- 1Jiangsu Key Laboratory of Phylogenomics and Comparative Genomics, School of Life Sciences, Jiangsu Normal University, Xuzhou, China
- 2Department of Applied Biology, College of Agriculture and Life Sciences, Chonnam National University, Gwangju, South Korea
Salinity, as a major environmental stressor, limits plant growth, development, and crop yield remarkably. However, plants evolve their own defense systems in response to salt stress. Recently, microRNA (miRNA) has been broadly studied and considered to be an important regulator of the plant salt-stress response at the post-transcription level. In this review, we have summarized the recent research progress on the identification, functional characterization, and regulatory mechanism of miRNA involved in salt stress, have discussed the emerging manipulation of miRNA to improve crop salt resistance, and have provided future direction for plant miRNA study under salt stress, suggesting that the salinity resistance of crops could be improved by the manipulation of microRNA.
Introduction
Salinity, as a major environmental stress factor, restricts crop growth and yield globally. It is reported that salinity affected a land area as large as 800 million hectares across the globe, accounting for 6% of the land (Abdel Latef et al., 2020; Attia et al., 2021). Approximately 20% of the irrigated soils are affected by salinity stress (Zhao et al., 2013), and 50% of arable land will be affected by 2050 (Butcher et al., 2016). Salt stress leads to changes in metabolic activity, cell wall damage, and cytoplasmic dissolution; it reduces the photosynthetic efficiency, accelerates aging, increases respiratory consumption and toxin accumulation, and eventually results in plant death (Osman et al., 2020; Abdel Latef et al., 2021). It is estimated that salinity can result in $27.3 billion in agricultural damage every year (Qadir et al., 2014). On the other hand, regional food scarcity will persist continually, particularly in South Asia, sub-Saharan Africa, the Middle East, and where population increase is rapid but agricultural outputs are low (FAO, 2017). Therefore, breeding and growing salt-tolerant crops to utilize the marginal and high-salinity soils are one of the most important strategies to meet the increase in food demand required by the estimated population in 2050 of 10 billion people (Mekonnen and Hoekstra, 2016; FAO, 2017; Morton et al., 2019).
MicroRNA (miRNA) is a non-coding single-stranded small RNA with a length of 21–24 nucleotides, and it acts as gene regulators to control the transcript abundance of its target gene. In the wild, miRNA exists in diverse organisms, including plants, animals, and microorganisms, and it regulates growth, development, signal transduction, response to adversity, and other biological processes. It was firstly discovered in Caenorhabditis (Lee and Ambros, 2001) and was then detected in four laboratories at approximately the same time in Llave et al. (2002), Mette et al. (2002), Park et al. (2002), Reinhart et al. (2002). After that, more and more plant miRNAs have been identified and functionally characterized in various plant species. MiRNA family names are listed in the order of publication, and miRNAs with similar sequences (usually fewer than 3 nt in difference) and common functions are classified as members of the same miRNA family (Wang Q. et al., 2014). Both the intraspecific conservation and interspecific differences of miRNAs are environmentally adaptive and evolve with the change in environment (Zhang et al., 2018). However, the evolution of miRNAs is conservative because some key target genes of miRNAs are conservative (Gramzow and Theißen, 2019).
Various enzymes and functional proteins are involved in the plant’s miRNA biosynthesis and functions. The primary miRNA transcripts for plants are produced by RNA polymerase II from miRNA genes, and these then pair with complementary bases to form special hairpin structures (Budak and Akpinar, 2015). Then, the stem ring secondary structure is generated by the DICER-LIKE1 (Bielewicz et al., 2013). After the methylation catalyzed by HUA Enhance 1 at the 3′ end, the double strand was transferred to the cytoplasm with the help of the transport protein HST. In the cytoplasm, this double-stranded miRNA is decomposed into mature single-stranded miRNA and integrated into RNA-induced silencing complex (RISC) cells, where miRNA interacts with the complementary target mRNA and activates the catalytic RISC with the assistance of Argonaute 1 (AGO1) (Koroban et al., 2016). There are two modes for miRNA to regulate gene expression: RNA cleavage and translation inhibition. The first mode is that miRNAs guide the Argonaute component of RISC to cleave a single phosphodiester bond opposite to the 10th and 11th nucleotides of the miRNA within complementary RNA. Then, the RISC will be free by releasing the fragments, and it then subsequently recognizes and cleaves another transcript (Jones-Rhoades et al., 2006). Afterward, the cleavage fragments are released to make the RISC competent for other RNA recognization and cleavage (Jones-Rhoades et al., 2006). MiRNA-mediated translational repression requires the participation of P-body components, a microtubule-severing enzyme, AGO1, and AGO10 (Brodersen et al., 2008). In addition, miRNA possibly prevents translation by triggering the sequestration of miRNA target in P-bodies (Chen, 2009). In addition, each miRNA can control multiple target genes (Haas et al., 2012). For instance, miR156 promotes floral meristem identity transformation by targeting SPL3, SPL4, and SPL5 in Arabidopsis thaliana (Xu et al., 2016). A gene can also be regulated by multiple miRNAs. For example, miR31 and miR143 affect steroid hormone synthesis by targeting the FSHR receptor (Zhang et al., 2019).
MiRNAs can regulate plant growth, development, pathogens, and abiotic stress responses. MiR160, miR169, peu-miRn68, and 477b are involved in the hormone signaling crosstalk model of root growth and development in apple rootstock, A. thaliana and Populus (Sorin et al., 2014; Lian et al., 2018; Meng et al., 2020). Cs-miR414 and cs-miR828 are involved in tea bud dormancy (Jeyaraj et al., 2014). For pathogen stress regulations, miR397 plays a negative regulatory role in apple resistance to hepatitis B virus (Yu et al., 2020), miR396 affects the susceptibility to rice blast (Chandran et al., 2019), and miR528 increases the viral defense ability of Oryza sativa (Wu et al., 2017). In the aspect of abiotic stress regulations, miR399 and miR827 are important for the resistance to phosphorus deficiency (Hackenberg et al., 2013; Du et al., 2018). The lack of sulfur induces the expression of miR395 for the regulation of genes in the sulfur assimilation pathway (Kawashima et al., 2009). The expression of miR319 is crucial for the cold tolerance of rice (Yang et al., 2013). MiR399 regulates Arabidopsis flowering at different temperatures (Kim et al., 2011). Recently, the comparative antagonistic expression profile of miR169 indicates that the miR169 family is a general regulator of various abiotic stresses (Rao et al., 2020). In addition, the over-expression of miR156 changes the expression level of other miRNAs, thus increasing the contents of anthocyanins, flavonoids, and flavonols and decreasing the total lignin content, suggesting the essential role of miRNAs in nutritional processes (Wang et al., 2020).
Noticeably, it is demonstrated that miRNA plays important roles in plant salinity responses and adaptation through various miRNA-mediated biological processes, including signal transduction, membrane transport, protein biosynthesis and degradation, photosynthesis, and transcription. In the present review, we mainly discuss the recent research progress on salt-stress-related miRNA in plants and the future research direction about miRNA in the salinity stress research field to come up with a strategy to improve the agronomic traits of stress tolerance through the manipulation of miRNAs.
