- 1Biotechnology Center in Southern Taiwan of Agricultural Biotechnology Research Center, Academia Sinica, Tainan, Taiwan
- 2Institute of Tropical Plant Science, National Cheng Kung University, Tainan, Taiwan
Oryza sativa BRASSINOSTEROID UPREGULATED1 LIKE1 (OsBUL1) positively affects lamina inclination and grain size. OsBUL1 knock-out (osbul1) plants as well as transgenic rice with reduced level of OsBUL1 expression produce erect leaves and small grains. Here, we identified a putative downstream gene of OsBUL1, OsBUL1 DOWNSTREAM GENE1 (OsBDG1) encoding a small protein with short leucine-rich-repeats by cDNA microarray analyses in the lamina joint and panicles of wild-type and osbul1 plants. Transgenic rice plants with increased OsBDG1 expression exhibit increased leaf angle and grain size, which is similar to an OsBDG1 activation tagging line whereas double stranded RNA interference (dsRNAi) lines for OsBDG1 knock-down generate erect leaves with smaller grains. Moreover, transgenic rice expressing OsBDG1 under the control of OsBUL1 promoter also shows enlarged leaf bending and grain size phenotypes. Two genes, OsAP2 (OsAPETALA2) and OsWRKY24 were identified as being upregulated transcriptional activators in the lamina joint of pOsBUL1:OsBDG1 plants and induced expression of the two genes driven by OsBUL1 promoter caused increased lamina inclination and grain size in rice. Thus, our work demonstrates that a series of genes showing expression cascades are involved in the promotion of cell elongation in lamina joints and functionally cause increased lamina inclination.
Introduction
Crops are generally grown at high planting density in the field. For better yield and resistance against lodging, cereal crops, such as rice, with semi-dwarf and/or erect leaf phenotypes are strongly desired (Van Camp, 2005; Sakamoto et al., 2006). A rice leaf is composed of a leaf sheath, leaf blade, and a lamina joint (area between leaf blade and sheath, also called the collar) with auricles and ligule (Hoshikawa, 1989). In particular, leaf angle (the degree of bending between the leaf blade and leaf sheath) is one of the key agronomic traits that affects crop architecture and grain yields (Sinclair and Sheehy, 1999). Crops with erect leaves capture more light for photosynthesis and are also desirable for dense planting, all of which increase yields (Sakamoto et al., 2006).
Previously, it was shown that both phytohormone and non-hormone-related genes are involved in controlling the lamina inclination.
The brassinosteroid (BR)-deficient mutant dwarf4-1, ebisu dwarf (d2), and dwarf1 (brd1), the BR signaling mutant d61-7, and plants with suppressed expression of OsBRASSINAZOLE-RESISTANT 1 (OsBZR1) encoding a transcription factor involved in the BR signaling cascade display erect leaves (Hong et al., 2002, 2003; Sakamoto et al., 2006; Bai et al., 2007) while overexpression of BR biosynthesis genes or signaling components resulted in large leaf inclination (Yamamuro et al., 2000; Bai et al., 2007). For example, transgenic rice plants overexpressing sterol C-22 hydroxylase (a rate-limiting enzyme in BR biosynthesis) showed increased lamina angles (Wu et al., 2008). In addition to BR, other phytohormones are also involved in controlling the lamina inclination of rice. Ethylene participates in the response of BR-induced lamina joint inclination, and auxin (IAA) influences the lamina joint inclination at high concentrations and has a synergistic interaction with BR (Cao and Chen, 1995; Nakamura et al., 2009). Reduced expression of SPINDLY, a negative regulator of gibberellin (GA) signaling, also causes increased lamina inclination (Shimada et al., 2006). In addition, transgenic rice plants with overexpression of LAX PANICLE (LAX) (Komatsu et al., 2003), OsILI-BINDING HLH1 (OsIBH1) (Zhang et al., 2009) and T-DNA insertion mutants of OsWRKY11 (Wang et al., 2005), OsLIGULELESS1 (OsLG1) encoding a SQUAMOSA promoter binding domain protein (Lee et al., 2007) exhibited erect leaves, whereas the double stranded RNA interference (dsRNAi) transgenic lines for rice MADS-box genes belonging to the SHORT VEGETATIVE PHASE (SVP) group such as OsMADS22, OsMADS47, and OsMADS55 showed increased lamina angles (Lee et al., 2008). Decreased expression of OsLIC encoding a CCCH-type zinc-finger protein also results in increased lamina inclination through regulating BR signaling (Wang et al., 2008). It was also reported that LC2 encoding a VERNALIZATION INSENSITIVE 3 (VIN3)-like protein acts as a repressor of cell division for regulation of collar development and the lc2 mutation caused increased leaf angles (Zhao et al., 2010). A recent report showed that an activation tagging line of rice, slender grain Dominant (slg-D), with enhanced expression of a gene encoding BAHD acyltransferase-like protein, produced slender grains with enlarged leaf angles (Feng et al., 2016), and a gain-of-function epiallele of rice RELATED TO ABSCISIC ACID INSENSITIVE3 (ABI3)/VIVIPAROUS1 (VP1) 6 (RAV6) encoding a B3 DNA-binding domain-containing protein caused larger lamina inclination but smaller grain size by modulating BR homeostasis (Zhang et al., 2015). Moreover, induced expression of genes encoding atypical Helix-Loop-Helix (HLH) proteins such as BRASSINISTEROID UPREGULATED1 (BU1; Tanaka et al., 2009), Oryza sativa BU1 like 1 (OsBUL1; Jang et al., 2017), INCREASED LAMININAR INCLINATION (ILI; Zhang et al., 2009) and POSITIVE REGULATOR OF GRAIN LENGTH 1 (PGL1; Heang and Sassa, 2012) or basic HLH (bHLH) proteins including OsBUL1 Complex1 (OsBC1; Jang et al., 2017) conferred rice plants higher lamina angle degree with increased grain size.
In this work, transcriptomes in collars and panicles of OsBUL1 null mutants and WT plants were analyzed and OsBUL1 DOWNSTREAM GENE1 (OsBDG1) was identified as a putative downstream gene of OsBUL1. It encodes a small leucine rich repeat (LRR) protein possessing cell elongation activity. Sequentially, OsAP2 and OsWRKY24 are identified as putative downstream genes of OsBDG1, and functional activities are assessed with respect to rice lamina inclination and grain size. Both genes are able to increase lamina inclination and grain size under the control of OsBUL1 promoter. We, therefore, provide a sequential flow of acting genes from OsBUL1 for positive effects on rice lamina joint inclination and grain size likely through cell elongation.
Materials and Methods
Plant Materials and Growth Conditions
The activation tagging line of OsBDG1, PFG_1B_05536 is in Dongjin background (Jeong et al., 2006). Transgenic rice plants were produced with Tainung67 (TNG67) japonica rice cultivar. Rice plants (Oryza sativa) were grown in the field under natural long days or in the greenhouse with 28°C day/25°C night cycles. To assess the leaf angles of mature rice plants in the paddy field, leaves were numbered from top to bottom. Lamina angles were measured between a stem and a leaf blade at the second or the third leaf from eight to twelve individual plants per line when panicles come out of flag-leaf sheaths. Transgenic plants used for analyses in this work are all T3 or T4 independent lines.
Lamina Joint Inclination Bioassay
Sterilized seeds were germinated and grown for 10 days in a dark chamber. Lamina joint inclination bioassays were performed as previously described (Jeong et al., 2007). Seedlings were sampled by excising approximately 2 cm segments that contained lamina joints at the same position from each plant under dim light condition. They were floated on distilled water containing various concentrations of BL. After incubation in a dark chamber at 28°C for 2 days, the angle induced between the lamina and the sheath was measured.
Vector Construction and Transformation
Each open reading frame (ORF) of OsBDG1, OsAP2, and OsWRKY24 was cloned into pGA3426 (Kim et al., 2009) or its derivatives for overexpression and/or dsRNAi purposes in rice. For expression by OsBUL1 promoter, the ubiquitin promoter of pGA3426 was replaced with the 2.2 kb-OsBUL1 promoter (Jang et al., 2017). Constructed plasmids were individually transformed into embryonic calli of TNG67 rice cultivars by Agrobacterium-LBA4404 mediation as described previously (Jeon et al., 2000).
Hormone Treatment
Ten-day-old rice (O. sativa cv. TNG67) seedlings grown in the growth chamber were treated with brassinolide (1 μM, BL from Sigma–Aldrich) or GA (100 μM GA3 from Sigma–Aldrich). Whole parts above the roots were harvested for RNA extraction at 24 h time point after treatment.
