- 1Center for Desert Agriculture, Biological and Environmental Sciences Division, King Abdullah University of Science and Technology, Thuwal, Saudi Arabia
- 2Institute of Life Sciences, Scuola Superiore Sant’Anna, Pisa, Italy
- 3School of Plant Sciences, Arizona Genomics Institute, University of Arizona, Tucson, AZ, United States
As the world’s population expands from 7.6 billion to 10 billion over the next 30 years, scientists and farmers across the globe must explore every angle necessary to provide a safe, stable and sustainable food supply for generations to come. Rice, and its wild relatives in the genus Oryza, will play a significant role in helping to solve this 10 billion people question due to its place as a staple food for billions. The genus Oryza is composed of 27 species that span 15 million years of evolutionary diversification and have been shown to contain a plethora of untapped adaptive traits, e.g., biotic and abiotic resistances, which can be used to improve cultivated rice. Such traits can be introduced into cultivated rice, in some cases by conventional crossing, and others via genetic transformation and gene editing methods. In cases where traits are too complex to easily transfer to cultivated rice [e.g., quantitative trait loci (QTL)], an alternative strategy is to domesticate the wild relative that already contains the desired adaptive traits – i.e., “neodomestication”. To utilize the Oryza genus for crop improvement and neodomestication, we first need a set of genomic resources that can be used to efficiently identify, capture, and guide molecular crop improvement. Here, we introduce the concept of platinum standard reference genome sequences (PSRefSeq) – a new standard by which contiguous near-gap free reference genomes can now be produced. By having a set of PSRefSeqs for every Oryza species we set a new bar for how crop wild relatives can be integrated into crop improvement programs.
Introduction
As the world population grows and climate change dictates new conditions for agriculture worldwide, finding new solutions to ensure a sustainable food supply is crucial. Rice is the staple food in the regions of the world with the largest projected population growth. Moreover, these regions and their available arable land are expected to be most negatively affected by climate change in the future (Milovanovic and Smutka, 2017). Cultivated rice, as for most modern crops, has undergone domestication and genetic bottlenecks over millennia, giving this crop limited genetic potential to adapt to rapid environmental changes. In contrast, the wild relatives (WRs) of rice have been shown to possess the genetic potential to enhance yield, nutrient content, and resistance to several environmental stressors (Brar and Khush, 2018).
For decades, rice breeders have utilized genetic traits from the WRs of the genus Oryza, first via conventional crossing and now more recently using plant transformation and gene editing tools, to produce new, more resilient varieties of cultivated rice (Brar and Khush, 2018). Although these WRs contain novel alleles and variations with the potential to improve cultivated rice, issues such as hybrid-viability, -sterility, and -weakness hinder their introgression into cultivated rice, especially from the more distant relatives (Nadir et al., 2018). However, a toolbox of high-quality WR genome assemblies and gene-editing tools could help address these issues by providing technology for the precise transfer of newly discovered and advantageous wild alleles and genes into cultivated backgrounds.
Gene editing would also enable researchers to leverage these WRs with adaptive traits through neodomestication, i.e., the domestication of plants that have not previously been used for agriculture (Zsögön et al., 2018). As most traits in domesticated ideotypes arose from loss-of-function mutations, inducing such mutations in the orthologs of domestication-related genes in the WRs could result in a neodomesticated plant that maintains its desired adaptive trait(s) (e.g., abiotic and biotic resistance) (Doebley et al., 2006; Fernie and Yan, 2019). Though very far from a trivial task, neodomestication of the WRs of rice could potentially solve the challenge of feeding the planet’s rapidly growing population in the face of climate change. Neodomestication untangles the relationships between plant architecture, physiological mechanisms, and cellular processes that produce the complex and polygenic traits necessary to provide resistance or tolerance to environmental stressors.
Thus, in this review, we will discuss the following topics:
1. The untapped pool of genetic diversity buried within the genomes of the wild relatives of rice
2. The use of high-quality near gap-free reference genome sequences to aid in crop improvement
3. Barriers to genetic introgression and alternative strategies to introgression via neodomestication
