Abstract
Root–root interaction research gained more and more attention over the past few years. Roots are pivotal for plant survival because they ensure uptake of water and nutrients. Therefore, detection of adjacent roots might lead to competitive advantages. Several lines of experimental evidence suggest that roots have ways to discriminate non-related roots, kin, and—importantly—that they can sense self/non-self roots to avoid intra-plant competition. In this mini-review, the existence of self/non-self recognition in plant roots will be discussed and the current knowledge on the mechanisms that could be involved will be summarized. Although the process of identity recognition is still not completely understood, interesting data are available and emerging new technologies will certainly aid to better understand this research field that can have an important biological, ecological, and agricultural impact.
INTRODUCTION
Competition among coexisting plants—most restrictively defined as a negative interaction among individuals with reduced growth, survival, or fecundity of neighbors as a consequence (Casper and Jackson, 1997)—is all about the availability of space, nutrients, water, and light. This contest is thought to be, at least in part, responsible for the plant diversity in different ecosystems (Goldberg and Barton, 1992; Wilson and Tilman, 1993). Moreover, it is, if anything, a showcase for the remarkable adaptive plasticity of plants, i.e., their ability to alter their morphology and physiology in response to environmental stimuli (Bradshaw, 1965; reviewed in Hodge, 2009; Ford, 2014).
Roots are pivotal for plant survival because they ensure the uptake of nutrients and water and they secure fixation in the soil; hence, the growing interest in the study of belowground plant competition. Plants that grow together in one soil volume depend on the same resources and rearrange their root systems to gain access to these limited supplies (Robinson, 1994). Indeed, root systems develop differently when neighboring roots are present and their growth responses vary. These responses are determined by species, relatedness, even genotype, and by self or non-self identity of the competing roots (reviewed in Schenk et al., 1999; Chen et al., 2012). The latter indicate interactions among roots of the same individual plant (“self”) or of different plants (“non-self”), whereby plants also seem to be able to recognize kin (Dudley and File, 2007). Thus, roots possess a so-called “identity recognition.” However, how do roots recognize other roots? Although root–root interaction studies are extremely complex due to the many factors that influence root competition and the inaccessibility of the belowground root system, recent efforts addressed this still open question. Nevertheless, the exploration of molecular mechanisms of root identity recognition is limited. With next generation sequencing methods becoming more available in research practices, it seems only timely to address this question by using such state of the art techniques, for which proteomics and metabolomics approaches could also prove useful.
Here, knowledge on root–root dynamics between interacting plants will be summarized and new advances will be discussed that cannot only enhance the understanding of plant evolution and biology, but can also have an impact on ecology and agriculture.
ROOT–ROOT INTERACTIONS: HOW THE ROOT SYSTEM RESPONDS TO NEIGHBORING ROOTS
A lot of experimental evidence suggests that plants alter their root growth in the presence of other plants (for a review, see Schenk et al., 1999). Pioneering work on root interaction focused mainly on spatial segregation, such as intraspecifically in Parthenium argentatum (guayule; Muller, 1946) or Prunus persica (peach) trees (Bini and Chisci, 1961), or interspecifically, such as Juglans nigra (black walnut) roots that exclude Solanum lycopersicum (tomato) roots (Massey, 1925). Roots can also be attracted to other roots; for instance, Fragaria vesca (wild strawberry) roots are drawn to Glechoma hederacea (ground ivy) roots, whereas the ivy roots avoid the strawberry roots (de Kroon, 2007). In addition, root elongation responses also occur: for instance, elongation of Fragaria chiloensis (beach strawberry) roots is stimulated upon contact with ground ivy (Semchenko et al., 2007b). Analysis of the overall root biomass of natural grassland systems revealed overyielding, no effect, or even underyielding when mixtures are compared with monocultures (Faget et al., 2013). In crops as well, effects on root growth by neighboring roots are clear, not only intraspecifically, such as for Glycine max (soybean) and Allium cepa (onion; Raper and Barber, 1970; Baldwin and Tinker, 1972), but also when intercropped. For instance, when certain Zea mays (maize) and soybean species are grown together, the roots of each plant tend to keep away from each other and become shallower than those in systems intercropped with their kin. Remarkably, not every maize variety responds in the same manner to the presence of the same soybean species (Fang et al., 2011). Similarly, roots of a Beta vulgaris (beet) variety grow faster and deeper than legume roots grown in the same soil, providing a competitive advantage (Tosti and Thorup-Kristensen, 2010). Furthermore, roots can accumulate in the top soil, such as in mixed grassland species (Mamolos et al., 1995). The root density in the top soil of Acacia saligna (orange wattle) trees intercropped with Sorghum bicolor (sorghum) is also higher than that of monocultures (Lehmann et al., 1998). Intriguingly, root allocations might be influenced by kin recognition, i.e., the ability to discriminate siblings from strangers. In Cakile edentula (sea rocket) and Impatiens pallida (pale touch-me-not), root allocation is larger and smaller in groups of strangers than of siblings, respectively (Dudley and File, 2007; Murphy and Dudley, 2009). In conclusion, responses (Figure 1A) vary in several experiments, indicating that roots sense the presence of other roots and that identity recognition might be important in altering root growth.