Identification and Expression of Plant miRNAs Under Salt Stress
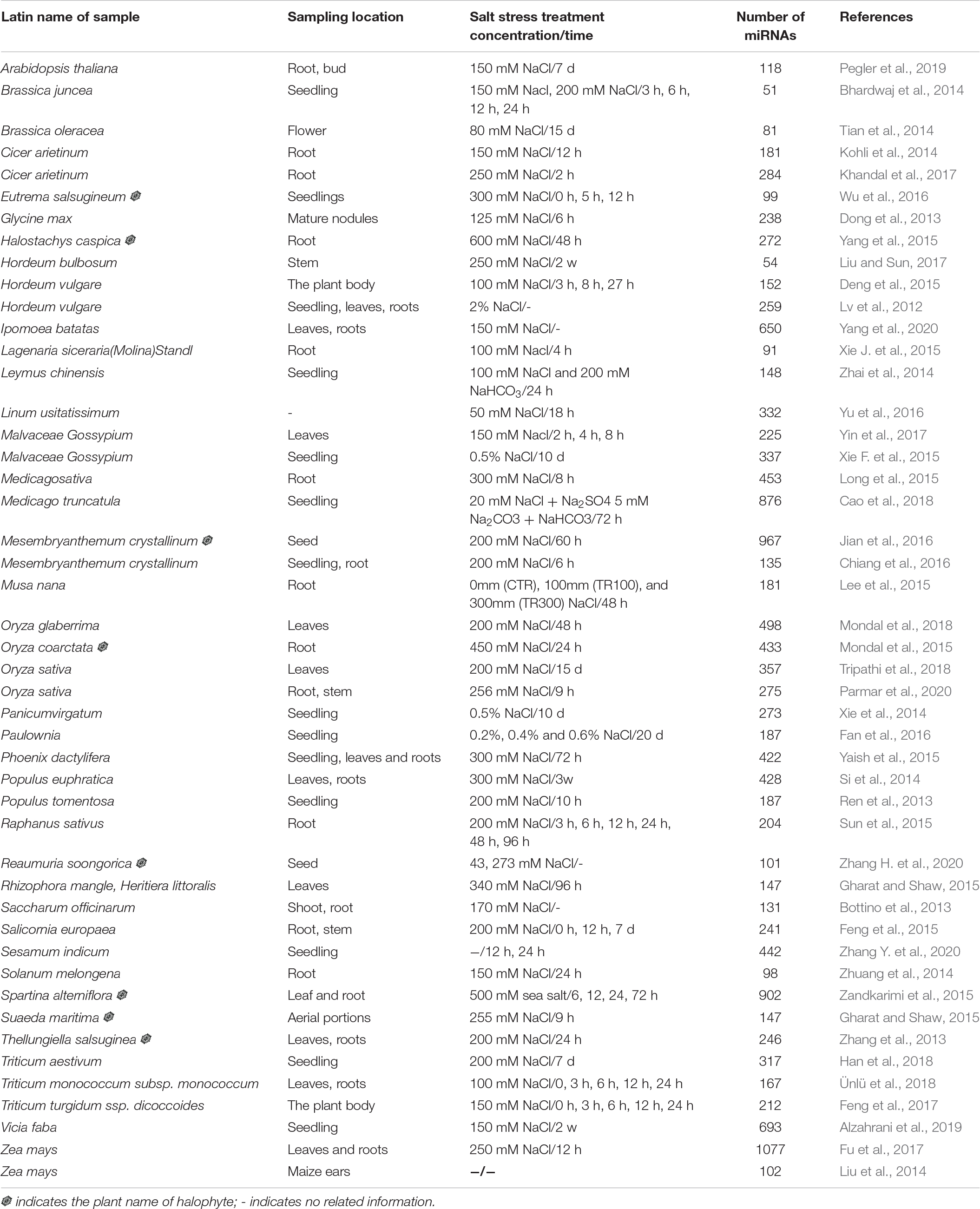
In recent years, with the rapid development of biotechnology, such as microarray and high-throughput deep sequencing, thousands of plant miRNAs were identified under salt stress. As shown in Table 1, different concentrations (80–600 mM) of NaCl and treatment time (3 h to 15 days) were applied for salt stress treatments for identifying salt-responsive miRNA (Table 1). MiRNAs were detected in leaf, root, stem, and flower separately or in the whole seedling (Table 1). Fu et al. identified 1,077 miRNAs in Zea mays, comprising the highest number of identified miRNAs in various crops among the reports (Fu et al., 2017). Moreover, 882, 876, 693, and 650 miRNAs were identified in Mesembryanthemum crystallinum, Medicago truncatula, Vicia faba, and Ipomoea batatas, respectively (Jian et al., 2016; Cao et al., 2018; Alzahrani et al., 2019; Yang et al., 2020). The numbers of identified miRNA vary from dozens to hundreds, which may be due to the plant species, tissue specificity, development stage, and salt stress treatment methods. However, the large-scale identification of miRNAs under salt stress is very necessary and essential, and it lays a solid foundation for the further illumination of the miRNA network.
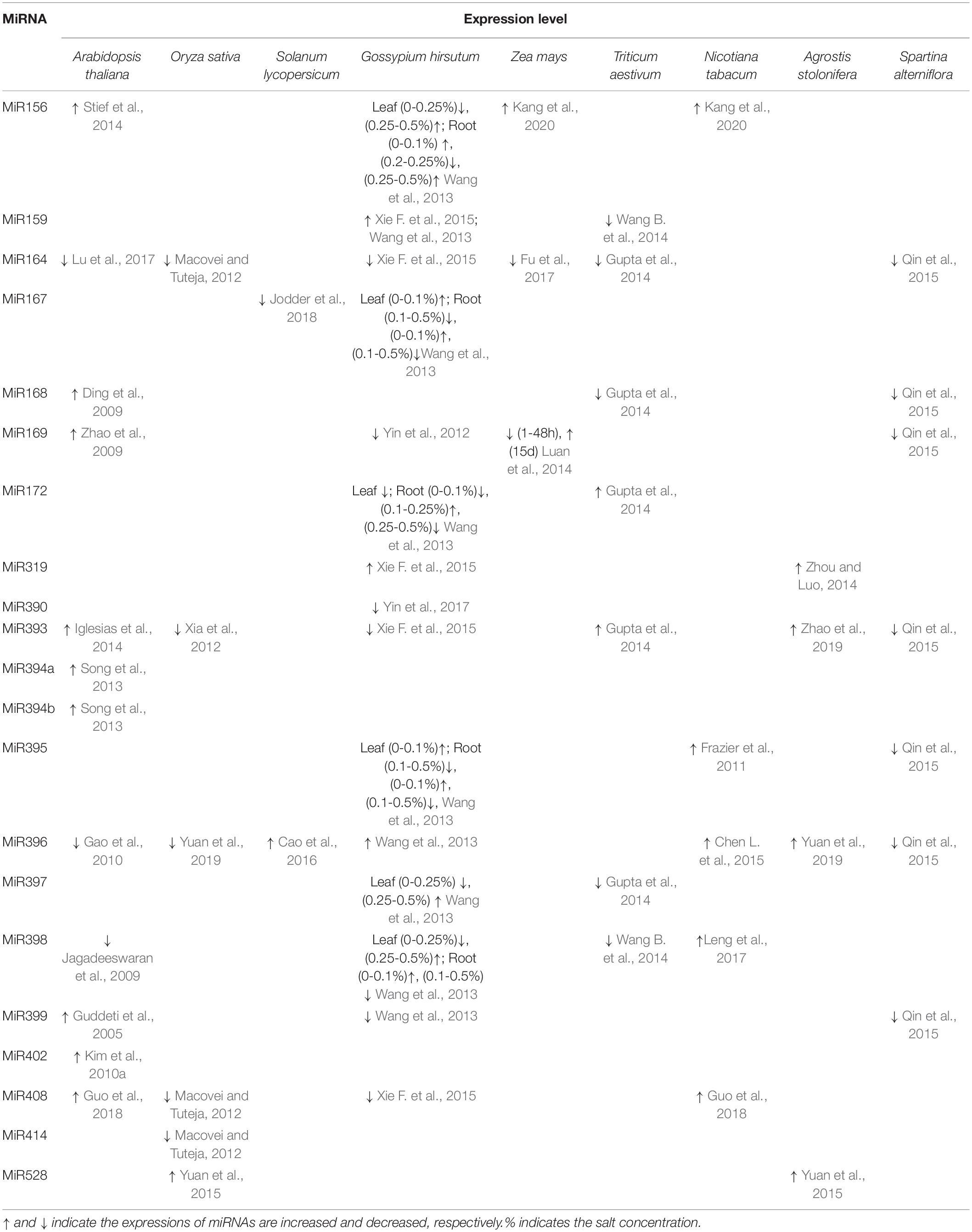
The expression levels of miRNA are up- or down-regulated by salinity stress. For instance, the expression of miR167 in panicle is negatively correlated with the increase of salt concentration (Jodder et al., 2018). In cotton, miR156, miR157, and miR172 are up-regulated at 0.25% NaCl, but their expression decreases with increasing salt concentration (Wang et al., 2013). The expression of miR164 also decreases with the increase of salt stress in maize (Shan et al., 2020). Macovei et al. found that the expression levels of Osa-miR414, -miR164e, and -miR408 significantly decrease with increased salt stress and further regulate the occurrence of genes to resist external salt stress by increasing the content of helicases (Macovei and Tuteja, 2012). In addition, some miRNAs are expressed differently in the early and late stages of salt stress treatment. For example, zma-miR169 displays initial up-regulation and subsequent down-regulation under salt stress (Luan et al., 2015). MiRNAs and their targets, such as cotton miR156-SPL2, miR159-TCP3, miR162-DCL1, miR395-APS1, and miR396-GRF1, exhibit negative correlation on expression levels (Wang et al., 2013).
