- 1Department of Physiology, Anatomy and Microbiology, School of Life Sciences, La Trobe University, Melbourne, VIC, Australia
- 2Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, VIC, Australia
Throughout evolution, plants and pathogenic fungi have been in a constant battle where fungi have developed new mechanisms to infect plants while plants have co-evolved to combat the infection. The early stages of plant-pathogen interactions occur in the intercellular spaces of the plant tissue and thus involve a myriad of secreted factors. Traditionally, all proteins released into the extracellular space were thought to be transported via the ER-Golgi dependent classical secretory pathway. However, non-classical secretion of proteins/RNA through extracellular vesicles (EVs) has recently been reported to contribute to the milieu of extracellular molecules that mediate plant-fungal interactions (Rodrigues et al., 2007; Meyer et al., 2009). EVs can be broadly classified into exosomes and ectosomes (Keerthikumar et al., 2015). Exosomes are secreted microvesicles (30–150 nm in diameter) of endocytic origin that are released by multiple cell types and are conserved across various species (Lotvall et al., 2014; Gangoda et al., 2015). In contrast, ectosomes or shedding microvesicles are larger (100–1000 nm in diameter) and bud off directly from the plasma membrane (Keerthikumar et al., 2015). For clarity, we will collectively refer to both types of membranous vesicles as EVs in this article.
Recent studies on mammalian systems have highlighted the role of EVs in cell-cell communication and the intercellular transport of cargo (proteins, nucleic acids, and carbohydrates) (Batista et al., 2011; Cossetti et al., 2014). Whilst the role of EVs in plant-fungal interactions is still poorly defined, this non-canonical secretory pathway has been proposed as an alternative route for the secretion of virulence and defense molecules by fungi and plants, respectively (Robatzek, 2007; Rodrigues et al., 2011). The basic requirement for successful host colonization is the establishment of a parasitic relationship between the fungal pathogen and the host. This requires the induction of specific defense mechanisms in the fungus for protection against the plant innate immune system (Hayes et al., 2013). Evasion or suppression of the plant defense response is thought to be regulated by virulence factors that are secreted from the fungus and act at the plasma membrane or in the cytoplasm of the plant cell (Rodrigues et al., 2008a). Interestingly, recent studies allude to the EV-mediated transport of virulence factors from the fungus into the host cell as a more efficacious delivery mechanism than simple diffusion (Rodrigues et al., 2008a; Silverman and Reiner, 2011). Similarly, in plants, when the integrity of the cell wall is threatened by a fungal pathogen, a response is mediated, at least in part, by multivesicular bodies (MVBs) (An et al., 2006b). In mammalian cells, it is well documented that fusion of MVBs with the plasma membrane results in the secretion of exosomes (Boukouris and Mathivanan, 2015; Gangoda et al., 2015). Though the production of MVBs may not always result in the secretion of EVs, the observation that plants produce MVBs in response to a fungal infection leads to the speculation that EVs may play a critical role in plant-fungal interactions. Here, we will discuss the current knowledge on EVs in the context of human-fungal interactions and their potential roles in plant-fungal interactions.
Role of EVs in Human-fungal Interactions
Fungal EVs were first isolated from the human fungal pathogen Cryptococcus neoformans (Rodrigues et al., 2007). These EVs contained well known virulence factors such as the capsular polysaccharide glucuronoxylomannan (GXM) and the virulence regulator, glucosylceramide (Rodrigues et al., 2008a). Rodrigues and colleagues also reported the presence of several other pathogenicity-associated components that are delivered into the host via EVs. Furthermore, the isolated EVs were biologically active as they could invigorate phagocytes in the host and enhance their antimicrobial activity (Oliveira et al., 2010a). Other mammalian fungal pathogens including Histoplasma capsulatum, Candida parapsilosis, Sporothrix schenckii, and Candida albicans also deliver a variety of effector molecules in a similar manner (Albuquerque et al., 2008; Vargas et al., 2015; Gil-Bona et al., 2015b). Interestingly, the serum from patients with H. capsulatum infections contains antibodies to proteins that are present in the EVs produced by the pathogen indicating involvement of EVs in the host-pathogen interaction. Moreover, characterization of EVs from the human pathogens C. neoformans, H. capsulatum and Malassezia sympodialis has implicated them in the modulation of the host immune system and regulation of the host-pathogen interaction in favor of the fungus (Rodrigues et al., 2008b; Gehrmann et al., 2011).