FIGURE 1
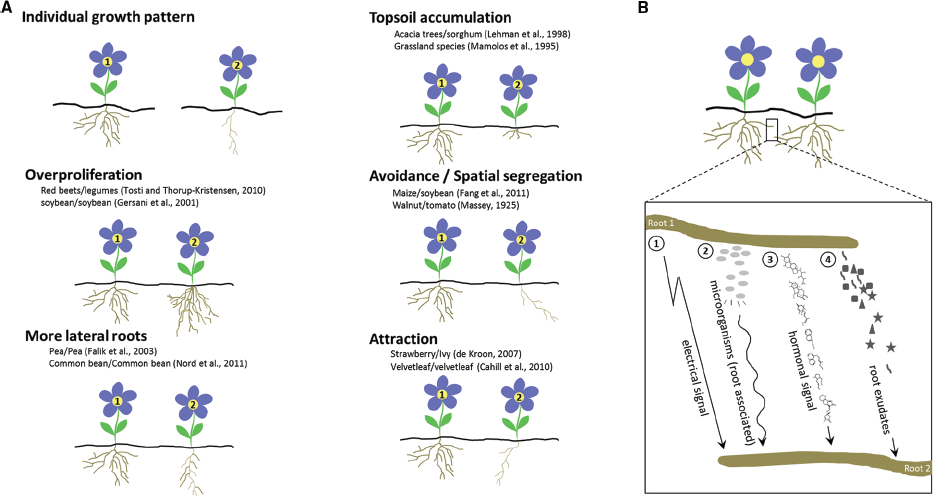
Figure 1. Common root responses to neighboring plants and possible mechanisms of root identity recognition. (A) Range of root responses toward neighboring roots. Two plants are depicted (plant 1 and 2) of which plant 2 shows a differential root response because it is neighbored with plant 1. Plants 1 and 2 can be either of the same species or of different species. Examples (both intra- and interspecifically) of interacting species, are provided for each response: overproliferation of the root system, i.e., root biomass changes which may include main root length increases, more adventitious rooting, more and higher order lateral roots, etc.; increased lateral rooting; accumulation in the top soil; spatial segregation; root attraction. (B) Four proposed mechanisms in self/non-self root recognition: resonant amplification of electrical or hormonal signals (Schenk et al., 1999; Falik et al., 2003); root associated microorganisms and/or their secreted substances (Steenhoudt and Vanderleyden, 2000); and soluble metabolites in root exudates (Biedrzycki et al., 2010; Caffaro et al., 2011; Fang et al., 2013; Semchenko et al., 2014). A non-self root–root interaction is shown.
IDENTITY RECOGNITION AND ITS IMPORTANCE
In plants, identity recognition has been unequivocally demonstrated for several biological processes that affect plant fitness, reproduction, and/or survival. For instance approximately 60% of the angiosperms show self-incompatibility which ultimately serves to prevent self-fertilization (for reviews, see Kitashiba and Nasrallah, 2014; Sawada et al., 2014). Moreover, host recognition systems of parasitic plants (Cardoso et al., 2011) and recognition of potential pathogens basically relies on the ability to discriminate “self” and “non-self” (Sanabria et al., 2008).
As shown above, identity recognition is also of great importance for the outcome of belowground interactions. Self/non-self recognition had first been reported for the desert shrub Ambrosia dumosa (burro-weed). Roots of Ambrosia stop growing when root systems from other Ambrosia plants (i.e., the same plant population) are encountered, seemingly as an avoidance response, but not when roots from the same physiological individual (i.e., self roots) are sensed (Mahall and Callaway, 1991, 1992). In contrast, roots of Larrea tridentata (creosote bush), also a desert shrub, tolerate neither other Larrea nor Ambrosia roots in their proximity (Mahall and Callaway, 1991, 1992). Since these first findings, self/non-self identity recognition has been studied in various species, but no uniform responses are observed. In some species, non-self roots seem to promote root growth traits, whereas root growth is not enhanced by self roots (Table 1). Nonetheless, a clear conclusion could be drawn, namely that to be recognized as self roots, they must be physiologically attached. Detached roots, even when they originate from the same and, thus, genetically identical individual, are recognized as non-self (Mahall and Callaway, 1991; Gruntman and Novoplansky, 2004; Falik et al., 2006; Nord et al., 2011). In contrast, kin recognition or recognition of the same species/population, but not of the same individual would occur via different mechanisms. In Arabidopsis, photosensory receptors distinguish between light signals from kin and other neighbors and allow leaf repositioning to decrease light competition (Crepy and Casal, 2014). The distinction between strangers and siblings could also be based on genetic similarity, although both in Pisum sativum (pea) and Buchloe dactyloides (buffalograss; Falik et al., 2003; Gruntman and Novoplansky, 2004) intermediate responses and phenotypes during self/non-self root experiments hint at certain overlaps.