Total RNA Isolation and Quantitative RT-PCR Analysis
Total RNAs of all the materials harvested were isolated using RNeasy plant mini kit (Qiagen) or Trizol solution (Invitrogen) according to the manufacturer’s instructions. RNAs after DNase treatment were subjected to reverse transcriptase reactions using oligo(dT) primer and Superscript III reverse transcriptase (Invitrogen) according to the manufacturer’s protocol. Subsequent PCR was conducted with the first-strand cDNA mixture and EX-Taq polymerase (Takara Bio). Quantitative PCR (qPCR) was carried out by a CFX96TM real-time system (Bio-Rad) using Maxima SYBR Green qPCR Master Mix (Thermo). For PCR, each sample was analyzed in triplicate. The run protocol was: denaturation at 95°C for 10 min and annealing/extension repeated 45 times (95°C for 15 s and 60°C for 30 s, data acquisition was performed). Housekeeping genes such as OsUBQ (Komiya et al., 2008) and OsAct (Caldana et al., 2007) were included in the reactions as internal controls for normalizing the variations in the amount of cDNA used (Guénin et al., 2009). The threshold cycle (CT) was automatically determined for each reaction by the system set with default parameters. The specificity of the qRT-PCR was determined by curve analysis of the amplified products using the standard method installed in the system. Information on primers used is presented in Supplementary Table S1.
Microarray Analyses
Collars of 80-day-old rice plants and young panicles less than 10 cm in length were harvested for RNA extraction for microarray. We used the Rice Whole Genome OneArray v1.1 (Phalanx Biotech Group, Taiwan) containing 22,003 DNA oligonucleotide probes. Each probe is a 60-mer designed in the sense direction. Among the probes, 21,179 probes corresponded to the annotated genes in the RGAP v.6.1 and BGI database. Additionally, we included 824 control probes. The detailed descriptions of the gene array list are available from http://www.phalanx.com.tw/products/RiOA_Probe.php. Fluorescent antisense RNA targets were prepared from 1 μg total RNA samples with the OneArray Amino Allyl aRNA Amplification Kit (Phalanx Biotech Group, Taiwan) and Cy5 dyes (Amersham Pharmacia, United States). Fluorescent targets were hybridized to the Rice OneArray with Phalanx hybridization buffer using Phalanx Hybridization System. After hybridization for 16 h at 50°C, non-specific binding targets were washed away by three washing steps (Washing I, 42°C 5 min; Washing II, 42°C, 5 min; 25°C, 5 min; Washing III, rinse 20 times), and the slides were dried by centrifugation and scanned by an Agilent G2505C scanner (Agilent Technologies, United States). The Cy5 fluorescent intensities of each spot were analyzed by GenePix 4.1 software (Molecular Devices). The signal intensity of each spot was introduced into Rosetta Resolver System (Rosetta Bio-software) to process data analysis. The error model of the Rosetta Resolver System could remove both systematic and random errors from the data. We filtered out the spots whose flag was less than 0. Spots that passed the criteria were normalized by 50% media scaling normalization method. The technical repeat data was tested by Pearson correlation coefficient calculation to check the reproducibility (R-value > 0.975). Normalized spot intensities were transformed to gene expression log2 ratios between the control and treatment groups. The interest spots which show significant differences were selected by log2 ratio ≥ 1 or log2 ratio ≤-1 and P < 0.05. Two independent biological replicates of hybridizations were performed.
Histological Analyses
Lamina joint samples were fixed by 4% paraformaldehyde in 0.1 M sodium phosphate buffer, dehydrated through a series of graded ethanol baths, replaced with xylene, and embedded in Paraplast plus (Sigma–Aldrich). Paraffin sections (12 μm) were cut and stained with filtered 1% toluidine blue. The sections were photographed under a light microscope (Olympus BX51).
Subcellular Localization of Proteins
For cellular localization of OsBC1, OsBDG1, OsBUL1, OsAP2.2, and OsWRKY24 in rice, either yellow florescence protein (YFP):gateway (GW) or cyan fluorescent protein (CFP):GW vector was used for the florescence fusion as described previously (Jang et al., 2015). Isolation and transfection of rice protoplasts were followed as described by Zhang et al. (2011) and images of cells with fluorescence were taken by confocal microscopy (LSM 510 META NLO DuoScan, Carl Zeiss).
Scanning Electron Microscopy
To observe epidermal cells of lemma in rice grains by scanning electron microscopy (SEM), whole grains were coated with gold by the ion sputter machine (E-1010, Hitachi) and examined with SEM (Quanta 250, FEI, Hillsboro, OR, United States).
Results
OsBDG1 May Act Downstream of OsBUL1
To identify the downstream genes of OsBUL1 in the expressional hierarchy, comparison of transcriptomes of collars and panicles between WT and osbul1 plants was conducted using rice 22k-oligo microarrays1. The osbul1 is a mRNA null mutant caused by a T-DNA insertion in the first exon of OsBUL1 gene (Jang et al., 2017).
Fourteen genes were upregulated, while 221 genes were downregulated in both collars and panicles of osbul1 plants (Figure 1A). Among candidate genes, OsBUL1 DOWNSTREAM GENE (OsBDG1) was selected for further study since expression level of OsBDG1 was reduced both in collars and panicles of osbul1 plants (Figure 1B), and OsBDG1 encodes a novel small protein containing a conserved LRR N-terminal domain (LRRNT) at the N-terminal part and three leucin-rich-repeat (LRR) domains toward its carboxyl terminus. In addition, its expression is known to be induced in transgenic rice plants overexpressing Arabidopsis CYP90B1 which encodes for a sterol C-22 hydroxylase active in BR biosynthesis (Wu et al., 2008). To verify the microarray results, we performed expressional analyses on OsBDG1 in collars and panicles of osbul1 and WT plants, and found similar expression patterns to the microarray results (Figure 1C). On the contrary, the level of OsBDG1 expression was higher in OsBUL1-overexpressing rice plants compared to the WT control (Figure 1D) indicating OsBUL1 may act upstream of OsBDG1. Transgenic rice plants harboring pUbi:OsBDG1 show increased lamina inclination with elongated cells in the lamina joint and grain size (Figures 2A–C) whereas reduced expression of OsBDG1 results in erect leaves with reduced cell length in lamina joint and smaller grains (Figures 2D–F). Phenotypes of overexpressors and dsRNAi lines for OsBDG1 are similar to those of rice plants with increased OsBUL1 expression and osbul1, respectively, supporting the expressional relationship between OsBUL1 and OsBDG1.
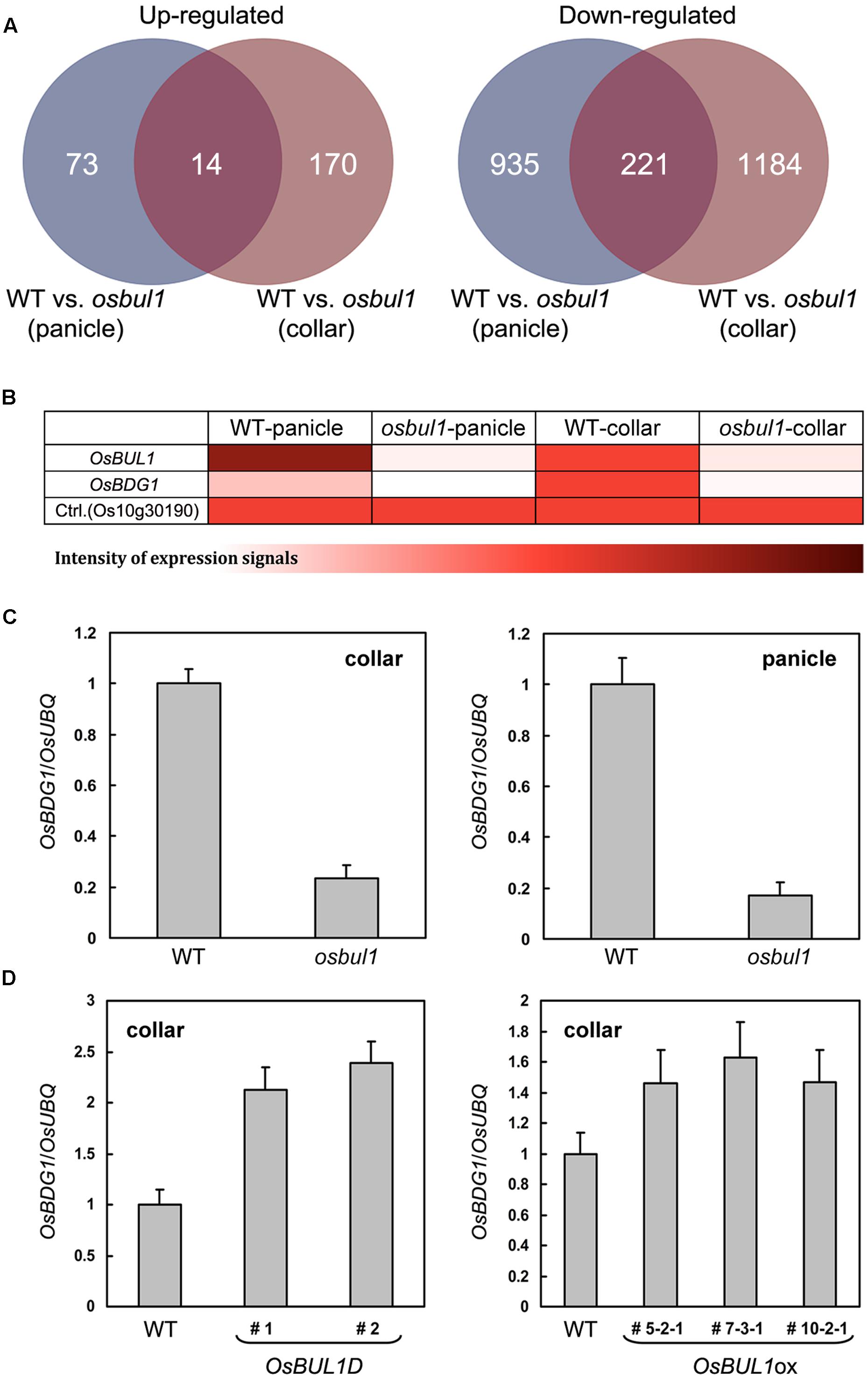
FIGURE 1. OsBDG1 is a putative downstream gene of OsBUL1 in expression. (A) Venn diagrams for upregulated and downregulated genes in the panicle and collar of osbul1. (B) A heat map based on microarray hybridization results shows that OsBDG1 expression is reduced in osbul1. Color scale represents log signal values and Os10g30190 is a control showing similar signal values in the microarray hybridization between WT and osbul1. (C) Quantitative RT-PCR confirmed the reduced expression of OsBDG1 in osbul1 in collars and panicles. Conversely, expression level of OsBDG1 is higher in an OsBUL1 activation tagging line, OsBUL1D (Jang et al., 2017) and OsBUL1 overexpressors (ox) than in the WT (D). Data are the average of three or four independent experiments and normalized by OsUBQ. Error bars indicate SD.