Role of EVs in Plant-fungal Interactions
A major component of plant-fungal interactions is the secretion of small proteins by both organisms. Plants produce pathogenesis related (PR) proteins, many of which inhibit fungal growth or directly kill fungal cells (Sels et al., 2008). Fungi secrete virulence factors encoded by the avirulence (AVR) genes (Stergiopoulos and De Wit, 2009; Rodrigues et al., 2014; Gil-Bona et al., 2015a). However, in spite of decades of research, it is still unclear as to how these proteins cross the plasma membranes and cell walls of both species. The AVR genes AVRa10 and AVRk1of the fungal pathogen Blumeria graminis f. sp. hordei encode proteins that lack signal peptides. Despite the lack of classical secretion signal, they still enter the cells in a susceptible host plant and are required for the pathogenicity of the fungus (Ridout et al., 2006). In mammalian systems, it is well established that certain proteins with and without signal peptides are transported via EVs (Kalra et al., 2012; Simpson et al., 2012). Hence, EVs could potentially mediate the transfer of these fungal virulence factors into plant hosts (Rodrigues et al., 2008b). However, further studies are required to understand this highly complex phenomenon.
The ability of a plant to mount a rapid defense response against potential pathogens is vital to its survival. Intercellular organelle rearrangements and structural modulation of the cytoskeleton with increased focal secretion of compounds lead to the formation of a physical barrier at the attack site that might prevent successful infection (Frey and Robatzek, 2009). These modifications may involve rapid and targeted delivery of molecules via EVs. Fungal infection enhances the formation of both intracellular MVBs and paramural vesicles between the plasma membrane and cell wall in the plant cells indicating the critical role of this secretory pathway in plant innate immune response (An et al., 2006a; Wang et al., 2014). For example, vesicular structures were associated with the accumulation of phenolic compounds and H2O2 that prevented pathogenic establishment of the powdery mildew fungus B. graminis in barley (Hordeum vulgare) leaves. Though the detection of plant MVBs at the site of infection provides indirect evidence of their role in plant defense (An et al., 2006a), more work is needed to define their molecular composition and whether they do indeed transport innate immunity proteins to the site of infection and/or into the fungal cell.
MVBs were first reported in the appressoria and haustoria of the powdery mildew fungus B. graminis (Hippe, 1985; Hippe-Sanwald et al., 1992). More recently, microscopic examination confirmed the presence of membrane bound vesicles at the biotrophic interface between B. graminis and the host plant (Micali et al., 2011). Furthermore, the haustorial complexes produced by Golovinomyces orontii in infected Arabidopsis leaves have vesicles and MVBs in the haustorial body (cytoplasm), paramural space, and extrahaustorial matrix. In addition, vesicle budding and fusion of MVB-like structures with the fungal plasma membrane has been observed in this interaction, although the microscopic evidence did not reveal whether the vesicles were derived from the plant or the fungus (Micali et al., 2011). In other studies, EVs from fungal pathogen Paracoccidioides brasiliensis and H. capsulatum were reported to transport antioxidants (superoxide dismutase and catalase B) and heat shock proteins (Hsp60 and Hsp70) which may have an essential role in the fungal defense mechanism (Albuquerque et al., 2008; Vallejo et al., 2012). Only recently, Hsp60 was also reported in the proteome of EVs from the fungus Alternaria infectoria. Several species of this genus Alternaria are also considered major plant pathogens (Silva et al., 2014).
Biochemical analyses of EVs from various human fungal pathogens has revealed the presence of a variety of lipids, proteins and RNA (Peres Da Silva et al., 2015). Although observed by microscopy, EVs have not been isolated and characterized from a plant pathogenic fungus. Nevertheless, upon uptake by the plant cell, it is possible that the contents could modulate the plant response to the invading fungal pathogen by attenuating the immune response. Similarly, plant exosomes are poorly characterized and their composition is largely unknown. However, it is plausible that some plant exosomes, particularly those produced in response to fungal threat, might contain small molecules and proteins that are toxic to the fungus.