Genetic Diversity of the Wild Relatives of Rice
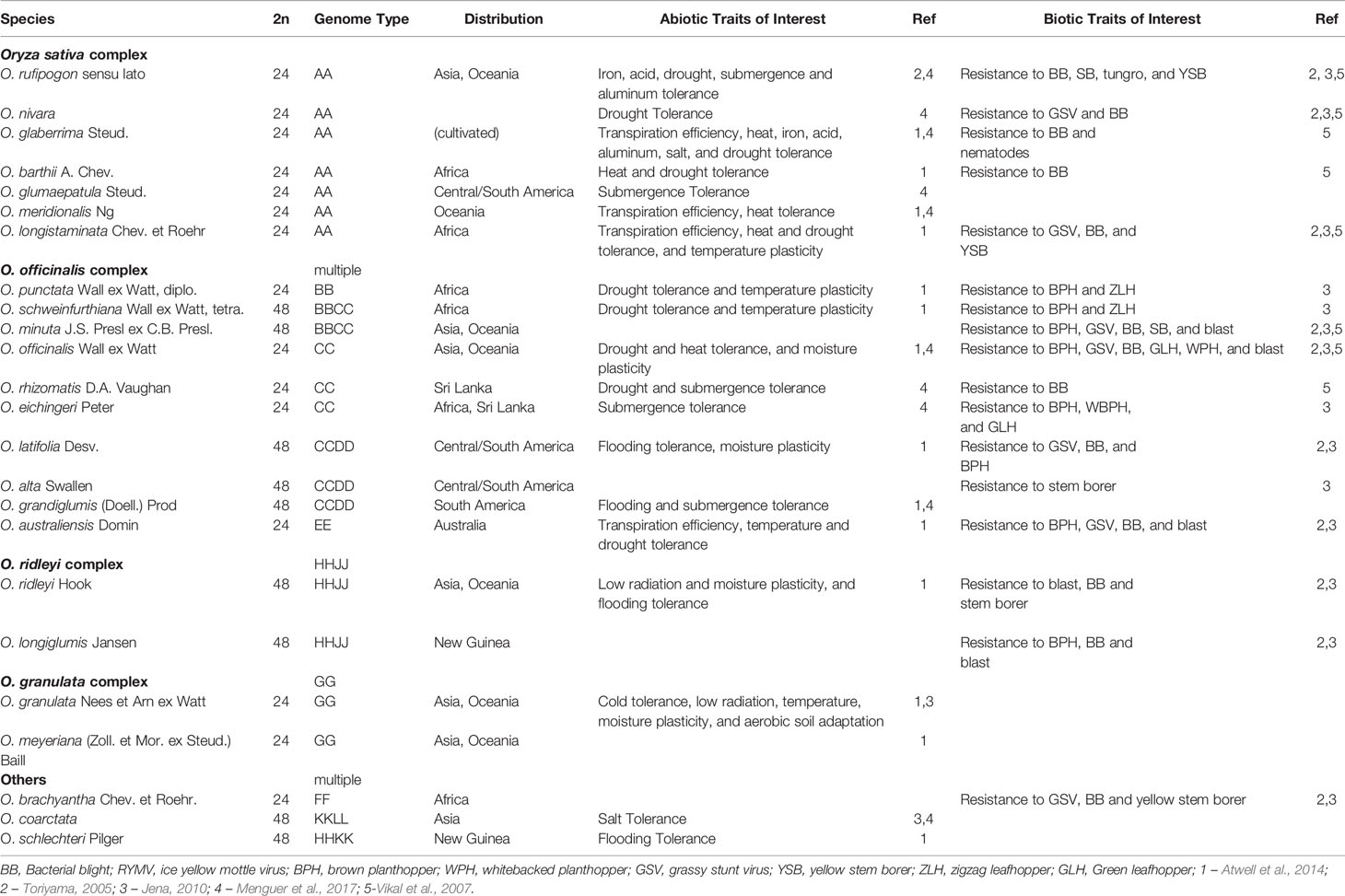
The Oryza genus contains 11 different genome types across 27 species, both diploid and polyploid, with a 3.6-fold difference in genome sizes (from around 300 Mb in the diploid FF genome of O. brachyantha to 1,283 Mb in the HHJJ polyploid genome of O. ridleyi, with the AA genome of cultivated rice falling in the 400Mb range). There are four species complexes within the Oryza genus: (1) O. sativa, (2) O. officinalis, (3) O. ridleyi, and (4) O. meyeriana (Vaughan, 1989). The WRs O. rufipogon and O. barthii have been identified as the progenitors of the domesticated rice species, O. sativa and O. glaberrima, respectively (Fuller et al., 2010; Wang et al., 2014; Chen et al., 2019). According to the classifications of gene pools initially defined by Harlan and de Wet, 1971, species in the O. sativa complex fall within the primary gene pool (Nadir et al., 2018). Despite reproductive barriers such as low cross-fertility, low F1 seed germinability and F2 hybrid weakness, common introgression within the O.sativa complex was observed and gene transfer requires traditional breeding methods (Jones et al., 1997; Jena, 2010; Zheng and Ge, 2010; Pusadee et al., 2016; Wang et al., 2017; Nadir et al., 2018). The secondary gene pool is represented by the officinalis complex and O. brachyantha, which is defined with regards to the primary gene pool as cross-incompatible with non-homologous chromosome pairing and require special techniques such as embryo rescue to achieve gene transfer (Jena, 2010). The tertiary gene pool complexes include the O. meyeriana, O. ridleyi, and unclassified O. schlecteri and O. coarctata species, which are highly cross-incompatible with O.sativa, producing anomalous, sterile or lethal hybrids (Jena, 2010; Brar and Khush, 2018). Taking into account their evolutionary history and known polyploidization events (both ancient and recent) (Ge et al., 1999; Lu et al., 2009), the WRs across the genus have successfully adapted to a wide range of habitats, which is reflected in their genomic content and associated phenotypes. A subset of these adaptive traits have been utilized for crop improvement, particularly the genes that control major biotic and abiotic stress resistances as well as yield-enhancement (Zhang K. et al., 2019).
The WRs of rice show a remarkable adaptive plasticity to a diverse set of habitats, often with extreme or suboptimal conditions (such as lack of freshwater, high temperatures, and flooding). This adaptivity has resulted in a range of beneficial traits that have been selected to “fine-tune” existing accessions of cultivated rice, which are crucial for adapting to changes in environmental conditions (Table 1A). For example, the first disease resistance gene cloned in rice was Xa21 from the AA genome WR - O. longistaminata (Song et al., 1995). This gene confers resistance to rice blast, a devastating disease of rice. Another gene, Bph18, which provides resistance to the brown planthopper, was cloned from a more distant EE genome WR – O. australiensis (Ji et al., 2016). Both of these resistance genes have been introduced into O. sativa through marker-assisted selection to produce at least 13 resistant varieties grown in India, Philippines, China, and Korea (Sanchez et al., 2013). Abiotic traits such as salt stress tolerance in O. coarctata, heat tolerance via evening flowering in O. australiensis, early morning flowering (EMF) during the cooler morning hours in O. officinalis, as well as genes involved in low temperature adaptation have been identified in O. rufipogon are also strong candidates for introgression into cultivated rice (Zeigler et al, 2014; Bheemanahalli et al., 2017; Biswal et al., 2019).
The WRs of rice have also been used as a source of other agronomic traits, such as yield. For example, O. rufipogon, the closest WR of Asian cultivated rice, was found to contain yield-enhancing QTL (yld1 and yld2). These QTL were found to be associated with increased yield (18% and 17%) relative to the high-yielding Chinese hybrid rice V64 (Xiao et al., 1996). Subsequent studies have identified other QTL in O. rufipogon linked to yield component traits, including increased grain per panicle (e.g., gpp3.1 identified in crosses with O. rufipogon and Jefferson which is a US japonica cultivar), and increased grain weight (e.g., gw1.1 and gw1.2 in crosses with Brazil Caiapo japonica cultivar) (Moncada et al., 2001; Thomson et al., 2003). Besides yield-component traits, a knowledge gap exists regarding WRs that may have superior nutritional and/or taste qualities. For example, seed from the KKLL genome WR O. coarctata is known for its sweet palatable taste and is consumed as a local delicacy in Bangladesh (Kabir and Humayun, 2012). Overall, such traits could potentially be utilized in cultivated rice to address the nutritional needs and flavor preferences of a growing population.
