- 1State Key Laboratory of Crop Stress Resistance and High-Efficiency Production, College of Agronomy, Northwest A&F University, Yangling, Shaanxi, China
- 2State Key Laboratory of Crop Gene Resources and Breeding/National Key Facility for Crop Gene Resources and Genetic Improvement, Institute of Crop Sciences, Chinese Academy of Agricultural Sciences, Beijing, China
Rice bacterial leaf blight (BB), caused by Xanthomonas oryzae pv. oryzae (Xoo), leads to severe yield losses in rice. Resistance breeding is a sustainable approach to mitigate the impact of this disease. In this study, a novel BB resistance gene, Xa50(t), was identified in the germplasm line CX315. Genetic analysis revealed that the resistance is conferred by a dominant resistant gene, tentatively named Xa50(t), which provides broad-spectrum and robust resistance to multiple Xoo strains. Using an F2 population derived from a cross between CX315 and IR24, Xa50(t) was fine-mapped to a 147.7 kb region on the long arm of chromosome 11 flanked by InDel markers M11–588 and M11-602. Gene expression analysis identified three candidate genes out of 13 open reading frames (ORFs) predicted in candidate region. ORF5 and ORF9 (Xa4), encoding a wall-associated kinase (WAK)-like protein, and ORF13, encoding a receptor-like kinase protein, were significantly upregulated in CX315 following Xoo inoculation. While ORF9 is predicted to encode the Xa4 resistance gene, CRISPR/Cas9-based knockout of Xa4 did not abolish Xa50(t)-mediated resistance in the CX315 line, indicating that Xa50(t) confers resistance in a complementary manner. Transcriptome analysis further revealed that oxidative stress response and immune signaling pathways were enriched in CX315 at 48 hours post-inoculation. Together, these findings highlight the potential of Xa50(t) as a valuable genetic resource for improving BB resistance in rice, and the transcriptome data provides molecular insight into the BB resistance response.
1 Introduction
Bacterial blight (BB), caused by Xanthomonas oryzae pv. oryzae (Xoo), is one of the most devastating diseases in rice production, leading to yield losses exceeding 50% in susceptible varieties (Timilsina et al., 2020; Wang et al., 2025). Genetic resistance offers a sustainable strategy for controlling Xoo infection, but pathogen evolution can overcome existing resistance mechanisms (Ji et al., 2016; Xu et al., 2022). Therefore, identifying new resources of resistance and development of genetically resistant cultivars is critical for controlling the disease. A thorough understanding of the genetic basis of resistance is essential to maximize its potential in rice breeding programs (Ji et al., 2018; Jiang et al., 2020). Thus, continuous efforts to discover new resistance genes, particularly those effective against emerging Xoo pathotypes, remained a priority.
Germplasm collections, including wild relatives, traditional landraces and elite accessions, offer a rich source of genetic diversity for resistance breeding efforts (Tirnaz et al., 2022). These materials often harbor unique resistance traits, making them valuable genetic resources for managing emerging pathogen strains and improving the durability of resistance in cultivated varieties. Recent studies have underscored the potential of rice germplasm in uncovering novel trait related and BB resistance genes (Wang et al., 2023). For example, a diverse panel of rice germplasm from China was evaluated and found to contain accessions with broad-spectrum resistance to Xoo strains, even in the absence of previously characterized resistance genes (Lu et al., 2021). Moreover, underutilized germplasm has revealed novel quantitative trait loci (QTLs) associated with disease resistance (Fang et al., 2023). These findings emphasize the unexplored potential of germplasm collections in overcoming the limitations of existing resistance genes.
Rice BB resistance is primarily controlled by resistance (R) genes that detect pathogens and activate defense responses (Kumar et al., 2020). Over the past decades, intensive efforts in rice genetics have led to the identification and functional characterization of multiple R genes through approaches such as positional cloning, genome-wide association studies (GWAS), mutant screening, and transcriptome profiling. These R genes operate via a range of mechanisms, including direct or indirect recognition of pathogen effectors, activation of transcriptional reprogramming, and modulation of defense-related hormone pathways. For example, Xa21 encodes a receptor-like kinase that perceives conserved pathogen-associated molecular patterns, while Xa23 and Xa27 act as executor genes triggered by pathogen effectors to induce localized cell death. To enhance resistance durability and counteract pathogen evolution, gene pyramiding strategies have been employed to combine multiple R genes such as Xa4, Xa21, and Xa23 into elite cultivars (Yugander et al., 2018; Wang et al., 2020). This has led to the development of rice varieties with broad-spectrum and long-lasting resistance. Another class, wall-associated kinase (WAK)-type R genes have shown strong potential for inclusion in such pyramiding strategies due to their unique extracellular sensing and broad defensive functions (Dossa et al., 2020). WAK family members are promising targets for understanding BB resistance and improving rice cultivars.
WAKs have emerged as important R genes in plant immunity, acting as molecular guards that monitor cell wall integrity during pathogen attack (Delteil et al., 2016; Harkenrider et al., 2016). These proteins integrate extracellular signals of pathogen invasion with intracellular defense responses, making them essential for both disease resistance and stress adaptation (Stephens et al., 2022). The Xa4 gene, one of the most widely deployed BB resistance genes in rice (Luo et al., 2012), encodes a WAK-like protein that provides durable resistance to multiple Xoo strains and enhances resistance by reinforcing the cell wall remodeling cellulose synthesis (Hu et al., 2017). However, while significant progress has been made in understanding its role in defense, the molecular mechanisms and functional diversity within the WAK family remain underexplored, highlighting the need for further investigation into their potential for improving both biotic and abiotic stress tolerance.
Recent advances in genome editing, particularly CRISPR/Cas9 technology, have revolutionized functional genomics by enabling precise knockout of candidate genes to validate their role in disease resistance (Langner et al., 2018). Coupled with high-throughput RNA sequencing (RNA-seq), these tools allow comprehensive profiling of transcriptional changes in response to pathogen infection and gene disruption (Du et al., 2024). This integrative approach provides deep insights into the molecular mechanisms underlying resistance, helping to distinguish gene-specific effects from broader immune responses. In present study, we utilized CRISPR-mediated knockout of the Xa4 gene and transcriptome analyses to dissect the independent function of the novel resistance gene Xa50(t) in the CX315 germplasm.
Our research screened rice germplasm and identified rice germplasm accession CAAS-11X315 (CX315), which exhibited strong resistance to multiple Xoo isolates. To fine map the genetic resistance, we utilized F2 population from CX315 × IR24 and fine-mapped the resistance gene to a 147.7 kb region on chromosome 11 and tentatively names it as Xa50(t). Quantitative real time PCR (qPCR) and RNA-seq validation showed three genes significantly upregulated after Xoo inoculation. Transcriptome analysis revealed enrichment of oxidative stress and immune signaling pathways at 48 hours post-inoculation. Importantly, knockout of Xa4 did not affect Xa50(t)-mediated resistance, indicating Xa50(t) acts independently of Xa4.
2 Material and methods
2.1 Plant materials and growth conditions
The rice germplasm line CAAS-11308X315 (CX315) was sourced from the Germplasm Resources Center of CAAS and identified through systematic resistance screening against diverse Xoo strains under controlled inoculation conditions. Two susceptible rice cultivars, IR24 and JG30 used from laboratory-maintained materials as controls in comparison with the resistant line CX315. Statistical analysis of the data was performed using a two-tailed Student’s t-test. Plants were cultivated at CAAS experimental fields located in Beijing and Sanya, Hainan, under net house or open field conditions, with conventional water and fertilizer management practices.
2.2 Bacterial strains and inoculation
The Xoo strains used to assess the resistance of CX315 to BB included races from China and other countries. These strains included PXO61, PXO86, PXO79, PXO71, PXO112, PXO99A, PXO280, PXO339, and PXO341 from the Philippines; T7174, T7147, and T7133 from Japan; KXO85 and KXO576 from Korea; and HLJ72, HB17, NX42, ZHE173, GD1358, LN57, JS49-6, GX15-2, Yun17-3, LN44, and others from China. Xoo strains were cultured in NA medium (Polypeptone 5g/L, sucrose 10g/L, yeast extract 1g/L and beef extract 3g/L, Bacto agar 15g/L (for solid media)) at 28°C for three days, then resuspended in sterile distilled water and maintained OD600 of 1.0 (measured using Eppendorf spectrometer 1000). Inoculation was performed using either the leaf-clipping method or needleless syringe infiltration, with lesion lengths measured at 14 days post-inoculation (dpi) for leaf clipping and 5 dpi for infiltration.