Table 2 shows the expression levels of some representative miRNAs in plants under salt stress. MiR156, miR319, and miR528 are induced by salinity stress (Wang et al., 2013; Stief et al., 2014; Zhou and Luo, 2014; Xie F. et al., 2015; Yuan et al., 2015), while miR164 and miR397 are repressed (Macovei and Tuteja, 2012; Wang et al., 2013; Gupta et al., 2014; Qin et al., 2015; Xie F. et al., 2015; Lu et al., 2017), which were confirmed at least in two plant species (Table 2). Interestingly, the expression levels of nine miRNAs (e.g., miR159, miR168, miR169, miR172, miR393, miR395, miR396, miR399, and miR408) were promoted in some plant species but were inhibited in the other plant species. For instance, salinity stress increases the expression of miR393 in Arabidopsis thaliana, Triticum aestivum, and Agrostis stolonifera, but decreases the expression of miR393 in Oryza sativa, Gossypium sp., and Spartina alterniflora (Xia et al., 2012; Gupta et al., 2014; Iglesias et al., 2014; Qin et al., 2015; Xie F. et al., 2015; Zhao et al., 2019). Similarly, the expression of miR396 is increased by salinity in Solanum lycopersicum, Nicotiana tabacum, and Agrostis stolonifera but decreased in Arabidopsis thaliana, Oryza sativa, and Spartina alterniflora (Gao et al., 2010; Chen L. et al., 2015; Qin et al., 2015; Cao et al., 2016; Yuan et al., 2019). Up- or down-regulated gene expression usually suggests potential positive or negative functional role. However, the same miRNA has an opposite expression pattern in different plant species under salinity stress conditions, suggesting the same miRNA may play a diverse role in different plant species under salt stress. Moreover, the expression levels of some miRNAs, including miR167, miR390, miR394, miR402, and miR414 were only investigated in very few plant species under salinity stress (Table 2). Considering some miRNAs displayed totally different expressions in different species, their expression patterns need to be investigated in more plant species under salinity stress conditions.
miRNA Studies in Halophyte Palnts
Glycophyte plants, such as Arabidopsis and rice, can only survive at salinity levels 0–100 mM NaCl without any capability to adapt to high salt stress (Horie et al., 2012), whereas some remarkable halophytes can tolerate salinity levels as high as >1000 mM NaCl (Flowers and Colmer, 2008; Munns and Tester, 2008). To an extent, the salt-sensitive glycophytes may not provide enough insights into salt tolerance mechanisms, and the halophytes may have more value for expanding our knowledge about salt resistance mechanisms. Therefore, the exploration of the role of halophyte miRNAs in salinity adaptation can offer compelling contributions for devising strategies of resistance improvement in crops through genetic engineering and plant selection programs. However, there are not many reports on the discovery of salt-responsive miRNAs in halophytes (Table 1).
The halophyte plant Suaeda maritima grows naturally along the seashore. The expression of S. maritima sma-miR2 and sma-miR5 increases under the influence of seawater, suggesting their metabolic regulatory roles specific to saline environments (Gharat and Shaw, 2015). Eutrema salsugineum, a close relative of A. thaliana, can thrive in high salt conditions ranging from 100 to 500 mM (Amtmann, 2009). E. salsugineum has been developed as a valuable model plant for salt stress-tolerance study because its salinity tolerance is extreme, its lifetime is short, its seed production is copious, and its transformation is easy (Zhu, 2000; Amtmann et al., 2005). Zhang et al. (2013) identified 246 miRNAs candidates in E. salsugineum. In addition, 26 conserved miRNAs and 4 novel miRNAs were found to display a significant response to salt stress in E. salsugineum (Zhang et al., 2013; Wu et al., 2016). Recently, 88 conserved miRNAs and 13 novel miRNAs were identified from Reaumuria soongorica seeds treated with various NaCl concentrations, providing a useful reference for salt resistance improvement of seed germination (Zhang H. et al., 2020). A total of 135 conserved miRNAs and the hairpin precursor of 12 novel mcr-miRNAs were found from M. crystallinum seedlings treated with 200 mM NaCl (Chiang et al., 2016). Oryza coarctata is a wild relative of rice and grown in saline water. Mondal et al. found 338 known and 95 novel miRNAs in salt-treated O. coarctata leaves, providing a miRNA-target networking that is involved in salt stress adaption (Mondal et al., 2015). Halostachys caspica (Bieb.), a salt-tolerant short shrub, can be naturally grown on the field with a salt concentration as high as 100 g/kg dry soil (Song et al., 2006). (Yang et al., 2015) found that 31 conserved miRNAs and 12 novel miRNAs were significantly up-regulated, and 48 conserved miRNAs and 13 novel miRNAs were significantly down-regulated by salinity stress in H. caspica. A set of miRNAs were also identified in a salt marsh monocot halophyte smooth cordgrass (Spartina alterniflora Loisel) and another plant named salt cress (Thellungiella salsuginea) (Zhang et al., 2013; Zandkarimi et al., 2015). These identified miRNAs in halophytes can be further projected as potential miRNAs for developing salt tolerance in glycophyte crops.
Functions of miRNA Under Salt Stress
Numerous plant miRNAs have been identified under salt stress, but not many miRNAs have been functionally characterized in detail. Table 3 shows us the miRNAs responsive to salt stress, and these which were functionally studied by transgenetic approaches, such as overexpression and knocked down/out of the miRNA itself or its targets (Table 3). For instance, miR394a/b over-expression and lcr (functional loss of miR394 target LCR) mutant plants are hypersensitive to salt stress, but LCR over-expressing plants display the salt-tolerant phenotype (Song et al., 2013). MiR393 is a comparative well-studied plant miRNA in different plant species, including Arabidopsis, rice, and creeping bentgrass. MiR393ab mutant shows reduced inhibition of LR (lateral root) number and length, increased levels of ROS in LRs, and reduced APX enzymatic activity (Iglesias et al., 2014). Over-expressing Osa-mR393 in rice and Arabidopsis reduces tolerance to salt and drought and increases tillers and early flowering (Gao et al., 2011; Xia et al., 2012), while over-expressing miR393-resistant form mTIR1 in Arabidopsis enhances salt tolerance in mTIR1 transgenic plant (Chen Z. et al., 2015). However, over-expressing Osa-miR393a in creeping bentgrass improves salt stress tolerance associated with the increased uptake of potassium (Zhao et al., 2019), suggesting that the same miRNA or different miRNA from the same miRNA family may have different promotion and inhibition effects on salt tolerance in different plants. A similar situation was found for miRNA396, that is, over-expressing Osa-miR396c reduced salt and alkali stress tolerance in rice and Arabidopsis (Gao et al., 2010), but enhanced salt tolerance associated with improved water retention, increased chlorophyll content, cell membrane integrity, and Na+ exclusion during high salinity exposure in creeping bentgrass (Yuan et al., 2019). Additionally, over-expressing Sp-miR396a-5p in tobacco enhanced its tolerance to salt, drought, and cold stresses (Chen L. et al., 2015). The overexpression of miR395c or miR395e retarded and accelerated, respectively, the seed germination of Arabidopsis under high salt or dehydration stress conditions (Kim et al., 2010b).
Over-expressing miR156a weakens salt resistance in apples, whereas its target gene MdSPL13 strengthens salt resistance (Ma et al., 2020). Transgenic Arabidopsis plants over-expressing the target gene PeNAC070 of miR164 exhibits promoted LR development, delayed stem elongation, and increased sensitivity to salt stress (Lu et al., 2017). Over-expressing the target gene GmNFYA3 of miR169 reduces leaf water loss, enhances drought tolerance, and increases sensitivity to high salinity and exogenous ABA (Ni et al., 2013). Over-expression of miR172c substantially increased the sensitivity of plant roots to salt stress, and the removal of miR172c would decrease the sensitivity of plant roots to salt stress, respectively (Li et al., 2016; Sahito et al., 2017). Osa-miR319a and mi319b positively regulate salt tolerance in creeping bentgrass and swithgrass, respectively (Zhou et al., 2013; Zhou and Luo, 2014; Liu et al., 2019). MiR390 increases LR growth under salt stress via the auxin pathway (He et al., 2018). Additionally, over-expressing miR399f, miR402, and miR408 in Arabidopsis, Tae-miR408 and Sm-MIR408 in tobacco, and Osa-miR528 in creeping bentgrass increases salinity tolerance (Kim et al., 2010a; Feng et al., 2013; Ma et al., 2015; Yuan et al., 2015; Baek et al., 2016; Bai et al., 2018; Guo et al., 2018), indicating that these miRNAs enhance plant salt stress adaptation. By contrast, over-expressing miR414c, miR417, and miRNVL5 increases sensitivity to salinity stress (Jung and Kang, 2007; Gao et al., 2016; Wang et al., 2019). Collectively, these results suggest that the agronomic trait of salinity stress tolerance could be enhanced by the manipulation of miRNA or its target.
