- Rubber Research Institute, Chinese Academy of Tropical Agricultural Sciences, Danzhou, China
As a key enzyme in the pentose phosphate pathway (PPP), glucose-6-phosphate dehydrogenase (G6PDH) provides nicotinamide adenine dinucleotide phosphate (NADPH) and intermediary metabolites for rubber biosynthesis, and plays an important role in plant development and stress responses. In this study, four Hevea brasiliensis (Para rubber tree) G6PDH genes (HbG6PDH1 to 4) were identified and cloned using a genome-wide scanning approach. All four HbG6PDH genes encode functional G6PDH enzymes as shown by heterologous expression in E. coli. Phylogeny analysis and subcellular localization prediction show that HbG6PDH3 is a cytosolic isoform, while the other three genes (HbG6PDH1, 2 and 4) are plastidic isoforms. The subcellular locations of HbG6PDH3 and 4, two latex-abundant isoforms were further verified by transient expression in rice protoplasts. Enzyme activity assay and expression analysis showed HbG6PDH3 and 4 were implicated in PPP during latex regeneration, and to influence rubber production positively in rubber tree. The cytosolic HbG6PDH3 is a predominant isoform in latex, implying a principal role for this isoform in controlling carbon flow and NADPH production in the PPP during latex regeneration. The expression pattern of plastidic HbG6PDH4 correlates well with the degree of tapping panel dryness, a physiological disorder that stops the flow of latex from affected rubber trees. In addition, the four HbG6PDHs responded to temperature and drought stresses in root, bark, and leaves, implicating their roles in maintaining redox balance and defending against oxidative stress.
Introduction
In plants, the pentose phosphate pathway (PPP) is a principal glycometabolism pathway that plays an important role in growth, development, and physiological stresses. The PPP produces nicotinamide adenine dinucleotide phosphate (NADPH) for reductive biosyntheses of metabolic products including fatty acids and amino acids, as well as many intermediary metabolites such as nucleotides (von Schaewen et al., 1995; Dennis et al., 1997; Wakao and Benning, 2005; Wakao et al., 2008). Glucose-6-phosphate dehydrogenase (G6PDH) is recognized as a key enzyme of the plant PPP pathway, and is involved in regulating the metabolic rates of many physiological processes (Kruger and von Schaewen, 2003). According to their subcellular locations, G6PDH molecular variants are divided into cytosolic and plastidic isoforms, both of which are encoded by nuclear genes (Schnarrenberger et al., 1973; von Schaewen et al., 1995; Kruger and von Schaewen, 2003). The G6PDH genes have been identified in several plants including potato (Graeve et al., 1994; von Schaewen et al., 1995; Wendt et al., 2000), Populus (Lin et al., 2005, 2013), wheat (Nemoto and Sasakuma, 2000), rice (Zhang et al., 2013), barley (Cardi et al., 2013, 2015), soybean (Liu et al., 2013), and Arabidopsis (Wakao and Benning, 2005; Wakao et al., 2008; Siddappaji et al., 2013), and their involvement in growth and development events including seed germination (Pu et al., 1994; Han et al., 1998) and oil accumulation (Wakao et al., 2008) has been reported. G6PDH genes also respond to various environmental stresses including salt (Nemoto and Sasakuma, 2000; Wang et al., 2008; Zhang et al., 2013; Cardi et al., 2015), drought (Scharte et al., 2009; Liu et al., 2013), heavy metals (Ślaski et al., 1996), and low temperature (Lin et al., 2013; Yang et al., 2014). For years, the studies of G6PDHs have been mainly focused on aspects of transcription and activity analysis under various stresses, stressing their roles in maintaining cell redox balance to enhance stress resistance in plants.
The Para rubber tree, Hevea brasiliensis, yields natural rubber (cis-1, 4-polyisoprene), which is an important industrial and strategic raw material. Latex (the cytoplasm of rubber-producing laticifers), flowing from severed laticifers in the bark of rubber tree after tapping (the method of rubber harvesting), is the harvested product (Metcalfe, 1967; Lewinsohn, 1991). During every tapping commonly with 2- or 3-day intervals, rubber tree discharges and loses a large amount of latex (more than 100 ml in high yielding cultivars), about one third of which is natural rubber. This lost rubber and other latex constituents including sugars, nucleic acids and proteins are regenerated within the laticifers by the time the tree is next tapped. In laticifers, glycometabolism supplies carbon and energy for rubber regeneration, and its efficiency affects yield production in rubber tree (Tupy, 1973; Jacob et al., 1988; Bouteau et al., 1991, 1992; Silpi et al., 2006, 2007). During latex regeneration, the PPP supplies NADPH for the reducing reactions in rubber biosynthesis, and supplies pentose for nucleic acid synthesis (Lynen, 1969). Various environmental factors such as cold, wind, disease, and especially wounding stresses from tapping affect rubber yield (Sookmark et al., 2002; Venkatachalam et al., 2009; Li et al., 2010; Gebelin et al., 2013; Long et al., 2015). The act of tapping the tree inevitably imposes a form of wounding stress that leads to peroxide accumulation in latex. As a reducing agent, NADPH is thought to act in clearing the superfluous peroxide to sustain the redox balance when trees experience physiological stress. As a rate-limiting enzyme, G6PDH plays an important role in maintaining the carbon flow and NADPH production in the PPP. Hitherto, there is little information available regarding the genes encoding G6PDHs in rubber tree. An analysis of the G6PDH family, including its expression and enzymatic activity will improve the understanding of the physiological roles of PPP in rubber tree.
In this study, four G6PDH genes of H. brasiliensis (HbG6PDH1 to 4) comprising one cytosolic and three plastidic isoforms were identified and cloned for the first time. Analyses of enzyme activity and expression patterns suggested that these HbG6PDHs are involved in PPP during latex regeneration, of which the cytosolic HbG6PDH3 is the major isoform that plays a role in the PPP of laticifers and in mitigating the effects of oxidative stress.
Materials and Methods
Plant Materials
Reyan7-33-97 (CATAS7-33-97 or RY7-33-97) rubber trees (H. brasiliensis) selected for this study were cultivated at the experimental plantation of the Rubber Research Institute of Chinese Academy of Tropical Agricultural Sciences (Danzhou, Hainan, China). These trees were tapped every three days for latex collection in a half spiral cut. To study the tissue expression patterns of the HbG6PDHs, seven tissues (latex, leaves, bud, seed, male flower, female flower, and bark) were collected for RNA extraction from 10-year-old mature trees of Reyan7-33-97 that had been tapped over the proceeding past 2 years. Trees of the same cultivar were used to examine the effects of various plant hormones on HbG6PDHs. To analyze the effect of tapping and wounding on HbG6PDHs expression levels, 8-year-old mature virgin trees (untapped trees) were selected.
Isolation of HbG6PDHs Genes
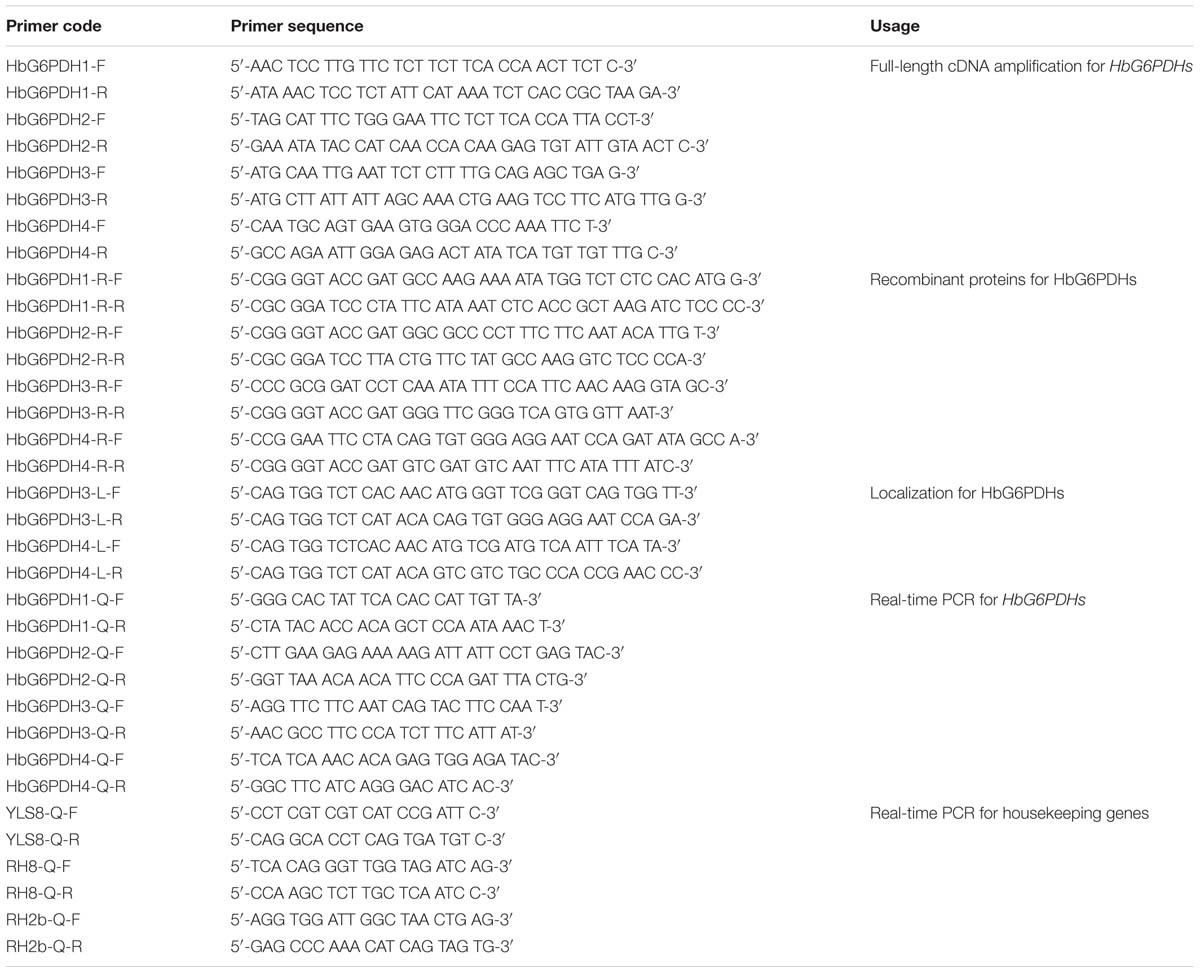
To isolate and identify the HbG6PDH sequences, the G6PDH genes of A. thaliana, P. trichocarpa, and O. sativa were used as queries to search against the transcriptome database of H. brasiliensis. Based on the sequences of resulting contigs, multiple pairs of primers were designed and used to amplify the cDNAs of HbG6PDH (Table 1). The PCR products were cloned into the pMD18-T cloning vectors (TaKaRa Biotechnology, Dalian, China), and then transformed into Escherichia coli DH5α cells. The obtained full-length cDNA sequences were used in BLAST search and other bioinformatic analysis using the NCBI database.
Construction of a Phylogenetic Tree
A phylogenetic tree of G6PDH was obtained by analyzing the deduced amino acid sequence from H. brasiliensis (AIE47266, AIE47267, AIE47268, AIE47269), P. trichocarpa (EEE79649, ERP53365, EEE94856, EEF03929), R. communis (EEF47431, EEF32168, EEF50009), A. thaliana (Q9FY99, Q8I743, Q43727, Q93ZW0, Q9IK23, Q9FJI5), Zea mays (AFW57831, DAA45780, XP008658752, ACG39996), O. sativa (BAC84352, ABF96582, ABF95637, AAL79959), and T. aestivum (BAA97662) using the Neighbor-Joining method in the MEG 5.05 software. A bootstrap analysis was performed using 1,000 replicates.