Table 1
| Parameter | Effect | Species | Reference |
|---|---|---|---|
| Root elongation rate | Decline for non-self roots, no effect for self roots | Ambrosia dumosa | Mahall and Callaway (1991) |
| Decline for both self and non-self roots | Larrea tridentata | Mahall and Callaway (1991) | |
| Root growth (length and/or number) | No effect | Andropogon gerardii | Markham and Halwas (2011) |
| Reduced for non-self, no effect for self | Arabidopsis thaliana | Biedrzycki et al. (2010) | |
| Fewer and shorter roots toward self | Buchloe dactyloides | Gruntman and Novoplansky (2004) | |
| Lateral roots | More and longer lateral roots toward non-self | Pisum sativum | Falik et al. (2003) |
| Root segregation | Roots avoid non-self roots, no effect for self roots | Arabidopsis thaliana | Caffaro et al. (2011) |
| Spatial segregation for self roots | Fragaria chiloensis | Holzapfel and Alpert (2003) | |
| Attraction for same genotype, avoidance for different genotypes | Oryza sativa | Fang et al. (2013) | |
| No effect | Fragaria vesca | Semchenko et al. (2007b) | |
| Avoidance for self and non-self | Glechoma hederacea | Semchenko et al. (2007b) | |
| Root biomass | No effect of neighboring plants | Avena sativa | Semchenko et al. (2007a) |
| Self-inhibition | Glycine max | Gersani et al. (2001) | |
| Less biomass in presence of self roots | Trifolium repens | Falik et al. (2006) | |
| Overproliferation toward non-self | Phaseolus varigaris | Maina et al. (2002) |
Overview of self/non-self root recognition studies: parameters analyzed, outcome, and used species.
The impact of root growth inhibition by other plants, whether they are self, strangers, or kin, can be intuitively explained in terms of “space defense” and resource availability. Indeed, inhibition is less demanding than direct competition for the same nutrients in the shared space (for a review, see Schenk et al., 1999). In contrast, root growth overproliferation might maximize the nutrient uptake, but could also affect propagation. In this so-called “tragedy of the commons” that is demonstrated in soybean (Gersani et al., 2001) and Phaseolus varigaris (Kenya beans; Maina et al., 2002), the root overproliferation response reduces the reproductive biomass without competitive advantages. However, other studies (Holzapfel and Alpert, 2003; Gruntman and Novoplansky, 2004) do not indicate shoot or reproductive mass changes, so root growth inhibition is not always paralleled by obvious aboveground modifications and could be species specific. Besides biomass alterations, quick physiological responses can be mediated by root identity recognition. In pea, root competition does not affect photosynthesis, although leaf dark respiration is halved, whereas root respiration increases in the vicinity of non-self roots (Meier et al., 2013).
Root overproduction, at the expense of reproductive or shoot biomass, suggests that regulation of the identity recognition can be an important means to increase crop yields. In the cases in which the tragedy of the commons had been observed, isolation of plants from each other could enhance yield (e.g., biomass, seeds, fruits, and flowers) with the same input of water and nutrients (Maina et al., 2002). Moreover, data derived from transcriptomics techniques, only recently applied in the field of root identity recognition, can prove useful. Thus far, differentially expressed gene sets have been reported for intraspecific and interspecific competition of Arabidopsis thaliana (thale cress) plants and for Centaurea maculosa (spotted knapweed; Broz et al., 2008; Biedrzycki et al., 2011; Masclaux et al., 2012; Schmid et al., 2013). The existence of a core gene set involved in identity recognition, as suggested by Schmid et al. (2013), merits further research. Moreover, identity recognition seems to be evolutionarily conserved because it has been reported already in spermatophytes (Gorelick and Marler, 2014). The molecular biology behind root identity recognition should be tested exhaustively, for example, by employing deep sequencing methods. Biedrzycki et al. (2011) and Schmid et al. (2013) demonstrated that the molecular responses of root and pathogen recognition overlap. Comparison of their datasets with datasets of plant growth promoting rhizobacteria that can accelerate the growth and vegetative phase of plants (Poupin et al., 2013) can be most relevant for crops. The results might have great applications in agricultural practices, in addition to the discovery of the mechanisms responsible for identity recognition that have long been elusive.
