- Department of Plant Agriculture, University of Guelph, Guelph, ON, Canada
Legumes are protein sources for billions of humans and livestock. These traits are enabled by symbiotic nitrogen fixation (SNF), whereby root nodule-inhabiting rhizobia bacteria convert atmospheric nitrogen (N) into usable N. Unfortunately, SNF rates in legume crops suffer from undiagnosed incompatible/suboptimal interactions between crop varieties and rhizobia strains. There are opportunities to test much large numbers of rhizobia strains if cost/labor-effective diagnostic tests become available which may especially benefit researchers in developing countries. Inside root nodules, fixed N from rhizobia is assimilated into amino acids including glutamine (Gln) for export to shoots as the major fraction (amide-exporting legumes) or as the minor fraction (ureide-exporting legumes). Here, we have developed a new leaf punch based technique to screen rhizobia inoculants for SNF activity following inoculation of both amide exporting and ureide exporting legumes. The assay is based on measuring Gln output using the GlnLux biosensor, which consists of Escherichia coli cells auxotrophic for Gln and expressing a constitutive lux operon. Subsistence farmer varieties of an amide exporter (lentil) and two ureide exporters (cowpea and soybean) were inoculated with different strains of rhizobia under controlled conditions, then extracts of single leaf punches were incubated with GlnLux cells, and light-output was measured using a 96-well luminometer. In the absence of external N and under controlled conditions, the results from the leaf punch assay correlated with 15N-based measurements, shoot N percentage, and shoot total fixed N in all three crops. The technology is rapid, inexpensive, high-throughput, requires minimum technical expertise and very little tissue, and hence is relatively non-destructive. We compared and contrasted the benefits and limitations of this novel diagnostic assay to methods.
Introduction
Legumes are critical to human and livestock agricultural systems (Broughton et al., 2003; Leigh, 2004; Foyer et al., 2016), as they fix atmospheric nitrogen (N) into a usable form of N (NH3/) through symbiotic association with rhizobia bacteria inside underground root nodule organs (Udvardi and Poole, 2013). The enzyme glutamine (Gln) synthetase assimilates into amino acids (Bernard and Habash, 2009; Betti et al., 2012). Therefore, symbiotic nitrogen fixation (SNF) provides the essential building block for amino acid biosynthesis (Prell and Poole, 2006; Udvardi and Poole, 2013), enabling legumes such as lentil and soybean to be a primary source of high-quality protein for billions of people especially in developing nations, and for livestock especially in wealthier societies (Graham and Vance, 2003). SNF also enables the deposition of organic fertilizer into soil during litter decomposition (Thilakarathna et al., 2015, 2016a), thus reducing the need for synthetic N fertilizers, of which subsistence farmers are primary beneficiaries. Unfortunately, some legume species and varieties show low SNF rates, in part due to incompatible or suboptimal interactions with strains of available soil rhizobia (Thilakarathna and Raizada, 2017). Although improved rhizobia can be introduced, screening a panel of rhizobia strains against local crop varieties/landraces for improved SNF is expensive especially for subsistence farmers in developing nations, in part because of the limitations of current SNF diagnostic methods (Unkovich and Pate, 2000; Howieson and Dilworth, 2016). Symbiotic nitrogen fixation activity is currently measured using different methods including counting the number of differentiated rhizobia (bacteroids) per nodule (Bourcy et al., 2013), N difference assay (Unkovich et al., 2008), ureide assay (Unkovich et al., 2008), acetylene reduction assay (Lodwig et al., 2003; Starker et al., 2006), hydrogen production (Kiers et al., 2003; Cabeza et al., 2015), and 15N techniques (Lodwig et al., 2003), of which 15N is the most commonly used and considered to be the most accurate (Howieson and Dilworth, 2016). These methods are challenging in terms of time, labor, accuracy, and the need for a non-fixing reference plant, destructive sampling, large amounts of tissue, technical expertise, expensive reagents, and/or equipment (Herridge et al., 2008; Howieson and Dilworth, 2016). Therefore, there is a need for alternative high-throughput SNF diagnostic methods.
