- Stress Signal Transduction Lab, Department of Plant Molecular Biology, University of Delhi South Campus, New Delhi, India
Introduction
A key node in number of essential cellular processes in eukaryotes, Armadillo was originally characterized in Drosophila as the component of Wingless/Wnt signal transduction pathway (Nusslein-Volhard and Wieschaus, 1980). β-catenin is the mammalian homolog of Armadillo playing dual role in structural and transcriptional regulation during embryonic development (Conacci-Sorrell et al., 2002). Even though initially characterized in animals, members of the Armadillo proteins are also known to exist in non-animals including slime mold (Dictyostelium discoideum) and plants (Wang et al., 1998; Barelle et al., 2006; Veses et al., 2009). The existence of Armadillo repeat family of proteins across species suggests ancient evolutionary origin and functional conservation of these proteins in multicellular organisms (Coates, 2003). The intricate role of β-catenin raises several doubts about the mechanism by which it mediates interaction with diverse partner proteins using common interface, and how this interaction influences adhesion and transcription?
The ARM family proteins have been identified with multiple functional domains in more than one species. Genome-wide studies in plants have shown the existence of large number of Armadillo homologs in Physcomitrella patens, Arabidopsis and Oryza sativa (Mudgil et al., 2004; Sharma et al., 2014). One assumption is that, Armadillo family being evolutionary conserved, perform similar role in all organisms. However, the existence of multigene Armadillo family with various subfamilies indicate novel species specific functions of these proteins in plants. Several recent studies have made known the function of numerous ARM proteins in Arabidopsis and rice. Apart from their analogous role in regulation of gene expression and developmental processes, various proteins were discovered to be predominantly involved in plant stress responses.
Thus, an intriguing and important question remains as in what way the similar effector proteins of Wnt pathway function and how similar canonical response is prevented or exist in plants. Recent progress in studies of ARM proteins in plants has suggested some possible answers to this question. However, the Wnt signaling mechanism regulated by ARM repeat proteins is still unknown. Regarding this, many underscoring questions are just beginning to emerge that remains to be answered.
Wnt Signaling—Developmental Regulation in Plants and Animals
Wnt proteins are one of the foremost signaling molecule essential for cell polarity, embryonic development and the determination of cell fate in metazoa (Cadigan and Nusse, 1997; Wodarz and Nusse, 1998; Logan and Nusse, 2004). A combination of molecular and genetic studies has provided evidences for how Wnt1, Wnt3a, and Wnt8 specifically induce the activation of “canonical β-catenin” pathway in animals (Du et al., 1995; Shimizu et al., 1997; Kuhl et al., 2000). However, no evidence for a Wnt, Frizzled (Fz) and low-density-lipoprotein-related protein receptors has been obtained in plants. Despite this, few homologs of proteins, which act as negative regulator of Wnt signaling has been unveiled in plants. Based on BLAST searches, the serine/threonine kinase GSK-3 (glycogen synthase kinase-3), CK1 (casein kinase 1) and APC (Adenomatous polyposis coli), which together form a destruction complex to stimulate degradation of β-catenin in animals were found to be conserved in plants (Figure 1) (Li et al., 2001). This has been proven in animals that activity of GSK3/CK1 complex is inhibited in response to Wnt signal perception at the cell surface to relieve its inhibitory effects on downstream β-catenin (He et al., 2004; Tamai et al., 2004; Nusse, 2005). The conservation of β-catenin destruction complex in plants points toward novel targets and modulation of Wnt signaling.
