- 1Department of Genetics and Genome Biology, University of Leicester, Leicester, United Kingdom
- 2Department of Genetics and Cell Physiology, Faculty of Biological Sciences, University of Wrocław, Wrocław, Poland
- 3Key Laboratory of Plant Resources Conservation and Sustainable Utilization/Guangdong Provincial, Key Laboratory of Applied Botany, South China Botanical Garden, Chinese Academy of Sciences, Guangzhou, China
Structural chromosome rearrangements involving translocations, fusions and fissions lead to evolutionary variation between species and potentially reproductive isolation and variation in gene expression. While the wheats (Triticeae, Poaceae) and oats (Aveneae) all maintain a basic chromosome number of x=7, genomes of oats show frequent intergenomic translocations, in contrast to wheats where these translocations are relatively rare. We aimed to show genome structural diversity and genome relationships in tetraploid, hexaploid and octoploid Avena species and amphiploids, establishing patterns of intergenomic translocations across different oat taxa using fluorescence in situ hybridization (FISH) with four well-characterized repetitive DNA sequences: pAs120, AF226603, Ast-R171 and Ast-T116. In A. agadiriana (2n=4x=28), the selected probes hybridized to all chromosomes indicating that this species originated from one (autotetraploid) or closely related ancestors with the same genomes. Hexaploid amphiploids were confirmed as having the genomic composition AACCDD, while octoploid amphiploids showed three different genome compositions: AACCCCDD, AAAACCDD or AABBCCDD. The A, B, C, and D genomes of oats differ significantly in their involvement in non-centromeric, intercalary translocations. There was a predominance of distal intergenomic translocations from the C- into the D-genome chromosomes. Translocations from A- to C-, or D- to C-genome chromosomes were less frequent, proving that at least some of the translocations in oat polyploids are non-reciprocal. Rare translocations from A- to D-, D- to A- and C- to B-genome chromosomes were also visualized. The fundamental research has implications for exploiting genomic biodiversity in oat breeding through introgression from wild species potentially with contrasting chromosomal structures and hence deleterious segmental duplications or large deletions in amphiploid parental lines.
Introduction
Polyploidy and whole genome duplication have been recognized as major evolutionary processes in plants (Soltis et al., 2015; Alix et al., 2017; Zwaenepoel and Van de Peer, 2019; Huang and Rieseberg, 2020; Heslop-Harrison et al., 2022). While all plants are known to have whole genome duplications within their ancestry, one or more post cretaceous-tertiary (K-T) polyploidy events have been found in about half of species, including crops and wild plants. Genes that have been duplicated during the polyploidization process may retain or change their original function and can be mutationally or epigenetically silenced. In new polyploids, many evolutionary processes occur above the organizational level of duplicated genes (Adams and Wendel, 2005). These include elimination of whole chromosomes or even whole genomes (Hordeum: Gernand et al., 2005; Nicotiana: Patel et al., 2011) as well as intra- and inter-genomic chromosome translocations (Badaeva et al., 2007; Tomaszewska, 2021; Tomaszewska and Kosina, 2021). Changes occurring in polyploid nuclei may be associated with high chromatin condensation, transposable element activity, satellite homogenization, modulation of DNA methylation and histone modification (McClintock, 1984; Comai et al., 2003; Naish et al., 2021), and lead to the stabilization of polyploid genomes, thus increasing vitality and fertility, and extending the adaptive potential of such plants (Alix et al., 2017).
Chromosome translocations are important for plant evolution (Martin et al., 2020; Biswas et al., 2022; Liu et al., 2022; Wang et al., 2022), and occur in diploids, and after hybrid or amphiploid formation; they lead to exchange of chromosome segments within and between the ancestral genomes. Comparative genetic analysis in the grass family, including major crops, showed that particular taxonomic units are characterized by different chromosome translocations in terms of their position on chromosomes, reciprocity, and number of breaks involved (Bardsley et al., 1999). Various intragenomic reciprocal translocations were observed in rye (Kubaláková et al., 2003) and barley (Søgaard and von Wettstein-Knowles, 1987) genomes, and single intragenomic translocations between the B-genome chromosomes were the most frequent in wheat species (Badaeva et al., 2007). A detailed analysis of 373 Chinese wheat varieties has shown 14 different structural chromosome rearrangements including single- and reciprocal- translocations (Huang et al., 2018). Only two types of intergenomic translocations between A and B-genome chromosomes were observed regularly in 4x and 6x wheats. Translocations between D- and A- or B-genome (including 1Ds chromosomes) were rare, and most translocation breakpoints were at or near the centromere, rarely interstitially. These data together with the analysis of pericentric inversions conducted by Qi et al. (2006) give insight into the evolutionary dynamics of pericentromeric regions in the Triticeae tribe. The outcomes of genome evolution are different in the sister tribe of Aveneae, also involving an invariant basic chromosome number of x=7 and various polyploids: the oat lineage originated before wheat, with the C genome appearing earlier than the A or D genomes, suggesting a longer evolution time for oat hexaploids than for wheat (Fu, 2018; Peng et al., 2022), certainly with the major hexaploid crop species Triticum aestivum.
The genus Avena comprises about thirty diploid, tetraploid and hexaploid species with a basic chromosome number x=7 (Baum, 1977). The genomes of oat species were classified into A, B, C and D genome groups and further subdivisions (Rajhathy and Thomas, 1974; designated by a subscript e.g., Al, As, Cp or Cv). Diploid species are characterized by the presence of A or C genomes; the B genome is found only in some tetraploids. The D genome is present in all hexaploids (Badaeva et al., 2005) and some tetraploids (Yan et al., 2016; Tomaszewska and Kosina, 2021; Yan et al., 2021). The literature on the number and localization of intergenomic chromosome translocations in oats mainly uses in situ hybridization with total genomic DNA or some abundant repetitive DNA sequences (Chen and Armstrong, 1994; Jellen et al., 1994; Leggett et al., 1994; Linares et al., 2000; Tomaszewska and Kosina, 2021). Many distal (terminal) and several interstitial (subterminal) intergenomic translocations were observed, and only a few of them were reciprocal. Although the detection of translocation between A/D and C or A and B genomes with the use of genomic in situ hybridization, or genome-specific repetitive DNA probes, is straightforward, it is much more difficult to determine the translocations between A and D genomes due to the lack of sequence differentiation (Katsiotis et al., 2000; Luo et al., 2014; Liu et al., 2019; see wheat results from Huang et al., 2018). Linares et al. (1998) distinguished A and D genomes with the use of satellite sequences specific for the A genome, and Liu et al. (2019) developed probes specific for the D-genome. However, the translocations between A and D genomes have only been observed in oat endosperm (Tomaszewska and Kosina, 2021).
The combination of molecular cytogenetic and genomic or bioinformatic methods are promising to identify the processes occurring when genomes come together in polyploids (Heslop-Harrison and Schwarzacher, 2011; Tomaszewska et al., 2022). We can then address features of polyploidy that may be related to evolutionary adaptation (Barker et al., 2016) and test the hypotheses that chromosomal rearrangements, including intergenomic translocations, occur early in polyploid evolution. For this purpose, it is worth using artificial hybrids and amphiploids to track changes in genomes shortly after hybridization and duplication, and in further progenies (Tomaszewska and Kosina, 2018; Tomaszewska and Kosina, 2021). Understanding the coexistence of genomes in polyploid nuclei and consequences of genome rearrangement in polyploids is important to answer how non-cultivated diploids can be used for germplasm enhancement. Following either natural selection, speciation, or laboratory crossing, diploid genomes may be stabilized in a polyploid, or in backcrosses regularly used to generate breeding lines including wild species (Newell et al., 2012) so lines made for research may be more widely valuable. Translocations and other rearrangements may be useful to ensure desirable gene combinations are retained during breeding or represent a challenge because they restrict recombination.
Here we aim to recognize genomes in oat diploids and identify sequence characteristics of genomes with respect to the repetitive DNA composition, to show genome-relationships in tetra- and hexaploid Avena species and establish genome composition of their synthetic hybrids and amphiploids (6x and 8x) using fluorescence in situ hybridization (FISH) with well-characterized repetitive DNA sequences. Then we aim to establish patterns of intergenomic translocations across different taxa and relate it to the evolutionary divergence of oat genomes.
Materials and methods
Plant materials
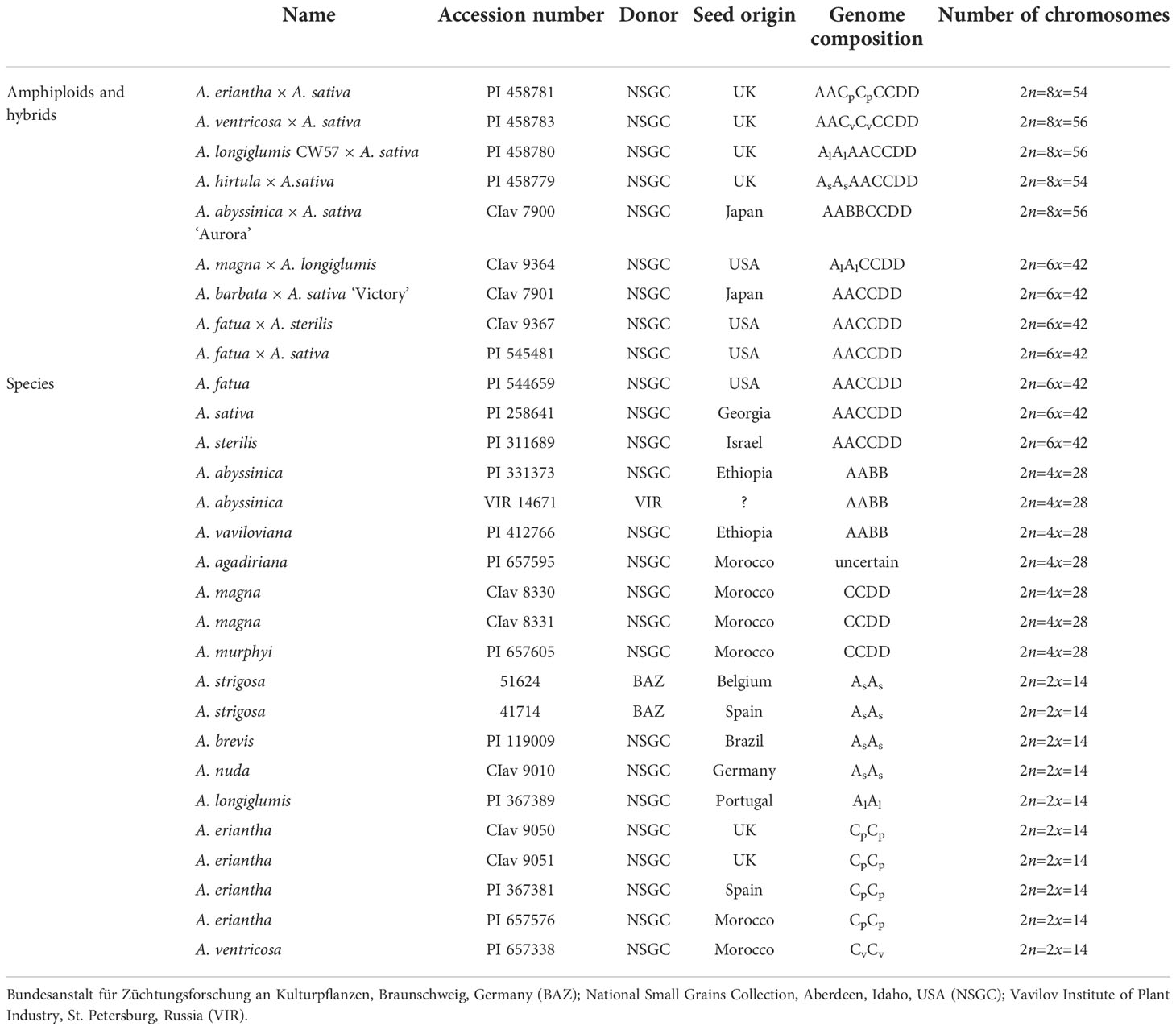
Species, hybrids and amphiploids of oats used in the study are listed in Table 1. Accessions were kindly provided by international germplasm collections. Some amphiploids and hybrids studied here were developed in United Kingdom, and also by Nishiyama in Japan mostly in the 1960-1970s (Nishiyama, 1962; Nishiyama et al., 1963; Nishiyama and Yabuno, 1979). The reproductive potential of oat species and their hybrid progeny (amphiploids) used in this study was already established by Tomaszewska and Kosina (2022), showing a high level of pollen grain viability. The botanical nomenclature of studied oat species was applied according to http://www.theplantlist.org/, accessed on 16 August 2022.
Chromosome preparation
Oat chromosomes were prepared according to the protocol described by Schwarzacher and Heslop-Harrison (2000). The root tips were treated with ice-cold water for 24 h to accumulate metaphases, and fixed in 96% ethanol: glacial acetic acid (3:1) for 48 h. Fixed root-tips were then washed in enzyme buffer (10mM citric acid/sodium citrate) for 15 min, and digested in an enzyme solution composed of 20U/ml cellulase (Sigma C1184), 10U/ml Onozuka RS cellulase (RPI C32400), and 20U/ml pectinase (Sigma P4716 from Aspergillus niger; solution in 40% glycerol) for 60 min at 37°C. Root tips were then squashed in 60% acetic acid. Cover slips were removed after freezing with dry ice. Slides were air-dried and used for in situ hybridization.
