- 1Departmentof Plant Nutrition, Institute of Crop Science and Resource Conservation, Rheinische Friedrich-Wilhelms-Universität Bonn, Bonn, Germany
- 2Department of Biochemistry, Indian Institute of Science, Bengaluru, India
- 3Department of Chemistry and Pharmacy & CIBSS – The Center of Biological Signaling Studies, Albert-Ludwigs University Freiburg, Freiburg, Germany
Inositol pyrophosphates (PP-InsPs), derivatives of inositol hexakisphosphate (phytic acid, InsP6) or lower inositol polyphosphates, are energy-rich signaling molecules that have critical regulatory functions in eukaryotes. In plants, the biosynthesis and the cellular targets of these messengers are not fully understood. This is because, in part, plants do not possess canonical InsP6 kinases and are able to synthesize PP-InsP isomers that appear to be absent in yeast or mammalian cells. This review will shed light on recent discoveries in the biosynthesis of these enigmatic messengers and on how they regulate important physiological processes in response to abiotic and biotic stresses in plants.
Introduction
Inositol phosphates (InsPs) belong to the multifaceted family of signaling molecules that control a plethora of physiological processes across the eukaryote landscape (Shears, 2015; Thota and Bhandari, 2015; Laha et al., 2021a). These molecules are based on a six-carbon ring structure, cis-1,2,3,5-trans-4,6-cyclohexanehexol, commonly referred to as myo-inositol (Shears, 2015). Combinatorial phosphorylation of the myo-inositol ring could generate a large array of InsP species, of which only a few were identified in cell extracts (Shears, 2015). The physiological functions of most of these InsP species are largely unexplored. Almost 40 years ago, inositol 1,4,5-trisphosphate [Ins(1,4,5)P3] was shown for the first time to act as a second messenger, by acting as a calcium release factor that stimulates its specific receptor/Ca2+-permeable ion channel on endomembranes in pancreatic acinar cells (Streb et al., 1983; Irvine, 2003). In land plants, changes in intracellular InsP3 levels were shown to be responsive to various factors, such as root gravitropism, heat shock signal transduction, mechanical wounding, osmotic stress, pollen dormancy and blue light perception (Knight et al., 1997; Liu et al., 2006; Chen et al., 2008; Mosblech et al., 2008; Wang et al., 2009, 2012). These responses to InsP3 were assumed to be mediated by cytosolic Ca2+, as several studies showed that either treatment with caged photoactivatable compounds to release cytosolic InsP3 or direct InsP3 microinjection resulted in a transient increase in cytosolic Ca2+ (Blatt et al., 1990; Gilroy et al., 1990; Allen and Sanders, 1994; Tucker and Boss, 1996; Monteiro et al., 2005).
Additionally, it was shown that tomato plants expressing the human type I inositol polyphosphate 5-phosphatase (InsP5-ptase), a key enzyme in the phosphoinositide pathway, were more tolerant to drought and light stress, a phenotype that was suggested to be caused by the decrease of InsP3 detected in those plants (Khodakovskaya et al., 2010; Alimohammadi et al., 2012). Notably, even though InsP1, InsP2, InsP3, and InsP4 levels were shown to be decreased in InsP5-ptase-expressing plants, the role of other InsP species including PP-InsPs were not considered in these studies.
Importantly, genomes of flowering plants do not encode homologs of mammalian InsP3 receptors, which appear to have been lost during the course of evolution (Krinke et al., 2007; Wheeler and Brownlee, 2008; Munnik and Testerink, 2009; Munnik and Vermeer, 2010; Munnik and Nielsen, 2011; Zhang et al., 2018). Therefore, the role of Ins(1,4,5)P3 in plants remains unresolved. InsP6, also known as myo-inositol 1,2,3,4,5,6 hexakisphosphate, phytic acid or phytate, is the most abundant form of InsPs in eukaryotes, with concentrations in the range of 10–100 μM in animal and yeast cells, and up to 500 μM in slime molds (Wundenberg and Mayr, 2012; Pisani et al., 2014). InsP6 is the fully phosphorylated version of myo-inositol and serves as a phosphate (Pi) storage molecule during seed development. In this process, InsP6 accumulates in storage microbodies in the form of mixed salts with cations, such as zinc, calcium, iron, potassium, magnesium and manganese (Raboy and Gerbasi, 1996; Otegui et al., 2002; Secco et al., 2017). The storage protein bodies are then degraded during seed germination, leading to the rapid hydrolysis of InsP6 by phytases to provide nutrients to the developing seedling (Raboy and Gerbasi, 1996; Loewus and Murthy, 2000). Due to its strong affinity toward different mineral cations, InsP6 is considered an antinutrient for humans and non-ruminant animals (McCance and Widdowson, 1942; Halsted et al., 1972). Since non-ruminant animals (e.g., pigs and poultry) lack phytases in their digestive tracts, excrements derived from phytate-rich diet contain phytate-bound phosphorus, which is often released in open water bodies, leading to eutrophication and environmental pollution (Rockström et al., 2009; Raboy, 2020).
InsP6 also serves as an important signaling molecule, directly or indirectly as a precursor of “di/pyro-phosphate”-containing inositol polyphosphates, commonly referred to as inositol pyrophosphates (PP-InsPs). These energy-rich InsP species are ubiquitous in eukaryotes, with InsP7 and InsP8 representing the most well-characterized species (Wilson et al., 2013; Shears, 2015). In plants, PP-InsPs control a range of important biological functions, including immune responses, hormone perception, and phosphate homeostasis (Zhang et al., 2007; Jadav et al., 2013; Laha et al., 2015, 2016, 2020; Jung et al., 2018; Kuo et al., 2018; Dong et al., 2019; Zhu et al., 2019; Gulabani et al., 2021; Land et al., 2021; Riemer et al., 2021).
The metabolic pathways leading to the production of PP-InsPs are well-established in metazoan and yeast. In these organisms, mammalian IP6K/yeast Kcs1-type kinases catalyze the phosphorylation of InsP6 or 1-InsP7 at the 5 position, resulting in the generation of 5-InsP7 or 1,5-InsP8, respectively (Saiardi et al., 1999; Draskovic et al., 2008). Furthermore, mammalian PPIP5K/yeast Vip1 kinases phosphorylate the 1 position of InsP6 and 5-InsP7 to generate 1-InsP7 and 1,5-InsP8, respectively (Mulugu et al., 2007; Lin et al., 2009; Wang et al., 2011; Zhu et al., 2019). The PP-InsP biosynthetic pathway is partially conserved in plants. For instance, while the Arabidopsis genome encodes Vip1 isoforms, genes encoding Kcs1-type kinase(s) could not be identified yet. However, recent studies have demonstrated that the Arabidopsis thaliana kinases ITPK1 and ITPK2 phosphorylate InsP6, which is first generated by the phosphorylation of InsP5 [2-OH] by IPK1, to synthesize 5-InsP7 in vitro (Adepoju et al., 2019; Laha et al., 2019; Whitfield et al., 2020) and in planta (Parvin Laha et al., 2020; Riemer et al., 2021). These proteins belong to the family of ATP-grasp fold proteins with the capability to bind ATP in a cleft between the β sheet toward the central and C-terminal domain (Miller et al., 2005; Josefsen et al., 2007). Notably, their homologs ITPK3 and ITPK4 do not appear to phosphorylate InsP6 in vitro or in vivo (Laha et al., 2019). The Arabidopsis Vip1 isoforms VIH1 and VIH2 harbor both an N-terminal ATP-grasp kinase domain, as well as a C-terminal phosphatase-like domain and are responsible for InsP8 production in planta (Figure 1A; Laha et al., 2015; Zhu et al., 2019).

Figure 1. Inositol pyrophosphate biosynthesis pathway in plants and protein architecture of kinases. (A) ITPK1/2 and VIH1/2 phosphorylate InsP6 to generate 5-InsP7 and 1-InsP7, respectively. Further, VIH1/2 use 5-InsP7 as substrate to generate InsP8. The isomer identity of InsP8 remains unresolved but presumably represents the 1,5 and/or the 3,5-InsP8 isomer. VIH1/2 are also able to dephosphorylate 1/3,5-InsP8 and 1/3-InsP7 to InsP6. At low adenylate charge, the ITPK1 kinase domain also catalyzes the reverse reaction from 5-InsP7 to InsP6 in the presence of ADP to locally generate ATP. The gray arrows and question marks denote alternative routes of 1/3-InsP7 and 1/3,5-InsP8 synthesis, and the responsible enzymes, respectively. (B) Schematic representation of ITPK1, ITPK2, VIH1 and VIH2 architectures. Kinase domains are shown in dark gray, phosphatase domains in light gray.
The recent establishment of novel methods for InsP analyses led to the emergence of several plant PP-InsP studies, which have been instrumental to establish PP-InsP as novel signaling molecules in plants. Remarkably, to date it is still challenging to separate different PP-InsP isomers. This leads to the open question, whether the enantiomers 1-InsP7 or 3-InsP7 and 1,5-InsP8 or 3,5-InsP8 are the main isomers in plants or, if both exist, which of them is the most abundant.
Inositol pyrophosphates play a crucial role in the adaption to several stress responses in plants. Previous work has demonstrated the relevance of InsP7 and InsP8 in responses to hormones such as auxin, salicylic acid or jasmonate (Laha et al., 2015, 2016; Parvin Laha et al., 2020; Gulabani et al., 2021).
In addition, Pi homeostasis in plants was shown to be regulated by kinases involved in InsP synthesis (Kuo et al., 2014, 2018), most likely due to their contribution to the synthesis of InsP8, which serves as a proxy for Pi (Dong et al., 2019; Zhu et al., 2019; Riemer et al., 2021). Interestingly, certain bacterial plant pathogens interfere with plant InsP6- and potentially PP-InsP-dependent hormone signaling by injecting XopH-like type III effectors that function as 1-phytases (Blüher et al., 2017). However, it is still unclear how this modulation of the host’s InsP and PP-InsP status benefits the pathogen (Blüher et al., 2017). Beyond that, recent studies demonstrated a link between pathogen defense and Pi starvation, displaying InsPs and PP-InsPs as crosstalk mediators of abiotic and biotic stresses (Gulabani et al., 2021).
In this review, we present in detail the latest findings of the roles of these phosphate-rich molecules in regulating different biotic and abiotic responses in plants.
Enzymatic activity of PP-InsP kinases
The function of InsP and PP-InsP kinases is not only limited to the generation of higher inositol pyrophosphates, as they can also shift their activity from PP-InsP synthases to ATP synthases in response to different ATP ratios (Voglmaier et al., 1996; Gu et al., 2017; Zhu et al., 2019; Riemer et al., 2021). It was already shown that mammalian IP6K kinases can transfer a phosphate group from InsP7 to ADP to generate ATP (Voglmaier et al., 1996). Furthermore, both mammalian and yeast IP6K/Kcs1 activities react to changes in cellular ATP levels with respect to the generation of 5-InsP7 (Saiardi et al., 1999; Gu et al., 2017).
In contrast to IP6K/Kcs1, which harbor only a kinase domain, mammalian and yeast PPIP5K/Vip1 harbor both an N-terminal kinase domain and a C-terminal phosphatase domain in the same protein, enabling them to act as bifunctional enzymes (Fridy et al., 2007; Mulugu et al., 2007; Wang et al., 2015; Zhu et al., 2019). As mentioned above, Arabidopsis VIH1 and VIH2 also possess an N-terminal kinase and a C-terminal phosphatase domain (Figure 1B). Similarly to the mammalian IP6K (Voglmaier et al., 1996), Arabidopsis ITPK1 does not only transfer phosphates to inositol polyphosphates but also acts as ATP synthase under varying ATP/ADP ratios or Pi concentrations (Figure 1B). The enzyme has a high KM of 520 μM for ATP and shifts its activity from kinase to ATP synthase at low adenylate energy charges by transferring the β-phosphate from 5-InsP7 to generate ATP from ADP (Riemer et al., 2021). In addition, ITPK1 exclusively uses 5-InsP7 and no other InsP7 isomer as a substrate for this ADP phosphotransferase activity in vitro, in agreement with a high substrate specificity (Riemer et al., 2021). Besides, ITPK1 was shown to act as an InsP(3,4,5,6)4 1-kinase/InsP5 [2-OH] 1-phosphotranferase to generate ATP from ADP in vitro (Whitfield et al., 2020).
Taken together, plant PP-InsP kinases catalyze both the generation and the removal of PP-InsPs. This raises the hypothesis that the enzymes might modulate energy reserves by shifting their activities, for instance, in response to environmental changes, such as phosphorus limitation or sufficiency (Saiardi et al., 1999; Riemer et al., 2021).
Discovery of new PP-InsP4 and InsP7 isomers in plants
The detection and quantification of plant PP-InsP species is challenging due to their low abundance, as well as the presence of high amounts of acid phosphatases in plant extracts, which leads to rapid degradation of PP-InsPs (Laha et al., 2021b). Until recently, Strong Anion Exchange High Performance Liquid Chromatography (SAX-HPLC) and Polyacrylamide Gel Electrophoresis (PAGE) were the most common methods used to analyze InsPs and PP-InsPs (Azevedo and Saiardi, 2006; Pisani et al., 2014). Owing to its easy set-up and low costs, PAGE is still widely employed to resolve higher inositol polyphosphates. The drawback of both of the above-mentioned methods is the inability to separate PP-InsP isomers. The first clarification of isomer identity of a particular PP-InsP species in plant tissue was possible via two-dimensional nuclear magnetic resonance spectroscopy (NMR), by taking advantage of an Arabidopsis mrp5 mutant (Laha et al., 2019). This mutant is defective in vacuolar loading of InsP6, leading to elevated PP-InsP cyto/nucleoplasmic levels (Nagy et al., 2009; Desai et al., 2014; Laha et al., 2019; Riemer et al., 2021). NMR analyses of mrp5 seed extracts and comparison with synthetic references demonstrated that 5-InsP7 is the major PP-InsP species present in mrp5 seeds (Laha et al., 2019).
Coupling of the two powerful tools “capillary electrophoresis” and “electrospray ionization mass spectrometry” (CE-ESI-MS) has enabled new insights into the abundance of InsP and PP-InsP isomers in mammalian cells, yeast, amoeba and plants (Qiu et al., 2020). Due to its high tolerance of complex sample matrices, the combined CE-ESI-MS enables separation of highly charged metabolites with compelling sensitivity. By employing this technique, the generation of 1/3-InsP7 and 5-InsP7 by VIH2 and ITPK1, respectively, was finally confirmed in planta (Riemer et al., 2021). Notably, this work also revealed for the first time the presence of 4/6-InsP7 in plants. In Arabidopsis, this InsP7 isomer was found to be more prominent than 5-InsP7 and 1/3-InsP7. However, in contrast to 1/3-InsP7 and 5-InsP7, the new isomer is less responsive to Pi deplete and replete conditions (Riemer et al., 2021) and the function(s) of 4/6-InsP7 and the potential kinase(s) that generate this new isomer in plants are still unknown.