Subcelluar Localization
For subcellular localization analysis, the HbG6PDH3 and 4 were sub-cloned with BsaI site into the pCAMBIA1302-derived pBWA(V)HS vector to produce HbG6PDH-GFP fusions (Supplementary Figures S1 and S2). The specific primer pairs used were listed in Table 1. Leaves from rice seedlings that were cultured at 30°C in darkness after germination for 7 to 15 days were sampled to prepare protoplasts used in transient expression analysis. The experimental procedures were essential according to Yu et al. (2015). The transformed protoplasts were observed with laser scanning confocal microscope (Olympus FV1000, Janpan). For GFP, the excitation and emission wavelengths were 480 and 510 nm, respectively. Chloroplast autofluorescence was visualized in a detection channel with excitation and emission wavelengths of 556 and 650 nm, respectively.
Hormones, Tapping, and Wounding Treatments
To determine the expression patterns of HbG6PDHs in response in hormone treatments, four batches of five trees were selected. Three batches were treated with each hormone in 1% carboxyl methyl cellulose (CMC) at 3, 12, and 24 h before tapping, and another batch was treated with 1% CMC as a control. The four batches were tapped at the same time, and latex was collected for RNA isolation. The hormones were jasmonic acid (JA) (0.005%), abscisic acid (ABA) (200 μmol/L), cytokinin (CTK) (200 μmol/L), salicylic acid (SA) (200 μmol/L), gibberellin (GA) (100 μmol/L), and 2, 4-dichlorophenoxyacetic acid (2, 4-D) (66 μmol/L). For the ethylene treatment, the trees were treated with 1% ethephon (2-chloroethylphosphonic acid, an ethylene generator) applied on the cut at 12, 24, and 48 h before tapping with the control group treated with 1% CMC. Latex was collected for RNA isolation and determination of enzyme activity. In the experiment where mature virgin trees were opened for tapping, fifteen trees were selected and tapped in a half spiral pattern every three days, and the latex was collected from the first eight tappings for RNA isolation and enzyme activity analysis. For the wounding treatment, four batches of 10 mature virgin trees were selected, three of which were wounded at 2, 12, and 24 h before tapping, with the fourth batch was unwounded as the control. For tapping panel dryness (TPD) experiment, four batches (five trees/each batch) were selected according to TPD severities (severity < 30%, 30% < severity < 60%, 60% < severity < 90%, with healthy trees as the control). RNA extraction for treatments involving plant hormone, tapping, wounding and TPD were carried out according to the protocol of Tang (Tang et al., 2010), while the latex collection and preparation for enzyme activity determination were as previously described (Liu et al., 2015).
Stress Treatments
Stress treatments were carried out as previously described (Xiao et al., 2014). For the low temperature stress treatment, the tissue cultured plants were transferred to conical flasks containing Hoagland’s solution, placed in a growth chamber at 5°C under continuous white light, and incubated for 0, 3, 12, and 24 h. For the high temperature stress treatment, the manipulations were similar, except that the temperature was adjusted to 40°C, and the relative humidity was maintained at 80%. For the drought stress treatment, the tissue cultured plants were transferred into basic Hoagland’s solution containing 20% PEG6000, and incubated for a range of different times (0 h, 3 h, 6 h, 12 h, 1 day, 3 days, 4 days, and 7 days). For all the stress treatments, roots, leaves, and barks were sampled for RNA extraction from stressed plants at each time point, and samples from the unstressed plants were used as controls.
RNA Isolation and cDNA Synthesis
Total RNA was extracted by the protocol as described previously (Tang et al., 2007, 2010). RNA samples were treated with DNase I (TaKaRa) to eliminate the trace contaminants of genomic DNA. Integrity of the RNA samples was checked by agarose gel electrophoresis, while their concentration and quality were examined by NanoDrop 2000 (Thermo, USA). Synthesis of cDNA was performed using the RevertAidTM First Strand cDNA Synthesis Kit (Fermentas, Canada) following the manufacturer’s protocol.
Real-Time PCR
Real-time PCR was performed to analyze the HbG6PDH expression pattern with YLS8 (mitosis protein), RH8 (DEAD/DEAH box helicase), and RH2b (DEAD box RNA helicase) as the internal control genes for normalization according to Li et al. (2011). The primers for the target and internal control genes were designed using freeware IDT DNA1 with default parameters, and were then synthesized by Invitrogen China (Shanghai, China) (Table 1). Real-time PCR was carried out using the SYBR® Premix Ex Taq II (Perfect Real Time) (Takara, Dalian, China) and CFX96 TouchTM Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA). The PCR reaction system and procedures were carried out as previously described (Long et al., 2015). The Bio-Rad CFX Manager Software 3.0 was used for visualizing and analyzing the data, including the quantification cycle values, PCR efficiency and correlation coefficients. The relative fold change of expression was calculated following Vandesompele (Vandesompele et al., 2002).
Recombinant Protein Expression
To identify the enzyme activity of target proteins, recombinant vectors expressing HbG6PDHs were constructed to prokaryotic expression in E. coli. Four HbG6PDH ORFs were amplified using primers with different restrict enzyme sites (Table 1), and then inserted into the plasmid pMAL-c5E (New England BioLabs) in the same translational frame as the malE gene that encodes a maltose-binding protein (MBP). The resulting recombinant vectors were transformed into the E. coli BL21 (DE3) to generate the recombinant proteins. The transformed clones were cultivated at 37°C in liquid LB medium supplemented with 100 μg/ml ampicillin and 25 μg/ml chloramphenicol. The isopropyl β-D-1-thiogalactopyranoside (IPTG) was added at 0.4–0.6 of OD600 nm to a final concentration of 1.0 mM. After shaking at 30°C for 2 h, the induced strains were broken by ultrasonic wave with 200 w for 5 min, and then centrifuged at 10,000g for 10 mins. The supernatant was collected and stored for protein electrophoresis detection and enzyme activity assay.
Enzyme Activity Assays
For recombinant proteins, three independent transformed E. coli clones for each recombinant vector were selected to measure the enzyme activity. To analyze enzyme activity for ethylene and tapping treatments, the cytosolic serum (C-serum) was separated and purified from the latex by ultracentrifugation for 2 h at 270,000 ×g at 4°C G6PDH catalyze glucose-6-phosphate to 6-phosphoglucnate concomitant with the reduction of NADP+ to NAPDH. The G6PDH activity was measured by a spectrophotometer at 340 nm according to the increasing rate of NAPDH. One unit (U) of enzyme activity was defined as the amount of enzyme that produced 1 nmol of NAPDH in 1 min at 25°C. G6PDH activity and protein concentration were assayed using the kits purchased from SuZhou Keming Bioengineer Company (SuZhou, China), following the manufacturer’s instruction (Koide and Oda, 1959).
Results
Identification of H. brasiliensis G6PDH Genes and Functional Expression in E. coli
Four G6PDH genes were identified from the transcriptome database of rubber tree, and their full-length cDNA sequences were then obtained and submitted to GeneBank (HbG6PDH1, KJ599636.1; HbG6PDH2, KJ599637.1; HbG6PDH3, KJ599638.1; HbG6PDH4, KJ599639.1). The four HbG6PDH genes represent the entire rubber tree G6PDH family that was verified by BLAST-searching the Hevea draft genome sequence (Rahman et al., 2013). A homology analysis of the four full-length cDNA sequences indicated that nucleotide identity among them ranged from 40.6 to 67.2%, with the highest identity between HbG6PDH1 and 2, and the lowest identity between HbG6PDH1 and 4.
The four HbG6PDH genes contained different length within the open reading frame (ORF) in range of 1,548 to 1,884 bp, encoding 515 to 627 amino acid residues. The predicted molecular weights of mature HbG6PDH proteins ranged from 58.9 to 70.5 kDa (Table 2). Reflecting what was found in the cDNA sequences, a high similarity, ranging from 45.3 to 76.1%, was shared among the four HbG6PDH protein sequences. Two sites, a substrate-binding site (IDHYLG) and NADP-binding site (NEFVIRLQP), were highly conserved in the protein sequences (Figure 1). A conserved Rossman fold (GXXGXXG/A) domain was also found in the four protein sequences (Figure 1). Nevertheless, high divergence was observed in some regions of the amino acid sequences.
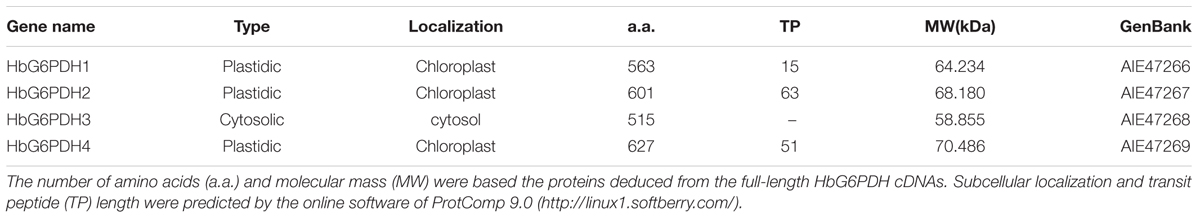
TABLE 2. Basic information for the four Hevea brasiliensis glucose-6-phosphate dehydrogenase (HbG6PDHs).
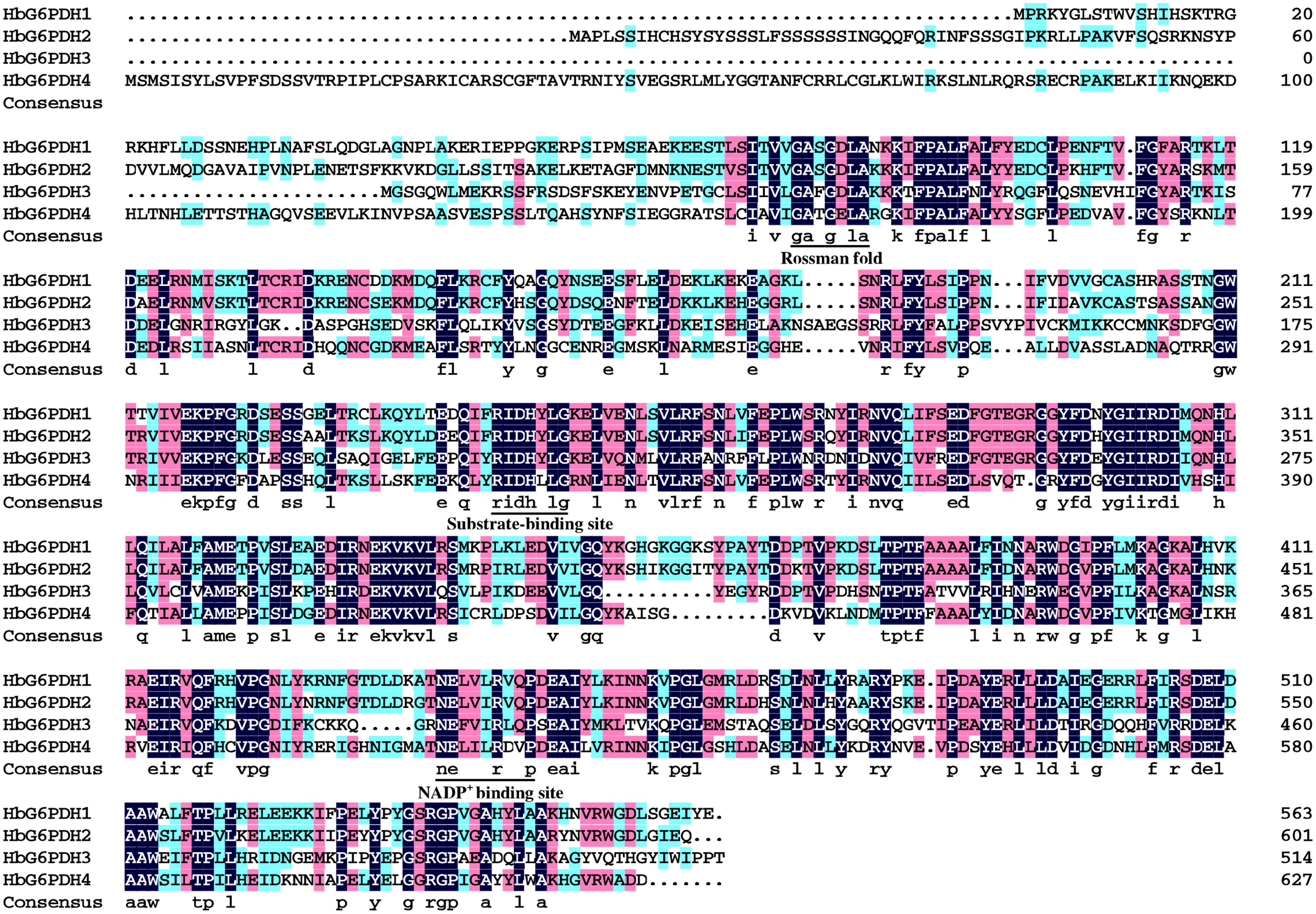

FIGURE 1. Amino acid sequence alignment of the four Hevea brasiliensis glucose-6-phosphate dehydrogenase (HbG6PDH) isoforms. Strictly conserved sequences are in white on black background; similar amino acids are in black on red and blue backgrounds. The conserved Rossman fold, substrate-binding site, NADP+ binding site were underlined.