Probes used for in situ hybridization
Four different probes were selected for fluorescence in situ hybridization to distinguish genomes in polyploids and recognize major intergenomic translocations:
1. A genome-specific pAs120 (Linares et al., 1998)
2. C genome-specific AF226063 (Ananiev et al., 2002; Liu et al., 2019)
3. D genome-specific Ast_R171 (Liu et al., 2019)
4. D genome-specific Ast_T116 (Liu et al., 2019)
Conserved regions were amplified in a standard Polymerase Chain Reaction (PCR) using genome-specific primers (Linares et al., 1998; Liu et al., 2019) synthesized commercially (Sigma-Aldrich). These probes were labelled with digoxigenin-11-dUTP, biotin-16-dUTP or tetramethyl-rhodamine-5-dUTP (Roche) using BioPrime Array CGH and purified using BioPrime Purification Module (Invitrogen). The 45bp AF226063 oligonucleotide probe (Liu et al., 2019) was synthesized commercially (Sigma-Aldrich) with TET fluorescent dye attached to oligonucleotides at the 5’-end.
Fluorescence in situ hybridization procedure
FISH was performed as described by Schwarzacher and Heslop-Harrison (2000) and Tomaszewska and Kosina (2021) with minor modifications. The hybridization mixture consisted of 50% deionised formamide, 10% dextran sulphate, 1% sodium dodecyl sulphate (SDS), 2x SSC (saline sodium citrate buffer), amplified and labelled probe(s) (2 ng μL−1 each), and 200 ng μL−1 salmon sperm DNA was predenatured for 10 min at 75°C and stabilized on ice for 10 min. The 45bp AF226063 oligonucleotide probe was not predenatured and was added to the hybridization mixture containing amplified probes shortly after the predenaturation step. The hybridization mixture and chromosomes were then denatured together in a hybridization oven for 7 min at 75°C. Hybridization was performed at 37°C overnight. Slides were washed at 42°C in 2x SSC for 2 min, in 0.1x SSC for 10 min, and 2x SSC for 20 min. Hybridization signals of probes labelled with digoxigenin-11-dUTP and biotin-16-dUTP were detected with antidigoxigenin conjugated to fluorescein isothiocyanate (FITC; Roche Diagnostics) and streptavidin conjugated to Alexa 594 or Alexa 647 (Life Technologies-Molecular Probes), respectively. Air-dried slides were counterstained with DAPI (4′,6-diamidino-2-phenylindole, 2μg mL) in antifade solution (AF1, Citifluor).
Microscopy and image capture
The slides were examined with a Nikon Eclipse 80i epifluorescence microscope (Nikon, Tokyo, Japan). Images were taken using a DS-QiMc monochromatic camera and NIS-Elements v.2.34 software. Karyotypes were prepared using IdeoKar 1.3 (Mirzaghaderi and Marzangi, 2015) and Adobe Photoshop.
Results
Chromosome numbers
The number of chromosomes of studied accessions is shown in Table 1. For FISH analysis, we used accessions with the number of chromosomes typical for a given species and thus omitted A. wiestii PI 299112 and A. barbata PI 337795, each having 42 chromosomes instead of the reported 14 and 28, respectively, suggesting uncontrolled crossing in breeding or involvement of 2n gametes in stocks. All species, hybrids and amphiploids of oat studied here were euploid (Table 1), except A. eriantha x A. sativa and A. hirtula x A. sativa where most metaphases had 2n=8x-2 = 54. Ploidy levels of hybrids and amphiploids were re-examined and compared with genebank databases. Ploidy level of A. barbata x A. sativa ‘Victory’ CIav 7901, previously recognized as octoploid, had to be revised to hexaploid.
FISH-based reference karyotypes of diploid oats
Two different probes, pAs120 (Linares et al., 1998) and AF226063 (Ananiev et al., 2002; Liu et al., 2019) repeats, were used for in situ hybridization on metaphases of diploid oat species (Table 1). The karyotypes were prepared, and the patterns of signals of the two genome-specific probes were determined (Figure 1). These patterns were helpful in establishing the genomic composition of polyploids and recognizing intergenomic translocations in tetra-, hexa- and octo-ploids. Probe pAs120 has been tested on chromosomes of different diploids having Al (A. longiglumis) and As genomes (A. brevis, A. nuda, A. strigosa). All studied species showed dispersed signals along chromosomes, except for telomeres and secondary constrictions. Probe pAs120 was not useful for distinguishing between Al and As genomes. Probe AF226063 has been used on chromosomes of different accessions of diploid A. eriantha (Cp genome) and A. ventricosa (Cv genome). Some chromosomes showed specific probe signals in the form of bands being good chromosome markers. Probe AF226063 was not useful for distinguishing between Cp and Cv genomes.
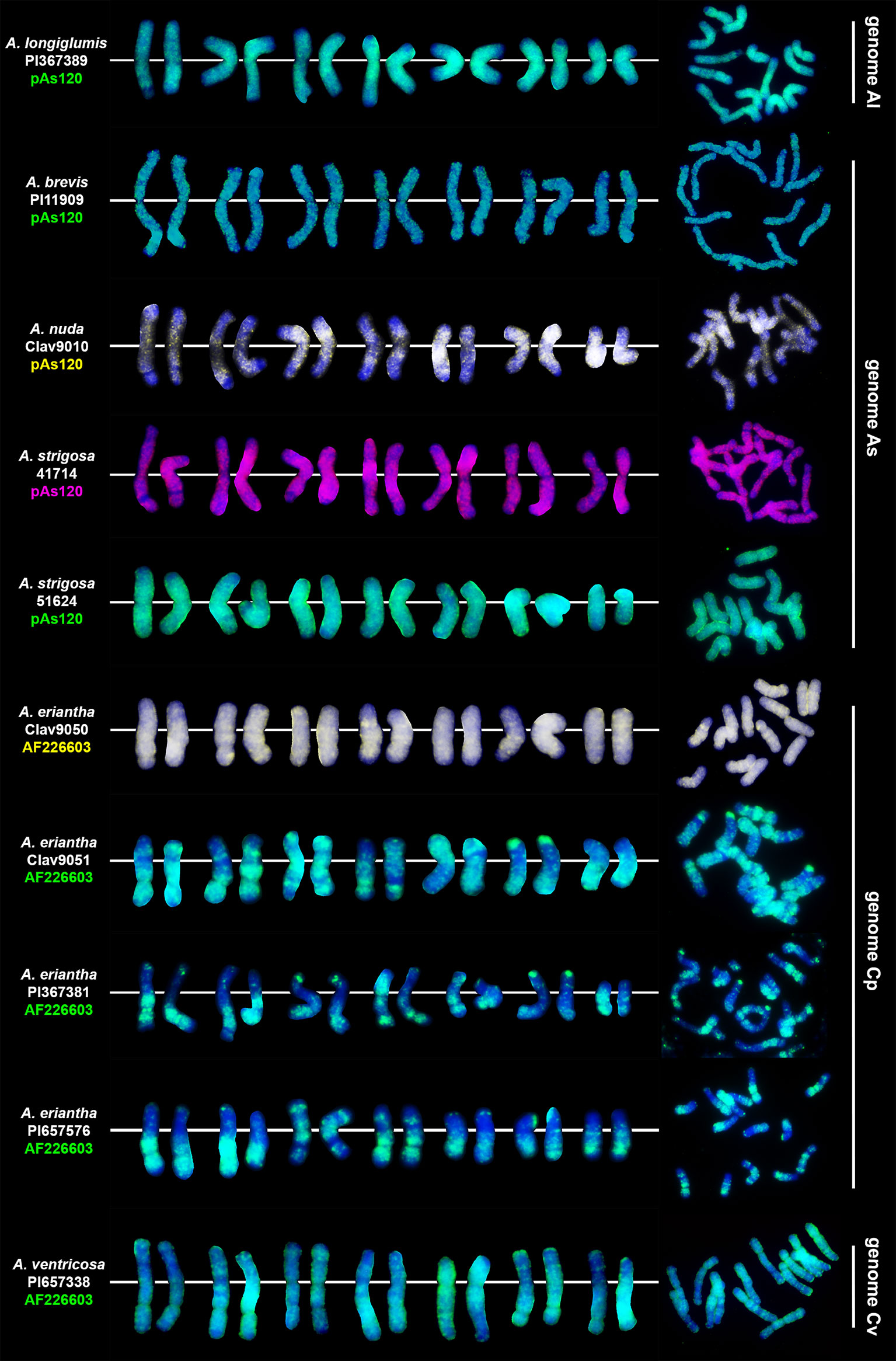
Figure 1 Chromosomal location of highly repetitive DNA motifs in diploid Avena species having genome constitution of Al, As, Cp and Cv. Scale bar = 10μm.
Chromosomal location of highly repetitive DNA motifs in tetraploid and hexaploid oat species
We studied three species belonging to the AABB Avena group: A. abyssinica, A. vaviloviana and A. agadiriana. Genome composition of A. abyssinica and A. vaviloviana was confirmed using pAs120 probe, which painted 14 chromosomes out of 28, designating the A genome (Figure 2). Chromosomes that were not painted by pAs120 were considered genome B. Hybridization of Ast-T116 probe to chromosomes of A. agadiriana gave strong dispersed signals along all of the 28 chromosomes. Another probe pAs120 gave weak dispersed signals along 28 chromosomes, hence the genomic composition of this species remains ambiguous. Genome composition of two tetraploid species previously classified into AACC Avena group, A. magna (syn. A. maroccana) and A. murphyi, was established using two different probes (Figure 3). In both species, AF226603 painted 14 chromosomes out of 28 while Ast-T116 showed dispersed signals along the remaining 14 chromosomes, proving that the genome composition of these species was CCDD. All hexaploid species studied here, including A. fatua, A. sterilis and A. sativa, had AACCDD genome composition, and this was confirmed using three different genome-specific probes: pAs120, AF226603 and Ast-T116 (Figure 4).
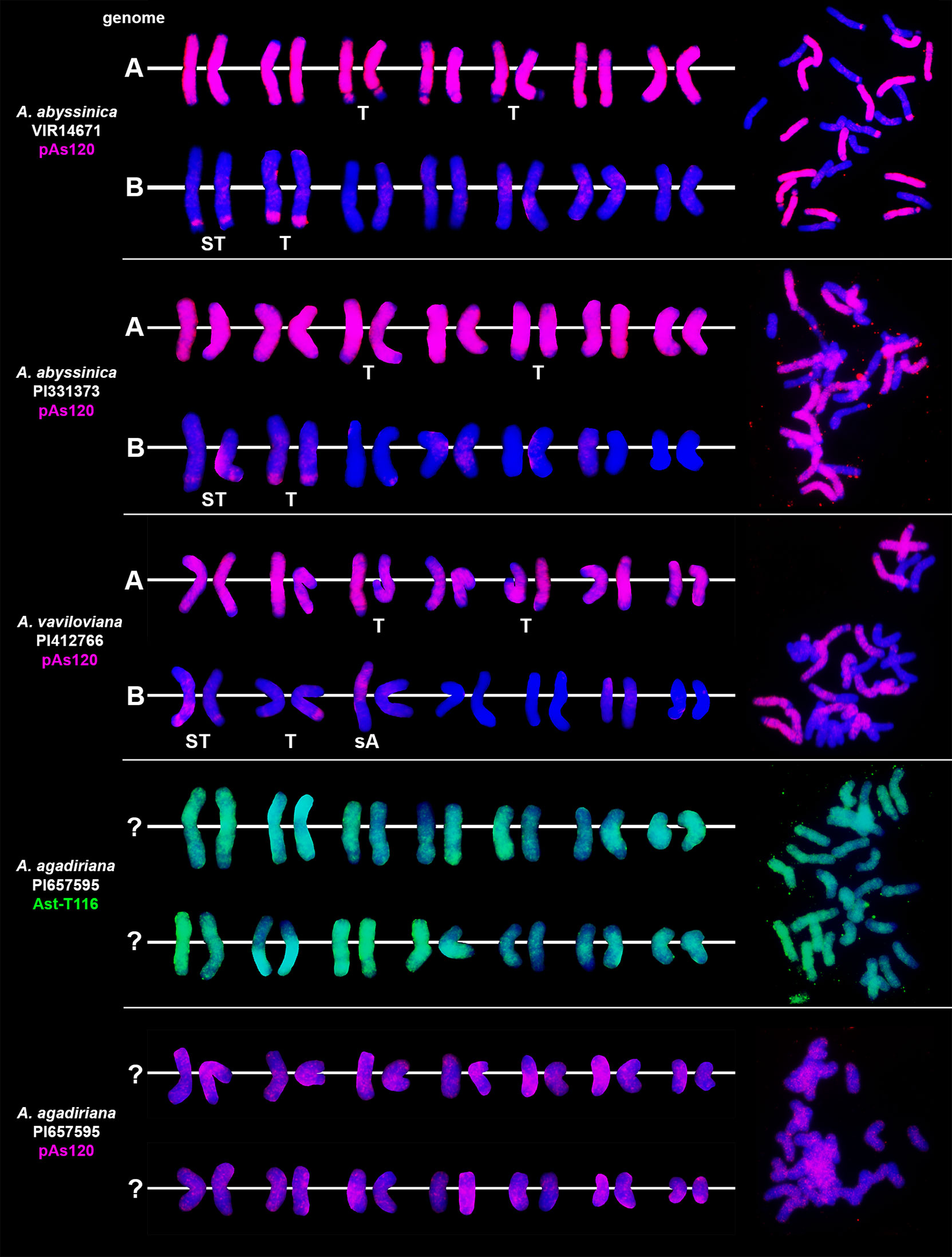
Figure 2 Genome composition and intergenomic translocations pattern in tetraploid oat species belonging to AABB Avena group. D- distal translocation, I- interstitial translocation, A - translocation of whole arm. Scale bar = 10μm.