Notably, not only InsP6 is converted to higher PP-InsPs, but also isomers of pentakisphosphates (InsP5) can serve as precursors for the generation of 5-diphosphoinositol tetrakisphosphate (5PP-InsP4) in yeast and mammalian cells (Wang et al., 2018). For instance, Saiardi et al. (2000) demonstrated that the yeast InsP6 kinase Kcs1 can generate PP-InsP4 from InsP5 [2-OH] in vitro. Interestingly, the affinity of Kcs1 for InsP5 was shown to be threefold higher (mean KM = 1.2 μM) than for InsP6 (mean KM = 3.3 μM). The mammalian IP6K1 also phosphorylates InsP5 [2-OH] to PP-InsP4, but in this case with similar affinities for InsP5 and InsP6 phosphorylation (Saiardi et al., 2000). While InsP5 levels in yeast are low and therefore probably do not represent the main Kcs1 substrate in vivo, this might be different in mammalian cells, where InsP5 [2-OH] and InsP6 levels are similar and represent physiologically relevant substrates of IP6K1 (Saiardi et al., 2000).
A recent study reported the identification of a novel PP-InsP4 isomer that does not co-migrate with a synthetic 5PP-InsP4 standard, suggesting a distinct structural identity as compared to PP- InsP4 isomers identified in yeast and mammalian cells (Riemer et al., 2021). CE-MS and PAGE data show that this plant PP-InsP4 isomer increases under Pi-starvation, as well as under Pi-resupply conditions, and is not detectable in nutrient-repleted plant roots. Interestingly, in roots of itpk1 loss-of-function mutants, this novel PP-InsP isomer seems to be less abundant, suggesting that ITPK1 might catalyze the generation of PP-InsP4 in planta (Riemer et al., 2021). Interestingly, ITPK1 catalyzes the generation of PP-InsP4 from InsP5 [6-OH] in vitro (Whitfield et al., 2020) but it remains to be shown whether plants possess the InsP5 [6-OH] isomer and whether the ITPK1-dependent PP-InsP4 derives from it.
The finding that other PP-InsP species than 1/3-InsP7 or 5-InsP7 were detected in plant extracts unveil an unexplored diversity of inositol pyrophosphates in plants. Also the involvement of putative unknown kinases responsible for the production of additional isomers in environmental responses still have to be investigated.
Inositol pyrophosphate kinases and their role in the adaption of plants to biotic and abiotic stress responses
Inositol pyrophosphate kinases are involved in salicylic acid-dependent immunity
The plant hormone salicylic acid (SA) regulates several processes like flower induction, stomatal closure and heat production mediated by alternative respiration in flowers (Rai et al., 1986; Raskin, 1992). Besides, SA is known to play a crucial role in defense mechanisms against bacteria, fungi, viruses and insects (Raskin et al., 1989; Chaerle et al., 1999; Martínez et al., 2004; Zarate et al., 2007; Kim and Hwang, 2014; Hao et al., 2018). Plant immune responses include the so-called PAMP-triggered immunity (PTI), characterized by the recognition of pathogen-associated molecular patterns (PAMPs, e.g., the bacterial peptide flagellin 22, or flg22), which triggers ion fluxes, ROS production and a series of signaling cascades that ultimately lead to local or systemic responses to restrict pathogen invasion (Seybold et al., 2014). Besides PTI, plants count on a second layer of protection, the effector triggered immunity (ETI), in which plants recognize effector proteins secreted by the pathogen. The ETI usually triggers fast defense reactions, such as hypersensitive response (HR), to promptly restrict pathogen colonization (Dodds and Rathjen, 2010).
Both PTI and ETI are modulated by SA, which is also key for the establishment of systemic acquired resistance (SAR), an additional layer of defense that protects plants from subsequent pathogen attacks (Hõrak, 2020). For instance, SA activates, via the regulatory protein NPR1, expression of several pathogenesis-related (PR) genes, which encode different types of proteins with antimicrobial properties (Van Loon et al., 2006; Hõrak, 2020).
A defined role of PP-InsPs in SA-signaling is still unclear. This is because studies showing an involvement of InsPs and PP-InsPs in SA-dependent immunity have in part contradictory outcomes. Arabidopsis mutants disrupted in InsP6 biosynthesis, for instance, showed increased susceptibility to bacterial, fungal and viral infections (Murphy et al., 2008; Poon et al., 2020), as well as to cyst nematode infestation (Jain, 2015). In fact, the Arabidopsis ips2 and ipk1 mutants defective in the activities of enzymes for the first and last steps in InsP6 biosynthesis, respectively, were similarly susceptible to microbial pathogens than NahG-transgenic lines and to sid2 mutants, both of which are unable to accumulate normal levels of SA (Murphy et al., 2008). The SA contents in ips2 and ipk1, however, did not differ from those of wild-type plants, and also increased, similarly to wild-type, after challenge with Pseudomonas syringae pv. tomato (Pst) DC3000 AvrB (Murphy et al., 2008). These results indicate that the enhanced susceptibility of ips2 and ipk1 is not related to low SA levels, but could be caused by the disruption of InsP6 biosynthesis (Murphy et al., 2008). Further studies of the ipk1 mutant and of loss-of-function mutants of another IPS isoform (IPS3) confirmed an involvement of InsP6 in basal pathogen responses (Poon et al., 2020). While displaying a higher susceptibility to Pst than wild-type plants, when the ips2, ips3, and ipk1 mutants were assessed for SAR acquirement, no impairment was detected. Besides, all mutants except ipk1 presented flg22-induced resistance to Pst, indicating that PTI was inhibited in ipk1 only (Poon et al., 2020). In this case, however, disruption of InsP6 synthesis in ipk1 did not affect typical responses to flg22, such as Ca2+ influx, oxidative burst, root growth inhibition and activation of PAMP-triggered genes. Taken together, these data suggest that InsP6 biosynthesis is important for maintaining basal resistance against various pathogens, contributing to defense mechanisms different from canonical PTI (Murphy et al., 2008; Poon et al., 2020).
In contrast to findings presented by those previous studies, a recent analysis of ipk1, itpk1, and vih2 mutants revealed that these enzymes act as negative regulators of SA-dependent immunity (Gulabani et al., 2021). Mutant plants, in which either InsP7 or InsP8 levels are impaired (Stevenson-Paulik et al., 2005; Sweetman et al., 2007; Desai et al., 2014; Laha et al., 2015; Kuo et al., 2018; Riemer et al., 2021), were significantly more resistant to bacterial infection by Pst in comparison to wild-type (Gulabani et al., 2021). Such a response was associated with an apparent constitutive activation of defenses observed in these plants. For instance, they showed a strong upregulation of SA biosynthesis genes, such as SID2/ICS1, and higher levels of free or glycolsyl moiety-conjugated SA (SAG) than wild-type plants. Along with these findings, an increase in the expression of PR1 and PR2, together with an accumulation of the respective proteins was observed. Also protein levels of ENHANCED DISEASE SUSCEPTIBILITY 1 (EDS1) and SUPPRESSOR OF nrp1-1 CONSTITUTIVE 1 (SNC1), both of which are required for basal defenses, were higher in these mutants than in wild-type plants, probably due to their elevated SA levels (Gulabani et al., 2021).
Although previous studies highlighted the importance of InsP6 in maintaining basal defenses against bacteria (Murphy et al., 2008; Ma et al., 2017; Poon et al., 2020), a set of mutants reduced in phytic acid levels, such as mik-1, ipk2β, or itpk4, displayed comparable PTI to wild-type (Gulabani et al., 2021). These findings suggest that InsP6 is not directly involved in triggering plant defenses but point toward a role of higher inositol pyrophosphates in regulating basal immunity (Gulabani et al., 2021).
Currently, it remains unclear whether InsP7 or InsP8 is the main player in SA-mediated defense. Both molecules might act indirectly by the regulation of an antagonistic crosstalk between auxin-SA and jasmonic-acid (JA)-SA, respectively. As described in details in section “As described in the section 5.5,” of this review, 5-InsP7 was proposed to regulate auxin signaling by acting as a co-ligand of the ASK1-TIR1-Aux/IAA auxin receptor complex (Parvin Laha et al., 2020), and exogenous application of auxin enhances the Pst infestation by interfering with SA-defenses (Navarro et al., 2006; Wang et al., 2007). Therefore, disruption of auxin signaling in itpk1 mutants might enhance basal immunity (Gulabani et al., 2021). On the other hand, the antagonism between JA-SA crosstalk in plants is well-described and even pathogens have the capability to secrete hormone-mimicking effectors to hijack host defense mechanisms (Zheng et al., 2012; Caarls et al., 2015). For instance, coronatine, a Pst-produced phytotoxin that mimics JA, triggers virulence by downregulating SA-dependent defenses in plants (Brooks et al., 2005; Zheng et al., 2012). Furthermore, several studies indicate that endogenous SA is antagonistic to JA-dependent defense mechanisms in plants, leading to a prioritized SA-driven resistance over JA-regulated defense (reviewed in Pieterse et al., 2012). Along these lines, ipk1 plants primed with injection of air or water to mimic wounding were less susceptible to Pst than corresponding wild-type plants that were primed in the same way (Poon et al., 2020). It remains unclear which ipk1-dependent inositol phosphate species might be responsible for this phenotype, as ipk1 mutants are defective in InsP6, InsP7, and InsP8 synthesis (Laha et al., 2015; Gulabani et al., 2021).
Taken together, elevated SA levels and expression of PTI-responsive genes in ipk1, itpk1 and vih2 might be related to disrupted JA signaling by low PP-InsP levels, causing enhanced SA-defense mechanisms (Gulabani et al., 2021). Further research is needed to unveil the involvement of specific PP-InsPs and other InsP species in regulating plant SA-dependent immunity.
The role of inositol pyrophosphates in jasmonate perception
Jasmonic acid and its derivates, collectively known as jasmonates (JA), play a crucial role in regulating plant development and defense against several necrotrophs and herbivores (Wasternack and Hause, 2013). In response to wounding or herbivory insects, the level of the bioactive JA conjugate jasmonic isoleucine (JA-Ile) is elevated (Fonseca et al., 2009; Koo et al., 2009), which in turn binds to the Coronatine Insensitive 1 (COI1) protein (Feys et al., 1994; Xie et al., 1998; Xu et al., 2002; Katsir et al., 2008), the F-box component of the SCF ubiquitin E3 ligase complex (Devoto et al., 2002). Binding of JA-Ile to COI1 recruits the Jasmonate ZIM Domain (JAZ) transcriptional repressor, which subsequently undergoes polyubiquitylation and SCFCOI1-mediated proteasome degradation (Chini et al., 2007; Thines et al., 2007; Yan et al., 2007). JAZ degradation then de-represses MYC2 and other transcription factors and consequently triggers the expression of JA-dependent genes (Boter et al., 2004; Browse, 2009).
Crystallization of the insect-purified auxin receptor TIR1/IAA complex that contained insect-derived InsP6 provided important information to better understand phytohormone-mediated signaling in plants (Tan et al., 2007). Nano-electrospray mass spectroscopy of the ASK1-COI1 complex that was purified from an insect cell line ectopically expressing the Arabidopsis ASK1-COI1 complex indeed revealed the existence of a molecule whose molar mass corresponded to InsP5 (Sheard et al., 2010). A multistep purification strategy followed by 1H NMR analysis and Total Correlation Spectroscopy (TOCSY) allowed to identify this ligand as either InsP5 [1-OH] or InsP5 [3-OH] (Sheard et al., 2010). In the crystal structure of the ASK1-COI1 complex, the presence of strong electron densities congregating in the core of the solenoid structure likely represents individual phosphates that replaced the insect-derived InsP5 ligand, probably due to high concentrations of ammonium phosphate, used as a precipitant during crystallization (Sheard et al., 2010). To further evaluate the functional role of InsPs in ASK1-COI1-JAZ1 co-receptor complex, a ligand-binding reconstitution assay revealed that both Ins(1,4,5,6)P4 and InsP5 [3-OH] can strongly induce ASK1-COI1-JAZ co-receptor complex formation in vitro, whereas InsP6 appeared to be less effective (Sheard et al., 2010). However, it is still unclear whether plants contain InsP5 [3-OH], its enantiomer InsP5 [1-OH] or both isomers.
Several studies pointed to an involvement of InsPs in wound response, as well as disease resistance in plants (Mosblech et al., 2008; Murphy et al., 2008). Arabidopsis plants heterologously expressing human inositol phosphate 5-phosphatase exhibit reduced levels of InsP3 (Perera et al., 2006; Hung et al., 2014) and are found to be susceptible to the cabbage moth Plutella xylostella (Mosblech et al., 2008). As previously mentioned, Murphy et al. (2008) also showed that Arabidopsis ipk1 and ics2 mutant plants with defects the in production of InsP6 are compromised in plant defense against bacterial (Pseudomonas syringae), viral (Tobacco Mosaic Virus), and necrotrophic fungal (Botrytis cinerea) pathogens.
To gain insights into the functional role of InsPs in JA signaling, mutant lines defective in putative inositol phosphate binding residues of COI1 were analyzed (Mosblech et al., 2011). Yeast two-hybrid (Y2H) analysis of the COI1 mutant variants revealed reduced interaction with JAZ proteins in presence of the JA analog coronatine, which suggests that InsP binding to the receptor complex might be important (Mosblech et al., 2011). Supporting this statement, Y2H studies using mutant lines defective in InsP biosynthesis revealed an enhanced COI1/JAZ interaction in the ipk1Δ yeast strain, which has high levels of InsP5 [2-OH] (Mosblech et al., 2011).
Although one cannot simply compare the yeast ipk1Δ mutant with Arabidopsis ipk1-1 lines, as yeast ipk1 strains show elevated levels of a specific PP-InsP4 isomer that cannot be detected in Arabidopsis (Saiardi et al., 2002; Laha et al., 2015; Riemer et al., 2021), these studies suggest a potential role of InsPs in regulating JA responses in plants. The enhanced interaction of COI1 and JAZ in ipk1Δ yeast strain could also be explained by the high levels of PP-InsP4. Additionally, Arabidopsis ipk1-1 plants are not only defective in InsP6 but are also severely compromised in InsP7 and InsP8 (Laha et al., 2015). To further evaluate the potential role of PP-InsPs in JA-dependent responses, VIH2-deficient plants defective in InsP8 synthesis were investigated (Laha et al., 2015). The mutant plants had unchanged levels of InsP5 [2-OH], but were shown to be severely susceptible to the generalist herbivore Mamestra brassicae and the Brassicaceae specialist Pieris rapae. This suggests that VIH2-dependent InsP8 but not InsP5 [2-OH] is critical for defense against these insects (Laha et al., 2015). In addition, vih2 mutants showed reduced expression of JA-dependent genes, despite an increase in JA. Therefore, the compromised resistance of Arabidopsis vih2 mutants against herbivory insects might be explained by a defect in JA perception, and not by compromised JA production (Laha et al., 2015). Additionally, in vitro binding assays of ASK1-COI1-JAZ1-coronatine with different radiolabeled InsPs indicated that higher inositol polyphosphates, such as InsP6 and InsP7, are capable to bind to the JA-receptor complex with higher efficiency than lower InsPs (Laha et al., 2015, 2016). Unfortunately, plant-purified InsP8 was not included in these binding assays due to its low amounts in plants and its high susceptibility to acid phosphatases present in plant extracts (Laha et al., 2015).
Taken together, it has been proposed that coincidence detection of VIH2-dependent InsP8 and JA is important for plant defense against necrotrophic and herbivorous pathogens (Laha et al., 2015, 2016). While these studies provide some mechanistic insights into the role of InsP8 in JA responses, future work is needed to clarify the molecular basis of VIH2 functions. For instance, it has not been established whether the catalytic activity of VIH2 solely contributes to JA responses, or whether VIH2 regulates JA responses through a yet unidentified mechanism. It might be also interesting to learn whether the phosphatase domain of VIH2 contributes to JA-related defense responses.