Discussion and Future Prospects
In the face of soil salinization, the cultivation of saline-tolerant plants is one of the most economical and effective technologies for biological improvement. Understanding the molecular mechanisms of miRNAs in abiotic stress provides an effective tool for plant breeding, especially in the context of climate and human-induced environmental changes. The essential regulating role of miRNAs in plant salt stress response reveals that miRNA could be applied for salt resistance improvement in crops. The salinity resistance of transgenic plants can be remarkably increased by over-expressing miRNA or knocking down/out the target gene of miRNA. Alternatively, the salinity resistance can be promoted by knocking down/out miRNA, which has a negative effect on salinity response, or over-expressing the target gene of the miRNA. Considering that one miRNA may have more than one targets that would cause totally different effects on plants, we should carefully consider the miRNA effects on crop growth, development, and the sensitivity to other abiotic stresses when optimizing the salinity resistance by miRNA manipulation.
The homologous tetraploid was more tolerant to salt stress than the diploid. Moreover, novel miRNAs induced by genome replication were identified, suggesting salt-responsive miRNAs could be screened by comparative analysis on the plant materials with different ploidy and salinity stress tolerance to explain the key roles of miRNA in achieving better salt stress tolerance. Generally, miRNAs are evolutionarily conserved in their functions in response to salt stress. However, the same miRNAs or different miRNAs from the same miRNA family may have different promotion and inhibition effects on salt tolerance in different plants. Therefore, the function of some miRNAs should be widely studied in different species, especially in crops.
Moreover, considering the significant number of salt- stress-responsive miRNAs identified by using powerful technology (such as high throughput sequencing), only a few miRNAs have been functionally characterized. Therefore, after the identification of plant miRNAs under salinity stress, further studies should be focused on the exploration of function, which will be very crucial for the salt tolerance improvement through miRNA manipulation in crops. Additionally, miRNAs may affect the plant stress tolerance through their interaction with ABA biosynthesis and the regulation of auxin response factors, The investigation of the crosstalk between miRNA and plant hormone will thus expand our knowledge and understanding of the role of plant miRNAs under stress conditions. Finally, the construction of the plant miRNA network in salt stress response will shed light on the salinity resistance improvement through miRNA manipulation in crops.
Author Contributions
TX conceived and designed this manuscript. TX, LZ, and ZY wrote the manuscript. YW and TD helped to revise the manuscript. All authors read and approved the manuscript.
Funding
This work was supported jointly by the projects of the National Natural Science Foundation of China (32072117 and 31701481), the Natural Science Foundation of Jiangsu Higher Education Institutions of China (19KJA510010), and the Key R&D Program of Xuzhou-Modern Agriculture (KC20039).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Abdel Latef, A. A., Abu Alhmad, M. F., Kordrostami, M. F., Abo-Baker, A. E., and Zakir, A. (2020). Inoculation with Azospirillum lipoferum or Azotobacter chroococcum reinforces maize growth by improving physiological activities under saline conditions. J. Plant Growth Regul. 39, 1293–1306. doi: 10.1007/s00344-020-10065-9
Abdel Latef, A. A., Omer, A. M., Badawy, A. A., Osman, M. S., and Ragaey, M. M. (2021). Strategy of salt tolerance and interactive impact of Azotobacter chroococcum and/or Alcaligenes faecalis inoculation on canola (Brassica napus L.). plants grown in saline soil. Plants 10:110. doi: 10.3390/plants10010110
Alzahrani, S. M., Alaraidh, I. A., Khan, M. A., Migdadi, H. M., Alghamdi, S. S., and Alsahli, A. A. (2019). Identification and characterization of salt-responsive microRNAs in Vicia faba by high-throughput sequencing. Genes 10:303. doi: 10.3390/genes10040303
Amtmann, A. (2009). Learning from evolution: thellungiella generates new knowledge on essential and critical components of abiotic stress tolerance in plants. Mol. Plant 2, 3–12. doi: 10.1093/mp/ssn094
Amtmann, A., Bohnert, H. J., and Bressan, R. A. (2005). Abiotic stress and plant genome evolution. Search for new models. Plant Physiol. 138, 127–130. doi: 10.1104/pp.105.059972
Attia, M. S., Osman, M. S., Mohamed, A. S., Mahgoub, H. A., Garada, M. O., Abdelmouty, E. S., et al. (2021). Impact of foliar application of chitosan dissolved in different organic acids on isozymes, protein patterns and physio-biochemical characteristics of tomato grown under salinity stress. Plants 10:388. doi: 10.3390/plants10020388
Baek, D., Chun, H. J., Kang, S., Shin, G., Park, S. J., Hong, H., et al. (2016). A Role for Arabidopsis miR399f in salt, drought, and ABA signaling. Mol. Cells 39, 111–118. doi: 10.14348/molcells.2016.2188
Bai, Q., Wang, X., Chen, X., Shi, G., Liu, Z., Guo, C., et al. (2018). Wheat miRNA TaemiR408 acts as an essential mediator in plant tolerance to Pi deprivation and salt stress via modulating stress-associated physiological processes. Front. Plant Sci. 9:499. doi: 10.3389/fpls.2018.00499
Bhardwaj, A. R., Joshi, G., Pandey, R., Kukreja, B., Goel, S., Jagannath, A., et al. (2014). A genome-wide perspective of miRNAome in response to high temperature, salinity and drought stresses in Brassica juncea (Czern) L. PLoS One 9:e92456. doi: 10.3389/fpls.2018.00499
Bielewicz, D., Kalak, M., Kalyna, M., Windels, D., Barta, A., Vazquez, F., et al. (2013). Introns of plant pri-miRNAs enhance miRNA biogenesis. EMBO Rep. 14, 622–628. doi: 10.1038/embor.2013.62
Bottino, M. C., Rosario, S., Grativol, C., Thiebaut, F., Rojas, C. A., Farrineli, L., et al. (2013). High-throughput sequencing of small RNA transcriptome reveals salt stress regulated microRNAs in sugarcane. PLoS One 8:e59423. doi: 10.1371/journal.pone.0059423
Brodersen, P., Sakvarelidze-Achard, L., Bruun-Rasmussen, M., Dunoyer, P., Yamamoto, Y. Y., Sieburth, L., et al. (2008). Widespread translational inhibition by plant miRNAs and siRNAs. Science 30 320, 1185–1190. doi: 10.1126/science.1159151
Budak, H., and Akpinar, B. A. (2015). Plant miRNAs: biogenesis, organization and origins. Funct. Integr. Genomics 15, 523–531. doi: 10.1007/s10142-015-0451-2
Butcher, K., Wick, A. F., Desutter, T., Chatterjee, A., and Harmon, J. (2016). Soil salinity: a threat to global food security. Agron. J. 108, 2189–2200. doi: 10.2134/agronj2016.06.0368
Cao, C., Long, R., Zhang, T., Kang, J., Wang, Z., Wang, P., et al. (2018). Genome-wide identification of microRNAs in response to salt/alkali stress in Medicago truncatula through high-throughput sequencing. Int. J. Mol. Sci. 19:4076. doi: 10.3390/ijms19124076
Cao, D., Wang, J., Ju, Z., Liu, Q., Li, S., Tian, H., et al. (2016). Regulations on growth and development in tomato cotyledon, flower and fruit via destruction of miR396 with short tandem target mimic. Plant Sci. 247, 1–12. doi: 10.1016/j.plantsci.2016.02.012
Chandran, V., Wang, H., Gao, F., Cao, X. L., Chen, Y. P., Li, G. B., et al. (2019). MiR396-OsGRFs module balances growth and rice blast disease-resistance. Front. Plant Sci. 9:1999. doi: 10.3389/fpls.2018.01999
Chen, L., Luan, Y., and Zhai, J. (2015). Sp-miR396a-5p acts as a stress-responsive genes regulator by conferring tolerance to abiotic stresses and susceptibility to Phytophthora nicotianae infection in transgenic tobacco. Plant Cell Rep. 34, 2013–2025. doi: 10.1007/s00299-015-1847-0
Chen, X. (2009). Small RNAs and their roles in plant development. Annu. Rev. Cell Dev. Biol. 25, 21–44. doi: 10.1146/annurev.cellbio.042308.113417
Chen, Z., Hu, L., Han, N., Hu, J., Yang, Y., Xiang, T., et al. (2015). Overexpression of a miR393-resistant form of transport inhibitor response protein 1 (mTIR1) enhances salt tolerance by increased osmoregulation and Na+ exclusion in Arabidopsis thaliana. Plant Cell Physiol. 56, 73–83. doi: 10.1093/pcp/pcu149
Chiang, C. P., Yim, W. C., Sun, Y. H., Ohnishi, M., Mimura, T., Cushman, J. C., et al. (2016). Identification of ice plant (Mesembryanthemum crystallinum L.) microRNAs using RNA-seq and their putative roles in high salinity responses in seedlings. Front. Plant Sci. 7:1143. doi: 10.3389/fpls.2016.01143
Deng, P., Wang, L., Cui, L., Feng, K., Liu, F., Du, X., et al. (2015). Global identification of microRNAs and their targets in barley under salinity stress. PLoS One 10:e0137990. doi: 10.1371/journal.pone.0137990
Ding, D., Zhang, L., Wang, H., Liu, Z., Zhang, Z., and Zheng, Y. (2009). Differential expression of miRNAs in response to salt stress in maize roots. Ann. Bot. 103, 29–38. doi: 10.1093/aob/mcn205
Dong, Z., Shi, L., Wang, Y., Chen, L., Cai, Z., Wang, Y., et al. (2013). Identification and dynamic regulation of microRNAs involved in salt stress responses in functional soybean nodules by high-throughput sequencing. Int. J. Mol. Sci. 14, 2717–2738. doi: 10.3390/ijms14022717
Du, Q. G., Wang, K., Zou, C., Xu, C., and Li, W. X. (2018). The PILNCR1-miR399 regulatory module is important for low phosphate tolerance in maize. Plant Physiol. 177, 1743–1753. doi: 10.1104/pp.18.00034
Fan, G., Li, X., Deng, M., Zhao, Z., and Yang, L. (2016). Comparative analysis and identification of miRNAs and their target genes responsive to salt stress in diploid and tetraploid Paulownia fortunei seedlings. PLoS One 11:e0149617. doi: 10.1371/journal.pone.0149617
FAO (2017). The Future of Food and Agriculture - Trends and Challenges. Rome: Food and Agriculture Organization of the United Nations.