Inside root nodules, fixed N from rhizobia is assimilated into Gln (Prell and Poole, 2006; Tegeder, 2014; Figure 1A) for export to shoots as amino acids as the major fraction (amide-exporting legumes) or as the minor fraction (10–20% in ureide-exporting legumes) (Pate et al., 1980; Atkins, 1987). Generally, legumes that originate in temperate regions (e.g., alfalfa, pea, clover) are amide exporters, whereas those that originate in the tropics and subtropics (e.g., soybean, cowpea) predominately export fixed N as ureide compounds (e.g., allantoin) (Unkovich et al., 2008). Whole cell biosensors have been engineered to detect metabolites and emit outputs in response (e.g., fluorescence, luminescence) that can be easily measured, substituting for more expensive analytical chemistry methods such as liquid chromatography mass spectrometry (LC-MS) (Goron and Raizada, 2014). We previously developed a rapid leaf punch-based assay for Gln using a whole cell Gln biosensor called GlnLux (Tessaro et al., 2012). GlnLux was created by transforming an Escherichia coli Gln auxotroph with a constitutive lux reporter (Tessaro et al., 2012). Advantages of an auxotroph compared to expression-based reporter fusions are that there is an absolute requirement for the metabolite (i.e., Gln) and significantly fewer artifacts caused by microbial or plant metabolism or signaling. We showed that GlnLux cells exposed to extracts of leaf punches taken across the veins of growing leaves can accurately report N availability in maize (Tessaro et al., 2012; Goron and Raizada, 2016; Goron et al., 2017).
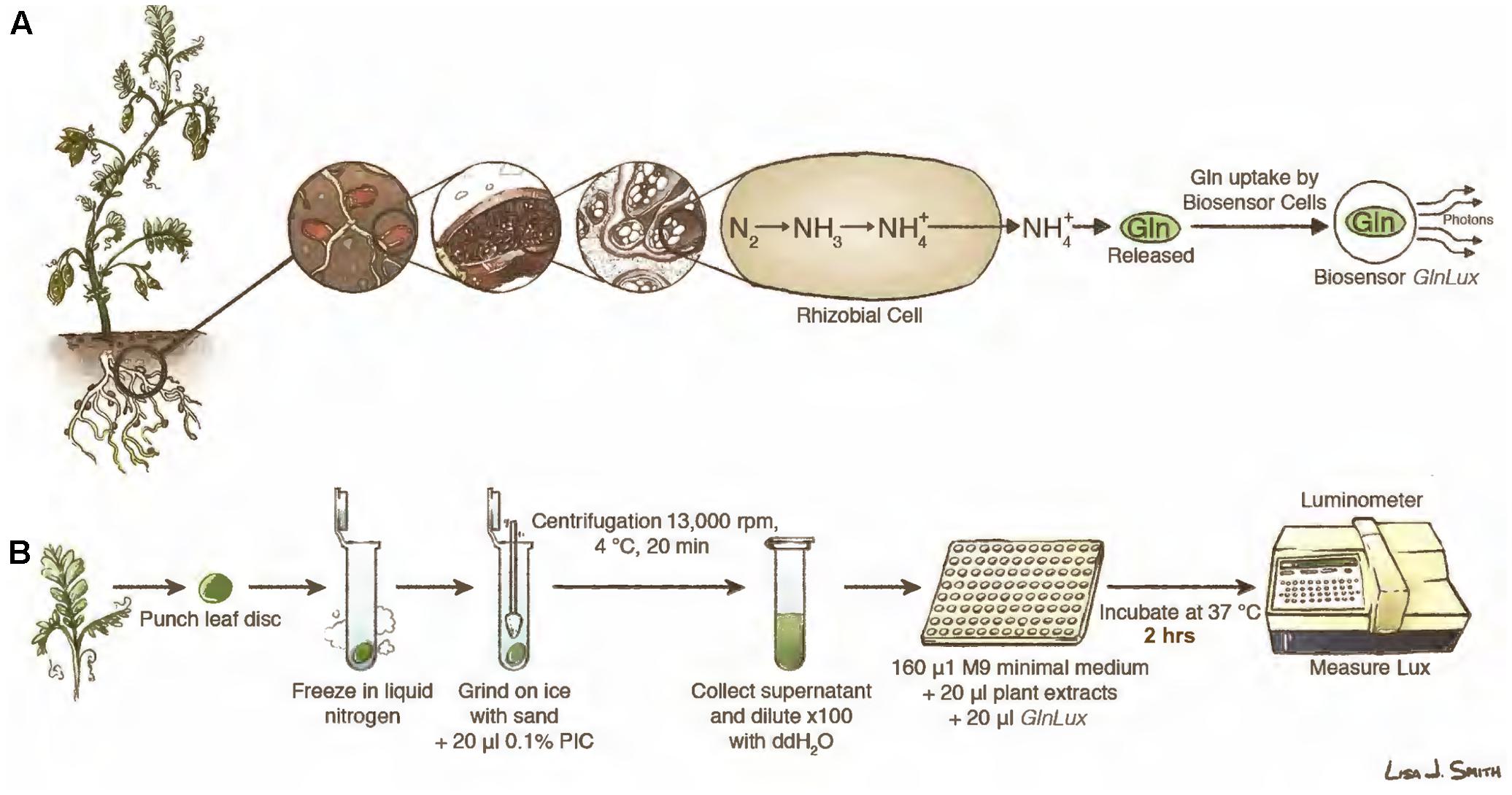
FIGURE 1. Leaf punch assay to measure symbiotic nitrogen fixation (SNF) in legumes using the GlnLux biosensor methodology. (A) Principle underlying the leaf punch assay. Legume plants have root nodules that host rhizobia, which following differentiation into bacteroids, convert atmospheric nitrogen (N) into , which is then assimilated into Gln. When Gln is released from plant tissues to the medium, it is taken up by GlnLux biosensor cells, causing them to become metabolically active and divide, which proportionally activates a lux operon to release photons that are measured using photon capture devices. (B) Overview of the assay using extracts of leaf punches incubated with GlnLux biosensor cells in 96-well plates and measured using a luminometer (see section Materials and Methods). Images courtesy of Lisa Smith (University of Guelph) can be re-used under the Creative Commons CC-BY License.