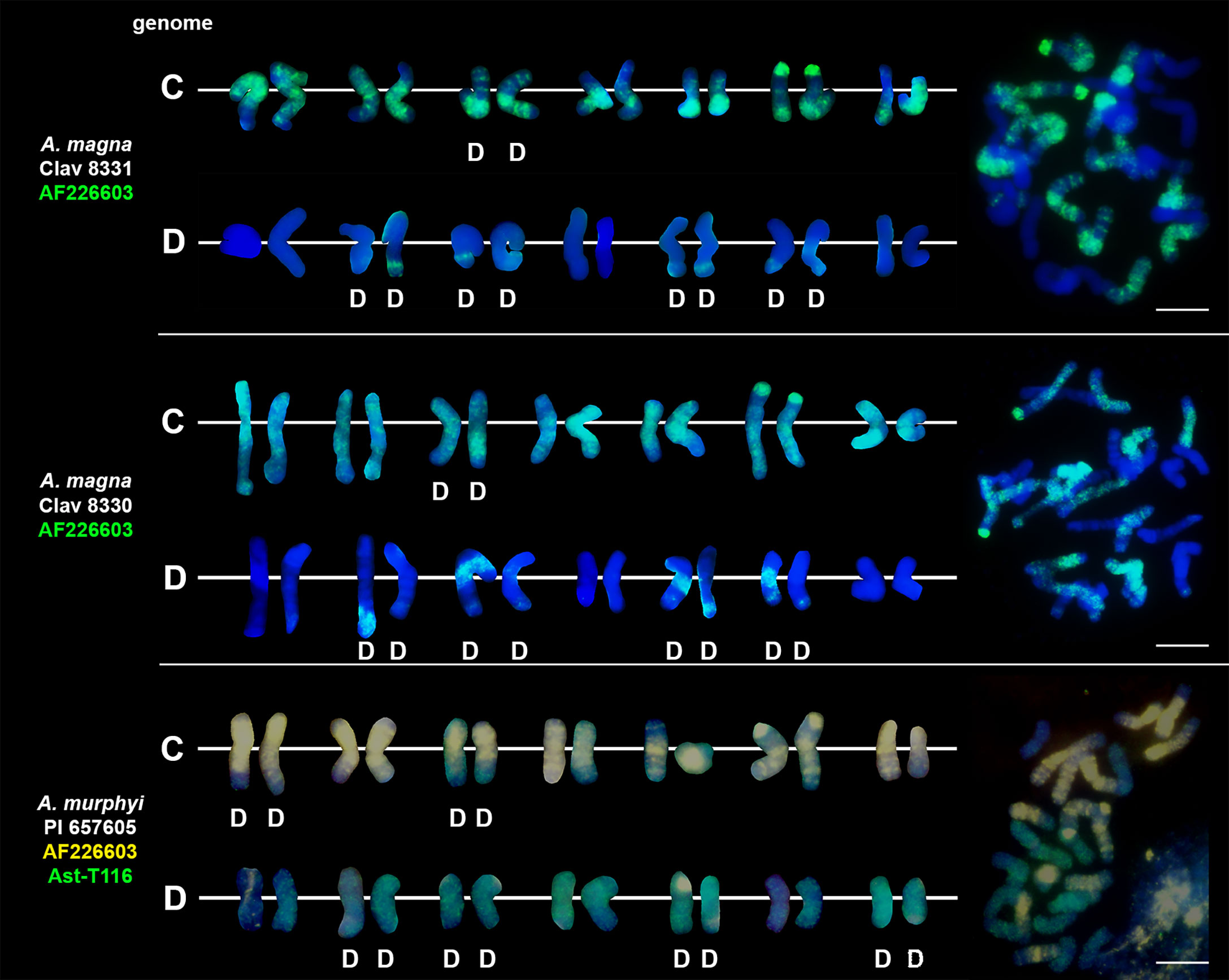
Figure 3 Genome composition and intergenomic translocations pattern in tetraploid oat species belonging to CCDD Avena group. D- distal translocation. Scale bar = 10μm.
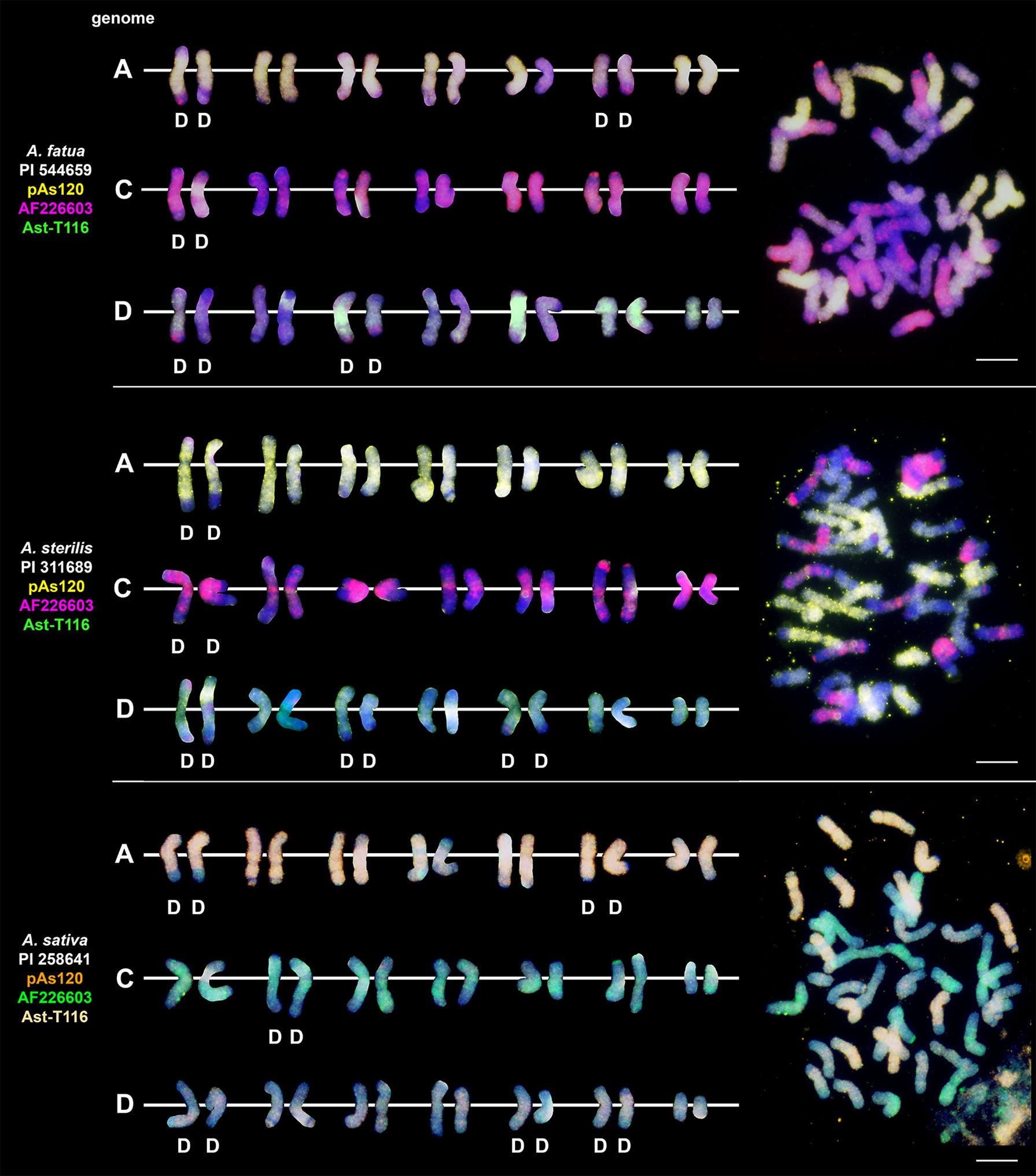
Figure 4 The diversity of intergenomic chromosome translocations across different hexaploid oat species. D- distal translocation. Scale bar = 10μm.
Chromosomal organization of specific repeats in artificial hybrids and amphiploids of oats
Probes pAs120, AF226603, Ast-R171 and Ast-T116 were used to establish genome composition of different artificial hybrids and amphiploids of oats. The probes produced multiple signals that were evenly distributed along 14 chromosomes each, enabling identification of A, C and D genomes. Chromosomes that were not painted by any of these probes were considered genome B. Each of the analyzed hexaploid hybrids and amphiploids had the genomic composition of AACCDD (Figure 5), as did hexaploid oat species (Figure 4). In the amphiploid A. magna x A. longiglumis, we were able to more accurately determine the genomic composition as AlAlCCDD by looking at the genomes of the parental species. We recognized 3 types of octoploid amphiploids, each having different genome composition: AACCCCDD, AAAACCDD or AABBCCDD. A. eriantha x A. sativa and A. ventricosa x A. sativa belonged to the first group having AACpCpCCDD and AACvCvCCDD genomes, respectively (Figure 6). No letter in the subscript was assigned to the second C genome of these amphiploids because this genome originated from A. sativa where the ancestors of this species have not been thoroughly investigated. A. longiglumis CW57 × A. sativa and A. hirtula x A. sativa belonged to the second group of amphiploids having AlAlAACCDD and AsAsAACCDD genomes, respectively (Figure 7). No letter in the subscript was assigned to the second A genome originated from A. sativa. The most likely genomic composition of A. abyssinica x A. sativa ‘Aurora’ is AABBCCDD (Figure 8). One of the genomes of this amphiploid has not been labelled with any probe and is believed to belong to genome B.
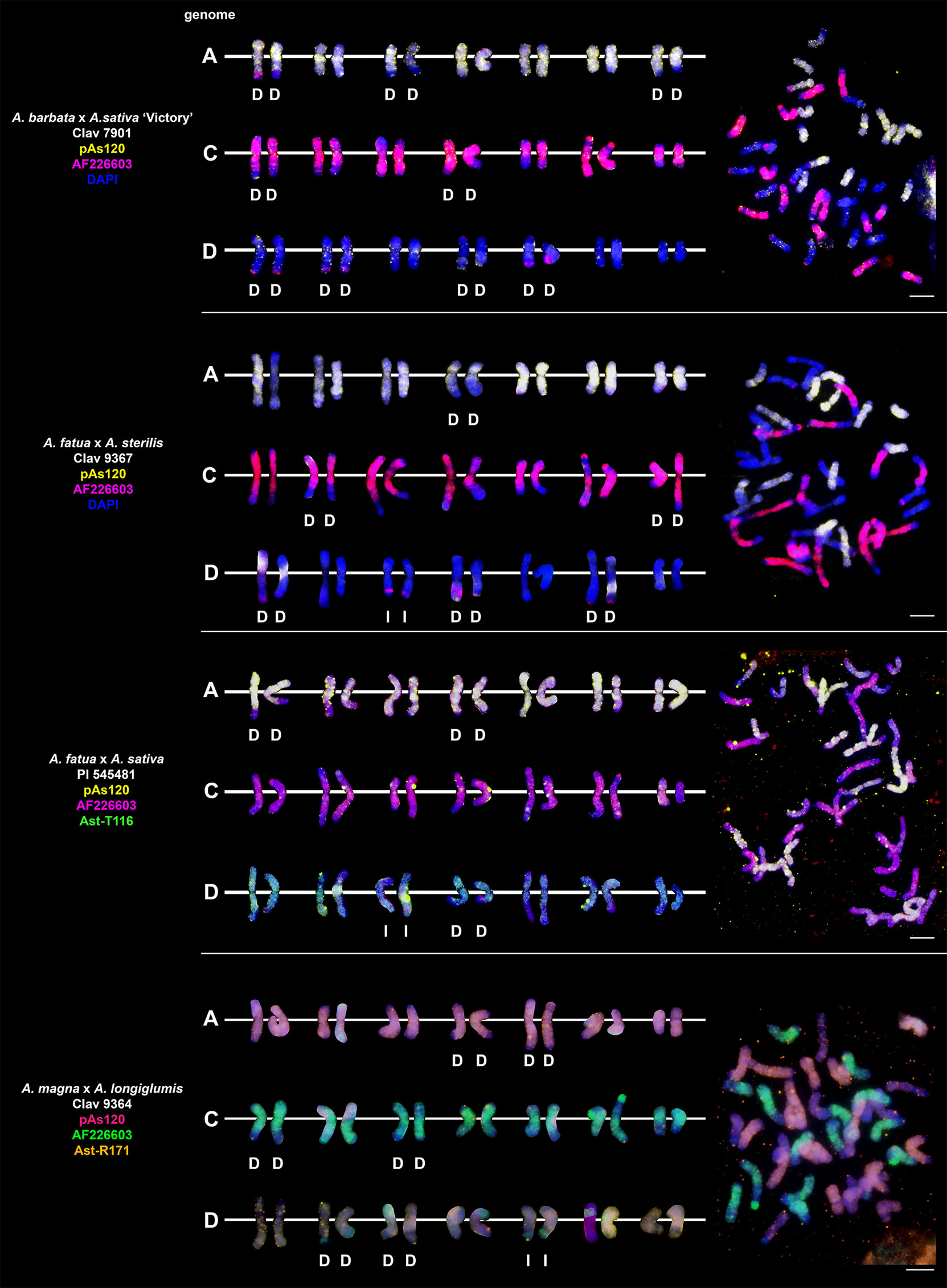
Figure 5 Chromosomal location of repetitive DNA motifs enabling identification of AACCDD genomes and intergenomic translocations in different hexaploid hybrids and amphiploids of oats. D- distal translocation, I- interstitial translocation. Scale bar = 10μm.