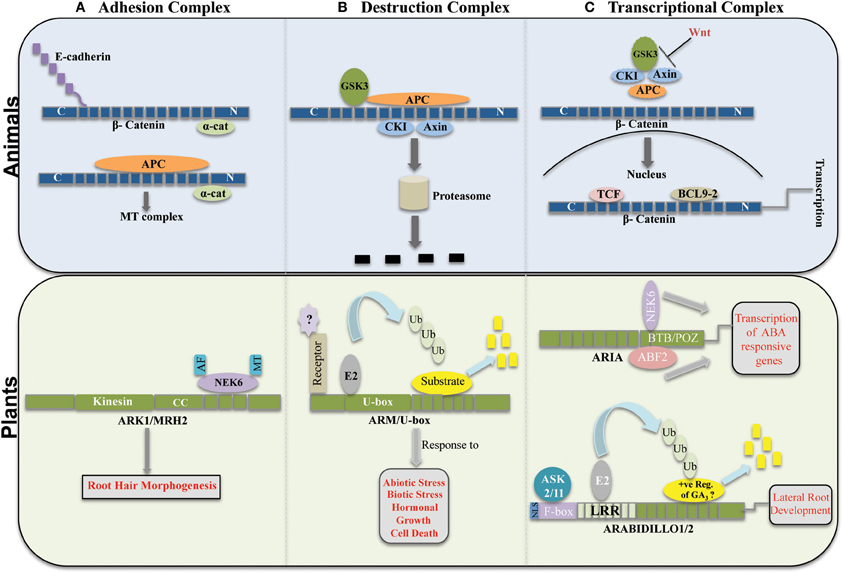
Figure 1. Functional comparison of β-cat like-ARM repeats protein in plants and animals. (A) Adhesion Complex: β-catenin in animals binds cytoplasmic tail of cadherin to link it with α-catenin. Additionally, β-catenin together with APC interacts with microtubule complexes. In plants, ARK1/MRH2 (ARM repeat kinesin1/morphogenesis of root hair 1) interacts with NEK6 (NIMA-related protein kinase 6) to mediate root epidermal cell morphogenesis. CC represent coiled coil domain. (B) Destruction Complex: β-catenin is targeted for proteasomal degradation by a GSK3, APC, CKI, and Axin complex in the cytoplasm. Similarly in plants, ARM/U-box proteins, in response to various stimuli target substrate protein for proteasomal degradation. (C) Transcriptional Complex: Wnt signals inhibits the destruction complex, free β-catenin enters the nucleus where it links with the transcriptional regulators to activate transcription of target genes. In plants, ARIA an ARM protein with BTB/POZ domain binds with ABF2 and NEK6 transcription factors to stimulate transcription of ABA responsive genes. Additionally, ARABIDILLO1/2 interacts with ASK2/11 through their F-box domain to mediate degradation of possibly a positive regulator of GA3 signaling to promote transcription of genes related to lateral root development.
Potential “Wnt-Like” Signaling Functions for Plant ARM Family Proteins
Arabidopsis comprises a multigene SHAGGY-related protein kinase (ASK) gene family, which is 70% identical to glycogen synthase kinase-3 from mammals, (Bourouis et al., 1990; Siegfried et al., 1990; Woodgett, 1990) classified into four distinct subfamilies (Jonak and Hirt, 2002). In the past few years, significant progress has been made in understanding how GSK3s perform their diverse functions in plants. The diverged biological functions of these members in signal transduction, cell patterning, cytokinesis and determination of cell fate has been established and credited to their diversity within plants (Dornelas et al., 1998). Most of the plants GSKs are found to be involved in brassinosteroid signaling and salt stress response (Dornelas et al., 2000; Kim et al., 2009). Brassinosteroids (BRs) are plant hormones, which signal through a plasma membrane localized receptor kinase BRI1. BRI1 interacts with BAK1 (BRI1 associated receptor kinase 1) to mediate plant steroid signaling (Nam and Li, 2002). BES1 has been identified as a suppressor of BRI1, which in turn is negatively regulated by a kinase BIN2 (Yin et al., 2002). Interestingly, the BR signaling pathway mechanism is analogous to the Wnt signaling pathway. In the proposed model, BIN2 which shares sequence homology with GSK-3 (Li and Nam, 2002), phosphorylate and destabilize its substrate BES-1. In response to brassinosteroids, BES-1 is stabilized and accumulates in the nucleus to activate target gene expression (Yin et al., 2002).
It is important to note that both BES-1 and β-catenin does not share homology at the protein sequence level. Similarly, BRI1 and Wnt are the two different receptors and does not belong to the same family (He et al., 2002; Yin et al., 2002; Zhao et al., 2002). However, it will be interesting to know if any of the protein in multigene Armadillo family in plants, gets regulated in the same manner or it is simply the way in which the pathway is conserved.