2.3 Genetic analysis and fine mapping of Xa50(t)
The susceptible rice cultivar IR24 was crossed with the resistant germplasm line CX315 to develop an F2 mapping population through selfing of heterozygous F1 plants. Both F1 and F2 populations were phenotypically screened for resistance or susceptibility following inoculation with T7174 strain. Segregation patterns in the F2 population were analyzed using the Chi-square test (χ²)to determine the genetic basis of resistance. Polymorphic insertion-deletion (InDel) markers were developed based on the Shuhui498 reference indica genome (https://mbkbase.org/R498/) and used to identify genetic differences between the parents IR24 and CX315 (Supplementary Table 2). Specific primers targeting these polymorphic regions were designed using the Shuhui498 genome sequence. Genomic DNA was extracted from F2 individuals using the CTAB method. PCR amplification with polymorphic InDel markers was performed on these DNA samples, and PCR products were resolved by electrophoresis on 5% agarose gels to detect marker segregation. Linkage analysis was then conducted to map candidate genes associated with resistance.
2.4 RNA extraction and quantitative real-time PCR
Total RNA was extracted from plant samples using the Trizol reagent according to the manufacturer’s protocol (Rio et al., 2010). The extracted RNA was then reverse-transcribed into first-strand cDNA using the TIANScript-II RT Kit (Tiangen Biotech, Beijing). Quantitative real-time PCR (qRT-PCR) was performed using the Taq Pro Universal SYBR qPCR Master Mix (Vazyme) on an ABI 7500 Real-Time PCR System (Thermo Fisher Scientific, USA). The sequences of primers used for qRT-PCR are given in Supplementary Table 5. The 20μL reaction mixture contained 10μL of 2X SYBR Master Mix, 0.4μL each of forward and reverse primer, 2μL of cDNA template, and 7.2μL of nuclease-free water. The PCR program consisted of an initial denaturation at 95°C for 3 minutes, followed by 40 cycles of 95°C for 5 seconds and 60°C for 34 seconds. Relative gene expression levels were calculated using the 2-ΔΔCt method (Livak and Schmittgen, 2001), with the rice Actin gene serving as the internal reference.
2.5 Development of Xa4-KO CRISPR vector and transformation
Target sequences within the exon of the Xa4 gene were selected using the TargetDesign tool (http://skl.scau.edu.cn/targetdesign/). These sequences were ligated into sgRNA expression cassettes via overlapping PCR to generate two sgRNAs, sgRNA-U6a-T1 and sgRNA-U6b-T2 (Supplementary Table 4). The sgRNA cassettes were cloned into the binary vector pYLCRISPR-Cas9-Xa4-KO using Golden Gate cloning (Ma et al., 2015). Agrobacterium-mediated transformation was performed to introduce the CRISPR construct into rice calli. Genomic DNA was extracted from transformed plant leaves using the CTAB method and used as template for PCR amplification with Xa4-KO-seq primers listed in Supplementary Table 4. Sanger sequencing of the PCR products used for identification of knockout mutations in Xa4.
2.6 RNA-sequencing
Total RNA was extracted from rice leaf tissues using the Trizol reagent, and RNA quality was assessed using a Nanodrop spectrophotometer and Bioanalyzer. RNA libraries were prepared using the TrueSeq RNA Library Prep Kit (Illumina) with fragmentation, reverse transcription, and amplification steps, followed by sequencing on an Illumina NovaSeq 6000 platform with paired-end 150 bp reads. Raw reads were processed to remove low-quality bases and adapter sequences using Trimmomatic and aligned to the rice reference genome Shuhui498 using STAR. Gene expression levels were quantified with feature Counts and analyzed for differential expression using DESeq2, considering a fold change ≥ 2 and an FDR < 0.05. Functional annotation of differentially expressed genes was performed using the NCBI Nr and GO databases, with KEGG pathway analysis to identify enriched biological processes. RNA-seq data was submitted to NCBI SRA database under project PRJNA1209632.
3 Results
3.1 CX315 confers broad-spectrum and genetically controlled resistance to Xoo
Initial screening of germplasm lines identified CX315 as highly resistant line to Xoo strains from the Philippines, China, Japan, and Korea. To ascertain the genetic resistance, CX315 and the susceptible cultivar JG30 were inoculated at the tillering stage with 21 representative Xoo strains. CX315 exhibited significantly robust resistance to all strains, with lesion lengths ranging from 0.1 to 6 cm, whereas JG30 showed consistent susceptibility, with lesion lengths exceeding 7 cm (Figure 1A). These findings highlight CX315 as a promising source of broad-spectrum resistance for bacterial blight in rice.
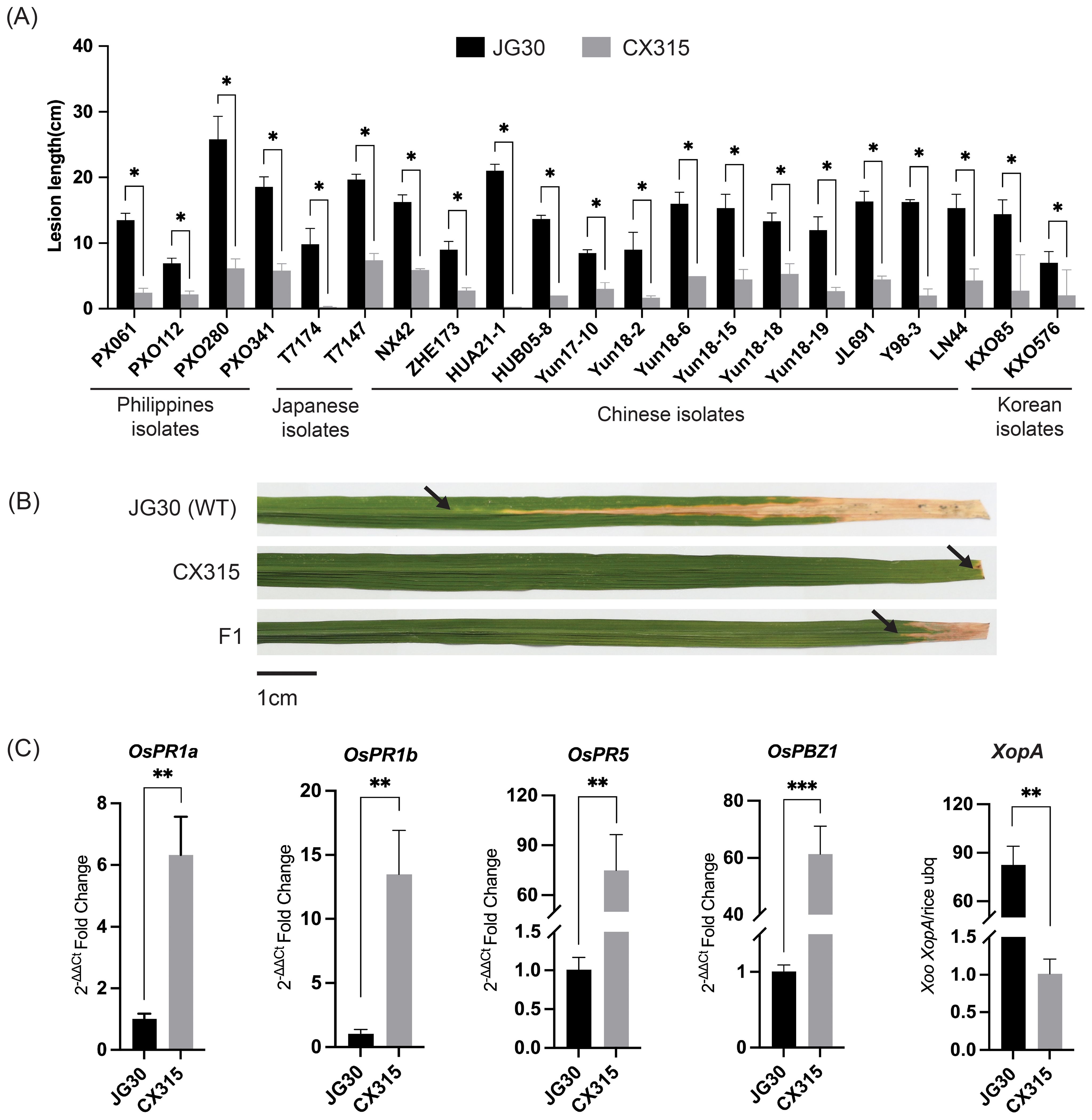
Figure 1. Phenotypic and molecular characterization of resistance conferred by CX315 to Xoo. (A) Lesion length comparison between the resistant line CX315 and the susceptible parent JG30 across different Xoo strains. CX315 exhibited significantly reduced lesion lengths compared to JG30 for all tested strains. Asterisks (*) indicate significant differences (*p< 0.05, **p< 0.01) between CX315 and JG30 as determined by Student’s t-test. (B) Representative images of lesion development on leaves of JG30, CX315, and the F1 hybrid plants 14 days post inoculation with Xoo strain T7174. JG30 leaves exhibit severe lesions, while CX315 and F1 plants demonstrate resistance with shorter lesions. Black arrows highlight the lesion sites, and a scale bar of 1 cm is provided for reference. (C) Differential expression of defense-related genes (OsPR1a, OsPR1b, OsPR5, OsPBZ1) and the Xoo effector gene XopA in JG30 and CX315 after inoculation with T7174. CX315 shows significantly higher upregulation of these genes compared to JG30, indicating activation of defense pathways. Conversely, the reduced abundance of XopA in CX315 suggests a significantly lower bacterial load in the resistant line.