Beyond the association of traits to genes-of-interest, polyploidy in crops often confers greater genome plasticity and adaptation. The polyploid WRs of rice are likely to be more adaptive than their extinct diploid progenitors and may exhibit higher adaptability to extreme habitats, different climates, soil types, drought/submergence conditions, and biotic stresses (Van de Peer et al., 2017). In rice, hormone responses and differential expression of stress-response genes due to polyploidy confer autotetraploid rice with better drought-tolerance characteristics than diploid rice. Autotetraploid O. sativa (produced by colchicine-treatment to cause genome doubling) was less affected under severe drought stress, with less effect on net photosynthetic rates and peroxidation levels of its cell membranes demonstrated higher activity of the enzymes (superoxide dismutase, peroxidase, catalase) implicated in decreasing the amount of reactive oxygen species, which limited membrane lipid peroxidation (Yang et al., 2014). The authors hypothesized that the gene dosage effects due to polyploidy are the reason for drought tolerance in autotetraploid O. sativa. Similarly, polyploidy was found to confer rice growth and survival advantages under salt stress. When exposed to salt stress, four pairs of polyploid accessions, created from diploid progenitors, presented higher salt tolerance via dry weight, chlorophyll a/b content, and mortality rates of the tetraploids (Jiang et al., 2013). In addition, root ultrastructure imaging revealed decreased membrane damage and increased stability of nuclei and membrane organelles in the roots of tetraploid rice compared to diploid rice under salt stress (Tu et al., 2014). As the salt-induced production of reactive oxygen species affects membrane integrity via lipid/protein peroxidation, it is hypothesized that increased stability of membranes, nuclei and membrane organelles is an indication of normal metabolism under salt stress in roots of the tetraploid rice (Tu et al., 2014). Previously, tetraploid rootstocks in citrus have been demonstrated to be more tolerant to salt stress as compared to the corresponding diploid rootstocks (Saleh et al., 2008). These examples show that polyploidy can be advantageous due to potential dosage effects. However, a better understanding of genome content and structure is required, besides the trait-associated genes and QTL, to identify advantageous traits in the WRs for their introgression into cultivated backgrounds.
Use of High-Quality Reference Genomes to Aid in Crop Improvement
For crop improvement of the genus Oryza, breeders require access to a comprehensive set of genomic tools that can bridge the 27 species and 11 genome types that Oryza has accumulated over 15 million years of evolutionary history. The International Oryza Map Alignment Project (I-OMAP), established in 2003, set out to create a genus-level comparative genomics platform for the interrogation of the genus Oryza and provide information on rice-related genome evolution and organization, comparative genomics, physiology, biochemistry, and crop improvement. I-OMAP began with the generation of a set of publicly available deep-coverage BAC libraries and manually edited physical maps for cultivated rice and WRs used in active breeding programs (Jacquemin et al., 2013). These resources led to many early discoveries on the genome biology of Oryza, and grasses in general, and facilitated the generation of several draft genome assemblies, including that of African rice (Wang et al., 2014; Stein et al., 2018).
Recently, I-OMAP generated a set of 15 PSRefSeqs from one representative accession for each of the 15 sub-populations of O. sativa, based on the 3,000 rice genome (3K-RG) dataset, to be used as a reference guide to characterize all standing natural variation within Asian cultivated rice (Zhou et al., 2020). A PSRefSeq is defined as a high-quality near gap-free chromosome-level reference genome validated with optical maps. The O. sativa PSRefSeq dataset is composed of 12 newly sequenced genomes (Zhou et al., 2020), and 3 previously published genomes for Minghui 63 (MH63), Zhenshan 97 (ZS97) and N22 (Zhang et al., 2016a; Zhang et al., 2016b; Stein et al., 2018). The average number of contigs for these 15 assemblies is 113 contigs (i.e., 912, 237, and 181 for N22, MH63, and ZS97, respectively, and an average of 30 contigs for the remaining 12). The average number of gaps across the 15 assemblies is 46, with 8 out of 15 having less than 10 gaps (Zhou et al., 2020).
For ZS97 and MH63, genome completeness was estimated at ~92.7% (ZS97), ~94.8% (MH63) using the CEGMA pipeline with a core set of 248 eukaryotic genes (Parra et al., 2009). BUSCO evaluations for the remaining 13 genomes, which interrogate a much larger core gene set (N=956 for N22 and N=1,427 for the 12 newly sequenced assemblies), averaged 98.6%, and demonstrates the high contiguity and completeness of the majority of the O. sativa PSRefSeq dataset (Simão et al., 2015). When compared with previously released rice genomes, the O. sativa PSRefSeq data set, described here, provides more contiguous and accurate sequence data which, thereby, will improve further downstream analysis such as TE (transposable element), centromere/telomere and gene annotations.
The O. sativa 15 PSRefSeq dataset, combined with the original IRGSP RefSeq (International Rice Genome Sequencing Project, 2005; Kawahara et al., 2013), presents a multiple reference pan-genome template for the primary gene pool of Asian cultivated rice for mapping resequencing data that can accurately characterize genetic variation at the subpopulation level (Zhou et al., 2020). Given that this PSRefSeq dataset is now publicly available, the next logical step is to produce PSRefSeqs that represent the secondary and tertiary gene pools of the genus Oryza, and when combined, will create an unprecedented pan-genome for the entire genus – i.e., a “pan-genus-genome” or Oryza-PGG. Currently, I-OMAP is producing a set of PSRefSeqs from the representatives of all 25 WRs and O. glaberrima, using a combination of long-read sequencing technologies and optical maps (Udall and Dawe, 2018), with a target release date of December 2020.
The Oryza PGG will provide the full range of short, medium, and long-range structural variations that exist across the genus. Moreover, the Oryza PGG will confirm previously discovered structural variants (SVs) and the presence/absence of variants (PAVs). Zhou et al. (2020) recently showed that the majority of PAVs in the subpopulations of O. sativa are comprised of transposable elements (TEs). The Oryza PGG will allow us to better annotate and understand TEs in the WRs of rice and shed light on the contribution of TEs to genome size variation and genome plasticity of adaptive traits. For instance, the TE content of O. coarctata was estimated to be the lowest among Oryza species, despite a predicted genome size of 665 Mb (Zuccolo et al., 2007; Mondal et al., 2018).
Analyses of the Oryza PGG will also reveal the vast majority of single gene and gene family content and evolution across the genus. Previously, Zhang L. et al. (2019) leveraged the genomic data available for 13 Oryza species (domesticated and wild) to study the formation of de novo protein-coding genes. They identified 175 candidate de novo genes and estimated the rate of de novo gene origination as 51.5 de novo genes per MYA. De novo originated genes can provide novel biological functions and have previously been shown to play a key role in pathogen resistance in plants (Chen et al., 2013). For instance, OsDR10 is an Oryza-lineage specific de novo originated gene that confers broad-spectrum bacterial blight resistance when suppressed in rice (Xiao et al., 2009). Currently, OsDR10 is characterized as an “orphan” gene and there is currently little information on its evolution in the Oryza lineage. Once the Oryza PGG becomes available, the evolutionary mechanisms contributing to the generation of OsDR10 may be revealed and thus, provide insights into new gene evolution across the genus.
