Abstract
Sweet sorghum is a C4 crop with the characteristic of fast-growth and high-yields. It is a good source for food, feed, fiber, and fuel. On saline land, sweet sorghum can not only survive, but increase its sugar content. Therefore, it is regarded as a potential source for identifying salt-related genes. Here, we review the physiological and biochemical responses of sweet sorghum to salt stress, such as photosynthesis, sucrose synthesis, hormonal regulation, and ion homeostasis, as well as their potential salt-resistance mechanisms. The major advantages of salt-tolerant sweet sorghum include: 1) improving the Na+ exclusion ability to maintain ion homeostasis in roots under salt-stress conditions, which ensures a relatively low Na+ concentration in shoots; 2) maintaining a high sugar content in shoots under salt-stress conditions, by protecting the structures of photosystems, enhancing photosynthetic performance and sucrose synthetase activity, as well as inhibiting sucrose degradation. To study the regulatory mechanism of such genes will provide opportunities for increasing the salt tolerance of sweet sorghum by breeding and genetic engineering.
Introduction
Soil salinization is a compelling environmental problem worldwide (Landi et al., 2017). Over ~6% of the world’s lands are saline and ~20% of irrigated lands are currently salt-affected (Song and Wang, 2015; Yuan et al., 2016a). Salt stress is an important factor determining plant growth and yields (Mahajan and Tuteja, 2005; Deng et al., 2015), by substantially affecting key biological processes, such as photosynthesis (Feng et al., 2014), energy metabolism (Song et al., 2016), protein synthesis, and lipid metabolism (Sui et al., 2010; Sui and Han, 2014; Sui et al., 2018). A limited number of crops can survive in saline land, which can lead to desertification (Flowers and Colmer, 2010). Another two limiting factors for the development of the economy in many developing countries are the shortage of cultivated land and energy crisis (IEA, 2019). At present, these developing countries are facing the problem of “large population and less land”. As a consequence, the development of biomass energy and feed crops is restricted by the reduction of cultivated land resources. If energy crops and feed crops with high salt tolerance can be widely planted on saline soils, it is of strategical importance to provide solutions for addressing the challenges of energy and food security.
Plants have evolved various adaptive mechanisms in response to salt stress, leading to survival in saline land. These mechanisms occur at multiple levels, the molecular level (Yuan et al., 2016b) and the physiological and biochemical levels (Guo et al., 2012; Kong et al., 2016; Leng et al., 2018). At the molecular level, many genes related to salt-stress responses are differentially expressed after plants sense the external salt-stress signals (Zhu, 2001). For example, OsCCC1 plays a significant role in K+ and Cl− homeostasis (Kong et al., 2011), overexpression of SsHKT1;1 (Shao et al., 2014), AtZFP1 (Han et al., 2014), SsCHLAPXs (Pang et al., 2011), and PcAPX (Cao et al., 2017) enhances plant salt tolerance. The differential expressions of these genes then drive the salt-stress responses at the physiological and biochemical levels (Cui et al., 2018). The first step is to block Na+ transport to shoots through apoplastic barriers, including Casparian bands and suberin lamellae (Krishnamurthy et al., 2011). The second step is to dilute Na+ in the cytoplasm through Na+ compartmentalization by transporting Na+ into the vacuole (Blumwald, 2000). The third step is to exclude Na+ outside using a Na+/H+ antiporter in the plasma membrane. The transport of Na+ from roots to leaves can also be restricted by a high-affinity K+ transporter (HKT) (Byrt et al., 2007; Byrt et al., 2014).
Sweet sorghum [Sorghum bicolor (L.) Moench], which was generated from grain sorghum, has been widely cultivated for animal feed production in tropical and subtropical regions and, in particular, has a large planting area in Asia, Africa, and many developing countries (Antonopoulou et al., 2008). India is the largest sorghum grower in the world, with 7.5 million hectares of lands for growing sweet sorghum, followed by Nigeria (7.6 million hectares) and Sudan (6.6 million hectares) (Rao et al., 2013). In America, about 2.6 million hectares were devoted to sorghum for grain and 153,780 hectares were used for sorghum silage (Getachew et al., 2016). In China, about 100,000 hectares of lands were used to plant sweet sorghum. Sweet sorghum has the characteristics of high photosynthetic efficiency (Guo et al., 2018a), high fermentable sugar content in stalks, and high salt tolerance (Vasilakoglou et al., 2011; Sui et al., 2015). As a C4 crop, it has the ability to produce high biomass under stress conditions. Its high nitrogen-use efficiency enables it to produce two to three times more biomass per harvest than the more commonly used silage crop such as maize (Xie and Xu, 2019). Compared with the original feed crops, sweet sorghum presents the advantages of possessing high temperature and drought tolerance levels, high yields, low soil fertility requirements, and good palatability and digestibility (Chen L. et al., 2018). It has been reported that the protein content in sweet sorghum leaves is up to 22% (Ferraris and Charles-Edwards, 1986). A meta-analysis of feeding experiments found that cows fed with sweet sorghum silage could produce more milk (1.64 kg/day/cow) with an improved quality compared with those fed with maize silage (Sánchez-Duarte et al., 2018). Additionally, most sorghum accumulates high amounts of tannins in the seeds. It has been shown that tannins can promote milk production, lambing percentage, and growth of wool, as well as reducing the risk of rumen bloat (Patra and Saxena, 2011). In an in vitro experiment, the supplement of 15 mg tannins in 500 mg of total oven-dried guinea grass reduced methane (CH4) production by 47% (Tan et al., 2011). These findings imply another important advantage of using sweet sorghum silage over other crops: lower methane production and lower damage to the atmosphere. Salt-tolerant sweet sorghum species have stable or increased Brix scores in their stems under salt-stress conditions (Sui et al., 2015). However, salt-sensitive sweet sorghum species often exhibit the decreased net photosynthetic rates and comprehensive utilization degrees of Brix. Salt resistance of sweet sorghum can be improved by the pivotal process of salt exclusion, which induces the accumulation of Na+ in the vacuoles of root parenchymata and limits Na+ transportation into shoots (Yang et al., 2018).
This review will focus on the physiological and biochemical responses of sweet sorghum to salt stress and the salt-tolerance mechanisms, and discuss how this crop plant can be used to improve saline land.
Physiological and Biochemical Responses of Metabolic Processes to Salt Stress
Many important metabolic processes are affected by salt stress at the physiological and biochemical levels. Generally, photosynthesis and growth rate of most plants would be inhibited by salt stress. The regulation of these metabolic processes in response to salt stress could modulate the ability of salt tolerance among sweet sorghum accessions.
Photosynthesis
Photosynthesis is the most important process occurs in the chloroplasts of higher plants. Solar energy can be converted into chemical energy by plants through photosynthesis. Photosynthesis includes light and dark reactions, each of which consists of a series of redox reactions related to energy capture and conversion (Liu, 2016). At the initial step of photosynthesis, light energy is captured by the peripheral light-harvesting complexes, which contain most of the chlorophyll and carotenoid pigments and are peripherally associated with photosystems (PSs) I and II (Figure 1A) (Galka et al., 2012). For many plants such as tomato, Zygophyllum xanthoxylum, and Phaseolus vulgaris, salt stress could affect the chloroplast structures and decrease the chlorophyll content resulting in the reduced photosynthetic rate (Sudhir and Murthy, 2004; Wydrzynski, 2008; Ma et al., 2012).
Figure 1
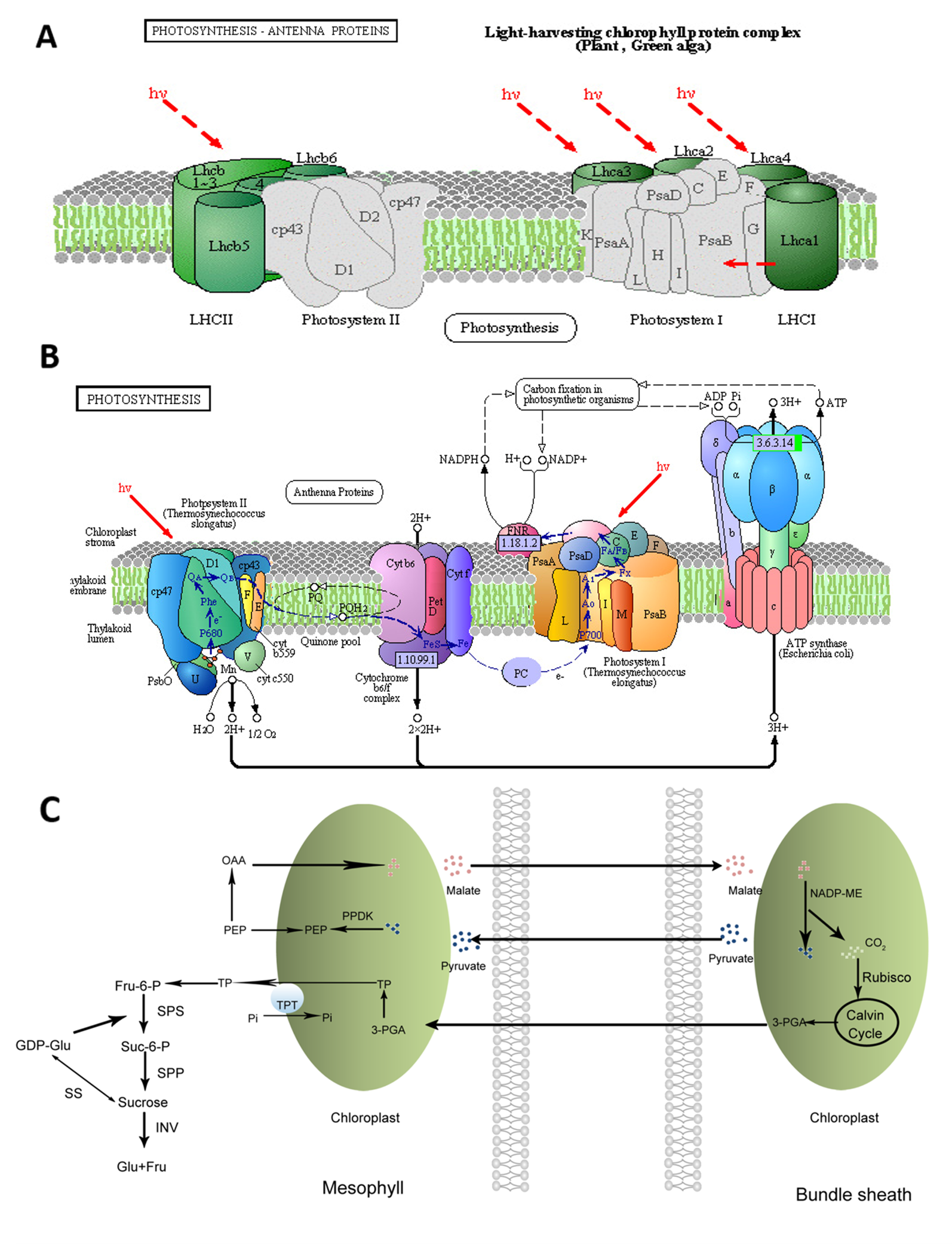
Visualization of pathways related to the accumulation of sucrose in sweet sorghum under salt stress (Repaint refers to Sui et al., 2015). (A): Photosynthetic antenna systems and the light-harvesting process; (B): photosynthetic pathway (Sui et al., 2015); (C): pathway of the carbon fixation in photosynthetic organisms and sucrose biosynthesis.