To determine whether CX315’s resistance is genetically controlled, we crossed CX315 (♂) and the susceptible cultivar IR24 (♀), and the resulting F1 plants were evaluated after inoculation with the Xoo strain T7174. The F1 plants exhibited resistance comparable to CX315, as evidenced by shorter lesion lengths (Figure 1B). Evaluation of F2 population informed 1,767 plants exhibited resistance, while 555 plants were susceptible, fitting a 3:1 Mendelian segregation ratio (χ² = 1.4935, p = 0.222), thereby confirming that resistance in CX315 is controlled by a single dominant gene (Supplementary Table 1). To further validate that this resistance is not only genetically inherited but also associated with active defense responses, we analyzed the expression of key pathogenesis-related (PR) genes following pathogen challenge. PR genes are molecular markers of plant immune activation, and their upregulation reflects the activation of basal and systemic defense pathways. qRT-PCR results showed significant induction of OsPR1a, OsPR1b, OsPR5, and OsPBZ1 in CX315 compared to JG30 (Figure 1C), indicating that CX315 confers a strong defense response upon infection. Additionally, the bacterial population of Xoo in CX315 was significantly lower than in JG30, as shown by reduced expression of the bacterial effector gene XopA (Figure 1C). These findings collectively support the hypothesis that CX315 harbors a genetically controlled resistance mechanism that actively suppresses pathogen proliferation through enhanced immune signaling.
3.2 Fine mapping of Xa50(t) identifies a candidate region on chromosome 11
To fine-map the R gene in CX315 and elucidate the genetic basis of its resistance, an F2 mapping population was generated by selfing heterozygous F1 plants derived from a cross between CX315 and IR24. This population was phenotyped for resistance to the Xoo strain T7174. To locate the resistance locus, 256 insertion-deletion (InDel) markers distributed across all 12 rice chromosomes were screened for polymorphism between CX315, IR24, and their F2 progeny. Totally, 112 markers were polymorphic and used for genotyping. Linkage analysis based on recombination frequencies revealed that markers ID11–7 and M11-540, located on chromosome 11, were tightly associated with the resistance phenotype, with recombination events observed in 30 and 25 individuals, respectively (Figure 2). Subsequent fine mapping narrowed the candidate interval to a 147.7 kb region between markers M11–588 and M11-602, with M11–489 showing complete co-segregation with the resistance trait. This high-resolution mapping region harbors 13 predicted open reading frames (ORFs) according to the reference genome (Shuhui498). Gene annotation within the 147.7 kb mapping interval on chromosome 11 identified a high-confidence candidate region for the novel resistance gene Xa50(t) (Supplementary Table 3). While ORF9 is predicted to be Xa4 gene locus we named ORF9 to Xa4 subsequently. The presence of WAK, receptor-like kinase, and leucine-rich repeat encoding ORFs within the candidate region warrants further investigation to precisely identify the Xa50(t) gene.
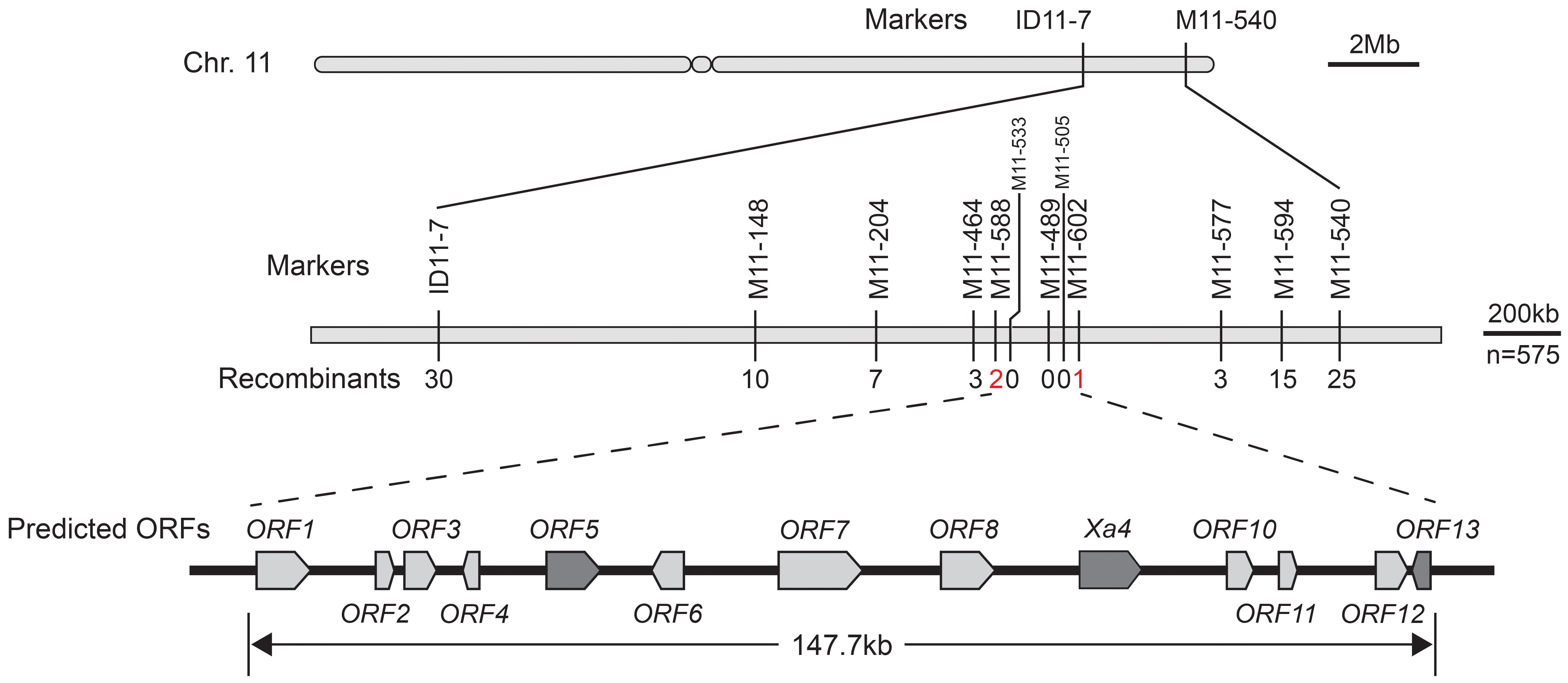
Figure 2. Fine mapping of Xa50(t) on Chromosome 11. Genetic mapping of the resistance gene Xa50(t) using recombinants from the F2 population. The region was narrowed down to a 147.7 kb interval flanked by markers M11–588 and M11–602 on Chromosome 11. The recombination frequency is displayed for key markers, and the final co-segregating marker (M11-489) is indicated in red. Within the interval, 14 predicted open reading frames were identified.