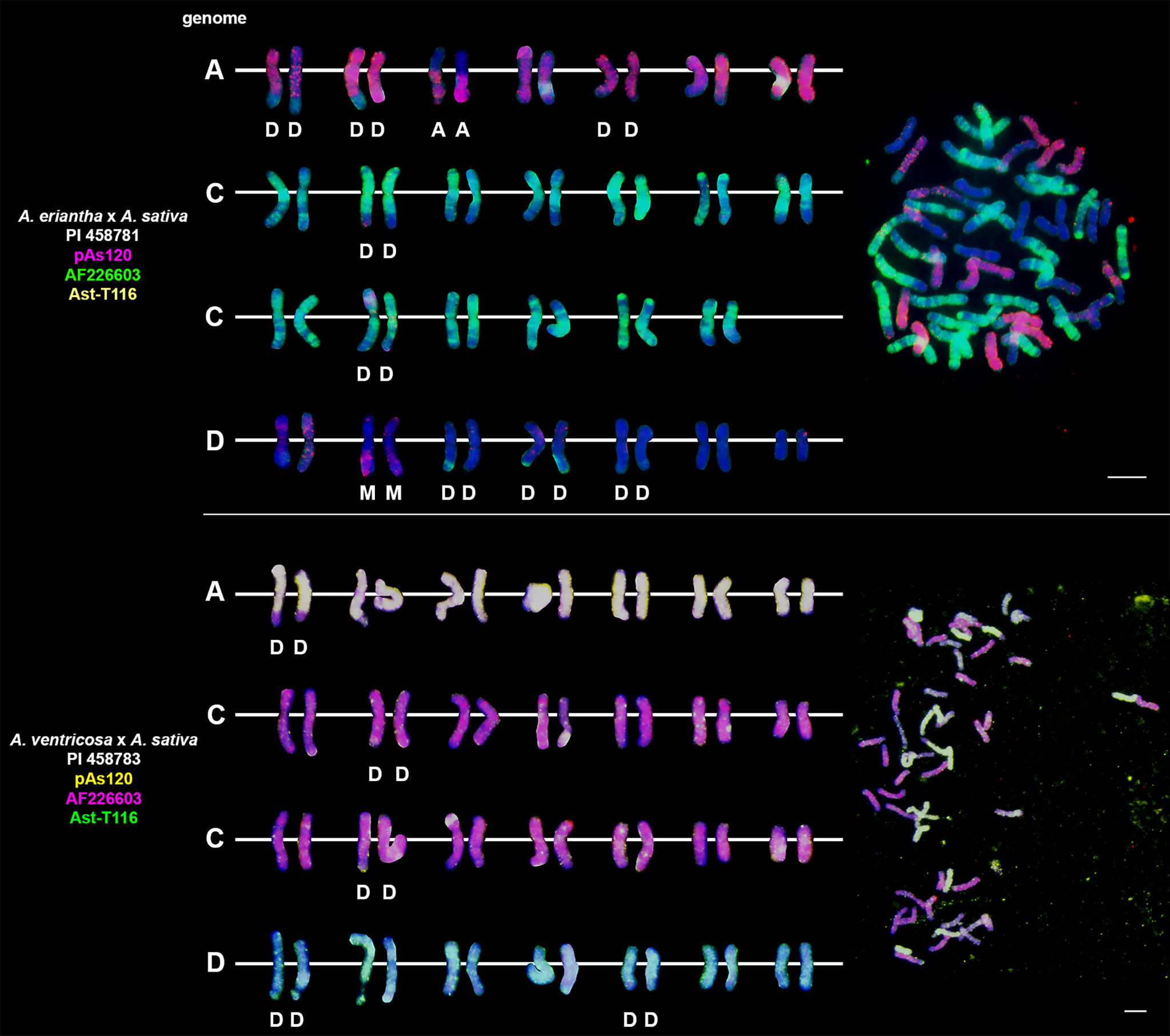
Figure 6 Fluorescence in situ hybridization with repetitive DNA probes to chromosomes of amphiploids formed by crossing diploid species having CC genomes with A. sativa. Intergenomic translocations were visualized. D- distal translocation, I- interstitial translocation, M - multiple translocation or insertion. Scale bar = 10μm.
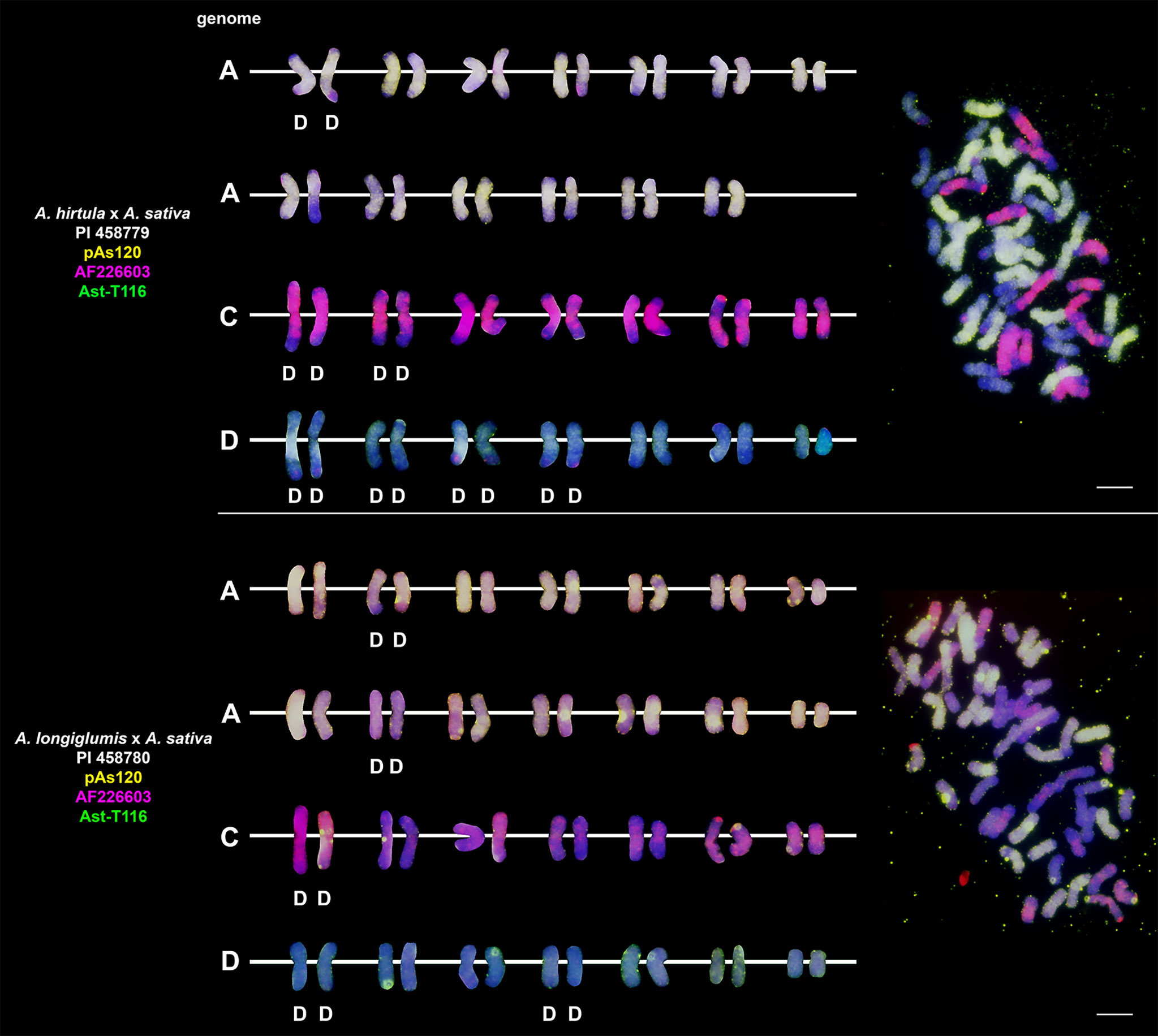
Figure 7 Fluorescence in situ hybridization with repetitive DNA probes to chromosomes of amphiploids formed by crossing diploid species having AA genomes with A. sativa. Intergenomic translocations were visualized. D- distal translocation. Scale bar = 10μm.
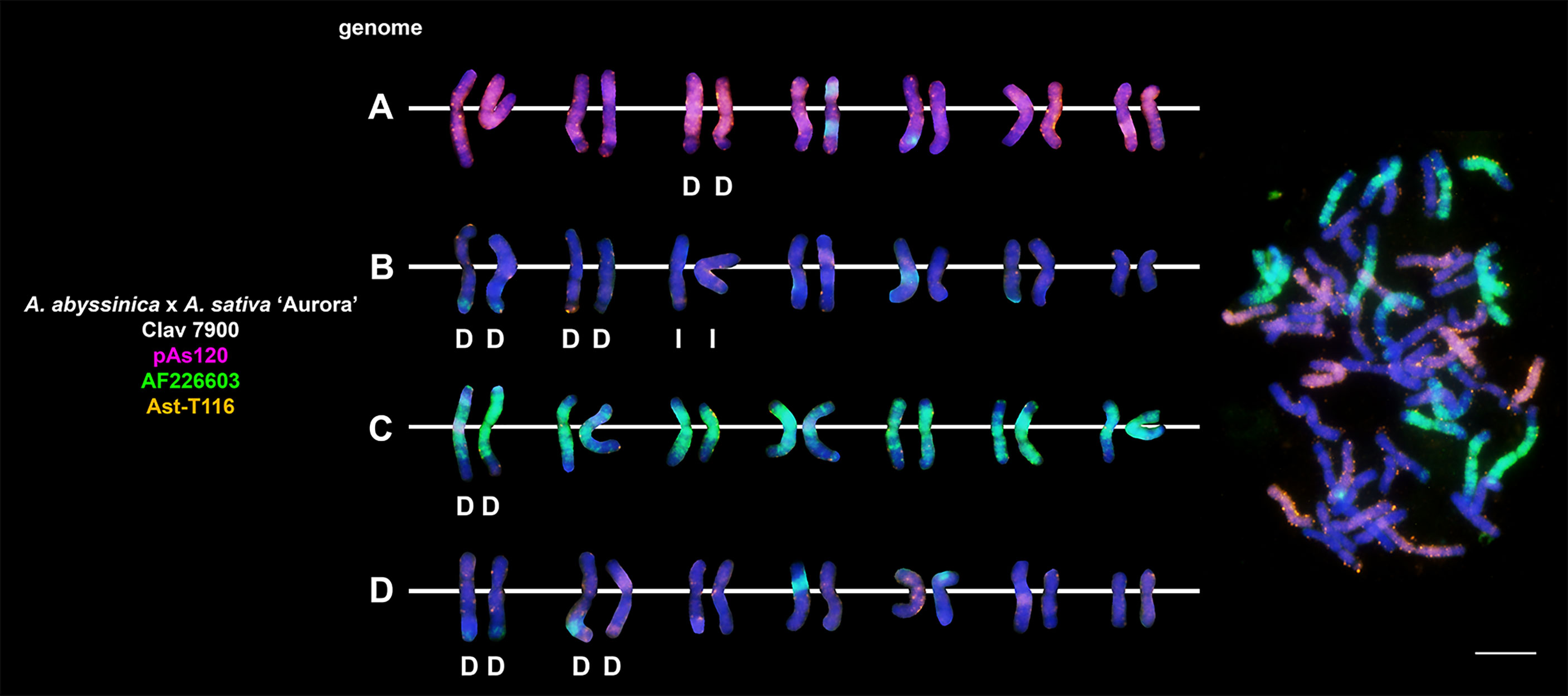

Figure 8 Fluorescence in situ hybridization with repetitive DNA probes to chromosomes of an amphiploid formed by crossing tetraploid A. abyssinica with A. sativa. Intergenomic translocations were visualized. D- distal translocation, I- interstitial translocation. Scale bar = 10μm.
Visualization of intergenomic translocations using genome-specific repeats
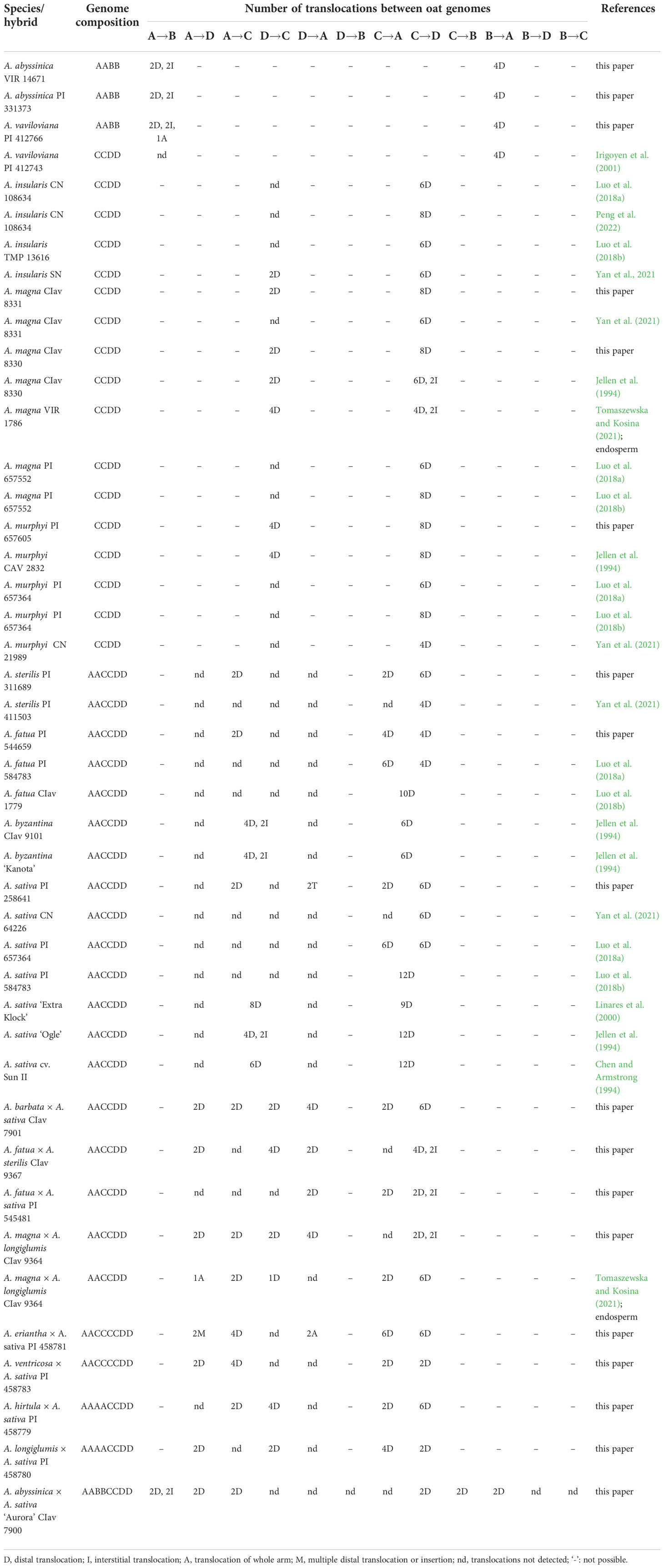
Fluorescent in situ hybridization with the use of repetitive DNA motifs to chromosomes of different polyploids (4x, 6x, 8x) enabled identification of intergenomic translocations (Figures 2–8). Using probes specific to genomes A, C and D, we were able to discriminate C→A, C→D and A→C, D→C translocations in hexaploids (Figures 4, 5) and octoploids (Figures 6–8). Rare A→D, D→A and C→B translocations were also visualized. The translocations involving B and D genomes were not detected in A. abyssinica x A. sativa ‘Aurora’ octoploid (Figure 8).