The decrease in photosynthesis induced by salt stress could be relevant to the stomatal and/or nonstomatal factors in soybean seedlings (Jiao et al., 2017). Salt stress has direct and indirect effects on the chlorophyll content and photosynthetic efficiency of plants. The direct effects are achieved by regulating the activity and expression levels of enzymes involved in chlorophyll biosynthesis and photosynthesis (Figure 1B). The indirect effects are achieved by specific regulating pathways such as antioxidant enzyme systems. It has been reported that the maximum quantum yield of photosystem II (PSII; Fv/Fm), photochemical quenching coefficient (qP), and electron transport rate (ETR) substantially decreased, whereas non-photochemical quenching (qN) increased in sorghum under saline conditions (Netondo et al., 2004). In poplar (Populus spp.), salt stress could inhibit Fv/Fm due to the salt-induced increase of the minimal fluorescence (Fo) and the notable decline of the maximal fluorescence (Fm). It could also result in the decrease of qP but could greatly elevate the coefficient of nonphotochemical qN in the light-adapted state (Wang et al., 2007). In contrast, Fv/Fm was almost not affected by salt stress in rice, whereas qN increased in sensitive cultivars with the increasing salt stress (Dionisio-Sese and Tobita, 2000).
It was also reported that salt stress can affect water potential, stomatal conductance, and sugar accumulation to induce inhibition and ion toxicity (Feng et al., 2014). In salt-sensitive sweet sorghum, salt stress was revealed to significantly decrease the net photosynthetic rate, PSII photochemical efficiency, stomatal conductance, and intercellular CO2 concentration (Sui et al., 2015). In contrast, salt-tolerant sweet sorghum genotypes can still exploit certain mechanisms to minimize these responses throuth apoplastic barriers (Figure 2B). The mechanism of salt-tolerant sweet sorghum for maintaining high photosynthetic efficiency under salt stress may be related to the contributions of several pathways, such as maintaining the stability of the photosynthetic system and enhancing the efficiency of CO2 fixation. In the salt-tolerance and salt-sensitive genotypes, Fv/Fm and the actual PSII efficiency (ΦPSII) were reduced with increasing NaCl concentration, and the decrease was more significant in Roma which is sensitive to salt stress. The chlorophyll content in M-81E sweet sorghum was not altered greatly by 50 mM NaCl, whereas it decreased in Roma (Sui et al., 2015). No significant changes in the photosynthetic rate, stomatal conductance, and intercellular CO2 concentration were observed in M-81E under salt stress. However, these photosynthesis-related indicators in Roma were significantly influenced by salt stress. Additionally, many genes mapped to the photosynthesis pathway showed different expressions in Roma and all of them were down-regulated under salt stress (Sui et al., 2015). Most of these genes are related to the photosystem complexes, the connection between photosystems and light-harvesting proteins, and electron transport chain. Only a few genes in M-81E mapped to the photosynthesis pathway were affected by salt stress, and 2 genes related to the stable assembly of the oxygen-evolving complex and ATP synthase were up-regulated under salt stress (Sui et al., 2015). These results suggest that salt stress could affect the assembly of photosystems and reduce the efficiency of electron transport, resulting in the decrease of ATP and NADPH in plants under salt stress. These effects are particularly critical in salt-sensitive species, whereas salt-tolerant species can protect the formation of photosynthetic assemblies by increasing the expression of particular genes.
Figure 2
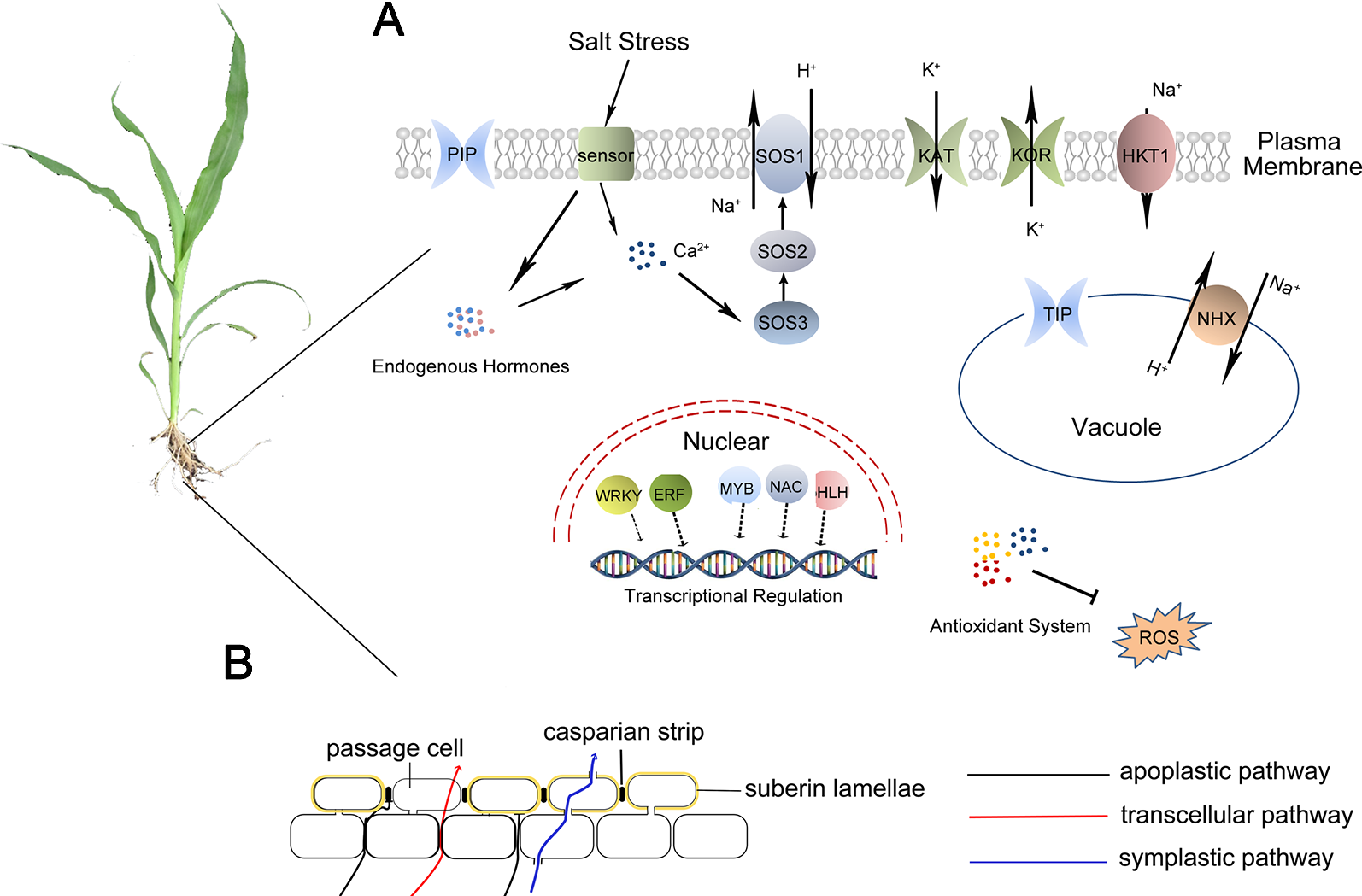
Possible roles of salt resistance in sweet sorghum. (A): Cells of sweet sorghum firstly sense Na+ by an unknown sensor and change the content of intracellular hormones and Ca2+. The expression of some transcription factors, such as NAC, bHLH, MYB, is also initiated. These transcription factors then activate the expression of genes encoding proteins related to salt stress response such as SOS1, HKT1, NHX, ROS scavenging proteins, aquaporins, and some ion channels. (B): Physical barrier effect of root apoplastic barriers can block the apoplastic transpiration bypass flow of water and solutes.
