fer (Yuan et al., 2005). The FIT-null mutant fit-1 and fit-3 generated by T-DNA insertion showed similar phenotypes as tomato T3238fer. The transcription of FRO2 and IRT1 in fit-1 and fit-3 was severely impaired (Colangelo and Guerinot, 2004; Jakoby et al., 2004). Microarray analysis showed that 72 genes upregulated under iron deficiency in wild type were completely or partially deregulated in the mutant fit-1 (such as AHA2, IRT1, IRT2, NRAMP1, and NAS1), some of which were proposed to be its direct targets (Colangelo and Guerinot, 2004; Jakoby et al., 2004; Yuan et al., 2008; Ivanov et al., 2011; Sivitz et al., 2012; Wang et al., 2013). Recently, the list of FIT-regulated genes was expanded (Mai et al., 2016). As a key factor in the transcriptional regulation of Fe uptake and homeostasis, the expression of FIT is tightly regulated at transcriptional and posttranscriptional levels.
In this review, we summarize the recent progress in the studies of FIT and several of its interaction partners in Arabidopsis thaliana (a model plant of strategy I plants), and give an overview of FIT-related regulation pathway in iron deficiency responses and homeostasis of strategy I plants.
Regulators Interacted With Fit in Initiating Iron Deficiency Responses
Ib Subgroup of bHLH Transcription Factors
As shown in introduction, FIT, an ortholog of tomato FER, is a key transcription factor in the control of iron deficiency responses and homeostasis in Arabidopsis thaliana (Colangelo and Guerinot, 2004; Jakoby et al., 2004; Yuan et al., 2005). However, overexpression of FIT didn’t result in obvious phenotypical and physiological alterations, compared to wild type (Colangelo and Guerinot, 2004; Jakoby et al., 2004; Yuan et al., 2005; Meiser et al., 2011; Sivitz et al., 2011; Gratz et al., 2019). Based on these, it was suggested that FIT is essential, but it alone is not sufficient for the activation of iron deficiency responses and the effective iron uptake system in Arabidopsis. Indeed, Yuan et al. (2008) and Wang et al. (2013) revealed that the Ib subgroup of bHLH transcription factors bHLH038, bHLH039, bHLH100, and bHLH101 were able to interact with FIT to form the heterodimer of FIT/bHLH38, FIT/bHLH39, FIT/bHLH100, and FIT/bHLH101 in plant cells. Further, they did the transcriptional activation assay, and showed that the co-expression of FIT with one of the Ib subgroup of bHLH transcription factors enabled to activate GUS expression driven by either the FRO2 or the IRT1 promoter in yeast cells, suggesting that the formed heterodimer complex could bind to the promoter of iron uptake genes FRO2 and IRT1, and directly activated their expression (Yuan et al., 2008; Wang et al., 2013). The expression of the four Ib bHLH genes was dependent on iron status, and strongly upregulated in root and leaf under iron starvation (Vorwieger et al., 2007; Wang et al., 2007). The plants double overexpressing FIT with bHLH38, bHLH39, bHLH100, or bHLH101 changed the expression patterns of FRO2 and IRT1 from induced to a constitutive expression regardless iron status (Yuan et al., 2008; Wang et al., 2013). Phenotypically, the transgenic plants revealed a higher ferric-chelate reductase activity in root and accumulated more iron in shoot than wild type and the plants overexpressing of bHLH38, bHLH39, bHLH100, or bHLH101 alone. The four Ib bHLH genes have a high sequence identity, and function redundantly in the regulation of iron uptake and homeostasis. Therefore, the single and double mutants of the four genes did not show obviously phenotypical and physiological alterations, whereas their triple mutant bHLH38/100/101 and bHLH39/100/101 exhibited sensitivity to iron limitation and chlorotic phenotype, compared to wild type (Wang et al., 2013). Additionally, they found that the four bHLH proteins possessed different significance in regulation of iron-deficiency responses and uptake (bHLH 39 > bHLH101 > bHLH38 > bHLH100). The similar results were also reported by Sivitz et al. (2012) and Maurer et al. (2014). They did not observed an obvious effect on the expression of iron acquisition genes in double knocking out-mutant bhlh100bhlh101 and triple knocking-out mutant bhlh39bhlh100bhlh101 (3xbhlh).
In addition to the activation function, Cui et al. (2018) recently reported that Ib bHLH proteins have a stabilization function for FIT in plant cells. They detected abundant FIT protein accumulation in roots of plants overexpressing FIT and bHLH38, but not in the roots of plants overexpressing FIT alone under iron-sufficient conditions. Further, they also observed that the roots of bHLH38-overexpressing plants accumulated much more FIT under iron sufficiency and protected JA-induced FIT degradation under JA treatment, compared to wild type. Based on the data, they speculated that the Ib subgroup of bHLH transcription factors might play a role in the protection of FIT from degradation. Alternatively, the phenomenon may be caused by higher expression of FIT in bHLH38-overexpressing plants and FIT and bHLH38 double overexpressing plants. However, as reported by Yuan et al. (2008), over expression of bHLH038 did not change the expressional level of FIT, compared with wild type. Anyway, more experiments are needed to confirm it.