Most of the translocation events were at the distal position of long chromosome arms (terminal or subterminal), rarely interstitially (Figures 2–8). Some of the distal translocations involved SAT chromosomes, as observed in A. abyssinica and A. vaviloviana (Figure 2). In some of the accessions examined, we detected whole chromosome arm translocations, i.e. A. vaviloviana (Figure 2; one chromosome from a pair) or A. eriantha x A. sativa (Figure 6; two chromosomes from a pair). In the latter, we additionally revealed a pair of D genome chromosomes having red C genome signals at the ends of both arms. It suggests multiple distal translocations or insertion, presumably involving chromosome 3 from genome A showing large whole arm translocation (Figure 6).
Pattern of intergenomic translocations across different oat taxa
Our detailed studies of genome relationships in polyploid species and interspecific amphiploids and hybrids of oats, including both FISH (Figures 2–8; Table 2) and some literature data (Table 2) analyses, indicated that the chromosomes of A, B, C and D genomes differ significantly in their involvement in translocations. There is a predominance of distal intergenomic translocations from the C- into the D-genome chromosomes in CCDD-tetraploid and AACCDD-hexaploid species. A→C or D→C translocations are less frequent, proving that at least some of the translocations in oat polyploids are non-reciprocal. The number of A→B and B→A translocations in tetraploid A. abyssinica was the same, but their position on the chromosomes (distal versus interstitial) indicated their non-reciprocity.
Comparative analysis of species and artificial hybrids/amphiploids having genomic composition of AACCDD (Figures 4, 5; Table 2) showed that these two groups differ in the number and position of translocations. Multitude of translocations covering all genomes in hybrid octoploids (Figures 6–8), and probably other extensive changes including large insertions (A. eriantha x A. sativa, Figure 6) have occurred soon after amphiploid formation.
Our analysis across different oat taxa (Figures 2–8; Table 2) showed that some species, such as A. abyssinica and A. vaviloviana (Figure 2), share some of the intergenomic translocations (most likely they are identical-by-descent, but re-iteration of same events in different lineages cannot be ruled out). The translocation of the entire arm of a single chromosome observed in A. vaviloviana may be species-specific or specific for this particular accession. Both our karyotype analysis (Figure 3) and the data obtained by different authors (Table 2) indicated that some translocations between the C and D genomes in hexaploid oat species (Figure 4) involved the same chromosomes as in tetraploid CCDD species (Figure 3). The translocation pattern seen in amphiploids created by crossing different maternal diploids with paternal A. sativa indicated that some translocations were transferred from A. sativa to the hybrid generation, and new translocations were also appeared (Figures 5–7).
Discussion
Repetitive DNA elements are rapidly evolving major components of plant genomes, thus becoming important tools for studying the large-scale organization and evolution of plant genomes (Heslop-Harrison and Schwarzacher, 2011; Mehrotra and Goyal, 2014; Biscotti et al., 2015; Naish et al., 2021; Zhang et al., 2022a). Here, by using identified repetitive sequences that were unique to the A, C and D genomes of oats (Linares et al., 1998; Ananiev et al., 2002; Liu et al., 2019), we could reveal the genome composition of polyploids and define the nature of major evolutionary changes in oat genomes. We can then discuss intergenomic translocations, their consequences for phylogeny, speciation and polyploidy, and creation of new hybrids and breeding.
Re-evaluation of genome composition of tetraploid and hexaploid oat species and potential speciation in the genus Avena
Although the genome composition of hexaploid oat species is well known (Chen and Armstrong, 1994; Jellen et al., 1994; Liu et al., 2019), and also confirmed by our analysis using A, C and D genome-specific repetitive DNA probes, there are still debates about the classification and origin of genomes in tetraploid species. The genomic composition of tetraploid A. magna, A. murphyi and A. insularis, previously assigned as AACC (Leggett et al., 1994; Shelukhina et al., 2007), has recently been revised using molecular and cytological analysis (Peng et al., 2010; Yan et al., 2014; Tomaszewska and Kosina, 2021). Here, we presented new evidence confirming the CCDD genomic constitutions of the tetraploid A. magna and A. murphyi using published C genome-specific 45bp AF226603 probe (Ananiev et al., 2002; Liu et al., 2019) as well as D genome-specific Ast-T116 and Ast-R171 probes (Liu et al., 2019). These data together with high-density genetic markers analysis (Fominaya et al., 2021) suggested that the CCDD-genome tetraploids could contribute to the evolution of hexaploid oats (Figure 9). Our in situ hybridization with pAs120 probe confirmed a clear dissimilarity of the two genomes in tetraploid A. abyssinica and A. vaviloviana, consistent with Irigoyen et al. (2001) and Yan et al. (2014), proving AABB genome composition of these species. Karyotype and molecular data revealed distinction of A. agadiriana from other species classified as AABB-genome tetraploids (Jellen and Gill, 1996; Badaeva et al., 2010; Peng et al., 2010; Yan et al., 2014; Chew et al., 2016). Our analysis showed evenly distributed signals of the D genome-specific Ast-T116 probe on 28 chromosomes of A. agadiriana suggesting that the genome composition of this species might be DDDD, not AABB as previously stated. On the other hand, pAs120 probe showed weak dispersed signals along all of the 28 chromosomes, thus, the evidence provided here should be taken as putative and encouraging for further exploration of this species. The phylogenetic analysis supported A. agadiriana being closely related to A. canariensis and A. longiglumis (Chew et al., 2016; Peng et al., 2018). Some authors speculated that the genome composition of this species might be AADD due to some similarities to CCDD and AACCDD Avena groups (Badaeva et al., 2010; Luo et al., 2018c), but our FISH signals of A and D genome-specific probes rather indicated an autopolyploid origin of A. agadiriana. Our results may also suggest that the genomes of A. agadiriana are different from those previously identified in other oat species as A, B, C and D and may require a different nomenclature (Yan et al., 2016), especially that the analysis of chloroplast genome demonstrated potential independent evolution of this species (Figure 9; Fu, 2018). Correct recognition of the A. agadiriana genomes could complement the data on the origin of the D genome in AACCDD-hexaploid oats (Luo et al., 2018c), supported by easy crossing of these two species (Badaeva et al., 2010).
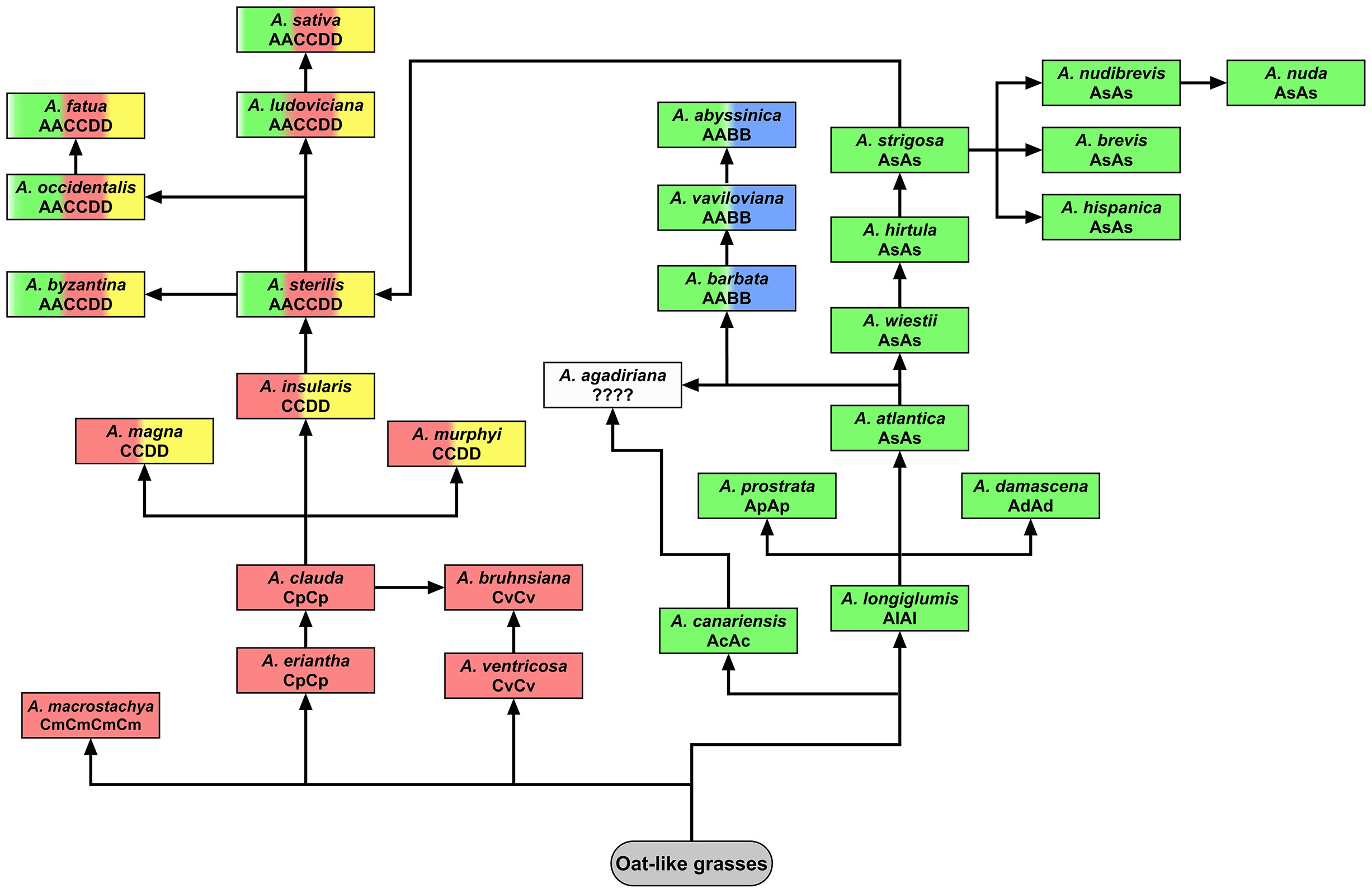
Figure 9 Model of potential evolutionary pathways leading to the development of modern tetraploid and hexaploid oat species based on the latest phylogenetic analyses. C and A genomes developed independently from oat-like grasses within the subgenus Avenastrum (Loskutov, 2007). According to Fu (2018), C-genome species diverged 19.9-21.2 million years ago (Mya), while A. canariensis, considered the oldest species having A genome, diverged 13-15 Mya. These data are contradictory to the analysis of chloroplast genome conducted by Liu et al. (2020), indicating that the diverged A genomes originate from A. longiglumis. A. bruhnsiana is an endemic species closely related to A. ventricosa, with A. clauda being a potential second ancestor (Gnutikov et al., 2022). Presumably different diploid species having the A and C genomes may have been involved in the evolution of polyploid oats (Peng et al., 2010). Species having AABB genomes (A. barbata, A. vaviloviana, A. abyssinica) show a close relationship with A. wiestii or A. hirtula. Peng et al. (2018) suggested that these species derived through autopolyploidization of As genome, contrary to Badaeva et al. (2010) and Irigoyen et al. (2001) who proposed allotetraploid origin. According to Peng et al. (2018), A. agadiriana, which developed independently, is closely related to the As and Ac genomes, but the genome composition of this species remains debatable (this paper; Badaeva et al., 2010; Yan et al., 2016; Luo et al., 2018c). Phylogenetic analysis conducted by Peng et al. (2018) indicated that Cp genome (A. eriantha, A. clauda) is more closely related to CCDD-tetraploids and AACCDD-hexaploids than Cv genome (A. ventricosa). Probably not one, but several of the extant CCDD-tetraploids were involved in the formation of hexaploid oat species (Yan et al., 2021), but A. insularis and diploid species having As genome seem to be putative ancestors of oat hexaploids (Drossou et al., 2004; Badaeva et al., 2005; Peng et al., 2018; Fominaya et al., 2021; Jiang et al., 2021). Molecular cytogenetic analysis revealed that none of the modern diploid oat species having A or C genome was the direct ancestor of polyploids, however, A. insularis showed a relatively high number of transcribed spacers, displaying sequence similarity to one accession of A. hirtula (Loskutov et al., 2021). On the other hand, the principal coordinates analysis revealed that A. eriantha and A. longiglumis are closer to CCDD oat tetraploids than other diploids (Yan et al., 2016). This was also proved by analysis of chloroplast and mitochondrial genomes (Fu, 2018), and easy crossing of A. longiglumis with CCDD tetraploids (see A. magna x A. longiglumis amphiploid in this paper; Drossou et al., 2004; Tomaszewska and Kosina, 2021).