Finally, cell wall remodeling is a key process on both sides of the plant-fungus interaction (Bellincampi et al., 2014). Proteomics studies highlighted the presence of various enzymes (Endochitinase 1 precursor, Beta-glucosidase 4, Beta-1,3-glucanosyltransferase 3, and Chitin synthase B) in EVs secreted by H. capsulatum. Similarly, S. cerevisiae secreted more than 20 proteins implicated in cell wall assembly including glucanases and glucanosyl transferases (Oliveira et al., 2010b). These enzymes have the capacity to regulate synthesis and hydrolysis of cell wall components highlighting the potential role of EVs in cell wall remodeling (Albuquerque et al., 2008). Fungal cell wall synthesis is also known to be mediated by chitosomes, small vesicles containing chitin microfibrils, in Neurospora crassa (Riquelme et al., 2007). Chitosomes follow an unconventional secretory pathway to transport various components of the chitin synthase family required for fungal cell wall synthesis. Whilst the differences between chitosomes and EVs are not clearly understood, it can be speculated that EVs can also play a pivotal role in fungal cell wall remodeling. In plants, reinforcement of the cell wall is one of the major strategies of the host to restrain further invasion by the pathogen (Lionetti and Métraux, 2014). The delivery of the cell wall carbohydrates to the extending chains of insoluble polysaccharides that make up majority of the cell wall is relatively poorly understood. The role of EVs in cell wall remodeling in both the fungus and plant is understudied and needs further research.
Conclusion
Recent findings pertaining to the role of EVs in the interaction between fungal pathogens and humans have led us to ask whether EVs also have a major role in plant pathogen interactions. It is still unknown how effectors and defense molecules are packaged and transported across the plasma membranes and cell walls of the plant and fungal cells. We propose that proteins lacking secretion signals could be packaged into EVs for passage through the plasma membrane and the cell wall (Figure 1). Alternatively, proteins containing a secretion signal could be secreted into the matrix of the cell wall and then bind to EVs via a lipid binding motif. The protein then transits the cell wall as a passenger on the outer leaflet of the vesicle. Based on the discovery that EVs aid disease progression (Boukouris and Mathivanan, 2015; Gangoda et al., 2015), we propose that EVs can mediate/aid in fungal infection. This could be achieved via the transfer of effectors via EVs and/or by modulating the host cells response in favor of the fungal pathogen. Similarly, we propose that plant EVs can aid in the protection against pathogenic infections. Upon infection, defense molecules can be packaged and delivered to the site of infection to protect against the invading pathogen. Further to this, we propose that the molecular cargo present in EVs is specific to the type of insult or infection. For instance, molecular cargo present in EVs of plants during stress can be significantly different to that produced during fungal infection. Thus the molecular cargo contained within EVs of plant or fungus can serve as indicators of health, stress, and disease. Investigation of the role of EVs in plant-fungal interactions is likely to uncover a new mechanism for delivery and identification of molecules required for a productive infection and/or defense response. This knowledge will enhance our ability to protect agricultural crops against the damaging effects of fungal pathogens and securing our food sources for generations to come.