The expression of the Ib bHLH genes was controlled genetically and epigenetically at transcription level. Shk1 binding protein 1 (SKB1/AtPRMT5), which catalyzes the symmetric dimethylation of histone H4R3 (H4R3sme2), was identified to be involved in iron homeostasis in Arabidopsis, while the SKB1 lesion mutant exhibited higher iron content in shoots and more tolerance to iron deficiency, compared to wild type (Fan et al., 2013). The expression of SKB1 was not dependent on iron, but the level of H4R3sme2 mediated by SKB1 was clearly related to iron status in plants. Chromatin immunoprecipitation (ChIP) and genome-wide ChIP-seq results showed that SKB1 associated with the chromatin in the promoter region of bHLH38, bHLH39, bHLH100 and bHLH101, and symmetrically dimethylated histone H4R3 (Fan et al., 2013). Iron deficiency may cause an increase in the disassociation of SKB1 from chromatin of the four Ib bHLH genes and resulted in a decrease of H4R3sme2 level, thereby elevating their transcription and enhancing iron uptake.
At genetic control, a recent report showed that the expression of bHLH38, bHLH39, bHLH100, and bHLH101 was positively regulated by bHLH034, bHLH104, bHLH105, and bHLH115, which are classified as the IVc subgroup of bHLH transcription factors. The mutants of IVc bHLH transcription factors displayed sensitivity to iron deficiency, including reduced root length, chlorotic leaves, and decreased ferric chelate reductase activity (Zhang et al., 2015; Li et al., 2016; Liang et al., 2017). Under both Fe-sufficient and Fe-deficient conditions, the transcript levels of bHLH38, bHLH39, bHLH100, and bHLH101 were always lower in the mutants of IVc bHLH transcription factors than in wild-type plants. Transient expression assays and ChIP experiments confirmed that IVc bHLH transcription factors were able to directly activate the promoters of bHLH38, bHLH39, bHLH100, and bHLH101 (Zhang et al., 2015; Li et al., 2016; Liang et al., 2017).
EIN3 and EIL1
As known for a long time, ethylene as a phytohormone plays important roles in iron deficiency responses. Its molecular mechanism has been elucidated recently. Lingam et al. (2011) reported that EIN3 (Ethylene Insensitive 3) and EIL1 (EIN3-Like 1), two transcription factors involved in the ethylene signaling pathway, can interact with FIT, and showed that the interaction of FIT with EIN3 and EIL1 was required for FIT accumulation, and contributes to the expression of FIT-downstream genes. Loss function of EIN3 and EIL1 led to the significant reduction of FIT accumulation even under iron deficiency condition.
MED16 and MED25
During the transcription process, Mediator, a large protein complex comprising more than 20 subunits, is required and functions as a bridge connecting the RNA polymerase II (Pol II) complex and specific transcriptional activators (Asturias et al., 1999). Through interaction with specific transcription activators, Mediator coordinates and transfers the developmental and environmental signals to the transcriptional machinery to regulate the expression of corresponding genes (Bäckström et al., 2007; Mathur et al., 2011; Zhang et al., 2012). MED16 (Mediator subunit 16) and MED25 (Mediator subunit 25), organized in the tail module of the Mediator, have been recently identified to be involved in the regulation of Fe-dependent gene expression (Yang et al., 2014; Zhang et al., 2014). Loss of MED16 or MED25 led to a declined expression of some iron deficiency response genes, such as FRO2, IRT1 and AHA2, under iron deficiency condition. The mutant med16 and med25 showed hypersensitivity to the stress of iron limitation. Zhang et al. (2014) demonstrated that MED16 functioned in the interaction with FIT to enhance the expression of FIT-dependent genes by recruiting FIT-bHLH complex to their promoters. Additionally, Yang et al. (2014) found that MED25 was able to interact with MED16, EIN3 and EIL1, which may play some roles in FIT stabilization.