Finally, PSRefSeqs of each species will also aid conservation efforts by serving as “platinum standard” templates for characterizing the diversity of seed bank materials. This information will guide the management of genetic resources through in-situ conservation efforts to capture the genetic diversity in the wild or ex-situ collections of the newly identified wild populations (Brozynska et al., 2016). Some of the WRs in the genus are endangered or, as in the case of O. neocaledonica (Molla et al., 2018) and O.schlecteri absent from known genebank collections (Germplasm Resources Information Network, n.d.). From an evolutionary perspective, gained or lost SVs and copy number variations (CNVs) can be indicative of local adaptation of wild populations to unique habitats. The information gained from PSRefSeqs will help in the long-term maintenance of wild populations in their native habitats.
Barriers to Genetic Introgression and Alternative Strategies to Introgression via Neodomestication of WR Rice Species
For the successful introgression of desired traits from the WRs of rice, incompatibility barriers between the secondary, tertiary, and primary Oryza gene pools must be overcome (Harlan and de Wet, 1971; Vincent et al., 2013). The introgression of desirable traits within the AA genomes is relatively straightforward compared to the introgression of traits from more distant complexes. For instance, crossing WRs from the secondary/tertiary gene pools is more laborious and can require embryo rescue and chromosome doubling to produce interspecific hybrids and alien introgression lines (AILs). For the closely related O. officinalis complex species, monosomic alien addition lines (MAALs) and interspecific hybrids, with O. sativa as the recurrent parent, have been successfully produced to carry the following biotic stress-resistance traits: brown planthopper resistance, whitebacked planthopper, and bacterial blight resistances from O. officinalis, O. minuta, and O. latifolia, respectively (Brar and Khush, 2018).
Despite the successes achieved by introgression from the O. officinalis complex, crossing O. sativa with more distant WRs is difficult and time-consuming, and can result in progeny depression or complete progeny sterility. Moreover, linkage drag of loci, related to negative traits such as low yield, and shattering can also occur (Xiao et al., 1998). For the meyeriana complex, breeding efforts resulted in 40 derived lines of O. sativa x O. granulata, some of which were found to contain partial alien chromosome introgressions from two out of six O. granulata chromosomes analyzed. However, no favorable traits (such as cold and low radiation tolerances) were found to be introgressed from O. granulata (Brar et al., 1996). Similarly, attempts to produce hybrids from O. sativa x O. ridleyi and O. sativa x O. coarctata crosses failed due to necrosis in the progeny or progeny sterility, respectively, thereby demonstrating the difficulty of hybrid formation between distant WRs of rice with O. sativa (Brar and Khush, 2018).
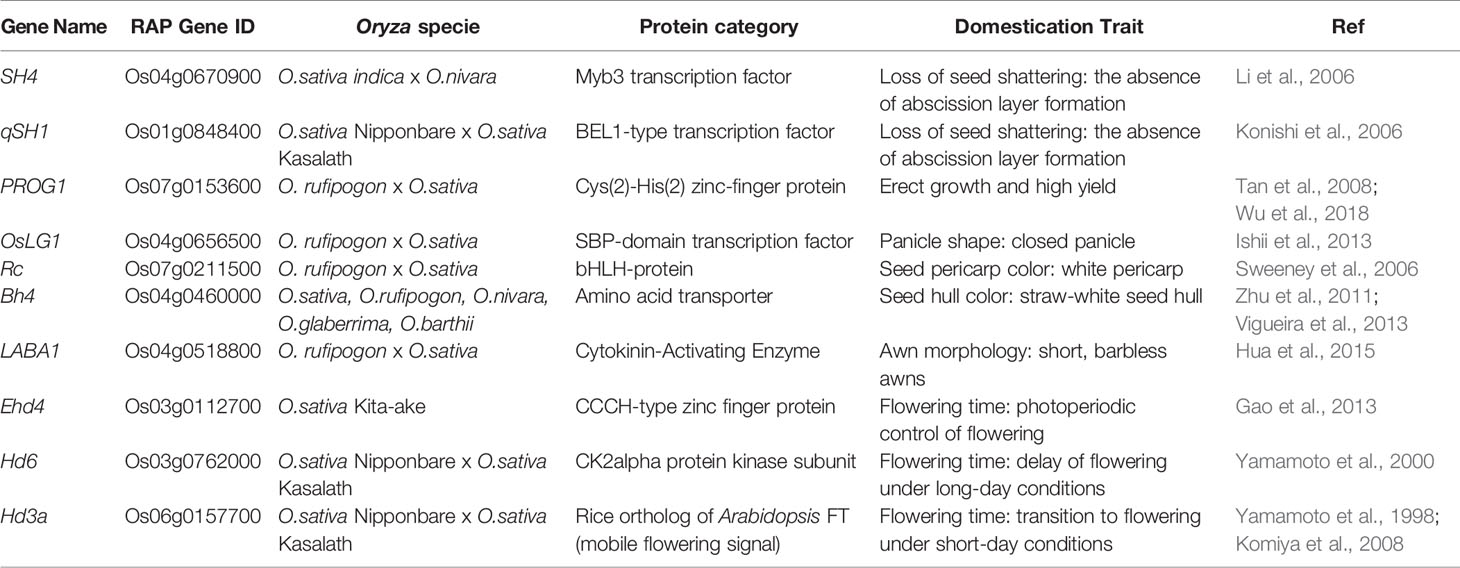
“Neodomestication” represents an alternative strategy to conventional trait introgression from a WR into a domesticated species. This method preserves adaptations to biotic/abiotic stresses while exhibiting the traits of a domesticated crop (e.g., loss of shattering, erect growth, larger seeds). For example, homologues of genes involved in the domestication of rice in the WRs could be targeted for neodomestication to produce a domesticated ideotype in a WR. Shapter et al. (2013) presented an early example of an accelerated neodomestication process using random EMS mutagenesis to initiate the domestication of weeping rice grass (Microlaena stipoides), which resulted in the generation of two non-shattering mutant plants in the orthologs of qSH1 and sh4. Now, with gene editing technology such as CRISPR-Cas9 (Bortesi and Fischer, 2015), it is possible to precisely edit homologues of well-studied domestication genes in crop WRs, such as those listed in Table 1B, to activate genes, modify alleles, introduce targeted base substitutions, delete large genomic segments, or introduce complete genes previously unavailable in a given species (Fernie and Yan, 2019). Besides targeting orthologues of domestication genes for neodomestication of WR Oryza candidates, CRISPR/Cas9 gene editing could be used to introduce WR candidate genes implicated in favourable traits such as abiotic stress tolerances into the cultivated rice background by overcoming the reproductive Oryza gene pool barriers (Harlan and de Wet, 1971).