The studies presented here led to the assumption that plants are able to use different InsPs to cope against several pathogens and herbivores (Mosblech et al., 2011; Laha et al., 2015). However, whether InsPs allow plants to differentially respond against pathogens and how this process takes place remains an interesting question. The precise mechanism by which the JA co-receptor complex discriminates and specifically binds to a particular InsP isomer is still unclear. Furthermore, the physiological relevance of various InsP isomers in the context of the JA signaling pathway is still unsolved. It would be interesting to explore the possibility that different InsPs could form a series of distinctive JA-co-receptor complexes, which would help plants to induce specific immune responses against distinct pathogens.
The 1-phytase activity of Xanthomonas type III effector XopH
Several Gram-negative bacteria of the genus Xanthomonas cause diseases in different plant hosts, such as pepper, rice, wheat, tomato, citrus, cabbage, and banana, leading to substantial crop yield losses (Ryan et al., 2011; Jacques et al., 2016). A broad range of factors influence host specificity and pathogenicity. These include bacterial lipopolysaccharides, adhesins, transcription factors and TonB-dependent receptors, as well as the type III secretion system (T3SS) (Raetz and Whitfield, 2002; Ghosh, 2004; Blanvillain et al., 2007; Das et al., 2009; Büttner, 2016). The latter is responsible for the translocation of effector proteins into the plant cell cytosol (Büttner, 2016; Constantin et al., 2017; Newberry et al., 2019). The tomato and pepper pathogen Xanthomonas campestris pv. vesicatoria (Xcv) encodes more than 30 T3S effector proteins, which are generally designated as Xops (Xanthomonas outer proteins) and are known to cause characteristic bacterial spot disease symptoms (Thieme et al., 2005; Teper et al., 2015). In resistant plants, the effectors are recognized by immune receptors, often leading to HR at the infected area to suppress spreading of biotrophic pathogens from the site of infection (Goodman and Novacky, 1994; Mur et al., 2008). One member of the Xops effector family, XopH, depicts typical features of dual-specific protein phosphatases and can dephosphorylate the generic substrate p-nitrophenyl phosphate (pNPP) (Potnis et al., 2012).
Blüher et al. (2017) reported a novel phytate-degrading activity of XopH in vitro and in planta, which is assumed to account for the activation of HR in resistant plants. Using a novel NMR method coupled with spiking experiments, as well as biochemical studies with recombinant XopH, the authors identified XopH as a 1-phytase that cleaves the phosphate from the C1 hydroxy group of InsP6, resulting in the generation of InsP5 [1-OH] (Blüher et al., 2017). HPLC data of S. cerevisiae and N. benthamiana ectopically expressing XopH revealed a reduction of InsP6 and a strong accumulation of InsP5 [1/3-OH] also in vivo (Blüher et al., 2017). To confirm whether XopH executes 1-phytase activity in planta, the authors performed XopH digestion of InsP5 [1/3-OH] species purified from [3H]-myo-inositol-labeled transgenic N. benthamiana overexpressing xopH. The plant-purified InsP5 [1/3-OH] was resistant to XopH degradation and was not phosphorylated by plant enzymes, supporting the idea that this PP-InsP isomer is absent in plants and is more likely a product of XopH phytase activity (Blüher et al., 2017). Strikingly, the XopH-induced HR in pepper plants harboring the Bs7 resistance (R) gene seems to be dependent on the effector’s phytase activity. This led to the assumption that Bs7 more likely recognizes the result of XopH activity, such as changes in inositol polyphosphate levels, but not the protein itself (Blüher et al., 2017). It was also observed that heterologous expression of XopH in N. benthamiana resulted in a strong reduction of InsP7 and InsP8, presumably interfering with InsP7- and InsP8-dependent hormone signaling. In agreement with this, qRT-PCR analysis of N. benthamiana leaves constitutively expressing xopH showed an induction of the JA marker genes PR1b, PR4, and PI-II after wounding, strengthening the involvement of the effector protein in JA signaling (Blüher et al., 2017). Since those genes are also responsive to ethylene (ET), a hormone acting synergistically to JA, it cannot be excluded that XopH also affects the ET pathway. Indeed, virus-induced gene silencing of EIN2 and EBF1, which are the positive and negative regulators of the ET pathway, respectively, caused the suppression of xopH-induced upregulation of PR4 and PI-II in N. benthamiana (Donnell et al., 1996; Adie et al., 2007; Zhu and Lee, 2015; Blüher et al., 2017).
It still remains unclear for what purpose Xanthomonas secretes XopH into the host cells. One possibility is that the XopH phytase activity might release phosphate from the plant tissue, which could enhance the nutritional status of the pathogen. A similar activity was observed for the phytase PhyA, which is secreted by the rice pathogen X. oryzae pv. Oryzae. It was suggested that this bacterial pathogen uses phytate as the sole phosphate source and that this activity contributes to its virulence (Chatterjee et al., 2003). In addition, XopH might also degrade higher inositol pyrophosphates and thereby influence hormone signaling pathways of the host, leading to manipulation of JA- or ET-mediated defense responses to the pathogen’s benefit (Blüher et al., 2017).
Inositol pyrophosphates are involved in phosphate homeostasis
Phosphorus is an essential element and a key determinant for growth and development of all living organisms, as it composes essential molecules such as ATP, nucleic acids and phospholipids (Marschner, 1995). Plants take up phosphorus in the form of Pi, which is highly immobile, chemically fixated, as well as unevenly distributed in soils, causing a very limited access of available Pi (Holford, 1997; Seidel et al., 2021). Plants respond to low Pi levels by metabolic changes such as an increase of sulfo- and galactolipids at the expense of phospholipids (Essigmann et al., 1998; Härtel et al., 2000) and by increasing RNA degradation to release phosphate for other cellular processes (Taylor et al., 1993; Bariola et al., 1994). Furthermore, Pi-starved plants increase Pi acquisition via the production and secretion of phosphatases, exudation of organic acids, modification of root architecture and development of root hairs, as well as enhanced expression of Pi transporters (Karthikeyan et al., 2002; Mudge et al., 2002; Rausch and Bucher, 2002; Vance et al., 2003; Shin et al., 2004; Plaxton and Tran, 2011; Péret et al., 2011). These metabolic, morphological and transcriptional mechanisms belong to the so called phosphate starvation response (PSR), which is interrupted upon Pi replenishment (Vance et al., 2003; Chiou and Lin, 2011; Secco et al., 2013).
The majority of Pi starvation-induced (PSI) genes in plants is regulated by the MYB-CC transcription factor PHOSPHATE STARVATION RESPONSE REGULATOR 1 (PHR1) and its homolog PHR1-LIKE 1 (PHL1) (Rubio et al., 2001). PHR1 is expressed under Pi-sufficient conditions and controls Pi signaling and homeostasis through binding as a dimer to an imperfect palindromic sequence (PHR1-binding sequence, or P1BS) present in the promoters of Pi starvation-induced genes (Rubio et al., 2001).
Recent studies showed that a class of stand-alone SPX (SYG1/Pho81/XPR1-domain containing protein 1) proteins negatively regulates the activity of PHR transcription factors by high affinity binding to PHRs under sufficient Pi supply (Puga et al., 2014). The formation of the SPX-PHR complex in turn prevents the binding of the transcription factors to the P1BS motifs, thereby repressing the expression of PSI genes. Under low Pi conditions, the binding affinity of SPX to the PHRs is decreased, leading to the activation of their transcriptional targets (Puga et al., 2014; Qi et al., 2017).
Structural studies of SPX domains from proteins of different organisms indicate that PP-InsPs bind to SPX domains on a conserved cluster of basic residues and regulate the activity of such proteins, as shown for an SPX-containing component of the Vacuolar Transporter Chaperone (VTC) complex that mediates polyphosphate synthesis in baker’s yeast (Wild et al., 2016). Similar conserved clusters of basic residues at the surface of the SPX N-terminus were also identified in plant SPX proteins (Wild et al., 2016).
Recently, Dong et al. (2019) demonstrated that InsP8 binds to the rice OsSPX1 domain with a Kd of approximately 5.7 μM in vitro. In addition, Co-IP results revealed that SPX1 is not able to interact with PHR1 under Pi starvation conditions but can be restored by adding 1 μM InsP8 (Dong et al., 2019). On the other hand, the SPX-PHR interaction cannot be restored by the addition of InsP7, corroborating the idea that InsP8 but not InsP7 acts as the key regulator of Pi starvation responses in plants (Dong et al., 2019).
Several in vivo studies confirmed the involvement of PP-InsPs in PSR in plants. Arabidopsis mutants defective in IPK1 activity exhibit a disturbed phosphate starvation phenotype (Kuo et al., 2014). This results in an increased Pi overaccumulation when grown under Pi sufficient conditions and Pi accumulation in response to increasing external Pi concentrations (Stevenson-Paulik et al., 2005; Kuo et al., 2014, 2018). In addition, the mutants displayed reduced levels of InsP6, InsP7, and InsP8 (Laha et al., 2015; Kuo et al., 2018; Land et al., 2021).
Under Pi-replete conditions, the loss of ITPK1 but not of ITPK2 causes a robust overaccumulation of Pi similar to what was observed in ipk1 plants, even though only a decrease in 5-InsP7 and not in InsP8 was observed in itpk1 plants under such conditions (Riemer et al., 2021). In contrast, Pi-starved itpk1 plants that were resupplied with Pi displayed strong defects in both 5-InsP7 and InsP8 synthesis, again coinciding with a robust PSR phenotype (Riemer et al., 2021). An earlier study reported reduced InsP8 levels of itpk1 plants, as revealed by PAGE analyses also under Pi-replete conditions (Wang et al., 2021). The difference between the works of Riemer et al. (2021) and Wang et al. (2021) might be explained by different growth conditions employed by these two independent studies, including different Pi-availabilities at the Pi-replete condition.
While disruption of VIH1 and VIH2 did not cause any PSR phenotype, such as PSR gene expression and Pi-accumulation under Pi-replete conditions (Kuo et al., 2018; Land et al., 2021; Riemer et al., 2021), loss of VIH2 caused a mild PSR phenotype upon Pi-resupply to Pi-starved plants (Riemer et al., 2021). Importantly, vih1 vih2 double mutants (in which the respective kinase domains are defective) are seedling lethal (Zhu et al., 2019).
This is explained by the severe PSR phenotype of the double mutant seedlings caused by the strong overaccumulation of Pi, confirmed by the high expression of Pi starvation marker genes (Dong et al., 2019; Zhu et al., 2019). On the PP-InsP level, an increase in 5-InsP7 was observed in the double mutant, while InsP8 was below the limit of detection (Zhu et al., 2019). In contrast, HPLC profiles of a vih1 vih2 double mutant shown in Land et al. (2021) displayed reduced InsP7 and InsP8 levels. It is worth mentioning that the T-DNA insertion in this particular vih2 allele (vip1-2) is positioned outside the core VIH2 kinase domain. Taken together, the disruption of both VIH1 and VIH2 appears to result in the loss of the plant’s ability to maintain intracellular Pi levels due to defective InsP8 synthesis (Dong et al., 2019; Zhu et al., 2019). Notably, the itpk1 vih2 double mutant displays inhibited plant growth and an increase of approximately 27% in shoot P levels (Riemer et al., 2021). This strongly suggests that the combined activities of ITPK1 and VIH2 are critical for maintaining Pi homeostasis in plants, by concomitantly generating both the precursor (5-InsP7) as well as the main substrate (InsP8) of Pi sensing (Figure 2; Riemer et al., 2021). Lack of a PSR phenotype of ITPK4-deficient plants that display reduced levels of InsP6, InsP7, and InsP8 (Kuo et al., 2018; Wang et al., 2021) suggests that regulation of phosphate homeostasis by InsP and PP-InsPs might be even more complex. Future work should try to clarify the identities of InsP7 and InsP8 isomers by CE-ESI-MS analyses and, by taking advantage of chiral selectors, to address the question which PP-InsP species are altered. Besides, whether certain InsP7 or InsP8 isomers, or even enantiomers, play antagonistic roles in regulating the interaction of free standing SPX proteins with PHR1/PHL1 still needs clarification.

Figure 2. Model for the ITPK1-dependent phosphorylation of InsP6 and 5-InsP7 removal and possible link of ITPK1 with VIHs and phosphate homeostasis. Upon Pi-deficiency, ATP levels drop and stimulate ITPK1 to transfer the P-phosphate from 5-InsP7 to ADP, leading to the local generation of ATP and decreased 5-InsP7 levels. Additionally, low ATP/ADP ratios (i.e., low adenylate charge) and low Pi levels cause the switch from kinase to phosphatase activity of VIH proteins to hydrolyze InsP8. Lacking PP-InsPs, the interaction between PHR1 and SPX1 is destabilized, which promotes the binding of PHR1 to the P1BS motif in the promoter region of PSI genes. As a result, the Pi starvation response is activated. When plant cells regain sufficient Pi, ATP levels increase and the kinase activity of ITPK1 is activated, leading to the generation of 5-InsP7, which further serves as substrate for the kinase-activated VIH proteins to produce InsP8. Consequently, the accumulation of PP-InsPs facilitates the binding of SPX proteins to PHR1 to suppress Pi starvation responses.
Recent studies have pointed to a connection between plant’s Pi status and immune responses (Campos-Soriano et al., 2020; Val-Torregrosa et al., 2022). These findings are based on the involvement of a miRNA species (miR399) in the regulation of Pi homeostasis in Arabidopsis (Chiou et al., 2006; Paul et al., 2015). Upon Pi starvation, miR399 accumulates and represses its target gene PHOSPHATE2 (PHO2, encoding an E2 ubiquitin-conjugating enzyme) that is responsible for phosphate transporter degradation, leading to an enhanced Pi uptake in plants (Fujii et al., 2005; Kraft et al., 2005; Chiou et al., 2006; Liu et al., 2012; Huang et al., 2016). In rice, miR399 overexpression resulted in Pi accumulation in leaves and higher susceptibility to the fungal pathogen Magnaporthe oryzae, which was also observed upon high Pi fertilization (Campos-Soriano et al., 2020). In contrast, Val-Torregrosa et al. (2022) demonstrated an enhanced resistance to necrotrophic and hemibiotrophic fungal pathogens in Arabidopsis lines overexpressing miR399, as well as in pho2 loss-of-function lines. The high Pi accumulation in Arabidopsis leaves caused by miR399 overexpression and lack of functional PHO2 was linked to an elevated ROS production. This was assumed to be related to an increased HR in these plants during pathogen infection. Besides the changes in ROS levels, the mutant lines showed elevated SA and JA levels, combined with the upregulation of SA- and JA-dependent defense genes (Val-Torregrosa et al., 2022). Intriguingly, pho2 mutants were also shown to accumulate high levels of InsP8 (Riemer et al., 2021). As previously mentioned, InsP8 is a key player in JA- and Pi signaling (Laha et al., 2015, 2016; Dong et al., 2019; Riemer et al., 2021), raising the hypothesis that Pi homeostasis and pathogen defense mechanisms might be linked by the plant’s PP-InsP status.