CIPK11
Cytosolic Ca2+ as the second messenger is an important signaling molecule to transduce developmental and environmental cues to activate physiological processes in plants. Recently, Tian et al. (2016) and Gratz et al. (2019) used the method of fluorescence resonance energy transfer (FRET), and observed an increased cytoplasm Ca2+ concentration in primary roots under iron deficiency. Further, they physiologically characterized the loss-of-function mutants of CIPK11, CIPK23, and CBL1/CBL9 on iron deficiency medium, and demonstrated that calcium-dependent CBL-CIPK pathway was involved in response to iron deficiency in Arabidopsis. With yeast-two-hybrid and BiFC assay, Gratz et al. (2019) confirmed that CIPK11 was positively involved in iron deficiency response through directly interaction with FIT. CIPK11 transcript abundance was significantly elevated under iron deficiency, and its expression pattern was similarly to the FIT in the early differentiation root zone, in which iron uptake occurs predominantly (Gratz et al., 2019). Loss of CIPK11 function resulted in the partial inactivation of the iron deficiency responses and reduced the iron content in seed. In vitro assay displayed that CIPK11 as a kinase enabled phosphorylation of FIT at Ser271 amino acid (Gratz et al., 2019). Further, using a series of molecular methods including fit-3 mutant complementation assay, protein homo- and hetero-dimerization assay in yeast and planta, transcription self-activation capacity assay and fluorescence recovery after photobleaching analysis (Gratz et al., 2019), they revealed that phosphorylation of Ser272 amino acid modulated FIT nuclear accumulation, homo-dimerization, interaction with bHLH039 and transcriptional activity, and proposed that FIT phosphorylation status determines its activity.
Regulators Interacted With Fit in Repressing Iron Deficiency Responses
ZAT12
As known, H2O2 as a signal is involved in many abiotic stresses (Mittler and Blumwald, 2015). Increase in H2O2 accumulation was observed in plant roots under the deficiency of potassium, nitrogen, phosphorus and iron (Ranieri et al., 2001; Shin et al., 2005; Sun et al., 2007). Recently, Le et al. (2016) reported that zinc finger of Arabidopsis thaliana 12 (ZAT12) functioned in the negative regulation of plant responses to prolonged iron deficiency stress. ZAT12 is a marker of abiotic stress and its expression is induced by oxidative stress. Thus, a molecular connection between Fe deficiency and the oxidative stress response was illustrated. ZAT12 was expressed mainly in the root early differentiation and elongation zone, and responded to iron deficiency in the late stage, compared with Fe sufficiency. Using yeast-two-hybrid approach and BiFC assay in plant, they demonstrated that the EAR motif of ZAT12 interacted with FIT-C terminus. Based on the result that FIT expression was down-regulated and ZAT12 expression was up-regulated under iron deficiency with H2O2 stress, ZAT12 should be a repressor of FIT. The formation of FIT-ZAT12 complexes in root cells would inactivate the FIT under prolonged Fe deficiency condition.
DELLA
Iron deficiency will trigger obvious changes in root architecture, such as reduced primary root length and increased frequency of root hairs, as well as high affinity iron uptake mechanism. Exogenous gibberellin (GA) treatment has been shown to promote the expression of IRT1, FRO2, bHLH38, and bHLH39 in the GA-deficient mutant ga3ox1ga3ox2 (the double-knockout mutant of GA biosynthetic gene GA3OX1 and GA3OX2 which are involved in later steps of the gibberellic acid biosynthesis), although the expression of FIT was not induced by the application of GA (Matsuoka et al., 2014). Recently, Wild et al. (2016) demonstrated that GA signaling was involved in the iron deficiency response by interaction of DELLA with protein FIT, bHLH38, and bHLH39. DELLA-FIT interaction did not interfere with the formation of FIT-Ib bHLH heterodimer, but impaired the binding capacity of the FIT-Ib bHLH complex to corresponding promoters. DELLA protein exists in all tissues of root under normal condition. Under iron deficiency, DELLA protein will accumulate in the root meristem to repress the root growth, and degrade in the epidermis cells of the differentiation zone, relieving FIT from the DELLA-FIT complex for activating the expression of iron uptake genes (Wild et al., 2016).
FBP
Under iron deficiency, plants will upregulate the expression of Fe transporter IRT1 in strategy I plants. Due to the low substrate specificity of IRT1 protein, other divalent transition metals, such as Zn, will be transported into root cells. Thus, the excessive Zn will be acquired into root cells from the environment, and result in toxicity to the plant under the condition of iron limitation. To keep the Zn balance and detoxify in cells, plant will up-regulate the expression of some genes related to sequestration and chelation of Zn. Chen et al. (2018) used yeast-two-hybrid approach and identified a novel FIT-binding protein (FBP), which is involved in the distribution and balance of Fe and Zn homeostasis. They confirmed that FBP sequestered FIT to obstruct the heterodimer formation of FIT with Ib bHLH protein in the root stele and negatively regulated the expression of NAS1, NAS2, and NAS4 genes to balance the Fe and Zn homeostasis in Arabidopsis. FBP transcript abundance significantly reduced along with increased Zn concentration. The plants with loss function of FBP displayed the elevated expression of NAS genes, and tolerance to excessive Zn stress (Chen et al., 2018). Further, they did DNA-protein binding and transcriptional activation analysis, and confirmed that FBP inhibited the binding of FIT-Ib bHLH heterodimer to the promoters of NAS genes. The negative regulation caused by sequestration of FIT is restricted in the root stele, but not in epidemic cells, because FBP only expressed in the root stele. Considering that FIT protein accumulation was comparable in roots of FBP-null mutant fbp with wild type under high Zn condition (Chen et al., 2018), the main function of FBP looks like to inactivate FIT by blocking heterodimer formation with Ib bHLH proteins, but not for its degradation.