As a proof-of-concept study for neodomestication using gene editing, Zsögön et al. (2018) targeted six key domestication traits for loss of function in the wild tomato S. pimpinellifolium (i.e., SP, SELF-PRUNING; O, OVATE; FW2.2, FRUIT WEIGHT; LYCOPENE BETA CYCLASE, FASCIATED, MULTIFLORA). Four of the six genes (i.e., all except FASCIATED and MULTIFLORA) were successfully targeted and contained indel mutations, which resulted in a WR with a domesticated “phenotype”, i.e., an altered morphology, a three-fold increase in fruit size, a ten-fold increase in fruit number compared to the WR, and a 500% increase in lycopene accumulation in the fruit, relative to the cultivated Solanum lycopersicum (Zsögön et al., 2018).
Similarly to wild tomato, neodomestication of the WRs of rice is theoretically possible, albeit non-trivial. O. coarctata is an attractive candidate for neodomestication because it is the only halophyte in the Oryza genus and can thrive in salinity levels of up to 40 E.CedS m−1 (i.e., brackish water salinity levels) along the coastal regions from Pakistan to Myanmar (Bal and Dutt, 1986). In India, the habitat of this species is similar to that of mangrove forests where O. coarctata can be found submerged in saline water for up to 12 h a day due to lunar tides (Bal and Dutt, 1986; Mondal et al., 2018). For decades, rice breeders have tried to introgress the halophyte characteristics of O. coarctata into cultivated rice with limited or no success (Brar and Khush, 2018; Prusty et al., 2018). Thus, O. coarctata is an ideal candidate for neodomestication and, if successful, would allow farmers to grow rice on land that otherwise is unable to support conventional rice farming practices. A proposed methodology for the neodomestication of O. coarctata includes the targeting of orthologues of domestication genes in O. coarctata (Table 1B) with precise gene editing. An O. coarctata PSRefSeq will enable less off-target CRISPR/Cas9 enzyme effects (not attributed to the enzyme mode of action) as the contiguity and fidelity of the PSRefSeq allows for better precision for single guide RNA (sgRNA) selection (Biswal et al., 2019). Moreover, O. coarctata, unlike S. pimpinellifolium, is not a direct progenitor of the cultivated O. sativa and is a polyploid which, besides affecting the number of domestication-related genes, also translates to a vast difference in the genome structure, thereby making gene editing more challenging due to the potential linkage of domestication genes to non-desirable traits (i.e., linkage drag). Given the paucity of available genome assembly resources, a PSRefSeq for O. coarctata would play a vital role in the precise identification of the orthologues of domestication genes that could be targeted for neodomestication.
Author Contributions
SM and NA-B contributed equally to this work and share co-first authorship. SM and NA-B wrote the manuscript. AZ contributed to writing the manuscript, editing and organizing the structure of the manuscript. RW reviewed and edited the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by KAUST to RW.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Atwell, B. J., Wang, H., Scafaro, A. P. (2014). Could abiotic stress tolerance in wild relatives of rice be used to improve Oryza sativa? Plant Sci. 215–216, 48–58. doi: 10.1016/j.plantsci.2013.10.007
Bal, A. R., Dutt, S. K. (1986). Mechanism of salt tolerance in wild rice (Oryza coarctata Roxb). Plant Soil 92 (3), 399–404. doi: 10.1007/BF02372487
Bheemanahalli, R., Sathishraj, R., Manoharan, M., Sumanth, H. N., Muthurajan, R., Ishimaru, T., et al. (2017). Is early morning flowering an effective trait to minimize heat stress damage during flowering in rice? Field Crops Res. 203, 238–242. doi: 10.1016/j.fcr.2016.11.011
Biswal, A. K., Mangrauthia, S. K., Reddy, M. R., Yugandhar, P. (2019). CRISPR mediated genome engineering to develop climate smart rice: Challenges and opportunities. Semin. Cell Dev. Biol. 96, 100–106. doi: 10.1016/j.semcdb.2019.04.005
Bortesi, L., Fischer, R. (2015). The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 33 (1), 41–52. doi: 10.1016/j.biotechadv.2014.12.006
Brar, D. S., Khush, G. S. (2018). “Wild Relatives of Rice: A Valuable Genetic Resource for Genomics and Breeding Research,” in The Wild Oryza Genomes. Eds. Mondal, T. K., Henry, R. J. Compendium of Plant Genomes. (Cham: Springer International Publishing), 1–25. doi: 10.1007/978-3-319-71997-9_1
Brar, D. S., Dalmacio, R., Elloran, R., Aggarwal, R., Angeles, R., Khush, G. S. (1996). “Gene transfer and molecular characterization of introgression from wild Oryza species into rice,” in Rice genetics III (Manila, Philippines: IRRI), 477–486.