MECHANISMS OF IDENTITY RECOGNITION IN PLANTS
For self-incompatibility, specific ligands are involved (Sawada et al., 2014). Volatile cues from self cuttings of Artemisia tridentata (sagebrush) increase herbivore resistance when compared to volatiles from non-self cuttings (Karban and Shiojiri, 2009). In addition, light signals mediate discrimination between kin and neighbors, leading to leaf repositioning which requires auxin biosynthesis (Crepy and Casal, 2014). Regarding root communication (Figure 1B), mediation through electrical signals has been proposed (Schenk et al., 1999). Furthermore, experimental data in pea demonstrate that hormonal rhythms might be implicated (Falik et al., 2003), as corroborated by Gruntman and Novoplansky (2004) who concluded that an unknown physiological mechanism (i.e., electrical or hormonal rhythm) might be responsible for root discrimination in buffalograss. Differential internal oscillatory signals and their resonant amplification would lead to the recognition of a non-self root. Alternatively, perception of neighboring roots has been proposed to be attributed to associated microorganisms and their secreted substances (Steenhoudt and Vanderleyden, 2000). Transcriptomics data have confirmed this hypothesis by the striking overlap of genes associated with plant reactions to neighbors and with responses to pathogens (Biedrzycki et al., 2011; Schmid et al., 2013). Surprisingly, however, in axenic cultures of Arabidopsis plants exposed only to root exudates—i.e., the mixture of compounds that are actively secreted or passively released by roots (Bais et al., 2006)—of strangers, the induction of lateral root formation is higher than that after exposure to sibling exudates. This observation suggests that a soluble chemical, originating from root exudation, might be responsible for identity recognition (Biedrzycki and Bais, 2010a,b; Biedrzycki et al., 2010). By means of the root secretion inhibitor sodium orthovanadate, which blocks active root secretion of several phenolic compounds in Arabidopsis, seedlings no longer recognize strangers, implying that active secretion by roots is required for kin recognition (Biedrzycki et al., 2010). However, self/non-self recognition is not influenced by the secretion inhibitor, confirming that two separate identity recognition mechanisms exist. Caffaro et al. (2011) demonstrated that self and non-self exudates similarly reduce root growth, but that addition of activated charcoal, specifically reverses the effect of self roots on root growth, indicating that reduced amounts of secondary metabolites in the medium affected self/non-self recognition. Consistently, root proximity is important for interaction responses in Oryza sativa (rice). Exclusion of aerial interactions by shoot separation experiments hinted at the induction of interactions by root exudates that diffuse into the medium rather than by physical contact (Fang et al., 2013). In Deschampsia cespitosa (tufted-hair grass), root exudates have also been demonstrated as cues of neighbor identity that control root mass and morphology (Semchenko et al., 2014). Noteworthy, root-object recognition might occur via allelopathic root exudates, as shown in pea (Falik et al., 2005), but, according to recent evidence in rice, could also be mediated via different processes that require physical contact of the root tip with the obstacle (Fang et al., 2013). Unraveling the exact nature of the signals that trigger identity recognition would be a gigantic leap forward in root–root interaction studies.
SELF/NON-SELF RECOGNITION IN ROOTS: PITFALLS
The interpretation of some of the experiments concerning self/non-self recognition remains somewhat controversial (for a review, see Chen et al., 2012). As already mentioned above (see also Table 1), different species are used to study self/non-self identity recognition in plants, making it difficult to draw clear conclusions due to likely species-specific and genotype-specific effects (Fang et al., 2011). Moreover, the root growth strategies of the species under study might influence the outcome of the experiments. For instance, in the strawberry/ivy experiments, strawberries grow clonally and always spread widely within plant communities, which may well affect whether a neighboring root will be attracted or avoided (de Kroon, 2007; Faget et al., 2013).
Moreover, several parameters have been analyzed during root recognition research, such as root biomass, adaptation of root architecture/morphology, and root length (Table 1). Most studies focus on root biomass, but root architecture may well be the primary and quickest response that does not necessarily impose an altered photosynthate allocation when compared to mere root growth, as demonstrated in Phaseolus vulgaris (common bean; Nord et al., 2011).
Split-root experimental systems have been used to study self/non-self recognition in root (Gersani et al., 2001; Maina et al., 2002; O’Brien et al., 2005), but the effects of pot volume and nutrient levels, which are important factors determining root growth, are difficult to correct and will influence responses to strangers and/or identity recognition. Indeed, several results can be rationalized as responses to soil volume (Schenk, 2006; Hess and de Kroon, 2007; Markham and Halwas, 2011). Nonetheless, other experiments have unequivocally demonstrated root mass changes that depend solely on the identity of the interacting root, as, for instance, in buffalograss (Gruntman and Novoplansky, 2004). In addition, plants would react to available resources rather than to the presence of a neighbor that will, while growing, deplete the same soil zone from soil nutrients (Semchenko et al., 2007a). Indeed, roots grow preferentially where supplies are most accessible (Gersani et al., 1998; Hodge, 2009), the probable reason for avoidance of other root systems. Therefore, nutrient levels and detection of the presence of other root systems are often confounded as well (O’Brien et al., 2005; Klemens, 2008; Fang et al., 2011). For example, common bean plants will change their root system architecture and produce fewer roots in soil patches that are occupied by neighboring roots (Nord et al., 2011), possibly in relation to the phosphorus concentration in the soil, which is nearly immobile and influences the developmental plasticity of roots (e.g., Borch et al., 1999; reviewed in Ticconi and Abel, 2004). Nonetheless, transcriptomics analysis of Arabidopsis in the presence or absence of competing Hieracium pilosella (mouse-ear hawkweed) clearly indicate that sensing neighboring roots occurs before resource depletion is discovered (Schmid et al., 2013).