Identification of genomes in artificial hybrids and amphiploids of oats
Recognition of genome composition and genomic changes in hybrids is important in the context of plant speciation and evolution to explore drivers that played a major role in genome divergence (Alix et al., 2008; Patokar et al., 2016; Martin et al., 2020; Tomaszewska, 2021). The genome composition and intergenomic rearrangements of artificial hybrids and amphiploids of oats were investigated here for the first time using repetitive DNA sequences. A certain advantage of the repetitive DNA probes over the whole genomic DNA probes used for in situ hybridization is observed (Tomaszewska et al., 2022). Despite the often perfect discrimination of genomes in allopolyploids/amphiploids using total genomic DNA isolated from a diploid species (Tomaszewska and Kosina, 2021), such probes are not applicable to distinguish closely related genomes such as those observed in Brassica (Alix et al., 2008), Triticum (Tang et al., 2018), Cenchrus and Urochloa tropical forage grasses (Rathore et al., 2022; Tomaszewska et al., 2022) or highly homologous A and D genomes in Avena (Linares et al., 1998). The parental species of amphiploids studied here are meiotically compatible in the crossing process, exhibiting sometimes partial sterility (Loskutov, 2001; Tomaszewska and Kosina, 2022), and these data together with established genome composition of hybrids and amphiploids contribute to a better understanding of the complex evolutionary processes within the genus Avena.
Rapid genome changes in allopolyploids: intergenomic translocations in polyploid plant species and synthetic hybrids
Intra- and inter-genomic chromosome translocations have been recognized as significant processes accompanied natural evolution of diploid and polyploid plant species (Martin et al., 2020). The exchange of segments of non-homologous chromosomes occurred soon after allopolyploid formation, shape their genomes giving them an adaptive and evolutionary advantage (Tomaszewska, 2021). In some allopolyploid species, intergenomic translocations seem to be common, as in Nicotiana (Kenton et al., 1993), Festulolium species (Kopecký et al., 2006), Solanum tuberosum and S. caripense (Braz et al., 2018), Anthoxanthum (Chumová et al., 2021), Hordeum secalinum (Bustos et al., 1996) and H. capense (Taketa et al., 1999) or wild wheats (Jiang and Gill, 1994; Badaeva et al., 1995), contrary to wheat cultivars (Badaeva et al., 2007) or Secale (Kubaláková et al., 2003) where intragenomic translocations located at pericentromeric regions of chromosomes, rarely interstitially, are the most common. In Musaceae, there are many translocations at any place in the chromosome arm (Wang et al., 2022).
Intergenomic translocations are a relatively common phenomenon in artificially induced allopolyploids (amphiploids). In newly synthesized wheat-rye allopolyploids, the transfer of rye telomeric chromatin involving the short arm of chromosome 1 (Kubaláková et al., 2003) to wheat centromere was observed in two continuous generations (Fu et al., 2010). Through distant crossings the yield of wheat cultivars has been improved and resistance to pathogens and insects has been induced (Mondal et al., 2016). Numerous distal (terminal) intergenomic translocations were also detected in wheat-barley lines (Prieto et al., 2001; Nagy et al., 2002), wheat-Aegilops biuncialis amphiploids (Molnár et al., 2009), amphiploid trigeneric hybrid involving Triticum, Thinopyrum and Lophopyrum (Kosina and Heslop-Harrison, 1996) or in endosperm of Avena amphiploid (Tomaszewska and Kosina, 2021).
Evolutionary dynamics in genus Avena in the context of intergenomic translocations
Comparative linkage mapping of oat species suggested extensive chromosome rearrangement in ancestral diploids, both involving SAT and non-SAT chromosomes (Badaeva et al., 2010; Latta et al., 2019); and intergenomic translocations occurred between non-homologous chromosomes were considered to be significant evolutionary forces leading to divergence of the tetra- and hexaploid oat species (Linares et al., 1998). Polyploid oat genomes are particularly rearranged showing multiple distal, rarely interstitial, intergenomic translocations. Our analysis of oat species and artificial hybrids having genomic composition of AACCDD showed that these two groups differ in the number and position of translocations, meaning that a new pattern of intergenomic translocations emerged soon after hybridization and was passed on and maintained over the next hybrid generations. The translocation pattern seen in amphiploids created by crossing different maternal diploids with paternal A. sativa indicated that some translocations were transferred from A. sativa to the hybrid generation. New translocations also appeared, supporting the hypothesis that at least some of the intergenomic translocations contributed significantly to the divergence of oat species. However, the analysis of A. sativa indicated ongoing genomic exchange in this hexaploid (Latta et al., 2019), thus, it is not only hybridization and genome duplication that affect genome rearrangement. In the hybrids involving A. strigosa, A. eriantha, and A. magna, a poor chromosome pairing was observed during meiosis, and the translocations may be the major factor limiting fertility rather than the lack of chromosome homology (Tomaszewska and Kosina, 2021). Thus, potentially, the presence of intra- as well as inter-genomic translocations in Avena would limit backcrossing in breeding (Leggett, 1998; Nikoloudakis and Katsiotis, 2015).
Our study showed that A. abyssinica and A. vaviloviana share some of the translocations between A and B genomes which indicates that translocations were one of the factors driving the divergence of AABB Avena group. The translocation of the entire arm of a single chromosome observed in A. vaviloviana may be species-specific or specific for this particular accession. Intergenomic translocations have not been identified in A. agadiriana using our selected probes, and the FISH pattern suggested that this species may be of autopolyploid origin thus its affiliation to the AABB group and potential drivers shaping this species remain debatable. Both our karyotype analysis and the data obtained by different authors (Jellen et al., 1994; Luo et al., 2018a; Luo et al., 2018b; Tomaszewska and Kosina, 2021; Yan et al., 2021; Peng et al., 2022) indicated that some intergenomic translocations could also influence the divergence of the CCDD Avena group. Some translocations between the C and D genomes in hexaploid oat species involved the same chromosomes as in tetraploid CCDD species (this paper; Badaeva et al., 2010; Yan et al., 2021) supporting the hypothesis that one of the species belonging to this oat group contributed to the evolution of hexaploid Avena species. Unfortunately, without knowing the exact ancestors of hexaploids, it is difficult to verify whether translocations which appeared in CCDD Avena species were transferred to AACCDD species through hybridization. Analysis of intergenomic translocations across different hexaploid oat species and cultivars indicated that the pattern of intergenomic translocation of hexaploid A. byzantina is different from that observed in other wild, weedy or cultivated hexaploid oat species (Badaeva et al., 2011), and also ‘Kanota’ and ‘Ogle’ cultivars differed from each other in at least some of the intergenomic translocations (Jellen et al., 1994). Therefore, in oats, three types of intergenomic translocations should be distinguished: common or group-specific, species-specific, and cultivar- or accession-specific (Sanz et al., 2010).
Intergenomic translocations and their implications for polyploid plant evolution
While some translocations are common for the Panicoideae group, Setaria, Saccharum, Sorghum, and Zea show further chromosome rearrangements which distinguish them from other grasses (Devos and Gale, 1997), and reciprocal translocations separating species within the Secale genus were revealed (Singh and Röbbelen, 1977). Some of the intergenomic translocations observed in wheat group are species-specific, as those observed in Triticum dicoccon, T. timopheevii and T. turgidum (Jiang and Gill, 1994; Levy and Feldman, 2002), supporting the diphyletic hypothesis of the evolution of tetraploid wheats. Another examples of species-specific intergenomic translocations are those seen in tetraploid Nicotiana tabacum (Kenton et al., 1993). A nucleocytoplasmic interaction (NCI) hypothesis of polyploid plant genome evolution explains the presence of species-specific intergenomic translocations (Jiang and Gill, 1994). It was speculated that the newly formed polyploid has to overcome a bottleneck of hybrid sterility resulting from cytoplasmic-nuclear interactions of paternal and maternal species. Some chromosomal and/or genome changes must occur to stabilize genomes and restore fertility in hybrids. Intergenomic chromosome translocations seem to be one of the major mechanisms stabilizing non-homologous genomes that come together in polyploid nuclei.
Intra- and inter-genomic translocations in plant breeding
The allopolyploid condition enables intra- and inter-genomic reshuffling of chromosomal segments and genes through translocations, recombination, transposition or introgressions (Kenton et al., 1993; Levy and Feldman, 2002; Patokar et al., 2016; Svačina et al., 2020; Li et al., 2021a; Martin et al., 2020; Tomaszewska and Kosina, 2021; Tomaszewska et al., 2022). The opportunity for chromosomal reshuffling has proved important in hexaploid wheat breeding to introduce alien chromosome segments from related species, creating new genetic combinations and genome arrangements (e.g. Ali et al., 2016). The fundamental research has implications for breeding oats if F1 hybrid combinations include undetected translocations leading to poor performance and restricting recombination, particularly in crosses made in oat breeding programmes between different accessions. In wheat, there are some reciprocal translocations which restrict some crosses (Zhang et al., 2022b). With interspecific crosses for exploitation of biodiversity in the Avena genus by introgression from wild species and recombination between chromosomes, chromosomal translocations between the species may make selection of genetically balanced lines more difficult. Agrawal et al. (2020) have developed pools of 20,000 oligonucleotides for use as probes in Brassica. By allowing identification of chromosome arms in chromosome preparations, these probes identified a reciprocal translocation present in one cultivar of Brassica rapa, cv. Purple Top Milan turnip, compared to cv. Chiifu-401 pak choi, which would make use of a hybrid difficult. It is likely that the translocations detected in the oat lines will mean care is required to use F1 hybrids in a breeding programme in case there are undetected translocations.
Genome sequence assemblies
New long-molecule sequencing approaches are allowing assembly of Avena genome sequences, covering nearly 4,000Mbp in the diploids (Maughan et al., 2019; Li et al., 2021b; Liu et al., 2022; Peng et al., 2022). High coverage shows the reciprocal translocations which are also revealed with the chromosomal probes. Currently it is not possible to survey large numbers of lines, and too complex to reconstruct with high coverage of the recombinant chromosomes of multiple species (Figure 9; chromosomal studies using in situ hybridization as Agrawal et al., 2020 or Huang et al., 2018 to identify translocations within species). It will be interesting to identify with base-pair resolution the chromosomal breakpoints, and further characterize the non-reciprocal nature of the translocations. We can then address key concern: what are the consequences of additional copies of alleles if segments are duplicated?
Conclusions
Tetraploid and hexaploid oat species, as well as hexaploid and octoploid synthetic hybrids and amphiploids are characterized by multiple distal intergenomic translocations, in contrast to the wheat-group species (wheat, rye, barley) that show mainly intragenomic translocations located in centromeric and interstitial regions of chromosomes. The presence of distal, mostly non-reciprocal translocations in the Avena group makes oat chromosomes particularly rearranged. It suggests that intergenomic translocations were a major mechanism of divergence in the evolution of oat species, and a new pattern of translocations is established in synthetic hybrids and amphiploids. The consequences of distal and interstitial intergenomic translocations for hybrid stability and gene expression from both genomes involved in translocation remain little known.
Data availability statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
Author contributions
PT and PH-H: funding acquisition. PT: investigation and methodology. PH-H: supervision. All authors: conceptualization, writing original draft review and editing, and have read and agreed to the published version of the manuscript.
Funding
This project was funded under the RCUK-CIAT Newton-Caldas Initiative, with funding from UK’s Official Development Assistance Newton Fund awarded by UK Biotechnology and Biological Sciences Research Council (BB/R022828/1), and by the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No 844564, individual fellowship to PT. This project has also received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No 101006417.
Acknowledgments
We are grateful to Bundesanstaltfür Züchtungsforschung an Kulturpflanzen (Braunschweig, Germany), National Small Grains Collection (Aberdeen, Idaho, USA) and Vavilov Institute of Plant Industry (St. Petersburg, Russia) for their generous provision of seeds.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Adams, K. L., Wendel, J. F. (2005). Polyploidy and genome evolution in plants. Curr. Opin. Plant Biol. 8 (2), 135–141. doi: 10.1016/j.pbi.2005.01.001
Agrawal, N., Gupta, M., Banga, S. S., Heslop-Harrison, J. S. (2020). Identification of chromosomes and chromosome rearrangements in crop brassicas and Raphanus sativus: A cytogenetic toolkit using synthesized massive oligonucleotide libraries. Front. Plant Sci. 11, 598039. doi: 10.3389/fpls.2020.598039
Ali, N., Heslop-Harrison, J. S., Ahmad, H., Graybosch, R. A., Hein, G. L., Schwarzacher, T. (2016). Introgression of chromosome segments from multiple alien species in wheat breeding lines with wheat streak mosaic virus resistance. Heredity 117 (2), 114–123. doi: 10.1038/hdy.2016.36
Alix, K., Gérard, P. R., Schwarzacher, T., Heslop-Harrison, J. S. (2017). Polyploidy and interspecific hybridization: Partners for adaptation, speciation and evolution in plants. Ann. Bot. 120 (2), 183–194. doi: 10.1093/aob/mcx079
Alix, K., Joets, J., Ryder, C. D., Moore, J., Barker, G. C., Bailey, J. P., et al. (2008). The CACTA transposon Bot1 played a major role in Brassica genome divergence and gene proliferation. Plant J. 56, 1030–1044. doi: 10.1111/j.1365-313X.2008.03660.x
Ananiev, E. V., Vales, M. I., Phillips, R. L., Rines, H. W. (2002). Isolation of A/D and c genome specific dispersed and clustered repetitive DNA sequences from Avena sativa. Genome 45 (2), 431–441. doi: 10.1139/g01-148
Badaeva, E. D., Dedkova, O. S., Gay, G., Pukhalskyi, V. A., Zelenin, A. V., Bernard, S., et al. (2007). Chromosomal rearrangements in wheat: Their types and distribution. Genome 50, 907–926. doi: 10.1139/G07-072
Badaeva, E. D., Jiang, J., Gill, B. S. (1995). Detection of intergenomic translocations with centromeric and noncentromeric breakpoints in Triticum araraticum: mechanism of origin and adaptive significance. Genome 38 (5), 976–981. doi: 10.1139/g95-128
Badaeva, E. D., Loskutov, I. G., Shelukhina, O. Y., Pukhalsky, V. A. (2005). Cytogenetic analysis of diploid Avena l. species containing the as genome. Russ. J. Genet. 41 (12), 1428–1433. doi: 10.1007/s11177-006-0018-3
Badaeva, E. D., Shelukhina, O. Y., Dedkova, O. S., Loskutov, I. G., Pukhalskyi, V. A. (2011). Comparative cytogenetic analysis of hexaploidy Avena l. species. Russ. J. Genet. 47 (6), 691–702. doi: 10.1134/S1022795411060068
Badaeva, E. D., Shelukhina, O. Y., Goryunova, S. V., Loskutov, I. G., Pukhalskiy, V. A. (2010). Phylogenetic relationships of tetraploid AB-genome Avena species evaluated by means of cytogenetic (C-banding and FISH) and RAPD analyses. J. Bot. 742307, 1–13. doi: 10.1155/2010/742307
Bardsley, D., Cuadrado, A., Jack, P., Harrison, G., Castilho, A., Heslop-Harrison, J. S. (1999). Chromosome markers in the tetraploid wheat Aegilops ventricosa analysed by in situ hybridization. Theor. Appl. Genet. 99 (1), 300–304. doi: 10.1007/s001220051236
Barker, M. S., Arrigo, N., Baniaga, A. E., Li, Z., Levin, D. A. (2016). On the relative abundance of autopolyploids and allopolyploids. New Phytol. 210, 391–398. doi: 10.1111/nph.13698
Baum, B. R. (1977). “Oats: wild and cultivated,” in A monograph of the genus avena l. (Poaceae) (Ottawa: Thorn Press Limited).