Online software, ProtComp 9.02, was used to evaluate the presence of a transit peptide, which suggested the localization of HbG6PDH3 in the cytosol, and HbG6PDH1, 2 and 4 in the chloroplast (Table 2).
To determine whether these HbG6PDHs encode functional G6PDH enzymes, they were expressed in E. coli using the pMAL-c5E expression vector to produce MBP fusion that is due to express cytoplasmically. As expected, the four HbG6PDH genes were highly expressed in the cytosol of E. coli, and the recombinant proteins showed the expected molecular mass totaling the MBP protein and the respective HbG6PDH proteins (Figure 2A). The supernatant from the sonicated E. coli cells were tested for the G6PDH enzyme activity. As shown in Figure 2B, the extracts from E. coli cells expressing the four HbG6PDH genes showed significantly higher G6PDH activity than the control E. coli cells harboring an empty pMAL-c5E vector. These results indicated that all four HbG6PDH genes encode enzymatically active G6PDHs.
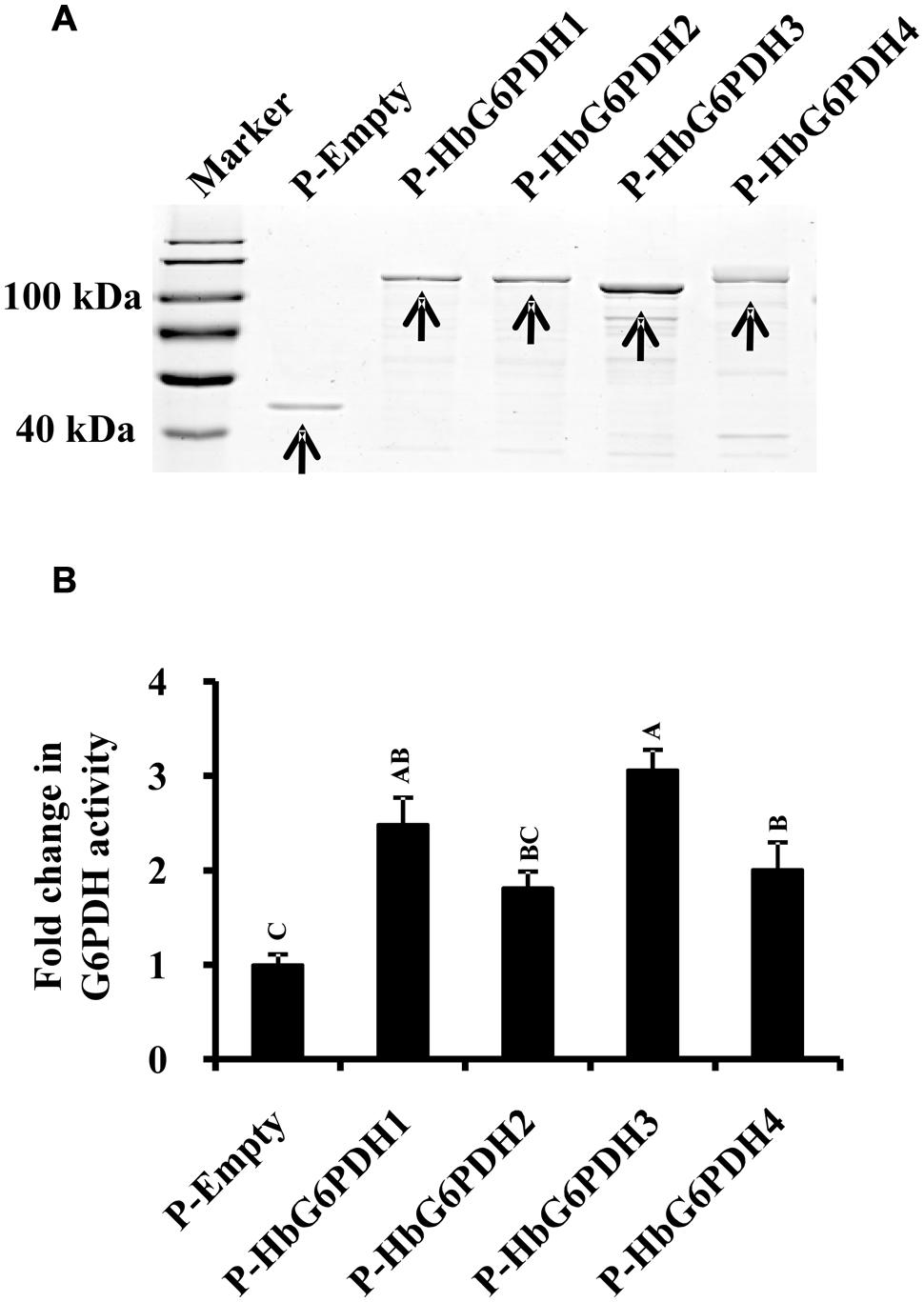
FIGURE 2. Expression of the four HbG6PDH genes in Escherichia coli. (A) Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) profile of protein extracts from ITPG-induced E. coli cells harboring the pMAL-c5E-Empty and pMAL-c5E-HbG6PDHs (HbG6PDH1 to 4). The arrowheads showed the protein bands of maltose binding protein (MBP) (P-Empty) and MBP-HbG6PDH fusions (P-HbG6PDH1 to 4). (B) Enzyme activity of the recombinant HbG6PDH proteins from pMAL-c5E-Empty and pMAL-c5E-HbG6PDHs (HbG6PDH1 to 4).
Phylogeny and Subcellular Localization of HbG6PDHs
Multiple sequence alignment was performed among four HbG6PDHs and other twenty-two G6PDH proteins of six plants, viz. P. trichocarpa, R. communis, A. thaliana, Z. mays, O. stativa, and T. aestivum. The results showed 35.4–92.4% homology in amino acid sequence among the G6PDH members, and the substrate-binding site and NADP-binding site were both highly conserved in these G6PDHs plants. However, the transit peptides associated with the different proteins were diverse, leading to varying predicted locations of the G6PDH family members. A phylogenic tree was constructed on the basis of sequence similarity of the mature proteins, and classified them into two groups (I and II) (Figure 3). Group I corresponded to plastidic G6PDH isoforms including four Arabidopsis plastidic isoforms (AtG6PDH1 to 4) (Wakao and Benning, 2005; Née et al., 2009). Group I was further divided into three clusters (A, B, and C), into which the three HbG6PDHs (HbG6PDH1, 2 and 4) were equally scattered. It is worth noting that in each cluster, the HbG6PDH isoform was closely related to its homologue from R. communis under the same spurge (Euphorbiaceae) family with rubber tree. Group II corresponded to the cytosolic isoforms, including HbG6PDH3 and the two Arabidopsis cytosolic isoforms (AtG6PDH5 and 6) which have been experimentally verified to locate in cytosol (Wakao and Benning, 2005; Wakao et al., 2008). The phylogenic analysis is consistent with the previous online prediction for the HbG6PDHs. To further verify the subcellular locations of the HbG6PDHs, two latex-abundant isoforms, HbG6PDH3 and 4 were translationally fused to GFP, and transiently expressed in rice protoplasts. As shown in Figure 4, HbG6PDH3 and 4 proteins were unequivocally located in cytosol and chloroplast, respectively.
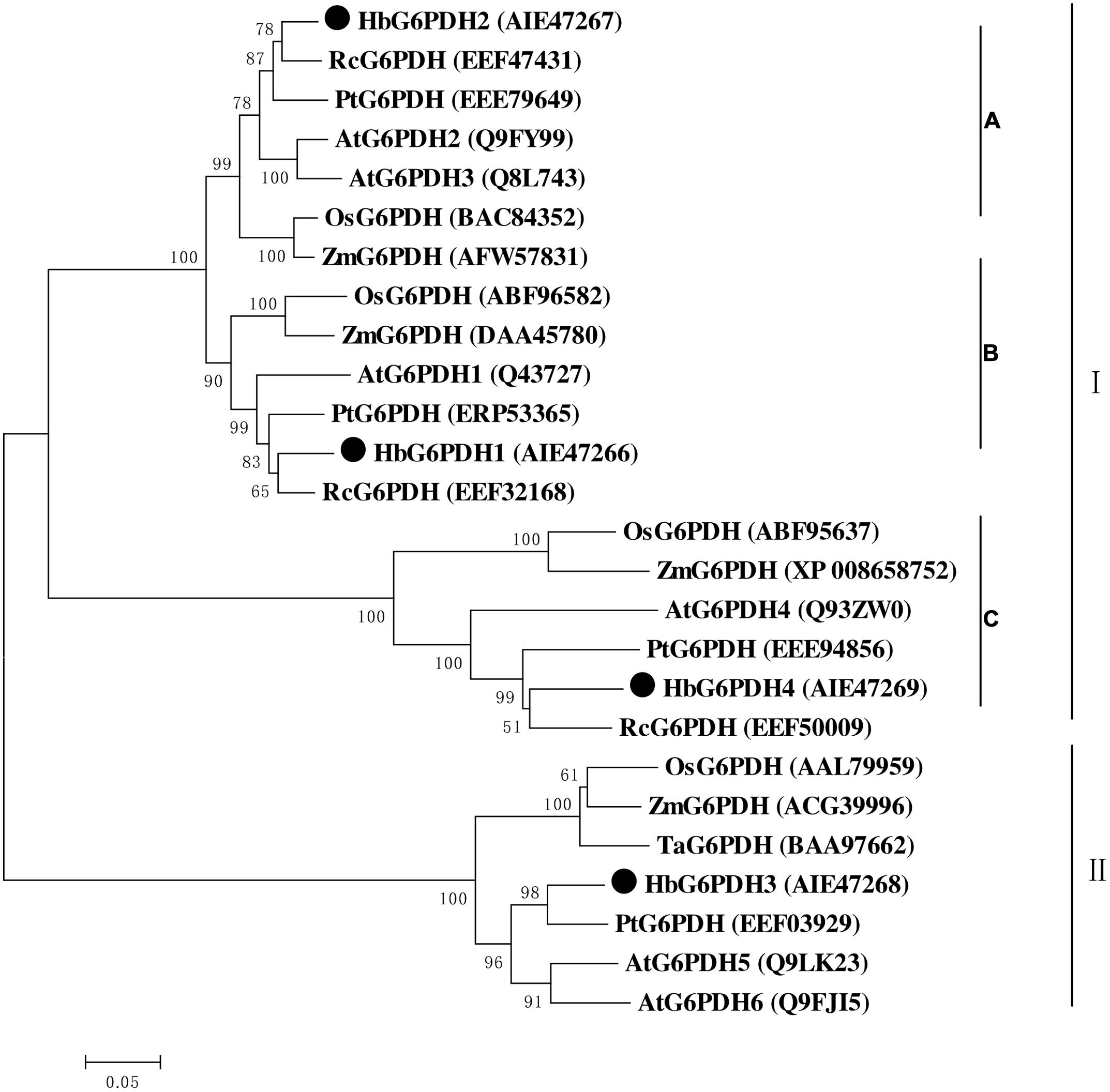
FIGURE 3. Phylogenetic analysis of 26 plant glucose-6-phosphate dehydrogenase (G6PDH) isoforms from public databases. The unrooted phylogenetic tree of 26 G6PDH proteins including 4 H. brasiliensis G6PDHs was constructed using the neighbor-joining method in the MEGA 5.05 program. Except the H. brasiliensis, the six other plant species are Populus trichocarpa, Ricinus communis, Arabidopsis thaliana, Zea mays, Oryza sativa, and Triticum aestivum. The group I were divided into three clusters (A–C).
