- State Key Laboratory of Agrobiotechnology and National Maize Improvement Center of China, Department of Plant Genetics and Breeding, China Agricultural University, Beijing, China
VQ motif-containing proteins play crucial roles in abiotic stress responses in plants. Recent studies have shown that some VQ proteins physically interact with WRKY transcription factors to activate downstream genes. In the present study, we identified and characterized genes encoding VQ motif-containing proteins using the most recent version of the maize genome sequence. In total, 61VQ genes were identified. In a cluster analysis, these genes clustered into nine groups together with their homologous genes in rice and Arabidopsis. Most of the VQ genes (57 out of 61 numbers) identified in maize were found to be single-copy genes. Analyses of RNA-seq data obtained using seedlings under long-term drought treatment showed that the expression levels of most ZmVQ genes (41 out of 61 members) changed during the drought stress response. Quantitative real-time PCR analyses showed that most of the ZmVQ genes were responsive to NaCl treatment. Also, approximately half of the ZmVQ genes were co-expressed with ZmWRKY genes. The identification of these VQ genes in the maize genome and knowledge of their expression profiles under drought and osmotic stresses will provide a solid foundation for exploring their specific functions in the abiotic stress responses of maize.
Introduction
Maize is one of the most important crops worldwide, and has recently become the most widely planted crop in China. Like any other crop, maize is greatly affected by various abiotic stresses such as drought. Generally, plants respond to abiotic stresses through several different strategies such as avoidance and tolerance. Many stress-related genes have been isolated from maize and other plants. Among these stress-related genes are those coding transcription factors that are involved in multiple stress response pathways. Some of the best-characterized stress-responsive transcription factors are C-repeat-binding factor (CBF)/dehydration-responsive element-binding (DREB) (Liu et al., 2013), NF-YB1 (Nelson et al., 2007), NAC (Hu et al., 2006), bZIP (Zhang et al., 2015), and WRKY proteins (Hu et al., 2013). Among these transcription factors, WRKYs are encoded by one of the largest gene families involved in abiotic stress responses (Eulgem et al., 2000; Dong et al., 2003; Chen et al., 2012). Genes encoding WRKY transcription factors have been identified in many species including maize, rice and Arabidopsis (Rushton et al., 2010; Wei et al., 2012). Members of the WRKY gene family have been shown to participate in several biotic and abiotic stress responses (Jiang and Deyholos, 2009; Li et al., 2009, 2013), leaf senescence (Robatzek and Somssich, 2002; Miao et al., 2004), hormone signaling (Chen et al., 2010; Shang et al., 2010), and seed development (Luo et al., 2005).
Several studies have shown that VQ domain-containing proteins can physically interact with WRKY transcription factors (Chi et al., 2013; Lin and Jing, 2015). The VQ proteins contain 50–60 conserved amino acids in the VQ (FxxxVQxLTG) motif, and interact with WRKY transcription factors via the conserved residues V and Q (Lai et al., 2011). AtVQ9 was reported to act antagonistically with WRKY8, which directly bind to the promoter of RDA29A in response to salt stress. Transgenic Arabidopsis plants with reduced AtVQ9 expression or overexpressing WRKY8 showed tolerance to osmotic stress (Hu et al., 2013). Two other VQ domain-containing proteins, SIB1 and SIB2, were shown to interact with WRKY33 by recognizing the C-terminal WRKY domain. The expression levels of SIB1 and SIB2 were up-regulated in response to the necrotrophic pathogen Botrytis cinerea. Overexpression of SIB1 in Arabidopsis showed enhanced resistance to B.cinerea. Further analyses indicated that the N-terminus of both SIB1 and SIB2 contained dual subcellular localization signals for the chloroplast and nucleus (Lai et al., 2011). MSK1, also known as AtVQ21, acts as a substrate of MAPK4 (MAP kinase 4), interacting directly with WRKY25 and WRKY33 to activate the MAPK4-regulated defense response (Andreasson et al., 2005). AtCAMBP25, which encodes another VQ motif-containing protein, was shown to function as a negative regulator in the osmotic stress response (Perruc et al., 2004). AtVQ14 was shown to regulate endosperm growth through its interaction with MINISEED3, which encodes the WRKY transcription factor WRKY10 (Luo et al., 2005; Wang et al., 2010). A large-scale analysis of the interactions between Arabidopsis VQ and WRKY proteins in yeast cells showed that VQ proteins could act as cofactors for group I and IIc WRKY proteins (Cheng et al., 2012). VQ motif-containing proteins could also partner with other transcription factors such as PIF, a bHLH-type transcription factor (Li et al., 2014c).
In recent years, 34, 40, and 74 VQ family genes have been identified in Arabidopsis (Cheng et al., 2012), rice (Kim et al., 2013; Li et al., 2014b), and soybean (Wang et al., 2014), respectively. More recently, 18 VQ genes were identified in grapevine (Vitis vinifera L.) (Wang et al., 2015). The expression profiles of VQ genes were analyzed in response to pathogen infection and drought stress in rice, and in response to low-nitrogen stress in soybean. However, the VQ genes in maize have remained largely uncharacterized, although large number of genomic and RNA-seq datasets are available for maize (Schnable et al., 2009; Lai et al., 2010; Jiao et al., 2012; Chen et al., 2014; Li et al., 2014a). In this study, we searched the maize genome to identify VQ genes identification. We conducted a systematic phylogenetic analysis comparing the maize VQ genes with those in rice and Arabidopsis. To infer the potential functions of these genes, we analyzed the expression profiles of ZmVQ genes using RNA-seq data and qRT-PCR obtained from maize seedling shoots under long-term drought stress and salt treatment, respectively. The identification of these ZmVQ genes and knowledge of their transcription patterns under abiotic stress will be useful for further studies on the molecular mechanisms of these important transcription factors in the abiotic stress tolerance of maize.
Materials and Methods
Searches for VQ Proteins in Maize Database and Phylogenetic Analysis
The VQ motif (Pfam05678) was used as a query to scan the B73 filter gene database. BLASTP searches of protein libraries1 were also performed using the full-length amino acid sequence predicted for each newly identified gene. Only predicted protein sequences containing VQ motifs were defined as ZmVQ proteins. Multiple sequence alignment of the 61 ZmVQ proteins was conducted using the MUSCLE method (Arsova et al., 2010; Grant et al., 2010; Shi et al., 2013; Li et al., 2015). Then, a phylogenetic tree was constructed using the neighbor-joining method with MEGA6 software and bootstrap analysis of 1,000 replicates.
Analysis of Gene Transcription Patterns in Different Tissues Using Public Datasets
To analyze differences in transcriptional patterns among various tissues, the expression values of ZmVQ genes were downloaded from our previous work (Chen et al., 2014). The samples used in that work included seed tissues (embryo and endosperm), root, shoot, shoot apical meristem, leaf, cob, tassel, silk, ovule, and immature ear. The detailed descriptions of the 78 different tissues are summarized in Supplementary Table S1. The methods for normalizing transcript levels and constructing heat maps were described previously (Chen et al., 2014).
Identification of cis-elements in Promoters of ZmVQ and ZmWRKY Genes
The distribution of seven cis-elements was determined by analyzing the promoter regions (2000-bp DNA sequences upstream of gene start codons) of ZmVQ and ZmWRKY genes. The cis-elements are binding sites for the following classes of transcription factors: WRKY transcription factor (W-box, TTGAC[C/T]) (Chen et al., 2012), drought response element (DRE, [A/G]CCGAC) (Sakuma et al., 2002), and calmodulin-binding transcription activator (CG-box, [A/C/G]CGCG[C/G/T]), all of which are involved in various abiotic stress responses (Yang and Poovaiah, 2002); MYB transcription factor, which is involved in drought inducibility (MBS, CAACTG) (Urao et al., 1993); abscisic acid responsive element (ABRE, [C/T]ACGTG[G/T]) (Osakabe et al., 2014), salicylic acid responsive element (SARE, TGACG) (Pieterse and Van Loon, 2004), and environmental stimuli responsive element (G-box, CACGTG) (Williams et al., 1992). The sequences of ZmVQ promoters were downloaded from the Phytozome v.10.1.
Plant Materials, Growth Conditions and Drought Stress Treatment
Seeds of the maize inbred line B73 were germinated in an incubator, and then transferred to pots (10 cm diameter, 10 seedlings per pot) containing a vermiculite: soil mixture (1:1, v/v). The seedlings were grown under the following conditions: 25°C ± 2°C, 60–70% humidity, 18-h light/6-h dark photoperiod, with natural sunlight. Each plot had 10 seedlings. For the drought treatment, the pots containing seedlings were fully irrigated until seedlings reached the three-leaf stage. Then, the seedlings were subjected to drought stress by withholding water for 6 days. The seedlings were re-watered on day 6. The seedling shoots were sampled at three time points; day 3, 6, and 7 (24 h after re-watering), respectively. The controls were watered as necessary and sampled at the same time points. Three biological replicates were collected. All samples were wrapped in aluminum foil, frozen in liquid nitrogen, and stored at -80°C until use.