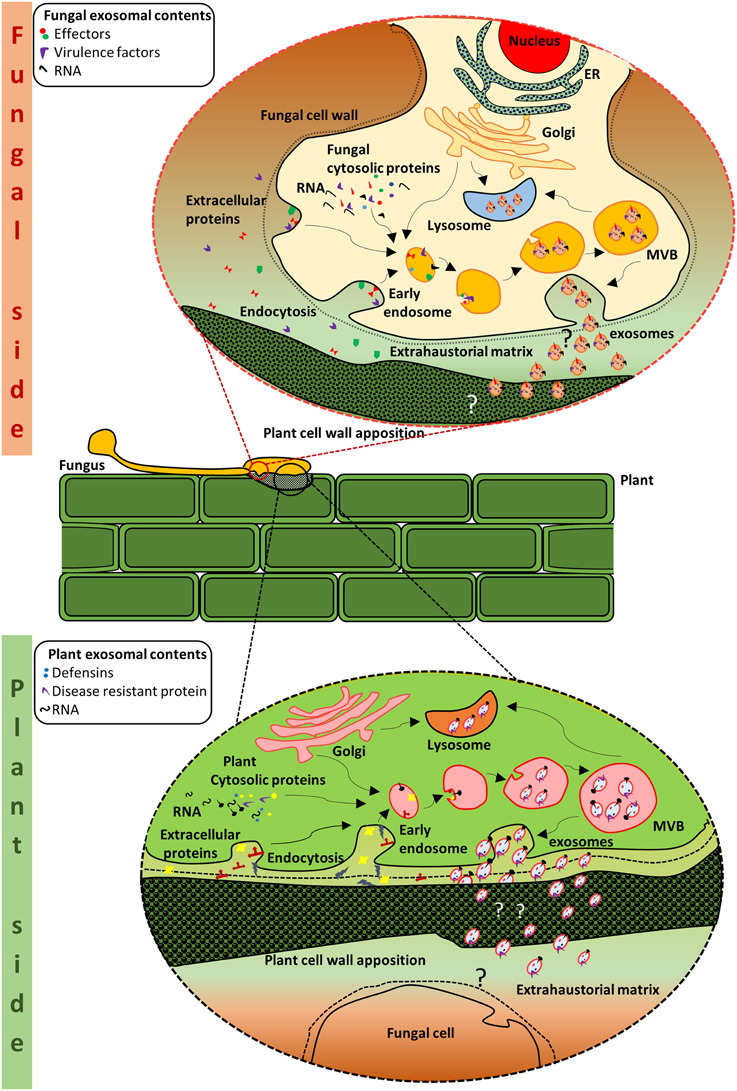
Figure 1. Schematic representation of putative crosstalk via EVs at the plant-fungal interface. Exosome biogenesis and secretion in the fungal side: MVBs are formed from the early endosomes. Within the MVBs, invagination of the limiting membrane results in the formation of intraluminal vesicles which are packaged with protein and RNA cargo from the cell. The MVBs either fuse with the plasma membrane or with the lysosome for degradation. When the MVBs fuse with the plasma membrane, the intraluminal vesicles are released as exosomes. The exosomes are considered to contain various molecules including effectors that are required for the establishment of the pathogen and/or infection. Exosome biogenesis and secretion in the plant side: Similarly, on the plant side, vesicles from the MVBs may contain innate immunity proteins and defense molecules that can impede fungal growth or lead to alterations in the fungal cell wall. Thus, the plant and its fungal counterpart could utilize the exosomes as one of the many strategies in their mutual struggle for survival.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
SM is supported by the Australian Research Council Discovery project grant (DP130100535) and Australian Research Council DECRA (DE150101777). MA is supported by the Australian Research Council Discovery project grant (DP150104386). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
Albuquerque, P. C., Nakayasu, E. S., Rodrigues, M. L., Frases, S., Casadevall, A., Zancope-Oliveira, R. M., et al. (2008). Vesicular transport in Histoplasma capsulatum: an effective mechanism for trans-cell wall transfer of proteins and lipids in ascomycetes. Cell. Microbiol. 10, 1695–1710. doi: 10.1111/j.1462-5822.2008.01160.x
An, Q., Ehlers, K., Kogel, K.-H., van Bel, A. J. E., and Hückelhoven, R. (2006a). Multivesicular compartments proliferate in susceptible and resistant MLA12-barley leaves in response to infection by the biotrophic powdery mildew fungus. New Phytol. 172, 563–576. doi: 10.1111/j.1469-8137.2006.01844.x
An, Q., Hückelhoven, R., Kogel, K. H., and van Bel, A. J. (2006b). Multivesicular bodies participate in a cell wall−associated defence response in barley leaves attacked by the pathogenic powdery mildew fungus. Cell. Microbiol. 8, 1009–1019. doi: 10.1111/j.1462-5822.2006.00683.x
Batista, B. S., Eng, W. S., Pilobello, K. T., Hendricks-Munoz, K. D., and Mahal, L. K. (2011). Identification of a conserved glycan signature for microvesicles. J. Proteome Res. 10, 4624–4633. doi: 10.1021/pr200434y
Bellincampi, D., Cervone, F., and Lionetti, V. (2014). Plant cell wall dynamics and wall-related susceptibility in plant-pathogen interactions. Front. Plant Sci. 5:228. doi: 10.3389/fpls.2014.00228