Here, we tested the hypothesis that the GlnLux leaf punch assay can be used to infer SNF output for high-throughput screening of legume inoculants under controlled conditions with minimal exogenous N (Figure 1A). Extracts of single leaf punches spanning veins of young legume leaves were incubated with GlnLux cells in 96-well liquid culture plates, then photon emissions were measured using a plate luminometer (Figure 1B). In order to validate the GlnLux leaf punch bioassay, two approaches were used: evaluation of different rhizobia strains using both amide and ureide exporting legumes. The crops used were subsistence Nepalese varieties of lentil (Lens culinaris) (an amide exporter), a legume of critical importance to South Asia, the Middle East, and East Africa (Sharpe et al., 2013); cowpea (Vigna unguiculata L. walp.) (a ureide exporter), a drought-tolerant legume grown in Sub-Saharan Africa, South Asia, and the Caribbean (Ehlers and Hall, 1997); and soybean (Glycine max) (a ureide exporter), the world’s most important legume in terms of production (Herridge et al., 2008). The results of GlnLux were compared to traditional SNF diagnostic methods including the standard 15N dilution technique. Plants were grown with minimal N fertilizer to facilitate N-isotope analysis.
Materials and Methods
Plant Materials and Rhizobium Strains
For the lentil × rhizobia strain experiment, the crop variety was Nepalese Simal, and there were four Rhizobium leguminosarum biovar viciae strains (wild type: 3841, VF39, 248; mutant 17-B) (Yost et al., 2006; Vanderlinde et al., 2011). For the cowpea × rhizobia strain experiment, the crop variety was Nepalese Surya and there were two strains of Bradyrhizobium yuanmingense (TTC9, TSC10), one strain of B. japonicum (DTB4) and one strain of B. elkanii (DTC9) (Sarr et al., 2009). For the soybean × rhizobia strain experiment, the crop variety was Nepalese Puja and there were three strains of B. japonicum (USDA 110, USDA 123, USDA 510) (Sadowsky et al., 1987; Bhagwat et al., 1991; Kaneko et al., 2002) and two strains of B. elkanii (USDA 94, USDA 76) (Kuykendall et al., 1992). Different rhizobia strains for each crop were selected to capture a diversity of nodulation and N fixation efficiency.