Sweet sorghum possesses the C4 pathway of carbon fixation. CO2 is fixed initially in the mesophyll cells by phosphoenolpyruvate carboxylase (PEPC) to form malic acid. Malic acid can then diffuse into bundle sheath cells, where they are decarboxylated, and the released CO2 is then fixed by Rubisco (Hatch, 1987), the key enzyme in photosynthetic CO2 assimilation (Whitney et al., 2011). The phosphorylation of PEPC depends on the interplay of phosphoenolpyruvate carboxylase kinase (PEPC-k) and a 2A-type protein phosphatase (Carter et al., 1990; Vidal and Chollet, 1997). It has been reported that the PEPC-k content in leaves of sorghum could be induced by salt stress under both light and dark conditions, which contributes to the improvement of carbon fixation efficiency (Echevaria et al., 2001; García-Mauriño et al., 2003). NADP-Malate dehydrogenase (NADP-ME) is a critical enzyme of the C4 pathway, which catalyzes the oxidative decarboxylation of malic acid to provide CO2 to Rubisco. It has been shown that the expression of the gene encoding NADP-ME could be activated by salt stress (Sun et al., 2003; Liu et al., 2007). Recently, it has been revealed that NADP-ME in sweet sorghum is associated with salt tolerance. NADP-ME in M-81E sweet sorghum was activated by salt stress and overexpression of SbNADP-ME increased the photosynthetic capacity in Arabidopsis under salt stress (Guo et al., 2018b). The increase of NADP-ME may lead to an increase of CO2 and pyruvate levels, which enhanced the CO2 fixation efficiency. Additionally, NADPH can also participate in the reactive oxygen species (ROS) metabolism by providing power (Mittler, 2002). Møller reported that the NADPH-specific glutathione reductase (GR) can use NADPH to catalyze glutathione reduction for scavenging ROS (Møller, 2001).
Sucrose Synthesis, Metabolism, and Transportation
Sweet sorghum represents an excellent model for fermentative production due to the fact that its stem contains rich fermentable sugars, such as sucrose, fructose, and glucose (Almodares and Hadi, 2009). Sucrose is the main sugar accumulated through photosynthesis, accounting for ~85% of the total content, in the stems of sweet sorghum (Gnansounou et al., 2005; Kühn and Grof, 2010). The flow direction of triose phosphate formed during photosynthesis determines the distribution of photosynthetic products. It remains in the chloroplast stroma for the completion of the Calvin cycle or to be converted to starch or enters the cytoplasm via triose phosphate translocator and synthesizes sucrose under the action of a series of enzymes, such as sucrose phosphate synthetase (SPS), sucrose synthetase (SS), and invertase (INV). Sucrose is temporarily stored in the vacuoles or is exported using long-distance transport through the phloem (Figure 1C).
Plant organs are generally divided into the source and sink organs (Turgeon, 1989). A source organ possessing photosynthetic activities usually has a net output of light assimilation, which occurs in the mature leaf. Sucrose is the main source of carbon and energy in the epidermal tissue of plants. It is synthesized in the cytoplasm of leaves (source tissues) where it is loaded into the phloem and is transported to sink tissues (Guimaraes et al., 1997; Mccormick et al., 2010). Sucrose can be transported from the phloem to sink tissues through two pathways (Qazi et al., 2012). In sugarcane, the radial transfer of sucrose from the vascular bundles to parenchymal cells of the sucrose-storing internodes follows a symplastic pathway (Rae et al., 2005). While, in sorghum, it follows an apoplastic pathway (Tarpley and Vietor, 2007). Additionally, sucrose transporters (SUTs) are involved in the long-distance transport of sucrose through the phloem, fulfilling the loading and unloading functions. Among these SUTs, SUT1 is highly specific for sucrose transport in monocotyledonous plants (Scofield et al., 2007; Reinders et al., 2010; Slewinski and Robert and Braun, 2010). The expression of SUT1 in the internodes of sweet sorghum was lower than that in grain sorghum, which may reflect the reduced retrieval of sucrose in the phloem and consequently, an enhanced efflux into storage parenchymal cells (Qazi et al., 2012).
Many enzymes, such as SPS, SS, and INV, are tightly involved in sugar metabolism (Turgeon, 1989). SS can catalyze both the synthesis and the decomposition of sucrose (Schäfer et al., 2005; Hoffmann-Thoma et al., 2010). SPS takes part in the synthesis of sucrose phosphate, which can be converted into sucrose by the catalysis of sucrose phosphate phosphatase (Lunn and Macrae, 2003). INV plays a key role in the process of sucrose degradation and mainly catalyzes the decomposition of sucrose to monosaccharides (Lunn and Macrae, 2003).
Hormones
Hormones play essential roles in the germination, growth, and development of plants (Chen et al., 2013a; Li et al., 2015). Hormones and their signaling pathways are integrated to allow the triggering of appropriate cellular and physiological responses to adapt to stress circumstances (Ryu and Cho, 2015). To cope with abiotic stresses, several plant hormones regulate defense response reactions (Kazuo et al., 2009; Ahmad and Prasad, 2012). Plant hormones play important roles in responses to salt stress (Javid et al., 2011; Zeng et al., 2018). At the physiological and biochemical levels, sweet sorghum triggers an endogenous hormone-related signaling cascade after sensing external salt stress, which provokes downstream salt-stress responses. Under salt-stress conditions, sweet sorghum can restrict the transport of Na+ to the aboveground plant parts by salt exclusion and enhance the antioxidant system simultaneously, and resist the secondary oxidative stress caused by salt stress (Sui et al., 2015; Yang et al, 2018). These protective mechanisms allow sweet sorghum to maintain high photosynthetic efficiency under salt-stress conditions. Maintenance of high photosynthetic performance, up-regulation of genes functioning in sucrose synthesis (SPS and SS encoding genes), and down-regulation of genes related to sucrose degradation (INV-encoding genes) triggered by salt stress lead to the accumulation of sucrose of salt-tolerant sweet sorghum lines.
Among the known hormones, abscisic acid (ABA), jasmonate (JA), ethylene, and cytokinin (CK) have been demonstrated to mediate plant defense responses against abiotic stresses (Nakashima and Yamaguchishinozaki, 2013). One of the fastest responses to abiotic stress is the accumulation of ABA, which is a key regulator in the activation of plant cellular adaptation to drought and salt stresses (Chen et al., 2013b). The increased levels of ABA facilitate the binding to its receptor to initiate signal transduction, leading to cellular responses to stresses (Ng et al., 2014; Sah et al., 2016). Additionally, many other salt-stress-related pathways, such as proline accumulation and calcium signaling, are also physiologically linked to the ABA signaling pathway (Yang et al., 2017). CKs play roles in the regulation of plant responses to stress (Ha et al., 2012). Unlike ABA, CKs can play positive or negative roles under stress conditions (Zwack and Rashotte, 2015; Yang et al., 2017). Colebrook et al. (2014) illustrated that the gibberellin levels affect plant growth under several stress conditions. Although hormones are critical in the salt-stress responses of sweet sorghum, the key hormones functioning in the response reactions vary among sweet sorghum genotypes and possess distinct levels of salt tolerance. Yang et al. (2017) reported that in the salt-tolerant inbred line M-81E, ABA may play a key role in the salt-stress response. However, in salt-sensitive inbred lines, JA may act as the key regulatory hormone of salt stress. It is presumably that ABA and JA regulate the growth and development of sweet sorghum in different tissues of sweet sorghum. ABA plays roles in predominantly leaves whereas JA is active mainly in roots (Yang et al., 2017). It has been described that ABA modulates the degradation of PEPC-k in sorghum leaves by increasing kinase activity in the illuminated leaf of the non-stressed plant and decreasing PEPC-k activity in the dark (Monreal et al., 2007). These results indicated that the changes of ABA content in the leaves of salt-tolerant sweet sorghum variants may be related to the maintenance of high photosynthetic efficiency of sweet sorghum under salt stress.
Maintenance of Ion Homeostasis
Under salt stress, sorghum can operate osmotic regulation by transporting ions into vacuoles or accumulating soluble substances (Lacerda et al., 2003). Sweet sorghum can scavenge ROS through enzymatic and non-enzymatic antioxidant systems, adapting to secondary oxidative stress caused by salt stress (Chai et al., 2010). In addition, sweet sorghum can also accumulate Na+ in roots and limit the transportation of Na+ up to shoots under salt stress, which is called salt exclusion (Dai et al., 2014), an important salt-tolerance-related process in monocotyledonous crops, including rice, maize, and sweet sorghum. Plants having the characteristics of salt exclusion can accumulate Na+ in the vacuoles of root parenchyma and suppress Na+ transportation into shoots. Root apoplastic barriers consist of Casparian bands and suberin lamellae. The formation of apoplastic barriers can block the apoplastic transpirational bypass flow of Na+ (Krishnamurthy P et al., 2009; Yang et al., 2018) (Figure 2). Additionally, the high-affinity potassium transporter (HKT) plays important roles in retrieving Na+ transport from roots to shoots. Salt stress induces the expression of SbHKT1;4 in salt-tolerant inbred sorghum, and it is vital for balancing the ratio of Na+/K+ to improve plant growth (Wang et al., 2014). The combining effects of apoplastic barriers in roots and Na+ transport by HKTs lead to Na+ accumulation in roots. The accumulated Na+ is finally transported into the vacuoles through NHXs or out of root cells by the Na+/H+ antiporters in the plasma membrane (Yang et al., 2018) (Figure 2A).