3.3 Transcriptome analysis of CX315 and JG30 in response to Xoo
Transcriptome profiling revealed significant differential gene expression (DEG) patterns in CX315 and JG30 at both 0 and 48 hpi with Xoo strain T7174, highlighting key molecular mechanisms of resistance. Functional enrichment analyses emphasized the activation of defense-related pathways in CX315, along with suppression of susceptibility-associated processes. A total of 6,221 differentially expressed genes (DEGs) were identified between CX315 and JG30 at 48 hpi, with 2,275 upregulated and 3,946 downregulated in CX315 (Figure 3C). The Venn diagram (Figure 3A) shows the distribution and overlap of DEGs across genotypes and timepoints, revealing a distinct set of CX315-specific DEGs at 48 hpi. The volcano plot (Figure 3B) illustrates the magnitude of differential expression, highlighting genes with strong pathogen-responsive activation or repression.
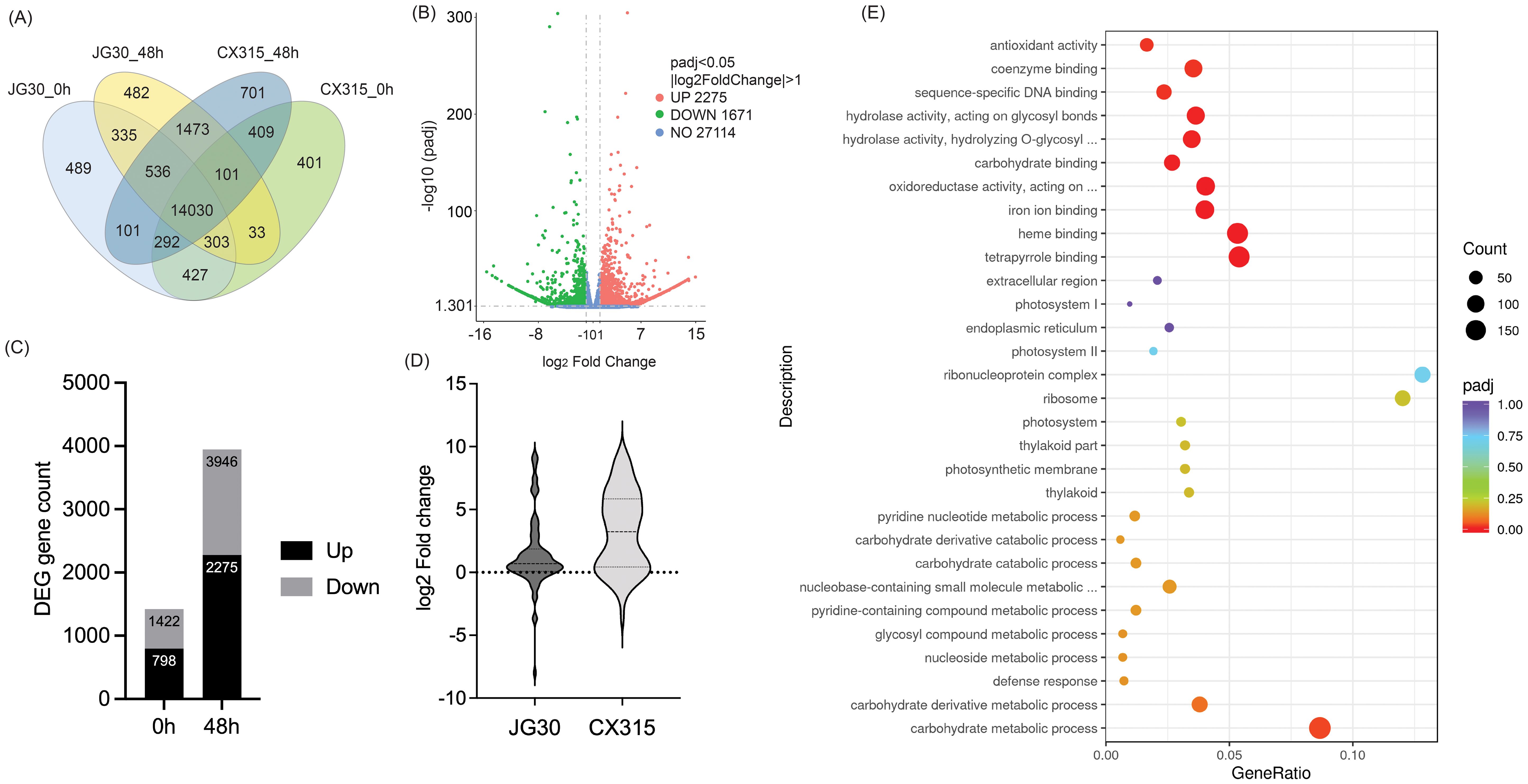
Figure 3. Differential gene expression and enrichment analysis in CX315 and JG30 during Xoo infection. (A) Venn diagram representing overlapping and unique differentially expressed genes (DEGs) across CX315 and JG30 at 0 hours (0h) and 48 hours post-inoculation (48h). CX315 shows a unique set of DEGs at 48h, highlighting its distinct transcriptional response to Xoo infection. (B) Bar graph showing the distribution of upregulated (black) and downregulated (gray) DEGs in CX315 compared to JG30 at baseline (0h) and post-inoculation (48h). The number of DEGs increases significantly at 48h, reflecting an enhanced transcriptional reprogramming in CX315 during infection. (C) Volcano plot of DEGs between CX315 and JG30 at 48h. Genes with significant upregulation (log2FoldChange > 1, padj < 0.05) are shown in pink, while significantly downregulated genes are shown in green. Non-significant genes are represented in blue. This plot highlights key genes strongly involved in defense and metabolic processes. (D) Violin plot illustrating the distribution of log2 fold change in WAK gene expression at 48h between CX315 and JG30. (E) Gene Ontology (GO) enrichment analysis of DEGs in CX315 at 48h. Significantly enriched GO terms are shown for biological processes, molecular functions, and cellular components. The size of the dots indicates the number of DEGs associated with each term, while the color represents statistical significance (adjusted p-value).
At baseline (0 hpi), 2,220 DEGs were identified between CX315 and JG30, including 798 upregulated and 1,422 downregulated genes in CX315. GO enrichment of these genes indicated that upregulated transcripts were predominantly involved in oxidative stress response and carbohydrate derivative metabolic processes, suggesting that CX315 maintains a primed defense state even prior to infection. Pathway analysis further revealed enrichment in phenylpropanoid biosynthesis and MAPK signaling—two key components of plant immune responses.
At 48 hpi, GO enrichment of CX315 DEGs demonstrated activation of defense-related processes, including “response to biotic stimulus,” “oxidoreductase activity,” and “cell wall organization,” while downregulated genes were enriched in photosynthesis-related pathways (Figure 3E). This shift reflects a strategic reallocation of metabolic resources from growth to defense under pathogen attack. In line with this, members of the wall-associated kinase (WAK) family showed higher expression in CX315 compared to JG30 at 48 hpi (Figure 3D), consistent with their involvement in Xa50(t)-mediated immune signaling.
Temporal comparison within CX315 (0 h vs. 48 hpi) identified 5,321 DEGs, of which 2,491 were upregulated and 2,830 were downregulated post-inoculation. Enriched biological processes among upregulated genes included oxidative stress and chitin metabolic responses, while downregulated transcripts were associated with photosynthetic pathways such as photosystem I and thylakoid membrane components.
Overall, these findings demonstrate that CX315 mounts a rapid and coordinated transcriptional defense in response to Xoo, involving the activation of key immune pathways and suppression of susceptibility-associated and energy-intensive metabolic processes. This dynamic transcriptional reprogramming provides insight into the molecular mechanisms by which Xa50(t) confers enhanced resistance to bacterial blight.