Feng, H., Zhang, Q., Wang, Q., Wang, X., Liu, J., Li, M., et al. (2013). Target of tae-miR408, a chemocyanin-like protein gene (TaCLP1), plays positive roles in wheat response to high-salinity, heavy cupric stress and stripe rust. Plant Mol. Biol. 83, 433–443. doi: 10.1007/s11103-013-0101-9
Feng, J., Wang, J., Fan, P., Jia, W., Nie, L., Jiang, P., et al. (2015). High-throughput deep sequencing reveals that microRNAs play important roles in salt tolerance of euhalophyte Salicornia europaea. BMC Plant Biol. 15:63. doi: 10.1186/s12870-015-0451-3
Feng, K., Nie, X., Cui, L., Deng, P., Wang, M., and Song, W. (2017). Genome-Wide identification and characterization of salinity stress-responsive miRNAs in wild emmer wheat (Triticum turgidum ssp. dicoccoides). Genes 8:156. doi: 10.3390/genes8060156
Flowers, T. J., and Colmer, T. D. (2008). Salinity tolerance in halophytes. New Phytol. 179, 945–963. doi: 10.1111/j.1469-8137.2008.02531.x
Frazier, T. P., Sun, G., Burklew, E., and Zhang, B. (2011). Salt and drought stresses induce the aberrant expression of microRNA genes in tobacco. Mol. Biotechnol. 49, 159–165. doi: 10.1007/s12033-011-9387-5
Fu, R., Zhang, M., Zhao, Y., He, X., Ding, C., Wang, S., et al. (2017). Identification of salt tolerance-related microRNAs and their targets in maize (Zea mays L.) using high-throughput sequencing and degradome analysis. Front. Plant Sci. 8:864. doi: 10.3389/fpls.2017.00864
Gao, P., Bai, X., Yang, L., Lv, D., Li, Y., Cai, H., et al. (2010). Over-expression of osa-MIR396c decreases salt and alkali stress tolerance. Planta 231, 991–1001. doi: 10.1007/s00425-010-1104-2
Gao, P., Bai, X., Yang, L., Lv, D., Pan, X., Li, Y., et al. (2011). Osa-MIR393: a salinity- and alkaline stress-related microRNA gene. Mol. Biol. Rep. 38, 237–242. doi: 10.1007/s11033-010-0100-8
Gao, S., Yang, L., Zeng, H. Q., Zhou, Z. S., Yang, Z. M., Li, H., et al. (2016). A cotton miRNA is involved in regulation of plant response to salt stress. Sci. Rep. 6:19736. doi: 10.1038/srep19736
Gharat, S. A., and Shaw, B. P. (2015). Novel and conserved miRNAs in the halophyte Suaeda maritima identified by deep sequencing and computational predictions using the ESTs of two mangrove plants. BMC Plant Biol. 15:301. doi: 10.1186/s12870-015-0682-3
Gramzow, L., and Theißen, G. (2019). Plant miRNA conservation and evolution. Methods Mol. Biol. 1932, 41–50. doi: 10.1007/978-1-4939-9042-9_3
Guddeti, S., Zhang, D. C., Li, A. L., Leseberg, C. H., Kang, H., Li, X. G., et al. (2005). Molecular evolution of the rice miR395 gene family. Cell Res. 15, 631–638. doi: 10.1038/sj.cr.7290333
Guo, X., Niu, J., and Cao, X. (2018). Heterologous expression of salvia miltiorrhiza microRNA408 enhances tolerance to salt stress in Nicotiana benthamiana. Int. J. Mol. Sci. 19:3985. doi: 10.3390/ijms19123985
Gupta, O. P., Meena, N. L., Sharma, I., and Sharma, P. (2014). Differential regulation of microRNAs in response to osmotic, salt and cold stresses in wheat. Mol. Biol. Rep. 41, 4623–4629. doi: 10.1007/s11033-014-3333-0
Haas, U., Sczakiel, G., and Laufer, S. D. (2012). MicroRNA-mediated regulation of gene expression is affected by disease-associated SNPs within the 3′-UTR via altered RNA structure. RNA Biol. 9, 924–937. doi: 10.4161/rna.20497
Hackenberg, M., Shi, B. J., Gustafson, P., and Langridge, P. (2013). Characterization of phosphorus-regulated miR399 and miR827 and their isomirs in barley under phosphorus-sufficient and phosphorus-deficient conditions. BMC Plant Biol. 13:214. doi: 10.1186/1471-2229-13-214
Han, H., Wang, Q., Wei, L., Liang, Y., Dai, J., Xia, G., et al. (2018). Small RNA and degradome sequencing used to elucidate the basis of tolerance to salinity and alkalinity in wheat. BMC Plant Biol. 18:195. doi: 10.1186/s12870-018-1415-1
He, F., Xu, C., Fu, X., Shen, Y., Guo, L., Leng, M., et al. (2018). The microRNA390/trans- acting short interfering RNA3 module mediates lateral root growth under salt stress via the auxin pathway. Plant Physiol. 177, 775–791. doi: 10.1104/pp.17.01559
Horie, T., Karahara, I., and Katsuhara, M. (2012). Salinity tolerance mechanisms in glycophytes: an overview with the central focus on rice plants. Rice 5:11. doi: 10.1186/1939-8433-5-11
Iglesias, M. J., Terrile, M. C., Windels, D., Lombardo, M. C., Bartoli, C. G., Vazquez, F., et al. (2014). MiR393 regulation of auxin signaling and redox-related components during acclimation to salinity in Arabidopsis. PLoS One 9:e107678. doi: 10.1371/journal.pone.0107678
Jagadeeswaran, G., Saini, A., and Sunkar, R. (2009). Biotic and abiotic stress down-regulate miR398 expression in Arabidopsis. Planta 229, 1009–1014. doi: 10.1007/s00425-009-0889-3
Jeyaraj, A., Chandran, V., and Gajjeraman, P. (2014). Differential expression of microRNAs in dormant bud of tea [Camellia sinensis (L.) O. Kuntze]. Plant Cell Rep. 33, 1053–1069. doi: 10.1007/s00299-014-1589-4
Jian, H., Wang, J., Wang, T., Wei, L., Li, J., and Liu, L. (2016). Identification of rapeseed microRNAs involved in early stage seed germination under salt and drought stresses. Front. Plant Sci. 7:658. doi: 10.3389/fpls.2016.00658
Jodder, J., Das, R., Sarkar, D., Bhattacharjee, P., and Kundu, P. (2018). Distinct transcriptional and processing regulations control miR167a level in tomato during stress. RNA Biol. 15, 130–143. doi: 10.1080/15476286.2017.1391438
Jones-Rhoades, M. W., Bartel, D. P., and Bartel, B. (2006). MicroRNAS and their regulatory roles in plants. Annu. Rev. Plant Biol. 57, 19–53. doi: 10.1146/annurev.arplant.57.032905.105218
Jung, H. J., and Kang, H. (2007). Expression and functional analyses of microRNA417 in Arabidopsis thaliana under stress conditions. Plant Physiol. Biochem. 45, 805–811. doi: 10.1016/j.plaphy.2007.07.015
Kang, T., Yu, C. Y., Liu, Y., Song, W. M., Bao, Y., Guo, X. T., et al. (2020). Subtly manipulated expression of zmmiR156 in tobacco improves drought and salt tolerance without changing the architecture of transgenic plants. Front. Plant Sci. 10:1664. doi: 10.3389/fpls.2019.01664