ROS Scavenging
In plants, dynamic balance of the ROS content is essential for plant growth. However, when the ROS accumulation exceeds the tolerance of the cell, it will cause damage to plants by attacking membrane structure or mediating apoptosis (Xue et al., 2013). Salt stress can induce the accumulation of ROS, which could greatly affect plant photosynthesis, metabolism, signal transduction, and other physiological and biochemical processes. The antioxidant system is adopted in plants to scavenge ROS and free radicals to prevent damages caused by ROS (Sui et al., 2015). Antioxidant enzymes serve as important components of the antioxidant system to remove ROS in plants and increase plant resistance, allowing the maintenance of physiological growth and development. For instance, superoxide dismutase is used to scavenge O2−. by converting it to H2O2. Catalase is used to scavenge H2O2 by catalyzing it to H2O and O2 (Mittler, 2002). The expression of ROS-associated antioxidant enzymes is commonly induced by salt stress (Evelin et al., 2009; Xue et al., 2013). Sorghum could use these antioxidant enzymes to adapt to salt and drought stresses (Jogeswar et al., 2006).
Osmotic Stress
The responses of plants to salt stress are characterized by two phases. The first phase is determined by osmotic stress and the second phase is ionic stress (Munns, 1993). Plants are capable of adjusting their water balance in response to salt stress. Plants adapt to osmotic stress mainly by reducing transpiration and accumulating osmotic adjustment substances (Chen M. et al., 2018). It has been reported that sorghum accumulated proline and soluble carbohydrates after NaCl treatment for 20 days (Heidari, 2009). The accumulation of these substances reduced the water potential of sorghum and maintained the water-absorbing capacity under salt stress. The aquaporin proteins play important roles in transportation of water across biological membranes, which is crucial for plants to combat salt stress (Xin et al., 2014). In sweet sorghum, the aquaporin-encoding genes in both salt-tolerant and salt-sensitive lines were up-regulated under salt stress, which may promote water transport efficiency (Yang et al., 2018). Additionally, the stomatal conductance in leaves of sweet sorghum was notably decreased under salt stress, which plays an important role in reducing water loss through transpiration (Sui et al., 2015).
Responses to Salt Stress At Different Developmental Stages of Sweet Sorghum
Sweet sorghum has high salt tolerance (Vasilakoglou et al., 2011; Sui et al., 2015). However, the salt effects on sweet sorghum vary among species and occur in different developmental stages. Germination is the starting point for the growth and developmental processes for all crops. Therefore, high germination ability is important for crops in saline soils. The inhibition effects of salt stress on plant germination are mainly through two ways. Primarily, salt stress can drastically decrease the osmotic potential of the soil solution to retard water absorption of seeds. Additionally, salt stress can also cause the sodium and/or chloride toxicity to the embryo and alter protein synthesis (Panuccio et al., 2014). Seed germination is a major factor limiting the establishment of plant populations under saline conditions. Salt-tolerant sweet sorghum germplasms can maintain a good germination state under salt stress (Almodares et al., 2007; Patanè et al., 2009; Patanè et al., 2013). However, under salt-stress conditions, the germination of salt-sensitive sweet sorghum germplasms is significantly inhibited. The germination rate, germination index, germination energy, and fresh weights of roots and shoots were all decreased under salt stress, which result in low densities and low yields (Ding et al., 2018).
It has been reported that the increase in the Na+ concentration and the decrease in the K+ concentration in salt-sensitive sweet sorghum germplasms during salt stress could lead to the decreased levels of several synthesis and metabolism actions, such as photosynthesis and nutrient transport (Sui et al., 2015; Yang et al., 2018), impeding the growth of both roots and shoots (Almodares et al., 2014). These effects appear to be more significant in salt-sensitive sweet sorghum inbred (Yang et al., 2018). These implied that tolerance to salt stress in sweet sorghum may be related to the decrease of Na+ and Cl+ accumulation and the maintenance of K+ level.
Many of the screening experiments on sweet sorghum have been performed under controlled environmental conditions during the early developmental stages, such as germination and seedling stages. The salt tolerance of only a few germplasms under field conditions has been reported (Vasilakoglou et al., 2011; Fan et al., 2013) as the variation in environment (such as alternations in temperature and rainfall) would easily lead to the variation in the growth of sweet sorghum. A study of four sweet sorghum species in Northern Greece has revealed that sorghum grown in intermediate-salt soil (3.2 dS m−1) produced more juice and sugar, and have greater crop and ethanol yields than that grown in high-salt soil (6.9 dS m−1) (Vasilakoglou et al., 2011). Moreover, the physiological features of sweet sorghum grown in saline soil vary greatly among germplasms. As it has been reported, some physiological features such as seed emergence rate and growth rate of sweet sorghum decreased under salt stress. However, the plant height, stem width, seed emergence rate, and the total leaf area were significantly higher in salt-tolerant sweet sorghum germplasm than that in salt-sensitive counterparts (Ding et al., 2013). In addition, photosynthetic parameters and sugar content were also less affected in salt-tolerant sweet sorghum germplasm than that in salt-sensitive lines (Fan et al., 2013).
Responses of Metabolic Processes to Salt Stress At the Molecular Level
The responses of metabolic processes to salt stress at the physiological and biochemical levels are closely related to the regulatory expression of some encoding genes. Nowadays, many genes related to salt response in sweet sorghum have been identified and characterized (Buchanan et al., 2005; Sun et al., 2016; Casto et al., 2018). ABA is a key regulator of the activation of plant cellular adaptation to drought and salt stresses. Examination of the differential gene expression in Sorghum after exposure of seedlings to high salinity, osmotic stress, and ABA revealed that 89 genes were induced with one-fold changes bigger than 2 in shoots, and 84 genes were induced with two-fold changes in roots by salt and ABA treatments (Buchanan et al., 2005). Moreover, there is a great overlap in gene expression in response to salt and ABA treatments, which indicated that ABA signaling pathway plays an important role in sorghum in response to salt stress. It has been reported that salt stress can regulate the accumulation of endogenous ABA in sorghum and SbABI5 expression in sorghum roots is a key step in the ABA signaling pathway to improve salt tolerance (Sun et al., 2016). Under salt stress, the ABA synthesis-related gene Sb04g030640 was up-regulated in the leaves of salt-tolerant sweet sorghum inbred lines and was down-regulated in salt-sensitive inbred lines. However, genes related to ABA metabolism showed the opposite trends in the two inbred lines. Additionally, genes related to the ABA signaling pathway, such as PYR/PYL and PP2C, also possess different expression patterns. After treatment with NaCl, the PYL8-encoding gene Sb09g006700 was down-regulated in the roots of salt-sensitive Roma. By contrast, the DEGs-encoding gene PP2C was up-regulated in salt-sensitive Roma but was down-regulated in salt-tolerant M-81E (Yang et al., 2017). Genes involved in the synthesis, metabolism, and signal transduction of JA, indole-3-acetic acid, and CK in sweet sorghum were also expressed with various regulatory patterns under salt-stress conditions (Yang et al., 2017).
Root apoplastic barriers, consisting of Casparian bands and suberin lamellae, play pivotal roles in blocking the apoplastic bypass flow of water and ions into the stele and Na+ transport into shoots (Pignocchi and Foyer, 2003). Salt stress induces the strengthening of the root apoplastic barriers (Krishnamurthy P et al., 2009). The expression of genes related to apoplastic barriers in sweet sorghum was up-regulated under high salinity conditions (Yang et al., 2018), suggesting that the apoplastic barriers may take part in the salt response of sweet sorghum. HKTs have been proven taking part in retrieving Na+ from the xylem vessels thus restricting the transport of Na+ from roots to leaves. They are involved in the regulation of Na+ and K+ transportation and maintenance of Na+/K+ balance (Byrt et al., 2014). It has been reported that the expression of SbHKT1;4 was strongly up-regulated in salt-tolerant sorghum, correlating with an optimal balanced Na+/K+ ratio and enhanced plant growth (Wang et al., 2014). In sweet sorghum, the expression of SbHKT1;5 was enhanced both in salt-tolerant and salt-sensitive inbred lines after NaCl treatments, and SbHKT1;5 has a greater expression in the salt-tolerant M-81E (Yang et al., 2018).
In addition, the genes related to ROS scavenging, heat shock proteins, and osmoregulation solutes also showed differential expression in sweet sorghum under NaCl treatment (Yang et al., 2018). With the expression of these genes, the contents of proline, soluble protein, catalase, and peroxidase increased in sweet sorghum under NaCl treatment (Oliveira et al., 2011; Ngara et al., 2012). These accumulations may contribute to the increase of salt-resistance capability of sweet sorghum.