3.4 Differential expression of candidate genes in response to Xoo inoculation
It was indicated that pathogen-triggered induction of Xa4 expression contributes to its disease resistance (Hu et al., 2017). To prioritize candidate genes within the 147.7 kb interval linked to Xa50(t), qRT-PCR was conducted on all 13 predicted ORFs in CX315 and the susceptible cultivar JG30 at 0- and 48-hours post-inoculation with Xoo strain T7174. Three genes ORF5, Xa4, and ORF13 were significantly upregulated in CX315 at 48 hpi, while expression of ORF5 and Xa4 remained unchanged in JG30 (Figure 4). Interestingly, ORF13 displayed moderate induction in JG30, suggesting it may be partially responsive to pathogen inoculation even in the absence of functional resistance (Figure 4). This pathogen-responsive expression in the resistant background suggests their potential roles in Xa50(t)-mediated defense. The expression of ORF4 could not be reliably determined, due to technical limitations in primer design.
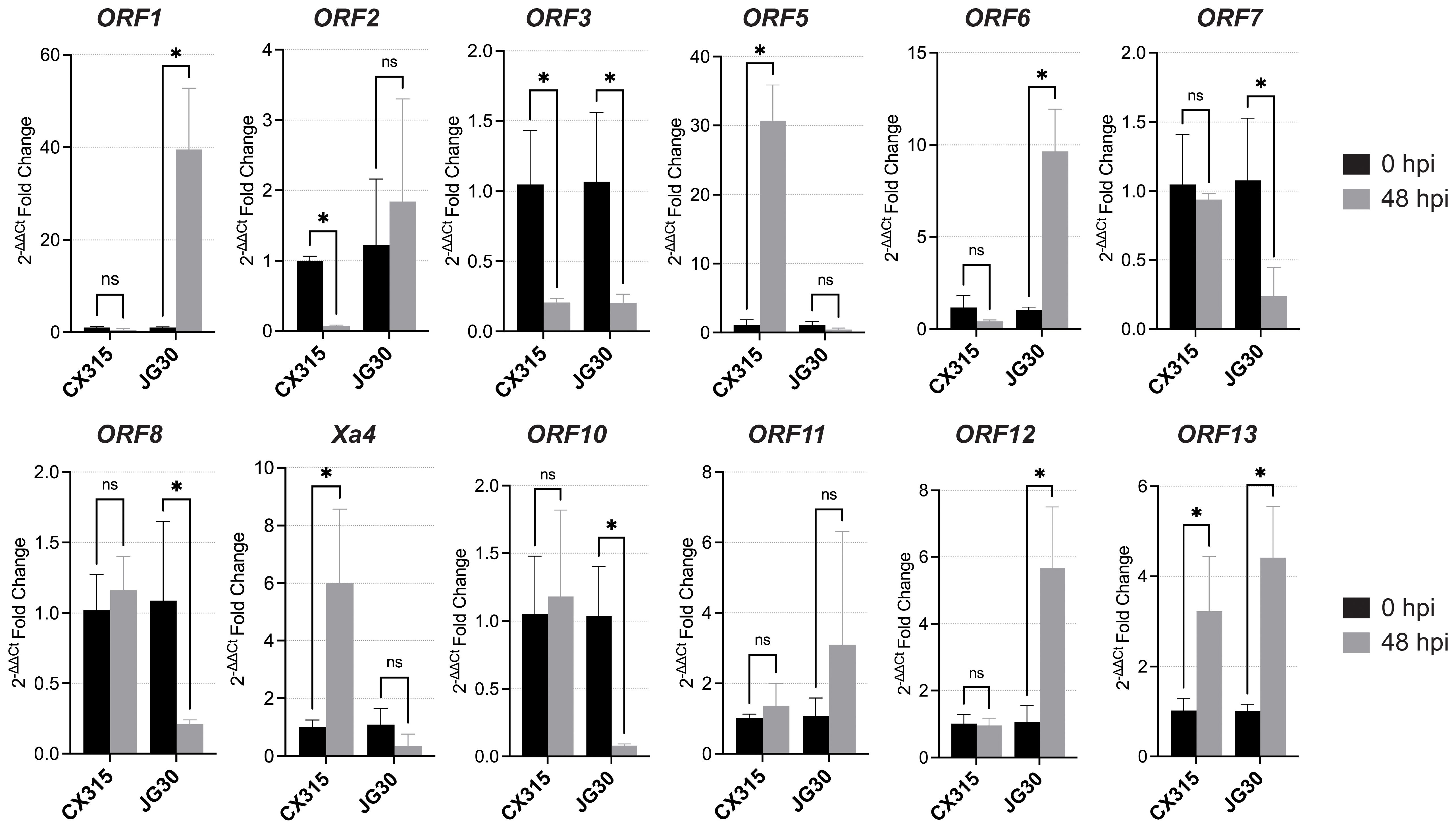
Figure 4. Expression analysis of the 12 candidate ORFs in CX315 (resistant) and JG30 (susceptible) plants at 0 and 48 hours post inoculation (hpi) with Xoo strain T7174. Significant upregulation was observed in several ORFs (e.g., ORF5, Xa4 and ORF13) in CX315, while JG30 exhibited no significant induction. Statistical significance was determined using a Student’s t-test (*p<0.05; ns, not significant).
ORF5 encode wall-associated kinase (WAK)-like proteins, which act as sensors of cell wall perturbation or pathogen-derived signals, initiating immune responses. ORF13 encodes a receptor-like kinase (RLK) with WAK-related domains, suggesting a role in downstream signal transduction or receptor complex formation. The co-induction of these kinase-domain genes points to a multilayered signaling cascade contributing to CX315 resistance. Their selective activation in response to Xoo highlights them as promising candidates underlying Xa50(t) and warrants further functional analysis.
3.5 Xa50(t) provide distinct resistance than Xa4
Since the fine-mapped interval of Xa50(t) overlapped with the known Xa4 locus, comparative pathotyping was performed to determine whether Xa50(t) is allelic to Xa4 or confers resistance through an independent mechanism. For this purpose, the Xa4-containing line IRBB4 and the Xa50(t)-harboring line CX315 were inoculated with the Xoo strain T7174 at the seedling stage and further challenged with a panel of diverse Xoo strains at the tillering stage. At the seedling stage CX315 exhibited a hypersensitive-like response and significantly shorter lesions compared to IRBB4 (Figure 5A). At the tillering stage, the resistance responses significantly varied between CX315 and IRBB4 for several strains, indicating that the underlying mechanisms differ between Xa50(t) and Xa4 (Figures 5B, C).