Kawashima, C. G., Yoshimoto, N., Maruyama-Nakashita, A., Tsuchiya, Y. N., Saito, K., Takahashi, H., et al. (2009). Sulphur starvation induces the expression of microRNA-395 and one of its target genes but in different cell types. Plant J. 57, 313–321. doi: 10.1111/j.1365-313X.2008.03690.x
Khandal, H., Parween, S., Roy, R., Meena, M. K., and Chattopadhyay, D. (2017). MicroRNA profiling provides insights into post-transcriptional regulation of gene expression in chickpea root apex under salinity and water deficiency. Sci. Rep. 7:4632. doi: 10.1038/s41598-017-04906-z
Kim, J. Y., Kwak, K. J., Jung, H. J., Lee, H. J., and Kang, H. (2010a). MicroRNA402 affects seed germination of Arabidopsis thaliana under stress conditions via targeting demeter- like protein3 mRNA. Plant Cell Physiol. 51, 1079–1083. doi: 10.1093/pcp/pcq072
Kim, J. Y., Lee, H. J., Jung, H. J., Maruyama, K., Suzuki, N., and Kang, H. (2010b). Overexpression of microRNA395c or 395e affects differently the seed germination of Arabidopsis thaliana under stress conditions. Planta 232, 1447–1454. doi: 10.1007/s00425-010-1267-x
Kim, W., Ahn, H. J., Chiou, T. J., and Ahn, J. H. (2011). The role of the miR399-PHO2 module in the regulation of flowering time in response to different ambient temperatures in Arabidopsis thaliana. Mol. Cells 32, 83–88. doi: 10.1007/s10059-011-1043-1
Kohli, D., Joshi, G., Deokar, A. A., Bhardwaj, A. R., Agarwal, M., Katiyar-Agarwal, S., et al. (2014). Identification and characterization of Wilt and salt stress-responsive microRNAs in chickpea through high-throughput sequencing. PLoS One 9:e108851. doi: 10.1371/journal.pone
Koroban, N. V., Kudryavtseva, A. V., Krasnov, G. S., Sadritdinova, A. F., Fedorova, M. S., Snezhkina, A. V., et al. (2016). The role of microRNA in abiotic stress response in plants. Mol. Biol. 50, 387–394. doi: 10.7868/S0026898416020105
Lee, R. C., and Ambros, V. (2001). An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862–864. doi: 10.1126/science.1065329
Lee, W. S., Gudimella, R., Wong, G. R., Tammi, M. T., Khalid, N., and Harikrishna, J. A. (2015). Transcripts and microRNAs responding to salt stress in musa acuminata colla (AAA Group) cv. berangan roots. PLoS One 10:e0127526. doi: 10.1371/journal.pone.0127526
Leng, X., Wang, P., Zhu, X., Li, X., Zheng, T., Shangguan, L., et al. (2017). Ectopic expression of CSD1 and CSD2 targeting genes of miR398 in grapevine is associated with oxidative stress tolerance. Funct. Integr. Genomics 17, 697–710. doi: 10.1007/s10142-017-0565-9
Li, W., Wang, T., Zhang, Y., and Li, Y. (2016). Overexpression of soybean miR172c confers tolerance to water deficit and salt stress, but increases ABA sensitivity in transgenic Arabidopsis thaliana. J. Exp. Bot. 67, 175–194. doi: 10.1093/jxb/erw404
Lian, C. L., Yao, K., Duan, H., Li, Q., Liu, C., Yin, W. L., et al. (2018). Exploration of ABA responsive miRNAs reveals a new hormone signaling crosstalk pathway regulating root growth of populus euphratica. Int. J. Mol. Sci. 19:1481. doi: 10.3390/ijms19051481
Liu, B., and Sun, G. (2017). MicroRNAs contribute to enhanced salt adaptation of the autopolyploid Hordeum bulbosum compared with its diploid ancestor. Plant J. 91(1):57–69. doi: 10.1111/tpj.13546
Liu, H., Qin, C., Chen, Z., Zuo, T., Yang, X., Zhou, H., et al. (2014). Identification of miRNAs and their target genes in developing maize ears by combined small RNA and degradome sequencing. BMC Genomics 15:25. doi: 10.1186/1471-2164-15-25
Liu, Y., Li, D., Yan, J., Wang, K., Luo, H., and Zhang, W. (2019). MiR319 mediated salt tolerance by ethylene. Plant Biotechnol. J. 17, 2370–2383. doi: 10.1111/pbi.13154
Llave, C., Kasschau, K. D., Rector, M. A., and Carrington, J. C. (2002). Endogenous and silencing-associated small RNAs in plants. Plant Cell 14, 1605–1619. doi: 10.1105/tpc.003210
Long, R. C., Li, M. N., Kang, J. M., Zhang, T. J., Sun, Y., and Yang, Q. C. (2015). Small RNA deep sequencing identifies novel and salt-stress-regulated microRNAs from roots of Medicago sativa and Medicago truncatula. Physiol. Plant 154, 13–27. doi: 10.1111/ppl.12266
Lu, X., Dun, H., Lian, C., Zhang, X., Yin, W., and Xia, X. (2017). The role of peu-miR164 and its target PeNAC genes in response to abiotic stress in Populus euphratica. Plant Physiol. Biochem. 115, 418–438. doi: 10.1016/j.plaphy.2017.04.009
Luan, M., Xu, M., Lu, Y., Zhang, L., Fan, Y., and Wang, L. (2015). Expression of zma-miR169 miRNAs and their target ZmNF-YA genes in response to abiotic stress in maize leaves. Gene 55, 178–185. doi: 10.1016/j.gene.2014.11.001
Luan, M., Xu, M., Lu, Y., Zhang, Q., Zhang, L., Zhang, C., et al. (2014). Family-wide survey of miR169s and NF-YAs and their expression profiles response to abiotic stress in maize roots. PLoS One 9:e91369. doi: 10.1371/journal.pone.0091369
Lv, S., Nie, X., Wang, L., Du, X., Biradar, S. S., Jia, X., et al. (2012). Identification and characterization of microRNAs from barley (Hordeum vulgare L.) by high-throughput sequencing. Int. J. Mol. Sci. 13, 2973–2984. doi: 10.3390/ijms13032973
Ma, C., Burd, S., and Lers, A. (2015). MiR408 is involved in abiotic stress responses in Arabidopsis. Plant J. 84, 169–187. doi: 10.1111/tpj.12999
Ma, Y., Xue, H., Zhang, F., Jiang, Q., Yang, S., Yue, P., et al. (2020). The miR156/SPL module regulates apple salt stress tolerance by activating MdWRKY100 expression. Plant Biotechnol. J. 9, 311–323. doi: 10.1111/pbi.13464
Macovei, A., and Tuteja, N. (2012). MicroRNAs targeting DEAD-box helicases are involved in salinity stress response in rice (Oryza sativa L.). BMC Plant Biol. 12:183. doi: 10.1186/1471-2229-12-183
Mekonnen, M. M., and Hoekstra, A. Y. (2016). Four billion people facing severe water scarcity. Sci. Adv. 2:e1500323. doi: 10.1126/sciadv.1500323
Meng, Y., Mao, J. P., Tahir, M. M., Wang, H., Wei, Y. H., Zhao, C. D., et al. (2020). Mdm-miR160 participates in auxin-induced adventitious root formation of apple rootstock. Sci. Horticuamsterdam 270:109442.