Brozynska, M., Furtado, A., Henry, R. J. (2016). Genomics of crop wild relatives: expanding the gene pool for crop improvement. Plant Biotechnol. J. 14 (4), 1070–1085. doi: 10.1111/pbi.12454
Chen, S., Krinsky, B. H., Long, M. (2013). New genes as drivers of phenotypic evolution. Nat. Rev. Genet. 14 (9), 645–660. doi: 10.1038/nrg3521
Chen, E., Huang, X., Tian, Z., Wing, R. A., Han, B. (2019). The Genomics of Oryza Species Provides Insights into Rice Domestication and Heterosis. Annu. Rev. Plant Biol. 70, 639–665. doi: 10.1146/annurev-arplant-050718-100320
Doebley, J. F., Gaut, B. S., Smith, B. D. (2006). The Molecular Genetics of Crop Domestication. Cell 127 (7), 1309–1321. doi: 10.1016/j.cell.2006.12.006
Fernie, A. R., Yan, J. (2019). De Novo Domestication: An Alternative Route toward New Crops for the Future. Mol. Plant 12 (5), 615–631. doi: 10.1016/j.molp.2019.03.016
Fuller, D. Q., Sato, Y.-I., Castillo, C., Qin, L., Weisskopf, A. R., Kingwell-Banham, E. J., et al. (2010). Consilience of genetics and archaeobotany in the entangled history of rice. Archaeol. Anthropol. Sci. 2 (2), 115–131. doi: 10.1007/s12520-010-0035-y
Gao, H., Zheng, X.-M., Fei, G., Chen, J., Jin, M., Ren, Y., et al. (2013). Ehd4 encodes a novel and Oryza-genus-specific regulator of photoperiodic flowering in rice. PloS Genet. 9 (2), e1003281. doi: 10.1371/journal.pgen.1003281
Ge, S., Sang, T., Lu, B.-R., Hong, D.-Y. (1999). Phylogeny of rice genomes with emphasis on origins of allotetraploid species. Proc. Natl. Acad. Sci. 96 (25), 14400–14405. doi: 10.1073/pnas.96.25.14400
Germplasm Resources Information Network GRIN-Global Web v 1.10.6.1. (n.d.). Available at: https://gringlobal.irri.org/ ( Retrieved August 26, 2020).
Harlan, J. R., de Wet, J. M. J. (1971). Toward a Rational Classification of Cultivated Plants. Taxon. 20 (4), 509–517. doi: 10.2307/1218252. JSTOR.
Hua, L., Wang, D. R., Tan, L., Fu, Y., Liu, F., Xiao, L., et al. (2015). LABA1, a Domestication Gene Associated with Long, Barbed Awns in Wild Rice. Plant Cell 27 (7), 1875–1888. doi: 10.1105/tpc.15.00260
International Rice Genome Sequencing Project (2005). The map-based sequence of the rice genome. Nature 436, 793–800. doi: 10.1038/nature03895
Ishii, T., Numaguchi, K., Miura, K., Yoshida, K., Thanh, P. T., Htun, T. M., et al. (2013). OsLG1 regulates a closed panicle trait in domesticated rice. Nat. Genet. 45 (4), 462–465, 465e1-2. doi: 10.1038/ng.2567
Jacquemin, J., Bhatia, D., Singh, K., Wing, R. A. (2013). The International Oryza Map Alignment Project: development of a genus-wide comparative genomics platform to help solve the 9 billion-people question. Curr. Opin. Plant Biol. 16 (2), 147–156. doi: 10.1016/j.pbi.2013.02.014
Jena, K. K. (2010). The species of the genus Oryza and transfer of useful genes from wild species into cultivated rice, O. sativa. Breed. Sci. 60 (5), 518–523. doi: 10.1270/jsbbs.60.518
Ji, H., Kim, S.-R., Kim, Y.-H., Suh, J.-P., Park, H.-M., Sreenivasulu, N., et al. (2016). Map-based Cloning and Characterization of the BPH18 Gene from Wild Rice Conferring Resistance to Brown Planthopper (BPH) Insect Pest. Sci. Rep. 6 (1), 34376. doi: 10.1038/srep34376
Jiang, A., Gan, L., Tu, Y., Ma, H., Zhang, J., Song, Z., et al. (2013). The effect of genome duplication on seed germination and seedling growth of rice under salt stress. Aust. J. Crop Sci. 7, 1814–1821.
Jones, M. P., Dingkuhn, M., Aluko, G. K., Semon, M. (1997). Interspecific Oryza Sativa L. X O. Glaberrima Steud. progenies in upland rice improvement. Euphytica 94 (2), 237–246. doi: 10.1023/A:1002969932224
Kabir, S. M., Humayun (2012). “Rice,” in Banglapedia: National Encyclopedia of Bangladesh, 2nd ed. Eds. Islam, S., Jamal, A. A. (Dhaka, Bangladesh: Asiatic Society of Bangladesh).
Kawahara, Y., de la Bastide, M., Hamilton, J. P., Kanamori, H., McCombie, W. R., Ouyang, S., et al. (2013). Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6 (1), 4. doi: 10.1186/1939-8433-6-4
Komiya, R., Ikegami, A., Tamaki, S., Yokoi, S., Shimamoto, K. (2008). Hd3a and RFT1 are essential for flowering in rice. Development 135 (4), 767–774. doi: 10.1242/dev.008631
Konishi, S., Izawa, T., Lin, S. Y., Ebana, K., Fukuta, Y., Sasaki, T., et al. (2006). An SNP Caused Loss of Seed Shattering During Rice Domestication. Science 312 (5778), 1392–1396. doi: 10.1126/science.1126410
Li, C., Zhou, A., Sang, T. (2006). Rice Domestication by Reducing Shattering. Science 311 (5769), 1936–1939. doi: 10.1126/science.1123604
Lu, F., Ammiraju, J. S. S., Sanyal, A., Zhang, S., Song, R., Chen, J., et al. (2009). Comparative sequence analysis of MONOCULM1-orthologous regions in 14 Oryza genomes. Proc. Natl. Acad. Sci. 106 (6), 2071–2076. doi: 10.1073/pnas.0812798106
Menguer, P. K., Sperotto, R. A., Ricachenevsky, F. K. (2017). A walk on the wild side: Oryza species as source for rice abiotic stress tolerance. Genet. Mol. Biol. 40 (1 Suppl 1), 238–252. doi: 10.1590/1678-4685-GMB-2016-0093
Milovanovic, V., Smutka, L. (2017). Asian Countries in the Global Rice Market. Acta Universitatis Agric. Silvic. Mendelianae Brunensis 65 (2), 679–688. doi: 10.11118/actaun201765020679
Molla, K. A., Karmakar, S., Molla, J., Azharudheen, T. P. M., Datta, K. (2018). “Oryza Neocaledonica Morat,” in The Wild Oryza Genomes. Eds. Mondal, T. K., Henry, R. J. Compendium of Plant Genomes. (Cham: Springer International Publishing), 203–206. doi: 10.1007/978-3-319-71997-9_19