In addition to nutrients and soil volume, water availability must be considered as well. In Ambrosia dumosa, intraspecific water competition is thought to be the reason for growth reduction when self roots are recognized (Mahall and Callaway, 1992). Hence, local changes in the microclimate should also be taken into account; for instance, root temperature gradients affect root productivity and lead to top soil accumulation (Füllner et al., 2012). Indeed, these problems have been recognized and highly controlled experiments have been set up in which plants are grown in preconditioned liquid media (Biedrzycki et al., 2010; Caffaro et al., 2011). Nevertheless, because these systems remain artificial, their relevance in natural soil systems can be questioned. Alternatively, the use of clonal ramet pairs (Holzapfel and Alpert, 2003; Gruntman and Novoplansky, 2004; Semchenko et al., 2007b) was suggested to circumvent the above mentioned problems. However, although pot volume and nutrient levels could indeed be kept constant, disconnected ramets would still be considered to access only half the amount of nutrients (i.e., two one-root system plants) as compared to connected plants (a single two-root system plant) that have access to the full amount of nutrients. These approaches have thus been criticized as well (Hess and de Kroon, 2007).
Moreover, quantification of belowground interactions is difficult, certainly at the level of the individual root. Although beyond the scope of this minireview, recent advances in imaging technology might be helpful. A transparent gel system is now developed that allows imaging and three-dimensional reconstruction to quantitatively assess root growth parameters during interaction studies (Fang et al., 2013). Likewise, fluorescent markers and horizontal minirhizotrons imaging systems (Faget et al., 2009, 2012) have proven successful to study maize, Lolium multiflorum (Italian ryegrass), and soybean interactions. As a drawback, genetically modified plants are required that, hence, hamper ecological applications. These non-destructive technologies outcompete the mere analysis of root biomass and are promising alternatives for root–root interaction and root identity recognition studies.
In conclusion, exciting advances in the field of self/non-self recognition of roots have been made over the recent years. New imaging technologies will not only aid to analyze the root response in a non-destructive way, but will also allow kinetics studies that will help to understand the mechanisms of root identity recognition and to avoid the confusion of the effects of root interactions with those of nutrients and root volume. The identification of a core set of genes involved in neighbor detection merits further research and functional analyses. New and high-resolution chemical analysis techniques, besides state-of-the-art techniques used to measure electrical signals in planta, as well as molecular biological approaches should be utilized to clarify root–root identity recognition. The obtained results can have an enormous impact on the research in plant biology and development as well as on the agricultural and ecological research fields and practice.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Statements
Acknowledgments
The author wishes to thank Sofie Goormachtig for helpful comments on the manuscript, and Kirsten De Wilde, Hilde Nelissen, Costas Stathopoulos, and Martine De Cock for help in preparing it. Stephen Depuydt was a postdoctoral fellow of the Research Foundation-Flanders.
Conflict of interest
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1
Bais H. P. Weir T. L. Perry L. G. Gilroy S. Vivanco J. M. (2006). The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol.57, 233–266. 10.1146/annurev.arplant.57.032905.105159
2
Baldwin J. P. Tinker P. B. (1972). A method for estimating the lengths and spatial patterns of two interpenetrating root systems. Plant Soil37, 209–213. 10.1007/BF01578497
3
Biedrzycki M. L. Bais H. P. (2010a). Kin recognition in plants: a mysterious behaviour unsolved. J. Exp. Bot.61, 4123–4128. 10.1093/jxb/erq250
4
Biedrzycki M. L. Bais H. P. (2010b). Kin recognition: another biological function for root secretions. Plant Signal. Behav.5, 401–402. 10.4161/psb.5.4.10795
5
Biedrzycki M. L. Jilany T. A. Dudley S. A. Bais H. P. (2010). Root exudates mediate kin recognition in plants. Commun. Integr. Biol.3, 28–35. 10.4161/cib.3.1.10118
6
Biedrzycki M. L. Lakshmannan V. Bais H. P. (2011). Transcriptome analysis of Arabidopsis thaliana plants in response to kin and stranger recognition. Plant Signal. Behav.10, 1515–1524. 10.4161/psb.6.10.16525
7
Bini G. Chisci P. (1961). Some observations on the effects of roots of peach and pear. Riv. Ortoflorofrutt. Ital.45, 345–352[in Italian with English summary].