Meanwhile, several lines of evidence suggest the role of Wnt signaling proteins i.e., Armadillo repeats containing proteins in the developmental regulation in both animals and plants (Amador et al., 2001). p120ctn is an Armadillo repeat protein identified as a component of E-cadherin-catenin cell adhesion complex (Daniel et al., 2002). The signaling and cell adhesion co-factor p120ctn is the only known binding partner for Kaiso, a novel BTB/POZ domain zinc finger transcription factor (Daniel et al., 2002). Another possible candidate mediating interaction within actin and microtubule filaments in plants is ARK/MRH2 kinesin (ARM repeat kinesin/Morphogenesis of root hair). ARK/MRH2 interacts with NIMA-related protein kinase NEK6, to regulate epidermal cell morphogenesis by modulating microtubule dynamics (Sakai et al., 2008).
In relation to this, Arabidopsis (AT5G13060) and rice (LOC_Os05G33050) also possess homologous proteins comprising ARM repeats and a BTB/POZ domain (Figure 1). The Arabidopsis BTB/POZ ARM protein also known as ABAP1 has been shown to be involved in DNA replication and gene transcription controls (Masuda et al., 2008).
Arabidillo-1/-2 and Oryzadillo are the closest homolog of β-catenin in Arabidopsis and Oryza sativa respectively, consisting of an F-box motif near their N-terminal, and several presumed sites for GSK-3 phosphorylation (Gagne et al., 2002; Kuroda et al., 2002; Coates, 2003). Remarkably, Arabidillo's are closest to the β-catenin homolog in Dictyostelium' Aar protein that consists of an F-box domain and is required for the differentiation and expression of prespore specific genes (Grimson et al., 2000). Besides, analogous to animals, physical interaction of Arabidillo-1/-2 proteins through their F-box domain with ASKs (SHAGGY-like protein kinase) lead to the formation of SCF complexes that target various substrates for ubiquitn/26S proteasome–mediated proteolysis has been proven in plants (Changjun et al., 2010). This suggest an evolutionary conservation of signal transduction pathway elements and their site of action within animals and plants.
Beyond Wnt Signaling: Role of Plant ARM Proteins
Exposure to abiotic and biotic stress results in alteration of cellular homeostasis in plants. The first response to stress factors, is to activate the signal transduction pathways that stimulate cell defense and adaptive mechanisms. Ubiquitination is a unique protein degradation mechanism utilized by plants to effectively degrade detrimental cellular proteins and components specific to these stress signalings. A majority of U-box E3 ubiquitin ligase encoding ARM proteins related to biotic and abiotic stress have been identified in plants. We can certainly anticipate new insight into the molecular mechanism of plant β-catenin-like proteins function in the context of abiotic stress signals.
There are 41 and 47 predicted U-box/ARM proteins in the genome of Arabidopsis and rice respectively (Mudgil et al., 2004; Sharma et al., 2014). A few of them have been functionally characterized in Arabidopsis. Many of these proteins have now been linked to specific stress and hormonal responses.
A biological role for the U-box/ARM protein AtPUB9 has been proposed in ABA (Abscisic acid) signaling (Samuel et al., 2008). In Arabidopsis, ATPUB18 and ATPUB19 are the two homologous proteins. Molecular analysis of AtPUB19 showed that it is upregulated in response to drought, salt, cold and ABA (Liu et al., 2011). In the consecutive year, role of ATPUB18 as a negative regulator has been put forward in ABA-mediated stomatal closure and drought responses (Seo et al., 2012). A different homologous pair of PUB proteins, AtPUB22 and 23 have been shown to play a combinatory role in the negative regulation of drought stress (Cho et al., 2008; Seo et al., 2012). A closely related ortholog of ATPUB22/23 in Capsicum annum known as CaPUB1 was found to be highly inducible in response to various abiotic stresses such as drought, cold and salt (Cho et al., 2006).