Boukouris, S., and Mathivanan, S. (2015). Exosomes in bodily fluids are a highly stable resource of disease biomarkers. Proteomics Clin. Appl. 9, 358–367. doi: 10.1002/prca.201400114
Cossetti, C., Iraci, N., Mercer, T. R., Leonardi, T., Alpi, E., Drago, D., et al. (2014). Extracellular vesicles from neural stem cells transfer IFN-gamma via Ifngr1 to activate Stat1 signaling in target cells. Mol. Cell 56, 193–204. doi: 10.1016/j.molcel.2014.08.020
Frey, N. F. D., and Robatzek, S. (2009). Trafficking vesicles: pro or contra pathogens? Curr. Opin. Plant Biol. 12, 437–443. doi: 10.1016/j.pbi.2009.06.002
Gangoda, L., Boukouris, S., Liem, M., Kalra, H., and Mathivanan, S. (2015). Extracellular vesicles including exosomes are mediators of signal transduction: are they protective or pathogenic? Proteomics 15, 260–271. doi: 10.1002/pmic.201400234
Gehrmann, U., Qazi, K. R., Johansson, C., Hultenby, K., Karlsson, M., Lundeberg, L., et al. (2011). Nanovesicles from Malassezia sympodialis and host exosomes induce cytokine responses – novel mechanisms for host-microbe interactions in atopic eczema. PLoS ONE 6:e21480. doi: 10.1371/journal.pone.0021480
Gil-Bona, A., Llama-Palacios, A., Parra, C. M., Vivanco, F., Nombela, C., Monteoliva, L., et al. (2015a). Proteomics unravels extracellular vesicles as carriers of classical cytoplasmic proteins in Candida albicans. J. Proteome Res. 14, 142–153. doi: 10.1021/pr5007944
Gil-Bona, A., Monteoliva, L., and Gil, C. (2015b). Global proteomic profiling of the secretome of Candida albicans ecm33 cell wall mutant reveals the involvement of Ecm33 in Sap2 secretion. J. Proteome Res. doi: 10.1021/acs.jproteome.5b00411. [Epub ahead of print].
Hayes, B. M. E., Bleackley, M. R., Wiltshire, J. L., Anderson, M. A., Traven, A., and van der Weerden, N. L. (2013). Identification and mechanism of action of the plant defensin NaD1 as a new member of the antifungal drug arsenal against Candida albicans. Antimicrob. Agents Chemother. 57, 3667–3675. doi: 10.1128/AAC.00365-13
Hippe, S. (1985). Ultrastructure of haustoria oferysiphe graminis f. sp.hordei preserved by freeze-substitution. Protoplasma 129, 52–61. doi: 10.1007/BF01282305
Hippe-Sanwald, S., Hermanns, M., and Somerville, S. C. (1992). Ultrastructural comparison of incompatible and compatible interactions in the barley powdery mildew disease. Protoplasma 168, 27–40. doi: 10.1007/BF01332648
Kalra, H., Simpson, R. J., Ji, H., Aikawa, E., Altevogt, P., Askenase, P., et al. (2012). Vesiclepedia: a compendium for extracellular vesicles with continuous community annotation. PLoS Biol. 10:e1001450. doi: 10.1371/journal.pbio.1001450
Keerthikumar, S., Gangoda, L., Liem, M., Fonseka, P., Atukorala, I., Ozcitti, C., et al. (2015). Proteogenomic analysis reveals exosomes are more oncogenic than ectosomes. Oncotarget 6, 15375–15396.