Osmotic Stress Treatment
Seeds of the B73 inbred line were used in this experiment. Seeds from the middle of the ear were sterilized using H2O2 (30%), and washed three times with sterile water. Then, the seeds were transferred to filter paper moistened with sterile water on a plastic tray, and covered with two pieces of filter paper that were also moistened with sterile water. The seed trays were kept in an incubator at 28°C, 60% humidity, in the dark. After 2 days, germinated seeds were transplanted into pots (10-cm diameter, 10 seedlings per pot) containing a vermiculite:soil mixture (1:1, v/v). The seedlings were grown in the incubator at 28°C, 60% humidity, under an 18-h light/6-h dark photoperiod. RNA was isolated from 2-week-old seedlings subjected to a NaCl osmotic stress treatment. For the NaCl treatment, the pots containing seedlings were irrigated with 250 mM NaCl (Hu et al., 2013), while the controls were irrigated with water. Seedling shoots were collected at 0, 24, and 48 h after the treatment. The experiment was repeated three times, with three plants per repeat. All samples were wrapped in aluminum foil, frozen in liquid nitrogen, and stored at -80°C until use.
Total RNA Extraction and Construction of Sequencing Libraries
Two biological replicates were used for RNA isolation, construction of libraries, and RNA sequencing. Total RNAs were isolated from maize seedling shoots using Trizol reagent (Invitrogen2) according to the manufacturer’s protocol. RNA-seq libraries were constructed as described previously (Chen et al., 2014). The libraries were sequenced using the Illumina HiSeq2000 platform3. After sequencing, raw reads were aligned to the B73 reference genome (v2) using Tophat 2.0.6 (Trapnell et al., 2009) with default settings for all parameters. In a correlation analysis, the average R2 between the two biological replicates was 0.94 (Supplementary Figure S1). The unique mapped reads were used to measure the transcript abundance of every gene using Cufflinks (Trapnell et al., 2012).
qRT-PCR Analysis
Total RNAs were isolated from maize tissue with TRIzol reagent (Invitrogen). DNase treatment was performed on 4 μg total RNA using RQ1 RNase-free DNase (Promega4) prior to first-strand cDNA synthesis using M-MLV Reverse Transcriptase (Promega). Real-time quantitative RT-PCR was performed using SYBR® Premix Ex TaqTM (Perfect Real Time, TaKaRa5) on an ABI7500 instrument (Applied Biosystems6). Each 20 μL PCR reaction mixture contained 10 μL 2× SYBR Green Premix, 2 μL 10-fold diluted cDNA product, 0.4 μL ROX Reference Dye II, and 0.4 μL forward and reverse primers. The PCRs were performed with a holding step at 95°C for 30 s, followed by 40 cycles of denaturation at 95°C for 30 s, annealing at 55–60°C for 30 s and extension at 72°C for 30 s. Quantitative assays were performed on each cDNA sample three times. Relative gene expression levels were calculated using the 2-ΔΔCT method (Livak and Schmittgen, 2001). β-Actin mRNA levels were determined with specific primers to allow normalization of transcript levels among samples. Supplementary Table S2 lists the primers used for qRT-PCR.
To validate the RNA-seq data, we isolated RNA from stored samples of drought-treated plants (days 3, 6, and 7-rewatered) and their corresponding controls and conducted qRT-PCR analyses. We evaluated the expression levels of ZmVQ genes in drought-treated plants and control plants of the same age in the qRT-PCR analyses to ensure that differences in gene expression were due to the treatment and not differences in the developmental stage of the samples. Seven different tissues were used for tissue-specific expression analysis in the B73 genetic background. Ear (7 cm), tassel (16 cm), seed [16 days after planting (DAP)], endosperm (16 DAP), and embryo (16 DAP) tissues were harvested from plants at the tasseling stage in the field in the summer of 2015. Root and leaf samples were obtained from 2-week-old seedlings that were grown in the laboratory under the same conditions as those described in the osmotic treatment section. All samples were wrapped in aluminum foil, frozen in liquid nitrogen, and stored at -80°C until use.
Data were analyzed by one-way ANOVA using Microsoft Excel software. The student’s t-test at a significance level of 0.05 was used to detect significant differences between the treatment and control values. All expression data were obtained from three biological repeats. Values shown in figures are means of three repeats with standard deviation (SD).
Analyses of ZmVQ Expression Profiles and Co-expression with ZmWRKY Based on RNA-seq Data
The relative expression levels of ZmVQ genes were calculated using the inferred formula (Yuan et al., 2006), where Ctreatment is the expression level of genes under drought treatment and Ccontrol is the expression level of genes in the control. Then, the relative expression level (Crelative = Ctreatment - Ccontrol) of ZmVQ genes was plotted to show the expression profiles (Figure 4). We analyzed co-expression patterns of ZmVQ and ZmWRKY genes (Supplementary Table S3) using RNA-seq data obtained from plants under drought at three time points. Genes whose expression levels were positively correlated (Pearson Correlation Coefficient > 0.90; (Li et al., 2014c) with those of other genes were selected for map drawing using Cytoscape (Shannon et al., 2003).
Results
Maize VQ Domain-Containing Protein Family has 61 Members
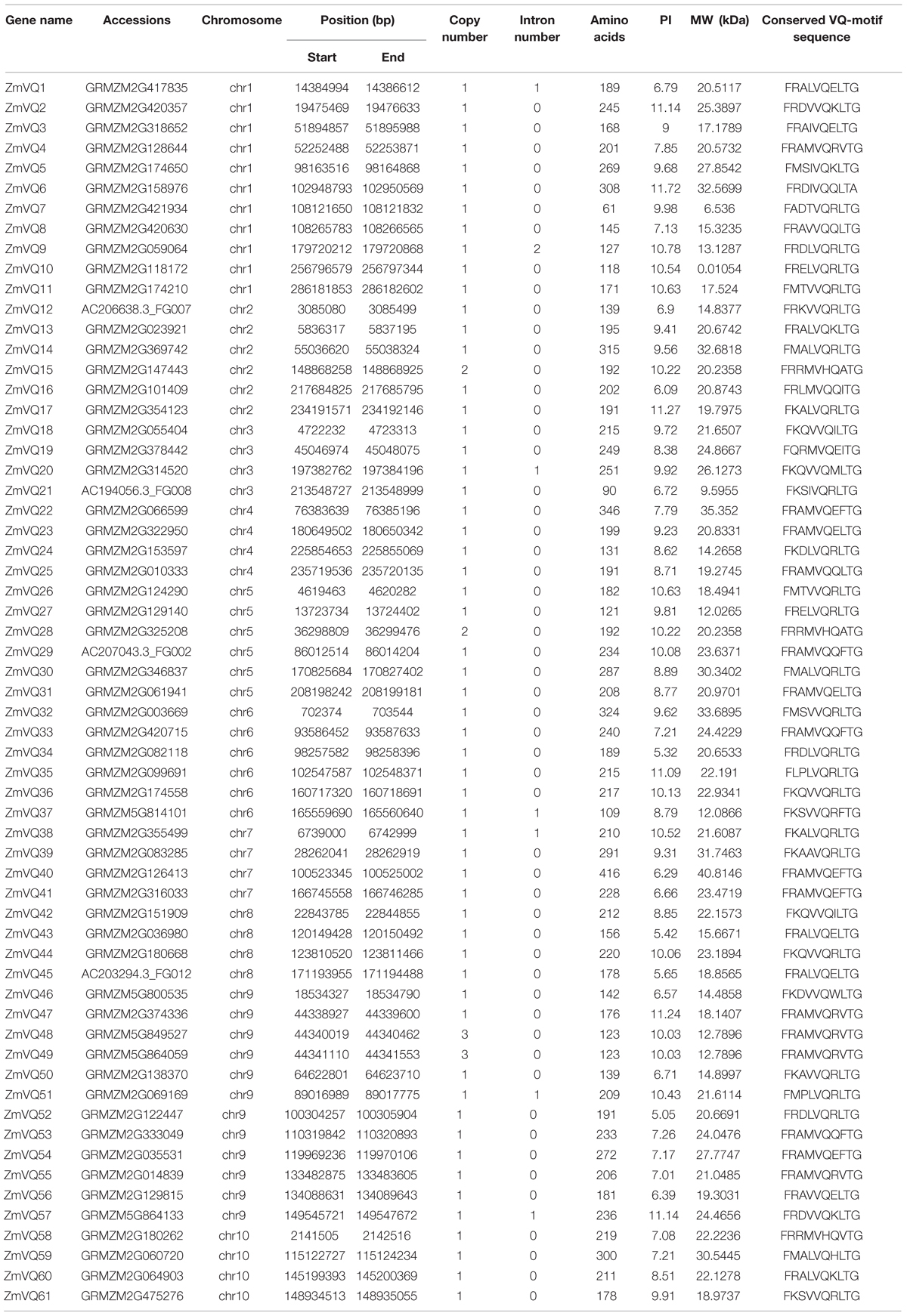
In total, we identified 61 VQ motif-containing proteins, which were numbered from ZmVQ1-ZmVQ61 based on the locations of their encoding genes on the chromosomes (Table 1; Figure 1; Supplementary Figure S2). Of the 61 proteins identified, 42 contained the conserved motif FxxxVQxLTG (42/61), and the other 19 contained FxxxVQxxTG or FxxxVQxLTx motifs (Figure 1). The core amino acids of three identified proteins (ZmVQ15, ZmVQ28, and ZmVQ58) were VH instead of VQ, similar to OsVQ37 and OsVQ39 in rice (Kim et al., 2013). Most of the ZmVQ proteins had fewer than 300 amino acid residues and five (ZmVQ6, ZmVQ14, ZmVQ22, ZmVQ32, and ZmVQ40) had more than 300. The predicted isoelectric points of the VQ proteins varied from 5.05 (ZmVQ52) to 11.72 (ZmVQ6). Gene structure analyses showed that 54 out of 61 ZmVQ genes lacked introns, while the other seven (ZmVQ1, ZmVQ9, ZmVQ20, ZmVQ37, ZmVQ38, ZmVQ51, and ZmVQ57) had only one or two introns. Most of the ZmVQ genes were single-copy genes in the B73 reference genome, except for ZmVQ15, ZmVQ28, ZmVQ48, and ZmVQ49, which had two or three copies. Table 1 shows detailed information for the ZmVQ genes, including their accession number, copy number, chromosome location, number of amino acid residues, molecular weight, and conserved motifs.