Plant Growth Conditions and Treatments
Seeds of all three crops were surface-sterilized with 70% ethanol for 2 min, 4% sodium hypochlorite for 3 min, and washed with six changes of sterile distilled water. Seeds were pre-germinated on sterilized, wetted filter paper in the dark at 28°C for 2 days, and individual seeds were transferred to germination pouches (17.8 × 16.5 cm, Mega International, Minneapolis, MN, United States) on light shelves containing 50 ml of double distilled water. Each plant was grown in a separate germination pouch. There was only one treatment (one rhizobia strain or non-inoculated control) per pouch, and each germination pouch was considered to be a single replicate. To further reduce cross-contamination, pouches were used that were sealed at the bottom, and nutrient solutions were not shared between treatments. For the lentil and cowpea studies, per treatment, five to six germination pouches were placed in a stand and placed in an open tupperware container. Different rhizobia treatments were not placed within the same container to avoid cross contamination of rhizobia strains. There were three containers per treatment, which were continuously randomized within and across light shelves. For the soybean study, five rhizobia treatments and the non-inoculated control (in individual germination pouches) were randomly allocated within a plastic container. There were eight trays (n = 8), which were continuously randomized within and across light shelves. After 1 week of germination, the water was removed and plants were supplied with 50 (lentil) and 100 ml (cowpea and soybean) of one-fourth strength N-free Hoagland’s nutrient solution (pH = 6.6), containing 0.5 mM K15NO3 (98 atom% 15N; 335134-1G; Sigma Aldrich, Oakville, ON, Canada) as starter N. One week later, this nutrient solution was removed and replaced with 50 ml of the one-fourth strength N-free Hoagland’s solution; the nutrient solution was replaced each week. Lentil plants were grown at 23 ± 2°C with supplemental lighting (range: 180–200 μmol m-2 s-1 at the top of the growth pouch, EcoLux SP65, 40W, F40SP65ECO), maintaining a photoperiod of 16 h/8 h light/dark cycles. Cowpea and soybean plants were grown at 23 ± 2°C with supplemental lighting (range: 250–300 μmol m-2 s-1 at the top of the growth pouch, EcoLux SP65, 40W, F40SP65ECO), maintaining a photoperiod of 16 h/8 h light/dark cycles. For the rhizobia treatments, strains for lentil (R. leguminosarum biovar viciae strains wild type: 3841, VF39, 248; mutant 17-B), cowpea (B. yuanmingense TTC9, B. yuanmingense TSC10, B. japonicum DTB4, B. elkanii DTC9), and soybean (B. japonicum USDA 110, B. japonicum USDA 123, B. japonicum USDA 510, B. elkanii USDA 94, B. elkanii USDA 76) were grown in tryptone-yeast (TY) extract and modified arabinose-gluconate medium (MAG) (cowpea and soybean) agar media, respectively, in Petri dishes for 3 days at 30°C. To prepare each inoculum, the agar plates were scraped and cells added to sterilized ddH2O, adjusted to a cell density of OD595 = 0.1. Plants were inoculated 1 week after plant growth with 1 ml of each inoculum, placed directly on roots. All the three legume species had a negative control, comprised of plants that were not inoculated with rhizobia.
GlnLux Leaf Punch Sampling
Eight healthy representative plants per treatment were sampled from lentil, cowpea, and soybean 4 weeks after inoculation with rhizobia as indicated. Leaf punches were collected from the fully expanded youngest leaf, across the leaflet main vein from lentil (from one leaflet at the tip), cowpea, and soybean using a 3 mm (lentil) and 6.35 mm (cowpea and soybean) hand puncher (235270975; Fiskars Brands Inc., Middleton, WI, United States) and immediately frozen in liquid N. Leaf punches were stored at -80°C.
Plant Morphometric Analysis
The same eight plants per treatment harvested for GlnLux measurements were used. The number of nodules per plant was counted manually. Roots were scanned using an Epson Expression 1640× scanner (Epson Canada Ltd., Markham, ON, Canada), and a detailed root morphological analysis was undertaken using WinRHIZO software (Regent Instruments Inc., Quebec City, QC, Canada) including root volume, total length, surface area, and average diameter. Shoot and root dry weights were measured after drying the plant materials in an oven at 60°C for 3 days.
Shoot Nitrogen and 15N Analysis
The same eight plants per treatment harvested for GlnLux measurements were used. The dried shoot samples (see above) were ground using a coffee grinder followed by a Bead Ruptor 12 Homogenizer (OMNI International, Kennesaw, GA, United States) and analyzed for 15N and total N% using a mass spectrometer (Costech ECS4010 Elemental Analyzer coupled to a Delta V mass spectrometer, Costech, CA, United States) at the Stable Isotope Facility, University of Saskatchewan, and University of British Columbia, Canada, using a standard protocol (Thilakarathna et al., 2012, 2016b). The %Ndfa of the lentil and soybean was calculated using the following formula according to the isotope dilution technique:
where atom % 15N excess = atom % 15N(lentil/soybean) - 0.3663.