Another report suggested the role of AtCHIP, an Arabidopsis U-box/ARM protein in response to extreme temperature conditions. Subsequently, AtCHIP was reported to be involved in the ABA stress signaling pathway by mediating interaction with protein phosphatase 2A (Yan et al., 2003). In rice, SPL11 was identified as a U-box containg ARM protein that functions as a negative regulator in the control of cell death and pathogen defense (Zeng et al., 2004). The Arabidopsis ortholog of SPL11, ATPUB13 is a functionally conserved protein regulating plant defense, cell death and flowering time (Li et al., 2012a,b). In Nicotiana, two U-box/ARM proteins NtCMPG1 and tobacco ACRE276 and their functional homolog in Arabidopsis, AtPUB17 has been implicated as positive mediators of plant defense and stress signaling (Gonzalez-Lamothe et al., 2006; Yang et al., 2006). Apart from this, expression analysis in rice has confirmed many of the ARM proteins without any associated domain to be differentially regulated under abiotic stress conditions suggesting a role of ARM repeats in the stress regulation (Sharma et al., 2014).
On the basis of facts described above, it can be concluded that animal and plant ARM repeat proteins share many resemblances. Therefore, it is possible that at least some transcription effectors involved in Wnt signaling are evolutionary conserved. These elements include nuclear accumulation in response to extracellular signal, phosphorylation and degradation. Apart from the common response, plants possess specific signaling pathways mediated by ARM proteins. In plants, ubiquitination is critically involved in the function of ARM proteins. The proliferation of β-catenin-like ARM proteins in plants suggest their significance in the regulation of diverse biological fuctions in them. Further study of these proteins in plants would contribute to our understanding of the molecular factors involved in response to abiotic stress.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We are thankful to research grants from Delhi University and Department of Biotechnology (DBT), India.
References
Amador, V., Monte, E., Garcia-Martinez, J. L., and Prat, S. (2001). Gibberellins signal nuclear import of PHOR1, a photoperiod-responsive protein with homology to Drosophila armadillo. Cell 106, 343–354. doi: 10.1016/S0092-8674(01)00445-7
Barelle, C. J., Richard, M. L., Gaillardin, C., Gow, N. A., and Brown, A. J. (2006). Candida albicans VAC8 is required for vacuolar inheritance and normal hyphal branching. Eukaryotic Cell 5, 359–367. doi: 10.1128/EC.5.2.359-367.2006
Bourouis, M., Moore, P., Ruel, L., Grau, Y., Heitzler, P., and Simpson, P. (1990). An early embryonic product of the gene shaggy encodes a serine/threonine protein kinase related to the CDC28/cdc2+ subfamily. EMBO J. 9, 2877–2884.
Cadigan, K. M., and Nusse, R. (1997). Wnt signaling: a common theme in animal development. Genes Dev. 11, 3286–3305. doi: 10.1101/gad.11.24.3286
Changjun, M., Ni, C., Xiaofeng, L., Pengfei, J., Zhaoyan, W., and Heng, L. (2010). F-box protein arabidillo-1 promotes lateral root development by depressing the functioning of GA3 in Arabidopsis. J. Plant Biol. 53, 374–380. doi: 10.1007/s12374-010-9125-8
Cho, S. K., Chung, H. S., Ryu, M. Y., Park, M. J., Lee, M. M., Bahk, Y. Y., et al. (2006). Heterologous expression and molecular and cellular characterization of CaPUB1 encoding a hot pepper U-Box E3 ubiquitin ligase homolog. Plant Physiol. 142, 1664–1682. doi: 10.1104/pp.106.087965