Moncada, P., Martínez, C. P., Borrero, J., Chatel, M., Gauch, H., Jr., Guimaraes, E., et al. (2001). Quantitative trait loci for yield and yield components in an Oryza sativa×Oryza rufipogon BC2F2 population evaluated in an upland environment. Theor. Appl. Genet. 102 (1), 41–52. doi: 10.1007/s001220051616
Mondal, T. K., Rawal, H. C., Chowrasia, S., Varshney, D., Panda, A. K., Mazumdar, A., et al. (2018). Draft genome sequence of first monocot-halophytic species Oryza coarctata reveals stress-specific genes. Sci. Rep. 8 (1), 13698. doi: 10.1038/s41598-018-31518-y
Nadir, S., Khan, S., Zhu, Q., Henry, D., Wei, L., Lee, D. S., et al. (2018). An overview on reproductive isolation in Oryza sativa complex. AoB Plants 10 (6). doi: 10.1093/aobpla/ply060
Parra, G., Bradnam, K., Ning, Z., Keane, T., Korf, I. (2009). Assessing the gene space in draft genomes. Nucleic Acids Res. 37 (1), 289–297. doi: 10.1093/nar/gkn916
Prusty, M. R., Kim, S.-R., Vinarao, R., Entila, F., Egdane, J., Diaz, M. G. Q., et al. (2018). Newly Identified Wild Rice Accessions Conferring High Salt Tolerance Might Use a Tissue Tolerance Mechanism in Leaf. Front. Plant Sci. 9, 43–57. doi: 10.3389/fpls.2018.00417
Pusadee, T., Jamjod, S., Rerkasem, B., Schaal, B. A. (2016). Life-history traits and geographical divergence in wild rice (Oryza rufipogon) gene pool in Indochina Peninsula region. Ann. Appl. Biol. 168 (1), 52–65. doi: 10.1111/aab.12242
Saleh, B., Allario, T., Dambier, D., Ollitrault, P., Morillon, R. (2008). Tetraploid citrus rootstocks are more tolerant to salt stress than diploid. Comptes Rendus Biol. 331 (9), 703–710. doi: 10.1016/j.crvi.2008.06.007
Sanchez, P. L., Wing, R. A., Brar, D. S. (2013). “The Wild Relative of Rice: Genomes and Genomics,” in Genetics and Genomics of Rice. Eds. Zhang, Q., Wing, R. A. Plant Genetics and Genomics: Crops and Models. (New York, NY: Springer), 9–25. doi: 10.1007/978-1-4614-7903-1_2
Shapter, F. M., Cross, M., Ablett, G., Malory, S., Chivers, I. H., King, G. J., et al. (2013). High-throughput sequencing and mutagenesis to accelerate the domestication of Microlaena stipoides as a new food crop. PloS One 8 (12), e82641. doi: 10.1371/journal.pone.0082641
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V., Zdobnov, E. M. (2015). BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31 (19), 3210–3212. doi: 10.1093/bioinformatics/btv351
Song, W.-Y., Wang, G.-L., Chen, L.-L., Kim, H.-S., Pi, L.-Y., Holsten, T., et al. (1995). A Receptor Kinase-Like Protein Encoded by the Rice Disease Resistance Gene, Xa21. Science 270 (5243), 1804–1806. doi: 10.1126/science.270.5243.1804
Stein, J. C., Yu, Y., Copetti, D., Zwickl, D. J., Zhang, L., Zhang, C., et al. (2018). Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover and innovation across the genus Oryza. Nat. Genet. 50 (2), 285–296. doi: 10.1038/s41588-018-0040-0
Sweeney, M. T., Thomson, M. J., Pfeil, B. E., McCouch, S. (2006). Caught Red-Handed: Rc Encodes a Basic Helix-Loop-Helix Protein Conditioning Red Pericarp in Rice. Plant Cell 18 (2), 283–294. doi: 10.1105/tpc.105.038430
Tan, L., Li, X., Liu, F., Sun, X., Li, C., Zhu, Z., et al. (2008). Control of a key transition from prostrate to erect growth in rice domestication. Nat. Genet. 40 (11), 1360–1364. doi: 10.1038/ng.197
Thomson, M. J., Tai, T. H., McClung, A. M., Lai, X.-H., Hinga, M. E., Lobos, K. B., et al. (2003). Mapping quantitative trait loci for yield, yield components and morphological traits in an advanced backcross population between Oryza rufipogon and the Oryza sativa cultivar Jefferson. Theor. Appl. Genet. 107 (3), 479–493. doi: 10.1007/s00122-003-1270-8
Toriyama, K. (2005). “Rice Is Life: Scientific Perspectives for the 21st Century,” in World Rice Research Conference eng 4-7 Nov 2004 Tsukuba (Japan). Eds. Heong, K. L., Hardy, B. (Los Banos (Philippines: eng International Rice Research Inst.). Available at: https://agris.fao.org/agris-search/search.do?recordID=XF2015014688.
Tu, Y., Jiang, A., Gan, L., Hossain, M., Zhang, J., Peng, B., et al. (2014). Genome duplication improves rice root resistance to salt stress. Rice 7 (1):15. doi: 10.1186/s12284-014-0015-4
Udall, J. A., Dawe, R. K. (2018). Is It Ordered Correctly? Validating Genome Assemblies by Optical Mapping. Plant Cell 30 (1), 7–14. doi: 10.1105/tpc.17.00514
Van de Peer, Y., Mizrachi, E., Marchal, K. (2017). The evolutionary significance of polyploidy. Nat. Rev. Genet. 18 (7), 411–424. doi: 10.1038/nrg.2017.26
Vaughan, D. A. (1989). Genus Oryza L. current status of taxonomy (Philippines: IRRI Research Paper Series). Available at: https://agris.fao.org/agris-search/search.do?recordID=PH9010360.
Vigueira, C. C., Li, W., Olsen, K. M. (2013). The role of Bh4 in parallel evolution of hull colour in domesticated and weedy rice. J. Evol. Biol. 26 (8), 1738–1749. doi: 10.1111/jeb.12171
Vikal, Y., Das, A., Patra, B., Goel, R. K., Sidhu, J. S., Singh, K. (2007). Identification of new sources of bacterial blight (Xanthomonas oryzae pv. oryzae) resistance in wild Oryza species and O. glaberrima. Plant Genet. Resources: Character. Util. 5 (02), 108–112. doi: 10.1017/S147926210777661X
Vincent, H., Wiersema, J., Kell, S., Fielder, H., Dobbie, S., Castañeda-Álvarez, N. P., et al. (2013). A prioritized crop wild relative inventory to help underpin global food security. Biol. Conserv. 167, 265–275. doi: 10.1016/j.biocon.2013.08.011
Wang, M., Yu, Y., Haberer, G., Marri, P. R., Fan, C., Goicoechea, J. L., et al. (2014). The genome sequence of African rice ( Oryza glaberrima ) and evidence for independent domestication. Nat. Genet. 46 (9), 982–988. doi: 10.1038/ng.3044.