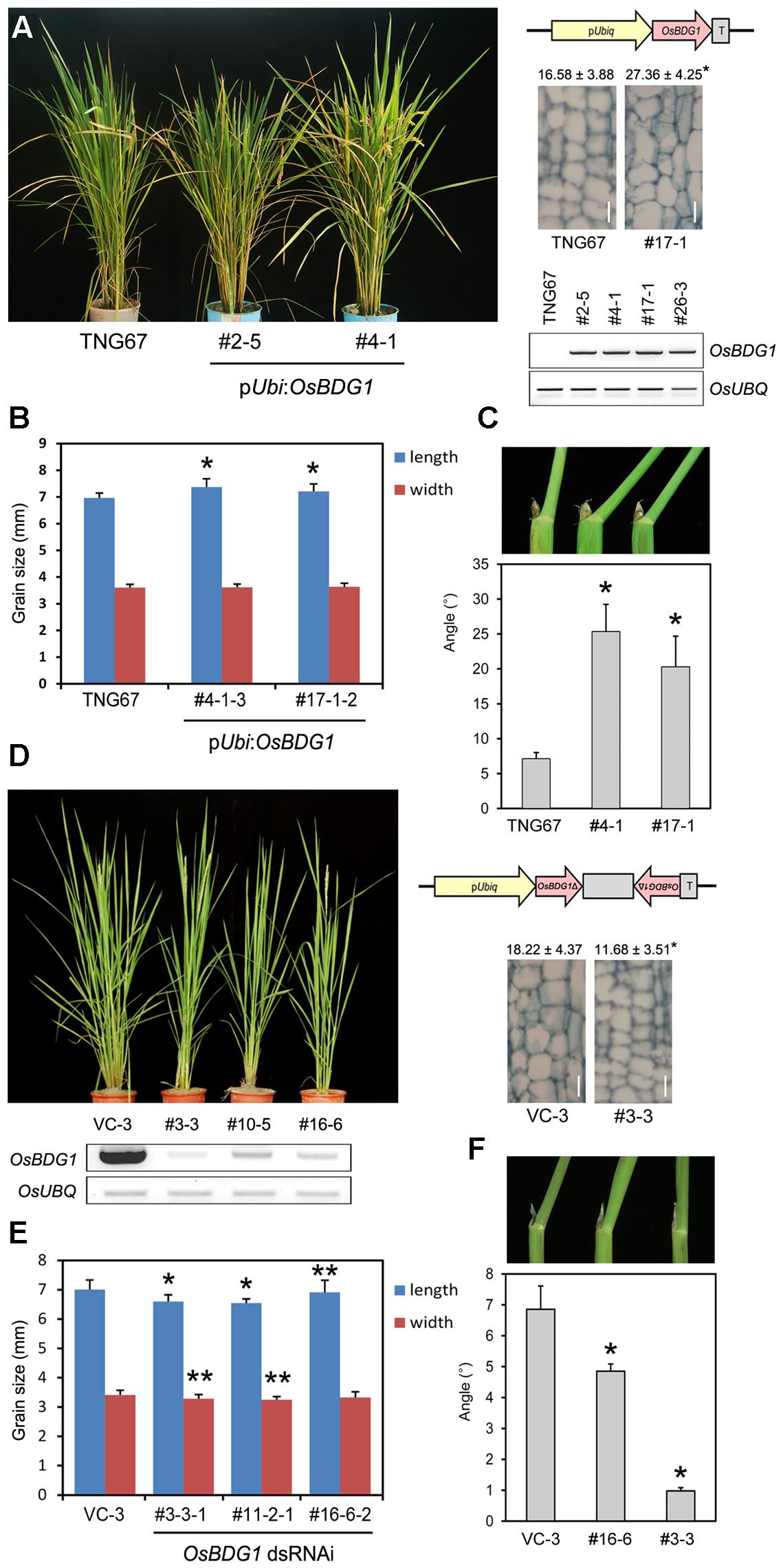
FIGURE 2. Transgenic rice plants with pUbi:OsBDG1 and OsBDG1-dsRNAi constructs. (A–C) Overexpression of OsBDG1 driven by ubiquitin promoter caused increase of leaf angles with cell length in the lamina joint and grain size. Longitudinal sections of the second leaf lamina joint are shown in (A) with measured cell length. Values are means ± SD (μm, n > 20) as instructed by Zhang et al. (2009). (∗P < 0.01, Student’s t-test). Bar = 20 μm. Semi-quantitative RT-PCR was conducted for OsBDG1 expression in transgenic rice plants together with WT. Twenty-four PCR cycles for both OsBDG1 and OsUBQ were applied. Values of grain size and leaf angles are presented as means ± SD in (B) (n = 35; ∗P < 0.01, Student’s t-test) and (C) (n > 10; ∗P < 0.01, Student’s t-test), respectively. (D–F) Reduced expression of OsBDG1 by dsRNAi-OsBDG1 approaches resulted in erect leaves with shorter cells in lamina joint and smaller grains. The 384 bp-fragment of OsBDG1 amplified by primers 5′ ATGGGGGCTCATTCTGCAGCGGCAGCTC 3′ and 5′ GTGCCACTCAGTGAATTCTTCTGAAGCTC 3′ was used for the dsRNAi construct. Semi-quantitative RT-PCR was conducted for OsBDG1 expression in transgenic rice plants together with vector control (VC). Thirty-five and 24 PCR cycles were used for OsBDG1 and OsUBQ, respectively. Longitudinal sections of the second leaf lamina joint are shown in (D) with measured cell length (μm, n > 20; ∗P < 0.01, Student’s t-test). Bar = 20 μm. Values of grain size and leaf angles are presented in (E) (n > 25; ∗P < 0.01; ∗∗P < 0.05, Student’s t-test) and (F) (n > 8; ∗P < 0.01, Student’s t-test), respectively.
Identification of an Activation Tagging Line for OsBDG1
A putative activation tagging line for OsBDG1, PFG_1B_05536 has been identified through the rice T-DNA database2 (An et al., 2003). Phenotypic analysis of the heterozygous population displayed a 3:1 ratio of OsBDG1D mutant phenotype to wild-type indicating OsBDG1D is a dominant mutant. T-DNA flanking sequence of the mutant was confirmed by PCR and comparison of the sequence through blastn search showed that the T-DNA was inserted in the 4.1 Kb downstream of Os11g31530 near Os11g31520 on chromosome 11 (Figure 3A). The expression of Os11g31530 was obviously influenced whereas the expression level of other genes located near the insertion position was not significantly affected (Figure 3B). The lamina angle increased more than two-fold in the activation tagging line with an increase of grain size (Figures 3C–E), which is similar to OsBDG1 overexpressors.