Cho, S. K., Ryu, M. Y., Song, C., Kwak, J. M., and Kim, W. T. (2008). Arabidopsis PUB22 and PUB23 are homologous U-Box E3 ubiquitin ligases that play combinatory roles in response to drought stress. Plant Cell 20, 1899–1914. doi: 10.1105/tpc.108
Coates, J. C. (2003). Armadillo repeat proteins: beyond the animal kingdom. Trends Cell Biol. 13, 463–471. doi: 10.1016/S0962-8924(03)00167-3
Conacci-Sorrell, M., Zhurinsky, J., and Ben-Ze'ev, A. (2002). The cadherin-catenin adhesion system in signaling and cancer. J. Clin. Invest. 109, 987–991. doi: 10.1172/JCI200215429
Daniel, J. M., Spring, C. M., Crawford, H. C., Reynolds, A. B., and Baig, A. (2002). The p120(ctn)-binding partner Kaiso is a bi-modal DNA-binding protein that recognizes both a sequence-specific consensus and methylated CpG dinucleotides. Nucleic Acid Res. 30, 2911–2919. doi: 10.1093/nar/gkf398
Dornelas, M. C., Lejeune, B., Dron, M. and Kreis, M. (1998). The Arabidopsis SHAGGY-related protein kinase (ASK) gene family: structure, organization and evolution. Gene 212, 249–257. doi: 10.1016/S0378-1119(98)00147-4
Dornelas, M. C., Van Lammeren, A. A., and Kreis, M. (2000). Arabidopsis thaliana SHAGGY-related protein kinases (AtSK11 and 12) function in perianth and gynoecium development. Plant J. 21, 419–429. doi: 10.1046/j.1365-313x.2000.00691.x
Du, S. J., Purcell, S. M., Christian, J. L., McGrew, L. L., and Moon, R. T. (1995). Identification of distinct classes and functional domains of Wnts through expression of wild type and chimeric proteins in Xenopus embryos. Mol. Cell. Biol. 15, 2625–2634.
Gagne, J. M., Downes, B. P., Shiu, S. H., Durski, A. M., and Vierstra, R. D. (2002). The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 99, 11519–11524. doi: 10.1073/pnas.162339999
Gonzalez-Lamothe, R., Tsitsigiannis, D. I., Ludwig, A. A., Panicot, M., Shirasu, K., and Jones, J. D. (2006). The U-box protein CMPG1 is required for efficient activation of defense mechanisms triggered by multiple resistance genes in tobacco and tomato. Plant Cell 18, 1067–1083. doi: 10.1105/tpc.106.040998
Grimson, M. J., Coates, J. C., Reynolds, J. P., Shipman, M., Blanton, R. L., and Harwood, A. J. (2000). Adherens junctions and beta-catenin-mediated cell signalling in a non-metazoan organism. Nature 408, 727–731. doi: 10.1038/35047099
He, J. X., Gendron, J. M., Yang, Y., Li, J., and Wang, Z. Y. (2002). The GSK3-like kinase BIN2 phosphorylates and destabilizes BZR1, a positive regulator of the brassinosteroid signaling pathway in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 99, 10185–10190. doi: 10.1073/pnas.152342599
He, X., Semenov, M., Tamai, K., and Zeng, X. (2004). LDL receptor-related proteins 5 and 6 in Wnt/beta-catenin signaling: arrows point the way. Development 131, 1663–1677. doi: 10.1242/dev.01117
Jonak, C., and Hirt, H. (2002). Glycogen synthase kinase 3/SHAGGY-like kinases in plants: an emerging family with novel functions. Trends Plant Sci. 7, 457–461. doi: 10.1016/S1360-1385(02)02331-2
Kim, T. W., Guan, S., Sun, Y., Deng, Z., Tang, W., Shang, J. X., et al. (2009). Brassinosteroid signal transduction from cell-surface receptor kinases to nuclear transcription factors. Nat. Cell Biol. 11, 1254–1260. doi: 10.1038/ncb1970
Kuhl, M., Sheldahl, L. C., Park, M., Miller, J. R., and Moon, R. T. (2000). The Wnt/Ca2+ pathway: a new vertebrate Wnt signaling pathway takes shape. Trends Genet. 16, 279–283. doi: 10.1016/S0168-9525(00)02028-X