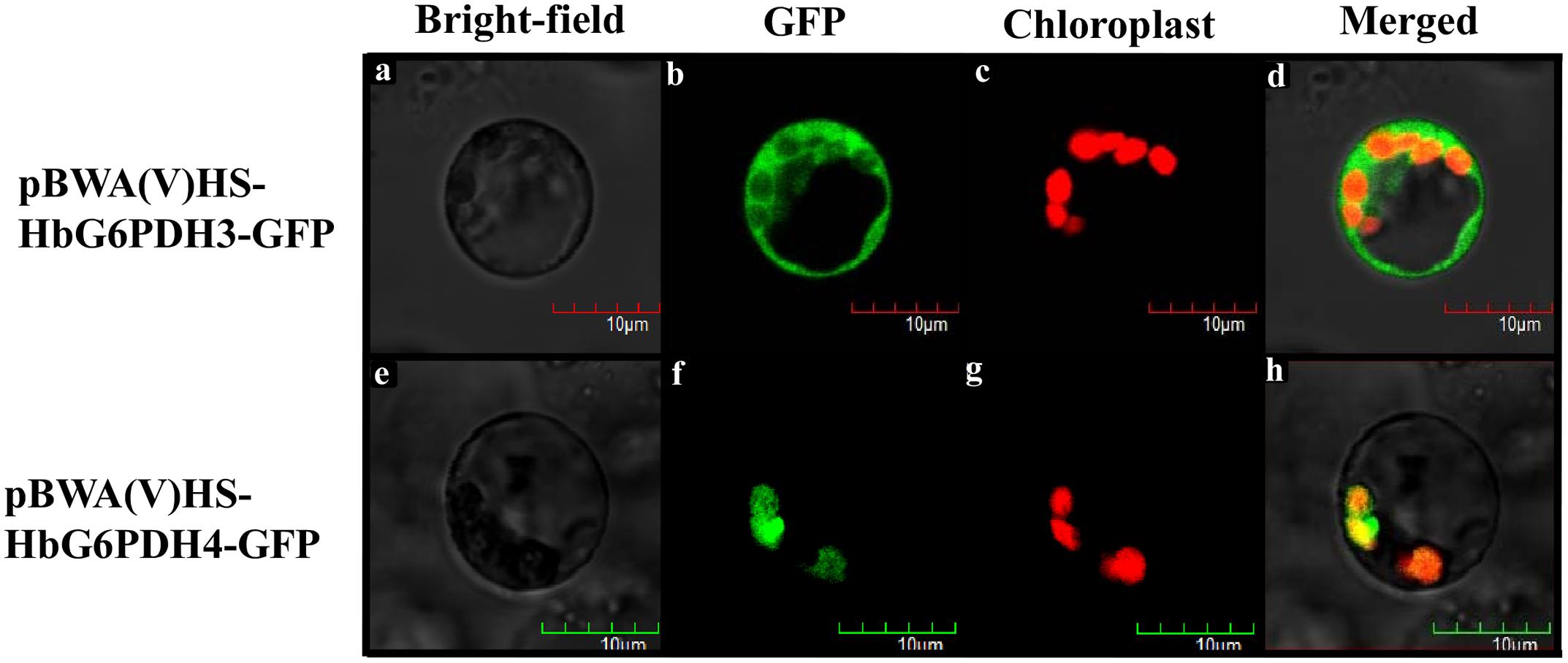
FIGURE 4. Localization of pCAMBIA1300-HbG6PDH3-GFP and pCAMBIA1300-HbG6PDH4-GFP in rice protoplasts. Scale bar = 10 μm. (a,e), bright field image; (b,f), transient expression of GFP; (c,g), chloroplast autofluorescence; (d,h), merged GFP and chloroplast image.
Tissue Expression Analysis of HbG6PDHs
The transcript levels of HbG6PDH genes were investigated in seven Hevea tissues (Figures 5A–D). Among the four genes, HbG6PDH3 and HbG6PDH4 showed a basically similar pattern of expression (Figures 5C,D). They were much higher (5- to 10-folds) expressed in latex than in other tissues. HbG6PDH1 presented somewhat a leaf-specific expression pattern, with the highest expression level in leaves, but with low or very low levels in the other tissues (Figure 5A). HbG6PDH2 showed a relatively constitutive expression; having much less variation among different tissues compared with other HbG6PDH genes (Figure 5B). Further, the relative expression abundance of the four HbG6PDH genes in latex was also analyzed. The results showed that HbG6PDH3 was the major isoform in latex, its expression being 50 fold more than that in HbG6PDH1 and 2 (Figure 5E). Although HbG6PDH4 expression in latex was only about 20% of that in HbG6PDH3, it was still much higher than the expression of the other two genes. Hereafter, the gene expression studies in latex, cytoplasm of rubber-producing laticifers will be focused on HbG6PDH3 and 4.
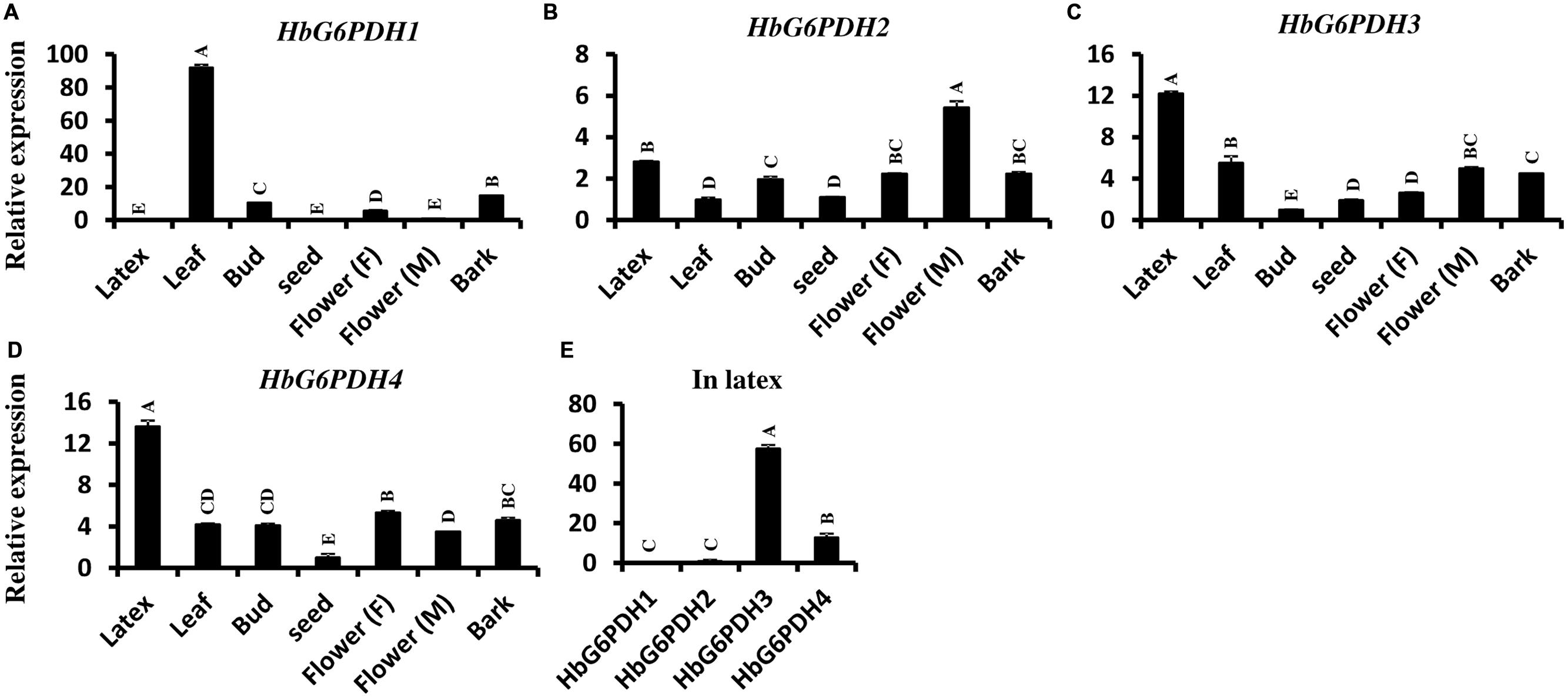
FIGURE 5. Expression of four HbG6PDH isoforms in different tissues. (A) HbG6PDH1; (B) HbG6PDH2; (C) HbG6PDH3; (D) HbG6PDH4; (E) Relative expression of four HbG6PDH isoforms in the latex. Flower (M) = male flower; Flower (F) = female flower. Different capital letters indicate a significant difference at 0.01 level.
Expression of HbG6PDH3 and 4 in Latex in Response to Tapping, Ethylene, Wounding, and TPD
In the latex of rubber trees, the expression profile of HbG6PDH3 and 4 were investigated after tapping, wounding, and ethylene application. Comparisons were also made between latex from trees suffering from TPD and latex from healthy trees. In the tapping experiment, HbG6PDH expression levels sharply declined in the first four tappings, and remained low thereafter. Compared with HbG6PDH4, the expression of HbG6PDH3 was more obviously affected by tapping, showing a 6-fold change in transcript levels in HbG6PDH3 as compared with a 3-fold with HbG6PDH4 after a series of tappings (Figure 6A). Following ethylene simulation, HbG6PDH3 and HbG6PDH4 showed different expression patterns, and the expression changes were much more obvious in HbG6PDH3 (Figure 6B). The expression of HbG6PDH3 was increased at 12 h treatment, and then declined abruptly thereafter. In comparison, there was little variation in the expression of HbG6PDH4 across treatments. After the wounding treatment, HbG6PDH3 and HbG6PDH4 expressions exhibited a similar profile of down-regulation, except that the extent of HbG6PDH3 down-regulation was more marked (5.8-folds) (Figure 6C). TPD is a complex physiological disorder that affects latex regeneration (Venkatachalam et al., 2009). In rubber trees with different degrees of TPD, the transcripts of HbG6PDH4 were significantly higher in healthy trees than in TPD ones (Figure 6D). However, there was no clear correlation between the expressional levels of the HbG6PDH3 and TPD severity.