The amount of shoot N derived from SNF was calculated based on the shoot N content and %Ndfa (shoot N content × %Ndfa/100).
Biosensor Strain
Construction of the GlnLux biosensor strain was previously reported (Tessaro et al., 2012). Briefly, a Gln-auxotrophic E. coli strain (JW3841-1, KanR) was generated by inserting a kanamycin cassette into GlnA [glnA732(del)::kan] (Baba et al., 2006). The strain was subsequently transformed with ampicillin-resistant plasmid pT7-lux (Meighen and Szittner, 1992) which contained a constitutive T7 promoter from Xenorhabdus luminescens driving expression of the luxCDABE operon from Vibrio fischeri to create strain GlnLux.
GlnLux Bacterial Growth Media
GlnLux bacteria were cultured in LB medium, which consisted of 5 g/l NaCl (BP358-212, Fisher Scientific), 5 g/l yeast extract (DF0127179, Fisher Scientific), and 10 g/l tryptone (BP1421-500, Fisher Scientific), with or without 12 g/l Bacto-Agar (BD; DF0140010, Fisher Scientific), pH 7.2. M9 minimal medium consisted of 20 ml/l 20% (w/v) D-(+)-glucose (G5767, Sigma), 100 μl/l 1 M CaCl2 (C-79, Fisher Scientific), 2 ml/l 1 M MgSO4 (230391, Sigma), and 200 ml/l 5× M9 salts (A-0171, Sigma), pH 7.0. All liquid and solid plate media were supplemented with 50 μg/ml kanamycin monosulfate (K378, PhytoTech, United States) and 100 μg/ml carbenicillin disodium salt (C346, PhytoTech, United States) to select for the disrupted glnA chromosome and reporter plasmid, respectively.
Leaf Punch Luminometer Assays – GlnLux Cell Preparation
Cells were prepared as previously described (Tessaro et al., 2012). Briefly, GlnLux bacteria were inoculated into 15 ml of LB medium in a 50 ml Falcon tube, and incubated overnight at 37°C with shaking at 250 × g. The culture was spun down at 2500 × g at 21°C for 10 min and the supernatant was decanted. The culture was then washed 3× in sterile M9 minimal medium with centrifugation as above. Finally, the culture was resuspended in 15 ml sterile M9 minimal medium in a 50 ml Falcon tube, and the GlnLux density was adjusted to OD595 = 0.025 using sterile M9 minimal medium. The resuspended GlnLux culture was incubated at 37°C with shaking at 250 × g for 14 h to deplete any endogenous Gln.
GlnLux Leaf Punch Luminometer Measurements of Plant Gln
The procedure was adapted from a previous protocol (Tessaro et al., 2012). Individual frozen leaf punches were ground in a 2 ml conical bottom microcentrifuge tube placed on ice using a micropestle (K7495150000, Kimble Chase, Fisher Scientific) with silica sand in 20 μl of 0.1% final (v/v) protease inhibitor cocktail (PIC) for plant cell extracts (100% stock; #P9599, Sigma). Plant extracts were centrifuged for 20 min at 4°C at 13,000 × g, and the supernatant was transferred to a microcentrifuge tube placed on ice. The plant extracts were diluted 100-fold in ddH2O. The diluted plant extracts were used for luminometer assays.
For luminometer assays, white opaque 96-well reader plates (#07-200-589, Fisher Scientific) were loaded with 160 μl/well M9 minimal medium followed by 20 μl/well of each plant extract. Finally, 20 μl of 14 h-Gln-depleted GlnLux culture (pre-depletion OD595 = 0.025) was added to each well. Plates were sealed with non-breathable sterile film (#361006008, Fisher Scientific) to prevent media evaporation, and centrifuged for 5 s at 2000 × g to mix the GlnLux bacteria with plant extracts and M9 minimal medium. The plates were incubated at 37°C without shaking. For lux quantification, plates were read in a MicoLumatPlus LB96V luminometer with WinGlow Software (Berthold Technologies, Germany). Samples were read after 2 h of incubation for 1 s in a luminometer chamber temperature of 37°C in an endpoint assay using the integrate software function. Readings were taken every hour after incubation until lux values saturated. The plates were transferred back and forth from a 37°C incubator, and non-breathable films were replaced following each read. The 0 μg/ml Gln standard reading was subtracted from all lux values.