Kuroda, H., Takahashi, N., Shimada, H., Seki, M., Shinozaki, K. and Matsui, M. (2002). Classification and expression analysis of Arabidopsis F-box-containing protein genes. Plant Cell Physiol. 43, 1073–1085. doi: 10.1093/pcp/pcf151
Li, J., and Nam, K. H. (2002). Regulation of brassinosteroid signaling by a GSK3/SHAGGY-like kinase. Science 295, 1299–1301. doi: 10.1126/science.1065769
Li, J., Nam, K. H., Vafeados, D., and Chory, J. (2001). BIN2, a new brassinosteroid-insensitive locus in Arabidopsis. Plant Physiol. 127, 14–22. doi: 10.1104/pp.127.1.14
Li, W., Ahn, I. P., Ning, Y., Park, C. H., Zeng, L., Whitehill, J. G., et al. (2012a). The U-Box/ARM E3 ligase PUB13 regulates cell death, defense, and flowering time in Arabidopsis. Plant Physiol. 159, 239–250. doi: 10.1104/pp.111.192617
Li, W., Dai, L., and Wang, G. L. (2012b). PUB13, a U-box/ARM E3 ligase, regulates plant defense, cell death, and flowering time. Plant Signal. Behav. 7, 898–900. doi: 10.4161/psb.20703
Liu, Y. C., Wu, Y. R., Huang, X. H., Sun, J., and Xie, Q. (2011). AtPUB19, a U-box E3 ubiquitin ligase, negatively regulates abscisic acid and drought responses in Arabidopsis thaliana. Mol. Plant 4, 938–946. doi: 10.1093/mp/ssr030
Logan, C. Y., and Nusse, R. (2004). The Wnt signaling pathway in development and disease. Annu. Rev. Cell Dev. Biol. 20, 781–810. doi: 10.1146/annurev.cellbio.20.010403.113126
Masuda, H. P., Cabral, L. M., De Veylder, L., Tanurdzic, M., de Almeida, E. J., Geelen, D., et al. (2008). ABAP1 is a novel plant Armadillo BTB protein involved in DNA replication and transcription. EMBO J. 27, 2746–2756. doi: 10.1038/emboj.2008.191
Mudgil, Y., Shiu, S. H., Stone, S. L., Salt, J. N., and Goring, D. R. (2004). A large complement of the predicted Arabidopsis ARM repeat proteins are members of the U-box E3 ubiquitin ligase family. Plant Physiol. 134, 59–66. doi: 10.1104/pp.103.029553
Nam, K. H., and Li, J. (2002). BRI1/BAK1, a receptor kinase pair mediating brassinosteroid signaling. Cell 110, 203–212. doi: 10.1016/S0092-8674(02)00814-0
Nusse, R. (2005). Wnt signaling in disease and in development. Cell Res. 15, 28–32. doi: 10.1038/sj.cr.7290260
Nusslein-Volhard, C., and Wieschaus, E. (1980). Mutations affecting segment number and polarity in Drosophila. Nature 287, 795–801. doi: 10.1038/287795a0
Sakai, T., Honing, H., Nishioka, M., Uehara, Y., Takahashi, M., Fujisawa, N., et al. (2008). Armadillo repeat-containing kinesins and a NIMA-related kinase are required for epidermal-cell morphogenesis in Arabidopsis. Plant J. Cell Mol. Biol. 53, 157–171. doi: 10.1111/j.1365-313X.2007.03327.x
Samuel, M. A., Mudgil, Y., Salt, J. N., Delmas, F., Ramachandran, S., Chilelli, A., et al. (2008). Interactions between the S-domain receptor kinases and AtPUB-ARM E3 ubiquitin ligases suggest a conserved signaling pathway in Arabidopsis. Plant Physiol. 147, 2084–2095. doi: 10.1104/pp.108.123380
Seo, D. H., Ryu, M. Y., Jammes, F., Hwang, J. H., Turek, M., Kang, B. G., et al. (2012). Roles of four Arabidopsis U-box E3 ubiquitin ligases in negative regulation of abscisic acid-mediated drought stress responses. Plant Physiol. 160, 556–568. doi: 10.1104/pp.112.202143
Sharma, M., Singh, A., Shankar, A., Pandey, A., Baranwal, V., Kapoor, S., et al. (2014). Comprehensive expression analysis of rice armadillo gene family during abiotic stress and development. DNA Res. doi: 10.1093/dnares/dst056. [Epub ahead of print].