Recently, Gulabani et al. (2021) demonstrated that the products of IPK1, ITPK1 and VIH2 kinase activities also function as crosstalk mediators between pathogen defense and Pi homeostasis, and that these enzymes act as suppressors of SA-dependent defense mechanisms. Strikingly, previous studies indicated the suppression of SA-responsive genes by PHR1 and as a consequence, phr1 phl1 double mutants appear to be more resistant to infections with PstDC3000 (Castrillo et al., 2017).
While PR1 and PR2 transcripts are upregulated in ipk1, itpk1, and vih2 lines, which are compromised in InsP8 levels or disrupted in functional PSR, the opposite was observed with the introduction of ipk1 and itpk1 into the phr1 phl1 mutant background (Gulabani et al., 2021). In this case, a reduced expression of both SA marker genes in comparison to Col-0 and phr1 phl1 was observed (Gulabani et al., 2021). The authors assumed that the downregulation of SA-associated defense genes in PP-InsP-compromised mutants is stimulated by a PHR1/PHL1-dependent increase in PSR. Furthermore, it is known that PSI genes might harbor SA-inducible elements in their promoters (Baek et al., 2017). Double mutants of the SA-biosynthesis gene SID2 in the ipk1 and itpk1 backgrounds, respectively, indeed resulted in decreased Pi overaccumulation phenotypes. Exogenous application of SA to wild-type plants led to increased transcripts of the PSI gene SPX1 or the PAMP-responsive gene WRKY38, both shown to be reduced in the ipk1 sid2 and itpk1 sid2 mutants, strengthening the hypothesis that SA may directly activate the transcription of PSI-genes (Gulabani et al., 2021). The phenotypes observed in ipk1, itpk1 and vih2 mutants are assumed to be related to the low InsP8 concentration that was observed at least in ipk1 and vih2 lines (Gulabani et al., 2021; Riemer et al., 2021), supporting a further putative link between PP-InsP-driven PSR and the capability to defend against pathogens.
To summarize, the connection of PSR and pathogen defense might give another perspective of how Pi is managed in crops. By having a deeper understanding of the factors affecting Pi homeostasis in plants, a more precise adjustment of fertilizer conditions may be employed in the field. This might help, for instance, to avoid strong pathogen infestation caused by excessive application of Pi, as well as to reduce environmental pollution and the depletion of global Pi-deposits, all of which will improve sustainability in crop production.
The role of inositol phosphates in auxin signaling
Auxin regulates a multitude of plant functions, including cell division, elongation, differentiation, embryonic development, root and stem tropisms, apical dominance, and flower formation (Young et al., 1990; Woodward and Bartel, 2005; Tanaka et al., 2006; Möller and Weijers, 2009; Leyser, 2010; Müller and Leyser, 2011; Christie and Murphy, 2013; Gallavotti, 2013; Geisler et al., 2014). This phytohormone coordinates those physiological processes by modulating the transcription of auxin-responsive genes through the action of the three protein families: TRANSPORT INHIBITOR RESPONSE1 (TIR1) and AUXIN- SIGNALING F-BOX proteins (AFB1-5), Aux/indole-3-acetic acid (IAA) transcriptional repressors, and the AUXIN RESPONSE FACTORS (ARFs) (Dharmasiri et al., 2005; Kepinski and Leyser, 2005; Tan et al., 2007). Auxin mediates their functions by binding to TIR1/AFB F-box proteins, enhancing the interaction of TIR1/AFB with Aux/IAAs repressors, which are in turn degraded by the Skp, Cullin, F-box-containing complex (SCF) ubiquitin ligase to activate ARF transcription factors (Wang and Estelle, 2014; Salehin et al., 2015). The Arabidopsis genome encodes 6 TIR1/AFBs, 29 Aux/IAA proteins, and 23 ARFs, which act combinatorically to regulate a wide range of auxin-dependent processes (Calderón Villalobos et al., 2012; Shimizu-Mitao and Kakimoto, 2014; Dinesh et al., 2016). The auxin co-receptors TIR1/AFB proteins comprise an F-box domain in the N terminus and 18 Leucine-rich repeat (LRR) domains at the C terminus (Tan et al., 2007). Structural analyses of the auxin co-receptor complexes purified from an insect cell line ectopically expressing Arabidopsis ASK1-TIR1 were instrumental in unveiling the molecular basis of auxin perception (Tan et al., 2007). The TIR1 crystal structure contained insect-derived InsP6 as a cofactor (Tan et al., 2007). InsP6 interacts with a highly basic surface area formed by 10 positively charged residues of TIR1, supporting the formation of the auxin binding pocket. These residues are also conserved in Arabidopsis AFBs, suggesting its binding importance in this subfamily of F-box proteins. When TIR1 is mutated in three residues that are involved in the coordination with InsP6, it fails to interact with either IAA7 or ASK1, implying a key role of InsP6 in the structural architecture of TIR1 (Calderón Villalobos et al., 2012). InsP6 interacts primarily with halves of the TIR1-LRR solenoids, loop-2, and the Arg403 residue. The Arg403 residue also binds to the carboxy group of auxin and is essential for the structural function of TIR1 (Calderón Villalobos et al., 2012). The authors demonstrated further that the mutation in His78, Arg403, and Ser438 residues of TIR1, which are involved in both auxin and InsP6 binding, failed to reconstitute the interaction between TIR1 and IAA7 in the presence of auxin (Calderón Villalobos et al., 2012). While these findings suggest an important function of InsP binding to TIR1, it needs to be investigated whether the designated InsP6 binding pocket of TIR1 can accommodate also other InsPs or is specific to InsP6. Even before InsP6 was identified as cofactor for the auxin receptor complex, inositol polyphosphates have been linked with several auxin-dependent physiological processes (Xu et al., 2005; Zhang et al., 2007). In Arabidopsis, two Inositol 1,4,5-Trisphosphate 3-Kinases (IPK2α and IPK2β) were found to harbor 6-/3-kinase activities and sequentially phosphorylate Ins(1,4,5)P3 to generate InsP5 [2-OH] via an Ins(1,3,4,6)P4 intermediate in vitro (Stevenson-Paulik et al., 2002; Xia et al., 2003). Expression analyses of IPK2α and IPK2β in different tissues of Arabidopsis plants pointed to a role of those kinases in plant growth and development (Xia et al., 2003; Xu et al., 2005). Silencing of IPK2α through antisense gene expression led to enhanced root growth and pollen germination in transgenic Arabidopsis plants (Xu et al., 2005), both of which are auxin-regulated processes (Fu and Harberd, 2003; Wu et al., 2008). Subsequent work investigating the physiological functions of IPK2β kinase uncovered that IPK2β is an early responsive gene that regulates axillary branching by an auxin signaling pathway. Furthermore, the application of exogenous IAA induced IPK2β expression and overexpressing of IPK2β results in altered auxin responses such as lateral root formation and primary root development (Zhang et al., 2007). Reverse Transcription-Polymerase chain reaction analysis (RT-PCR) of IPK2β overexpression lines revealed decreased expression of CYP83B1, a regulator of auxin production (Bartel et al., 2001; Woodward and Bartel, 2005; Zhang et al., 2007), and enhanced expression of PIN4, which mediates auxin transport (Friml, 2003; Zhang et al., 2007). Moreover, the expression levels of MAX4 and SPS, which are required for auxin-mediated shoot branching, was downregulated in IPK2β overexpression lines (Tantikanjana et al., 2001; Sorefan et al., 2003; Bainbridge et al., 2005; Zhang et al., 2007). Future work on auxin responses using ipk2β knockout lines will provide more insight into the IPK2β functions in auxin signaling. IPK2α and IPK2β are homologous genes with high sequence similarities (Stevenson-Paulik et al., 2002), and deletion of a single gene might not reveal its biological function due to redundancy. To date, the generation of ipk2α ipk2β double mutants was not successful because the homozygous double knockout appears to be lethal probably due to defects in pollen development, pollen tube guidance, and embryogenesis (Zhan et al., 2015). As such, the catalytic dead variants of IPK2β could not complement ipk2α ipk2β–associated lethality, suggesting an essential role of inositol polyphosphate signaling in plant reproduction (Zhan et al., 2015).
Other InsP kinases were found to be also involved in auxin-dependent physiological processes. Notably, transcriptome analysis of ipk1-1 plants showed that genes involved in root hair differentiation and root system development were misregulated in the mutant line. Moreover, ipk1-1 plants display a phenotype similar to the mrp5 mutant (Kuo et al., 2014). MRP5 encodes an ABC-type transporter mediating InsP6 loading into the vacuole (Nagy et al., 2009). In consequence, mrp5 mutant plants display reduced levels of InsP6, as well as elevated cytoplasmic levels of InsP7 and InsP8 (Desai et al., 2014; Laha et al., 2019), and exhibit a root system architecture (RSA) phenotype in response to elevated auxin (Gaedeke et al., 2001). Further, ipk1-1 plants having reduced levels of InsP6 also exhibit an altered RSA, which might be caused by compromised auxin signaling (Kuo et al., 2014). In line with this, the ipk1-1 mutant exhibited defects in gravitropic responses. Both ipk1-1 and mrp5 mutant plants were also insensitive to exogenous auxin supply, as evidenced by an increase in relative primary root length (Gaedeke et al., 2001; Laha et al., 2020). Taken together, these findings put forward the importance of IPK1 function in auxin signaling. The fact that the mrp5 mutant has elevated levels of InsP7 and InsP8 (Desai et al., 2014; Laha et al., 2019; Riemer et al., 2021), whereas ipk1-1 is severely compromised in those PP-InsP species (Laha et al., 2015), raise the possibility that the decreased levels of InsP6 or PP-InsP might contribute to auxin signaling. To further corroborate the role of PP-InsPs in auxin responses, the itpk1 and vih2 mutant lines were investigated. The itpk1 plants were shown to be defective in primary root elongation, leaf venation and compromised gravitropic root curvature, as well as thermomorphogenic adaptation, all of which are reminiscent of auxin deficient phenotypes (Laha et al., 2020). In auxin sensitivity assays, itpk1 plants displayed resistance to exogenous auxin, which could be fully rescued by itpk1 lines carrying a genomic ITPK1 fragment. This reinforces the idea that phenotypic defects of itpk1 mutant lines might be related to impaired auxin perception (Laha et al., 2020). ITPK1-deficient plants are defective not only in 5-InsP7 synthesis but are also perturbed in lower inositol phosphates homeostasis (Laha et al., 2019, 2020; Riemer et al., 2021), and their role in building auxin receptor complexes cannot be ignored (Laha et al., 2020). Specifically, HPLC profiles of both itpk1 and ipk1 mutants show reduced levels of InsP5 [1/3-OH], InsP7, and InsP8 and an increase in InsP4a, an unknown InsP4 isomer (Stevenson-Paulik et al., 2005; Laha et al., 2015).
Taken together, these results suggest that one or several inositol polyphosphate isomers might be important for auxin signaling. Future work is needed to clarify whether the control of auxin responses depends on the catalytic activity of ITPK1. Furthermore, the vih2 mutant lacking detectable InsP8 levels, as revealed by SAX-HPLC, did not exhibit auxin-related phenotypes, suggesting that InsP8 might not be critical for auxin responses (Laha et al., 2020). To further clarify the role of InsP8 in auxin signaling, future work on vih1 vih2 double knockout lines is necessary to account for a potential redundancy of the two VIH homologs.
Notably, competitive binding assays revealed that InsP6 and 5-InsP7 bind with similar affinities to the TIR1-ASK1-Aux/IAA7 auxin receptor complex (Laha et al., 2020). Considering the large amount of InsP6 present in plant cell extracts, an obvious question is how InsP7 (which comprises around 3% of global InsP6) could specifically control auxin perception. As mentioned earlier, several studies established that the major pool of InsP6 is stored in the vacuole (Nagy et al., 2009; Desai et al., 2014; Laha et al., 2019; Riemer et al., 2021), suggesting that the cyto/nucleo-plasmic concentration of InsP6 and InsP7 is distinct from the global cellular pool of InsP6 and InsP7. Investigating the localization of InsP6 and InsP7 at different compartments with the development of InsP6- and InsP7-specific sensors might clarify many of these open questions. Interestingly, a previous study reported that an InsP6 kinase interacts with certain protein complexes to generate InsP7 in close proximity to dedicated effector proteins (Rao et al., 2014). Similarly, recent work in Arabidopsis suggests that ITPK1 physically interacts with TIR1, presumably to channel 5-InsP7 to the auxin receptor complex (Laha et al., 2020). In addition, the potential of InsP molecules to induce a conformational change in TIR1 and promote the degradation of AUX/IAA is another conjecture to be solved. Knowing that different inositol phosphates have different affinities toward the auxin receptor complex is intriguing and raises the question whether these InsP molecules act differentially by forming distinct sets of auxin receptor complexes to regulate diverse auxin-related physiological processes. Altogether, many unsolved puzzles demand further research to identify the mechanism behind these phosphate-rich molecules playing a pivotal role in auxin-mediated plant growth and development.
In addition to the role of auxin in plant developmental and growth processes, several studies have also implicated a role of auxin in abiotic and biotic stresses (Cheong et al., 2002; Dowd et al., 2004; Hannah et al., 2005; Navarro et al., 2006; Wang et al., 2007; Jain and Khurana, 2009). The expression profiles of auxin-responsive genes of plants subjected to different biotic and abiotic stresses have pointed to a potential role of auxin in regulating plant defense responses, suggesting a possible crosstalk between auxin, abiotic and biotic stress signaling pathways (Ghanashyam and Jain, 2009). Recent findings revealed a potential role of auxin in regulating host-pathogen interaction. Auxin produced by different plant-associated microbes promotes disease susceptibility and antagonizes plant defense responses (Kunkel and Harper, 2018). Furthermore, Arabidopsis thaliana mutant lines defective in auxin signaling and perception showed increased levels of bacterial growth and suppressed host defenses, highlighting the role of auxin in biotic stress modulation (Djami-Tchatchou et al., 2020). Future work is needed to clarify whether inositol polyphosphates are involved in auxin-mediated pathogen defense responses.
Outlook
Here we pointed out the several roles InsPs and PP-InsPs play in regulating biotic and abiotic stress responses, and highlight these molecules as supporting modulators of plant metabolism to adapt to several environmental conditions (Figure 3). The recent development of more sensitive tools for the detection and quantification of low abundant PP-InsPs like CE-ESI-MS provides new insights into the large network of these molecules in eukaryotic systems (Qiu et al., 2020). However, the separation of enantiomeric PP-InsPs such as 1/3-InsP7 and 4/6-InsP7 still remains challenging with current chromatographic and electrophoretic methods. Future research needs to develop methods to distinguish between the mirror images to delineate which isomers are relevant in living plants. Besides the identification of the enantiomeric identity of these PP-InsP species, it will be a milestone to determine the responsible kinase for the newly identified 4/6-InsP7 and to determine the physiological processes this isomer regulates. We speculate that this might involve also responses to biotic stresses. Furthermore, the involvement of PP-InsPs in hormone signaling still remains enigmatic. Besides the role of these small molecules in auxin-, JA- and SA-dependent functions (Laha et al., 2015, 2020; Gulabani et al., 2021), a direct involvement in ethylene or brassinosteroid responses should be addressed, since a role of myo-inositol phosphate synthase in regulating plant growth and stress responses via ethylene- mediated signaling has been observed in Arabidopsis and wheat (Sharma et al., 2020a,b).