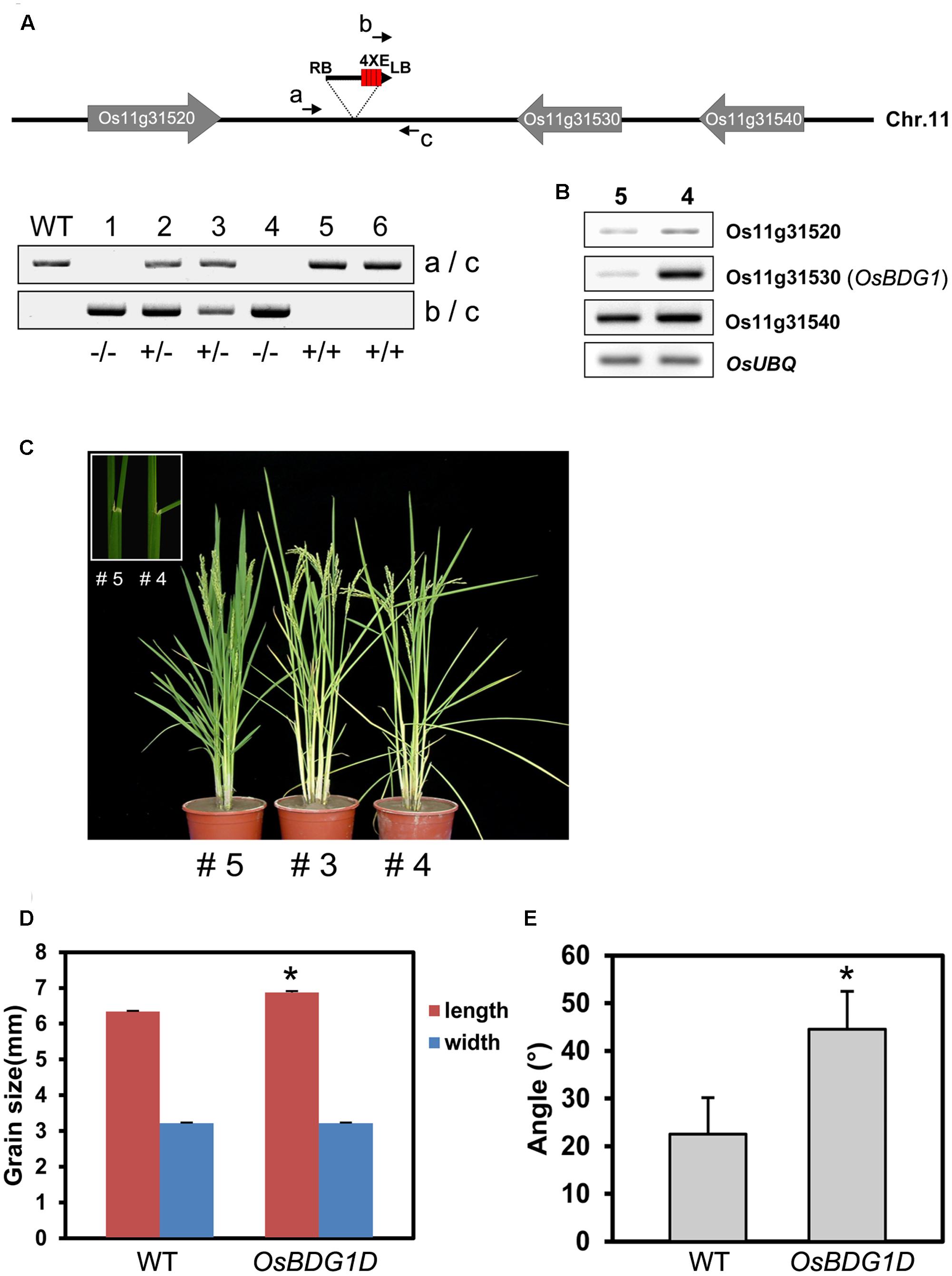
FIGURE 3. Identification and characterization of an OsBDG1 activation tagging line. (A) T-DNA is inserted in the intergenic region between Os11g31520 and Os11g31530. Genotyping was conducted by primers, (a) 5′ CATAGGAACAGAAGGAGTAC 3′, (b) 5′ GACGAGAGTGTCGTGCTCCACCATG 3′ and (c) 5′ GAGGAGATTGTGGGCTCATG 3′. (B) Expression of genes located near the T-DNA insertion. (C) A gain-of-function line of OsBDG1 showed increased lamina inclination. Lamina joint area of a WT segregant (#5; left) and a homozygous OsBDG1D line (#4) is shown in the box. (D) OsBDG1D plants produced grains with increased size and (E) caused increased leaf angles (the third leaf from the top of main stems). Error bars indicate SD. (∗P < 0.01, Student’s t-test).
Expression of OsBDG1 under the Control of OsBUL1 Promoter
To confirm the expression cascade of OsBUL1 and OsBDG1, we made a construct for expression of OsBDG1 under the control of the 2.2 kb-OsBUL1 promoter (Figure 4A) preferentially active in lamina joints and panicles of rice (Jang et al., 2017). Transgenic rice containing the construct exhibited a dramatic increase in lamina inclination and grain size (Figures 4B–E), which was similar to the lamina inclination and grain phenotypes of the OsBDG1 activation tagging line, PFG_1B_05536 and OsBDG1-overexpressing lines driven by ubiquitin promoter. Thus, expression of OsBDG1 under the control of OsBUL1 promoter phenocopies OsBUL1 overexpression by ubiquitin promoter or activation tagging systems (Jang et al., 2017) suggesting that OsBDG1 may act downstream of OsBUL1.
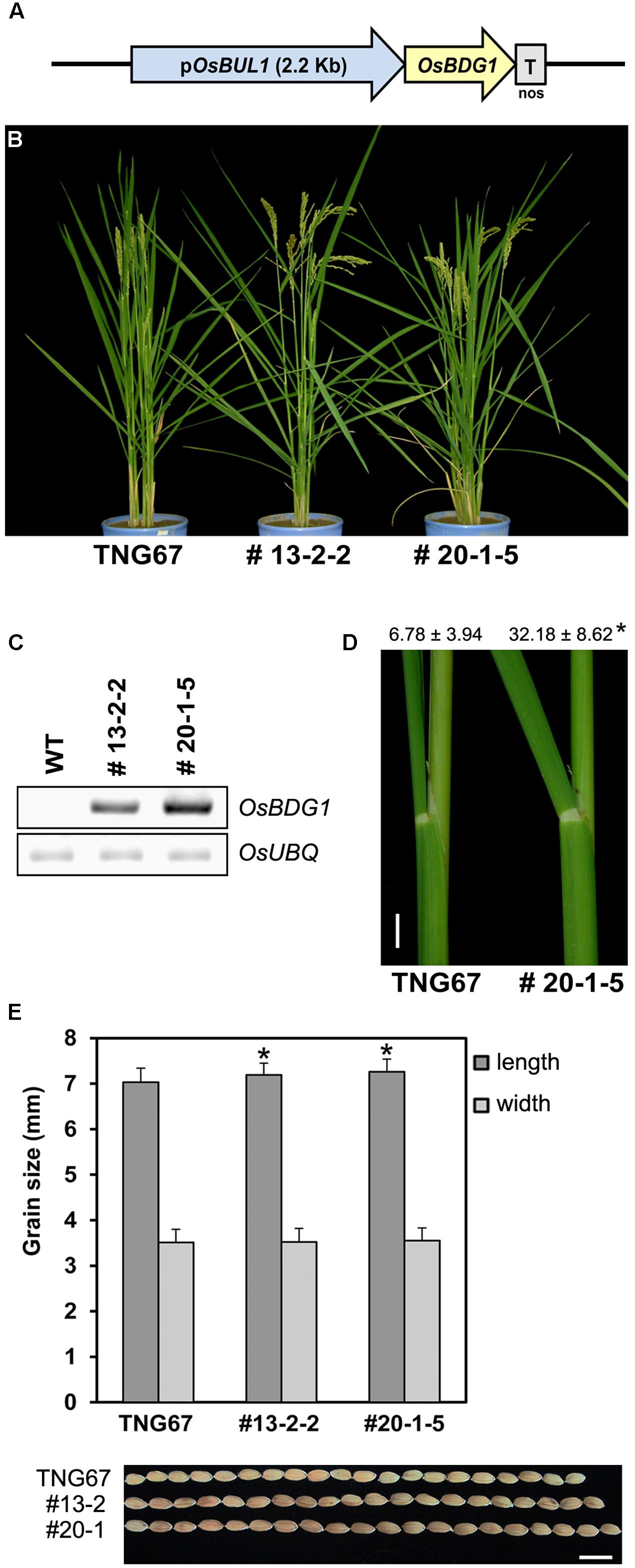
FIGURE 4. Phenotypic alterations of pOsBUL1:OsBDG1 transgenic rice plants. (A) A simplified structure of pOsBUL1:OsBDG1 construct. (B) Transgenic lines with pOsBUL1:OsBDG1 construct exhibited conspicuous increase of leaf angles. (C) Expression of OsBDG1 in collar between WT and transgenic plants. Semi-quantitative RT-PCR was conducted for OsBDG1 expression in transgenic rice plants together with WT. Twenty-six PCR cycles for both OsBDG1 and OsUBQ were applied. (D) Degree of leaf angles between WT and transgenic line, #20-1-5 at the second leaf from the top of the main stem. Values are given as means ± SD (degree; n > 14). (∗P < 0.01, Student’s t-test). Bar = 1 cm. (E) Transgenic lines produced grains with increased size. (∗P < 0.01, Student’s t-test). Bar = 1 cm.