Biscotti, M. A., Olmo, E., Heslop-Harrison, J. S. (2015). Repetitive DNA in eukaryotic genomes. Chromosome Res. 23, 415–420. doi: 10.1007/s10577-015-9499-z
Biswas, M. K., Schwarzacher, T., Heslop-Harrison, J. S. (2022). “Large scale genome analysis: Genome sequences, chromosomal reorganization, and repetitive DNA in Brassica juncea and relatives,” in The Brassica juncea genome. Eds. Kole, C., Mohapatra, T. (Switzerland: Springer).
Braz, G. T., He, L., Zhao, H., Zhang, T., Semrau, K., Rouillard, J.-M., et al. (2018). Comparative oligo-FISH mapping: an efficient and powerful methodology to reveal karyotypic and chromosomal evolution. Genetics 208, 513–523. doi: 10.1534/genetics.117.300344
Bustos, A., Cuadrado, A., Soler, C., Jouve, N. (1996). Physical mapping of repetitive DNA sequences and 5S and 18S-26S rDNA in five wild species of the genus Hordeum. Chromosome Res. 4, 491–499. doi: 10.1007/BF02261776
Chen, Q., Armstrong, K. (1994). Genomic in situ hybridization in A. sativa. Genome 37, 607–612. doi: 10.1139/g94-086
Chew, P., Meade, K., Hayes, A., Harjes, C., Vao, Y., Beattie, A. D., et al. (2016). A study on the genetic relationshios of Avena taxa and the origins of hexaploid oat. Theor. Appl. Genet. 129, 1405–1415. doi: 10.1007/s00122-016-2712-4
Chumová, Z., Mandáková, T., Trávníček, P. (2021). On the origin of tetraploid vernal grasses (Anthoxanthum) in Europe. Genes 12, 966. doi: 10.3390/genes12070966
Comai, L., Madlung, A., Josefsson, C., Tyagi, A. (2003). Do the different parental 'heteromes' cause genomic shock in newly formed allopolyploids? Phil. Trans. R. Soc Lond. B 358, 1149–1155. doi: 10.1098/rstb.2003.1305
Devos, K. M., Gale, M. D. (1997). Comparative genetics in the grasses. Plant Mol. Biol. 35, 3–15. doi: 10.1023/A:1005820229043
Drossou, A., Katsiotis, A., Leggett, J. M., Loukas, M., Tsakas, S. (2004). Genome and species relationships in genus Avena based on RAPD and AFLP molecular markers. Theor. Appl. Genet. 109, 48–54. doi: 10.1007/s00122-004-1615-y
Fominaya, A., Loarce, Y., González, J. M., Ferrer, E. (2021). Cytogenetic evidence supports Avena insularis being closely related to hexaploid oats. PloS One 16 (10), e0257100. doi: 10.1371/journal.pone.0257100
Fu, Y.-B. (2018). Oat evolution revealed in the maternal lineages of 25 Avena species. Sci. Rep. 8, 4252. doi: 10.1038/s41598-018-22478-4
Fu, S., Tang, Z., Ren, Z. (2010). Inter- and intra-genomic transfer of small chromosomal segments in wheat-rye allopolyploids. J. Plant Res. 123, 97–103. doi: 10.1007/s10265-009-0264-2
Gernand, D., Rutten, T., Varshney, A., Rubtsova, M., Prodanovic, S., Brüss, C., et al. (2005). Uniparental chromosome elimination at mitosis and interphase in wheat and pearl millet crosses involves micronucleus formation, progressive heterochromatinization, and DNA fragmentation. Plant Cell 17, 2431–2438. doi: 10.1105/tpc.105.034249
Gnutikov, A. A., Nosov, N. N., Loskutov, I. G., Machs, E. M., Blinova, E. V., Probatova, N. S., et al. (2022). New insights into the genomic structure of the oats (Avena l., poaceae): Intragenomic polymorphism of ITS1 sequences of rare endemic species Avena bruhnsiana Gruner and its relationship to other species with C-genomes. Euphytica 218, 3. doi: 10.1007/s10681-021-02956-z
Heslop-Harrison, J. S., Schwarzacher, T. (2011). Organisation of the plant genome in chromosomes. Plant J. 66, 18–33. doi: 10.1111/j.1365-313X.2011.04544.x
Heslop-Harrison, J. S., Schwarzacher, T., Liu, Q. (2022). Polyploidy: its consequences and enabling role in plant diversification and evolution. Ann. Bot. mcac132. doi: 10.1093/aob/
Huang, K., Rieseberg, L. H. (2020). Frequency, origins, and evolutionary role of chromosomal inversions in plants. Front. Plant Sci. 11, 296. doi: 10.3389/fpls.2020.00296
Huang, X., Zhu, M., Zhuang, L., Zhang, S., Wang, J., Chen, X., et al. (2018). Structural chromosome rearrangements and polymorphisms identified in Chinese wheat cultivars by high-resolution multiplex oligonucleotide FISH. Theor. Appl. Genet. 131 (9), 1967–1986. doi: 10.1007/s00122-018-3126-2
Irigoyen, M. L., Loarce, Y., Linares, C., Ferrer, E., Leggett, M., Fominaya, A. (2001). Discrimination of the closely related A and B genomes in AABB tetraploid species of Avena. Theor. Appl. Genet. 103, 1160–1166. doi: 10.1007/s001220100723
Jellen, E. N., Gill, B. S. (1996). C-banding variation in the Moroccan oat species Avena agadiriana (2n=4x=28). Theor. Appl. Genet. 92, 726–732. doi: 10.1007/BF00226095
Jellen, E. N., Gill, B. S., Cox, T. S. (1994). Genomic in situ hybridization differentiates between A/D- and C-genome chromatin and detects intergenomic translocations in polyploid oat species (genus Avena). Genome 37, 613–618. doi: 10.1139/g94-087
Jiang, J., Gill, B. S. (1994). Different species-specific chromosome translocations in Triticum timopheevii and T. turgidum support the diphyletic origin of polyploid wheats. Chromosome Res. 2, 59–64. doi: 10.1007/BF01539455
Jiang, W., Jiang, C., Yuan, W., Zhang, M., Fang, Z., Li, Y., et al. (2021). A universal karyotypic system for hexaploid and diploid Avena species brings oat cytogenetics into the genomics era. BMC Plant Biol. 21, 213. doi: 10.1186/s12870-021-02999-3
Katsiotis, A., Loukas, M., Heslop-Harrison, J. S. (2000). Repetitive DNA, genome and species relationships in Avena and Arrhenatherum. Ann. Bot. 86, 1135–1142. doi: 10.1006/anbo.2000.1284
Kenton, A., Parokonny, A. S., Gleba, Y. Y., Bennett, M. D. (1993). Characterization of the Nicotiana tabacum L. genome by molecular cytogenetics. Mol. Gen. Genet. 240, 159–169. doi: 10.1007/BF00277053
Kopecký, D., Loureiro, J., Zwierzykowski, Z., Ghesquière, M., Doležel, J. (2006). Genome constitution and evolution in Lolium × Festuca hybrid cultivars (Festulolium). Theor. Appl. Genet. 113, 731–742. doi: 10.1007/s00122-006-0341-z
Kosina, R., Heslop-Harrison, J. S. (1996). Molecular cytogenetics of an amphiploid trigeneric hybrid between Triticum durum, Thinopyrum distichum and Lophopyrum elongatum. Ann. Bot. 78 (5), 583–589. doi: 10.1006/anbo.1996.0164
Kubaláková, M., Valárik, M., Bartoš, J., Vrána, J., Cíhalíková, J., Molnár-Láng, M., et al. (2003). Analysis and sorting of rye (Secale cereale l.) chromosomes using flow cytometry. Genome 46, 893–905. doi: 10.1139/g03-054
Latta, R. G., Bekele, W. A., Wight, C. P., Tinker, N. A. (2019). Comparative linkage mapping of diploid, tetraploid, and hexaploidy Avena species suggests extensive chromosome rearrangement in ancestral diploids. Sci. Rep. 9, 12298. doi: 10.1038/s41598-019-48639-7
Leggett, J. M. (1998). Chromosome and genomic relationships between the diploid species Avena strigosa, A. eriantha and the tetraploid A. maroccana. Heredity 80, 361–363. doi: 10.1046/j.1365-2540.1998.00305.x
Leggett, J. M., Thomas, H. M., Meredith, M. R., Humphreys, M. W., Morgan, W. G., Thomas, J. M., et al. (1994). Intergenomic translocations and the genomic composition of Avena maroccana Gdgr. revealed by FISH. Chromosome Res. 2, 163–164. doi: 10.1007/BF01553495
Levy, A. A., Feldman, M. (2002). The impact of polyploidy on grass genome evolution. Plant Physiol. 130, 1587–1593. doi: 10.1104/pp.015727
Li, G., Zhang, T., Yu, Z., Wang, H., Yang, E., Yang, Z. (2021a). An efficient oligo-FISH painting system for revealing chromosome rearrangements and polyploidization in Triticeae. Plant J. 105, 978–993. doi: 10.1111/tpj.15081
Li, Y., Leveau, A., Zhao, Q., Feng, Q., Lu, H., Miao, J., et al. (2021b). Subtelomeric assembly of a multi-gene pathway for antimicrobial defense compounds in cereals. Nat. Commun. 12 (1), 1–13. doi: 10.1038/s41467-021-22920-8
Linares, C., Ferrer, E., Fominaya, A. (1998). Discrimination of the closely related a and d genomes of the hexaploid oat Avena sativa l. Proc. Natl. Acad. Sci. U.S.A. 95 (21), 12450–12455. doi: 10.1073/pnas.95.21.12450
Linares, C., Irigoyen, M. L., Fominaya, A. (2000). Identification of c-genome chromosomes involved in intergenomic translocations in Avena sativa l., using cloned repetitive DNA sequences. Theor. Appl. Genet. 100, 353–360. doi: 10.1007/s001220050046
Liu, Q., Li, X., Li, M., Xu, W., Schwarzacher, T., Heslop-Harrison, J. S. (2020). Comparative chloroplast genome analyses of Avena: insights into evolutionary dynamics and phylogeny. BMC Plant Biol. 20, 406. doi: 10.1186/s12870-020-02621-y
Liu, Q., Li, X., Zhou, X., Li, M., Zhang, F., Schwarzacher, T., et al. (2019). The repetitive DNA landscape in Avena (Poaceae): chromosome and genome evolution defined by major repeat classes in whole-genome sequence reads. BMC Plant Biol. 19, 226. doi: 10.1186/s12870-019-1769-z
Liu, Q., Yuan, H., Li, M., Wang, Z., Cui, D., Ye, Y., et al. (2022). Chromosome-scale genome assembly of the diploid oat Avena longiglumis reveals the landscape of repetitive sequences, genes and chromosome evolution in grasses. doi: 10.1101/2022.02.09.479819
Loskutov, I. G. (2001). Interspecific crosses in the genus Avena l. Russ. J. Genet. 37, 467–475. doi: 10.1023/A:1016697812009
Loskutov, I. G. (2007). On evolutionary pathways of Avena species. Genet. Resour. Crop Evol. 55, 211–220. doi: 10.1007/s10722-007-9229-2
Loskutov, I. G., Gnutikov, A. A., Blinova, E. V., Rodionov, A. V. (2021). The origin and resource potential of wild and cultivated species of the genus of oats (Avena l.). Russ. J. Genet. 57 (6), 642–661. doi: 10.1134/S1022795421060065
Luo, X., Tinker, N. A., Zhou, Y., Liu, J., Wan, W., Chen, L. (2018a). A comparative cytogenetic study of 17 Avena species using Am1 and (GAA)6 oligonucleotide FISH probes. Acta Physiol. Plant 40, 145. doi: 10.1007/s11738-018-2721-9
Luo, X., Tinker, N. A., Zhou, Y., Liu, J., Wan, W., Chen, L. (2018b). Chromosomal distributions of oligo-Am1 and (TTG)6 trinucleotide and their utilization in genome association analysis of sixteen Avena species. Genet. Resour. Crop Evol. 65, 1625–1635. doi: 10.1007/s10722-018-0639-0
Luo, X., Tinker, N. A., Zhou, Y., Wight, C. P., Liu, J., Wan, W., et al. (2018c). Genomic relationship among sixteen species of Avena based on (ACT)6 trinucleotide repeat FISH. Genome 61, 63–70. doi: 10.1139/gen-2017-0132
Luo, X., Zhang, H., Kang, H., Fan, X., Wang, Y., Sha, L., et al. (2014). Exploring the origin of the d genome of oat by fluorescence in situ hybridization. Genome 57 (9), 469–472. doi: 10.1139/gen-2014-0048
Martin, G., Baurens, F.-C., Hervouet, C., Salmon, F., Delos, J.-M., Labadie, K., et al. (2020). Chromosome reciprocal translocations have accompanied subspecies evolution in bananas. Plant J. 104, 1698–1711. doi: 10.1111/tpj.15031
Maughan, P. J., Lee, R., Walstead, R., Vickerstaff, R. J., Fogarty, M. C., Brouwer, C. R., et al. (2019). Genomic insights from the first chromosome-scale assemblies of oat (Avena spp.) diploid species. BMC Biol. 17, 92. doi: 10.1186/s12915-019-0712-y