FIGURE 1. Sequence alignment of 61 conserved VQ motif regions in maize. Highly conserved amino acids in VQ motif are shown in red.
Phylogenetic Analysis of VQ Domain-Containing Genes in Maize
To explore the relationships of ZmVQ genes in maize, we conducted a phylogenetic analysis of the VQ genes from maize, rice and Arabidopsis. This analysis was conducted using the neighbor-joining method and 61, 34, and 40 VQ genes from maize, Arabidopsis and rice, respectively. The genes clustered into nine distinct groups based on the structural features of their protein sequences (ZmVQI–V; Figure 2). Among the nine groups, there were 20 ZmVQ genes in group I, 2 ZmVQ genes in group II, 5 ZmVQ genes in group III, 9 ZmVQ genes in group IV, 14 ZmVQ genes in group V, 9 ZmVQ genes in group VI, and 2 ZmVQ genes in group VII. Groups VIII or IX were largely rice-specific, with no maize VQ genes and only one Arabidopsis VQ gene (AtVQ21) in group VIII.
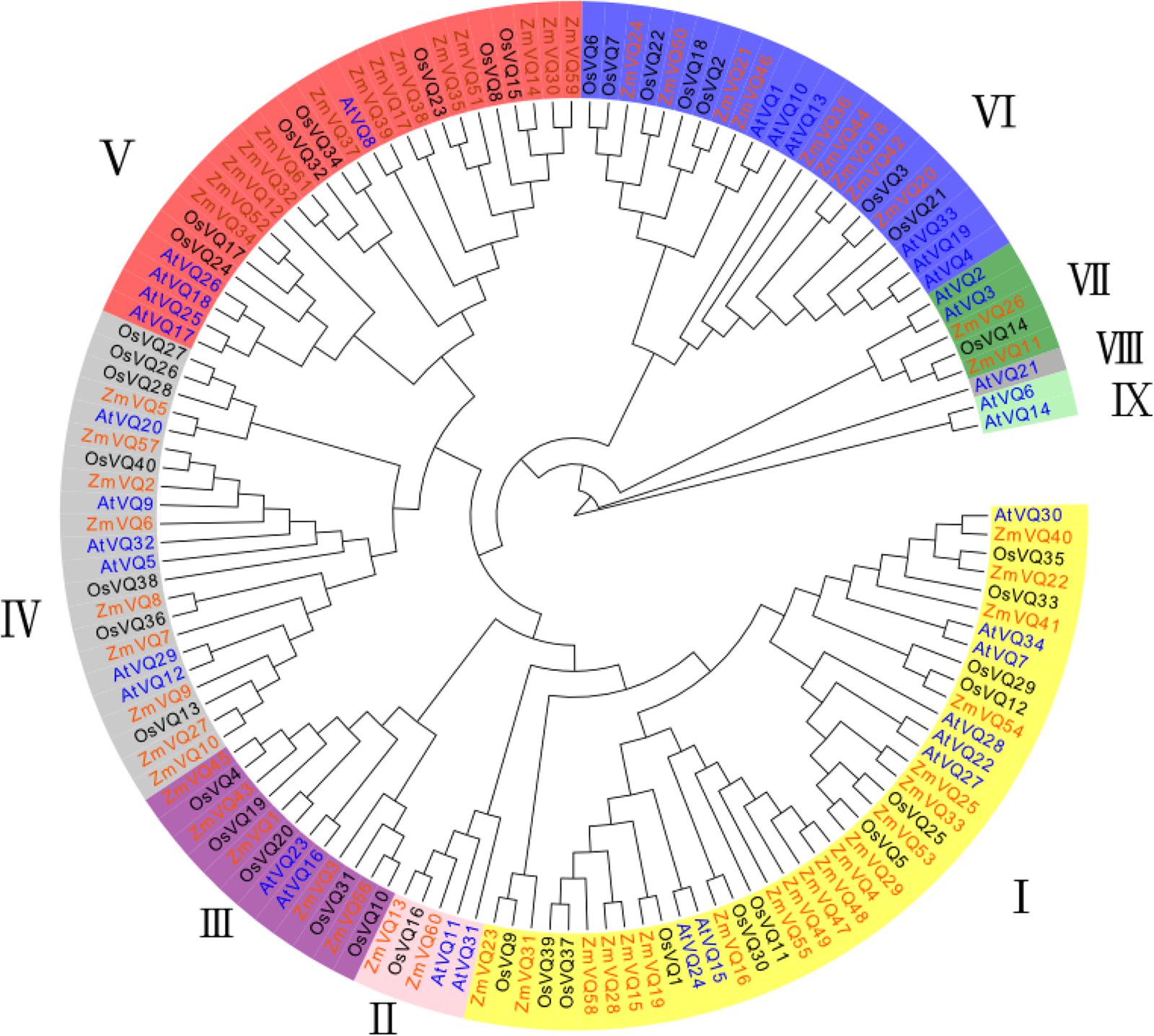
FIGURE 2. Phylogenetic tree of VQ genes from maize, Arabidopsis and rice. Sixty-one ZmVQ genes, 34 AtVQ genes, and 40 OsVQ genes clustered into nine groups (groups I–IX, represented by different colors).
All of the VQ domain-containing proteins contained the conserved motif of FxxxVxxxTx (Figure 1). Among the 61 ZmVQ proteins, 45 had the amino acid “L” next to the “TG,” eight had the amino acid “F” (ZmVQ22, 29, 33, 37, 40, 41, 53, and 54), six had the amino acid “V”(ZmVQ4, 47, 48, 49, 55, and 58), two had the amino acid “I” (ZmVQ16 and 19), and two had the amino acid “A” (ZmVQ15 and 28). The proteins with an “L” in this conserved domain were distributed into among all nine groups, but those with the rarer residues (F, V, I, and A) were restricted to groups I and V. Rice had 11 VQ proteins with residues other than “L” next to the “TG”: “F” (OsVQ5, 12, 25, 29, 33, 34, and 35), “V” (OsVQ11, 30, and 37), “I” (OsVQ1), while Arabidopsis had seven: “F” (AtVQ7, 27, 28, 30), “V” (AtVQ15 and 24), and “Y” (AtVQ22). Those with rarer amino acids in this domain also clustered into groups I and V, similar to those in maize (Figure 2).
Expression Patterns of 61 ZmVQ Genes in Different Maize Tissues
The expression patterns of the ZmVQ genes were investigated using available RNA-seq datasets from 78 different tissues (Supplementary Table S1). The 61 ZmVQ genes were clearly divided into four groups based on their expression patterns. Groups A, B, and C each contained six ZmVQ genes, and group D contained 43 ZmVQ genes. Genes in groups A and C tended to be expressed at higher levels in reproductive tissues such as the cob, tassel, and ovule (Figure 3). For example, ZmVQ13 and ZmVQ40 showed the highest expression levels in the cob, and ZmVQ42, ZmVQ43, and ZmVQ45 showed the highest expression levels in the ovule. Genes in group B showed no detectable expression in the RNA-seq libraries analyzed. Genes in group D were expressed at high levels in vegetative organs such as shoots, roots, and leaves. Some genes in group D showed tissue-specific expression patterns. For example, ZmVQ3, 7, 10, 21, 34, 35, and 38 showed the highest expression levels in roots, while ZmVQ2, 4, 6,11,20,22, 26, 41, 39, 44, 55, 56, and 59 were expressed at higher levels in leaves. Transcripts of ZmVQ41 were detected in leaf tissues, based on our RNA-seq data. The tissue-specific expression patterns of 12 ZmVQ genes were validated by qPCR analysis using seven different tissues (Supplementary Figure S3; Supplementary Table S6).