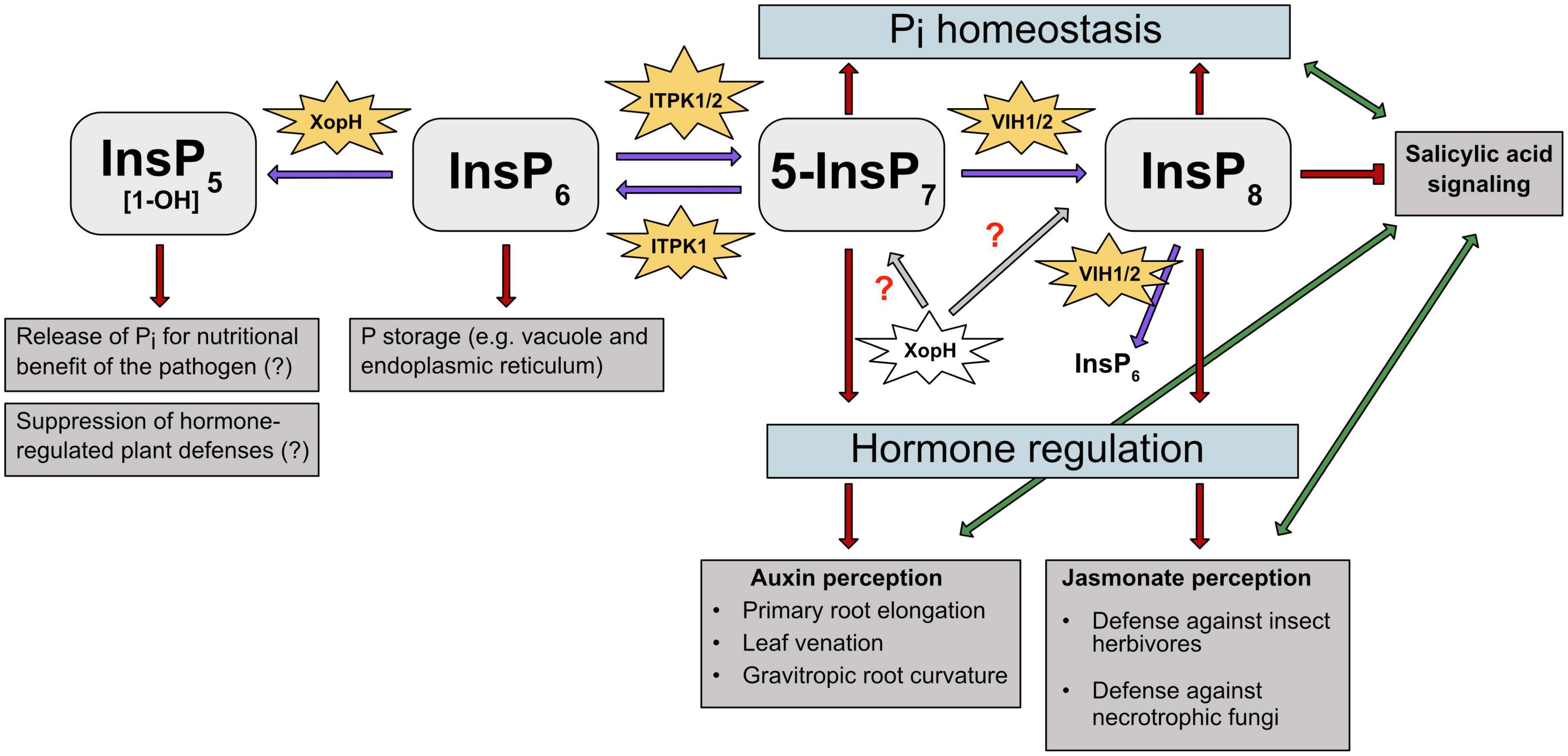
Figure 3. PP-InsPs and their kinases are involved in different abiotic and biotic stress responses in plants. PP-InsPs’ involvement in Pi homeostasis, hormone perception and regulation is depicted. Purple arrows indicate the kinase/phosphatase activity of the respective enzymes on InsPs and PP-InsPs. Gray arrows and red question marks depict a putative effect of XopH on PP-InsPs. Red arrows and T-shaped line indicate promotion and suppression of specific InsPs and PP-InsPs in regulating stress responses, respectively. Green arrows depict the interplay between plant hormones auxin, JA and SA, respectively. ITPK1 phosphorylates InsP6 to 5-InsP7. The latter serves as precursor for InsP8, which plays a crucial role in adaption to changing Pi levels. Additionally, InsP6 is degraded by XopH to potentially release Pi for the pathogen’s nutritional benefit. ITPK1 and VIH2 interaction is needed to maintain Pi homeostasis. Higher PP-InsPs are also involved in hormone perception and regulation. The ITPK1-generated 5-InsP7 is speculated to be involved in auxin and SA signaling. The VIH2- generated InsP8 has been proposed to represent a critical co-ligand of the JA receptor complex and is also assumed to regulate SA signaling. The bacterial type III effector XopH displays 1- phytase activity but may also have hydrolytic activities against PP-InsPs that might disrupt hormone-regulated defense mechanisms.
Finally, the identification of PP-InsPs and their different isomers will help to understand plant-pathogen interactions, which will be useful for improving crop growth and yield under abiotic and biotic stresses.
Author contributions
ER and DL conceived and prepared the outline of this review. ER, NP, RY, PR, HJ, MK, GS, and DL wrote the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was funded by grants from the Department of Biotechnology (DBT) for HGK-IYBA award (BT/13/IYBA/2020/04), SERB SRG/2021/000951, and Indian Institute of Science (IISc) start-up fund to DL. GS acknowledges funding from the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany’s Excellence Strategy, EXC-2070-390732324, PhenoRob and grants SCHA 1274/4-1, SCHA 1274/5-1). NP acknowledges CSIR for research fellowship. RY and PR were recipient of IISc research fellowship. HJ acknowledges funding from the German Research Foundation (DFG) under Germany’s Excellence Strategy (CIBSS – EXC-2189 – Project ID 390939984).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Adepoju, O., Williams, S. P., Craige, B., Cridland, C. A., Sharpe, A. K., Brown, A. M., et al. (2019). Inositol trisphosphate kinase and diphosphoinositol pentakisphosphate kinase enzymes constitute the inositol pyrophosphate synthesis pathway in plants. bioRxiv [Preprint] doi: 10.1101/724914
Adie, B., Rubio-Somoza, I., and Solano, R. (2007). Modulation of plant defenses by ethylene. J. Plant Growth Regul. 26, 160–177. doi: 10.1007/s00344-007-0012-6
Alimohammadi, M., de Silva, K., Ballu, C., Ali, N., and Khodakovskaya, M. V. (2012). Reduction of inositol (1,4,5)-trisphosphate affects the overall phosphoinositol pathway and leads to modifications in light signalling and secondary metabolism in tomato plants. J. Exp. Bot. 63, 825–835. doi: 10.1093/jxb/err306
Allen, G. J., and Sanders, D. (1994). Osmotic stress enhances the competence of Beta vulgaris vacuoles to respond to inositol 1,4,5-trisphosphate. Plant J. 6, 687–695. doi: 10.1046/j.1365-313X.1994.6050687.x
Azevedo, C., and Saiardi, A. (2006). Extraction and analysis of soluble inositol polyphosphates from yeast. Nat. Protoc. 1, 2416–2422. doi: 10.1038/nprot.2006.337
Baek, D., Chun, H. J., Yun, D. J., and Kim, M. C. (2017). Cross-talk between phosphate starvation and other environmental stress signaling pathways in plants. Mol. Cells 40, 697–705. doi: 10.14348/molcells.2017.0192
Bainbridge, K., Sorefan, K., Ward, S., and Leyser, O. (2005). Hormonally controlled expression of the Arabidopsis MAX4 shoot branching regulatory gene. Plant J. 44, 569–580. doi: 10.1111/j.1365-313X.2005.02548.x
Bariola, P. A., Howard, C. J., Taylor, C. B., Verburg, M. T., Jaglan, V. D., and Green, P. J. (1994). The Arabidopsis ribonuclease gene RNS1 is tightly controlled in response to phosphate limitation. Plant J. 6, 673–685. doi: 10.1046/j.1365-313x.1994.6050673.x
Bartel, B., LeClere, S., Magidin, M., and Zolman, B. K. (2001). Inputs to the active indole-3-acetic acid pool: De novo synthesis, conjugate hydrolysis, and indole-3-butyric acid β-oxidation. J. Plant Growth Regul. 20, 198–216. doi: 10.1007/s003440010025
Blanvillain, S., Meyer, D., Boulanger, A., Lautier, M., Guynet, C., Denancé, N., et al. (2007). Plant carbohydrate scavenging through tonB-dependent receptors: A feature shared by phytopathogenic and aquatic bacteria. PLoS One 2:e224. doi: 10.1371/journal.pone.0000224
Blatt, M., Thiel, G., and Trentham, D. (1990). Reversible inactivation of K+ channels of Vcia stomatal guard cells following the photolysis of caged inositol 1,4,5-trisphosphate. Nature 346, 766–769. doi: 10.1038/346766a0
Blüher, D., Laha, D., Thieme, S., Hofer, A., Eschen-Lippold, L., Masch, A., et al. (2017). A 1-phytase type III effector interferes with plant hormone signaling. Nat Commun. 8:2159. doi: 10.1038/s41467-017-02195-8
Boter, M., Ruı’z-Rivero, O., Abdeen, A., and Prat, S. (2004). Conserved MYC transcription factors play a key role in jasmonate signaling both in tomato and Arabidopsis. Genes Dev. 18, 1577–1591. doi: 10.1101/gad.297704
Brooks, D. M., Bender, C. L., and Kunkel, B. N. (2005). The Pseudomonas syringae phytotoxin coronatine promotes virulence by overcoming salicylic acid-dependent defences in Arabidopsis thaliana. Mol. Plant Pathol. 6, 629–639. doi: 10.1111/j.1364-3703.2005.00311.x
Browse, J. (2009). Jasmonate passes muster: A receptor and targets for the defense hormone. Annu. Rev. Plant Biol. 60, 183–205. doi: 10.1146/annurev.arplant.043008.092007
Büttner, D. (2016). Behind the lines-actions of bacterial type III effector proteins in plant cells. FEMS Microbiol. Rev. 40, 894–937. doi: 10.1093/femsre/fuw026
Caarls, L., Pieterse, C. M., and Van Wees, S. C. (2015). How salicylic acid takes transcriptional control over jasmonic signaling. Front. Plant Sci. 6:170. doi: 10.3389/fpls.2015.00170
Calderón Villalobos, L. I., Lee, S., De Oliveira, C., Ivetac, A., Brandt, W., Armitage, L., et al. (2012). A combinatorial TIR1/AFB–Aux/IAA co-receptor system for differential sensing of auxin. Nat. Chem. Biol. 8, 477–485. doi: 10.1038/nchembio.926
Campos-Soriano, L., Bundó, M., Bach-Pages, M., Chiang, S. F., Chiou, T. J., and San Segundo, B. (2020). Phosphate excess increases susceptibility to pathogen infection in rice. Mol. Plant Pathol. 21, 555–570. doi: 10.1111/mpp.12916
Castrillo, G., Teixeira, P. J., Paredes, S. H., Law, T. F., de Lorenzo, L., Feltcher, M. E., et al. (2017). Root microbiota drive direct integration of phosphate stress and immunity. Nature 543, 513–518. doi: 10.1038/nature21417
Chaerle, L., Van Caeneghem, W., Messens, E., Lambers, H., Van Montagu, M., and Van Der Straeten, D. (1999). Presymptomatic visualization of plant-virus interactions by thermography. Nat. Biotechnol. 17, 813–816. doi: 10.1038/11765
Chatterjee, S., Sankaranarayanan, R., and Sonti, R. V. (2003). PhyA, a secreted protein of Xanthomonas oryzae pv. oryzae, is required for optimum virulence and growth on phytic acid as a sole phosphate source. Mol. Plant Microbe Interact. 16, 973–982. doi: 10.1094/MPMI.2003.16.11.973
Chen, X., Lin, W., Wang, Y., Luan, S., and Xue, H. W. (2008). An inositol polyphosphate 5-phosphatase functions in PHOTOTROPIN1 signaling in Arabidopis by altering cytosolic Ca2+. Plant Cell 20, 353–366. doi: 10.1105/tpc.107.052670
Cheong, Y. H., Chang, H. S., Gupta, R., Wang, X., Zhu, T., and Luan, S. (2002). Transcriptional profiling reveals novel interactions between wounding, pathogen, abiotic stress, and hormonal responses in Arabidopsis. Plant Physiol. 129, 661–677. doi: 10.1104/pp.002857
Chini, A., Fonseca, S., Fernandez, G., Adie, B., Chico, J., Lorenzo, O., et al. (2007). The JAZ family of repressors is the missing link in jasmonate signalling. Nature 448, 666–671. doi: 10.1038/nature06006
Chiou, T. J., and Lin, S. I. (2011). Signaling network in sensing phosphate availability in plants. Annu. Rev. Plant Biol. 62, 185–206. doi: 10.1146/annurev-arplant-042110-103849
Chiou, T. J., Aung, K., Lin, S. I., Wu, C. C., Chiang, S. F., and Su, C. L. (2006). Regulation of phosphate homeostasis by microRNA in Arabidopsis. Plant Cell 18, 412–421. doi: 10.1105/tpc.105.038943
Christie, J. M., and Murphy, A. S. (2013). Shoot phototropism in higher plants: New light through old concepts. Am. J. Bot. 100, 35–46. doi: 10.3732/ajb.1200340
Constantin, E. C., Haegeman, A., Van Vaerenbergh, J., Baeyen, S., Van Malderghem, C., Maes, M., et al. (2017). Pathogenicity and virulence gene content of Xanthomonas strains infecting Araceae, formerly known as Xanthomonas axonopodis pv. dieffenbachiae. Plant Pathol. 66, 1539–1554. doi: 10.1111/ppa.12694
Das, A., Rangaraj, N., and Sonti, R. V. (2009). Multiple adhesin-like functions of Xanthomonas oryzae pv. oryzae are involved in promoting leaf attachment, entry, and virulence on rice. Mol. Plant Microbe Interact. 22, 73–85. doi: 10.1094/MPMI-22-1-0073
Desai, M., Rangarajan, P., Donahue, J. L., Williams, S. P., Land, E. S., Mandal, M. K., et al. (2014). Two inositol hexakisphosphate kinases drive inositol pyrophosphate synthesis in plants. Plant J. 80, 642–653. doi: 10.1111/tpj.12669
Devoto, A., Nieto-Rostro, M., Xie, D., Ellis, C., Harmston, R., Patrick, E., et al. (2002). COI1 links jasmonate signalling and fertility to the SCF ubiquitin–ligase complex in Arabidopsis. Plant J. 32, 457–466. doi: 10.1046/j.1365-313x.2002.01432.x
Dharmasiri, N., Dharmasiri, S., and Estelle, M. (2005). The F-box protein TIR1 is an auxin receptor. Nature 435, 441–445. doi: 10.1038/nature03543
Dinesh, D. C., Villalobos, L. I. A. C., and Abel, S. (2016). Structural biology of nuclear auxin action. Trends Plant Sci. 21, 302–316. doi: 10.1016/j.tplants.2015.10.019
Djami-Tchatchou, A. T., Harrison, G. A., Harper, C. P., Wang, R., Prigge, M. J., and Estelle, M. (2020). Dual role of auxin in regulating plant defense and bacterial virulence gene expression during Pseudomonas syringae PtoDC3000 pathogenesis. Mol. Plant Microbe Interact. 33, 1059–1071. doi: 10.1094/MPMI-02-20-0047-R
Dodds, P. N., and Rathjen, J. P. (2010). Plant immunity: Towards an integrated view of plant–pathogen interactions. Nat. Rev. Genet. 11, 539–548. doi: 10.1038/nrg2812
Dong, J., Ma, G., Sui, L., Wei, M., Satheesh, V., Zhang, R., et al. (2019). Inositol pyrophosphate InsP8 acts as an intracellular phosphate signal in Arabidopsis. Mol. Plant 12, 1463–1473. doi: 10.1016/j.molp.2019.08.002
Donnell, P., Calvert, C., Atzorn, R., and Wasternack, C. (1996). Ethylene as a signal mediating the wound response of tomato plants. Science 274, 1914–1917. doi: 10.1126/science.274.5294.1914