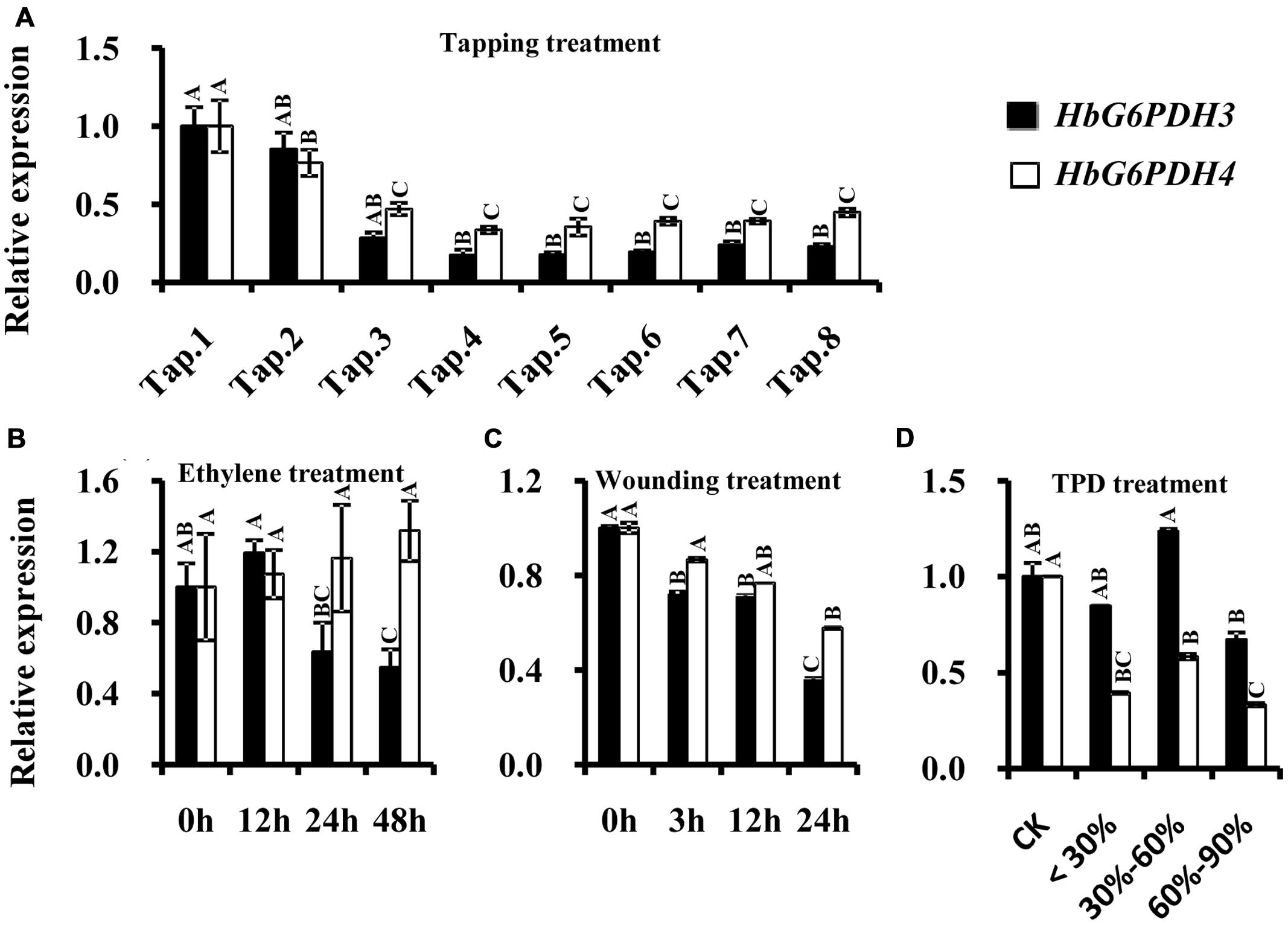
FIGURE 6. Responses of HbG6PDH3 and HbG6PDH4 to tapping (A), ethylene (B), wounding (C), and different severities of tapping panel dryness (TPD), (D). Different capital letters indicate a significant difference at 0.01 level for the same gene.
G6PDH Enzyme Activity in Latex Following Ethylene and Tapping Treatments
Glucose-6-phosphate dehydrogenase enzyme activity was measured in latex when trees were first tapped, and also following treatment with ethylene. In tapping experiment carried out on previously untapped trees, the enzyme activity gradually decreased with tapping, reaching about half of the enzyme activity recorded at the first tapping (Figure 7A). The trend in G6PDH enzyme activity with successive tappings was consistent with that of HbG6PDH3 and 4 gene expression (Figures 6A and 7A). With ethylene treatment, G6PDH enzyme activity decreased markedly after 12 h, but recovered gradually over 24 to 48 h from the time of ethylene application (Figure 7B). The change of enzyme activity did not show any obvious relationship with gene expression of either HbG6PDH3 or 4 (Figures 6B and 7B), suggesting a posttranscriptional and/or posttranslational regulation of ethylene treatment on the G6PDH activity.
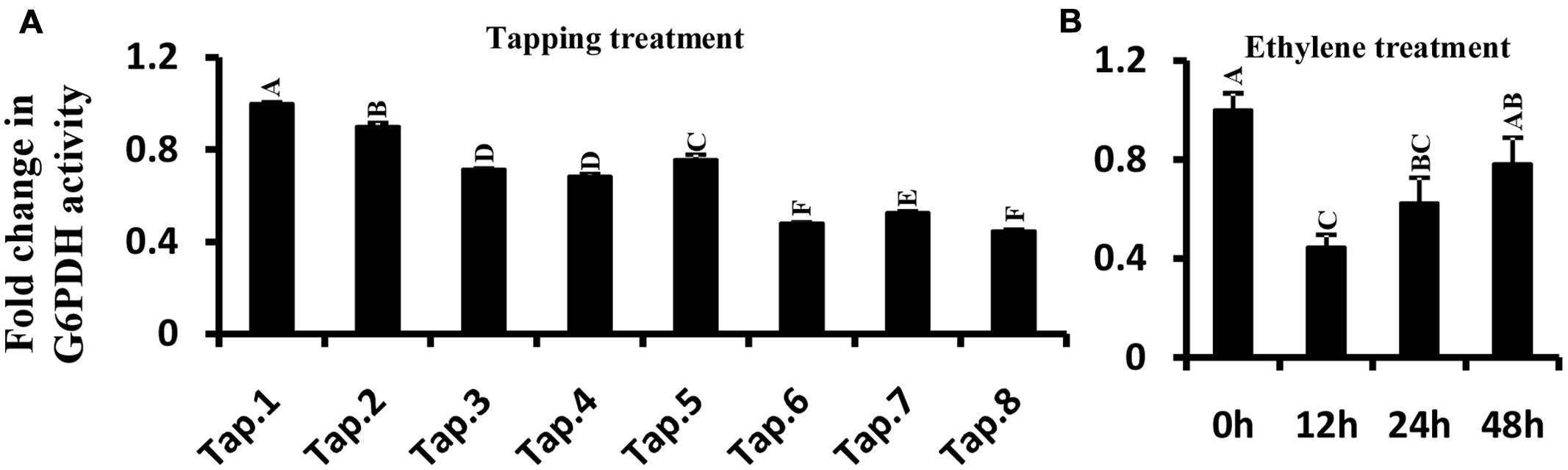
FIGURE 7. Effect of tapping (A) and ethylene treatment (B) on Hevea brasiliensis G6PDH enzyme activity in latex. Different capital letters indicate a significant difference at 0.01 level.
Expression of HbG6PDH3 and 4 in Latex Following Treatments with Plant Hormones
The expression patterns of HbG6PDH3 and 4 were investigated in latex after treatment with six hormones, viz. 2, 4-dichlorophenoxyacetic acid (2, 4-D), ABA, CTK, GA, JA, and SA (Figure 8). The two G6PDH genes presented generally similar expression patterns after ABA and JA treatments: down-regulation by ABA and up-regulation by JA, respectively. In the case of GA, the expression of HbG6PDH4 showed an initial down-regulation followed by a reversal, whereas HbG6PDH3 expression fluctuated during 24h treatment. After SA treatment, the expression of HbG6PDH3 increased for the first 12 h, and decreased thereafter; HbG6PDH4 expression showed an irregular response. In comparison, JA treatment showed a much higher stimulatory effect on the expression of HbG6PDH3 than on HbG6PDH4. In addition, HbG6PDH3 and 4 showed different expression patterns in response to the treatment of 2, 4-D and CTK. After 2, 4-D treatment, the expression of HbG6PDH3 showed a transient up-regulation at 3 h, whereas the expression of HbG6PDH4 decreased with the treatment, reaching the lowest level after 24 h. The expression of HbG6PDH3 displayed an increase for 12 h following CKT treatment, and then an obvious decrease after 24 h. In contrast, the expression of HbG6PDH4 showed a down-regulation after CTK treatment. The distinct response patterns of the HbG6PDH genes to hormone treatments suggest their differing roles in hormone-regulated metabolism.
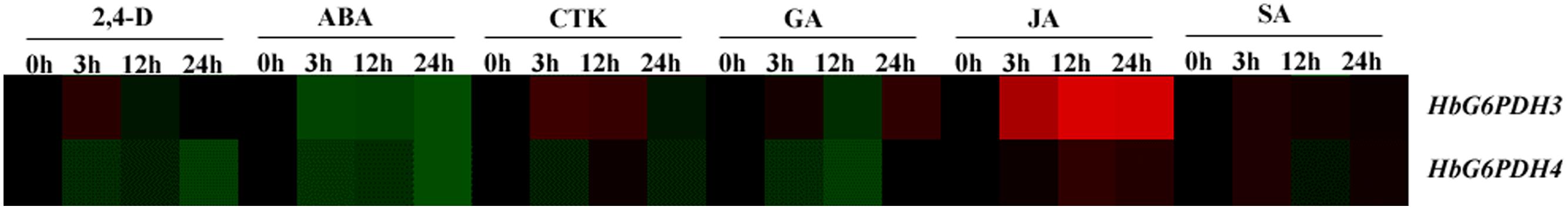
FIGURE 8. Responses of HbG6PDH3 and HbG6PDH4 to six plant hormones, viz. 2, 4-dichlorophenoxyacetic acid (2, 4-D), abscisic acid (ABA), cytokinin (CTK), gibberellic (GA), jasmonic acid (JA), and salicylic acid (SA). The expression data are shown as fold change (log2) as calculated by the ratio between the control (0 h) and treatments (3, 12, and 24 h) for each gene. Different colors represent different changes in the HbG6PDH gene expression levels: black represents no change, green represents down-regulation and red represents up-regulation. The brightness indicates the degree of expression change.
Expression of HbG6PDHs in Hevea Seedlings Responding to Abiotic Stresses
To understand the responses of HbG6PDHs to temperature and drought stresses, the expression profiles of four genes were examined in root, bark, and leaf of Hevea seedlings after stress treatment.
In root, the four HbG6PDH genes showed obvious responses to stresses of low temperature, high temperature, and drought. During low temperature treatment, HbG6PDH1, 2 and 3 showed similar patterns of an initial falling and then gradual rising expression (Figure 9A). However, HbG6PDH4 expression was strongly induced quickly after low temperature treatment, reaching the highest level at 6 h, and decreasing slightly thereafter (Figure 9A). After high temperature treatment, HbG6PDH1 to 3 showed similar expression dynamics: initial up-regulation at 3 h, then abrupt down-regulation at 6 h, and finally a recovery, although the extent of their expression change differed (Figure 9B). HbG6PDH4 expression showed a different response to the high temperature treatment, which was up-regulated at 3 h, and then decreased gradually. After drought treatment, HbG6PDH1 and HbG6PDH4 expression gradually rose with the duration of treatment (Figure 9C). The other two genes, HbG6PDH2 and HbG6PDH3, were up-regulated first and then down-regulated by drought treatment (Figure 9C).