To test the biosensor response to exogenous Gln, six different concentrations of Gln standards (L-Gln G229, PhytoTech, 0, 625 × 10-9, 125 × 10-8, 25 × 10-7, 5 × 10-6, 1 × 10-5 M) were also tested in the same plates along with plant samples, following the same method (Supplementary Figure S1).
Statistical Analyses
The effect of different rhizobia strains was analyzed using analysis of variance (ANOVA) set at p < 0.05. Means were compared using Tukey. To test for the linearity of the biosensor response, linear regression analysis was performed using a Goodness of Fit (R2) test. Total shoot N%, amount of shoot N fixed, and %Ndfa versus GlnLux correlation analysis were performed using the Pearson correlation test. All statistical analyses were performed using GraphPad Prism Software (v5, GraphPad Software, United States).
Results
Leaf Punch Screening of Inoculants Applied to Lentil
To test the GlnLux leaf punch assay with lentil, four strains of R. leguminosarum bv. viciae (VF39, 248, 3841, and 17-B), which differ in their N-fixation capacities, were inoculated onto a variety of lentil grown by smallholder Nepalese hillside farmers (Figure 2A). The different rhizobia strains all produced a similar number of root nodules (Figure 2B and Supplementary Figure S2A). A single punch per plant from a young leaf (from low-N grown lentils) was ground in PIC, co-incubated for 2 h with GlnLux cells in 96-well plates, and then the lux signal quantified after a 2-min read (Figure 1B). The GlnLux output (Figure 2E) showed a similar trend as the shoot N% (Figure 2F), and the percentage N derived from the atmosphere (%Ndfa) (Figure 2G). GlnLux output strongly correlated with SNF measured using the 15N stable isotope assay (%Ndfa) (Pearson correlation coefficient R2 = 0.9972) (Figure 2H), plant shoot N% (R2 = 0.8910) (Figure 2I), and amount of shoot N fixed (R2 = 0.890) (Figure 2J). The GlnLux result was not an artifact of altered Gln demand, as significant differences were not observed among the different rhizobia treatments for normalized shoot dry weight (Figure 2C), normalized root dry weight (Figure 2D), normalized total dry weight, shoot/root biomass ratio, root length, root surface area, or volume (Supplementary Figures S2B–F).
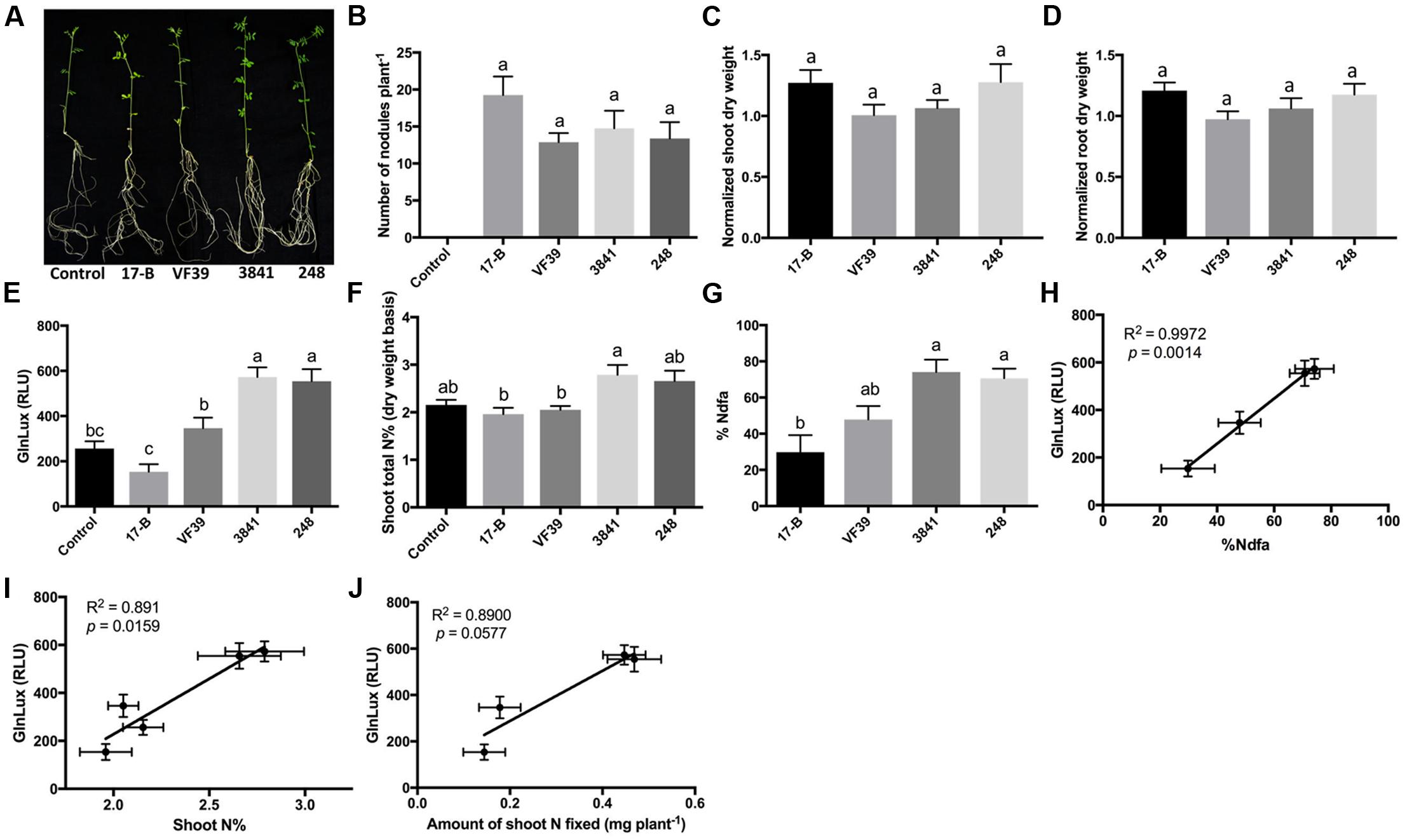