Dowd, C., Wilson, I. W., and McFadden, H. (2004). Gene expression profile changes in cotton root and hypocotyl tissues in response to infection with Fusarium oxysporum f. sp. vasinfectum. Mol. Plant Microbe Interact. 17, 654–667. doi: 10.1094/MPMI.2004.17.6.654
Draskovic, P., Saiardi, A., Bhandari, R., Burton, A., Ilc, G., and Kovacevic, M. (2008). Inositol hexakisphosphate kinase products contain diphosphate and triphosphate groups. Chem. Biol. 15, 274–286. doi: 10.1016/j.chembiol.2008.01.011
Essigmann, B., Gueler, S., Narang, R. A., Linke, D., and Benning, C. (1998). Phosphate availability affects the thylakoid lipid composition and the expression of SQD1, a gene required for sulfolipid biosynthesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. U.S.A. 95, 1950–1955. doi: 10.1073/pnas.95.4.1950
Feys, B., Benedetti, C. E., Penfold, C. N., and Turner, J. G. (1994). Arabidopsis mutants selected for resistance to the phytotoxin coronatine are male sterile, insensitive to methyl jasmonate, and resistant to a bacterial pathogen. Plant Cell 6, 751–759. doi: 10.1105/tpc.6.5.751
Fonseca, S., Chini, A., Hamberg, M., Adie, B., Porzel, A., Kramell, R., et al. (2009). (+)-7-iso-Jasmonoyl-Lisoleucine is the endogenous bioactive jasmonate. Nat. Chem. Biol. 5, 344–350. doi: 10.1038/nchembio.161
Fridy, P. C., Otto, J. C., Dollins, D. E., and York, J. D. (2007). Cloning and characterization of two human VIP1-like inositol hexakisphosphate and diphosphoinositol pentakisphosphate kinases. J. Biol. Chem. 292, 30754–30762. doi: 10.1074/jbc.M704656200
Friml, J. (2003). Auxin transport—shaping the plant. Curr. Opin. Plant Biol. 6, 7–12. doi: 10.1016/S1369526602000031
Fu, X., and Harberd, N. P. (2003). Auxin promotes Arabidopsis root growth by modulating gibberellin response. Nature 421, 740–743. doi: 10.1038/nature01387
Fujii, H., Chiou, T.-J., Lin, S.-I., Aung, K., and Zhu, J.-K. (2005). A miRNA involved in phosphate-starvation response in Arabidopsis. Curr. Biol. 15, 2038–2043. doi: 10.1016/j.cub.2005.10.016
Gaedeke, N., Klein, M., Kolukisaoglu, U., Forestier, C., Müller, A., Ansorge, M., et al. (2001). The Arabidopsis thaliana ABC transporter AtMRP5 controls root development and stomata movement. EMBO J. 20, 1875–1887. doi: 10.1093/emboj/20.8.1875
Gallavotti, A. (2013). The role of auxin in shaping shoot architecture. J. Exp. Bot. 64, 2593–2608. doi: 10.1093/jxb/ert141
Geisler, M., Wang, B., and Zhu, J. (2014). Auxin transport during root gravitropism: Transporters and techniques. Plant Biol. 16, 50–57. doi: 10.1111/plb.12030
Ghanashyam, C., and Jain, M. (2009). Role of auxin-responsive genes in biotic stress responses. Plant Signal. Behav. 4, 846–848. doi: 10.4161/psb.4.9.9376
Ghosh, P. (2004). Process of protein transport by the type III secretion system. Microbiol. Mol. Biol. Rev. 68, 771–795. doi: 10.1128/MMBR.68.4.771-795.2004
Gilroy, S., Read, N. D., and Trewavas, A. J. (1990). Elevation of cytoplasmic calcium by caged calcium or caged inositol trisphosphate initiates stomatal closure. Nature 346, 769–771. doi: 10.1038/346769a0
Goodman, R. N., and Novacky, A. J. (1994). The Hypersensitive Reaction in Plants to Pathogens: A Resistance Phenomenon. Chicago: American Phytopathological Society.
Gu, C., Nguyen, H.-N., Hofer, A., Jessen, H. J., Dai, X., Wang, H., et al. (2017). The significance of the bifunctional kinase/phosphatase activities of diphosphoinositol pentakisphosphate kinases (PPIP5Ks) for coupling inositol pyrophosphate cell signaling to cellular phosphate homeostasis. J. Biol. Chem. 292, 4544–4555. doi: 10.1074/jbc.M116.765743
Gulabani, H., Goswami, K., Walia, Y., Roy, A., Jameeta Noor, J., Ingole, K. D., et al. (2021). Arabidopsis inositol polyphosphate kinases IPK1 and ITPK1 modulate crosstalks between SA-dependent immunity and phosphate-starvation responses. Plant Cell Rep. 41, 347–363. doi: 10.1007/s00299-021-02812-3
Halsted, J. A., Ronaghy, H. A., Abadi, P., Haghshenass, M., Amirhakemi, G. H., Barakat, R. M., et al. (1972). Zinc deficiency in man. The Shiraz experiment. Am. J. Med. 53, 277–284. doi: 10.1016/0002-9343(72):90169-90166
Hannah, M. A., Heyer, A. G., and Hincha, D. K. (2005). A global survey of gene regulation during cold acclimation in Arabidopsis thaliana. PLoS Genet. 1:e26. doi: 10.1371/journal.pgen.0010026
Hao, Q., Wang, W., Han, Q., Wu, J., Lyo, B., Chen, F., et al. (2018). Isochorismate-based salicylic acid biosynthesis confers basal resistance to Fusarium graminearum in barley. Mol. Plant Pathol. 19, 1995–2010. doi: 10.1111/mpp.12675
Härtel, H., Dörmann, P., and Benning, C. (2000). DGD1-independent biosynthesis of extraplastidic galactolipids after phosphate deprivation in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 97, 10649–10654. doi: 10.1073/pnas.180320497
Holford, I. C. R. (1997). Soil phosphorus: Its measurement, and its uptake by plants. Aust. J. Soil Res. 35, 227–239.
Hõrak, H. (2020). Zones of defense? SA receptors have it under control. Plant Cell 32, 3658–3659. doi: 10.1105/tpc.20.00848
Huang, J., Yang, M., Lu, L., and Zhang, X. (2016). Diverse functions of smallRNAs in different plant-pathogen communications. Front. Microbiol. 4:1552. doi: 10.3389/fmicb.2016.01552
Hung, C. Y., Aspesi, P. Jr., Hunter, M. R., Lomax, A. W., and Perera, I. Y. (2014). Phosphoinositide-signaling is one component of a robust plant defense response. Front. Plant Sci. 5:267. doi: 10.3389/fpls.2014.00267
Irvine, R. F. (2003). 20 years of Ins(1,4,5)P3 and 40 years before. Nat. Rev. Mol. Cell Biol. 4, 586–590. doi: 10.1038/nrm1152
Jacques, M.-A., Arlat, M., Boulanger, A., Boureau, T., Carrère, S., Cesbron, S., et al. (2016). Using ecology, physiology, and genomics to understand host specificity in Xanthomonas. Ann. Rev. Phytopathol. 54, 163–187. doi: 10.1146/annurev-phyto-080615-100147
Jadav, R. S., Chanduri, M. V., Sengupta, S., and Bhandari, R. (2013). Inositol pyrophosphate synthesis by inositol hexakisphosphate kinase 1 is required for homologous recombination repair. J. Biol. Chem. 288, 3312–3321. doi: 10.1074/jbc.M112.396556
Jain, M., and Khurana, J. P. (2009). Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J. 276, 3148–3162. doi: 10.1111/j.1742-4658.2009.07033.x
Jain, R. (2015). Reduction of phytate by down-regulation of Arabidopsis thaliana MIPS and IPK1 genes alters susceptibility to beet cyst nematodes. Nematology 17, 401–407. doi: 10.1163/15685411-00002874
Josefsen, L., Bohn, L., Sørensen, M. B., and Rasmussen, S. K. (2007). Characterization of a multifunctional inositol phosphate kinase from rice and barley belonging to the ATP-grasp superfamily. Gene 397, 114–125. doi: 10.1016/j.gene.2007.04.018
Jung, J.-Y., Ried, M. K., Hothorn, M., and Poirie, R. Y. (2018). Control of plant phosphate homeostasis by inositol pyrophosphates and the SPX domain. Curr. Opin. Biotechnol. 49, 156–162. doi: 10.1016/j.copbio.2017.08.012
Karthikeyan, A. S., Varadarajan, D. K., Mukatira, U. T., D’Urzo, M. P., Damsz, B., and Raghothama, K. G. (2002). Regulated expression of Arabidopsis phosphate transporters. Plant Physiol. 130, 221–233. doi: 10.1104/pp.020007
Katsir, L., Schilmiller, A. L., Staswick, P. E., He, S. Y., and Howe, G. A. (2008). COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. Proc. Natl. Acad. Sci. U.S.A. 105, 7100–7105. doi: 10.1073/pnas.0802332105
Kepinski, S., and Leyser, O. (2005). The Arabidopsis F-box protein TIR1 is an auxin receptor. Nature 435, 446–451.
Khodakovskaya, M., Sword, C., Wu, Q., Perera, I. Y., Boss, W. F., Brown, C. S., et al. (2010). Increasing inositol (1,4,5)-trisphosphate metabolism affects drought tolerance, carbohydrate metabolism and phosphate-sensitive biomass increases in tomato. Plant Biotechnol. J. 8, 170–183. doi: 10.1111/j.1467-7652.2009.00472.x
Kim, D. S., and Hwang, B. K. (2014). An important role of pepper phenylalanine ammonia-lyase gene (PAL1) in salicylic acid-dependent signalling of the defence response to microbial pathogens. J. Exp. Bot. 65, 2295–2306. doi: 10.1093/jxb/eru109
Knight, H., Trewavas, A. J., and Knight, M. R. (1997). Calcium signalling in Arabidopsis thaliana responding to drought and salinity. Plant J. 12, 1067–1078. doi: 10.1046/j.1365-313x.1997.12051067.x
Koo, A. J., Gao, X., Jones, A. D., and Howe, G. A. (2009). A rapid wound signal activates the systemic synthesis of bioactive jasmonates in Arabidopsis. Plant J. 59, 974–986. doi: 10.1111/j.1365-313X.2009.03924.x
Kraft, E., Stone, S. L., Ma, L., Su, N., Gao, Y., Lau, O. S., et al. (2005). Genome analysis and functional characterization of the E2 and RING-type E3 ligase ubiquitination enzymes of Arabidopsis. Plant Physiol. 139, 1597–1611. doi: 10.1104/pp.105.067983
Krinke, O., Novotná, Z., Valentová, O., and Martinec, J. (2007). Inositol trisphosphate receptor in higher plants: is it real? J. Exp. Bot. 58, 361–376. doi: 10.1093/jxb/erl220
Kunkel, B. N., and Harper, C. P. (2018). The roles of auxin during interactions between bacterial plant pathogens and their hosts. J. Exp. Bot. 69, 245–254. doi: 10.1093/jxb/erx447
Kuo, H. F., Chang, T. Y., Chiang, S. F., Wang, W. D., Charng, Y. Y., and Chiou, T. J. (2014). Arabidopsis inositol pentakisphosphate 2-kinase, AtIPK1, is required for growth and modulates phosphate homeostasis at the transcriptional level. Plant J. 80, 503–515. doi: 10.1111/tpj.12650
Kuo, H. F., Hsu, Y. Y., Lin, W. C., Chen, K. Y., Munnik, T., Brearley, C. A., et al. (2018). Arabidopsis inositol phosphate kinases, IPK1 and ITPK1, constitute a metabolic pathway in maintaining phosphate homeostasis. Plant J. 95, 613–630. doi: 10.1101/270355
Laha, D., Johnen, P., Azevedo, C., Dynowski, M., Weiss, M., Capolicchio, S., et al. (2015). VIH2 regulates the synthesis of inositol pyrophosphate InsP(8) and jasmonate-dependent defenses in Arabidopsis. Plant Cell 27, 1082–1097. doi: 10.1105/tpc.114.135160
Laha, D., Kamleitner, M., Johnen, P., and Schaaf, G. (2021b). “Analyses of inositol phosphates and phosphoinositides by Strong Anion Exchange (SAX)-HPLC,” in Plant Lipids. Methods in Molecular Biology, Vol. 2295, eds D. Bartels and P. Dörmann (New York, NY: Humana), doi: 10.1007/978-1-0716-1362-7_20
Laha, D., Parvin, N., Dynowski, M., Johnen, P., Mao, H., Bitters, S. T., et al. (2016). Inositol polyphosphate binding specificity of the jasmonate receptor complex. Plant Physiol. 171, 2364–2370. doi: 10.1104/pp.16.00694
Laha, D., Parvin, N., Hofer, A., Giehl, R. F. H., Fernandez-Rebollo, N., von Wiren, N., et al. (2019). Arabidopsis ITPK1 and ITPK2 have an evolutionarily conserved phytic acid kinase activity. ACS Chem. Biol. 14, 2127–2133. doi: 10.1021/acschembio.9b00423
Laha, D., Portela-Torres, P., Desfougères, Y., and Saiardi, A. (2021a). Inositol phosphate kinases in the eukaryote landscape. Adv. Biol. Regul. 79:100782. doi: 10.1016/j.jbior.2020.100782
Laha, N. P., Dhir, Y. W., Giehl, R. F. H., Schaefer, E. M., Gaugler, P., Shishavan, Z. H., et al. (2020). ITPK1-dependent inositol polyphosphates regulate auxin responses in Arabidopsis thaliana. bioRxiv [Preprint].