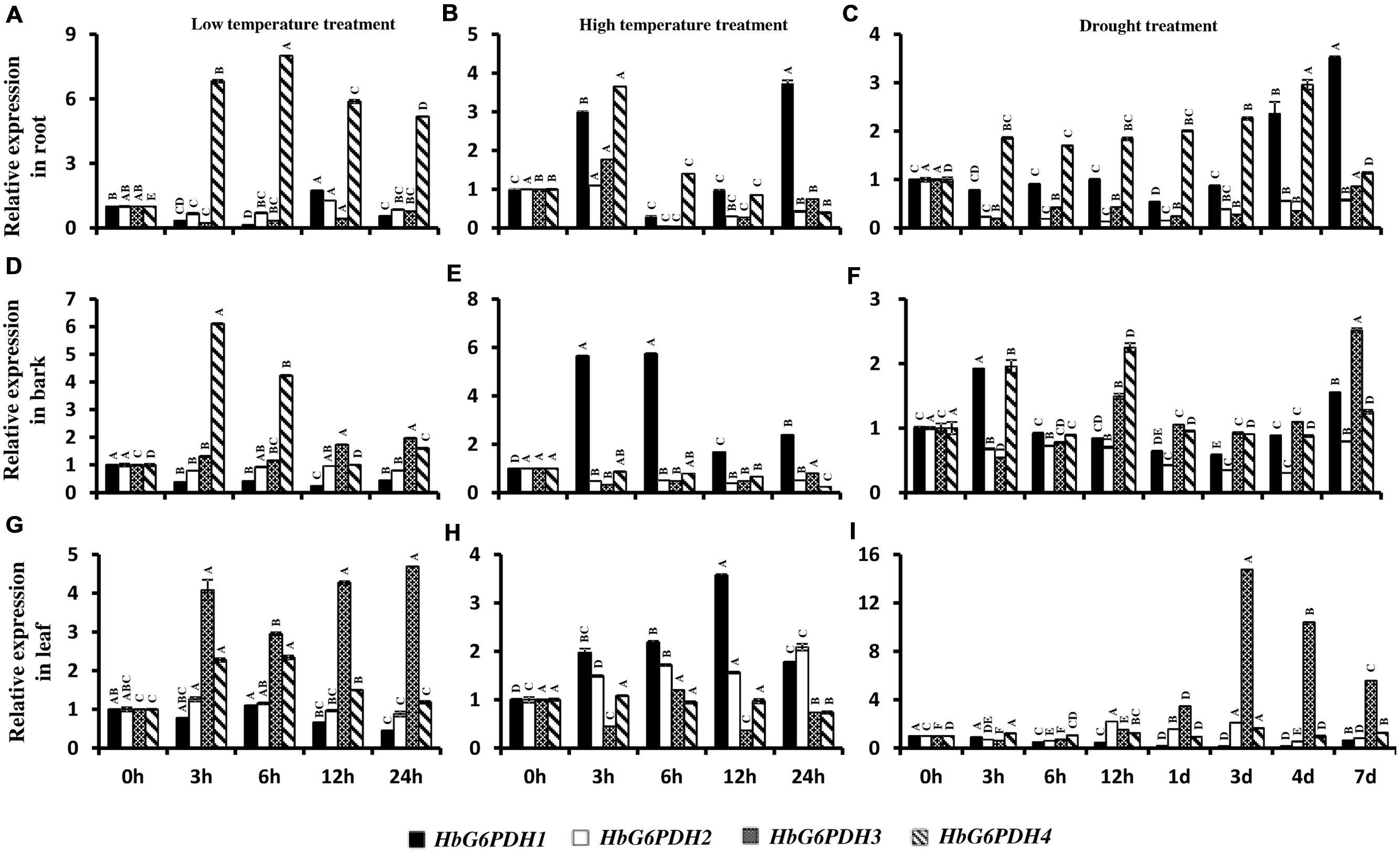
FIGURE 9. Expression of HbG6PDH isoforms in the root, bark, and leaves in response to various stress treatments, viz. low temperature (A,D,G), high temperature (B,E,H) and drought (C,F,I). Readings are expressed as values relative to the unstressed control. Different capital letters indicate a significant difference at 0.01 level among different samples for the same gene.
In bark, the four HbG6PDH genes also presented obvious responses to temperature and drought treatments. After low temperature treatment, HbG6PDH1 expression declined steadily, and HbG6PDH4 expression first increased, then decreased (Figure 9D). In comparison, the other two genes (HbG6PDH2 and 3) showed little response to low temperature treatment. After high temperature treatment, the expressions of HbG6PDH2 to 4 were down-regulated, whereas the expression of HbG6PDH1 was bolstered and peaked at 6 h before sharply receding thereafter (Figure 9E). After drought treatment, the expressions of the four HbG6PDH genes fluctuated through the duration of 7 day treatment (Figure 9F).
In leaves, the four HbG6PDH genes showed distinct expression patterns in response to three types of stresses. After low temperature treatment (Figure 9G), HbG6PDH3 and 4 expressions were up-regulated, although the extent of up-regulation was much higher for the former. Over the duration of treatment, the expression of HbG6PDH1 was decreased, whereas HbG6PDH2 was little affected. During high temperature treatment, HbG6PDH1 expression was up-regulated with peaking at 12 h, but the other three genes had no response or irregular fluctuation in expression (Figure 9H). HbG6PDH3 was obviously induced by drought treatment; its expression remained at a low level during the early stage of treatment but sharply increased from 12 h and peaked at 3 days (Figure 9I). HbG6PDH1 expression exhibited a progressive down-regulation, reaching its lowest level at 4 days. The expression trend of HbG6PDH2 was inconsistent, falling, and rising over time. Compared with the other three genes, the expression of HbG6PDH4 was little affected by drought treatment (Figure 9I).
Discussion
In plants, the PPP is an important carbohydrate metabolic pathway that generates NADPH and pentoses (5-carbon sugars), in parallel to the glycolytic pathway (Dennis et al., 1997; Kruger and von Schaewen, 2003). As one of key enzymes in the PPP, G6PDH controls carbon flow and NADPH production, playing an important role in plant growth and development. The pathway also plays a role in the response of plants to biotic and abiotic stresses.
Rubber Tree has Four G6PDH Genes
Consistent with other plants, rubber tree has a small G6PDH gene family. In this study, the four rubber tree G6PDH genes (HbG6PDH1 to 4) were first identified basing on a comprehensive H. brasiliensis transcriptome, and then verified as the entire G6PDH gene family after blasting against the recently released rubber draft genome sequence (Rahman et al., 2013). The deduced protein sequences have two highly conserved sites for substrate-binding (IDHYLG) and NADP-binding (NEFVIRLQP) (Figure 1), suggesting the corresponding genes encode active G6PDH enzymes. This kind of prediction was later demonstrated by investigating the enzymatic activities of the recombinant HbG6PDH proteins expressed in E. coli (Figure 2). Two groups of G6PDH isoforms, plastidic, and cytosolic G6PDH, are found in rubber tree of which HbG6PDH3 is cytosolic, and the other three HbG6PDHs (HbG6PDH1, 2 and 4) are plastidic. The larger number of plastidic isoforms indicates that most PPP steps take place in plastids rather than the cytosol in rubber tree, a phenomenon commonly observed in the plant kingdom (Kruger and von Schaewen, 2003). The subcellular localizations of these HbG6PDHs were initially predicted by an online software platform and consolidated by a phylogenetic analysis (Table 2; Figure 3), and subsequently demonstrated by transient expression of HbG6PDH3 and 4 in rice protoplasts using GFP fusions (Table 2; Figures 3 and 4). The divergence of G6PDH into cytosolic and plastidic types may have occurred before species divergence in plants, and plays a role in adaptation of plants to various habitats.
Different isoforms of G6PDH function in respective functional areas in plants. For example, the cytosolic G6PDHs are widespread in all tissues, especially high expression in sink tissues, whereas plastidic ones are highly expressed in green tissues and less so in sink tissues (von Schaewen et al., 1995). In our study, the plastidic HbG6PDH1 had a tissue-specific expression pattern with the highest expression level in leaves, but was low or very low expressed in a variety of sink tissues (Figure 5A), consistent with its Arabidopsis and Nicotiana orthologues (Knight et al., 2001; Wakao and Benning, 2005). However, the other two plastidic isoforms, HbG6PDH2 and 4, and the cytosolic HbG6PDH3 were abundantly expressed in different Hevea tissues. These results suggest that HbG6PDH1 functions mainly in green tissue (leaves), whereas the other three HbG6PDH genes function in diverse source and sink tissues (Figures 5B–D). Besides, the expression of HbG6PDH3 and 4 in latex had varying responses to different plant hormones, suggesting their distinct roles in various signal transduction pathways (Figure 8). Generally, HbG6PDH3 is more responsive to hormones than HbG6PDH4, and the expressions of both genes display little change in fold to most hormones. Of the six hormones and two HbG6PDH genes examined, a >2-fold expressional change was observed only for JA on HbG6PDH3, suggesting the involvement of HbG6PDH3 in JA responses in laticifers.
HbG6PDH3 and 4 are Implicated in PPP During Latex Regeneration
Nicotinamide adenine dinucleotide phosphate is mainly derived from photosynthesis and widely used in several anabolic reactions such as carbon fixation, fatty acid synthesis, and nitrogen assimilation (Dennis et al., 1997). In non-photosynthetic tissues or under non-photosynthetic conditions, the PPP is regarded as major sources of NADPH used for basic metabolism during plant growth and development (Thom et al., 1998). Beyond that, the PPP also provides abundant intermediary metabolites for other anabolic metabolisms, including the synthesis of aromatic amino acids, cell wall, and pigments (von Schaewen et al., 1995; Hauschild and von Schaewen, 2003) and is involved in many stress responses (Yang et al., 2014). As a rate-limiting enzyme in the PPP, G6PDH studies are always mainly focused on carbohydrate metabolism. Previous research has shown that G6PDH plays an important role in seed development, where the enhancement of its enzyme activity lead to the breaking of seed dormancy and the improvement of seed germination (Pu et al., 1994; Han et al., 1998). Also, the expression and activity of G6PDH is closely related with young spike development in rice (Huang et al., 2003). In peach, the G6PDH provides precursors for metabolic processes responsible for the red coloration of fruit (Kong et al., 2007).
In rubber tree, synthesis of natural rubber in laticifers uses sucrose as the main raw material (Tupy, 1973). The uptake of sucrose into laticifers relies on sucrose transporters, of which HbSUT3 is the major responsible member (Tang et al., 2010), and its cleavage into fructose and glucose in latex is conducted by a cytoplasmic alkaline/neutral invertase (HbNIN2) (Liu et al., 2015). The resultant glucose is metabolized by two parallel pathways, the PPP and the glycolytic pathway for providing metabolites (NADPH and 5-carbon sugars) and energy exploited in latex regeneration (Tupy, 1973). Of the four HbG6PDH genes, HbG6PDH3 and HbG6PDH4 are the major isoforms expressed in latex, and both have much higher expression levels in latex than the other tissues (Figures 5C,D), suggesting their major roles in the PPP in latex. In comparison, expression of the cytosolic HbG6PDH3 is more than five-fold that of the plastidic HbG6PDH4 (Figure 5E), indicating the relative importance of cytosolic PPP in latex metabolism. The HbG6PDH4 protein was predicted to locate on Frey-Wyssling particles, the sole class of plastidic type organelles suspended in latex. Both G6PDH enzyme activity and expression of HbG6PDH3 and HbG6PDH4 displayed striking responses to tapping in latex of virgin rubber trees (Figures 6A and 7A), and the pattern of these responses are negatively correlated with the tapping-stimulated rubber production (Tang et al., 2010), which suggest the involvement of PPP in latex regeneration.