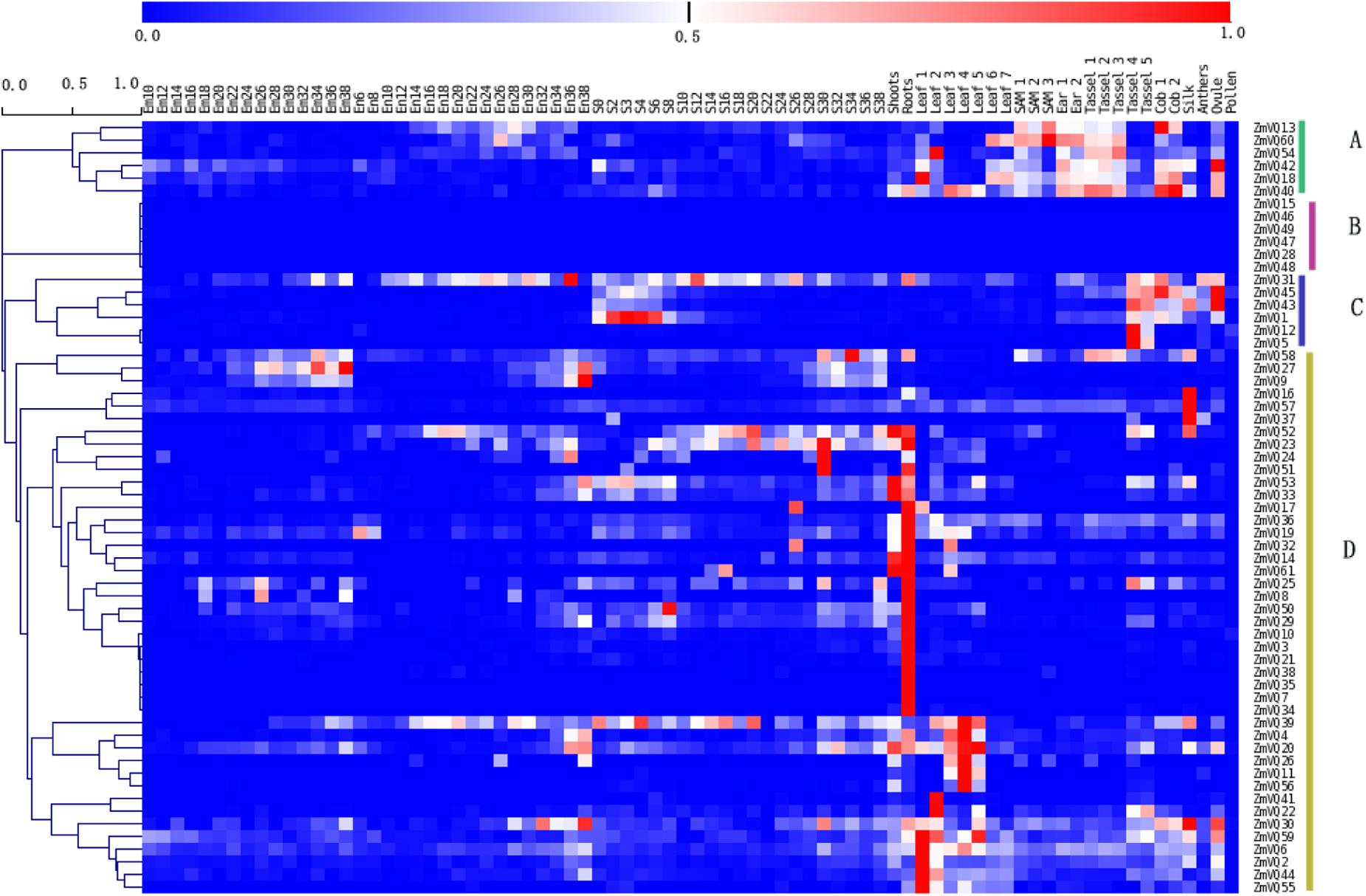
FIGURE 3. Heat map showing expression levels of 61 ZmVQ genes in 78 different tissues, based on RNA-seq data. Different colors in map represent gene transcript abundance values as shown in bar at top of figure.
Expression Patterns of ZmVQ Genes in Response to Drought Stress
Previous studies have shown that some VQ genes in Arabidopsis and rice were induced by NaCl, dehydration, and drought, based on qRT-PCR analyses (Hu et al., 2013; Kim et al., 2013). We analyzed the expression patterns of ZmVQ genes under drought stress. Total RNA was extracted from the drought treated seedlings and libraries were constructed for sequencing. First, the transcript levels of the drought-responsive marker gene ZmNAC111 (Mao et al., 2015) were quantified to evaluate the effect of our drought treatment. In our RNA-seq data, the expression level of ZmNAC111 changed dramatically in plants under drought stress (FPKM values of 38.80 and 100.03 on days 3 and 6 of drought stress, respectively) and decreased markedly after re-watering (FPKM value of 5.43 on day 7, after re-watering). These results confirmed that our drought treatment method was effective. Under these conditions, 41 ZmVQ genes were drought-responsive. According to their different expression patterns under drought treatment, the 41 drought-responsive ZmVQ genes clustered into four groups (Figure 4). The first group contained six genes that showed early drought-responsive expression, with decreased (ZmVQ1, 4, 42, 54) or induced (ZmVQ36, 40) transcript levels after 3 days of drought stress. Then, four genes of ZmVQ1, 4, 42, 54 were transcriptional up-regulated slightly at time point of 6 days (Figure 4), while it was opposed for ZmVQ36 and ZmVQ40.The second group consisted of 10 genes showing changed expression levels at both 3 and 6 days of drought stress. The third group contained 11 genes that showed a slower response to drought stress, with increased or decreased transcript levels after 6 days of drought stress. Among them, ZmVQ50 was expressed in response to re-watering. The fourth group consisted of 14 genes that did not show drought-responsive expression, but showed increased expression levels after re-watering. Together, our results showed that most of the VQ genes were responsive to drought stress, with complex expression patterns during drought stress and re-watering. The expression patterns of 10 ZmVQ genes were further validated by qRT-PCR (Supplementary Figure S4).
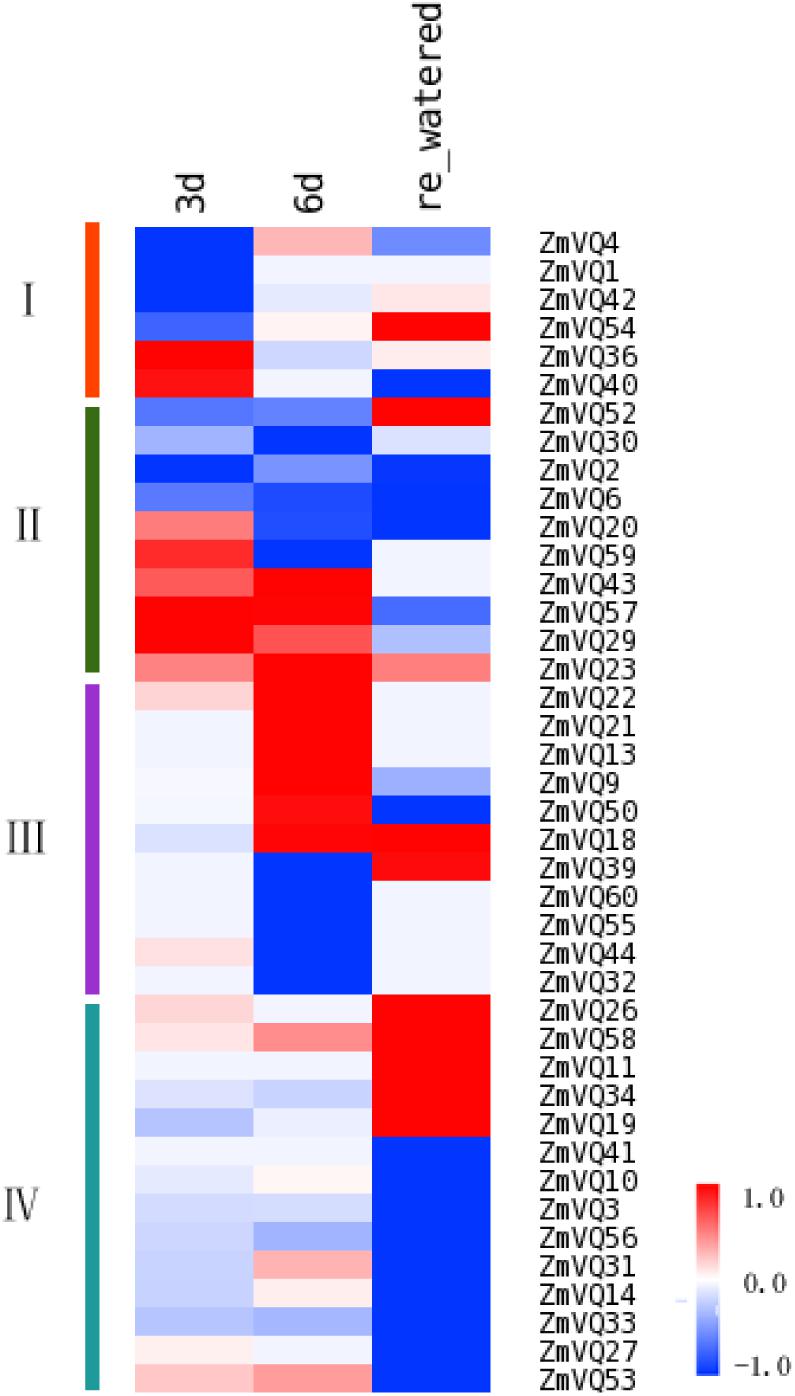
FIGURE 4. Heat map showing ZmVQ expression patterns in maize under long-term drought stress. ZmVQ gene expression levels were measured in tissues of seedlings at different stages of drought stress. Different colors in map represent gene transcript abundance values as shown in bar at top of figure. Color changes from blue to red in bar represent expression level changes from -1 to 1.