Mette, M. F., van der Winden, J., Matzke, M., and Matzke, A. J. (2002). Short RNAs can identify new candidate transposable element families in Arabidopsis. Plant Physiol. 130, 6–9. doi: 10.1104/pp.007047
Mondal, T. K., Ganie, S. A., and Debnath, A. B. (2015). Identification of novel and conserved miRNAs from extreme halophyte, Oryza coarctata, a wild relative of rice. PLoS One 10:e0140675. doi: 10.1371/journal.pone.0140675
Mondal, T. K., Panda, A. K., Rawal, H. C., and Sharma, T. R. (2018). Discovery of microRNA-target modules of African rice (Oryza glaberrima) under salinity stress. Sci. Rep. 8:570. doi: 10.1038/s41598-017-18206-z
Morton, M. J. L., Awlia, M., Al-Tamimi, N., Saade, S., Pailles, Y., Negrão, S., et al. (2019). Salt stress under the scalpel-dissecting the genetics of salt tolerance. Plant J. 97, 148–163. doi: 10.1111/tpj.14189
Munns, R., and Tester, M. (2008). Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 59, 651–681. doi: 10.1146/annurev.arplant.59.032607.092911
Ni, Z., Hu, Z., Jiang, Q., and Zhang, H. (2013). GmNFYA3, a target gene of miR169, is a positive regulator of plant tolerance to drought stress. Plant Mol. Biol. 82, 113–129. doi: 10.1007/s11103-013-0040-5
Osman, M. S., Badawy, A. A., Osman, A. I., and Abdel Latef, A. A. (2020). Ameliorative impact of an extract of the halophyte Arthrocnemum macrostachyum on growth and biochemical parameters of soybean under salinity stress. J. Plant Growth Regul. 1–12. doi: 10.1007/s00344-020-10185-2
Park, W., Li, J. J., Song, R. T., Messing, J., and Chen, X. M. (2002). Carpel factory, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr. Biol. 12, 1484–1495. doi: 10.1016/s0960-9822(02)01017-5
Parmar, S., Gharat, S. A., Tagirasa, R., Chandra, T., Behera, L., Dash, S. K., et al. (2020). Identification and expression analysis of miRNAs and elucidation of their role in salt tolerance in rice varieties susceptible and tolerant to salinity. PLoS One 15:e0230958. doi: 10.1371/journal.pone.0230958
Pegler, J. L., Oultram, J. M. J., Grof, C. P. L., and Eamens, A. L. (2019). Profiling the abiotic stress responsive microRNA landscape of Arabidopsis thaliana. Plants 8:58. doi: 10.3390/plants8030058
Qadir, M., Quillerou, E., Nanjia, V., Murtaza, G., Singh, M., Thomas, R. J., et al. (2014). Economics of salt-induced land degradation and restoration. Nat. Resour. Forum. 38:282–295. doi: 10.1111/1477-8947.12054
Qin, Z., Chen, J., Jin, L., Duns, G. J., and Ouyang, P. (2015). Differential expression of miRNAs under salt stress in spartina alterniflora leaf tissues. J. Nanosci. Nanotechnol. 15, 1554–1561. doi: 10.1166/jnn.2015.9004
Rao, S., Balyan, S., Jha, S., and Mathur, S. (2020). Novel insights into expansion and functional diversification of MIR169 family in tomato. Planta 251:55. doi: 10.1007/s00425-020-03346-w
Reinhart, B. J., Weinstein, E. G., Rhoades, M. W., Bartel, B., and Bartel, D. P. (2002). MicroRNAs in plants. Genes Dev. 16, 1616–1626. doi: 10.1101/gad.1004402
Ren, Y., Chen, L., Zhang, Y., Kang, X., Zhang, Z., and Wang, Y. (2013). Identification and characterization of salt-responsive microRNAs in Populus tomentosa by high-throughput sequencing. Biochimie 95, 743–750. doi: 10.1016/j.biochi.2012.10.025
Sahito, Z. A., Wang, L., Sun, Z., Yan, Q., Zhang, X., Jiang, Q., et al. (2017). The miR172c-NNC1 module modulates root plastic development in response to salt in soybean. BMC Plant Biol. 17:229. doi: 10.1186/s12870-017-1161-9
Shan, T., Fu, R., Xie, Y., Chen, Q., Wang, Y., Li, P., et al. (2020). Regulatory mechanism of maize (Zea mays L.) miR164 in salt stress response. Russ. J. Genet. 56, 835–842.
Si, J., Zhou, T., Bo, W., Xu, F., and Wu, R. (2014). Genome-wide analysis of salt-responsive and novel microRNAs in Populus euphratica by deep sequencing. BMC Genet. 15, (Suppl. 1):S6. doi: 10.1186/1471-2156-15-S1-S6
Song, J., Feng, G., and Zhang, F. S. (2006). Salinity and temperature effects on germination for three salt-resistant euhalophytes, Halostachys caspica, Kalidium foliatum and Halocnemum strobilaceum. Plant Soil 279, 201–207. doi: 10.1007/s11104-005-1012-6
Song, J. B., Gao, S., Sun, D., Li, H., Shu, X. X., and Yang, Z. M. (2013). MiR394 and LCR are involved in Arabidopsis salt and drought stress responses in an abscisic acid-dependent manner. BMC Plant Biol. 13:210. doi: 10.1186/1471-2229-13-210
Sorin, C., Declerck, M., Christ, A., Blein, T., Ma, L., Lelandais-Brière, C., et al. (2014). A miR169 isoform regulates specific NF-YA targets and root architecture in Arabidopsis. New Phytol. 202, 1197–1211. doi: 10.1111/nph.12735
Stief, A., Altmann, S., Hoffmann, K., Pant, B. D., Scheible, W. R., and Baurle, I. (2014). Arabidopsis mir156 regulates tolerance to recurring environmental stress through SPL transcription factors. Plant Cell 26, 1792–1807. doi: 10.1105/tpc.114.123851
Sun, X., Xu, L., Wang, Y., Yu, R., Zhu, X., Luo, X., et al. (2015). Identification of novel and salt-responsive miRNAs to explore miRNA-mediated regulatory network of salt stress response in radish (Raphanus sativus L.). BMC Genomics 16:197. doi: 10.1186/s12864-015-1416-5
Tian, Y., Tian, Y., Luo, X., Zhou, T., Huang, Z., Liu, Y., et al. (2014). Identification and characterization of microRNAs related to salt stress in broccoli, using high-throughput sequencing and bioinformatics analysis. BMC Plant Biol. 14:226. doi: 10.1186/s12870-014-0226-2
Tripathi, A., Chacon, O., Singla-Pareek, S. L., Sopory, S. K., and Sanan-Mishra, N. (2018). Mapping the microRNA expression profiles in glyoxalase over-expressing salinity tolerant rice. Curr. Genomics 19, 21–35. doi: 10.2174/1389202918666170228134530
Ünlü, E. S., Bataw, S., Aslan, Ş.D., Şahin, Y., and Zencirci, N. (2018). Identification of conserved miRNA molecules in einkorn wheat (Triticum monococcum subsp. monococcum) by using small RNA sequencing analysis. Turk. J. Biol. 42, 527–536. doi: 10.3906/biy-1802-3
Wang, B., Sun, Y. F., Song, N., Wei, J. P., Wang, X. J., Feng, H., et al. (2014). MicroRNAs involving in cold, wounding and salt stresses in Triticum aestivum L. Plant Physiol. Biochem. 80, 90–96. doi: 10.1016/j.plaphy.2014.03.020
Wang, M., Wang, Q., and Zhang, B. (2013). Response of miRNAs and their targets to salt and drought stresses in cotton (Gossypium hirsutum L.). Gene 530, 26–32. doi: 10.1016/j.gene.2013.08.009
Wang, Q., Wei, L., Guan, X., Wu, Y., Zou, Q., and Ji, Z. (2014). Briefing in family characteristics of microRNAs and their applications in cancer research. Biochim. Biophys. Acta 1844(1 Pt B), 191–197. doi: 10.1016/j.bbapap.2013.08.002
Wang, W., Liu, D., Chen, D., Cheng, Y., Zhang, X., Song, L., et al. (2019). MicroRNA414c affects salt tolerance of cotton by regulating reactive oxygen species metabolism under salinity stress. RNA Biol. 16, 362–375. doi: 10.1080/15476286.2019.1574163
Wang, Y. M., Liu, W. W., Wang, X. W., Yang, R. J., Wu, Z. Y., Wang, H., et al. (2020). MiR156 regulates anthocyanin biosynthesis through SPL targets and other microRNAs in poplar. Hortic. Res. 7:118. doi: 10.1038/s41438-020-00341-w
Wen, F. L., Yue, Y., He, T. F., Gao, X. M., Zhou, Z. S., and Long, X. H. (2020). Identification of miR390-TAS3-ARF pathway in response to salt stress in Helianthus tuberosus L. Gene 738:144460. doi: 10.1016/j.gene.2020.144460
Wu, J. G., Yang, R. X., Yang, Z. R., Yao, S., Zhao, S. S., Wang, Y., et al. (2017). ROS accumulation and antiviral defence control by microRNA528 in rice. Nat. Plants 3:16203. doi: 10.1038/nplants.2016.203
Wu, Y., Guo, J., Cai, Y., Gong, X., Xiong, X., Qi, W., et al. (2016). Genome-wide identification and characterization of Eutrema salsugineum microRNAs for salt tolerance. Physiol. Plant 157, 453–468. doi: 10.1111/ppl.12419