Ethylene is a commonly used latex yield stimulant in rubber production. Our results showed that ethylene depressed both HbG6PDH3 expression and G6PDH enzyme activity (Figures 6B and 7B). However, compared with the effect of tapping on virgin rubber trees, Ethylene treatment posed a much lesser effect on G6PDH gene expression and enzyme activity, which is consistent with its lesser effect on the latex output. Compared with HbG6PDH4, the latex-predominant isoform HbG6PDH3 showed a much more marked response to either tapping or the ethylene treatment (Figures 6A,B). Previous studies have shown that cytosolic G6PDH is the major isoform in higher plants, accounting for 80–95% of the whole cellular activity (Cardi et al., 2013). Based on the above results, we conclude that the cytosolic HbG6PDH3 is the major isoform functioning in Hevea latex regeneration. Although the exact mechanisms remain to be investigated, we could speculate the contribution of HbG6PDH3 to latex regeneration at least from two aspects: (1) the synthesis of NADPH and 5-carbon sugars; (2) the regulation of reactive oxygen species (ROS) metabolism. NADPH and 5-carbon sugars are essential for latex regeneration that includes various anabolic metabolisms, e.g., rubber biosynthesis and nucleic acid synthesis. Rubber production is essentially a defense response, and moderate levels of ROS production will help induce defense responses and thus contribute to latex production (D’Auzac et al., 1997; Duan et al., 2010). The tapping- and ethylene-depressed HbG6PDH3 expression and G6PDH enzyme activity (Figures 6 and 7) will contribute to increased ROS production (Leopold et al., 2003; Mendez et al., 2011), thus stimulating latex production. The potential roles of HbG6PDH3 in latex regeneration were further supported by the marked simulating effect of JA on HbG6PDH3 expression during 24 h treatment (Figure 8). JA is the sole phytohormone known to induce laticifer differentiation, which constitutes one of the key factors restricting sustainable rubber production (Hao and Wu, 2000). The up-regulation of HbG6PDH3 by JA suggested its additional contribution to latex regeneration through the participation in JA-stimulated laticifer differentiation. Also, it is worthwhile to investigate the functions of HbG6PDH4 in the anabolic metabolisms occurring in the Frey-Wyssling particles that contain carotenoid pigments. The results will help address not only the biological roles of Frey-Wyssling particles, but their functions in the regulation of latex regeneration.
HbG6PDHs are Responsive to Various Stresses
In plants, G6PDH is induced by both biotic and abiotic stresses. In this regard, the enzyme plays a key role in maintaining redox balance and protecting cells from the harm caused by oxidative stress. Common environment stresses cause excessive ROS generation and accumulation, which are harmful to cellular structure and function in plant. NADPH not only acts to maintain the output of reduced glutathione (GSH), but is also involved in activity of plasma membrane (PM) NADPH oxidase which would result in hydrogen peroxide (H2O2) accumulation (Leopold et al., 2003; Mendez et al., 2011). Any decrease in G6PDH activity upsets internal redox balance, leading to disorders of signaling pathways and cell cycles in plant (Manco et al., 2011). G6DPH is a key enzyme in rice, responding to salt stress to maintain cell redox balance (Zhang et al., 2013). In soybean roots, G6PDH plays a central role in the process of H2O2 metabolism to maintain GSH and ascorbate (Asc) levels under drought stress (Liu et al., 2013). In sugarcane, assay of both activity and transcription level of ScG6PDH indicate that cytosolic ScG6PDH plays a positive role in response to various stresses, including salt, drought, heavy metal (CdCl2), and low temperature treatments (Yang et al., 2014).
In rubber production, every tapping is a wounding treatment for rubber tree, which causes oxidative stress that influences physiological metabolism in laticifers. The overtapping and/or overstimulation of rubber trees is thought to result in a complex physiological disorder known as TPD where latex coagulates in the laticifers and the tree stops yielding partially or completely (Sookmark et al., 2002; Venkatachalam et al., 2009). In this study, HbG6PDH 4 were implicated in defending TPD, with high expression levels in the healthy tree (Figure 6D). Both HbG6PDH3 and 4 showed an obvious decreasing expression response to wounding treatment, which might depress the metabolic rate of the PPP (Figure 6C). In latex, peroxides may build up when the PPP activity declines in the course of latex flow when tree is tapped. It is speculated that HbG6PDHs reduce peroxide in latex to maintain redox balance and relieve oxidative stress during every tapping. In present study, HbG6PDHs were induced under different environmental stresses, including drought and temperature stresses (Figure 9). Responses of HbG6PDHs to drought were more intense than to temperature stress, especially for HbG6PDH3 in leaves and HbG6PDH1 in root. Under low temperature stress, HbG6PDH4 had similar up-regulated expression pattern in root, bark and leaf, suggesting its roles in protecting against the effects of low temperature. Different HbG6PDHs responded differently to stresses, suggesting the regulation of stress responses in rubber tree by a variety of factors involving many signal pathways.
Author Contributions
XL designed the development of this study, carried out the experiments, and wrote the manuscript. BH participated and analyzed data in the experiment. YF carried out sequence alignment analysis. CT planned the study, formulated the question, and participated in the design of the experiments.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
This research was supported by National Natural Science Foundation of China (31300570) and the 863 Program (2013AA102605).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2016.00215
FIGURE S1 | The vector of HbG6PDH3 for subcelluar localization.
FIGURE S2 | The vector of HbG6PDH4 for subcelluar localization.
Footnotes
References
Bouteau, F., Bousquet, U., Lacrotte, R., Cornel, D., Monestiez, M., and Rona, J. P. (1992). Sucrose/H+ and glucose/H+ symports at the plasma membrane of laticiferous cells and protoplasts of Hevea brasiliensis. Indian J. Nat. Rubber Res. 5, 25–37.
Bouteau, F., Lacrotte, R., Cornel, D., Monestiez, M., Bousquet, U., Pennarun, A. M., et al. (1991). Electrogenic active proton pumps in Hevea brasiliensis laticiferous cells : its role in activating sucrose/H+ and glucose/H+ symports at the plasma membrane. Bioelectrochem. Bioenerg. 26, 223–236. doi: 10.1016/0302-4598(91)80026-Y
Cardi, M., Castiglia, D., Ferrara, M., Guerriero, G., Chiurazzi, M., and Esposito, S. (2015). The effects of salt stress cause a diversion of basal metabolism in barley roots: possible different roles for glucose-6-phosphate dehydrogenase isoforms. Plant Physiol. Biochem. 86, 44–54. doi: 10.1016/j.plaphy.2014.11.001
Cardi, M., Chibani, K., Castiglia, D., Cafasso, D., Pizzo, E., Rouhier, N., et al. (2013). Overexpression, purification and enzymatic characterization of a recombinant plastidial glucose-6-phosphate dehydrogenase from barley (Hordeum vulgare cv. Nure) roots. Plant Physiol. Biochem. 73, 266–273. doi: 10.1016/j.plaphy.2013.10.008
D’Auzac, J., Jacob, J. L., Prévot, J. C., Clément, A., Gallois, R., Crestin, H., et al. (1997). “The regulation of cis-polyisoprene production (natural rubber) from Hevea brasiliensis,” in Recent Research Developments in Plant Physiology, ed. S. G. Pandalai (Trivandrum: Research Singpost), 273–332.
Dennis, D. T., Huang, Y., and Negm, F. B. (1997). “Glycolysis, the pentose phosphate pathway and anaerobic respiration,” in Plant Metabolism, eds D. T. Dennis, D. H. Turpin, D. D. Lefebvre, and D. B. Layzell (Harlow, NY: Longman), 105–123.
Duan, C., Rio, M., Leclercq, J., Bonnot, F., Oliver, G., and Montoro, P. (2010). Gene expression pattern in response to wounding, methyl jasmonate and ethylene in the bark of Hevea brasiliensis. Tree Physiol. 30, 1349–1359. doi: 10.1093/treephys/tpq066
Gebelin, V., Leclercq, J., Hu, S., Tang, C., and Montoro, P. (2013). Regulation of MIR genes in response to abiotic stress in Hevea brasiliensis. Int. J. Mol. Sci. 14, 19587–19604. doi: 10.3390/ijms141019587
Graeve, K., von Schaewen, A., and Scheibe, R. (1994). Purification, characterization, and cDNA sequence of glucose-6-phosphate dehydrogenase from potato (Solanum tuberosum L.). Plant J. 5, 353–361. doi: 10.1111/j.1365-313X.1994.00353.x
Han, J. G., Pu, X. C., and Mao, P. S. (1998). Changes of some enzyme activities in tall fescue seed during germination under accelerated ageing. Acta Agrestia Sin. 6, 84–89.
Hao, B. Z., and Wu, J. L. (2000). Laticifer differentiation in Hevea brasiliensis:induction by exogenous jasmonic acid and linolenic acid. Ann. Bot. 85, 37–43. doi: 10.1006/anbo.1999.0995
Hauschild, R., and von Schaewen, A. (2003). Differential regulation of glucose-6-phosphate dehydrogenase isoenzyme activities in potato. Plant Physiol. 133, 47–62. doi: 10.1104/pp.103.025676
Huang, J., Zhang, H., Wang, J., and Yang, J. (2003). Molecular cloning and characterization of rice 6-phosphogluconate dehydrogenase gene that is up-regulated by salt stress. Mol. Biol. Rep. 30, 223–227. doi: 10.1023/A:1026392422995
Jacob, J. L., Serres, E., Prevot, J. C., Lacrotte, R., Vidal, A., Eschbach, J. M., et al. (1988). The development of Hevea latex diagnosis. Agritrop 12, 97–115.