Expression of ZmVQ Genes Under NaCl Osmotic Stress
Hu et al. (2013) reported that Arabidopsis VQ9 was responsive to osmotic stress (Hu et al., 2013). To study whether the maize VQ genes were also responsive to osmotic stress, we conducted qRT-PCR analyses of 53 ZmVQ genes, but not the eight ZmVQ genes for which there were no gene-specific PCR primers. Total RNA was isolated from maize seedlings under NaCl osmotic stress. The relative expression values of the 53 ZmVQ genes are shown in Supplementary Table S4, while the untreated seedlings were used as control. The expression level of most ZmVQ genes increased from 0 to 24 h after the NaCl stress treatment, and then decreased by 48 h after the treatment. In particular, the expression levels of ZmVQ1, ZmVQ11, ZmVQ25, ZmVQ37, ZmVQ51, and ZmVQ52 were more than 20-times higher in drought-stressed seedlings than in control seedlings. By 24 h after the NaCl treatment, 36 out of the 53 ZmVQ genes were significantly up-regulated, while only 2 ZmVQ genes (ZmVQ16, 18) were down-regulated (Figure 5). By 48 h after the NaCl treatment, 29 out of 53 ZmVQ genes showed down-regulated expression, while four genes (ZmVQ4, 7, 33, 59) showed up-regulated expression. These findings suggested that many ZmVQ genes were involved in the response to NaCl osmotic stress.
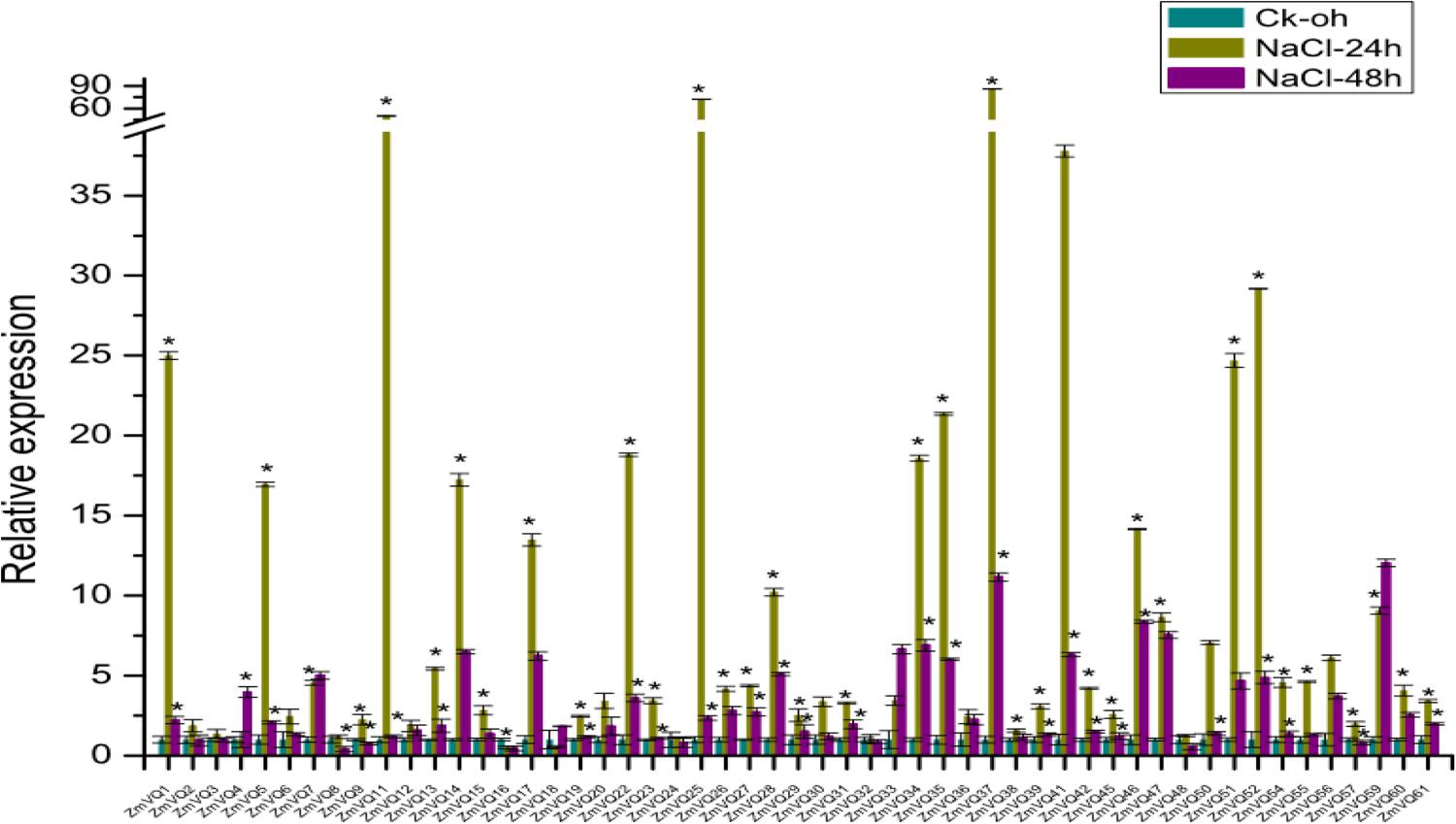
FIGURE 5. Expression of 53 ZmVQ genes in response to NaCl treatment. Gene expression in leaves of 2-week-old seedlings was analyzed by qRT-PCR. Expression levels in control (0 h) were normalized to 1. Asterisk indicates significant difference between treatment and control.
Co-expression of ZmVQ and ZmWRKY Genes
To investigate the potential interactions between WRKYs and VQs in maize, we conducted co-expression analysis for the 61 ZmVQ and ZmWRKY genes identified in previous studies (Figure 6; Supplementary Table S5) using the RNA-seq data. Twenty-seven ZmVQ genes were co-expressed with 49 ZmWRKY genes (Figure 6). Among these 27 ZmVQ genes, 21 were co-expressed with more than one ZmWRKY gene, while six ZmVQ genes (ZmVQ17, ZmVQ23, ZmVQ26, ZmVQ39, ZmVQ42, and ZmVQ59) were co-expressed with only one ZmWRKY gene (Figure 6). Moreover, some ZmVQ (ZmWRKY) genes showed co-expression patterns with other members of VQ (WRKY) genes. Therefore, it is possible that the co-expression of VQ genes is required for the normal functions of WRKY genes. Further verification and functional analyses are needed to explore the physical interactions among these transcription factors in maize.