IVa bHLH Transcription Factors
Jasmonic acid (JA) as a negative factor represses the upregulated expression of FIT, IRT1 and FRO2 under iron deficiency (Maurer et al., 2011). The depressed regulation mechanism has been elucidated recently. Cui et al. (2018) identified the IVa subgroup of bHLH transcription factors (bHLH18, bHLH19, bHLH20, and bHLH25), which are involved in the repressed regulation of iron deficiency responses under JA present. The four IVa bHLH genes expressed mainly in roots and responded to JA. Characterization of the single, double, triple and quadruple mutants of the four IVa bHLH genes revealed that the four bHLH transcription factors functioned redundantly in the JA-mediated inhibition of iron acquisition in Arabidopsis. Immunoblot analysis showed that bHLH18, bHLH19, bHLH20, and bHLH25 interacted with FIT and promoted its degradation mediated by JA under iron deficiency. Based on that the plants overexpressing Ib bHLH genes can alleviate JA-mediated FIT protein degradation, and exhibit more tolerance to iron deficiency under JA present, Cui et al. (2018) speculated that the main function of the four IVa bHLH proteins might be to compete with Ib bHLH proteins to bind FIT and stimulates its degradation. Further, they demonstrated that JA-promoted degradation of the FIT protein is through the 26S proteasome pathway. Additionally, they showed that MYC2 and JAR1, critical components of the JA signaling pathway, differentially regulated the expression of bHLH18, bHLH19, bHLH20, and bHLH25 to modulate FIT protein accumulation, and MYC2 depressed the expression of FIT and the four Ib bHLH genes at transcription level. Taken together, upregulated expression of IVa bHLH genes and downregulated expression of FIT and Ib bHLH genes under JA present result in lower accumulation of activators FIT and Ib bHLH proteins, consequently downregulating the expression of the iron-uptake genes IRT1 and FRO2. Considering that JA is an important integrator that balances plant growth and defense responses, the JA-mediated suppression of iron acquisition in plants should play some important roles in defense responses against biotic and abiotic stresses to promote plant survival.
Conclusion
As described above, the proteins encoded by 15 different genes have been confirmed to bind FIT during last several years in Arabidopsis. Their functions can be generally summarized into three categories, and outlined in Figure 1. The first group contains the four Ib bHLH proteins, CIPK11, MED16, EIN3, and EIL1. The eight proteins as positive regulators bind to FIT, and function as stabilizer and activator of FIT in iron deficiency responses and homeostasis. The second one is the four IVa bHLH proteins mediated by JA signal pathway and ZAT12. They play as negative regulators to enhance the degradation of FIT protein via interaction with FIT. The last group includes DELLA and FBP, which compete with Ib bHLH proteins to bind FIT, and inactivate its functions. These findings obviously reveal a new insight for understanding the regulation mechanisms of iron uptake and homeostasis in strategy I plants. Although a great progress has been made in studying the molecular regulation mechanisms of iron deficiency responses and homeostasis in strategy I plants in the past two decades, there are still many questions that are not answered. For example, iron uptake and homeostasis are tightly regulated dependent on iron status in plants, how does plant sense the iron status in cells and in environment, and how is the iron signal transduced in plants? As shown in Figure 1, some proteins, such as DELLA and FBP, involved in iron deficiency responses and homeostasis function only in a specific region or cell type of root, we still do not know what is the molecular mechanism(s) accurately controlling their spatial expression and functional patterns. As a key regulator, expression of FIT is controlled at transcriptional and posttranscriptional level. However, we do still not well know the regulation mechanisms.