McClintock, B. (1984). The significance of responses of the genome to challenge. Science 226, 792–801. doi: 10.1126/science.15739260
Mehrotra, S., Goyal, V. (2014). Repetitive sequences in plant nuclear DNA: types, distribution, evolution and function. Genom. Proteom. Bioinform. 12 (4), 164–171. doi: 10.1016/j.gpb.2014.07.003
Mirzaghaderi, G., Marzangi, K. (2015). IdeoKar: an ideogram constructing and karyotype analyzing software. Caryologia 68 (1), 31–35. doi: 10.1080/00087114.2014.998526
Molnár, I., Benavente, E., Molnár-Láng, M. (2009). Detection of intergenomic chromosome rearrangements in irradiated Triticum aestivum-Aegilops biuncialis amphiploids by multicolour genomic in situ hybridization. Genome 52, 156–165. doi: 10.1139/G08-114
Mondal, S., Rutkoski, J. E., Velu, G., Singh, P. K., Crespo-Herrera, L. A., Guzmán, C., et al. (2016). Harnessing diversity in wheat to enhance grain yield, climate resilience, disease and insect pest resistance and nutrition through conventional and modern breeding approaches. Front. Plant Sci. 7, 991. doi: 10.3389/fpls.2016.00991
Nagy, E. D., Molnár-Láng, M., Linc, G., Láng, L. (2002). Identification of wheat-barley translocations by sequential GISH and two-colour FISH in combination with the use of genetically mapped barley SSR markers. Genome 45, 1238–1247. doi: 10.1139/g02-068
Naish, M., Alonge, M., Wlodzimierz, P., Tock, A. J., Abramson, B. W., Schmücker, A., et al. (2021). The genetic and epigenetic landscape of the Arabidopsis centromeres. Science 376 (6588), 1–9. doi: 10.1126/science.abi7489
Newell, M. A., Asoro, F. G., Scott, M. P., White, P. J., Beavis, W. D., Jannink, J.-L. (2012). Genome-wide association study for oat (Avena sativa l.) beta-glucan concentration using germplasm of worldwide origin. Theor. Appl. Genet. 125, 1687–1696. doi: 10.1007/s00122-012-1945-0
Nikoloudakis, N., Katsiotis, A. (2015). Comparative molecular and cytogenetic methods can clarify meiotic incongruities in Avena allopolyploid hybrids. Caryologia 68, 84–91. doi: 10.1080/00087114.2015.1021170
Nishiyama, I. (1962). Cytogenetic studies in Avena, IX. new synthetic oats in the progenies of induced decaploid interspecific hybrids. Jap. J. Genet. 37, 118–130. doi: 10.1266/jjg.37.118
Nishiyama, I., Tabata, M., Iizuka, M. (1963). Cytogenetic studies in Avena, XI. chromosomal constitution of the inbred progeny of synthetic amphiploid oats. Jap. J. Genet. 38, 123–134. doi: 10.1266/jjg.38.123
Nishiyama, I., Yabuno, T. (1979). Triple fusion of the primary endosperm nucleus as a cause of interspecific cross-incompatibility in Avena. Euphytica 28, 57–65. doi: 10.1007/BF00029173
Patel, D., Power, J. B., Anthony, P., Badakshi, F., Heslop-Harrison, J. S., Davey, M. R. (2011). Somatic hybrid plants of Nicotiana x sanderae (+) N. debneyi with fungal resistance to Peronospora tabacina. Ann. Bot. 108 (5), 809–819. doi: 10.1093/aob/mcr197
Patokar, C., Sepsi, A., Schwarzacher, T., Kishii, M., Heslop-Harrison, J. S. (2016). Molecular cytogenetic characterization of novel wheat-Thinopyrum bessarabicum recombinant lines carrying intercalary translocations. Chromosoma 125, 163–172. doi: 10.1007/s00412-015-0537-6
Peng, Y.-Y., Wei, Y.-M., Baum, B. R., Yan, Z.-H., Lan, X.-J., Dai, S. F., et al. (2010). Phylogenetic inferences in Avena based on analysis of FL intron2 sequences. Theor. Appl. Genet. 121, 985–1000. doi: 10.1007/s00122-010-1367-9
Peng, Y., Yan, H., Guo, L., Deng, C., Wang, C., Wang, Y., et al. (2022). Reference genome assemblies reveal the origin and evolution of allohexaploid oat. Nat. Genet. 54, 1248–1258. doi: 10.1038/s41588-022-01127-7
Peng, Y., Zhou, P., Zhao, J., Li, J., Lai, S., Tinker, N. A., et al. (2018). Phylogenetic relationships in the genus Avena based on the nuclear Pgk1 gene. PloS One 13 (11), e0200047. doi: 10.1371/journal.pone.0200047
Prieto, P., Ramírez, M. C., Ballesteros, J., Cabrera, A. (2001). Identification of intergenomic translocations involving wheat, Hordeum vulgare and Hordeum chilense chromosomes by FISH. Hereditas 135, 171–174. doi: 10.1111/j.1601-5223.2001.t01-1-00171.x
Qi, L., Friebe, B., Gill, B. S. (2006). Complex genome rearrangements reveal evolutionary dynamics of pericentromeric regions in the Triticeae. Genome 49, 1628–1639. doi: 10.1139/g06-123
Rajhathy, T., Thomas, H. (1974). Cytogenetics of oats (Avena l.). Misc. Publ. Genet. Soc Can. 2, 1–90.
Rathore, P., Schwarzacher, T., Heslop-Harrison, J. S., Bhat, V., Tomaszewska, P. (2022). The repetitive DNA sequence landscape and DNA methylation in chromosomes of an apomictic tropical forage grass, Cenchrus ciliaris Front. Plant Sci. 13:952968. doi: 10.3389/fpls.2022.952968
Søgaard, B., von Wettstein-Knowles, P. (1987). Barley: Genes and chromosomes. Carlsberg Res. Commun. 52, 123–196. doi: 10.1007/BF02907531
Sanz, M. J., Jellen, E. N., Loarce, Y., Irigoyen, M. L., Ferrer, E., Fominaya, A. (2010). A new chromosome nomenclature system for oat (Avena sativa l. and A. byzantina C. Koch) based on FISH analysis of monosomic lines. Theor. Appl. Genet. 121, 1541–1552. doi: 10.1007/s00122-010-1409-3
Schwarzacher, T., Heslop-Harrison, J. S. (2000). Practical in situ hybridization (Oxford, UK: Bios).
Shelukhina, O. Y., Badaeva, E. D., Loskutov, I. G., Pukhalsky, V. A. (2007). A comparative cytogenetic study of the tetraploid oat species with the A and C genomes: Avena insularis, A. magna and A. murphyi. Russ. J. Genet. 43 (6), 613–626. doi: 10.1134/S102279540706004X
Singh, R. J., Röbbelen, G. (1977). Identification by Giemsa technique of the translocations separating cultivated rye from three wild species of. Secale. Chromosoma 59, 217–225. doi: 10.1007/BF00292779
Soltis, P. S., Marchant, D. B., Van de Peer, Y., Soltis, D. E. (2015). Polyploidy and genome evolution in plants. Curr. Opin. Genet. Dev. 35, 119−125. doi: 10.1016/j.gde.2015.11.003
Svačina, R., Sourdille, P., Kopecký, D., Bartoš, J. (2020). Chromosome pairing in polyploid grasses. Front. Plant Sci. 11, 1056. doi: 10.3389/fpls.2020.01056
Taketa, S., Ando, H., Takeda, K., von Bothmer, R. (1999). Detection of Hordeum marinum genome in three polyploid Hordeum species and cytotypes by genomic in situ hybridization. Hereditas 130, 185–188. doi: 10.1111/j.1601-5223.1999.00185.x
Tang, S., Tang, Z., Qiu, L., Yang, Z., Li, G., Lang, T., et al. (2018). Developing new oligo probes to distinguish specific chromosomal segments and the A, B, D genomes of wheat (Triticum aestivum L.) using ND-FISH. Front. Plant Sci. 9, 1104. doi: 10.3389/fpls.2018.01104
Tomaszewska, P. (2021). Understanding polyploid banana origins. a commentary on: 'Unravelling the complex story of intergenomic recombination in ABB allotriploid bananas'. Ann. Bot. 127 (1), iv–iv. doi: 10.1093/aob/mcaa183
Tomaszewska, P., Kosina, R. (2018). Instability of endosperm development in amphiploids and their parental species in the genus Avena l. Plant Cell Rep. 37, 1145–1158. doi: 10.1007/s00299-018-2301-x
Tomaszewska, P., Kosina, R. (2021). Cytogenetic events in the endosperm of amphiploid Avena magna x A. longiglumis. J. Plant Res. 134, 1047–1060. doi: 10.1007/s10265-021-01314-3
Tomaszewska, P., Kosina, R. (2022). Variability in the quality of pollen grains in oat amphiploids and their parental species. Braz. J. Bot. 45, 987–1000. doi: 10.1007/s40415-022-00822-3
Tomaszewska, P., Vorontsova, M. S., Renvoize, S. A., Ficinski, S. Z., Tohme, J., Schwarzacher, T., et al. (2022). Complex polyploid and hybrid species in an apomictic and sexual tropical forage grass group: Genomic composition and evolution in Urochloa (Brachiaria) species. Ann. Bot. doi: 10.1093/aob/mcab147 in press
Wang, Z., Rouard, M., Biswas, M. K., Droc, G., Cui, D., Roux, N., et al. (2022). A chromosome-level reference genome of ensete glaucum gives insight into diversity, chromosomal and repetitive sequence evolution in the Musaceae. GigaScience 11, 1–21. doi: 10.1093/gigascience/giac027
Yan, H.-H., Baum, B. R., Zhou, P.-P., Zhao, J., Wei, Y.-M., Ren, C. Z., et al. (2014). Phylogenetic analysis of the genus Avena based on chloroplast intergenic spacer psbA-trnH and single-copy nuclear gene Acc1. Genome 57, 267–277. doi: 10.1139/gen-2014-0075
Yan, H., Bekele, W. A., Wight, C. P., Peng, Y., Langdon, T., Latta, R. G., et al. (2016). High-density marker profiling confirms ancestral genomes of Avena species and identifies D-genome chromosomes of hexaploid oat. Theor. Appl. Genet. 129, 2133–2149. doi: 10.1007/s00122-016-2762-7
Yan, H., Ren, Z., Deng, D., Yang, K., Yang, C., Zhou, P., et al. (2021). New evidence confirming the CD genomic constitutions of the tetraploid Avena species in the section Pachycarpa Baum. PloS One 16 (1), e0240703. doi: 10.1371/journal.pone.0240703
Zhang, F., Chen, F., Schwarzacher, T., Heslop-Harrison, J. S., Teng, N. (2022a). The nature and genomic landscape of repetitive DNA classes in Chrysanthemum nankingense shows recent genomic changes. Ann. Bot. doi: 10.1093/aob/mcac066 in press
Zhang, W., Tang, Z., Luo, J., Li, G., Yang, Z., Labadie, K., Yang, M., et al. (2022b). Location of tandem repeats on wheat chromosome 5B and the breakpoint on the 5BS arm in wheat translocation T7BS. 7BL-5BS using single-copy FISH analysis. Plants 11 (18), 2394. doi: 10.3390/plants11182394
Keywords: intergenomic translocations, chromosome rearrangements, structural variation, genome identification, repetitive DNA motifs, polyploidy, amphiploids, Avena
Citation: Tomaszewska P, Schwarzacher T and Heslop-Harrison JS(P) (2022) Oat chromosome and genome evolution defined by widespread terminal intergenomic translocations in polyploids. Front. Plant Sci. 13:1026364. doi: 10.3389/fpls.2022.1026364
Received: 23 August 2022; Accepted: 14 October 2022;
Published: 22 November 2022.
Edited by:
André Luís Laforga Vanzela, State University of Londrina, BrazilReviewed by:
Zujun Yang, University of Electronic Science and Technology of China, ChinaAndras Cseh, ELKH Agricultural Institute, Centre for Agricultural Research, Hungary
Joanna Majka, Academy of Sciences of the Czech Republic, Czechia
Copyright © 2022 Tomaszewska, Schwarzacher and Heslop-Harrison. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Paulina Tomaszewska, cHQxODZAbGVpY2VzdGVyLmFjLnVr; cGF1bGluYS50b21hc3pld3NrYUB1d3IuZWR1LnBs; cGhoNEBsZWljZXN0ZXIuYWMudWs=
†ORCID: Paulina Tomaszewska, orcid.org/0000-0002-9596-7219
Trude Schwarzacher, orcid.org/0000-0001-8310-5489
Pat (J.S) Heslop-Harrison, orcid.org/0000-0002-3105-2167
 Paulina Tomaszewska
Paulina Tomaszewska Trude Schwarzacher
Trude Schwarzacher J. S. (Pat) Heslop-Harrison1,3†
J. S. (Pat) Heslop-Harrison1,3†