Under salt stress, the genes encoding photosynthetic proteins were also shown to present different expression regulations in distinct sweet sorghum genotypes. The genes encoding Lhca1 and Lhcb1-5 were down-regulated in both salt-tolerant and salt-sensitive sweet sorghum lines under salt stress (Sui et al., 2015). The expression levels of the genes encoding Lcha2-4 and Lchb6 dropped in salt-sensitive Roma under salt stress but did not change in salt-tolerant M-81E. Moreover, the genes Sb02g002830 and Sb09g021810 that are related to the stable assembly of the oxygen-evolving complex and ATP synthase were up-regulated, affecting the stability of the photosynthetic apparatus under salt-stress conditions (Sui et al., 2015). As a C4 plant, sweet sorghum uses NADP-malic enzyme, ribulose-bisphosphate carboxylase, phosphoenolpyruvate carboxylase, and pyruvate orthophosphate dikinase as the key enzymes in the dark reaction of photosynthesis. When treated with NaCl, the genes encoding these enzymes in sweet sorghum were down-regulated, indicating that salt stress reduced CO2 assimilation. However, genes encoding other enzymes involved in carbon fixation in photosynthetic organisms, such as NADP+-malate dehydrogenase, were extremely enhanced by salt stress in only salt-tolerant sweet sorghum genotypes. This could increase the levels of CO2, pyruvate, and NADPH, which enhance CO2 assimilation (Sui et al., 2015).
So far, the molecular mechanism of comprehensive salt responses in sweet sorghum has been largely unknown. It is necessary to draw great attention on the mechanism of salt tolerance and the functions of salt-tolerant genes/proteins in sorghum.
Genome-Wide and Genetic Engineering of Sweet Sorghum to Improve Salt Tolerance
The sequencing of the whole sorghum genome was completed in 2008 (Paterson et al., 2008). To date, 820 sorghum protein-encoding sequences have been registered (http://www.gramene.org). The molecular cytogenetic maps have also been completed using multiprobe FISH cocktails technology (Islamfaridi et al., 2002). Zheng et al. (2011) carried out the genome resequencing of sweet sorghum and Chinese sorghum strains using a second-generation high-throughput sequencing platform. In total, 95% of the sequences were mapped to the sorghum genome, and a large number of single-nucleotide polymorphisms were found (Zheng et al., 2011). The completion of the genome sequencing laid the foundation for in-depth study of functional genes in sweet sorghum. Many resistance-related genes, including those encoding key enzymes of some metabolic pathways, and genes important for biological productions have been characterized. Developing an efficient genetic transformation system is pivotal for molecular breeding and understanding the genetic control of plant physiology and development. It is especially necessary for the salt-tolerance breeding of sweet sorghum. Currently, the absence of an efficient sweet sorghum transformation protocol represents a major bottleneck in salt-tolerance genetic engineering (Raghuwanshi and Birch, 2010; Liu and Godwin, 2012).
Conclusion and Perspectives
To survive under salt stress, plants have developed multiple salt tolerance mechanisms. Briefly, sweet sorghum improves the Na+ exclusion ability to ensure a relatively low Na+ concentration in shoots. Additionally, salt-tolerant sweet sorghum can maintain a high sugar content in shoots, by protecting the assembly of photosystems, enhancing the biosynthesis of sucrose and inhibiting the degradation. However, many puzzles for understanding the sweet sorghum acclimation to salt stress remain to be elucidated, for example, the regulation of salt-tolerant pathways and their interactions, how to apply the theoretical research outputs to field for crop production. To address these puzzles, more advanced and in-depth studies are required. In the future, some techniques such as gene editing systems may be used to address these questions. It will promote biotechnological applications and molecular breeding of salt-tolerant crops, which can increase the usage of saline land and crop production.
Statements
Ethics statement
Written informed consent was obtained from the individual(s) for the publication of any potentially identifiable images or data included in this article.
Author contributions
ZY and J-LL initiated preparation of the manuscript. NS, QX, and L-NL finalized the manuscript. All authors have read and approved the final manuscript.
Funding
We are grateful for financial support from the National Key R&D Program of China (2018YFD1000700, 2018YFD1000704), the National Natural Science Research Foundation of China (31871538), Shandong Provincial Natural Science Foundation (ZR2016JL028), Major Program of Shandong Provincial Natural Science Foundation (2017C03), Shandong Province Key Research and Development Program (2019GSF107079), the Development Plan for Youth Innovation Team of Shandong Provincial (2019KJE012), Shandong Provincial Natural Science Foundation (ZR2015EM007), Royal Society (UF120411 and URF\R\180030 to L-NL), and the UK Biotechnology and Biological Sciences Research Council (BB/R003890/1 to L-NL).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1
AhmadP.PrasadM. (2012). Environmental adaptations and stress tolerance of plants in the era of climate change (New York:Springer Science & Business Media). doi: 10.1007/978-1-4614-0815-4_2
2
AlmodaresA.HadiM. R. (2009). Production of bioethanol from sweet sorghum: a review. Afr. J. Agric. Res.4, 772–780. doi: 10.1021/jf9024163
3
AlmodaresA.HadiM. R.DostiB. (2007). Effects of salt stress on germination percentage and seedling growth in sweet sorghum cultivars. J. Biol. Sci.7, 1492–1495. doi: 10.3923/jbs.2007.1492.1495
4
AlmodaresA.HadiM. R.KholdebarinB.SamedaniB.KharazianZ. A. (2014). The response of sweet sorghum cultivars to salt stress and accumulation of Na+, Cl– and K+ ions in relation to salinity. J. Environ. Biol.35, 733–739. doi: 10.1246/bcsj.44.99
5
AntonopoulouG.GavalaH. N.SkiadasI. V.AngelopoulosK.LyberatosG. (2008). Biofuels generation from sweet sorghum: fermentative hydrogen production and anaerobic digestion of the remaining biomass. Bioresource Technol.99 (1), 110–119. doi: 10.1016/j.biortech.2006.11.048
6
BlumwaldE. (2000). Sodium transport and salt tolerance in plants. Curr. Opin. In Cell Biol.12, 431–434. doi: 10.1016/S0955-0674(00)00112-5
7
BuchananC. D.LimS.SalzmanR. A.KagiampakisI.MorishigeD. T.WeersB. D.et al. (2005). Sorghum bicolor’s transcriptome response to dehydration, high salinity and ABA. Plant Mol. Biol.58, 699–720. doi: 10.1007/s11103-005-7876-2
8
ByrtC. S.PlattenJ. D.SpielmeyerW.JamesR. A.LagudahE. S.DennisE. S.et al. (2007). HKT1; 5-like cation transporters linked to Na+ exclusion loci in wheat, Nax2 and Kna1. Plant Physiol.143, 1918–1928. doi: 10.1104/pp.106.093476
9
ByrtC. S.XuB.KrishnanM.LightfootD. J.AthmanA.JacobsA. K.et al. (2014). The Na+ transporter, TaHKT1;5-D, limits shoot Na+ accumulation in bread wheat. Plant J. Cell Mol. Biol.80, 516–526. doi: 10.1111/tpj.12651
10
CaoS.DuX. H.LiL. H.LiuY. D.ZhangL.PanX.et al. (2017). Overexpression of Populus tomentosa cytosolic ascorbate peroxidase enhances abiotic stress tolerance in tobacco plants. Russian J. Plant Physiol.64 (2), 224–234. doi: 10.1134/s1021443717020029
11
CarterP. J.NimmoH. G.FewsonC. A.WilkinsM. B. (1990). Bryophyllum fedtschenkoi protein phosphatase 2A can dephosphorylate phosphoenolpyruvate carboxylase. FEBS Letters263, 233–236. doi: 10.1016/0014-5793(90)81381-W
12
CastoA. L.McKinleyB. A.YuK. M. J.RooneyW. L.MulletJ. E. (2018). Sorghum stem aerenchyma formation is regulated by SbNAC_D during internode development. Plant Direct2 (11), e00085. doi: 10.1002/pld3.85
13
ChaiY. Y.JiangC. D.ShiL.ShiT. S.GuW. B. (2010). Effects of exogenous spermine on sweet sorghum during germination under salinity. Biol. Plantarum54, 145–148. doi: 10.1007/s10535-010-0023-1
14
ChenM.ZhangW. H.LvZ. W.ZhangS. L.HidemaJ.ShiF. M.et al. (2013a). Abscisic acid is involved in the response of Arabidopsis mutant sad2-1 to ultraviolet-B radiation by enhancing antioxidant enzymes. South Afr. J. Botany85, 79–86. doi: 10.1016/j.sajb.2012.11.006
15
ChenX.ZhuW.AzamS.LiH.ZhuF.LiH.et al. (2013b). Deep sequencing analysis of the transcriptomes of peanut aerial and subterranean young pods identifies candidate genes related to early embryo abortion. Plant Biotechnol. J.11, 115–127. doi: 10.1111/pbi.12018
16
ChenL.YuanX.LiJ.DongZ.ShaoT. (2018). Effects of applying oil-extracted microalgae on the fermentation quality, feed-nutritive value and aerobic stability of ensiled sweet sorghum. J. Sci. Food Agric.98, 4462–4470. doi: 10.1002/jsfa.8970
17
ChenM.YangZ.LiuJ.ZhuT.WangB. (2018). Adaptation mechanism of salt excluders under saline conditions and its applications. Int. J. Mol. Sci.19, 3668. doi: 10.3390/ijms19113668