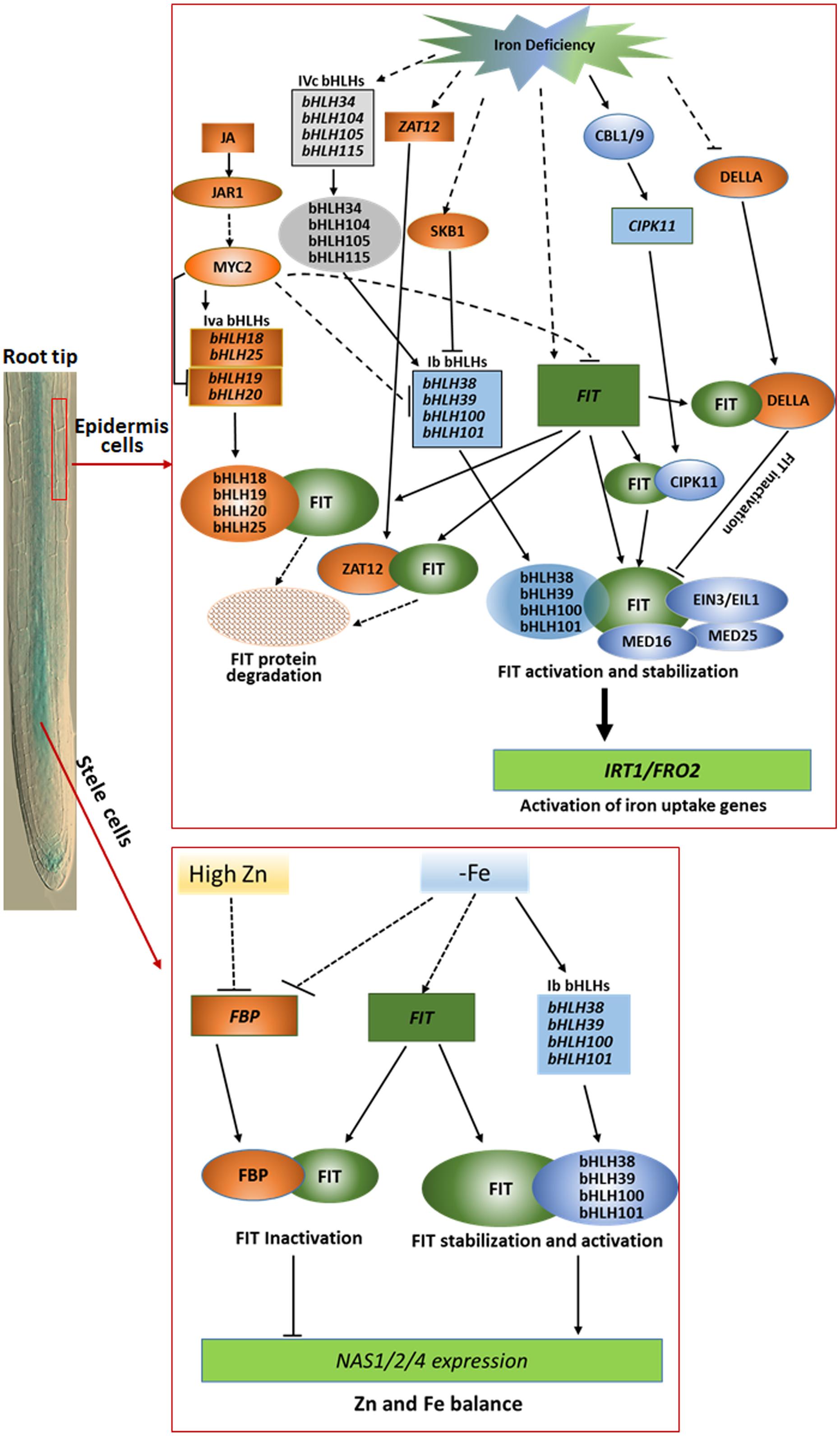
Figure 1. Outline of the regulation networks and functions of FIT-binding proteins in iron deficiency responses and homeostasis in root epidermis cells (upper part) and stele cells (lower part). In root epidermal cells, Ib bHLHs, MED16, MED25, EIN3/EIL1, and CIPK11 as positive modulators interacts with the key regulator FIT, functioning in activation of the expression of iron acquisition genes such as FRO2 and IRT1 under iron limitation condition, whereas DELLA, ZAT12, and IVa bHLHs as negative modulators compete with Ib bHLHs to bind FIT for negative regulation of FIT activity for avoidance of excessive iron uptake. In stele cells, FBP as a negative regulator sequestered FIT to obstruct the heterodimer formation of FIT with Ib bHLH protein to downregulate the expression of NAS1, NAS2, and NAS4 for balancing Fe and Zn homeostasis in Arabidopsis under low iron or high Zn stress condition. The rectangles represent transcripts of the corresponding genes, and the ovals mean proteins indicated. The blue and orange color depicted positive and negative regulators, respectively. The solid and dotted lines separately indicate the known and unknown regulation processes.