Land, E. S., Cridland, C. A., Craige, B., Dye, A., Hildreth, S. B., Helm, R. F., et al. (2021). A role for inositol pyrophosphates in the metabolic adaptations to low phosphate in Arabidopsis. Metabolites 11:601. doi: 10.3390/metabo11090601
Leyser, O. (2010). The power of auxin in plants. Plant Physiol. 154, 501–505. doi: 10.1104/pp.110.161323
Lin, H., Fridy, P. C., Ribeiro, A. A., Choi, J. H., Barma, D. K., Vogel, G., et al. (2009). Structural analysis and detection of biological inositol pyrophosphates reveal that the family of VIP/diphosphoinositol pentakisphosphate kinases are 1/3-kinases. J. Biol. Chem. 284, 1863–1872. doi: 10.1074/jbc.M805686200
Liu, H. T., Gao, F., Cui, S. J., Han, J. L., Sun, D. Y., and Zhou, R. G. (2006). Primary evidence for involvement of IP3 in heat-shock signal transduction in Arabidopsis. Cell Res. 16, 394–400. doi: 10.1038/sj.cr.7310051
Liu, T. Y., Huang, T. K., Tseng, C. Y., Lai, Y. S., Lin, S. I., Lin, W. Y., et al. (2012). PHO2-dependent degradation of PHO1 modulates phosphate homeostasis in Arabidopsis. Plant Cell 24, 2168–2183. doi: 10.1105/tpc.112.096636
Loewus, F. A., and Murthy, P. P. N. (2000). Myo-Inositol metabolism in plants. Plant Sci. 150, 1–19. doi: 10.1016/S0168-9452(99)00150-8
Ma, Y., Zhao, Y., and Berkowitz, G. A. (2017). Intracellular Ca2+ is important for flagellin-triggered defense in Arabidopsis and involves inositol polyphosphate signaling. J. Exp. Bot. 68, 3617–3628. doi: 10.1093/jxb/erx176
Martínez, C., Pons, E., Prats, G., and León, J. (2004). Salicylic acid regulates flowering time and links defence responses and reproductive development. Plant J. 37, 209–217. doi: 10.1046/j.1365-313x.2003.01954.x
McCance, R. A., and Widdowson, E. M. (1942). Mineral metabolism of healthy adults on white and brown bread dietaries. J. Physiol. 101, 44–85. doi: 10.1113/jphysiol.1942.sp003967
Miller, G. J., Wilson, M. P., Majerus, P. W., and Hurley, J. H. (2005). Specificity determinants in inositol polyphosphate synthesis: Crystal structure of inositol 1,3,4-trisphosphate 5/6-kinase. Mol. Cell 18, 201–212. doi: 10.1016/j.molcel.2005.03.016
Möller, B., and Weijers, D. (2009). Auxin control of embryo patterning. Cold Spring Harb. Perspect. Biol. 1:a001545. doi: 10.1101/cshperspect.a001545
Monteiro, D., Liu, Q., Lisboa, S., Scherer, G. E. F., Quader, H., and Malhó, R. (2005). Phosphoinositides and phosphatidic acid regulate pollen tube growth and reorientation through modulation of [Ca2+]c and membrane secretion. J. Exp. Bot. 56, 1665–1674. doi: 10.1093/jxb/eri163
Mosblech, A., König, S., Stenzel, I., Grzeganek, P., Feussner, I., and Heilmann, I. (2008). Phosphoinositide and inositolpolyphosphate signalling in defense responses of Arabidopsis thaliana challenged by mechanical wounding. Mol. Plant 1, 249–261. doi: 10.1093/mp/ssm028
Mosblech, A., Thurow, C., Gatz, C., Feussner, I., and Heilmann, I. (2011). Jasmonic acid perception by COI1 involves inositol polyphosphates in Arabidopsis thaliana. Plant J. 65, 949–957. doi: 10.1111/j.1365-313X.2011.04480.x
Mudge, S. R., Rae, A. L., Diatloff, E., and Smith, F. W. (2002). Expression analysis suggests novel roles for members of the Pht1 family of phosphate transporters in Arabidopsis. Plant J. 31, 341–353. doi: 10.1046/j.1365-313x.2002.01356.x
Müller, D., and Leyser, O. (2011). Auxin, cytokinin and the control of shoot branching. Ann. Bot. 107, 1203–1212. doi: 10.1093/aob/mcr069
Mulugu, S., Bai, W., Fridy, P. C., Bastidas, R. J., Otto, J. C., Dollins, D. E., et al. (2007). A conserved family of enzymes that phosphorylate inositol hexakisphosphate. Science 316, 106–109. doi: 10.1126/science.1139099
Munnik, T., and Nielsen, E. (2011). Green light for polyphosphoinositide signals in plants. Curr. Opin. Plant Biol. 14, 489–497. doi: 10.1016/j.pbi.2011.06.007
Munnik, T., and Testerink, C. (2009). Plant phospholipid signaling: “in a nutshell”. J. Lipid Res. 50 Suppl(Suppl.), S260–S265. doi: 10.1194/jlr.R800098-JLR200
Munnik, T., and Vermeer, J. E. (2010). Osmotic stress-induced phosphoinositide and inositol phosphate signalling in plants. Plant Cell Environ. 33, 655–669. doi: 10.1111/j.1365-3040.2009.02097
Mur, L. A., Kenton, P., Lloyd, A. J., Ougham, H., and Prats, E. (2008). The hypersensitive response; the centenary is upon us but how much do we know? J. Exp. Bot. 59, 501–520. doi: 10.1093/jxb/erm239
Murphy, A. M., Otto, B., Brearley, C. A., Carr, J. P., and Hanke, D. E. (2008). A role for inositol hexakisphosphate in the maintenance of basal resistance to plant pathogens. Plant J. 56, 638–652. doi: 10.1111/j.1365-313X.2008.03629.x
Nagy, R., Grob, H., Weder, B., Green, P., Klein, M., Frelet-Barrand, A., et al. (2009). The Arabidopsis ATP-binding cassette protein AtMRP5/AtABCC5 is a high affinity inositol hexakisphosphate transporter involved in guard cell signaling and phytate storage. J. Biol. Chem. 284, 33614–33622. doi: 10.1074/jbc.M109.030247
Navarro, L., Dunoyer, P., Jay, F., Arnold, B., Dharmasiri, N., Estelle, M., et al. (2006). A plant miRNA contribute to antibacterial resistance by repressing auxin signaling. Science 312, 436–439. doi: 10.1126/science.1126088
Newberry, E. A., Bhandari, R., Minsavage, G. V., Timilsina, S., Jibrin, M. O., Kemble, J., et al. (2019). Independent evolution with the gene flux originating from multiple Xanthomonas species explaiins genomic heterogeneity in Xanthomonas performance. Appl. Environ. Microbiol. 85:e00885–19. doi: 10.1128/AEM.00885-19
Otegui, M. S., Capp, R., and Staehelin, L. A. (2002). Developing seeds of Arabidopsis store different minerals in two types of vacuoles and in the endoplasmic reticulum. Plant Cell 14, 1311–1327. doi: 10.1105/tpc.010486
Parvin Laha, N., Dhir, Y. W., Giehl, R. F. H., Schäfer, E. M., Gaugler, P., Haghighat Shishavan, Z., et al. (2020). ITPK1-dependentinositol polyphosphates regulate auxin responses in Arabidopsis thaliana. bioRxiv [Prerpint] doi: 10.1101/2020.04.23.058487
Paul, S., Datta, S., and Datta, K. (2015). miRNA regulation of nutrient homeostasis in plants. Front. Plant Sci. 6:232. doi: 10.3389/fpls.2015.00232
Perera, I. Y., Hung, C. Y., Brady, S., Muday, G. K., and Boss, W. F. (2006). A universal role for inositol 1,4,5-trisphosphate-mediated signaling in plant gravitropism. Plant Physiol. 140, 746–760. doi: 10.1104/pp.105.075119
Péret, B., Clément, M., Nussaume, L., and Desnos, T. (2011). Root developmental adaptation to phosphate starvation: Better safe than sorry. Trends Plant Sci. 16, 442–450. doi: 10.1016/j.tplants.2011.05.006
Pieterse, C. M. J., Van der Does, D., Zamioudis, C., Leon-Reyes, A., and Van Wees, S. C. M. (2012). Hormonal modulation of plant immunity. Annu. Rev. Cell Dev. Biol. 28, 489–521. doi: 10.1146/annurev-cellbio-092910-154055
Pisani, F., Livermore, T., Rose, G., Chubb, J. R., Gaspari, M., and Saiardi, A. (2014). Analysis of Dictyostelium discoideum inositol pyrophosphate metabolism by gel electrophoresis. PLoS One 9:e85533. doi: 10.1371/journal.pone.0085533
Plaxton, W. C., and Tran, H. T. (2011). Metabolic adaptations of phosphate-starved plants. Plant Physiol. 156, 1006–1015. doi: 10.1104/pp.111.175281
Poon, J. S. Y., Le Fevre, R. E., Carr, J. P., Hanke, D. E., and Murphy, A. M. (2020). Inositol hexakisphosphate biosynthesis underpins PAMP-triggered immunity to Pseudomonas syringae pv. tomato in Arabidopsis thaliana but is dispensable for establishment of systemic acquired resistance. Mol. Plant Pathol. 21, 376–387. doi: 10.1111/mpp.12902
Potnis, N., Minsavage, G., Smith, J. K., Hurlbert, J. C., Norman, D., Rodrigues, R., et al. (2012). Avirulence proteins AvrBs7 from Xanthomonas gardneri andAvrBs1.1 from Xanthomonas euvesicatoria contribute to a novel gene-for-geneinteraction in pepper. Mol. Plant Microbe Interact. 25, 307–320. doi: 10.1094/MPMI-08-11-0205
Puga, M. I., Mateos, I., Charukesi, R., Wang, Z., Franco-Zorrilla, J. M., de Lorenzo, L., et al. (2014). SPX1 is a phosphate-dependent inhibitor of phosphate starvation response 1 in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 111, 14947–14952. doi: 10.1073/pnas.1404654111
Qi, W., Manfield, I. W., Muench, S. P., and Baker, A. (2017). AtSPX1 affects the AtPHR1-DNA-binding equilibrium by binding monomeric AtPHR1 in solution. Biochem. J. 474, 3675–3687. doi: 10.1042/BCJ20170522
Qiu, D., Wilson, M. S., Eisenbeis, V. B., Harmel, R., Riemer, E., Haas, T. M., et al. (2020). Analysis of inositol phosphate metabolism by capillary electrophoresis electrospray ionization mass spectrometry. Nat. Commun. 11:6035. doi: 10.1038/s41467-020-19928-x
Raboy, V. (2020). Low phytic acid Crops: Observations based on four decades of research. Plants 9:140. doi: 10.3390/plants9020140
Raboy, V., and Gerbasi, P. (1996). Genetics of myo-Inositol phosphate synthesis and accumulation. Subcell Biochem. 26, 257–285. doi: 10.1007/978-1-4613-0343-5_9
Raetz, C. R. H., and Whitfield, C. (2002). Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71, 635–700. doi: 10.1146/annurev.biochem.71.110601.135414
Rai, V. K., Sharma, S. S., and Sharma, S. (1986). Reversal of ABA-induced stomatal closure by phenolic compounds. J. Exp. Bot. 37, 129–134.
Rao, F., Cha, J., Xu, J., Xu, R., Vandiver, M. S., Tyagi, R., et al. (2014). Inositol pyrophosphates mediate the DNA-PK/ATM-p53 cell death pathway by regulating CK2 phosphorylation of Tti1/Tel2. Mol. Cell 54, 119–132. doi: 10.1016/j.molcel.2014.02.020
Raskin, I. (1992). Salicylate, a new plant hormone. Plant Physiol. 99, 799–803. doi: 10.1104/pp.99.3.799
Raskin, I., Turner, I. M., and Melander, W. R. (1989). Regulation of heat production in the inflorescences of an Arum lily by endogenous salicylic acid. Proc. Natl. Acad. Sci. U.S.A. 86, 2214–2218. doi: 10.1073/pnas.86.7.2214
Rausch, C., and Bucher, M. (2002). Molecular mechanisms of phosphate transport in plants. Planta 216, 23–37. doi: 10.1007/s00425-002-0921-3
Riemer, E., Qiu, D., Laha, D., Harmel, R. K., Gaugler, P., Gaugler, V., et al. (2021). ITPK1 is an InsP6/ADP phosphotransferase that controls phosphate signaling in Arabidopsis. Mol. Plant 14, 1864–1880. doi: 10.1016/j.molp.2021.07.011
Rockström, J., Steffen, W., Noone, K., Persson, A., Chapin, F. S., Lambin, E. F., et al. (2009). A safe operating space for humanity. Nature 461, 472–475. doi: 10.1038/461472a
Rubio, V., Linhares, F., Solano, R., Martín, A. C., Iglesias, J., Leyva, A., et al. (2001). A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Gene Dev. 15, 2122–2133. doi: 10.1101/gad.204401
Ryan, R. P., Vorhölter, F.-J., Potnis, N., Jones, J. B., Van Sluys, M.-A., Bogdanove, A. J., et al. (2011). Pathogenomics of Xathomonas: Understanding bacterium-plant interactions. Nat. Rev. Microbiol. 9, 344–355. doi: 10.1038/nrmicro2558
Saiardi, A., Caffrey, J. J., Snyder, S. H., and Shears, S. B. (2000). The inositol hexakisphosphate kinase family: Catalytic flexibility and function in yeast vacuole biogenesis. J. Biol. Chem. 275, 24686–24692. doi: 10.1074/jbc.M002750200
Saiardi, A., Erdjument-Bromage, H., Snowman, A. M., Tempst, P., and Snyder, S. H. (1999). Synthesis of diphosphoinositol pentakisphosphate by a newly identified family of higher inositol polyphosphate kinases. Curr. Biol. 9, 1323–1326. doi: 10.1016/s0960-9822(00)80055-x
Saiardi, A., Sciambi, C., McCaffery, J. M., Wendland, B., and Snyder, S. H. (2002). Inositol pyrophosphates regulate endocytic trafficking. Proc. Natl. Acad. Sci. U.S.A. 99, 14206–14211. doi: 10.1073/pnas.212527899
Salehin, M., Bagchi, R., and Estelle, M. (2015). SCFTIR1/AFB-based auxin perception: Mechanism and role in plant growth and development. Plant Cell 27, 9–19. doi: 10.1105/tpc.114.133744
Secco, D., Bouain, N., Rouached, A., Prom-u-thai, C., Hanin, M., Pandey, A. K., et al. (2017). Phosphate, phytate and phytases in plants: From fundamental knowledge gained in Arabidopsis to potential biotechnological applications in wheat. Crit. Rev. Biotechnol. 37, 898–910. doi: 10.1080/07388551.2016.1268089
Secco, D., Jabnoune, M., Walker, H., Shou, H., Wu, P., Poirier, Y., et al. (2013). Spatio-temporal transcript profiling of rice roots and shoots in response to phosphate starvation and recovery. Plant Cell 25, 4285–4304. doi: 10.1105/tpc.113.117325
Seidel, S. J., Gaiser, T., Ahrends, H. E., Hueging, H., Siebert, S., Bauke, S. L., et al. (2021). Crop response to P fertilizer omission under a changing climate-experimental and modeling results over 115 years of a long-term fertilizer experiment. Field Crops Res. 268:108174. doi: 10.1016/j.fcr.2021.108174
Seybold, H., Trempel, F., Ranf, S., Scheel, D., Romeis, T., and Lee, J. (2014). Ca2+ signaling in plant immune response: From pattern recognition receptors to Ca2+ decoding mechanisms. N. Phytol. 204, 782–790. doi: 10.1111/nph.13031
Sharma, N., Chaudhary, C., and Khurana, P. (2020a). Role of myo-inositol during skotomorphogenesis in Arabidopsis. Sci. Rep. 10:17329. doi: 10.1038/s41598-020-73677-x