Knight, J. S., Emes, M. J., and Debnam, P. M. (2001). Isolation and characterisation of a full-length genomic clone encoding a plastidic glucose 6-phosphate dehydrogenase from Nicotiana tabacum. Planta 212, 499–507. doi: 10.1007/s004250000419
Koide, H., and Oda, T. (1959). Pathological occurrence of glucose-6-phosphatase in serum in liver diseases. Clin. Chim. Acta 4, 554–561. doi: 10.1016/0009-8981(59)90165-2
Kong, W. F., Chen, J. Y., Hou, Z. X., Wen, P. F., Zhan, J. C., Pan, Q. H., et al. (2007). Activity and subcellular localization of glucose-6-phosphate dehydrogenase in peach fruits. J. Plant Physiol. 164, 934–944. doi: 10.1016/j.jplph.2006.06.001
Kruger, N. J., and von Schaewen, A. (2003). The oxidative pentose phosphate pathway: structure and organisation. Curr. Opin. Plant Biol. 6, 236–246. doi: 10.1016/S1369-5266(03)00039-6
Leopold, J. A., Zhang, Y. Y., Scribner, A. W., Stanton, R. C., and Loscalzo, J. (2003). Glucose-6-phosphate dehydrogenase overexpression decreases endothelial cell oxidant stress and increases bioavailable nitric oxide. Arterioscler. Thromb. Vasc. Biol. 23, 411–417. doi: 10.1161/01.ATV.0000056744.26901.BA
Lewinsohn, T. (1991). The geographical distribution of plant latex. Chemoecology 2, 64–68. doi: 10.1007/BF01240668
Li, D., Deng, Z., Chen, C., Xia, Z., Wu, M., He, P., et al. (2010). Identification and characterization of genes associated with tapping panel dryness from Hevea brasiliensis latex using suppression subtractive hybridization. BMC Plant Biol. 10:140. doi: 10.1186/1471-2229-10-140
Li, H., Qin, Y., Xiao, X., and Tang, C. (2011). Screening of valid reference genes for real-time RT-PCR data normalization in Hevea brasiliensis and expression validation of a sucrose transporter gene HbSUT3. Plant Sci. 181, 132–139. doi: 10.1016/j.plantsci.2011.04.014
Lin, Y., Lin, S., Guo, H., Zhang, Z., and Chen, X. (2013). Functional analysis of PsG6PDH, a cytosolic glucose-6-phosphate dehydrogenase gene from Populus suaveolens, and its contribution to cold tolerance improvement in tobacco plants. Biotechnol. Lett. 35, 1509–1518. doi: 10.1007/s10529-013-1226-2
Lin, Y. Z., Lin, S. Z., Zhang, W., Zhang, Z. Y., Guo, H., and Liu, W. F. (2005). Cloning and sequence analysis of a glucose-6-phosphate dehydrogenase gene PsG6PDH from freezing-tolerant Populus suaveolens. For. Stud. China 7, 1–6. doi: 10.1007/s11632-005-0028-6
Liu, J., Wang, X., Hu, Y., Hu, W., and Bi, Y. (2013). Glucose-6-phosphate dehydrogenase plays a pivotal role in tolerance to drought stress in soybean roots. Plant Cell Rep. 32, 415–429. doi: 10.1007/s00299-012-1374-1
Liu, S., Lan, J., Zhou, B., Qin, Y., Zhou, Y., Xiao, X., et al. (2015). HbNIN2, a cytosolic alkaline/neutral-invertase, is responsible for sucrose catabolism in rubber-producing laticifers of Hevea brasiliensis (para rubber tree). New Phytol. 206, 709–725. doi: 10.1111/nph.13257
Long, X., He, B., Wang, C., Fang, Y., Qi, J., and Tang, C. (2015). Molecular identification and characterization of the pyruvate decarboxylase gene family associated with latex regeneration and stress response in rubber tree. Plant Physiol. Biochem. 87, 35–44. doi: 10.1016/j.plaphy.2014.12.005
Manco, L., Pereira, J., Relvas, L., Rebelo, U., Crisostomo, A. I., Bento, C., et al. (2011). Chronic hemolytic anemia is associated with a new glucose-6-phosphate dehydrogenase in-frame deletion in an older woman. Blood Cells Mol. Dis. 46, 288–293. doi: 10.1016/j.bcmd.2011.02.001
Mendez, D., Linares, M., Diez, A., Puyet, A., and Bautista, J. M. (2011). Stress response and cytoskeletal proteins involved in erythrocyte membrane remodeling upon Plasmodium falciparum invasion are differentially carbonylated in G6PD A– deficiency. Free Radic. Biol. Med. 50, 1305–1313. doi: 10.1016/j.freeradbiomed.2011.02.024
Metcalfe, C. R. (1967). Distribution of latex in the plant kingdom. Econ. Bot. 21, 115–127. doi: 10.1007/BF02897859
Née, G., Zaffagnini, M., Trost, P., and Issakidis-Bourguet, E. (2009). Redox regulation of chloroplastic glucose-6-phosphate dehydrogenase: a new role for f-type thioredoxin. FEBS Lett. 583, 2827–2832. doi: 10.1016/j.febslet.2009.07.035
Nemoto, Y., and Sasakuma, T. (2000). Specific expression of glucose-6-phosphate dehydrogenase (G6PDH) gene by salt stress in wheat (Triticum aestivum L.). Plant Sci. 158, 53–60. doi: 10.1016/S0168-9452(00)00305-8
Pu, X. C., Han, J. G., and Li, M. (1994). Metabolic control of zoysiagrass seed when breaking dormancy. Grassland China 3, 20–24.
Rahman, A. Y., Usharraj, A. O., Misra, B. B., Thottathil, G. P., Jayasekaran, K., Feng, Y., et al. (2013). Draft genome sequence of the rubber tree Hevea brasiliensis. BMC Genomics 14:75. doi: 10.1186/1471-2164-14-75
Scharte, J., Schon, H., Tjaden, Z., Weis, E., and von Schaewen, A. (2009). Isoenzyme replacement of glucose-6-phosphate dehydrogenase in the cytosol improves stress tolerance in plants. Proc. Natl. Acad. Sci. U.S.A. 106, 8061–8066. doi: 10.1073/pnas.0812902106
Schnarrenberger, C., Oeser, A., and Tolbert, N. E. (1973). Two isoenzymes each of glucose-6-phosphate dehydrogenase and 6-phosphogluconate dehydrogenase in spinach leaves. Arch. Biochem. Biophys. 154, 438–448. doi: 10.1016/0003-9861(73)90077-5
Siddappaji, M. H., Scholes, D. R., Bohn, M., and Paige, K. N. (2013). Overcompensation in response to herbivory in Arabidopsis thaliana: the role of glucose-6-phosphate dehydrogenase and the oxidative pentose-phosphate pathway. Genetics 195, 589–598. doi: 10.1534/genetics.113.154351
Silpi, U., Chantuma, P., Kasemsap, P., Thaler, P., Thanisawanyangkura, S., Lacointe, A., et al. (2006). Sucrose and metabolism distribution patterns in the latices of three Hevea brasiliensis clones: effects of tapping and stimulation on the tree trunk. J. Rubber Res. 9, 115–131.
Silpi, U., Lacointe, A., Kasempsap, P., Thanysawanyangkura, S., Chantuma, P., Gohet, E., et al. (2007). Carbohydrate reserves as a competing sink: evidence from tapping rubber trees. Tree Physiol. 27, 881–889. doi: 10.1093/treephys/27.6.881
Ślaski, J. J., Zhang, G. C., Basu, U., Stephens, J. L., and Taylor, G. J. (1996). Aluminum resistance in wheat (Triticum aestivum) is associated with rapid, Al-induced changes in activities of glucose-6-phosphate dehydrogenase and 6-phosphogluconate dehydrogenase in root apices. Physiol. Plant. 98, 477–484. doi: 10.1034/j.1399-3054.1996.980307.x
Sookmark, U., Pujade-Renaud, V., Chrestin, H., Lacote, R., Naiyanetr, C., Seguin, M., et al. (2002). Characterization of polypeptides accumulated in the latex cytosol of rubber trees affected by the tapping panel dryness syndrome. Plant Cell Physiol. 43, 1323–1333. doi: 10.1093/pcp/pcf161
Tang, C., Huang, D., Yang, J., Liu, S., Sakr, S., Li, H., et al. (2010). The sucrose transporter HbSUT3 plays an active role in sucrose loading to laticifer and rubber productivity in exploited trees of Hevea brasiliensis (para rubber tree). Plant Cell Environ. 33, 1708–1720. doi: 10.1111/j.1365-3040.2010.02175.x
Tang, C., Qi, J., Li, H., Zhang, C., and Wang, Y. (2007). A convenient and efficient protocol for isolating high-quality RNA from latex of Hevea brasiliensis (para rubber tree). J. Biochem. Biophys. Methods 70, 749–754.
Thom, E., Möhlmann, T., and Quick, W. P. (1998). Sweet pepper plastids: enzymic equipment, characterisation of the plastidic oxidative pentose-phosphate pathway, and transport of phosphorylated intermediates across the envelope membrane. Planta 204, 226–233. doi: 10.1007/s004250050251
Tupy, J. (1973). The level and distribution pattern of fatex sucrose along the trunk of Hevea brasiliensis Mull. arg. as affected by the sink region induced by latex tapping. Physiol. Veg. 11, 1–11.
Vandesompele, J., De Preter, K., Pattyn, F., Poppe, B., Van Roy, N., De Paepe, A., et al. (2002). Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, RESEARCH0034. doi: 10.1186/gb-2002-3-7-research0034
Venkatachalam, P., Thulaseedharan, A., and Raghothama, K. (2009). Molecular identification and characterization of a gene associated with the onset of tapping panel dryness (TPD) syndrome in rubber tree (Hevea brasiliensis Muell.) by mRNA differential display. Mol. Biotechnol. 41, 42–52. doi: 10.1007/s12033-008-9095-y
von Schaewen, A., Langenkamper, G., Graeve, K., Wenderoth, I., and Scheibe, R. (1995). Molecular characterization of the plastidic glucose-6-phosphate dehydrogenase from potato in comparison to its cytosolic counterpart. Plant Physiol. 109, 1327–1335. doi: 10.1104/pp.109.4.1327
Wakao, S., Andre, C., and Benning, C. (2008). Functional analyses of cytosolic glucose-6-phosphate dehydrogenases and their contribution to seed oil accumulation in Arabidopsis. Plant Physiol. 146, 277–288. doi: 10.1104/pp.107.108423
Wakao, S., and Benning, C. (2005). Genome-wide analysis of glucose-6-phosphate dehydrogenases in Arabidopsis. Plant J. 41, 243–256. doi: 10.1111/j.1365-313X.2004.02293.x
Wang, X., Ma, Y., Huang, C., Li, J., Wan, Q., and Bi, Y. (2008). Involvement of glucose-6-phosphate dehydrogenase in reduced glutathione maintenance and hydrogen peroxide signal under salt stress. Plant Signal. Behav. 3, 394–395. doi: 10.4161/psb.3.6.5404
Wendt, U. K., Wenderoth, I., Tegeler, A., and Von Schaewen, A. (2000). Molecular characterization of a novel glucose-6-phosphate dehydrogenase from potato (Solanum tuberosum L.). Plant J. 23, 723–733. doi: 10.1046/j.1365-313x.2000.00840.x
Xiao, X., Tang, C., Fang, Y., Yang, M., Zhou, B., Qi, J., et al. (2014). Structure and expression profile of the sucrose synthase gene family in the rubber tree: indicative of roles in stress response and sucrose utilization in the laticifers. FEBS J. 281, 291–305. doi: 10.1111/febs.12595
Yang, Y., Fu, Z., Su, Y., Zhang, X., Li, G., Guo, J., et al. (2014). A cytosolic glucose-6-phosphate dehydrogenase gene, ScG6PDH, plays a positive role in response to various abiotic stresses in sugarcane. Sci. Rep. 4, 7090. doi: 10.1038/srep07090
Yu, H., Jiang, W., Liu, Q., Zhang, H., Piao, M., Chen, Z., et al. (2015). Expression pattern and subcellular localization of the ovate protein family in rice. PLoS ONE 10:e0118966. doi: 10.1371/journal.pone.0118966
Keywords: glucose-6-phosphate dehydrogenase, expression, latex regeneration, abiotic stresses, Hevea brasiliensis
Citation: Long X, He B, Fang Y and Tang C (2016) Identification and Characterization of the Glucose-6-Phosphate Dehydrogenase Gene Family in the Para Rubber Tree, Hevea brasiliensis. Front. Plant Sci. 7:215. doi: 10.3389/fpls.2016.00215
Received: 14 November 2015; Accepted: 08 February 2016;
Published: 25 February 2016.
Edited by:
David John Burritt, University of Otago, New ZealandReviewed by:
Agata Cieśla, Adam Mickiewicz University, PolandUpinder S. Gill, The Samuel Roberts Noble Foundation, USA
Copyright © 2016 Long, He, Fang and Tang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Chaorong Tang, Y2hhb3Jvbmd0YW5nQDEyNi5jb20=
 Xiangyu Long
Xiangyu Long Bin He
Bin He