18
ColebrookE. H.ThomasS. G.PhillipsA. L.HeddenP. (2014). The role of gibberellin signalling in plant responses to abiotic stress. J. Exp. Biol.217, 67–75. doi: 10.1242/jeb.089938
19
CuiF.SuiN.DuanG.LiuY.HanY.LiuS.et al. (2018). Identification of metabolites and transcripts involved in salt stress and recovery in peanut. Front. In Plant Sci.9, 217. doi: 10.3389/fpls.2018.00217
20
DaiL. Y.ZhangL. J.JiangS. J.YinK. D. (2014). Saline and alkaline stress genotypic tolerance in sweet sorghum is linked to sodium distribution. Acta Agric Scandinavica.64, 471–481. doi: 10.1080/09064710.2014.925574
21
DengY.FengZ.YuanF.GuoJ.SuoS.WangB. (2015). Identification and functional analysis of the autofluorescent substance in Limonium bicolor salt glands. Plant Physiol. Biochem.97, 20–27. doi: 10.1016/j.plaphy.2015.09.007
22
DingT. L.SongJ.GuoJ. R.SuiN.FanH.ChenM.et al. (2013). The cultivation technique for increasing the stalk sugar content of energy plant sweet sorghum in Yellow River delta. Adv Mat Res.724, 437–442. doi: 10.4028/www.scientific.net/AMR.724-725.437
23
DingT.YangZ.WeiX.YuanF.YinS.WangB. (2018). Evaluation of salt-tolerant germplasm and screening of the salt-tolerance traits of sweet sorghum in the germination stage. Funct. Plant Biol.45, 1073–1081. doi: 10.1071/FP18009
24
Dionisio-SeseM. L.TobitaS. (2000). Effects of salinity on sodium content and photosynthetic responses of rice seedlings differing in salt tolerance. J. Plant Physiol.157, 54–58. doi: 10.1016/S0176-1617(00)80135-2
25
EchevariaE.Garcia-MaurinoS. R.SolerA.VidalJ. (2001). Salt stress increases the Ca2+-independent phosphoenolpyruvate carboxylase kinase activity in sorghum leaves. Planta214 (2), 283–287. doi: 10.1007/s004250100616
26
EvelinH.KapoorR.GiriB. (2009). Arbuscular mycorrhizal fungi in alleviation of salt stress: a review. Ann. Botany104, 1263–1280. doi: 10.1093/aob/mcp251
27
FanH.ChengR. R.WuH. D.ChengS.YanJ.DingT. L.et al. (2013). Planting sweet sorghum in Yellow River Delta: agronomy characters of different varieties and the effects of sowing time on the yield and other biological traits. Adv Mat Res.726-731, 3–8. doi: 10.4028/www.scientific.net/AMR.726-731.3
28
FengZ. T.DengY. Q.FanH.SunQ. J.SuiN.WangB. S. (2014). Effects of NaCl stress on the growth and photosynthetic characteristics of Ulmus pumila L. seedlings in sand culture. Photosynthetica52, 313–320. doi: 10.1007/s11099-014-0032-y
29
FerrarisR.Charles-EdwardsD. A. (1986). A comparative analysis of the growth of sweet and forage sorghum crops. II. Accumulation of soluble carbohydrates and nitrogen. Aust. J. Agric. Res.37, 513. doi: 10.1071/ar9860513
30
FlowersT. J.ColmerT. D. (2010). Salinity tolerance in halophytes. New Phytol.179, 945–963. doi: 10.1111/j.1469-8137.2008.02531.x
31
GalkaP.SantabarbaraS.KhuongT. T. H.DegandH.MorsommeP.JenningsR. C.et al. (2012). Functional analyses of the plant photosystem I-light-harvesting complex II supercomplex reveal that light-harvesting complex II loosely bound to photosystem II is a very efficient antenna for photosystem I in state II. Plant Cell.24, 2963–2978. doi: 10.1105/tpc.112.100339
32
García-MauriñoS.MonrealJ.AlvarezR.VidalJ.EchevarríaC. (2003). Characterization of salt stress-enhanced phosphoenolpyruvate carboxylase kinase activity in leaves of Sorghum vulgare: Independence from osmotic stress, involvement of ion toxicity and significance of dark phosphorylation. Planta216 (4), 648–655. doi: 10.2307/23387685
33
GetachewG.PutnamD. H.BenC. M. D.PetersE. J. D. (2016). Potential of sorghum as an alternative to corn forage. Am. J. Plant Sci.7 (7), 1106–1121. doi: 10.4236/ajps.2016.77106
34
GnansounouE.DauriatA.WymanC. E. (2005). Refining sweet sorghum to ethanol and sugar: economic trade-offs in the context of North China. Bioresource Technol.96, 985–1002. doi: 10.1016/j.biortech.2004.09.015
35
GuimaraesC. T.SillsG. R.SobralB. W. S. (1997). Comparative mapping of Andropogoneae: Saccharum L. (sugarcane) and its relation to sorghum and maize. Proc. Natl. Acad. Sci.94, 14261–14266. doi: 10.1073/pnas.94.26.14261
36
GuoY. H.WangD.JiaW. J.SongJ.YangJ. C.WangB. S. (2012). Effects of seed vernalisation and photoperiod on flowering induction in the halophyte Thellungiella halophila. Aust. J. Botany60 (8), 743–748. doi: 10.1071/bt12180
37
GuoY.SongY.ZhengH.ZhangY.SuiN. (2018a). NADP-malate dehydrogenase of sweet sorghum improves salt tolerance of Arabidopsis thaliana. J. Agric. Food Chem.66, 5992–6002. doi: 10.1021/acs.jafc.8b02159
38
GuoY. Y.TianS. S.LiuS. S.WangW. Q.SuiN. (2018b). Energy dissipation and antioxidant enzyme system protect photosystem II of sweet sorghum under drought stress. Photosynthetica56, 861–872. doi: 10.1007/s11099-017-0741-0
39
HaS.VankovaR.Yamaguchi-ShinozakiK.ShinozakiK.TranL. S. (2012). Cytokinins: metabolism and function in plant adaptation to environmental stresses. Trends In Plant Sci.17, 172–179. doi: 10.1186/s12864-015-1760-5
40
HanG.WangM.YuanF.SuiN.SongJ.WangB. (2014). The CCCH zinc finger protein gene AtZFP1 improves salt resistance in Arabidopsis thaliana. Plant Mol. Biol.86, 237–253. doi: 10.1007/s11103-014-0226-5
41
HatchM. D. (1987). C4 photosynthesis: a unique elend of modified biochemistry, anatomy and ultrastructure. Bba Reviews Bioenergetics. 895, 81–106. doi: 10.1016/S0304-4173(87)80009-5
42
HeidariM. (2009). Antioxidant activity and osmolyte concentration of sorghum (Sorghum bicolor) and wheat (Triticum aestivum) genotypes under salinity stress. Asian J. Plant Sci.8 (3), 240–244. doi: 10.1118/1.598892
43
Hoffmann-ThomaG.HinkelK.NicolayP.WillenbrinkJ. (2010). Sucrose accumulation in sweet sorghum stem internodes in relation to growth. Physiol Plantarum97, 277–284. doi: 10.1034/j.1399-3054.1996.970210.x
44
IEA. (2019). Data from: Key World Energy Statistics 2018. https://webstore.iea.org/key-world-energy-statistics-2018
45
IslamfaridiM. N.ChildsK. L.KleinP. E.HodnettG.MenzM. A.KleinR. R.et al. (2002). A molecular cytogenetic map of sorghum chromosome 1. Fluorescence in situ hybridization analysis with mapped bacterial artificial chromosomes. Genetics161, 345. doi: 10.1023/A:1016080316076
46
JavidM. G.SorooshzadehA.MoradiF.SanavyS. A. M. M.AllahdadiI. (2011). The role of phytohormones in alleviating salt stress in crop plants. Aust. J. Crop Sci.32, 726–734. doi: 10.1163/156853811X610320
47
JiaoL.WangL.ZhouQ.HuangX. (2017). Stomatal and non-stomatal factors regulated the photosynthesis of soybean seedlings in the present of exogenous bisphenol A. Ecotoxicol Environ. Saf.145, 150–160. doi: 10.1016/j.ecoenv.2017.07.028
48
JogeswarG.PallelaR.JakkaN. M.ReddyP. S.RaoJ. V.SreenivasuluN. (2006). Antioxidative response in different sorghum species under short-term salinity stress. Acta Physiol Plantarum28, 465–475. doi: 10.1007/bf02706630
49
KühnC.GrofC. P. (2010). Sucrose transporters of higher plants. Curr. Opin. In Plant Biol.13, 287–297. doi: 10.1016/j.pbi.2010.02.001
50
KazuoN.YasunariF.NorihitoK.TakeshiK.TaishiU.SatoshiK.et al. (2009). Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/SnRK2.3, involved in ABA signaling are essential for the control of seed development and dormancy. Plant Cell Physiol.50, 1345–1363. doi: 10.1093/pcp/pcp083
51
KongX. Q.GaoX. H.SunW.AnJ.ZhaoY. X.ZhangH. (2011). Cloning and functional characterization of a cation-chloride cotransporter gene OsCCC1. Plant Mol. Biol.75 (6), 567–578. doi: 10.1007/s11103-011-9744-6
52
KongX. Q.WangT.LiW. J.TangW.ZhangD. M.DongH. Z. (2016). Exogenous nitric oxide delays salt-induced leaf senescence in cotton (Gossypium hirsutum L.). Acta Physiol Plantarum38 (3), 61. doi: 10.1007/s11738-016-2079-9
53
KrishnamurthyP.RanathungeK.FrankeR.PrakashH. S.SchreiberL.MathewM. K. (2009). The role of root apoplastic transport barriers in salt tolerance of rice (Oryza sativa L.). Planta230, 119–134. doi: 10.2307/23390598
54
KrishnamurthyP.RanathungeK.NayakS.SchreiberL.MathewM. K. (2011). Root apoplastic barriers block Na+ transport to shoots in rice (Oryza sativa L.). J. Exp. Botany62, 4215–4228. doi: 10.1093/jxb/err135
55
LacerdaC. F. DCambraiaJ.OliviaM. A.RuizH. A.PriscoJ. T. (2003). Solute accumulation and distribution during shoot and leaf development in two sorghum genotypes under salt stress. Environ. Exp. Botany. 49 (2), 107–120. doi: 10.1016/S0098-8472(02)00064-3
56
LandiS.HausmanJ. F.GuerrieroG.EspositoS. (2017). Poaceae vs. abiotic stress: focus on drought and salt stress, recent insights and perspectives. Front. In Plant Sci.8, 1214. doi: 10.3389/fpls.2017.01214
57
LengB. Y.YuanF.DongX. X.WangJ.WangB. S. (2018). Distribution pattern and salt excretion rate of salt glands in two recretohalophyte species of Limonium (Plumbaginaceae). South Afr. J. Botany115, 74–80. doi: 10.1016/j.sajb.2018.01.002