OsAP2 and OsWRKY24 Are Upregulated by OsBDG1 in Lamina Joints
To identify candidate genes affected by OsBDG1 in the lamina joint and also likely responsible for the increased lamina inclination, we compared transcriptomes of collars among two independent homozygous transgenic lines of pOsBUL1:OsBDG1, #13-2-2 and #20-1-5, and wild-type plants by microarray hybridization3. Since the expression level of OsBDG1 in the collar of a #13-2-2 plant is lower than that of a #20-1-5 plant (Figure 4C), we selected genes that showed the same increasing or decreasing patterns in the order of WT, #13-2-2 and #20-1-5. Five genes were identified as being upregulated by OsBDG1 in the lamina joint (Figures 5A,B) but no candidate was found to be downregulated in the comparison. Also, no significant difference in OsBUL1 expression was detected among WT, #13-2-2 and #20-1-5 lines (GEO accession no. GSE93817 for microarray results). We confirmed the expression patterns of each gene by qRT-PCR (Figure 5C). The Os10g41330 (OsAP2) produced two transcripts, Os10g41330.1 and Os10g41330.2 by alternative splicing (Supplementary Figure S1) and both transcripts showed similar accumulation patterns. Based on the increased expression level of each gene in the transgenic plants, we focused on two genes, Os10g41330 (OsAP2) and Os01g61080 (OsWRKY24) encoding putative transcription factors containing an AP2 domain and WRKY domain, respectively. Conversely, in collars of OsBDG1 dsRNAi lines, the expression of OsAP2.2 and OsWRKY24 is reduced without altered expression level of OsBUL1 supporting the notion that the two genes are downstream of OsBDG1 in expression (Supplementary Figure S2). However, the expression of OsBUL1 and OsBDG1 was positively affected by pOsBUL1:OsAP2 and/or pOsBUL1:OsWRKY24 (Supplementary Figure S3).
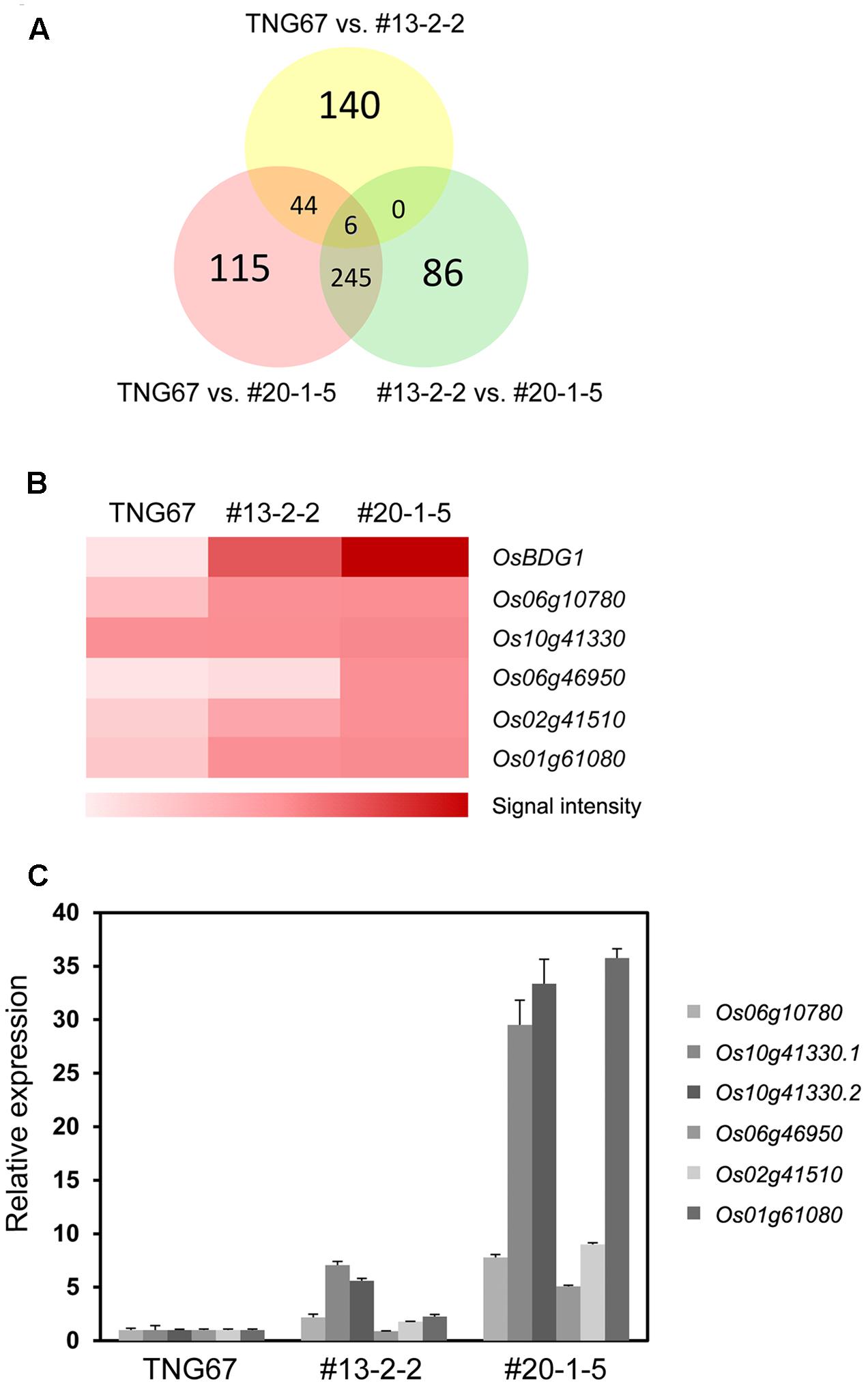
FIGURE 5. Identification of downstream genes of OsBDG1 acting in the lamina joint. (A) A Venn diagram showing numbers of genes upregulated by OsBDG1 in each comparison. Two independent T3 transgenic lines for pOsBUL1:OsBDG1, #13-2-2 and #20-1-5 were selected as mild and strong expressers of OsBDG1 in collars, respectively, as shown in Figure 4C. (B) A heat map generated by microarray hybridization shows expression level of the five candidate genes identified as being gradually increased by OsBDG1 expression. Color scale represents log signal values. (C) Verification of the expression level on each candidate gene in the collar of WT and transgenic lines, #13-2-2 and #20-1-5 by quantitative RT-PCR. Os10g41330 is known to produce two putative transcripts, Os10g41330.1 and Os10g41330.2 by alternative splicing (Supplementary Figure S1). Data are the average of three or four independent experiments and normalized by OsUBQ. Error bars indicate SD.
Molecular Characterization of OsBDG1, OsAP2, and OsWRKY24
Spatiotemporal expression of OsBDG1, OsAP2, and OsWRKY24 also overlapped in collars and growing panicles (Supplementary Figure S4A). Interestingly, the expression of the three genes was upregulated by phytohormones such as GA3 and BL, which affect cell elongation (Supplementary Figure S4B). Indeed, OsBDG1 dsRNAi lines exhibited reduced, but transgenic rice containing pOsBUL1:OsBDG1 showed increased sensitivity in BR response through lamina bending assays (Supplementary Figure S4C).
The OsBDG1 protein containing a short LRR motif was localized in the cytoplasm as well as the nucleus similar to OsBUL1 (Jang et al., 2017; Supplementary Figure S5). However, OsAP2.2 and OsWRKY24 proteins are localized in the nucleus and each protein shows a transcriptional activation activity in the yeast system (Supplementary Figures S5, S6C).
Increased Lamina Inclination and Grain Size Phenotypes Are Observed in pOsBUL1:OsAP2 and pOsBUL1:OsWRKY24 Plants
Transgenic rice plants containing pOsBUL1:OsAP2.2, a short transcript of Os10g41330 and pOsBUL1:genomic OsAP2 were generated for phenotypic analyses of lamina angles and grain size (Figure 6 and Supplementary Figure S1). Compared with the wild-type control, transgenic rice plants showed increased lamina inclination with increased amounts of OsAP2.2 transcripts (Figures 6A,B). However, we could not detect the long form of the transcript, OsAP2.1 in the transgenic plants containing pOsBUL1:genomic OsAP2. Transgenic rice for pOsBUL1:OsWRKY24 also exhibited a significant increase in lamina angles (Figure 6C) indicating both OsAP2.2 and OsWRKY24 are likely affected by OsBDG1 in the lamina joint. Of note, reduced expression level of both OsAP2.2 and OsWRKY24 was observed in panicles and lamina joints of osbul1 plants (Supplementary Figure S2). Additionally, panicle morphology of transgenic rice was affected by the two transgenes: panicle branches were spread and grain size was also increased (Supplementary Figure S3C). Elongated epidermal cells of lemma were also observed with higher expression level of genes involved in cell elongation such as OsExpansin (OsEXP) genes, OsXyloglucan endotransglucosylase/hydrolase1 (OsXTH1) and Osxyloglucan endotransglycosylase related1 (OsXTR1) in spikelets of transgenic rice plants (Supplementary Figure S6; Cho and Kende, 1997; Uozu et al., 2000; Hara et al., 2014).