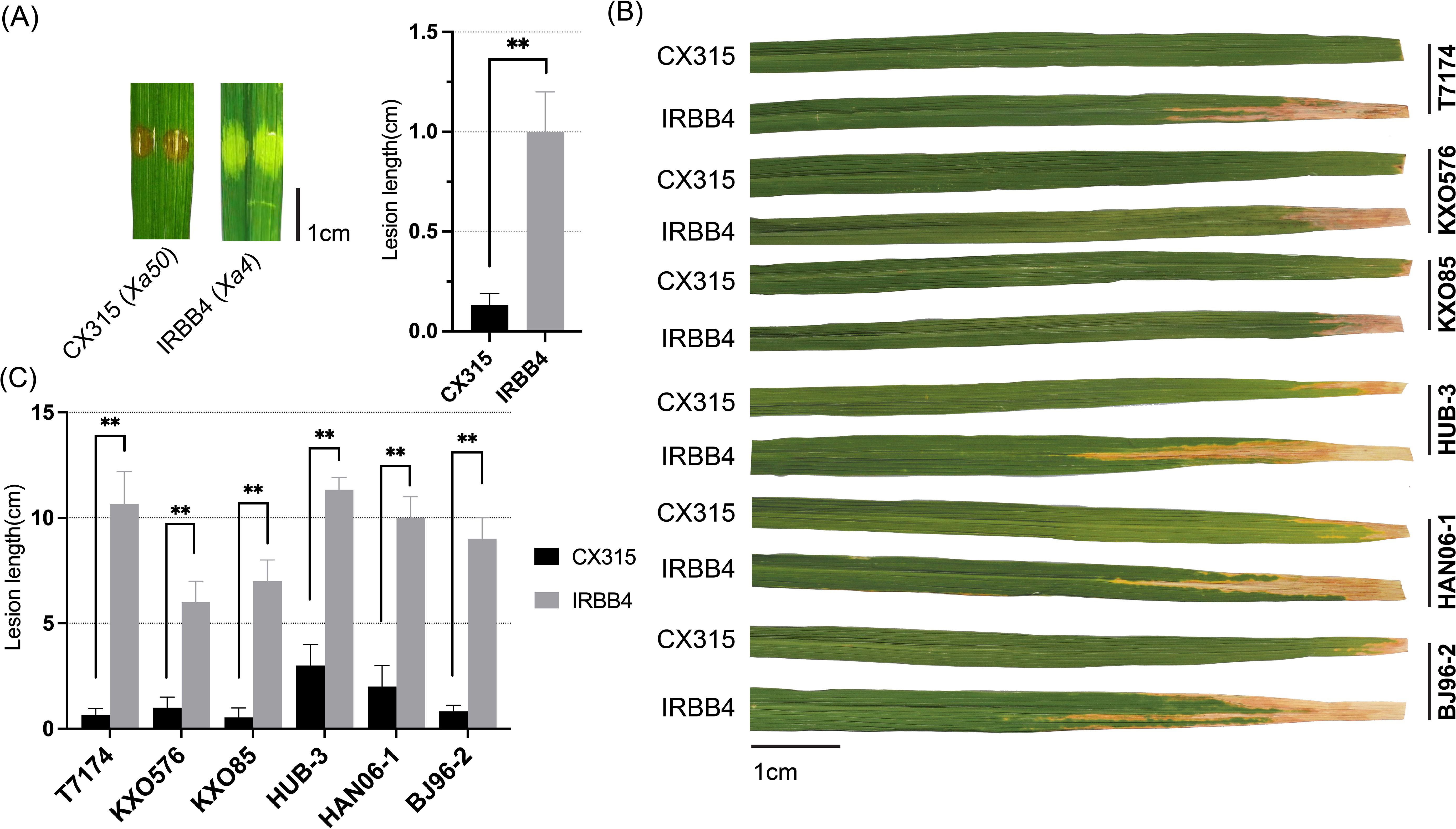
Figure 5. Xa50(t) confers resistance independently of Xa4 in rice. (A) Comparative resistance response of CX315 (Xa50(t)) and IRBB4 (Xa4) following seedling-stage inoculation with Xoo strain T7174. CX315 displayed significantly shorter lesion lengths than IRBB4. Representative leaf images and lesion quantification are shown. Data represent mean ± SD (n = 3); p < 0.01 (Student’s t-test). (B) Resistance profiles of CX315 and IRBB4 against six diverse Xoo strains at the tillering stage. Significant differences in lesion lengths between the two lines were observed for several strains, indicating differential resistance spectra. (C) Phenotype of leaves shown at right, with lesion length bar = 1cm. Data represent mean ± SD (n = 3). The asterisks (**) indicate a statistically significant difference (p < 0.01, Student’s t-test).
To confirm the presence of Xa4 in the CX315 background, we amplified and sequenced its coding region, which verified that Xa4 is intact in CX315 and suitable for targeted editing. To further validate whether Xa50(t) acts independently of Xa4, we employed CRISPR/Cas9-mediated knockout of Xa4 in the CX315 background. Two guide RNAs targeting the first exon of Xa4 were designed (Figure 6A), and the sgRNA expression cassettes were cloned into a CRISPR/Cas9 binary vector. Two homozygous T1 transgenic lines were obtained by Sanger sequencing confirmed indel mutations at the targeted sites, resulting in frameshift knockouts of Xa4 (Figure 6B, Supplementary Figure S1). Protein alignment confirmed that both Xa4-KO lines produce truncated proteins due to premature stop codons (Supplementary Figure S2). T1 homozygous knockout plants were inoculated with Xoo strain T7174 at the maturity stage. These Xa4-KO lines retained resistance phenotypes similar to the wild-type CX315 (Figure 6C), demonstrating that Xa50(t)-mediated resistance remains functional in the absence of Xa4.
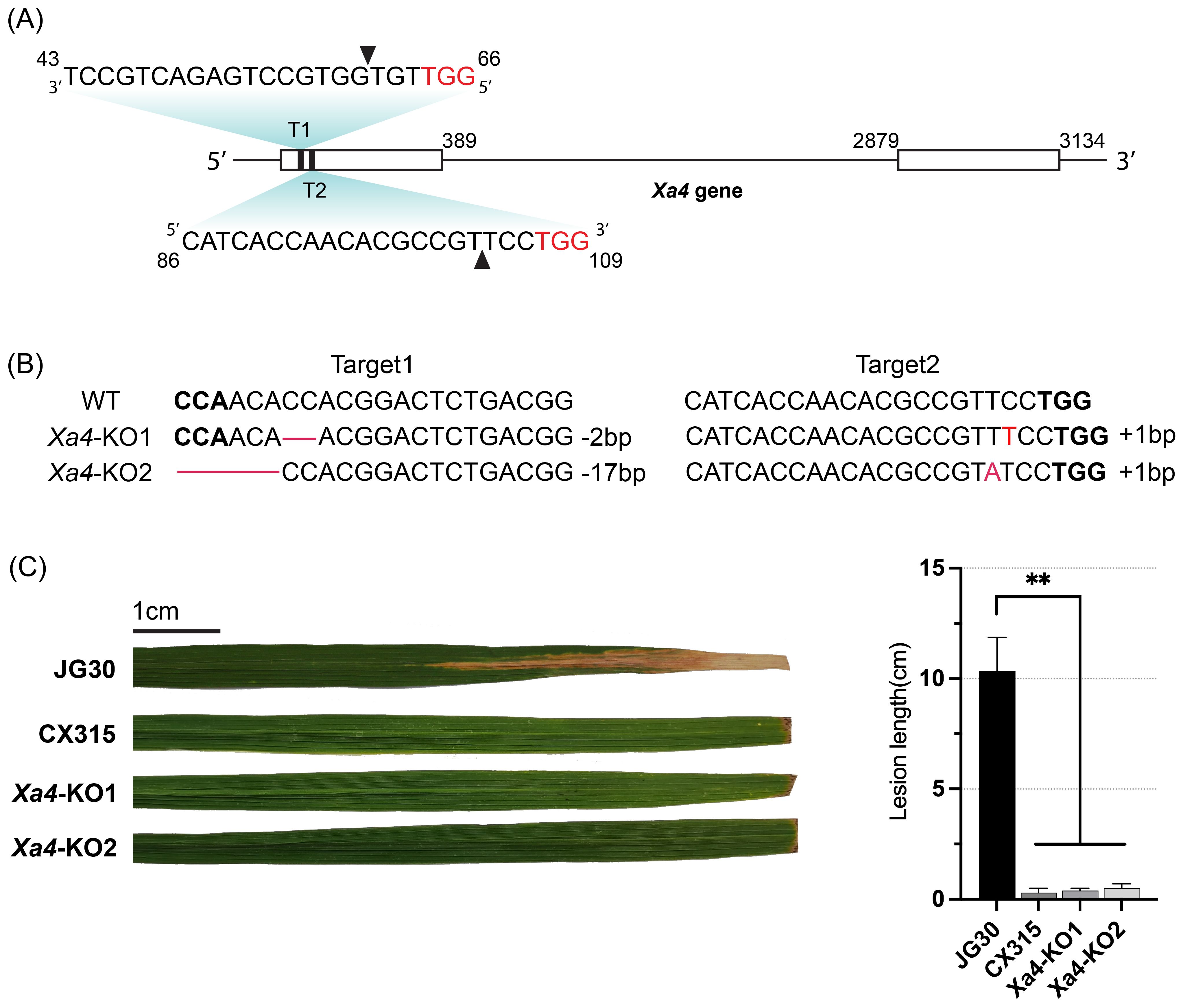
Figure 6. Knockout of Xa4 do not compromise resistance of CX315. (A) Schematic representation of the Xa4 gene and CRISPR/Cas9 target sites. Two target sequences (T1 and T2) located in the first exon were used for editing. Protospacer adjacent motif (PAM) sites are highlighted in red; cleavage sites are indicated by arrowheads. (B) Sanger sequencing of Xa4-KO1 and Xa4-KO2 lines confirmed small insertions/deletions (indels) at target loci. WT: wild-type JG30 sequence; red dashes denote deleted bases; PAM sequences in bold. (C) Lesion length analysis of JG30, CX315, and Xa4-KO lines at 14 days post-inoculation with Xoo strain T7174. The Xa4-KO lines maintained strong resistance, similar to CX315, whereas JG30 remained susceptible. Data are shown as mean ± SD (n = 3); p < 0.01 (Student’s t-test). Representative infected leaves are shown; scale bar = 1 cm.