58
LiW.ShinjiroY.AjmalK. M.PingA.LiuX.TranL. S. P. (2015). Roles of gibberellins and abscisic acid in regulating germination of Suaeda salsa dimorphic seeds under salt stress. Front. In Plant Sci.6, 1235. doi: 10.3389/fpls.2015.01235
59
LiuG.GodwinI. D. (2012). Highly efficient sorghum transformation. Plant Cell Rep.31, 999–1007. doi: 10.1007/s00299-011-1218-4
60
LiuS.ChengY.ZhangX.GuanQ.NishiuchiS.HaseK.et al. (2007). Expression of an NADP-malic enzyme gene in rice (Oryza sativa. L) is induced by environmental stresses; over-expression of the gene in Arabidopsis confers salt and osmotic stress tolerance. Plant Mol. Biol.64 (1-2), 49–58. doi: 10.1007/s11103-007-9133-3
61
LiuL. N. (2016). Distribution and dynamics of electron transport complexes in cyanobacterial thylakoid membranes. Biochim. Biophys. Acta (BBA) - Bioenergetics1857, 256–265. doi: 10.101 6/j.bbabio.2015.11.010
62
LunnJ. E.MacraeE. (2003). New complexities in the synthesis of sucrose. Curr. Opin. In Plant Biol.6, 208–214. doi: 10.1016/S1369-5266(03)00033-5
63
MøllerI. M. (2001). Plant mitochondria and oxidative stress: electron transport, NADPH turnover, and metabolism of reactive oxygen species. Annu. Rev. Plant Biol.52 (1), 561–591. doi: 10.1146/annurev.arplant.52.1.561
64
MaQ.YueL. J.ZhangJ. L.WuG. Q.BaoA. K.WangS. M. (2012). Sodium chloride improves photosynthesis and water status in the succulent xerophyte Zygophyllum xanthoxylum. Tree Physiol.32, 4–13. doi: 10.1093/treephys/tpr098
65
MahajanS.TutejaN. (2005). Cold, salinity and drought stresses: an overview. Arch. Biochem. Biophysics444, 139–158. doi: 10.1016/j.abb.2005.10.018
66
MccormickA. J.CramerM. D.WattD. A. (2010). Sink strength regulates photosynthesis in sugarcane. New Phytol.171 (4), 759–770. doi: 10.1111/j.1469-8137.2006.01785.x
67
MittlerR. (2002). Oxidative stress, antioxidants and stress tolerance. Trends In Plant Sci.7, 405–410. doi: 10.1016/S1360-1385(02)02312-9
68
MonrealJ. A.FeriaA. B.VinardellJ. M.VidalJ.EchevarríaC.García-MauriñoS. (2007). ABA modulates the degradation of phospho enol pyruvate carboxylase kinase in sorghum leaves. FEBS Letters581 (18), 3468–3472. doi: 10.1016/j.febslet.2007.06.055
69
MunnsR. (1993). Physiological processes limiting plant growth in saline soils: some dogmas and hypotheses. Plant Cell Environ.16 (1), 15–24. doi: 10.1111/j.1365-3040.1993.tb00840.x
70
NakashimaK.YamaguchishinozakiK. (2013). ABA signaling in stress-response and seed development. Plant Cell Rep.32, 959–970. doi: 10.1007/s00299-013-1418-1
71
NetondoG. W.OnyangoJ. C.BeckE. (2004). Sorghum and salinity: II. Gas exchange and chlorophyll fluorescence of sorghum under salt stress. Crop Sci.44, 806–811. doi: 10.2135/cropsci2004.8060
72
NgL. M.MelcherK.TehB. T.XuH. E. (2014.). Abscisic acid perception and signaling: structural mechanisms and applications. Acta Pharmacol. Sinica.35, 567–584. doi: 10.1038/aps.2014.5
73
NgaraR.NdimbaR.Borch-JensenJ.JensenO. N.NdimbaB. (2012). Identification and profiling of salinity stress-responsive proteins in Sorghum bicolor seedlings. J. Proteomics.75, 4139–4150. doi: 10.1016/j.jprot.2012.05.038
74
OliveiraA. B. D.AlencarN. L. M.PriscoJ. T.Gomes-FilhoE. (2011). Accumulation of organic and inorganic solutes in NaCl-stressed sorghum seedlings from aged and primed seeds. Scientia Agricola.68, 632–637. doi: 10.1590/S0103-90162011000600004
75
PangC. H.LiK.WangB. (2011). Overexpression of SsCHLAPXs confers protection against oxidative stress induced by high light in transgenic Arabidopsis thaliana. Physiol plantarum.143 (4), 355–366. doi: 10.1111/j.1399-3054.2011.01515.x
76
PanuccioM. R.JacobsenS. E.AkhtarS. S.MuscoloA. (2014). Effect of saline water on seed germination and early seedling growth of the halophyte quinoa. AoB Plants6, plu047. doi: 10.1093/aobpla/plu047
77
PatanèC.CavallaroV.CosentinoS. L. (2009). Germination and radicle growth in unprimed and primed seeds of sweet sorghum as affected by reduced water potential in NaCl at different temperatures. Industrial Crops Product.30, 1–8. doi: 10.1016/j.indcrop.2008.12.005
78
PatanèC.SaitaA.SortinoO. (2013). Comparative effects of salt and water stress on seed germination and early embryo growth in two cultivars of sweet sorghum. J. Agron. Crop Sci.199, 30–37. doi: 10.1111/j.1439-037X.2012.00531.x
79
PatersonA. H.BowersJ. E.FeltusF. A. (2008). “Genomics of sorghum, a semi-arid cereal and emerging model for tropical grass genomics,” in Genomics of Tropical Crop Plants. Eds. MooreP. H.MingR. (New York: Springer), 469–482. doi: 10.1007/978-0-387-71219-2_20
80
PatraA. K.SaxenaJ. (2011). Exploitation of dietary tannins to improve rumen metabolism and ruminant nutrition. J. Sci. Food Agric.91 (1), 24. doi: 10.1002/jsfa.4152
81
PignocchiC.FoyerC. H. (2003). Apoplastic ascorbate metabolism and its role in the regulation of cell signalling. Curr. Opin. In Plant Biol.6, 379–389. doi: 10.1016/S1369-5266(03)00069-4
82
QaziH. A.ParanjpeS.BhargavaS. (2012). Stem sugar accumulation in sweet sorghum-activity and expression of sucrose metabolizing enzymes and sucrose transporters. J. Plant Physiol.169, 605–613. doi: 10.1016/j.jplph.2012.01.005
83
RaeA. L.PerrouxJ. M.GrofC. P. L. (2005). Sucrose partitioning between vascular bundles and storage parenchyma in the sugarcane stem: a potential role for the ShSUT1 sucrose transporter. Planta220, 817–825. doi: 10.2307/23388789
84
RaghuwanshiA.BirchR. G. (2010). Genetic transformation of sweet sorghum. Plant Cell Rep.29, 997–1005. doi: 10.1007/s00299-010-0885-x
85
RaoP. S.KumarC. G.ReddyB. V. S. (2013). “Sweet sorghum: from theory to practice,” in Characterization of improved sweet sorghum cultivars. Springer Briefs in Agriculture. Eds. RaoP.KumarC. (India: Springer). doi: 10.1007/978-81-322-0783-2_1
86
ReindersA.SivitzA. B.HsiA.GrofC. P.PerrouxJ. M.WardJ. M. (2010). Sugarcane ShSUT1: analysis of sucrose transport activity and inhibition by sucralose. Plant Cell Environ.29, 1871–1880. doi: 10.1111/j.1365-3040.2006.01563.x
87
RyuH.ChoY.-G. (2015). Plant hormones in salt stress tolerance. J. Plant Biol. 58, 147–155. doi: 10.1007/s12374-015-0103-z
88
Sánchez-DuarteJ. I.KalscheurK. F.GarcíaA. D.Contreras-GoveaF. E. (2018). Short communication: Meta-analysis of dairy cows fed conventional sorghum or corn silages compared with brown midrib sorghum silage. J. Dairy Sci.102, 1–7. doi: 10.3168/jds.2018-14552
89
SahS. K.ReddyK. R.LiJ. (2016). Abscisic acid and abiotic stress tolerance in crop plants. Front. In Plant Sci.7, 571. doi: 10.3389/fpls.2016.00571
90
SchäferW. E.RohwerJ. M.BothaF. C. (2005). Partial purification and characterisation of sucrose synthase in sugarcane. J. Plant Physiol.162, 11–20. doi: 10.1016/j.jplph.2004.04.010
91
ScofieldG. N.HiroseT.AokiN.FurbankR. T. (2007). Involvement of the sucrose transporter, OsSUT1, in the long-distance pathway for assimilate transport in rice. J. Exp. Botany58, 3155–3169. doi: 10.1093/jxb/erm153
92
ShaoQ.HanN.DingT.ZhouF.WangB. (2014). SsHKT1; 1 is a potassium transporter of the C3 halophyte Suaeda salsa that is involved in salt tolerance. Funct. Plant Biol.41 (8), 790–802. doi: 10.1071/FP13265
93
SlewinskiT. L. M.Robert and BraunD. M. (2010). Sucrose transporter1 functions in phloem loading in maize leaves. J. Exp. Botany60, 881–892. doi: 10.1093/jxb/ern335
94
SongJ.WangB. (2015). Using euhalophytes to understand salt tolerance and to develop saline agriculture: Suaeda salsa as a promising model. Ann. Bot.115, 541. doi: 10.1093/aob/mcu194
95
SongJ.ZhouJ.ZhaoW.XuH.WangF.XuY.et al. (2016). Effects of salinity and nitrate on production and germination of dimorphic seeds applied both through the mother plant and exogenously during germination in Suaeda salsa. Plant Species Biol.31, 19–28. doi: 10.1111/1442-1984.12071
96
SudhirP.MurthyS. D. S. (2004). Effects of salt stress on basic processes of photosynthesis. Photosynthetica42, 481–486. doi: 10.1007/s11099-005-0001-6
97
SuiN.HanG. (2014). Salt-induced photoinhibition of PSII is alleviated in halophyte Thellungiella halophila by increases of unsaturated fatty acids in membrane lipids. Acta Physiol Plantarum36, 983–992. doi: 10.1007/s11738-013-1477-5
98
SuiN.LiM.LiK.SongJ.WangB. S. (2010). Increase in unsaturated fatty acids in membrane lipids of Suaeda salsa L. enhances protection of photosystem II under high salinity. Photosynthetica48, 623–629. doi: 10.1007/s11099-010-0080-x
99
SuiN.YangZ.LiuM.WangB. (2015). Identification and transcriptomic profiling of genes involved in increasing sugar content during salt stress in sweet sorghum leaves. BMC Genomics16, 534. doi: 10.3389/fpls.2018.00007
100
SuiN.WangY.LiuS.YangZ.WangF.WanS. (2018). Transcriptomic and physiological evidence for the relationship between unsaturated fatty acid and salt stress in peanut. Front. In Plant Sci.9, 7.