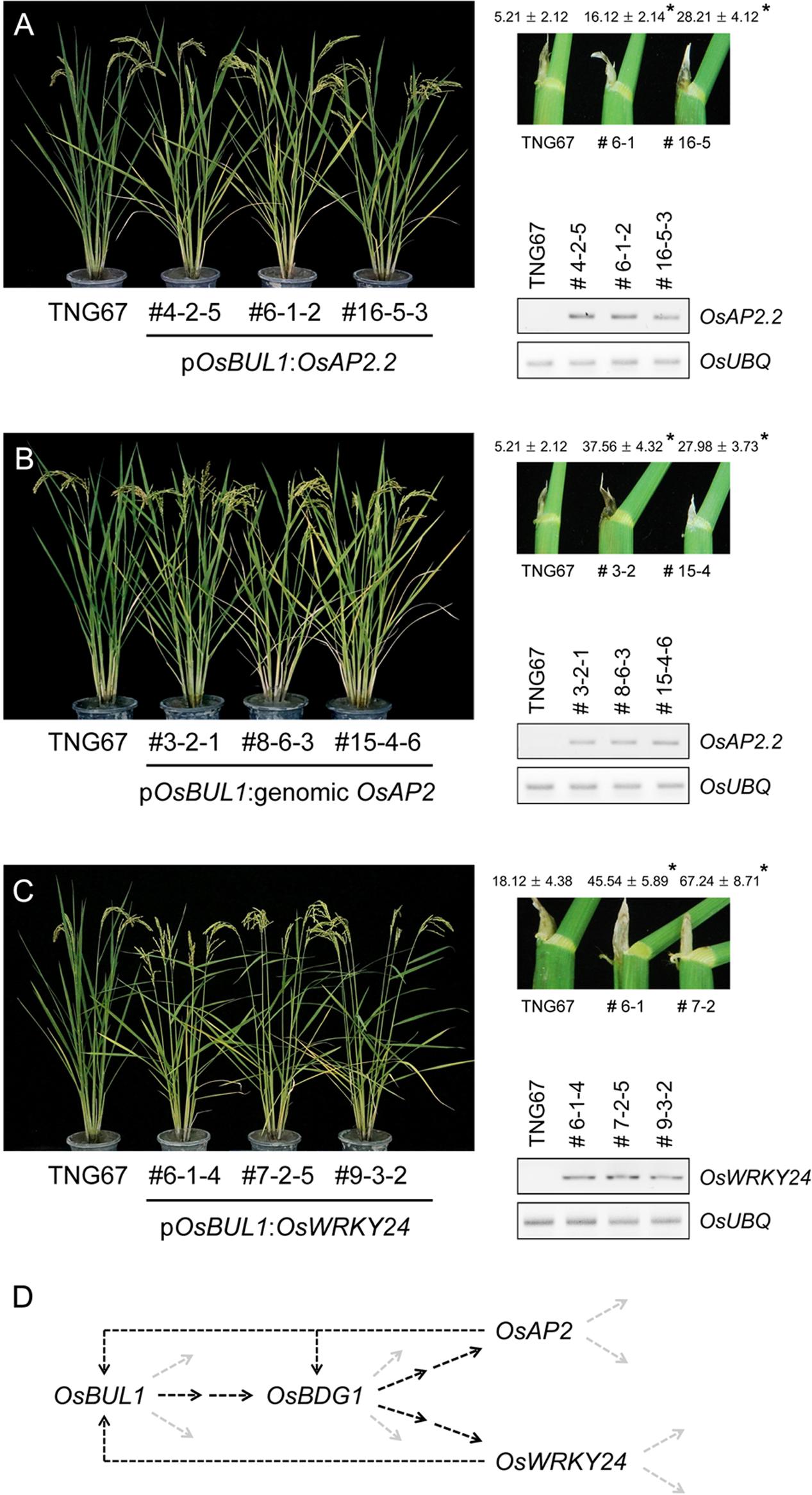
FIGURE 6. Generation of transgenic rice plants expressing OsAP2 and OsWRKY24 under the control of OsBUL1 promoter. (A,B) Transgenic rice containing pOsBUL1:OsAP2.2, a short transcript of OsAP2 (Os10g41330) or genomic OsAP2 showed increased lamina inclination. The longer OsAP2.1 transcript was not detected in pOsBUL1:genomic OsAP2 plants. Leaf angle was measured with the second leaf from the top of the main stem. (C) Transgenic rice with pOsBUL1:OsWRKY24 showed significant increase in leaf angles. The third leaf angles in the WT and transgenic lines were measured. RNAs were extracted from collars for cDNA synthesis and RT-PCR. Values for leaf angles are given as means ± SD (degree; n = 10 to 18). (∗P < 0.01, Student’s t-test). (D) Flow of genes acting in lamina inclination identified in this study. The dashed black lines were drawn based on results of expression analyses and transgenic approaches.
Discussion
Controlling leaf angle and grain size of crop plants occupies a key position in the generation of elite lines with desirable agronomic traits in crop breeding programs. However, the mechanisms regulating the traits remain largely unknown. In this work, we identified putative genes acting downstream of OsBUL1 for a positive effect on lamina inclination and grain size. First, with a view to investigating the downstream genes of OsBUL1, collection and comparison of transcriptomes of collars and panicles from WT and osbul1 were conducted through microarray hybridization and, OsBDG1 was selected as a putative downstream gene of OsBUL1. The expression of OsBDG1 was reduced both in collars and panicles of osbul1 and conversely increased in overexpressing plants and gain-of-function mutant of OsBUL1. Previously, OsBDG1 was shown to be upregulated in transgenic rice ectopically expressing a sterol C-22 hydroxylase that controls BR levels in plants (Wu et al., 2008). Also, OsBDG1 encodes a novel small protein with a rare structural feature; an LRR N-terminal domain (LRRNT) at the N-terminal part and three LRRs toward its carboxyl terminus. In this study, OsBDG1 transcripts were shown to accumulate in response to BL and transgenic rice plants with overexpression and reduced expression of OsBDG1 exhibited higher and lower sensitivities to BL, respectively, in lamina joint inclination bioassays (Supplementary Figure S4). Transgenic rice plants with increased expression of OsBDG1 driven by ubiquitin promoter or OsBUL1 promoter had phenotypes similar to those of rice plants including gain-of-function mutants of OsBDG1, OsBUL1 and OsBUL1 overexpressors (Jang et al., 2017) whereas OsBDG1-dsRNAi lines displayed similar phenotypes to osbul1 in lamina inclination and grain size implying correlation between the expression and functional cascade between the two genes, although we cannot exclude the possibility of their having parallel genetic pathways.
Sequentially, two genes encoding nuclear proteins, OsAP2 and OsWARKY24 were identified as being downstream of OsBDG1 in the lamina joint based on expression analyses of pOsBUL1:OsBDG1 plants. Moreover, the expression level of the two genes was lower in osbul1 as well as OsBDG1 knockdown lines demonstrating an expression cascade among OsBUL1, OsBDG1, OsAP2, and OsWRKY24 genes (Supplementary Figures S2). Recently, it was reported that SMALL ORGAN SIZE1 (SMOS1) encoding an AP2-type transcriptional factor acts as an auxin-dependent regulator for cell expansion during organ size control (Aya et al., 2014) and SHOEBOX (SHB), another AP2/ERF transcription factor directly activates transcription of the GA biosynthesis gene KS1 for the elongation of meristem cells in a developmental stage-specific manner (Li et al., 2015). Moreover, Wang et al. (2005) reported that OsWRKY11, a WRKY transcription factor, also regulates leaf inclination by analyzing a leaf angle mutant large leaf angles (lla), a T-DNA insertion mutant of OsWRKY11. Intriguingly, both OsAP2.2 and OsWARKY24 genes are upregulated by GA3 and slightly by BL and each protein exhibits transcriptional activation activity indicating that OsAP2.2 and OsWRKY24 may act as transcriptional activators to regulate the expression of downstream genes by phytohormones such as GA3 and/or BL. Functional characterization of the two genes by analyzing transgenic rice plants expressing each gene in the place where OsBUL1 is expressed also suggests that they influence cell elongation in a positive manner. Thus, we found expressional and putative functional relationships of genes involved in the promotion of cell elongation for increased lamina inclination and grain size of rice although it remains unknown whether direct regulation is available among these genes (Figure 6D). It would be of value to examine transcripts affected by the two transcription factors, OsAP2.2 and OsWRKY24 in the lamina joint of rice. The next challenge is to produce rice plants with erect leaves and larger grain size. A trial for reduced expression of genes such as OsBUL1 and/or OsBDG1 under the control of OsBC1 promoter (Jang et al., 2017) is worth conducting to test whether rice plants can exhibit an erect leaf trait without compromising on the grain size.