Author Contributions
Both authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
Funding
This work was supported by the National Natural Science Foundation of China (Grant No. 31870225).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Asturias, F. J., Jiang, Y. W., Myers, L. C., Gustafsson, C. M., and Kornberg, R. D. (1999). Conserved structures of mediator and RNA polymerase II holoenzyme. Science 283, 985–987. doi: 10.1126/science.283.5404.985
Bäckström, S., Elfving, N., Nilsson, R., Wingsle, G., and Bjorklund, S. (2007). Purification of a plant mediator from Arabidopsis thaliana identifies PFT1 as the Med25 subunit. Mol. Cell. 26, 717–729. doi: 10.1016/j.molcel.2007.05.007
Chen, C. L., Cui, Y., Cui, M., Zhou, W. J., Wu, H. L., and Ling, H. Q. (2018). A FIT-binding protein is involved in modulating iron and zinc homeostasis in Arabidopsis. Plant Cell Environ. 41, 1698–1714. doi: 10.1111/pce.13321
Cheng, X., Zhang, D., Cheng, Z., Keller, B., and Ling, H. Q. (2009). A new family of Ty1-copia-like retrotransposons originated in the tomato genome by a recent horizontal transfer event. Genetics 181, 1183–1193. doi: 10.1534/genetics.108.099150
Colangelo, E. P., and Guerinot, M. L. (2004). The essential basic helix-loop-helix protein FIT1 is required for the iron deficiency response. Plant Cell 16, 3400–3412. doi: 10.1105/tpc.104.024315
Cui, Y., Chen, C. L., Cui, M., Zhou, W. J., Wu, H. L., and Ling, H. Q. (2018). Four IVa bHLH transcription factors are novel interactors of FIT and mediate JA inhibition of iron uptake in Arabidopsis. Mol. Plant 11, 1166–1183. doi: 10.1016/j.molp.2018.06.005
Curie, C., Panaviene, Z., Loulergue, C., Dellaporta, S. L., Briat, J.-F., and Walker, E. L. (2001). Maize yellow stripe1 encodes a membrane protein directly involved in Fe (III) uptake. Nature 409, 346–349. doi: 10.1038/35053080
Fan, H., Zhang, Z., Wang, N., Cui, Y., Sun, H., Liu, Y., et al. (2013). SKB1/PRMT5 mediated histone H4R3 dimethylation of Ib subgroup bHLH genes negatively regulates iron homeostasis in Arabidopsis thaliana. Plant J. 77, 209–221. doi: 10.1111/tpj.12380
Fourcroy, P., Siso-Terraza, P., Sudre, D., Saviron, M., Reyt, G., Gaymard, F., et al. (2014). Involvement of the ABCG37 transporter in secretion of scopoletin and derivatives by Arabidopsis roots in response to iron deficiency. New Phytol. 201, 155–167. doi: 10.1111/nph.12471
Gratz, R., Manishankar, P., Ivanov, R., Koster, P., Mohr, I., Trofimov, K., et al. (2019). CIPK11-dependent phosphorylation modulates FIT activity to promote Arabidopsis iron acquisition in response to calcium signaling. Dev. Cell 48, 726–740. doi: 10.1016/j.devcel.2019.01.006
Hansch, R., and Mendel, R. R. (2009). Physiological functions of mineral micronutrients (Cu, Zn, Mn, Fe, Ni, Mo, B, Cl). Curr. Opin. Plant Biol. 12, 259–266. doi: 10.1016/j.pbi.2009.05.006
Ivanov, R., Brumbarova, T., and Bauer, P. (2011). Fitting into the harsh reality: regulation of iron deficiency responses in dicotyledonous plants. Mol. Plant 5, 27–42. doi: 10.1093/mp/ssr065
Jakoby, M., Wang, H. Y., Reidt, W., Weisshaar, B., and Bauer, P. (2004). FRU (BHLH029) is required for induction of iron mobilization genes in Arabidopsis thaliana. FEBS Lett. 577, 528–534. doi: 10.1016/j.febslet.2004.10.062
Jeong, J., and Guerinot, M. L. (2009). Homing in on iron homeostasis in plants. Trends Plant Sci. 14, 280–285. doi: 10.1016/j.tplants.2009.02.006
Le, C. T., Brumbarova, T., Ivanov, R., Stoof, C., Weber, E., Mohrbacher, J., et al. (2016). Zinc finger of Arabidopsis thaliana12 (ZAT12) interacts with fer-LIKE iron deficiency-induced transcription factor (FIT) linking iron deficiency and oxidative stress responses. Plant Physiol. 170, 540–557. doi: 10.1104/pp.15.01589
Li, L., Cheng, X., and Ling, H. Q. (2004). Isolation and characterization of Fe(III)-chelate reductase gene LeFRO1 in tomato. Plant Mol. Biol. 54, 125–136. doi: 10.1023/b:plan.0000028774.82782.16
Li, X., Zhang, H., Ai, Q., Liang, G., and Yu, D. (2016). Two bHLH transcription factors, bHLH34 and bHLH104, regulate iron homeostasis in Arabidopsis thaliana. Plant Physiol. 170, 2478–2493. doi: 10.1104/pp.15.01827