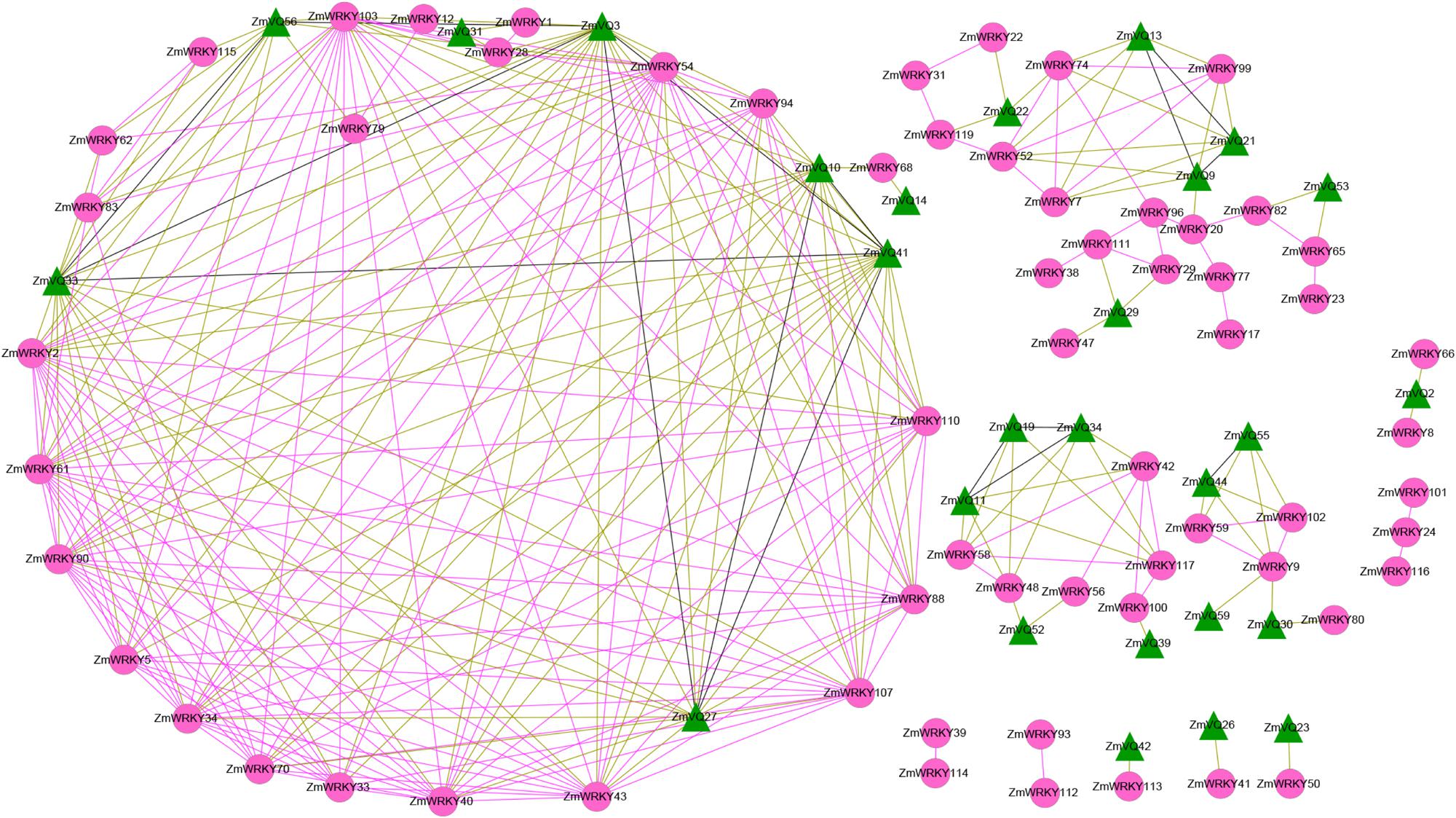
FIGURE 6. Co-expression of ZmVQ and ZmWRKY genes. Correlation coefficients of co-expression were >0.90. Red, green, and black lines indicate co-expression of two ZmWRKY genes, a ZmVQ gene and a ZmWRKY gene, and two ZmVQ genes, respectively. The circles with red color represent ZmWRKY genes, while the triangles with blue color represent ZmVQ genes.
Identification of cis-elements in ZmVQ and ZmWRKY Promoters
To explore the mechanisms by which these VQ genes responded to abiotic stresses, we searched for seven stress-related cis-acting regulatory elements (W-box, DRE, CG-box, MBS, ABRE, SARE, and G-box) in the ZmVQ and ZmWRKY promoters (2000-bp upstream sequences of start codons) (Supplementary Figures S5 and S6). The results revealed that 41∼92% of ZmVQ and 32∼90% ZmWRKY promoter regions enriched the cis-elements (Supplementary Figure S6). More than four of these cis-elements were present in 42 (69%) ZmVQ and 71 (60%) ZmWRKY promoter sequences. Some elements were detected in more than one location in the promoter sequences. For example, the promoter of ZmVQ22 gene contained eight CG-box sequences. At least one W-box (which binds WRKY transcription factors during abiotic stress responses) was present in 91% (55 out of 61) of ZmVQ promoters and 42% (50 out of 119) of ZmWRKY promoters. Supplementary Table S7 lists the cis-acting regulatory elements in the promoter regions of ZmVQ and ZmWRKY genes.
Discussion
Proteins containing the VQ motif play vital roles in stress responses and seed development in plants. Functional and expression profile analyses of VQ genes could help us to understand their regulation during specific developmental processes and/or stress responses. In this study, we isolated and characterized 61 ZmVQ genes using the latest maize genome data. Our analyses showed that there are more VQ genes in maize than in rice (40 OsVQ genes) (Li et al., 2014b) and in Arabidopsis (34 AtVQ genes) (Cheng et al., 2012), but fewer VQ genes in maize than in soybean (74 GmVQ genes) (Wang et al., 2014). The maize genome is larger than the rice and Arabidopsis genomes. Therefore, it is plausible that the maize genome contains more VQ genes than do the rice and Arabidopsis genomes. The soybean genome is smaller than the maize genome, but it contains more VQ genes. This may be because of ancient genome duplication events. It was reported that the soybean genome underwent two duplications 59 and 13 million years ago, which resulted in multiple copies of around 75% of its genes (Schmutz et al., 2010). Consistent with this explanation, there are two or three copies of most VQ genes in the soybean genome (except for the single-copy gene GmVQ55) (Wang et al., 2014), while most of the VQ genes in the maize, rice, and Arabidopsis genomes are single-copy genes.
Intronless genes are very common in the genomes of higher eukaryotes (Louhichi et al., 2011). The gene structure analysis revealed that most of the ZmVQ genes are intronless. Only six genes have one or two introns (Table 1). In the phylogenetic analysis of VQ genes in maize, rice and Arabidopsis, the few VQ genes containing intron(s) were distributed in several different groups. This result suggested that these introns arose relatively recently and independently in maize, rice, and Arabidopsis. The ZmVQ genes identified in our study, along with those identified in other species, will be useful for further research on intron evolution in plants.
The majority (67.21%) of ZmVQ genes showed drought- and osmotic-responsive expression, similar to VQ genes in rice and Arabidopsis. In rice, 22 out of 39 VQ genes were induced by biotic and abiotic stresses (Kim et al., 2013). Specifically, OsVQ2, 16, 20 were induced sharply by drought (Kim et al., 2013), as were their homologs in maize (ZmVQ21, 13, 1, Figure 4). Interestingly, ZmVQ13 and 1 were significantly induced by NaCl treatment (Figure 5), suggesting that these two ZmVQ genes are involved in both drought and osmotic stress responses. In Arabidopsis, AtVQ15 (also known as AtCaMBP25) and AtVQ9 were shown to be involved in osmotic stress responses (NaCl and mannitol) (Perruc et al., 2004; Hu et al., 2013). Transgenic plants overexpressing AtVQ15 showed increased sensitivity to osmotic stress induced by both NaCl and mannitol during the germination and seedling stages. In contrast, lines with down-regulated AtVQ15 expression showed increased osmotic stress tolerance, compared with that of wild type (Perruc et al., 2004). ZmVQ57, the homolog of AtVQ9, was induced under both drought and osmotic stresses (Figures 4 and 5). ZmVQ15, ZmVQ19, and ZmVQ28, orthologs of AtVQ15, were transcriptionally induced under salt stress (Figure 5). Moreover, ZmVQ19 was also induced by drought treatment (Figure 4), while ZmVQ28 was up-regulated in drought-stressed leaf meristems and ovaries of 1DAP (Kakumanu et al., 2012). Stress-related cis-acting elements are important clues for how gene expression is regulated in response to environmental stimuli. We identified many cis-elements in the promoters of drought-responsive genes (Supplementary Figure S7; Supplementary Table S8). In other studies, expression analyses showed that these genes were induced by drought stress at the seedling stage (Zheng et al., 2010; Mao et al., 2015). In our study, most ZmVQ genes were responsive to drought and NaCl, and their promoters contained more than three cis-elements. It is possible that the homologous genes between maize and rice, and between maize and Arabidopsis, have the same or similar functions in response to abiotic stresses.