8
Borch K. Bouma T. J. Lynch J. P. Brown K. M. (1999). Ethylene: a regulator of root architectural responses to soil phosphorus availability. Plant Cell Environ.22, 425–431. 10.1046/j.1365-3040.1999.00405.x
9
Bradshaw A. D. (1965). Evolutionary significance of phenotypic plasticity in plants. Adv. Genet.13, 115–155. 10.1016/S0065-2660(08)60048-6
10
Broz A. K. Manter D. K. Callaway R. M. Paschke M. W. Vivanco J. M. (2008). A molecular approach to understanding plant–plant interactions in the context of invasion biology. Funct. Plant Biol.35, 1123–1134. 10.1071/FP08155
11
Caffaro M. M. Vivanco J. M. Gutierrez Boem F. H. Rubio G. (2011). The effect of root exudates on root architecture in Arabidopsis thaliana. Plant Growth Regul.64, 241–249. 10.1007/s10725-011-9564-3
12
Cahill J. F. Jr. McNickle G. G. Haag J. J. Lamb E. G. Nyanumba S. M. St. Clair C. C. (2010). Plants integrate information about nutrients and neighbors. Science328, 1657. 10.1126/science.1189736
13
Cardoso C. Ruyter-Spira C. Bouwmeester H. J. (2011). Strigolactones and root infestation by plant-parasitic Striga, Orobanche and Phelipanche spp. Plant Sci.180, 414–420. 10.1016/j.plantsci.2010.11.007
14
Casper B. B. Jackson R. B. (1997). Plant competition underground. Annu. Rev. Ecol. Syst.28, 545–570. 10.1146/annurev.ecolsys.28.1.545
15
Chen B. J. W. During H. J. Anten N. P. R. (2012). Detect thy neighbor: identity recognition at the root level in plants. Plant Sci.195, 157–167. 10.1016/j.plantsci.2012.07.006
16
Crepy M. A. Casal J. J. (2014). Photoreceptor-mediated kin recognition in plants. New Phytol.10.1111/nph.13040[Epub ahead of print].
17
de Kroon H. (2007). How do roots interact?Science318, 1562–1563. 10.1126/science.1150726
18
Dudley S. A. File A. L. (2007). Kin recognition in an annual plant. Biol. Lett.3, 435–438. 10.1098/rsbl.2007.0232
19
Faget M. Herrera J. M. Stamp P. Aulinger-Leipner I. Frossard E. Liedgens M. (2009). The use of green fluorescent protein as a tool to identify roots in mixed plant stands. Funct. Plant Biol.36, 930–937. 10.1071/FP09125
20
Faget M. Liedgens M. Feil B. Stamp P. Herrera J. M. (2012). Root growth of maize in an Italian ryegrass living mulch studied with a non-destructive method. Eur. J. Agron.36, 1–8. 10.1016/j.eja.2011.08.002
21
Faget M. Nagel K. A. Walter A. Herrera J. M. Jahnke S. Schurr U. et al (2013). Root–root interactions: extending our perspective to be more inclusive of the range of theories in ecology and agriculture using in-vivo analyses. Ann. Bot.112, 253–266. 10.1093/aob/mcs296
22
Falik O. de Kroon H. Novoplansky A. (2006). Physiologically-mediated self/nonself root discrimination in Trifolium repens has mixed effects on plant performance. Plant Signal. Behav.1, 116–121. 10.4161/psb.1.3.2639
23
Falik O. Reides P. Gersani M. Novoplansky A. (2003). Self/non-self discrimination in roots. J. Ecol.91, 525–531. 10.1046/j.1365-2745.2003.00795.x
24
Falik O. Reides P. Gersani M. Novoplansky A. (2005). Root navigation by self inhibition. Plant Cell Environ.28, 562–569. 10.1111/j.1365-3040.2005.01304.x
25
Fang S. Clark R. T. Zheng Y. Iyer-Pascuzzi A. S. Weitz J. S. Kochian L. V. et al (2013). Genotypic recognition and spatial responses by rice roots. Proc. Natl. Acad. Sci. U.S.A.110, 2670–2675. 10.1073/pnas.1222821110
26
Fang S. Gao X. Deng Y. Chen X. Liao H. (2011). Crop root behavior coordinates phosphorus status and neighbors: from field studies to three-dimensional in situ reconstruction of root system architecture. Plant Physiol.155, 1277–1285. 10.1104/pp.110.167304
27
Ford E. D. (2014). The dynamic relationship between plant architecture and competition. Front. Plant Sci.5:275. 10.3389/fpls.2014.00275
28