Conclusion
We have identified a series of novel rice genes related to increased lamina inclination and grain size based on exploitation of their expressional relationships and evaluated their functional roles by molecular genetic approaches. These results indicate that they are good candidates that may be further studied or used together with proper promoters for improving crop productivity through desirable plant architecture in the future.
Accession Numbers
Genes in this article can be found in the GenBank/EMBL or RiceGE databases under the following accession numbers: OsAP2 (Os10g41330), OsBDG1 (Os11g31530), OsBUL1 (Os02g51320), OsEXPA1 (Os04g15840), OsEXPA2 (Os01g60770), OsEXPA3 (Os05g19570), OsEXPA4 (Os05g39990), OsWRKY24 (Os01g61080), OsXTH1 (Os04g51460), OsXTR1 (Os11g33270). GEO accession number for microarray data in this study is GSE93817.
Author Contributions
SJ designed the experiments. SJ and H-YL performed the experiments, analyzed the data. SJ wrote the article.
Funding
This research was supported by a core grant from the Biotechnology Center in Southern Taiwan (BCST) of the Agricultural Biotechnology Research Center (ABRC), Academia Sinica, Taiwan to SJ.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Dr. Gynheung An (Crop Biotech Institute, Kyung Hee University) for providing the OsBDG1 activation tagging mutant. We thank Ms. Pei-Chun Liao for rice transformation, Ms. Ching-Han Wang for lamina bending assays and Ms. Mei-Lin Kuo for analyses of microarray data. We also thank members of core facility laboratories of Academia Sinica for microscopy and DNA sequencing and Ms. Miranda Loney for help with English editing.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fpls.2017.01253/full#supplementary-material
FIGURE S1 | Sequence of OsAP2 genomic clone used for construction of pOsBUL1:genomic OsAP2. The long transcript of OsAP2 (OsAP2.1) is marked with uppercase and the yellow block is for OsAP2.2, a short transcript. Introns are shown in lowercase. Underlined sequences are quantitative PCR primers for OsAP2.1 and double underlined sequences are for OsAP2.2. Start and stop codons are presented with red color.
FIGURE S2 | Expression of OsAP2 and OsWRKY24 is reduced in osbul1 plants. (A) A heat map showing that the expression level of OsAP2 and OsWRKY24 as well as OsBDG1 is reduced in osbul1 plants. Color scale represents log signal values. (B,C) Experimental confirmation of the expression level of OsAP2.2 and OsWRKY24 compared with microarray data in (A). The expression of OsAP2.2 and OsWRKY24 is reduced in collars (D) and panicles (E) of OsBDG1-dsRNAi lines without significant alteration of OsBUL1 expression (F). Data are the average of two or three independent experiments and normalized by OsUBQ. Error bars indicate SD.
FIGURE S3 | Expression of OsBUL1 and OsBDG1 is affected by pOsBUL1:OsWRKY24 and/or pOsBUL1:OsAP2. (A) OsBUL1 expression is increased by pOsBUL1:OsWRKY24 and pOsBUL1:OsAP2.2 in panicles while OsBDG1 transcripts (B) are accumulated only in pOsBUL1:OsAP2.2 plants. Data are the average of three independent experiments and normalized by OsUBQ. Error bars indicate SD. (C) Panicle branches exhibit increased angles in pOsBUL1:OsAP2.2 and pOsBUL1:OsWRKY24 transgenic rice plants. Arrowheads indicate nodes for panicles. Bar = 5 cm. Length of grains from the transgenic lines also show a significant increase. (mm; n > 25). (∗P < 0.01, Student’s t-test).
FIGURE S4 | Expression pattern of OsBDG1, OsAP2.2 and OsWRKY24 analyzed by quantitative RT-PCR and lamina inclination of pOsBUL1:OsBDG1 and OsBDG1-dsRNAi plants in lamina bending assays. (A) Spatiotemporal expression patterns of genes studied. Error bars indicate SD of three technical repeats. (B) Expression analyses of genes with treatment of GA3 and BL at 24 h time points after treatment. Data are the average of three or four independent experiments and normalized by OsUBQ or OsAct. Error bars indicate SD of three biological replicates. Differences between the mock and hormone treated samples are highlighted with ∗P < 0.01; ∗∗P < 0.05 with Student’s t-test. (C) Lamina joint inclination bioassays with various concentrations of BL.
FIGURE S5 | Subcellular localization of proteins. (A) YFP:OsBDG1 and CFP:OsBC1 were co-transformed into rice protoplasts. OsBDG1 is localized in the cytoplasm as well as the nucleus while OsBC1 (Jang et al., 2017) is only found in the nucleus. (B) CFP:OsBDG1 and YFP:OsBUL1 were co-transformed into rice protoplasts. OsBDG1 and OsBUL1 are co-localized in the cell. (C,D) CFP:OsAP2.2 and CFP:OsWRKY24 are localized in the nucleus. Bar = 5 μm.
FIGURE S6 | Morphological alteration of epidermal cells of lemma from pOsBUL1:OsAP2.2 and pOsBUL1:OsWRKY24 and expression pattern of genes involved in cell elongation. (A) Elongated epidermal cells of lemma from pOsBUL1:OsAP2.2 and pOsBUL1:OsWRKY24 transgenic plants compared to those of WT. Bar = 100 μm. (B) Expression of genes involved in cell elongation in spikelets of transgenic lines. Data are the average of three independent experiments and normalized by OsAct. Error bars indicate SD. (C) Transcriptional activation activity analyses of OsAP2.2 and OsWRKY24. Unlike BD-OsBDG1, both BD-OsAP2.2 and BD-OsWARK24 fusion proteins displayed a transcriptional activation activity in the yeast system described by Jang et al. (2015).
TABLE S1 | Primers used for expression analyses in this study.
Footnotes
- ^ http://www.phalanxbiotech.com
- ^ http://signal.salk.edu/cgi-bin/RiceGE
- ^ http://www.phalanxbiotech.com
References
An, S., Park, S., Jeong, D.-H., Lee, D.-Y., Kang, H.-G., Yu, J.-H., et al. (2003). Generation and analysis of end sequence database for T-DNA tagging lines in rice. Plant Physiol. 133, 2040–2047. doi: 10.1104/pp.103.030478
Aya, K., Hobo, T., Sato-Izawa, K., Ueguchi-Tanaka, M., Kitano, H., and Matsuoka, M. (2014). A novel AP2-type transcription factor, SMALL ORGAN SIZE1, controls organ size downstream of an auxin signaling pathway. Plant Cell Physiol. 55, 897–912. doi: 10.1093/pcp/pcu023
Bai, M.-Y., Zhang, L.-Y., Gampala, S. S., Zhu, S.-W., Song, W.-Y., Chong, K., et al. (2007). Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc. Natl. Acad. Sci. U.S.A. 104, 13839–13844. doi: 10.1073/pnas.0706386104
Caldana, C., Scheible, W.-R., Mueller-Roeber, B., and Ruzicic, S. (2007). A quantitative RT-PCR platform for high-throughput expression profiling of 2500 rice transcription factors. Plant Methods 3:7. doi: 10.1186/1746-4811-3-7
Cao, H., and Chen, S. (1995). Brassinosteroid-induced rice lamina joint inclination and its relation to indole-3-acetic acid and ethylene. Plant Growth Regulat. 16, 189–196. doi: 10.1007/BF00029540
Cho, H. T., and Kende, H. (1997). Expression of expansin genes is correlated with growth in deepwater rice. Plant Cell 9, 1661–1671. doi: 10.1105/tpc.9.9.1661
Feng, Z., Wu, C., Wang, C., Roh, J., Zhang, L., Chen, J., et al. (2016). SLG controls grain size and leaf angle by modulating brassinosteroid homeostasis in rice. J. Exp. Bot. 67, 4241–4253. doi: 10.1093/jxb/erw204
Guénin, S., Mauriat, M., Pelloux, J., Van Wuytswinkel, O., Bellini, C., and Gutierrez, L. (2009). Normalization of qRT-PCR data: the necessity of adopting a systematic, experimental conditions-specific, validation of references. J. Exp. Bot. 60, 487–493. doi: 10.1093/jxb/ern305
Hara, Y., Yokoyama, R., Osakabe, K., Toki, S., and Nishitani, K. (2014). Function of xyloglucan endotransglucosylase/hydrolases in rice. Ann. Bot. 114, 1309–1318. doi: 10.1093/aob/mct292