Shimizu, H., Julius, M. A., Giarre, M., Zheng, Z., Brown, A. M. C., and Kitajewski, J. (1997). Transformation by Wnt family proteins correlates with regulation of β-catenin. Cell Growth Differ. 8, 1349–1358.
Siegfried, E., Perkins, L., Capaci, T., and Perrimon, N. (1990). Putative protein kinase product of the Drosophila segment polarity gene zeste-white. Nature 345, 825–829. doi: 10.1038/345825a0
Tamai, K., Zeng, X., Liu, C., Zhang, X., Harada, Y., Chang, Z., et al. (2004). A mechanism for Wnt coreceptor activation. Mol. Cell 13, 149–156. doi: 10.1016/S1097-2765(03)00484-2
Veses, V., Richards, A., and Gow, N. A. (2009). Vacuole inheritance regulates cell size and branching frequency of Candida albicans hyphae. Mol. Microbiol. 71, 505–519. doi: 10.1111/j.1365-2958.2008.06545.x
Wang, Y. X., Catlett, N. L., and Weisman, L. S. (1998). Vac8p, a vacuolar protein with armadillo repeats, functions in both vacuole inheritance and protein targeting from the cytoplasm to vacuole. J. Cell Biol. 140, 1063–1074. doi: 10.1083/jcb.140.5.1063
Wodarz, A., and Nusse, R. (1998). Mechanisms of Wnt signaling in development. Annu. Rev. Cell Dev. Biol. 14, 59–88. doi: 10.1146/annurev.cellbio.14.1.59
Woodgett, J. R. (1990). Molecular cloning and expression of glycogen synthase kinase-3/factor A. EMBO J. 9, 2431–2438.
Yan, J., Wang, J., Li, Q., Hwang, J. R., Patterson, C., and Zhang, H. (2003). AtCHIP, a U-box-containing E3 ubiquitin ligase, plays a critical role in temperature stress tolerance in Arabidopsis. Plant Physiol. 132, 861–869. doi: 10.1104/pp.103.020800
Yang, C. W., Gonzalez-Lamothe, R., Ewan, R. A., Rowland, O., Yoshioka, H., Shenton, M., et al. (2006). The E3 ubiquitin ligase activity of arabidopsis PLANT U-BOX17 and its functional tobacco homolog ACRE276 are required for cell death and defense. Plant Cell 18, 1084–1098. doi: 10.1105/tpc.105.039198
Yin, Y., Wang, Z. Y., Mora-Garcia, S., Li, J., Yoshida, S., Asami, T., et al. (2002). BES1 accumulates in the nucleus in response to brassinosteroids to regulate gene expression and promote stem elongation. Cell 109, 181–191. doi: 10.1016/S0092-8674(02)00721-3
Zeng, L. R., Qu, S., Bordeos, A., Yang, C., Baraoidan, M., Yan, H., et al. (2004). Spotted leaf11, a negative regulator of plant cell death and defense, encodes a U-box/armadillo repeat protein endowed with E3 ubiquitin ligase activity. Plant Cell 16, 2795–2808. doi: 10.1105/tpc.104.025171
Keywords: beta-catenin, Armadillo, abiotic stress, Wnt signaling, U-box
Citation: Sharma M, Pandey A and Pandey GK (2014) β-catenin in plants and animals: common players but different pathways. Front. Plant Sci. 5:143. doi: 10.3389/fpls.2014.00143
Received: 24 January 2014; Accepted: 25 March 2014;
Published online: 10 April 2014.
Edited by:
Mukesh Jain, National Institute of Plant Genome Research, IndiaReviewed by:
Maik Boehmer, Westfälische-Wihelms-Universität Münster, GermanyAshish Kumar Srivastava, Bhabha Atomic Research Centre, India
Copyright © 2014 Sharma, Pandey, and Pandey. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence:Z2twYW5kZXlAc291dGguZHUuYWMuaW4=
 Manisha Sharma
Manisha Sharma Amita Pandey
Amita Pandey Girdhar K. Pandey
Girdhar K. Pandey