Füllner K. Temperton V. M. Rascher U. Jahnke S. Rist R. Schurr U. et al (2012). Vertical gradient in soil temperature stimulates development and increases biomass accumulation in barley. Plant Cell Environ.35, 884–892. 10.1111/j.1365-3040.2011.02460.x
29
Gersani M. Abramsky Z. Falik O. (1998). Density-dependent habitat selection in plants. Evol. Ecol.12, 223–234. 10.1023/A:1006587813950
30
Gersani M. Brown J. S. O’Brien E. E. Maina G. M. Abramsky Z. (2001). Tragedy of the commons as a result of root competition. J. Ecol.89, 660–669. 10.1046/j.0022-0477.2001.00609.x
31
Goldberg D. E. Barton A. M. (1992). Patterns and consequences of interspecific competition in natural communities: a review of field experiments with plants. Am. Nat.139, 771–801. 10.1086/285357
32
Gorelick R. Marler T. E. (2014). Kin recognition by roots occurs in cycads and probably in conifers. Commun. Integr. Biol.7:e28009. 10.4161/cib.28009
33
Gruntman M. Novoplansky A. (2004). Physiologically mediated self/non-self discrimination in roots. Proc. Natl. Acad. Sci. U.S.A.101, 3863–3867. 10.1073/pnas.0306604101
34
Hess L. de Kroon H. (2007). Effects of rooting volume and nutrient availability as an alternative explanation for root self/non-self discrimination. J. Ecol.95, 241–251. 10.1111/j.1365-2745.2006.01204.x
35
Hodge A. (2009). Root decisions. Plant Cell Environ.32, 628–640. 10.1111/j.1365-3040.2008.01891.x
36
Holzapfel C. Alpert P. (2003). Root cooperation in a clonal plant: connected strawberries segregate roots. Oecologia134, 72–77. 10.1007/s00442-002-1062-x
37
Karban R. Shiojiri K. (2009). Self-recognition affects plant communication and defense. Ecol. Lett.12, 502–506[Erratum Ecol. Lett. 12, 999].10.1111/j.1461-0248.2009.01313.x
38
Kitashiba H. Nasrallah J. B. (2014). Self-incompatibility in Brassicaceae crops: lessons for interspecific incompatibility. Breed. Sci.64, 23–37. 10.1270/jsbbs.64.23
39
Klemens J. A. (2008). Kin recognition in plants?Biol. Lett.4, 67–68. 10.1098/rsbl.2007.0518
40
Lehmann J. Peter I. Steglich C. Gebauer G. Huwe B. Zech W. (1998). Below-ground interactions in dryland agroforestry. For. Ecol. Manage.111, 157–169. 10.1016/S0378-1127(98)00322-3
41
Mahall B. E. Callaway R. M. (1991). Root communication among desert shrubs. Proc. Natl. Acad. Sci. U.S.A.88, 874–876. 10.1073/pnas.88.3.874
42
Mahall B. E. Callaway R. M. (1992). Root communication mechanisms and intracommunity distributions of two Mojave desert shrubs. Ecology73, 2145–2151. 10.2307/1941462
43
Maina G. G. Brown J. S. Gersani M. (2002). Intra-plant versus inter-plant root competition in beans: avoidance, resource matching or tragedy of the commons. Plant Ecol.160, 235–247. 10.1023/A:1015822003011
44
Mamolos A. P. Elisseou G. K. Veresoglou D. S. (1995). Depth of root activity of coexisting grassland species in relation to N and P additions, measured using nonradioactive tracers. J. Ecol.83, 643–652. 10.2307/2261632
45
Markham J. Halwas S. (2011). Effect of neighbour presence and soil volume on the growth of Andropogon gerardii Vitman. Plant Ecol. Divers.4, 265–268. 10.1080/17550874.2011.618515
46
Masclaux F. G. Bruessow F. Schweizer F. Gouhier-Darimont C. Keller L. Reymond P. (2012). Transcriptome analysis of intraspecific competition in Arabidopsis thaliana reveals organ-specific signatures related to nutrient acquisition and general stress response pathways. BMC Plant Biol.12:227. 10.1186/1471-2229-12-227
47
Massey A. B. (1925). Antagonism of the walnuts (Juglans nigra and J. cinerea L.) in certain plant association. Phytopathology15, 773–784.
48
Meier I. C. Angert A. Falik O. Shelef O. Rachmilevitch S. (2013). Increased root oxygen uptake in pea plants responding to non-self neighbors. Planta238, 577–586. 10.1007/s00425-013-1910-4
49
Muller C. H. (1946). Root Development and Ecological Relations of Guayule. USDA Technical Bulletin 923. Washington, DC: United States Department of Agriculture, 1–114.