Together, the phenotypic divergence between IRBB4 and CX315, along with the retained resistance observed in Xa4-knockout lines, confirms that Xa50(t) operates independently and represents a functionally distinct resistance gene. These findings underscore the potential of Xa50(t) as a valuable genetic resource for breeding broad-spectrum and durable resistance in rice. However, the current mapping interval contains multiple candidate resistance genes, which prevents definitive identification of Xa50(t) at this stage. Further studies aimed at dissecting the regulatory networks and functional mechanisms associated with Xa50(t) will be essential to elucidate how this novel R gene contributes to immune specificity and complements existing resistance pathways in rice.
4 Discussion
The identification and mapping of Xa50(t) in this study provide a strong foundation for exploring BB resistant resource material in rice and represent a critical step towards breeding BB-resistant varieties. Map-based cloning facilitated the fine-mapping of Xa50(t) to a 147.7 kb region on chromosome 11, overlapping with the known resistance gene Xa4. Interestingly, this overlap revealed that the resistance observed in the germplasm CX315 was mediated by Xa50(t), while Xa4 was also present but functionally distinct. While, Xa4 gene belong to WAK-like family, which plays a central role in plant immunity by sensing cell wall perturbations and activating defense responses. Despite Xa4 and Xa50(t) proximity and functional similarities, the results of this study highlight the unique and complementary roles of Xa50(t) and Xa4 in combating Xoo.
The resistance responses of Xa50(t)-containing CX315 and Xa4-containing IRBB4 to a panel of several Xoo strains underscore the distinct and non-redundant nature of these genes. For example, both genes conferred resistance to strains like T7174, KXO576, and KXO85, but with significant differences in lesion lengths, with Xa50(t)-mediated resistance showing much shorter lesions (p < 0.01 and p < 0.05). This suggests that Xa50(t) provides a more robust resistance to certain strains compared to Xa4.
Transcriptomic analysis provided additional insights into the distinct defense mechanisms of Xa50(t). Upon Xoo inoculation, significant differences were observed in the gene expression profiles of CX315 (Xa50(t)) and IRBB4 (Xa4). Genes involved in phenylpropanoid, and flavonoid biosynthesis pathways were strongly upregulated in CX315, suggesting that Xa50(t) activates specific secondary metabolic pathways to reinforce cell walls and produce antimicrobial compounds. In contrast, Xa4-mediated resistance relies more on cell wall pectin sensing and brassinosteroid-regulated cell elongation. The differential expression of transcription factors, kinase-responsive genes, and hormone-responsive genes in CX315 highlights the unique regulatory network activated by Xa50(t). Notably, the upregulation of genes associated with ethylene and jasmonic acid signaling pathways indicates that Xa50(t) employs a broader hormonal crosstalk compared to Xa4, which predominantly regulates brassinosteroid signaling.
The structural and functional diversity within the WAK family, as exemplified by Xa4, underscores their central role in plant defense. WAKs serve as molecular bridges between the extracellular cell wall and intracellular signaling pathways, monitoring cell wall integrity and triggering immune responses upon pathogen attack. This divergence likely reflects an evolutionary strategy to expand the recognition repertoire of rice against a rapidly evolving pathogen like Xoo. Comparative studies of WAKs in rice and other crops could provide deeper insights into their evolutionary origins and functional diversification.
From a practical perspective, the distinct resistance profiles of Xa50(t) and Xa4 have significant implications for rice breeding. The ability of Xa50(t) to provide robust resistance to a wide range of Xoo strains makes it a valuable genetic resource for developing BB-resistant rice varieties. Moreover, the complementary resistance patterns of Xa50(t) and Xa4 highlight the potential for gene pyramiding to enhance the durability and breadth of resistance. Marker-assisted selection using molecular markers linked to Xa50(t) can accelerate its deployment in breeding programs. Furthermore, combining Xa50(t) with other resistance genes, including those targeting different pathogen effectors or stages of infection, could provide synergistic effects, reducing the risk of resistance breakdown due to pathogen adaptation.
Recent studies have highlighted the importance of structural variation (SV) including insertions, deletions, transpositions, and presence/absence variations in shaping disease resistance loci in rice. The Xa4 locus, in particular, resides within a WAK-rich genomic region known for high SV frequency and haplotype diversity across cultivated and wild rice accessions (Hu et al., 2017; Zhao et al., 2018). Analysis from the 3K Rice Genomes Project revealed extensive structural remodeling at resistance gene clusters, including transposon insertions, promoter rearrangements, and gene copy number variation, all of which can influence gene expression and specificity. The distinct resistance profile of CX315, despite overlapping the Xa4 locus, suggests that it harbors a unique haplotype shaped by such SV events. This structural differentiation likely underlies the independent function of Xa50(t), allowing it to confer resistance distinct from Xa4. Our findings reinforce the concept that SV is a key driver of novel R gene emergence and diversification, and that SV-rich regions such as the Xa4 locus are hotspots for mining durable resistance alleles in rice breeding programs. Due to the complexity of this region and the lack of long-read genome data for CX315, we were unable to reconstruct the complete sequence. Therefore, candidate gene identification relied on transcriptome profiling and qRT-PCR expression analysis. Future work involving high-resolution sequencing of this locus will be necessary to resolve the precise gene structure and distinguish Xa50(t) from closely linked homologs such as Xa4.
In conclusion, this study not only advances our understanding of the genetic and molecular basis of BB resistance in rice but also provides practical tools for crop improvement. The distinct roles of Xa50(t) and Xa4 in resistance, their complementary profiles, and their structural differences within the importance of exploring diverse genetic resources for sustainable disease management. The functional validation of ORF5 and ORF13 is warranted, and future studies involving gene knockout and overexpression will be essential to resolve their specific roles in Xa50(t)-mediated resistance.
Future research should focus on elucidating the molecular interactions between Xa50(t) and Xoo effectors, as well as exploring its potential role in regulating abiotic stress tolerance, given the multifunctional nature of WAKs. Such efforts will contribute to the broader goal of developing resilient rice varieties capable of withstanding both biotic and abiotic challenges.
Data availability statement
The RNA-seq data have been deposited in the NCBI Sequence Read Archive (SRA) under Project ID PRJNA1209632, with accession numbers SRR31970743, SRR31970744, SRR31971027, and SRR31971028.
Author contributions
ML: Investigation, Data curation, Software, Validation, Writing – review & editing, Writing – original draft, Methodology. MS: Software, Writing – original draft, Visualization, Writing – review & editing, Formal analysis. HZ: Writing – review & editing, Methodology, Data curation. JW: Methodology, Writing – review & editing, Data curation. YL: Conceptualization, Project administration, Supervision, Validation, Writing – review & editing. CW: Project administration, Data curation, Resources, Conceptualization, Validation, Writing – review & editing, Supervision. ZW: Validation, Methodology, Supervision, Writing – review & editing, Project administration. ZJ: Funding acquisition, Writing – review & editing, Conceptualization, Investigation, Resources, Writing – original draft, Validation, Project administration, Supervision.