Wang, H., Vieira, F. G., Crawford, J. E., Chu, C., Nielsen, R. (2017). Asian wild rice is a hybrid swarm with extensive gene flow and feralization from domesticated rice. Genome Res. 27 (6), 1029–1038. doi: 10.1101/gr.204800.116
Wu, Y., Zhao, S., Li, X., Zhang, B., Jiang, L., Tang, Y., et al. (2018). Deletions linked to PROG1 gene participate in plant architecture domestication in Asian and African rice. Nat. Commun. 9 (1), 4157. doi: 10.1038/s41467-018-06509-2
Xiao, J., Grandillo, S., Ahn, S. N., McCouch, S. R., Tanksley, S. D., Li, J., et al. (1996). Genes from wild rice improve yield. Nature 384 (6606), 223–224. doi: 10.1038/384223a0
Xiao, J., Li, J., Grandillo, S., Ahn, S. N., Yuan, L., Tanksley, S. D., et al. (1998). Identification of trait-improving quantitative trait loci alleles from a wild rice relative, Oryza rufipogon. Genetics 150 (2), 899–909.
Xiao, W., Liu, H., Li, Y., Li, X., Xu, C., Long, M., et al. (2009). A Rice Gene of De Novo Origin Negatively Regulates Pathogen-Induced Defense Response. PloS One 4 (2), e4603. doi: 10.1371/journal.pone.0004603
Yamamoto, T., Kuboki, Y., Lin, S. Y., Sasaki, T., Yano, M. (1998). Fine mapping of quantitative trait loci Hd-1, Hd-2 and Hd-3, controlling heading date of rice, as single Mendelian factors. Theor. Appl. Genet. 97 (1–2), 37–44. doi: 10.1007/s001220050864
Yamamoto, T., Lin, H., Sasaki, T., Yano, M. (2000). Identification of heading date quantitative trait locus Hd6 and characterization of its epistatic interactions with Hd2 in rice using advanced backcross progeny. Genetics 154 (2), 885–891.
Yang, P.-M., Huang, Q.-C., Qin, G.-Y., Zhao, S.-P., Zhou, J.-G. (2014). Different drought-stress responses in photosynthesis and reactive oxygen metabolism between autotetraploid and diploid rice. Photosynthetica 52 (2), 193–202. doi: 10.1007/s11099-014-0020-2
Zeigler, R. S. (2014). “Food security, climate change and genetic resources,” in Plant genetic resources and climate change. Eds. Jackson, M., Ford-Lloyd, B., Parry, M. (Wallingford: CABI), 1–15. doi: 10.1079/9781780641973.0001
Zhang, J., Chen, L.-L., Xing, F., Kudrna, D. A., Yao, W., Copetti, D., et al. (2016a). Extensive sequence divergence between the reference genomes of two elite indica rice varieties Zhenshan 97 and Minghui 63. Proc. Natl. Acad. Sci. 113 (35), E5163–E5171. doi: 10.1073/pnas.1611012113
Zhang, J., Chen, L.-L., Sun, S., Kudrna, D., Copetti, D., Li, W., et al. (2016b). Building two indica rice reference genomes with PacBio long-read and Illumina paired-end sequencing data. Sci. Data 3 (1), 160076. doi: 10.1038/sdata.2016.76
Zhang, K., Wang, X., Cheng, F. (2019). Plant Polyploidy: Origin, Evolution, and Its Influence on Crop Domestication. Hortic. Plant J. 5 (6), 231–239. doi: 10.1016/j.hpj.2019.11.003
Zhang, L., Ren, Y., Yang, T., Li, G., Chen, J., Gschwend, A. R., et al. (2019). Rapid evolution of protein diversity by de novo origination in Oryza. Nat. Ecol. Evol. 3 (4), 679–690. doi: 10.1038/s41559-019-0822-5
Zheng, X.-M., Ge, S. (2010). Ecological divergence in the presence of gene flow in two closely related Oryza species (Oryza rufipogon and O. nivara). Mol. Ecol. 19 (12), 2439–2454. doi: 10.1111/j.1365-294X.2010.04674.x.
Zhou, Y., Chebotarov, D., Kudrna, D., Llaca, V., Lee, S., Rajasekar, S., et al. (2020). A platinum standard pan-genome resource that represents the population structure of Asian rice. Sci. Data 7 (1), 1–11. doi: 10.1038/s41597-020-0438-2
Zhu, B.-F., Si, L., Wang, Z., Jingjie Zhu, Y. Z., Shangguan, Y., Lu, D., et al. (2011). Genetic Control of a Transition from Black to Straw-White Seed Hull in Rice Domestication1[C][W][OA]. Plant Physiol. 155 (3), 1301–1311. doi: 10.1104/pp.110.168500
Zsögön, A., Čermák, T., Naves, E. R., Notini, M. M., Edel, K. H., Weinl, S., et al. (2018). De novo domestication of wild tomato using genome editing. Nat. Biotechnol. 36 (12), 1211–1216. doi: 10.1038/nbt.4272
Keywords: neodomestication, rice wild relatives, platinum reference genomes, Oryza coarctata, Oryza
Citation: Mussurova S, Al-Bader N, Zuccolo A and Wing RA (2020) Potential of Platinum Standard Reference Genomes to Exploit Natural Variation in the Wild Relatives of Rice. Front. Plant Sci. 11:579980. doi: 10.3389/fpls.2020.579980
Received: 03 July 2020; Accepted: 04 September 2020;
Published: 23 September 2020.
Edited by:
Yohei Koide, Hokkaido University, JapanReviewed by:
Tomoyuki Furuta, Okayama University, JapanKanako Bessho-Uehara, Tohoku University, Japan
Copyright © 2020 Mussurova, Al-Bader, Zuccolo and Wing. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Andrea Zuccolo, YW5kcmVhLnp1Y2NvbG9Aa2F1c3QuZWR1LnNh; Rod A. Wing, cm9kLndpbmdAa2F1c3QuZWR1LnNh
 Saule Mussurova
Saule Mussurova Noor Al-Bader
Noor Al-Bader Andrea Zuccolo
Andrea Zuccolo Rod A. Wing
Rod A. Wing