Recently, genome-wide expression analyses showed that some rice VQ genes were co-expressed with WRKY transcription factors during the responses to attacks by three different pathogens (Li et al., 2014b; Wang et al., 2015). Thus, VQ and WRKY proteins might assemble to form one complex to regulate the target gene. In the present study, we found that 27 ZmVQ genes were co-expressed with 49 ZmWRKY transcription factors under drought stress (Figure 6; Supplementary Table S5). Furthermore, some of the co-expressed ZmWRKY genes were shown to be involved in abiotic stresses (Wei et al., 2012), implying that some VQ and WRKY genes are involved in the same biological pathway. In our RNA-seq data, most of the ZmWRKY genes co-expressed with ZmVQ genes belonged to groups I and II (Dong et al., 2003; Wei et al., 2012), whereas several studied has reported that most VQ genes interact with WRKY genes of groups I and II in response to environmental stimuli (Rushton et al., 2010; Cheng et al., 2012; Wang et al., 2015). The results of the cis-element analysis indicated that 55 out of 61 ZmVQ genes (91%) contained a W-box motif (TTGAC[C/T], binding sites for WRKY transcription factor) in their 2.0-kb promoter regions, while a W-box was present in the 1.5-kb promoter regions of 14 out of 18 VvVQ genes (78%) in grapevine (Wang et al., 2015). The presence of W-boxes in the promoters of ZmWRKY genes suggested that there is feedback regulation among ZmWRKY members, as reported in Arabidopsis (Dong et al., 2003). This indicates that WRKY transcription factors might act as binding factors to regulate the expression of both WRKY and VQ genes, resulting in the expression of WRKY cofactors to ensure appropriate responses to environmental stimuli (Cheng et al., 2012). Our results have shown which WRKY proteins interact with which VQ proteins. Further research should explore their physical interactions during the responses to various abiotic stresses in maize, to provide further molecular evidence for these interactions in plants.
Conclusion
We present here a comprehensive genome-wide study about the gene structure, phylogenetic relationship and tissue specificity of VQ domain-containing genes in maize. Expressional analysis of these VQ genes under drought and NaCl treatments suggested that some VQ genes are involved in abiotic stress responses. High co-expression correlation discovered between VQ genes and WRKY genes confirmed that many VQ genes and WRKY genes are likely functionally related. Information provided in this study will serve as a foundation for future exploration of the specific function of the VQ genes in the abiotic stress responses and their interaction with WRKY genes.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The reviewer Francisca Blanco and handling Editor Ariel Orellana declared that, despite having recently collaborated, the review process was conducted objectively.
Acknowledgments
This work was supported by the 863 Project (2012AA10A305), the 948 project (2011-G15) and the National Natural Science Foundation of China (31225020; 91435206; 31421005; 31301321).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2015.01177
TABLE S1 | Details of 78 different tissues used to generate RNA-seq data.
TABLE S2 | Premiers used for qPCR analyses of ZmVQ genes.
TABLE S3 | Accession number of ZmWRKY genes.
TABLE S4 | Transcript abundance (qPCR values) of genes in maize seedlings under NaCl stress.
TABLE S5 | The coefficient values of ZmVQ and ZmWRKY genes.
TABLE S6 | Stress-related cis-elements in promoters of ZmVQ and ZmWRKY genes.
TABLE S7 | Transcript abundance (qPCR values) of ZmVQ genes in seven maize different tissues.
TABLE S8 | Stress-related cis-elements in promoters of drought- responsive marker genes.
Footnotes
- ^http://www.ncbi.nlm.nih.gov/
- ^http://www.invitrogen.com/
- ^www.illumina.com
- ^www.promega.com
- ^http://www.takara-bio.com
- ^www.appliedbiosystems.com
References
Andreasson, E., Jenkins, T., Brodersen, P., Thorgrimsen, S., Petersen, N. H., Zhu, S., et al. (2005). The MAP kinase substrate MKS1 is a regulator of plant defense responses. EMBO J. 24, 2579–2589. doi: 10.1038/sj.emboj.7600737
Arsova, B., Hoja, U., Wimmelbacher, M., Greiner, E., Ustun, S., Melzer, M., et al. (2010). Plastidial thioredoxin z interacts with two fructokinase-like proteins in a thiol-dependent manner: evidence for an essential role in chloroplast development in Arabidopsis and Nicotiana benthamiana. Plant Cell 22, 1498–1515. doi: 10.1105/tpc.109.071001
Chen, H., Lai, Z., Shi, J., Xiao, Y., Chen, Z., and Xu, X. (2010). Roles of Arabidopsis WRKY18, WRKY40 and WRKY60 transcription factors in plant responses to abscisic acid and abiotic stress. BMC Plant Biol. 10:281. doi: 10.1186/1471-2229-10-281
Chen, J., Zeng, B., Zhang, M., Xie, S., Wang, G., Hauck, A., et al. (2014). Dynamic transcriptome landscape of maize embryo and endosperm development. Plant Physiol. 166, 252–264. doi: 10.1104/pp.114.240689
Chen, L., Song, Y., Li, S., Zhang, L., Zou, C., and Yu, D. (2012). The role of WRKY transcription factors in plant abiotic stresses. Biochim. Biophys. Acta 1819, 120–128. doi: 10.1016/j.bbagrm.2011.09.002
Cheng, Y., Zhou, Y., Yang, Y., Chi, Y. J., Zhou, J., Chen, J. Y., et al. (2012). Structural and functional analysis of VQ motif-containing proteins in Arabidopsis as interacting proteins of WRKY transcription factors. Plant Physiol. 159, 810–825. doi: 10.1104/pp.112.196816
Chi, Y., Yang, Y., Zhou, Y., Zhou, J., Fan, B., Yu, J. Q., et al. (2013). Protein-protein interactions in the regulation of WRKY transcription factors. Mol. Plant 6, 287–300. doi: 10.1093/mp/sst026
Dong, J., Chen, C., and Chen, Z. (2003). Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response. Plant Mol. Biol. 51, 21–37. doi: 10.1023/A:1020780022549
Eulgem, T., Rushton, P. J., Robatzek, S., and Somssich, I. E. (2000). The WRKY superfamily of plant transcription factors. Trends Plant Sci. 5, 199–206. doi: 10.1016/S1360-1385(00)01600-9
Grant, E. H., Fujino, T., Beers, E. P., and Brunner, A. M. (2010). Characterization of NAC domain transcription factors implicated in control of vascular cell differentiation in Arabidopsis and Populus. Planta 232, 337–352. doi: 10.1007/s00425-010-1181-2
Hu, H., Dai, M., Yao, J., Xiao, B., Li, X., Zhang, Q., et al. (2006). Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc. Natl. Acad. Sci. U.S.A. 103, 12987–12992. doi: 10.1073/pnas.0604882103
Hu, Y., Chen, L., Wang, H., Zhang, L., Wang, F., and Yu, D. (2013). Arabidopsis transcription factor WRKY8 functions antagonistically with its interacting partner VQ9 to modulate salinity stress tolerance. Plant J. 74, 730–745. doi: 10.1111/tpj.12159
Jiang, Y., and Deyholos, M. K. (2009). Functional characterization of Arabidopsis NaCl-inducible WRKY25 and WRKY33 transcription factors in abiotic stresses. Plant Mol. Biol. 69, 91–105. doi: 10.1007/s11103-008-9408-3
Jiao, Y., Zhao, H., Ren, L., Song, W., Zeng, B., Guo, J., et al. (2012). Genome-wide genetic changes during modern breeding of maize. Nat. Genet. 44, 812–815. doi: 10.1038/ng.2312
Kakumanu, A., Ambavaram, M. M. R., Klumas, C., Krishnan, A., Batlang, U., Myers, E., et al. (2012). Effects of drought on gene expression in maize reproductive and leaf meristem tissue revealed by RNA-seq. Plant Physiol. 160, 846–867. doi: 10.1104/pp.112.200444
Kim, D. Y., Kwon, S. I., Choi, C., Lee, H., Ahn, I., Park, S. R., et al. (2013). Expression analysis of rice VQ genes in response to biotic and abiotic stresses. Gene 529, 208–214. doi: 10.1016/j.gene.2013.08.023
Lai, J., Li, R., Xu, X., Jin, W., Xu, M., Zhao, H., et al. (2010). Genome-wide patterns of genetic variation among elite maize inbred lines. Nat. Genet. 42, 1027–1030. doi: 10.1038/ng.684
Lai, Z., Li, Y., Wang, F., Cheng, Y., Fan, B., Yu, J. Q., et al. (2011). Arabidopsis sigma factor binding proteins are activators of the WRKY33 transcription factor in plant defense. Plant Cell 23, 3824–3841. doi: 10.1105/tpc.111.090571
Li, G., Wang, D., Yang, R., Logan, K., Chen, H., Zhang, S., et al. (2014a). Temporal patterns of gene expression in developing maize endosperm identified through transcriptome sequencing. Proc. Natl. Acad. Sci. U.S.A. 111, 7582–7587. doi: 10.1073/pnas.1406383111