Funding
The author(s) declare financial support was received for the research and/or publication of this article. The research is supported by National Key R&D Program of China (2024YFD1200601), the National Natural Science Foundation of China (32401828) and the Basic Research Center of Innovation Program of Chinese Academy of Agricultural Sciences (CAAS-BRC-CS-2025-02).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Any alternative text (alt text) provided alongside figures in this article has been generated by Frontiers with the support of artificial intelligence and reasonable efforts have been made to ensure accuracy, including review by the authors wherever possible. If you identify any issues, please contact us.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2025.1657476/full#supplementary-material
References
Delteil, A., Gobbato, E., Cayrol, B., Estevan, J., Michel-Romiti, C., Dievart, A., et al. (2016). Several wall-associated kinases participate positively and negatively in basal defense against rice blast fungus. BMC Plant Biol. 16, 17. doi: 10.1186/s12870-016-0711-x
Dossa, G. S., Quibod, I., Atienza-Grande, G., Oliva, R., Maiss, E., Vera Cruz, C., et al. (2020). Rice pyramided line IRBB67 (Xa4/Xa7) homeostasis under combined stress of high temperature and bacterial blight. Sci. Rep. 10, 683. doi: 10.1038/s41598-020-57499-5
Du, Y., Liang, D., Qi, Z., Yu, J., Zhang, R., Song, T., et al. (2024). Transcriptome and differential expression analysis revealed the pathogenic-related genes in Magnaporthe oryzae during leaf and panicle infection. Phytopathol. Res. 6, 18. doi: 10.1186/s42483-024-00248-7
Fang, Y., Ding, D., Gu, Y., Jia, Q., Zheng, Q., Qian, Q., et al. (2023). Identification of QTLs conferring resistance to bacterial diseases in rice. Plants (Basel) 12, 2853. doi: 10.3390/plants12152853
Harkenrider, M., Sharma, R., De Vleesschauwer, D., Tsao, L., Zhang, X., Chern, M., et al. (2016). Overexpression of rice wall-associated kinase 25 (OsWAK25) alters resistance to bacterial and fungal pathogens. PloS One 11, e0147310. doi: 10.1371/journal.pone.0147310
Hu, K., Cao, J., Zhang, J., Xia, F., Ke, Y., Zhang, H., et al. (2017). Improvement of multiple agronomic traits by a disease resistance gene via cell wall reinforcement. Nat. Plants 3, 17009. doi: 10.1038/nplants.2017.9
Ji, Z., Ji, C., Liu, B., Zou, L., Chen, G., and Yang, B. (2016). Interfering TAL effectors of Xanthomonas oryzae neutralize R-gene-mediated plant disease resistance. Nat. Commun. 7, 13435. doi: 10.1038/ncomms13435
Ji, Z., Wang, C., and Zhao, K. (2018). Rice routes of countering xanthomonas oryzae. Int. J. Mol. Sci. 19, 3008. doi: 10.3390/ijms19103008
Jiang, N., Yan, J., Liang, Y., Shi, Y., He, Z., Wu, Y., et al. (2020). Resistance genes and their interactions with bacterial blight/leaf streak pathogens (Xanthomonas oryzae) in rice (Oryza sativa L.) - an updated review. Rice(NY) 13, 3. doi: 10.1186/s12284-019-0358-y
Kumar, A., Kumar, R., Sengupta, D., Das, S. N., Pandey, M. K., Bohra, A., et al. (2020). Deployment of Genetic and Genomic Tools Toward Gaining a Better Understanding of Rice-Xanthomonas oryzae pv. oryzae Interactions for Development of Durable Bacterial Blight Resistant Rice. Front. Plant Sci. 11. doi: 10.3389/fpls.2020.01152
Langner, T., Kamoun, S., and Belhaj, K. (2018). CRISPR Crops: Plant genome editing toward disease resistance. Annu. Rev. Phytopathol. 56, 479–512. doi: 10.1146/annurev-phyto-080417-050158
Livak, K. J. and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT Method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Lu, J., Wang, C., Zeng, D., Li, J., Shi, X., Shi, Y., et al. (2021). Genome-wide association study dissects resistance loci against bacterial blight in a diverse rice panel from the 3000 rice genomes project. Rice (N Y) 14, 22. doi: 10.1186/s12284-021-00462-3
Luo, Y., Sangha, J. S., Wang, S., Li, Z., Yang, J., and Yin, Z. (2012). Marker-assisted breeding of Xa4, Xa21 and Xa27 in the restorer lines of hybrid rice for broad-spectrum and enhanced disease resistance to bacterial blight. Mol. Breed. 30, 1601–1610. doi: 10.1007/s11032-012-9742-7
Ma, X., Zhang, Q., Zhu, Q., Liu, W., Chen, Y., Qiu, R., et al. (2015). A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284. doi: 10.1016/j.molp.2015.04.007
Rio, D. C., Ares, M., Jr., Hannon, G. J., and Nilsen, T. W. (2010). Purification of RNA using TRIzol (TRI reagent). Cold Spring Harb. Protoc. 2010, pdb.prot5439. doi: 10.1101/pdb.prot5439
Stephens, C., Hammond-Kosack, K. E., and Kanyuka, K. (2022). WAKsing plant immunity, waning diseases. J. Exp. Bot. 73, 22–37. doi: 10.1093/jxb/erab422
Timilsina, S., Potnis, N., Newberry, E. A., Liyanapathiranage, P., Iruegas-Bocardo, F., White, F. F., et al. (2020). Xanthomonas diversity, virulence and plant-pathogen interactions. Nat. Rev. Microbiol. 18, 415–427. doi: 10.1038/s41579-020-0361-8
Tirnaz, S., Zandberg, J., Thomas, W. J. W., Marsh, J., Edwards, D., and Batley, J. (2022). Application of crop wild relatives in modern breeding: An overview of resources, experimental and computational methodologies. Front. Plant Sci. 13. doi: 10.3389/fpls.2022.1008904
Wang, R., Ji, Z., Kou, Y., Yang, X., Huang, W., Sun, Z., et al. (2025). Occurrence and integrated control of major rice diseases in China. New Plant Prot. 2, e70004. doi: 10.1002/npp2.70004
Wang, Y., Li, X., Ishikawa, R., and Luo, X. (2023). Editorial: Mining and utilization of favorable gene resources in rice. Front. Plant Sci. 14. doi: 10.3389/fpls.2023.1289069
Wang, S., Liu, W., Lu, D., Lu, Z., Wang, X., Xue, J., et al. (2020). Distribution of bacterial blight resistance genes in the main cultivars and application of Xa23 in Rice Breeding. Front. Plant Sci. 11. doi: 10.3389/fpls.2020.555228
Xu, Z., Xu, X., Wang, Y., Liu, L., Li, Y., Yang, Y., et al. (2022). A varied AvrXa23-like TALE enables the bacterial blight pathogen to avoid being trapped by Xa23 resistance gene in rice. J. Adv. Res. 42, 263–272. doi: 10.1016/j.jare.2022.01.007
Yugander, A., Sundaram, R. M., Singh, K., Ladhalakshmi, D., Subba Rao, L. V., Madhav, M. S., et al. (2018). Incorporation of the novel bacterial blight resistance gene Xa38 into the genetic background of elite rice variety Improved Samba Mahsuri. PloS One 13, e0198260. doi: 10.1371/journal.pone.0198260
Keywords: bacterial blight resistance, Xa50(t), Xanthomonas oryzae pv. oryzae (Xoo), wall-associated kinase (WAK), Xa4
Citation: Li M, Shafique MS, Zhou H, Wang J, Liu Y, Wang C, Wang Z and Ji Z (2025) A new bacterial blight resistance gene Xa50(t) in the Xa4 locus confers resistance against Xanthomonas oryzae pv. oryzae in rice. Front. Plant Sci. 16:1657476. doi: 10.3389/fpls.2025.1657476
Received: 01 July 2025; Accepted: 07 August 2025;
Published: 29 August 2025.
Edited by:
Jianping Lu, Zhejiang University, ChinaReviewed by:
Wanwan Liang, Chinese Academy of Agricultural Sciences, ChinaBojun Ma, Zhejiang Normal University, China
Copyright © 2025 Li, Shafique, Zhou, Wang, Liu, Wang, Wang and Ji. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhiyuan Ji, aml6aGl5dWFuQGNhYXMuY24=; Zhonghua Wang, emh3YW5nbmV3QDEyNi5jb20=; Chunlian Wang, d2FuZ2NodW5saWFuQGNhYXMuY24=
†These authors have contributed equally to this work
 Man Li
Man Li Muhammad Sohaib Shafique
Muhammad Sohaib Shafique Houyu Zhou
Houyu Zhou Jialu Wang2
Jialu Wang2 Chunlian Wang
Chunlian Wang Zhiyuan Ji
Zhiyuan Ji