Lionetti, V., and Metraux, J. P. (2014). Plant cell wall in pathogenesis, parasitism and symbiosis. Front. Plant Sci. 5:612. doi: 10.3389/fpls.2014.00612
Lotvall, J., Hill, A. F., Hochberg, F., Buzas, E. I., Di Vizio, D., Gardiner, C., et al. (2014). Minimal experimental requirements for definition of extracellular vesicles and their functions: a position statement from the International Society for Extracellular Vesicles. J. Extracell. Vesicles 3:26913. doi: 10.3402/jev.v3.26913
Meyer, D., Pajonk, S., Micali, C., O'connell, R., and Schulze-Lefert, P. (2009). Extracellular transport and integration of plant secretory proteins into pathogen-induced cell wall compartments. Plant J. 57, 986–999. doi: 10.1111/j.1365-313X.2008.03743.x
Micali, C. O., Neumann, U., Grunewald, D., Panstruga, R., and O'connell, R. (2011). Biogenesis of a specialized plant–fungal interface during host cell internalization of Golovinomyces orontii haustoria. Cell. Microbiol. 13, 210–226. doi: 10.1111/j.1462-5822.2010.01530.x
Oliveira, D. L., Freire-De-Lima, C. G., Nosanchuk, J. D., Casadevall, A., Rodrigues, M. L., and Nimrichter, L. (2010a). Extracellular vesicles from Cryptococcus neoformans modulate macrophage functions. Infect. Immun. 78, 1601–1609. doi: 10.1128/IAI.01171-09
Oliveira, D. L., Nakayasu, E. S., Joffe, L. S., Guimaraes, A. J., Sobreira, T. J., Nosanchuk, J. D., et al. (2010b). Characterization of yeast extracellular vesicles: evidence for the participation of different pathways of cellular traffic in vesicle biogenesis. PLoS ONE 5:e11113. doi: 10.1371/journal.pone.0011113
Peres Da Silva, R., Puccia, R., Rodrigues, M. L., Oliveira, D. L., Joffe, L. S., César, G. V., et al. (2015). Extracellular vesicle-mediated export of fungal RNA. Sci. Rep. 5:7763. doi: 10.1038/srep07763
Ridout, C. J., Skamnioti, P., Porritt, O., Sacristan, S., Jones, J. D. G., and Brown, J. K. M. (2006). Multiple avirulence paralogues in cereal powdery mildew fungi may contribute to parasite fitness and defeat of plant resistance. Plant Cell 18, 2402–2414. doi: 10.1105/tpc.106.043307
Riquelme, M., Bartnicki-García, S., González-Prieto, J. M., Sánchez-León, E., Verdín-Ramos, J. A., Beltrán-Aguilar, A., et al. (2007). Spitzenkörper localization and intracellular traffic of green fluorescent protein-labeled CHS-3 and CHS-6 chitin synthases in living hyphae of Neurospora crassa. Eukaryotic Cell 6, 1853–1864. doi: 10.1128/EC.00088-07
Robatzek, S. (2007). Vesicle trafficking in plant immune responses. Cell. Microbiol. 9, 1–8. doi: 10.1111/j.1462-5822.2006.00829.x
Rodrigues, M. L., Nakayasu, E. S., Almeida, I. C., and Nimrichter, L. (2014). The impact of proteomics on the understanding of functions and biogenesis of fungal extracellular vesicles. J. Proteomics 97, 177–186. doi: 10.1016/j.jprot.2013.04.001
Rodrigues, M. L., Nakayasu, E. S., Oliveira, D. L., Nimrichter, L., Nosanchuk, J. D., Almeida, I. C., et al. (2008a). Extracellular vesicles produced by Cryptococcus neoformans contain protein components associated with virulence. Eukaryotic Cell 7, 58–67. doi: 10.1128/EC.00370-07