Heang, D., and Sassa, H. (2012). Antagonistic actions of HLH/bHLH proteins are involved in grain length and weight in rice. PLoS ONE 7:e31325. doi: 10.1371/journal.pone.0031325
Hong, Z., Ueguchi-Tanaka, M., Shimizu-Sato, S., Inukai, Y., Fujioka, S., Shimada, Y., et al. (2002). Loss-of-function of a rice brassinosteroid biosynthetic enzyme, C-6 oxidase, prevents the organized arrangement and polar elongation of cells in the leaves and stem. Plant J. 32, 495–508. doi: 10.1046/j.1365-313X.2002.01438.x
Hong, Z., Ueguchi-Tanaka, M., Umemura, K., Uozu, S., Fujioka, S., Takatsuto, S., et al. (2003). A rice brassinosteroid-deficient mutant, ebisu dwarf (d2), is caused by a loss of function of a new member of cytochrome P450. Plant Cell 15, 2900–2910. doi: 10.1105/tpc.014712
Jang, S., An, G., and Li, H.-Y. (2017). Rice leaf angle and grain size are affected by the OsBUL1 transcriptional activator complex. Plant Physiol. 173, 688–702. doi: 10.1104/pp.16.01653
Jang, S., Choi, S.-C., Li, H.-Y., An, G., and Schmelzer, E. (2015). Functional characterization of phalaenopsis aphrodite flowering genes PaFT1 and PaFD. PLoS ONE 10:e0134987. doi: 10.1371/journal.pone.0134987
Jeon, J.-S., Lee, S., Jung, K.-H., Jun, S.-H., Jeong, D.-H., Lee, J., et al. (2000). T-DNA insertional mutagenesis for functional genomics in rice. Plant J. 22, 561–570. doi: 10.1046/j.1365-313x.2000.00767.x
Jeong, D.-H., An, S., Park, S., Kang, H.-G., Park, G.-G., Kim, S.-R., et al. (2006). Generation of a flanking sequence-tag database for activation-tagging lines in japonica rice. Plant J. 45, 123–132. doi: 10.1111/j.1365-313X.2005.02610.x
Jeong, D.-H., Lee, S., Kim, S. L., Hwang, I., and An, G. (2007). Regulation of brassinosteroid responses by phytochrome B in rice. Plant Cell Environ. 30, 590–599. doi: 10.1111/j.1365-3040.2007.01644.x
Kim, S.-R., Lee, D.-Y., Yang, J.-I., Moon, S., and An, G. (2009). Cloning vectors for rice. J. Plant Biol. 52, 73–78. doi: 10.1007/s12374-008-9008-4
Komatsu, K., Maekawa, M., Ujiie, S., Satake, Y., Furutani, I., Okamoto, H., et al. (2003). LAX and SPA: major regulators of shoot branching in rice. Proc. Natl. Acad. Sci. U.S.A. 100, 11765–11770. doi: 10.1073/pnas.1932414100
Komiya, R., Ikegami, A., Tamaki, S., Yokoi, S., and Shimamoto, K. (2008). Hd3a and RFT1 are essential for flowering in rice. Development 135, 767–774. doi: 10.1242/dev.008631
Lee, J., Park, J.-J., Kim, S. L., Yim, J., and An, G. (2007). Mutations in the rice liguleless gene result in a complete loss of the auricle, ligule, and laminar joint. Plant Mol. Biol. 65, 487–499. doi: 10.1007/s11103-007-9196-1
Lee, S., Choi, S. C., and An, G. (2008). Rice SVP-group MADS-box proteins, OsMADS22 and OsMADS55, are negative regulators of brassinosteroid responses. Plant J. 54, 93–105. doi: 10.1111/j.1365-313X.2008.03406.x
Li, J., Zhao, Y., Chu, H., Wang, L., Fu, Y., Liu, P., et al. (2015). SHOEBOX modulates root meristem size in rice through dose-dependent effects of gibberellins on cell elongation and proliferation. PLoS Genetics 11:e1005464. doi: 10.1371/journal.pgen.1005464
Nakamura, A., Fujioka, S., Takatsuto, S., Tsujimoto, M., Kitano, H., Yoshida, S., et al. (2009). Involvement of C-22-hydroxylated brassinosteroids in auxin-induced lamina joint bending in rice. Plant Cell Physiol. 50, 1627–1635. doi: 10.1093/pcp/pcp106
Sakamoto, T., Morinaka, Y., Ohnishi, T., Sunohara, H., Fujioka, S., Ueguchi-Tanaka, M., et al. (2006). Erect leaves caused by brassinosteroid deficiency increase biomass production and grain yield in rice. Nat Biotechnol. 24, 105–109. doi: 10.1038/nbt1173
Shimada, A., Ueguchi-Tanaka, M., Sakamoto, T., Fujioka, S., Takatsuto, S., Yoshida, S., et al. (2006). The rice SPINDLY gene functions as a negative regulator of gibberellin signaling by controlling the suppressive function of the DELLA protein, SLR1, and modulating brassinosteroid synthesis. Plant J. 48, 390–402. doi: 10.1111/j.1365-313X.2006.02875.x
Sinclair, T. R., and Sheehy, J. E. (1999). Erect leaves and photosynthesis in rice. Science 283:1455. doi: 10.1126/science.283.5407.1455c
Tanaka, A., Nakagawa, H., Tomita, C., Shimatani, Z., Ohtake, M., Nomura, T., et al. (2009). BRASSINOSTEROID UPREGULATED1, encoding a helix-loop-helix protein, is a novel gene involved in brassinosteroid signaling and controls bending of the lamina joint in rice. Plant Physiol. 151, 669–680. doi: 10.1104/pp.109.140806
Uozu, S., Tanaka-Ueguchi, M., Kitano, H., Hattori, K., and Matsuoka, M. (2000). Characterization of XET-related genes of rice. Plant Physiol. 122, 853–860. doi: 10.1104/pp.122.3.853
Van Camp, W. (2005). Yield enhancement genes: seeds for growth. Curr. Opin. Biotechnol. 16, 147–153. doi: 10.1016/j.copbio.2005.03.002
Wang, D., Zhang, H., Hu, G., Fu, Y., Si, H., and Sun, Z. (2005). Genetic analysis and identification of alarge leaf angles (lla) mutant in rice. Chin. Sci. Bull. 50, 492–494. doi: 10.1007/bf02897468
Wang, L., Xu, Y., Zhang, C., Ma, Q., Joo, S.-H., Kim, S.-K., et al. (2008). OsLIC, a novel CCCH-type zinc finger protein with transcription activation, mediates rice architecture via brassinosteroids signaling. PLoS ONE 3:e3521. doi: 10.1371/journal.pone.0003521
Wu, C.-Y., Trieu, A., Radhakrishnan, P., Kwok, S. F., Harris, S., Zhang, K., et al. (2008). Brassinosteroids regulate grain filling in rice. Plant Cell 20, 2130–2145. doi: 10.1105/tpc.107.055087
Wu, F.-H., Shen, S.-C., Lee, L.-Y., Lee, S.-H., Chan, M.-T., and Lin, C.-S. (2009). Tape-Arabidopsis sandwich - a simpler Arabidopsis protoplast isolation method. Plant Methods 5:16 doi: 10.1186/1746-4811-5-16
Yamamuro, C., Ihara, Y., Wu, X., Noguchi, T., Fujioka, S., Takatsuto, S., et al. (2000). Loss of function of a rice brassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell 12, 1591–1606. doi: 10.1105/tpc.12.9.1591
Zhang, L.-Y., Bai, M.-Y., Wu, J., Zhu, J.-Y., Wang, H., Zhang, Z., et al. (2009). Antagonistic HLH/bHLH transcription factors mediate brassinosteroid regulation of cell elongation and plant development in rice and Arabidopsis. Plant Cell 21, 3767–3780. doi: 10.1105/tpc.109.070441
Zhang, X., Sun, J., Cao, X., and Song, X. (2015). Epigenetic mutation of RAV6 affects leaf angle and seed size in rice. Plant Physiol. 169, 2118–2128. doi: 10.1104/pp.15.00836
Zhang, Y., Su, J., Duan, S., Ao, Y., Dai, J., Liu, J., et al. (2011). A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods 7:30. doi: 10.1186/1746-4811-7-30
Zhao, S.-Q., Hu, J., Guo, L.-B., Qian, Q., and Xue, H.-W. (2010). Rice leaf inclination2, a VIN3-like protein, regulates leaf angle through modulating cell division of the collar. Cell Res. 20, 935–947. doi: 10.1038/cr.2010.109
Keywords: leaf inclination, grain size, plant architecture, transcriptional activator, transgenic rice, Oryza sativa
Citation: Jang S and Li H-Y (2017) Oryza sativa BRASSINOSTEROID UPREGULATED1 LIKE1 Induces the Expression of a Gene Encoding a Small Leucine-Rich-Repeat Protein to Positively Regulate Lamina Inclination and Grain Size in Rice. Front. Plant Sci. 8:1253. doi: 10.3389/fpls.2017.01253
Received: 16 May 2017; Accepted: 03 July 2017;
Published: 17 July 2017.
Edited by:
Hiroshi Takatsuji, National Agriculture and Food Research Organization (NARO), JapanReviewed by:
Eiji Nambara, University of Toronto, CanadaRobert Henry, The University of Queensland, Australia
Yukihiro Ito, Tohoku University, Japan
Copyright © 2017 Jang and Li. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Seonghoe Jang, ZmxvcmlnZW5AZ2F0ZS5zaW5pY2EuZWR1LnR3
 Seonghoe Jang
Seonghoe Jang Hsing-Yi Li
Hsing-Yi Li