Liang, G., Zhang, H., Li, X., Ai, Q., and Yu, D. (2017). bHLH transcription factor bHLH115 regulates iron homeostasis in Arabidopsis thaliana. J. Exp. Bot. 68, 1743–1755. doi: 10.1093/jxb/erx043
Ling, H. Q., Bauer, P., Bereczky, Z., Keller, B., and Ganal, M. (2002). The tomato fer gene encoding a bHLH protein controls iron-uptake responses in roots. Proc. Natl. Acad. Sci. U.S.A. 99, 13938–13943. doi: 10.1073/pnas.212448699
Lingam, S., Mohrbacher, J., Brumbarova, T., Potuschak, T., Fink-Straube, C., Blondet, E., et al. (2011). Interaction between the bHLH transcription factor FIT and ethylene insensitive3/ethylene insensitive3-like1 reveals molecular linkage between the regulation of iron acquisition and ethylene signaling in Arabidopsis. Plant Cell 23, 1815–1829. doi: 10.1105/tpc.111.084715
Mai, H. J., Pateyron, S., and Bauer, P. (2016). Iron homeostasis in Arabidopsis thaliana: transcriptomic analyses reveal novel FIT-regulated genes, iron deficiency marker genes and functional gene networks. BMC Plant Biol. 16:211. doi: 10.1186/s12870-016-0899-9
Marschner, H., Römheld, V., and Kissel, M. (1986). Different strategies in higher plants in mobilization and uptake of iron. J. Plant Nutr. 9, 695–713. doi: 10.1080/01904168609363475
Mathur, S., Vyas, S., Kapoor, S., and Tyagi, A. K. (2011). The mediator complex in plants: structure, phylogeny and expression profiling of representative genes in a dicot (Arabidopsis thaliana) and a monocot (Oryza sativa) during reproduction and abiotic stress. Plant Physiol. 157, 1609–1627. doi: 10.1104/pp.111.188300
Matsuoka, K., Furukawa, J., Bidadi, H., Asahina, M., Yamaguchi, S., and Satoh, S. (2014). Gibberellin-induced expression of Fe uptake-related genes in Arabidopsis. Plant Cell Physiol. 55, 87–98. doi: 10.1093/pcp/pct160
Maurer, F., Muller, S., and Bauer, P. (2011). Suppression of fe deficiency gene expression by jasmonate. Plant Physiol. Biochem. 49, 530–536. doi: 10.1016/j.plaphy.2011.01.025
Maurer, F., Naranjo Arcos, M. A., and Bauer, P. (2014). Responses of a triple mutant defective in three iron deficiency-induced basic helix-loop-helix genes of the subgroup Ib (2) to iron deficiency and salicylic acid. PLoS One 9:e99234. doi: 10.1371/journal.pone.0099234
Meiser, J., Lingam, S., and Bauer, P. (2011). Posttranslational regulation of the iron deficiency basic helix-loop-helix transcription factor FIT is affected by iron and nitric oxide. Plant Physiol. 157, 2154–2166. doi: 10.1104/pp.111.183285
Mittler, R., and Blumwald, E. (2015). The roles of ROS and ABA in systemic acquired acclimation. Plant Cell 27, 64–70. doi: 10.1105/tpc.114.133090
Mori, S., and Nishizawa, N. (1987). Methionine as a dominant precursor of phytosiderophores in graminaceae plants. Plant Cell Physiol. 28, 1081–1092.
Ranieri, A., Castagna, A., Baldan, B., and Soldatini, G. F. (2001). Iron deficiency differently affects peroxidase isoforms in sunflower. J. Exp. Bot. 52, 25–35. doi: 10.1093/jxb/52.354.25
Robinson, N. J., Procter, C. M., Connolly, E. L., and Guerinot, M. L. (1999). A ferric-chelate reductase for iron uptake from soils. Nature 397, 694–697. doi: 10.1038/17800
Römheld, V., and Marschner, H. (1986). Evidence for a specific uptake system for iron phytosiderophores in roots of grasses. Plant Physiol. 80, 175–180. doi: 10.1104/pp.80.1.175
Santi, S., and Schmidt, W. (2009). Dissecting iron deficiency-induced proton extrusion in Arabidopsis roots. New Phytol. 183, 1072–1084. doi: 10.1111/j.1469-8137.2009.02908.x
Schmidt, W. (2003). Iron solutions: acquisition strategies and signaling pathways in plants. Trends Plant Sci. 8, 188–193. doi: 10.1016/s1360-1385(03)00048-7
Shin, R., Berg, R. H., and Schachtman, D. P. (2005). Reactive oxygen species and root hairs in Arabidopsis root response to nitrogen, phosphorus and potassium deficiency. Plant Cell Physiol. 46, 1350–1357. doi: 10.1093/pcp/pci145
Sivitz, A., Grinvalds, C., Barberon, M., Curie, C., and Vert, G. (2011). Proteasome mediated turnover of the transcriptional activator FIT is required for plant iron-deficiency responses. Plant J. 66, 1044–1052. doi: 10.1111/j.1365-313X.2011.04565.x
Sivitz, A. B., Hermand, V., Curie, C. V., and Vert, G. (2012). Arabidopsis bHLH100 and bHLH101 control iron homeostasis via a FIT-independent pathway. PLoS One 7:e44843. doi: 10.1371/journal.pone.0044843