Rodrigues, M. L., Nimrichter, L., Oliveira, D. L., Frases, S., Miranda, K., Zaragoza, O., et al. (2007). Vesicular polysaccharide export in Cryptococcus neoformans is a eukaryotic solution to the problem of fungal trans-cell wall transport. Eukaryotic Cell 6, 48–59. doi: 10.1128/EC.00318-06
Rodrigues, M. L., Nimrichter, L., Oliveira, D. L., Nosanchuk, J. D., and Casadevall, A. (2008b). Vesicular trans-cell wall transport in fungi: a mechanism for the delivery of virulence-associated macromolecules? Lipid Insights 2, 27–40. doi: 10.2217/fmb.11.112
Rodrigues, M. L., Nosanchuk, J. D., Schrank, A., Vainstein, M. H., Casadevall, A., and Nimrichter, L. (2011). Vesicular transport systems in fungi. Future Microbiol. 6, 1371–1381. doi: 10.2217/fmb.11.112
Sels, J., Mathys, J., De Coninck, B. M. A., Cammue, B. P. A., and De Bolle, M. F. C. (2008). Plant pathogenesis-related (PR) proteins: a focus on PR peptides. Plant Physiol. Biochem. 46, 941–950. doi: 10.1016/j.plaphy.2008.06.011
Silva, B. M. A., Prados-Rosales, R., Espadas-Moreno, J., Wolf, J. M., Luque-Garcia, J. L., Gonçalves, T., et al. (2014). Characterization of Alternaria infectoria extracellular vesicles. Med. Mycol. 52, 202–210. doi: 10.1093/mmy/myt003
Silverman, J. M., and Reiner, N. E. (2011). Exosomes and other microvesicles in infection biology: organelles with unanticipated phenotypes. Cell. Microbiol. 13, 1–9. doi: 10.1111/j.1462-5822.2010.01537.x
Simpson, R. J., Kalra, H., and Mathivanan, S. (2012). ExoCarta as a resource for exosomal research. J. Extracell. Vesicles 1:18374. doi: 10.3402/jev.v1i0.18374
Stergiopoulos, I., and De Wit, P. J. (2009). Fungal effector proteins. Annu. Rev. Phytopathol. 47, 233–263. doi: 10.1146/annurev.phyto.112408.132637
Vallejo, M. C., Nakayasu, E. S., Matsuo, A. L., Sobreira, T. J. P., Longo, L. V. G., Ganiko, L., et al. (2012). Vesicle and vesicle-free extracellular proteome of Paracoccidioides brasiliensis: comparative analysis with other pathogenic fungi. J. Proteome Res. 11, 1676–1685. doi: 10.1021/pr200872s
Vargas, G., Rocha, J. D., Oliveira, D. L., Albuquerque, P. C., Frases, S., Santos, S. S., et al. (2015). Compositional and immunobiological analyses of extracellular vesicles released by Candida albicans. Cell. Microbiol. 17, 389–407. doi: 10.1111/cmi.12374
Keywords: exosomes, plant-fungal interaction, extracellular vesicles, cross kingdom regulation, secretome
Citation: Samuel M, Bleackley M, Anderson M and Mathivanan S (2015) Extracellular vesicles including exosomes in cross kingdom regulation: a viewpoint from plant-fungal interactions. Front. Plant Sci. 6:766. doi: 10.3389/fpls.2015.00766
Received: 29 July 2015; Accepted: 07 September 2015;
Published: 23 September 2015.
Edited by:
Dominique Job, Centre National de la Recherche Scientifique, FranceReviewed by:
Marcio L. Rodrigues, Oswaldo Cruz Foundation, BrazilDavid John Studholme, University of Exeter, UK
Copyright © 2015 Samuel, Bleackley, Anderson and Mathivanan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mark Bleackley, Marilyn Anderson and Suresh MathivananbS5ibGVhY2tsZXlAbGF0cm9iZS5lZHUuYXU=;bS5hbmRlcnNvbkBsYXRyb2JlLmVkdS5hdQ==;cy5tYXRoaXZhbmFuQGxhdHJvYmUuZWR1LmF1
 Monisha Samuel
Monisha Samuel Mark Bleackley
Mark Bleackley Marilyn Anderson
Marilyn Anderson Suresh Mathivanan
Suresh Mathivanan