101
SunS.-B.ShenQ.-R.WanJ.-M.LiuZ.-P. (2003). Induced expression of the gene for NADP-malic enzyme in leaves of Aloe vera L. under salt stress. Acta Biochim. Biophys. Sinica.35 (5), 423–429. doi: 10.1016/S0022-2836(03)00269-9
102
SunL.WangC.ZhouY. F.RuanY. Y.GongX.ZhangJ. (2016). Inhibition of sbabi5 expression in roots by ultra-high endogenous aba accumulation results in sorghum sensitivity to salt stress. Int. J. Agric. Biol.18, 146–154. doi: 10.17957/IJAB/15.0079
103
TanH. Y.SieoC. C.AbdullahN.LiangJ. B.HuangX. D.HoY. W. (2011). Effects of condensed tannins from Leucaena on methane production, rumen fermentation and populations of methanogens and protozoa in vitro. Anim. Feed Sci. Technol.169 (3-4), 185–193. doi: 10.1016/j.anifeedsci.2011.07.004
104
TarpleyL.VietorD. M. (2007). Compartmentation of sucrose during radial transfer in mature sorghum culm. BMC Plant Biol.7, 33. doi: 10.1186/1471-2229-7-33
105
TurgeonR. (1989). The Sink-Source Transition in Leaves. Annu. Rev. Plant Physiol. Plant Mol. Biol.40, 119–138. doi: 10.1146/annurev.pp.40.060189.001003
106
VasilakoglouI.DhimaK.KaragiannidisN.GatsisT. (2011). Sweet sorghum productivity for biofuels under increased soil salinity and reduced irrigation. Field Crops Res.120, 38–46. doi: 10.1016/j.fcr.2010.08.011
107
VidalJ.CholletR. (1997). Regulatory phosphorylation of C4 PEP carboxylase. Plant Physiol.2 (6), 230–237. doi: 10.1016/S1360-1385(97)89548-9
108
WangR.ChenS.DengL.FritzE.HüttermannA.PolleA. (2007). Leaf photosynthesis, fluorescence response to salinity and the relevance to chloroplast salt compartmentation and anti-oxidative stress in two poplars. Trees Struct. Funct.21 (5), 581–591. doi: 10.1007/s00468-007-0154-y
109
WangT. T.RenZ. J.LiuZ. Q.FengX.GuoR. Q.LiB. G.et al. (2014). SbHKT1;4, a member of the high-affinity potassium transporter gene family from Sorghum bicolor, functions to maintain optimal Na+/K+ balance under Na+ stress. J. Integr. Plant Biol.56, 315–332. doi: 10.1111/jipb.12144
110
WhitneyS. M.HoutzR. L.AlonsoH. (2011). Advancing our understanding and capacity to engineer nature’s CO2-sequestering enzyme, Rubisco. Plant Physiol.155 (1), 27–35. doi: 10.1104/pp.110.164814
111
WydrzynskiT. J. (2008). Water splitting by photosystem II—where do we go from here? Photosynthesis Res.98, 43–51. doi: 10.1007/s11120-008-9391-1
112
XieQ.XuZ. H. (2019). Sustainable sgriculture: from sweet sorghum planting and ensiling to ruminant feeding. Mol. Plant12, 603–606. doi: 10.1016/j.molp.2019.04.001
113
XinS.YuG.SunL.QiangX.XuN.ChengX. (2014). Expression of tomato SlTIP2;2 enhances the tolerance to salt stress in the transgenic Arabidopsis and interacts with target proteins. J. Plant Res.127 (6), 695–708. doi: 10.1007/s10265-014-0658-7
114
XueX.ZhangQ.WuJ. (2013). Research of reactive oxygen species in plants and its application on stress tolerance. Biotechnol. Bulletin.36, 6–11. doi: 10.13560/j.cnki.biotech.bull.1985.2013.10.033
115
YangZ.WangY.WeiX.ZhaoX.WangB.SuiN. (2017). Transcription profiles of genes related to hormonal regulations under salt stress in sweet sorghum. Plant Mol. Biol. Rep.35, 1–14. doi: 10.1007/s11105-017-1047-x
116
YangZ.ZhengH.WeiX.SongJ.WangB.SuiN. (2018). Transcriptome analysis of sweet Sorghum inbred lines differing in salt tolerance provides novel insights into salt exclusion by roots. Plant Soil.430, 423–439. doi: 10.1007/s11104-018-3736-0
117
YuanF.LengB.WangB. (2016a). Progress in studying salt secretion from the salt glands in recretohalophytes: how do plants secrete Salt? Front. In Plant Sci.7, 997. doi: 10.3389/fpls.2016.00977
118
YuanF.LyuM. J. A.LengB. Y.ZhuX. G.WangB. S. (2016b). The transcriptome of NaCl-treated Limonium bicolor leaves reveals the genes controlling salt secretion of salt gland. Plant Mol. Biol.91, 241–256. doi: 10.1007/s11103-016-0460-0
119
ZengY.ShenJ.LiB.JiangL. (2018). Hormone modulates protein dynamics to regulate plant growth. Proc. Natl. Acad. Sci.115, 3521–3523. doi: 10.1073/pnas.1802175115
120
ZhengL. Y.GuoX. S.HeB.SunL. J.PengY.DongS. S.et al (2011). Genome-wide patterns of genetic variation in sweet and grain sorghum (Sorghum bicolor). Genome Biol.12, R114. doi: 10.1186/gb-2011-12-11-r114
121
ZhuJ. K. (2001). Plant salt tolerance. Trends In Plant Sci.6 (2), 66–71. doi: 10.1016/S1360-1385(00)01838-0
122
ZwackP. J.RashotteA. M. (2015). Interactions between cytokinin signalling and abiotic stress responses. J. Exp. Botany66, 4863. doi: 10.1093/jxb/erv172
Summary
Keywords
sweet sorghum, salt-tolerance mechanism, Na+ exclusion, photosynthesis, sugar content
Citation
Yang Z, Li J-L, Liu L-N, Xie Q and Sui N (2020) Photosynthetic Regulation Under Salt Stress and Salt-Tolerance Mechanism of Sweet Sorghum. Front. Plant Sci. 10:1722. doi: 10.3389/fpls.2019.01722
Received
07 August 2019
Accepted
09 December 2019
Published
15 January 2020
Volume
10 - 2019
Edited by
Giovanni Stefano, Michigan State University, United States
Reviewed by
Jose Antonio Monreal, University of Seville, Spain; Suriyan Cha-um, National Science and Technology Development Agency, Thailand
Updates
Copyright
© 2020 Yang, Li, Liu, Xie and Sui.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Na Sui, suina800101@163.com
†These authors have contributed equally to this work.
This article was submitted to Plant Abiotic Stress, a section of the journal Frontiers in Plant Science
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.