Sharma, N., Chaudhary, C., and Khurana, P. (2020b). Wheat Myo-inositol phosphate synthase influences plant growth and stress responses via ethylene mediated signaling. Sci. Rep. 10:10766. doi: 10.1038/s41598-020-67627-w
Sheard, L. B., Tan, X., Mao, H., Withers, J., Ben-Nissan, G., Hinds, T. R., et al. (2010). Jasmonate perception by inositol phosphate-potentiated COI1-JAZ co-receptor. Nature 468, 400–405. doi: 10.1038/nature09430
Shears, S. B. (2015). Inositol pyrophosphates: Why so many phosphates? Adv. Biol. Regul. 57, 203–221. doi: 10.1016/j.jbior.2014.09.015
Shimizu-Mitao, Y., and Kakimoto, T. (2014). Auxin sensitivities of all Arabidopsis Aux/IAAs for degradation in the presence of every TIR1/AFB. Plant Cell Physiol. 55, 1450–1459. doi: 10.1093/pcp/pcu077
Shin, H., Shin, H. S., Dewbre, G. R., and Harrison, M. J. (2004). Phosphate transport in Arabidopsis: Pht1;1 and Pht1;4 play a major role in phosphate acquisition from both low- and high-phosphate environments. Plant J. 39, 629–642. doi: 10.1111/j.1365-313X.2004.02161.x
Sorefan, K., Booker, J., Haurogne, K., Goussot, M., Bainbridge, K., Foo, E., et al. (2003). MAX4 and RMS1 are orthologous dioxygenase-like genes that regulate shoot branching in Arabidopsis and pea. Genes Dev. 17, 1469–1474. doi: 10.1101/gad.256603
Stevenson-Paulik, J., Bastidas, R. J., Chiou, S. T., Frye, R. A., and York, J. D. (2005). Generation of phytate-free seeds in Arabidopsis through disruption of inositol polyphosphate kinases. Proc. Natl. Acad. Sci. U.S.A. 102, 12612–12617. doi: 10.1073/pnas.0504172102
Stevenson-Paulik, J., Odom, A. R., and York, J. D. (2002). Molecular and bioch, mical characterization of two plant inositol polyphosphate 6-/3-/5-kinases. J. Biol. Chem. 277, 42711–42718. doi: 10.1074/jbc.M209112200
Streb, H., Irvine, R. F., Berridge, M. J., and Schulz, I. (1983). Release of Ca2+ from a non mitochondrial intracellular store in pancreatic acinar cells by inositol-1,4,5- trisphosphate. Nature 306, 67–69. doi: 10.1038/306067a0
Sweetman, D., Stavridou, I., Johnson, S., Green, P., Caddick, S. E. K., and Brearley, C. A. (2007). Arabidopsis thaliana inositol 1,3,4-trisphosphate 5/6-kinase 4 (AtITPK4) is an outlier to a family of ATP-grasp fold proteins from Arabidopsis. FEBS Lett. 581, 4165–4171. doi: 10.1016/j.febslet.2007.07.046
Tan, X., Calderon-Villalobos, L. I. A., Sharon, M., Zheng, C., Robinson, C. V., Estelle, M., et al. (2007). Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645. doi: 10.1038/nature05731
Tanaka, H., Dhonukshe, P., Brewer, P. B., and Friml, J. (2006). Spatiotemporal asymmetric auxin distribution: A means to coordinate plant development. Cell Mol. Life Sci. 63, 2738–2754. doi: 10.1007/s00018-006-6116-5
Tantikanjana, T., Yong, J. W., Letham, D. S., Griffith, M., Hussain, M., Ljung, K., et al. (2001). Control of axillary bud initiation and shoot architecture in Arabidopsis through the supershoot gene. Genes Dev. 15, 1577–1588. doi: 10.1101/gad.887301
Taylor, C. B., Bariola, P. A., DelCardyre, S. B., Raines, R. T., and Green, P. J. (1993). RNS2: A senescence-associated RNase of Arabidopsis that diverged from the S-RNases before speciation. Proc. Natl Acad. Sci. U.S.A 90, 5118–5122. doi: 10.1073/pnas.90.11.5118
Teper, D., Sunitha, S., Martin, G. B., and Sessa, G. (2015). Five Xanthomonas type III effectors suppress cell death induced by components of immunity-associated MAP kinase cascades. Plant Signal. Behav. 10:e1064573. doi: 10.1080/15592324.2015.1064573
Thieme, F., Koebnik, R., Bekel, T., Berger, C., Boch, J., Büttner, D., et al. (2005). Insights into genome plasticity and pathogenicity of the plant pathogenic bacterium Xanthomonas campestris pv. vesicatoria revealed by the complete genome sequence. J. Bacteriol. 187, 7254–7266. doi: 10.1128/JB.187.21.7254-7266.2005
Thines, B., Katsir, L., Melotto, M., Niu, Y., Mandaokar, A., Liu, G., et al. (2007). JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signaling. Nature 448, 661–665. doi: 10.1038/nature05960
Thota, S. G., and Bhandari, R. (2015). The emerging roles of inositol pyrophosphates in eukaryotic cell physiology. J. Biosci. 40, 593–605. doi: 10.1007/s12038-015-9549-x
Tucker, E. B., and Boss, W. F. (1996). Mastoparan-induced intracellular Ca2+ fluxes may regulate cell-to-cell communication in plants. Plant Physiol. 111, 459–467. doi: 10.1104/pp.111.2.459
Val-Torregrosa, B., Bundó, M., Martín-Cardoso, H., Bach-Pages, M., Chiou, T.-J., Flors, V., et al. (2022). Segundo B.S. Phosphate-induced resistance to pathogen infection in Arabidopsis. Plant J. 110, 452–469. doi: 10.1111/tpj.15680
Van Loon, L. C., Rep, M., and Pieterse, C. M. J. (2006). Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 44, 135–162. doi: 10.1146/annurev.phyto.44.070505.143425
Vance, C. P., Uhde-Stone, C., and Allan, D. L. (2003). Phosphorus acquisition and use: Critical adaptations by plants for securing a nonrenewable resource. N. Phytol. 157, 423–447. doi: 10.1046/j.1469-8137.2003.00695.x
Voglmaier, S. M., Bembenek, M. E., Kaplin, A. I., Dormán, G., Olszewski, J. D., Prestwich, G. D., et al. (1996). Purified inositol hexakisphosphate kinase is an ATP synthase: Diphosphoinositol pentakisphosphate as a high-energy phosphate donor. Proc. Natl. Acad. Sci. U.S.A. 93, 4305–4310. doi: 10.1073/pnas.93.9.4305
Wang, D., Pajerowska-Mukhtar, K., Culler, A. H., and Dong, X. (2007). Salicylic acid inhibits pathogen growth in plants through repression of the auxin signaling pathway. Curr. Biol. 17, 1784–1790. doi: 10.1016/j.cub.2007.09.025
Wang, H., Falck, J. R., Hall, T. M., and Shears, S. B. (2011). Structural basis for an inositol pyrophosphate kinase surmounting phosphate crowding. Nat. Chem. Biol. 8, 111–116. doi: 10.1038/nchembio.733
Wang, H., Gu, C., Rolfes, R. J., Jessen, H. J., and Shears, S. B. (2018). Structural and biochemical characterization of Siw14: A protein-tyrosine phosphatase fold that metabolizes inositol pyrophosphates. J. Biol. Chem. 293, 6905–6910. doi: 10.1074/jbc.RA117.001670
Wang, H., Nair, V. S., Holland, A. A., Capolicchio, S., Jessen, H. J., Johnson, M. K., et al. (2015). Asp1 from Schizosaccharomyces pombe binds a [2Fe-2S](2+) cluster which inhibits inositol pyrophosphate 1-phosphatase activity. Biochemistry 54, 6462–6474. doi: 10.1021/acs.biochem.5b00532
Wang, R., and Estelle, M. (2014). Diversity and specificity: Auxin perception and signaling through the TIR1/AFB pathway. Curr. Opin. Plant Biol. 21, 51–58. doi: 10.1016/j.pbi.2014.06.006
Wang, Y., Chu, Y. J., and Xue, H. W. (2012). Inositol polyphosphate 5-phosphatase-controlled Ins(1,4,5)P3/Ca2+ is crucial for maintaining pollen dormancy and regulating early germination of pollen. Development 139, 2221–2233. doi: 10.1242/dev.081224
Wang, Y., Lin, W. H., Chen, X., and Xue, H. W. (2009). The role of Arabidopsis 5PTase13 in root gravitropism through modulation of vesicle trafficking. Cell Res. 19, 1191–1204. doi: 10.1038/cr.2009.105
Wang, Z., Kuo, H.-F., and Chiou, T.-J. (2021). Intracellular phosphate sensing and regulation of phosphate transport systems in plants. Plant Physiol. 187, 2043–2055. doi: 10.1093/plphys/kiab343
Wasternack, C., and Hause, B. (2013). Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. an update to the 2007 review in Annals of Botany. Ann. Bot. 111, 1021–1058. doi: 10.1093/aob/mct067
Wheeler, G. L., and Brownlee, C. (2008). Ca2+ signalling in plants and green algae - changing channels. Trends Plant Sci. 13, 506–514. doi: 10.1016/j.tplants.2008.06.004
Whitfield, H., White, G., Sprigg, C., Riley, A. M., Potter, B. V. L., Hemmings, A. M., et al. (2020). An ATP-responsive metabolic cassette comprised of inositol tris/tetrakisphosphate kinase 1 (ITPK1) and inositol pentakisphosphate 2-kinase (IPK1) buffers diphosphosphoinositol phosphate levels. Biochem. J. 477, 2621–2638. doi: 10.1042/BCJ20200423
Wild, R., Gerasimaite, R., Jung, J.-Y., Truffault, V., Pavlovic, I., Schmidt, A., et al. (2016). Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 352, 986–990. doi: 10.1126/science.aad9858
Wilson, M. S. C., Livermore, T. M., and Saiardi, A. (2013). Inositol pyrophosphates: Between signalling and metabolism. Biochem. J. 452, 369–379. doi: 10.1042/BJ20130118
Woodward, A. W., and Bartel, B. (2005). Auxin: Regulation, action, and interaction. Ann. Bot. 95, 707–735. doi: 10.1093/aob/mci083
Wu, J., Qin, Y., and Zhao, J. (2008). Pollen tube growth is affected by exogenous hormones and correlated with hormone changes in styles in Torenia fournieri L. Plant Growth Regul. 55, 137–148. doi: 10.1007/s10725-008-9268-5
Wundenberg, T., and Mayr, G. W. (2012). Synthesis and biological actions of diphosphoinositol phosphates (inositol pyrophosphates), regulators of cell homeostasis. Biol. Chem. 393, 979–998. doi: 10.1515/hsz-2012-0133
Xia, H. J., Brearley, C., Elge, S., Kaplan, B., Fromm, H., and Mueller-Roeber, B. (2003). Arabidopsis inositol polyphosphate 6-/3-kinase is a nuclear protein that complements a yeast mutant lacking a functional ArgR-Mcm1 transcription complex. Plant Cell 15, 449–463. doi: 10.1105/tpc.006676
Xie, D. X., Feys, B. F., James, S., Nieto-Rostro, M., and Turner, J. G. (1998). COI1: An Arabidopsis gene required for jasmonate-regulated defense and fertility. Science 280, 1091–1094. doi: 10.1126/science.280.5366.1091
Xu, J., Brearley, C. A., Lin, W. H., Wang, Y., Ye, R., Mueller-Roeber, B., et al. (2005). A role of Arabidopsis inositol polyphosphate kinase AtIpk2a, in pollen germination and root growth. Plant Physiol. 137, 94–103. doi: 10.1104/pp.104.045427
Xu, L., Liu, F., Lechner, E., Genschik, P., Crosby, W. L., Ma, H., et al. (2002). The SCF(COI1) ubiquitin–ligase complexes are required for jasmonate response in Arabidopsis. Plant Cell 14, 1919–1935. doi: 10.1105/tpc.003368
Yan, Y., Stolz, S., Chételat, A., Reymond, P., Pagni, M., Dubugnon, L., et al. (2007). A downstream mediator in the growth repression limb of the jasmonate pathway. Plant Cell 19, 2470–2483. doi: 10.115/tpc.107.050708
Young, L. M., Evans, M. L., and Hertel, R. (1990). Correlations between gravitropic curvature and auxin movement across gravistimulated roots of Zea mays. Plant Physiol. 92, 792–796. doi: 10.1104/pp.92.3.792
Zarate, S. I., Kempema, L. A., and Walling, L. (2007). Silverleaf whitefly induces salicylic acid defenses and suppresses effectual jasmonic acid defenses. Plant Physiol. 143, 866–875. doi: 10.1104/pp.106.090035
Zhan, H., Zhong, Y., Yang, Z., and Xia, H. (2015). Enzyme activities of Arabidopsis inositol polyphosphate kinases AtIPK2α and AtIPK2β are involved in pollen development, pollen tube guidance and embryogenesis. Plant J. 82, 758–771. doi: 10.1111/tpj.12846
Zhang, Q., van Wijk, R., Shahbaz, M., Roels, W., Schooten, B. V., Vermeer, J. E. M., et al. (2018). Arabidopsis phospholipase C3 is involved in lateral root initiation and ABA responses in seed germination and stomatal closure. Plant Cell Physiol. 59, 469–486. doi: 10.1093/pcp/pcx194
Zhang, Z. B., Yang, G., Arana, F., Chen, Z., Li, Y., and Xia, H. J. (2007). Arabidopsis inositol polyphosphate 6-/3-kinase (AtIpk2beta) is involved in axillary shoot branching via auxin signaling. Plant Physiol. 144, 942–951. doi: 10.1104/pp.106.092163
Zheng, X. Y., Spivey, N. W., Zeng, W., Liu, P. P., Fu, Z. Q., Klessig, D. F., et al. (2012). Coronatine promotes Pseudomonas syringae virulence in plants by activating a signaling cascade that inhibits salicylic acid accumulation. Cell Host Microbe 11, 587–596. doi: 10.1016/j.chom.2012.04.014
Zhu, J., Lau, K., Puschmann, R., Harmel, R. K., Zhang, Y. J., Pries, V., et al. (2019). Two bifunctional inositol pyrophosphate kinases/phosphatases control plant phosphate homeostasis. Elife 8:e43582. doi: 10.7554/eLife.43582
Keywords: inositol pyrophosphate, jasmonic acid signaling pathway, salicylic acid signaling pathway, plant biotic and abiotic stress responses, phosphate homeostasis, auxin response
Citation: Riemer E, Pullagurla NJ, Yadav R, Rana P, Jessen HJ, Kamleitner M, Schaaf G and Laha D (2022) Regulation of plant biotic interactions and abiotic stress responses by inositol polyphosphates. Front. Plant Sci. 13:944515. doi: 10.3389/fpls.2022.944515
Received: 15 May 2022; Accepted: 20 July 2022;
Published: 11 August 2022.
Edited by:
Fiona L. Goggin, University of Arkansas, United StatesReviewed by:
Catherine Freed, Virginia Tech, United StatesMariya Khodakovskaya, University of Arkansas at Little Rock, United States
Copyright © 2022 Riemer, Pullagurla, Yadav, Rana, Jessen, Kamleitner, Schaaf and Laha. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Esther Riemer, ZXJpZW1lckB1bmktYm9ubi5kZQ==; Debabrata Laha, ZGxhaGFAaWlzYy5hYy5pbg==
 Esther Riemer
Esther Riemer Naga Jyothi Pullagurla2
Naga Jyothi Pullagurla2 Henning J. Jessen
Henning J. Jessen Marília Kamleitner
Marília Kamleitner Gabriel Schaaf
Gabriel Schaaf Debabrata Laha
Debabrata Laha