Xia, K., Wang, R., Ou, X., Fang, Z., Tian, C., Duan, J., et al. (2012). OsTIR1 and OsAFB2 downregulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS One 7:e30039. doi: 10.1371/journal.pone.0030039
Xie, F., Stewart, C. N. Jr., Taki, F. A., He, Q., Liu, H., and Zhang, B. (2014). High-throughput deep sequencing shows that microRNAs play important roles in switchgrass responses to drought and salinity stress. Plant Biotechnol. J. 12, 354–366. doi: 10.1111/pbi.12142
Xie, F., Wang, Q., Sun, R., and Zhang, B. (2015). Deep sequencing reveals important roles of microRNAs in response to drought and salinity stress in cotton. J. Exp. Bot. 66, 789–804. doi: 10.1093/jxb/eru437
Xie, J., Lei, B., Niu, M., Huang, Y., Kong, Q., and Bie, Z. (2015). High throughput sequencing of small RNAs in the two cucurbita germplasm with different sodium accumulation patterns identifies novel microRNAs involved in salt stress response. PLoS One 10:e0127412. doi: 10.1371/journal.pone.0127412
Xu, M. L., Hu, T. Q., Zhao, J. F., Park, M. Y., Earley, K. W., Wu, G., et al. (2016). Developmental functions of miR156-regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) genes in Arabidopsis thaliana. PLoS Genet. 12:e1006263. doi: 10.1371/journal.pgen.1006263
Yaish, M. W., Sunkar, R., Zheng, Y., Ji, B., Al-Yahyai, R., and Farooq, S. A. (2015). A genome-wide identification of the miRNAome in response to salinity stress in date palm (Phoenix dactylifera L.). Front. Plant Sci. 6:946. doi: 10.3389/fpls.2015.00946
Yang, C. H., Li, D. Y., Mao, D. H., Liu, X., Ji, C. J., Li, X. B., et al. (2013). Overexpression of microRNA319 impacts leaf morphogenesis and leads to enhanced cold tolerance in rice (Oryza sativa L.). Plant Cell Environ. 36, 2207–2218. doi: 10.1111/pce.12130
Yang, R., Zeng, Y., Yi, X., Zhao, L., and Zhang, Y. (2015). Small RNA deep sequencing reveals the important role of microRNAs in the halophyte Halostachys caspica. Plant Biotechnol. J. 13, 395–408. doi: 10.1111/pbi.12337
Yang, Z., Zhu, P., Kang, H., Liu, L., Cao, Q., Sun, J., et al. (2020). High-throughput deep sequencing reveals the important role that microRNAs play in the salt response in sweet potato (Ipomoea batatas L.). BMC Genomics 21:164. doi: 10.1186/s12864-020-6567-3
Yin, Z., Han, X., Li, Y., Wang, J., Wang, D., Wang, S., et al. (2017). Comparative analysis of cotton small RNAs and their target genes in response to salt stress. Genes 8:369. doi: 10.3390/genes8120369
Yin, Z., Li, Y., Yu, J., Liu, Y., Li, C., Han, X., et al. (2012). Difference in miRNA expression profiles between two cotton cultivars with distinct salt sensitivity. Mol. Biol. Rep. 39, 4961–4970. doi: 10.1007/s11033-011-1292-2
Yu, X. Y., Gong, H. Y., Cao, L. F., Hou, Y. J., and Qu, S. C. (2020). MicroRNA397b negatively regulates resistance of Malus hupehensis to Botryosphaeria dothidea by modulating MhLAC7 involved in lignin biosynthesis. Plant Sci. 292:110390. doi: 10.1016/j.plantsci.2019.110390
Yu, Y., Wu, G., Yuan, H., Cheng, L., Zhao, D., Huang, W., et al. (2016). Identification and characterization of miRNAs and targets in flax (Linum usitatissimum) under saline, alkaline, and saline-alkaline stresses. BMC Plant Biol. 16:124. doi: 10.1186/s12870-016-0808-2
Yuan, S., Li, Z., Li, D., Yuan, N., Hu, Q., and Luo, H. (2015). Constitutive expression of rice microRNA528 alters plant development and enhances tolerance to salinity stress and nitrogen starvation in creeping bentgrass. Plant Physiol. 169, 576–593. doi: 10.1104/pp.15.00899
Yuan, S., Zhao, J., Li, Z., Hu, Q., Yuan, N., Zhou, M., et al. (2019). MicroRNA396-mediated alteration in plant development and salinity stress response in creeping bentgrass. Hortic. Res. 6:48. doi: 10.1038/s41438-019-0130-x
Zandkarimi, H., Bedre, R., Solis, J., Mangu, V., and Baisakh, N. (2015). Sequencing and expression analysis of salt-responsive miRNAs and target genes in the halophyte smooth cordgrass (Spartina alternifolia Loisel). Mol. Biol. Rep. 42, 1341–1350. doi: 10.1007/s11033-015-3880-z
Zhai, J., Dong, Y., Sun, Y., Wang, Q., Wang, N., Wang, F., et al. (2014). Discovery and analysis of microRNAs in Leymus chinensis under saline-alkali and drought stress using high-throughput sequencing. PLoS One 9:e105417. doi: 10.1371/journal.pone.0105417
Zhang, H., Liu, X., Yang, X., Wu, H., Zhu, J., and Zhang, H. (2020). MiRNA-mRNA integrated analysis reveals roles for miRNAs in a typical halophyte, Reaumuria soongorica, during seed germination under salt stress. Plants 9:351. doi: 10.3390/plants9030351
Zhang, Q., Zhao, C., Li, M., Sun, W., Liu, Y., Xia, H., et al. (2013). Genome-wide identification of Thellungiella salsuginea microRNAs with putative roles in the salt stress response. BMC Plant Biol. 13:180. doi: 10.1186/1471-2229-13-180
Zhang, Y., Gong, H., Li, D., Zhou, R., Zhao, F., Zhang, X., et al. (2020). Integrated small RNA and degradome sequencing provide insights into salt tolerance in sesame (Sesamum indicum L.). BMC Genomics 21:494. doi: 10.1186/s12864-020-06913-3
Zhang, Y., Yun, Z., Gong, L., Qu, H. X., Duan, X. W., Jiang, Y. M., et al. (2018). Comparison of miRNA evolution and function in plants and animals. Microrna 7, 4–10. doi: 10.2174/2211536607666180126163031
Zhang, Z., Chen, C. Z., Xu, M. Q., Zhang, L. Q., Liu, J. B., Gao, Y., et al. (2019). MiR-31 and miR-143 affect steroid hormone synthesis and inhibit cell apoptosis in bovine granulosa cells through FSHR. Theriogenology 123:45–53. doi: 10.1016/j.theriogenology.2018.09.020
Zhao, B., Ge, L., Liang, R., Li, W., Ruan, K., Lin, H., et al. (2009). Members of miR-169 family are induced by high salinity and transiently inhibit the NF-YA transcription factor. BMC Mol. Biol. 10:29. doi: 10.1186/1471-2199-10-29
Zhao, J., Yuan, S., Zhou, M., Yuan, N., Li, Z., Hu, Q., et al. (2019). Transgenic creeping bentgrass overexpressing Osa-miR393a exhibits altered plant development and improved multiple stress tolerance. Plant Biotechnol. J. 17, 233–251. doi: 10.1111/pbi.12960
Zhao, Q., Zhang, H., Wang, T., Chen, S. X., and Dai, S. J. (2013). Proteomics-based investigation of salt-responsive mechanisms in plant roots. J. Proteome 82, 230–253. doi: 10.1016/j.jprot.2013.01.024
Zhou, M., Li, D., Li, Z., Hu, Q., Yang, C., Zhu, L., et al. (2013). Constitutive expression of a miR319 gene alters plant development and enhances salt and drought tolerance in transgenic creeping bentgrass. Plant Physiol. 161, 1375–1391. doi: 10.1104/pp.112.208702
Zhou, M., and Luo, H. (2014). Role of microRNA319 in creeping bentgrass salinity and drought stress response. Plant Signal Behav. 9:e28700. doi: 10.4161/psb.28700
Zhu, J. K. (2000). Genetic analysis of plant salt tolerance using Arabidopsis. Plant Physiol. 124, 941–948. doi: 10.1104/pp.124.3.941
Keywords: miRNA, plant, salt stress, tolerance, salinity resistance, crop
Citation: Xu T, Zhang L, Yang Z, Wei Y and Dong T (2021) Identification and Functional Characterization of Plant MiRNA Under Salt Stress Shed Light on Salinity Resistance Improvement Through MiRNA Manipulation in Crops. Front. Plant Sci. 12:665439. doi: 10.3389/fpls.2021.665439
Received: 08 February 2021; Accepted: 29 April 2021;
Published: 17 June 2021.
Edited by:
Pasqualina Woodrow, University of Campania Luigi Vanvitelli, ItalyReviewed by:
Arafat Abdel Hamed Abdel Latef, South Valley University, EgyptTushar Suhas Khare, Savitribai Phule Pune University, India
Vinay Kumar, Pune University, India
Copyright © 2021 Xu, Zhang, Yang, Wei and Dong. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Tao Xu, eHV0YW9feXJAMTI2LmNvbQ==; Tingting Dong, ZHR0QGpzbnUuZWR1LmNu
†These authors have contributed equally to this work
 Tao Xu
Tao Xu Long Zhang1†
Long Zhang1†