Li, N., Li, X., Xiao, J., and Wang, S. (2014b). Comprehensive analysis of VQ motif-containing gene expression in rice defense responses to three pathogens. Plant Cell Rep. 33, 1493–1505. doi: 10.1007/s00299-014-1633-4
Li, Y., Jing, Y., Li, J., Xu, G., and Lin, R. (2014c). Arabidopsis VQ MOTIF-CONTAINING PROTEIN29 represses seedling deetiolation by interacting with PHYTOCHROME-INTERACTING FACTOR1. Plant Physiol. 164, 2068–2080. doi: 10.1104/pp.113.234492
Li, H., Gao, Y., Xu, H., Dai, Y., Deng, D., and Chen, J. (2013). ZmWRKY33, a WRKY maize transcription factor conferring enhanced salt stress tolerances in Arabidopsis. Plant Growth Regul. 70, 207–216. doi: 10.1007/s10725-013-9792-9
Li, P., Zhang, S., Zhang, S., Li, F., Zhang, H., Cheng, F., et al. (2015). Carotenoid biosynthetic genes in Brassica rapa: comparative genomic analysis, phylogenetic analysis, and expression profiling. BMC Genomics 16:492. doi: 10.1186/s12864-015-1655-5
Li, S., Fu, Q., Huang, W., and Yu, D. (2009). Functional analysis of an Arabidopsis transcription factor WRKY25 in heat stress. Plant Cell Rep. 28, 683–693. doi: 10.1007/s00299-008-0666-y
Lin, R., and Jing, Y. (2015). VQ-motif-containing protein family of plant-specific transcriptional regulators. Plant Physiol. 169, 371–378. doi: 10.1104/pp.15.00788
Liu, S., Wang, X., Wang, H., Xin, H., Yang, X., Yan, J., et al. (2013). Genome-wide analysis of ZmDREB genes and their association with natural variation in drought tolerance at seedling stage of Zea mays L. PLoS Genet. 9:e1003790. doi: 10.1371/journal.pgen.1003790
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Louhichi, A., Fourati, A., and Rebai, A. (2011). IGD: a resource for intronless genes in the human genome. Gene 488, 35–40. doi: 10.1016/j.gene.2011.08.013
Luo, M., Dennis, E. S., Berger, F., Peacock, W. J., and Chaudhury, A. (2005). MINISEED3 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine-rich repeat (LRR) KINASE gene, are regulators of seed size in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 102, 17531–17536. doi: 10.1073/pnas.0508418102
Mao, H., Wang, H., Liu, S., Li, Z., Yang, X., Yan, J., et al. (2015). A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat. Commun. 6, 8326. doi: 10.1038/ncomms9326
Miao, Y., Laun, T., Zimmermann, P., and Zentgraf, U. (2004). Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol. Biol. 55, 853–867. doi: 10.1007/s11103-005-2142-1
Nelson, D. E., Repetti, P. P., Adams, T. R., Creelman, R. A., Wu, J., Warner, D. C., et al. (2007). Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc. Natl. Acad. Sci. U.S.A. 104, 16450–16455. doi: 10.1073/pnas.0707193104
Osakabe, Y., Yamaguchi-Shinozaki, K., Shinozaki, K., and Tran, L. S. (2014). ABA control of plant macroelement membrane transport systems in response to water deficit and high salinity. New Phytol. 202, 35–49. doi: 10.1111/nph.12613
Perruc, E., Charpenteau, M., Ramirez, B. C., Jauneau, A., Galaud, J. P., Ranjeva, R., et al. (2004). A novel calmodulin-binding protein functions as a negative regulator of osmotic stress tolerance in Arabidopsis thaliana seedlings. Plant J. 38, 410–420. doi: 10.1111/j.1365-313X.2004.02062.x
Pieterse, C. M., and Van Loon, L. (2004). NPR1: the spider in the web of induced resistance signaling pathways. Curr. Opin. Plant Biol. 7, 456–464. doi: 10.1016/j.pbi.2004.05.006
Robatzek, S., and Somssich, I. E. (2002). Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev. 16, 1139–1149. doi: 10.1101/gad.222702
Rushton, P. J., Somssich, I. E., Ringler, P., and Shen, Q. J. (2010). WRKY transcription factors. Trends Plant Sci. 15, 247–258. doi: 10.1016/j.tplants.2010.02.006
Sakuma, Y., Liu, Q., Dubouzet, J. G., Abe, H., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2002). DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration– and cold-inducible gene expression. Biochem. Biophys. Res. Commun. 290, 998–1009. doi: 10.1006/bbrc.2001.6299
Schmutz, J., Cannon, S. B., Schlueter, J., Ma, J., Mitros, T., Nelson, W., et al. (2010). Genome sequence of the palaeopolyploid soybean. Nature 463, 178–183. doi: 10.1038/nature08670
Schnable, P. S., Ware, D., Fulton, R. S., Stein, J. C., Wei, F., Pasternak, S., et al. (2009). The B73 maize genome: complexity, diversity, and dynamics. Science 326, 1112–1115. doi: 10.1126/science.1178534
Shang, Y., Yan, L., Liu, Z. Q., Cao, Z., Mei, C., Xin, Q., et al. (2010). The Mg-chelatase H subunit of Arabidopsis antagonizes a group of WRKY transcription repressors to relieve ABA-responsive genes of inhibition. Plant Cell 22, 1909–1935. doi: 10.1105/tpc.110.073874
Shannon, P., Markiel, A., Ozier, O., Baliga, N. S., Wang, J. T., Ramage, D., et al. (2003). Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. doi: 10.1101/gr.1239303
Shi, Z., Maximova, S., Liu, Y., Verica, J., and Guiltinan, M. J. (2013). The salicylic acid receptor NPR3 is a negative regulator of the transcriptional defense response during early flower development in Arabidopsis. Mol. Plant 6, 802–816. doi: 10.1093/mp/sss091
Trapnell, C., Pachter, L., and Salzberg, S. L. (2009). TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111. doi: 10.1093/bioinformatics/btp120
Trapnell, C., Roberts, A., Goff, L., Pertea, G., Kim, D., Kelley, D. R., et al. (2012). Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578. doi: 10.1038/nprot.2012.016
Urao, T., Yamaguchishinozaki, K., Urao, S., and Shinozaki, K. (1993). An Arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5, 1529–1539. doi: 10.2307/3869736
Wang, A., Garcia, D., Zhang, H., Feng, K., Chaudhury, A., Berger, F., et al. (2010). The VQ motif protein IKU1 regulates endosperm growth and seed size in Arabidopsis. Plant J. 63, 670–679. doi: 10.1111/j.1365-313X.2010.04271.x
Wang, M., Vannozzi, A., Wang, G., Zhong, Y., Corso, M., Cavallini, E., et al. (2015). A comprehensive survey of the grapevine VQ gene family and its transcriptional correlation with WRKY proteins. Front. Plant Sci. 6:417. doi: 10.3389/fpls.2015.00417
Wang, X., Zhang, H., Sun, G., Jin, Y., and Qiu, L. (2014). Identification of active VQ motif-containing genes and the expression patterns under low nitrogen treatment in soybean. Gene 543, 237–243. doi: 10.1016/j.gene.2014.04.012
Wei, K. F., Chen, J., Chen, Y. F., Wu, L. J., and Xie, D. X. (2012). Molecular phylogenetic and expression analysis of the complete WRKY transcription factor family in maize. DNA Res. 19, 153–164. doi: 10.1093/dnares/dsr048
Williams, M. E., Foster, R., and Chua, N. H. (1992). Sequences flanking the hexameric G-box core CACGTG affect the specificity of protein binding. Plant Cell 4, 485–496. doi: 10.1105/tpc.4.4.485
Yang, T. B., and Poovaiah, B. W. (2002). A calmodulin-binding/CGCG box DNA-binding protein family involved in multiple signaling pathways in plants. J. Biol. Chem. 277, 45049–45058. doi: 10.1074/jbc.M207941200
Yuan, J. S., Reed, A., Chen, F., and Stewart, C. N. Jr. (2006). Statistical analysis of real-time PCR data. BMC Bioinformatics 7:85. doi: 10.1186/1471-2105-7-85
Zhang, L., Zhang, L., Xia, C., Zhao, G., Liu, J., Jia, J., et al. (2015). A novel wheat bZIP transcription factor, TabZIP60, confers multiple abiotic stress tolerances in transgenic Arabidopsis. Physiol. Plant. 153, 538–554. doi: 10.1111/ppl.12261
Keywords: maize, VQ domain-containing protein, drought stress, transcription factor, WRKY gene
Citation: Song W, Zhao H, Zhang X, Lei L and Lai J (2016) Genome-Wide Identification of VQ Motif-Containing Proteins and their Expression Profiles Under Abiotic Stresses in Maize. Front. Plant Sci. 6:1177. doi: 10.3389/fpls.2015.01177
Received: 21 August 2015; Accepted: 08 December 2015;
Published: 05 January 2016.
Edited by:
Ariel Orellana, Universidad Andrés Bello, ChileReviewed by:
Serena Varotto, University of Padova, ItalyFrancisca Blanco, Universidad Andrés Bello, Chile
Copyright © 2016 Song, Zhao, Zhang, Lei and Lai. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jinsheng Lai, amxhaUBjYXUuZWR1LmNu
†These authors have contributed equally to this work
 Weibin Song
Weibin Song Haiming Zhao
Haiming Zhao Xiangbo Zhang
Xiangbo Zhang Lei Lei
Lei Lei Jinsheng Lai*
Jinsheng Lai*