50
Murphy G. P. Dudley S. A. (2009). Kin recognition: competition and cooperation in Impatiens (Balsaminaceae). Am. J. Bot.96, 1990–1996. 10.3732/ajb.0900006
51
Nord E. A. Zhang C. Lynch J. P. (2011). Root responses to neighbouring plants in common bean are mediated by nutrient concentration rather than self/non-self recognition. Funct. Plant Biol.38, 941–952. 10.1071/FP11130
52
O’Brien E. E. Gersani M. Brown J. S. (2005). Root proliferation and seed yield in response to spatial heterogeneity of below-ground competition. New Phytol.168, 401–412. 10.1111/j.1469-8137.2005.01520.x
53
Poupin M. J. Timmermann T. Vega A. Zuñiga A. González B. (2013). Effects of the plant growth-promoting bacterium Burkholderia phytofirmans PsJN throughout the life cycle of Arabidopsis thaliana. PLoS ONE8:e69435. 10.1371/journal.pone.0069435
54
Raper C. D. Barber S. A. (1970). Rooting systems of soybeans. I. Differences in root morphology among varieties. Agron. J.62, 581–584. 10.2134/agronj1970.00021962006200050009x
55
Robinson D. (1994). The responses of plants to non-uniform supplies of nutrients. New Phytol.127, 635–674. 10.1111/j.1469-8137.1994.tb02969.x
56
Sanabria N. Goring D. Nürnberger T. Dubery I. (2008). Self/nonself perception and recognition mechanisms in plants: a comparison of self-incompatibility and innate immunity. New Phytol.178, 503–514. 10.1111/j.1469-8137.2008.02403.x
57
Sawada H. Morita M. Iwano M. (2014). Self/non-self recognition mechanisms in sexual reproduction: new insight into the self-incompatibility system shared by flowering plants and hermaphroditic animals. Biochem. Biophys. Res. Commun.450, 1142–1148. 10.1016/j.bbrc.2014.05.099
58
Schenk H. J. (2006). Root competition: beyond resource depletion. J. Ecol.94, 725–739. 10.1111/j.1365-2745.2006.01124.x
59
Schenk H. J. Callaway R. M. Mahall B. E. (1999). Spatial root segregation: are plants territorial?Adv. Ecol. Res.28, 145–180. 10.1016/S0065-2504(08)60032-X
60
Schmid C. Bauer S. Müller B. Bartelheimer M. (2013). Belowground neighbor perception in Arabidopsis thaliana studied by transcriptome analysis: roots of Hieracium pilosella cause biotic stress. Front. Plant Sci.4:296. 10.3389/fpls.2013.00296
61
Semchenko M. Hutchings M. J. John E. A. (2007a). Challenging the tragedy of the commons in root competition: confounding effects of neighbour presence and substrate volume. J. Ecol.95, 252–260. 10.1111/j.1365-2745.2007.01210.x
62
Semchenko M. John E. A. Hutchings M. J. (2007b). Effects of physical connection and genetic identity of neighbouring ramets on root-placement patterns in two clonal species. New Phytol.176, 644–654. 10.1111/j.1469-8137.2007.02211.x
63
Semchenko M. Saar S. Lepik A. (2014). Plant root exudates mediate neighbour recognition and trigger complex behavioural changes. New Phytol.204, 631–637. 10.1111/nph.12930
64
Steenhoudt O. Vanderleyden J. (2000). Azospirillum, a free-living nitrogen-fixing bacterium closely associated with grasses: genetic, biochemical and ecological aspects. FEMS Microbiol. Rev.24, 487–506. 10.1111/j.1574-6976.2000.tb00552.x
65
Ticconi C. A. Abel S. (2004). Short on phosphate: plant surveillance and countermeasures. Trends Plant Sci.9, 548–555. 10.1016/j.tplants.2004.09.003
66
Tosti G. Thorup-Kristensen K. (2010). Using coloured roots to study root interaction and competition in intercropped legumes and non-legumes. J. Plant Ecol.3, 191–199. 10.1093/jpe/rtq014
67
Wilson S. D. Tilman D. (1993). Plant competition and resource availability in response to disturbance and fertilization. Ecology74, 599–611. 10.2307/1939319
Summary
Keywords
root–root interaction, root competition, identity recognition, self/non-self recognition, root growth
Citation
Depuydt S (2014) Arguments for and against self and non-self root recognition in plants. Front. Plant Sci. 5:614. doi: 10.3389/fpls.2014.00614
Received
05 September 2014
Accepted
20 October 2014
Published
06 November 2014
Volume
5 - 2014
Edited by
Boris Rewald, University of Natural Resources and Life Sciences, Vienna, Austria
Reviewed by
Catharina Meinen, Georg-August-Universität Göttingen, Germany; Harsh Bais, University of Delaware, USA
Copyright
© 2014 Depuydt.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Stephen Depuydt, Ghent University Global Campus, 119 Songdomunhwa-Ro, Yeonsu-Gu, Incheon 406840, South Korea e-mail: stephen.depuydt@ghent.ac.kr
This article was submitted to Functional Plant Ecology, a section of the journal Frontiers in Plant Science.
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.