Sun, B., Jing, Y., Chen, K., Song, L., Chen, F., and Zhang, L. (2007). Protective effect of nitric oxide on iron deficiency-induced oxidative stress in maize (Zea mays). J. Plant Physiol. 164, 536–543. doi: 10.1016/j.jplph.2006.02.011
Takagi, S., Nomoto, K., and Takemoto, T. (1984). Physiological aspect of mugineic acid, a possible phytosiderophore of graminaceous plants. J. Plant Nutr. 7, 469–477. doi: 10.1080/01904168409363213
Tian, Q., Zhang, X., Yang, A., Wang, T., and Zhang, W. H. (2016). CIPK23 is involved in iron acquisition of Arabidopsis by affecting ferric chelate reductase activity. Plant Sci. 246, 70–79. doi: 10.1016/j.plantsci.2016.01.010
Vert, G., Grotz, N., Dedaldechamp, F., Gaymard, F., Guerinot, M. L., Briat, J. F., et al. (2002). IRT1, an Arabidopsis transporter essential for iron uptake from the soil and for plant growth. Plant Cell 14, 1223–1233. doi: 10.1105/tpc.001388
Vorwieger, A., Gryczka, C., Czihal, A., Douchkov, D., Tiedemann, J., Mock, H. P., et al. (2007). Iron assimilation and transcription factor controlled synthesis of riboflavin in plants. Planta 226, 147–158. doi: 10.1007/s00425-006-0476-9
Wang, H. Y., Klatte, M., Jakoby, M., Baumlein, H., Weisshaar, B., and Bauer, P. (2007). Iron deficiency-mediated stress regulation of four subgroup Ib BHLH genes in Arabidopsis thaliana. Planta 226, 897–908. doi: 10.1007/s00425-007-0535-x
Wang, N., Cui, Y., Liu, Y., Fan, H., Du, J., Huang, Z., et al. (2013). Requirement and functional redundancy of Ib subgroup bHLH proteins for iron deficiency responses and uptake in Arabidopsis thaliana. Mol. Plant 6, 503–513. doi: 10.1093/mp/sss089
Wild, M., Davie‘re, J.-M., Regnault, T., Sakvarelidze-Achard, L., Carrera, E., Lopez Diaz, I., et al. (2016). Tissue-specific regulation of gibberellin signaling fine-tunes Arabidopsis iron-deficiency responses. Dev. Cell 37, 190–200. doi: 10.1016/j.devcel.2016.03.022
Yang, Y., Ou, B., Zhang, J. Z., Si, W., Gu, H. Y., Qin, G. J., et al. (2014). The Arabidopsis mediator subunit MED16 regulates iron homeostasis by associating with EIN3/EIL1 through subunit MED25. Plant J. 77, 838–851. doi: 10.1111/tpj.12440
Yuan, Y., Wu, H., Wang, N., Li, J., Zhao, W., Du, J., et al. (2008). FIT interacts with AtbHLH38 and AtbHLH39 in regulating iron uptake gene expression for iron homeostasis in Arabidopsis. Cell Res. 18, 385–397. doi: 10.1038/cr.2008.26
Yuan, Y. X., Zhang, J., Wang, D. W., and Ling, H. Q. (2005). AtbHLH29 of Arabidopsis thaliana is a functional ortholog of tomato FER involved in controlling iron acquisition in strategy I plants. Cell Res. 15, 613–621. doi: 10.1038/sj.cr.7290331
Zhang, J., Liu, B., Li, M., Feng, D., Jin, H., Wang, P., et al. (2015). The bHLH transcription factor bHLH104 interacts with IAA-LEUCINE RESISTANT3 and modulates iron homeostasis in Arabidopsis. Plant Cell 27, 787–805. doi: 10.1105/tpc.114.132704
Zhang, X., Wang, C., Zhang, Y., Sun, Y., and Mou, Z. (2012). The Arabidopsis mediator complex subunit16 positively regulates salicylate-mediated systemic acquired resistance and jasmonate/ethylene-induced defense pathways. Plant Cell 24, 4294–4309. doi: 10.1105/tpc.112.103317
Zhang, Y., Wu, H. L., Wang, N., Fan, H. J., Chen, C. L., Cui, Y., et al. (2014). Mediator subunit 16 functions in the regulation of iron uptake gene expression in Arabidopsis. New Phytol. 203, 770–783. doi: 10.1111/nph.12860
Keywords: Arabidopsis, plant nutrition, iron, FIT, FIT-binding protein, transcriptional regulation
Citation: Wu H and Ling H-Q (2019) FIT-Binding Proteins and Their Functions in the Regulation of Fe Homeostasis. Front. Plant Sci. 10:844. doi: 10.3389/fpls.2019.00844
Received: 31 January 2019; Accepted: 12 June 2019;
Published: 26 June 2019.
Edited by:
Sebastien Thomine, Centre National de la Recherche Scientifique (CNRS), FranceReviewed by:
Gang Liang, Xishuangbanna Tropical Botanical Garden (CAS), ChinaPetra Bauer, Saarland University, Germany
Christian Dubos, Institut National de la Recherche Agronomique (INRA), France
Copyright © 2019 Wu and Ling. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hong-Qing Ling, aHFsaW5nQGdlbmV0aWNzLmFjLmNu
 Huilan Wu
Huilan Wu Hong-Qing Ling
Hong-Qing Ling