FIGURE 2. The GlnLux leaf punch assay to measure SNF in lentil. (A–D) Effect of rhizobia inoculants on morphological traits. (A) Representative pictures of lentil plants inoculated with four different strains of R. leguminosarum bv. viciae (wild types: 3841, VF39, 248; mutant 17-B) and the non-inoculated control. (B) Number of nodules per plant. The non-inoculated control had no nodules. (C) Normalized shoot dry matter per plant. Normalization of plant dry matter (rhizobia inoculated) was achieved by dividing dry matter of the non-inoculated plants. (D) Normalized root dry matter per plant. (E–J) Validation of the GlnLux luminometer assay as a method to measure SNF in an amide exporting legume (lentil) by comparing results to total N% and 15N dilution methods. (E) Corresponding GlnLux leaf punch outputs. A 0 μg/ml Gln standard reading was subtracted from all lux values, which were read in randomized replicates. RLU, relative lux units. (F) Shoot total N% analyzed using a mass spectrometer. (G) Percentage of N derived from the atmosphere (%Ndfa) calculated using the isotope dilution technique (see section Materials and Methods). (H) Correlation between GlnLux and %Ndfa (Pearson R2). (I) Correlation between GlnLux and shoot total N%. (J) Correlation between GlnLux and amount of shoot N fixed. For all graphs, the error bars represent the standard error of the mean (SEM) (N = 8). The different letters on top of each histogram indicate significant differences in the mean between treatments.
Leaf Punch Screening of Inoculants Applied to Cowpea
To test the GlnLux leaf punch assay with cowpea, a ureide exporter, two strains of B. yuanmingense (TTC9 and TSC10), one strain of B. japonicum (DTB4) and one strain of B. elkanii (DTC9), which differ in their nodulation capacity, were inoculated onto a variety of cowpea grown by smallholder Nepalese hillside farmers (Figure 3A). The different rhizobia strains produced a significantly different number of root nodules (P < 0.0001) (Figure 3B and Supplementary Figure S3A). The GlnLux output showed a similar trend (Figure 3E) as the shoot N% (Figure 3F) and the %Ndfa (Figure 3G). GlnLux output correlated with SNF measured using the 15N stable isotope assay (%Ndfa) (Pearson correlation coefficient R2 = 0.7038) (Figure 3H), plant shoot N% (R2 = 0.9873) (Figure 3I), and amount of shoot N fixed (R2 = 0.8641) (Figure 3J). Significant differences were not observed among the different rhizobia treatments for normalized root dry weight (Figure 3D), root length, root surface area, or volume (Supplementary Figures S3B–D). Plants inoculated with strain TTC9 had the highest shoot (Figure 3C) and total biomass (Supplementary Figure S3E), whereas the lowest biomass and shoot/root ratio were associated with strain DTC9 (Supplementary Figures S3E,F).
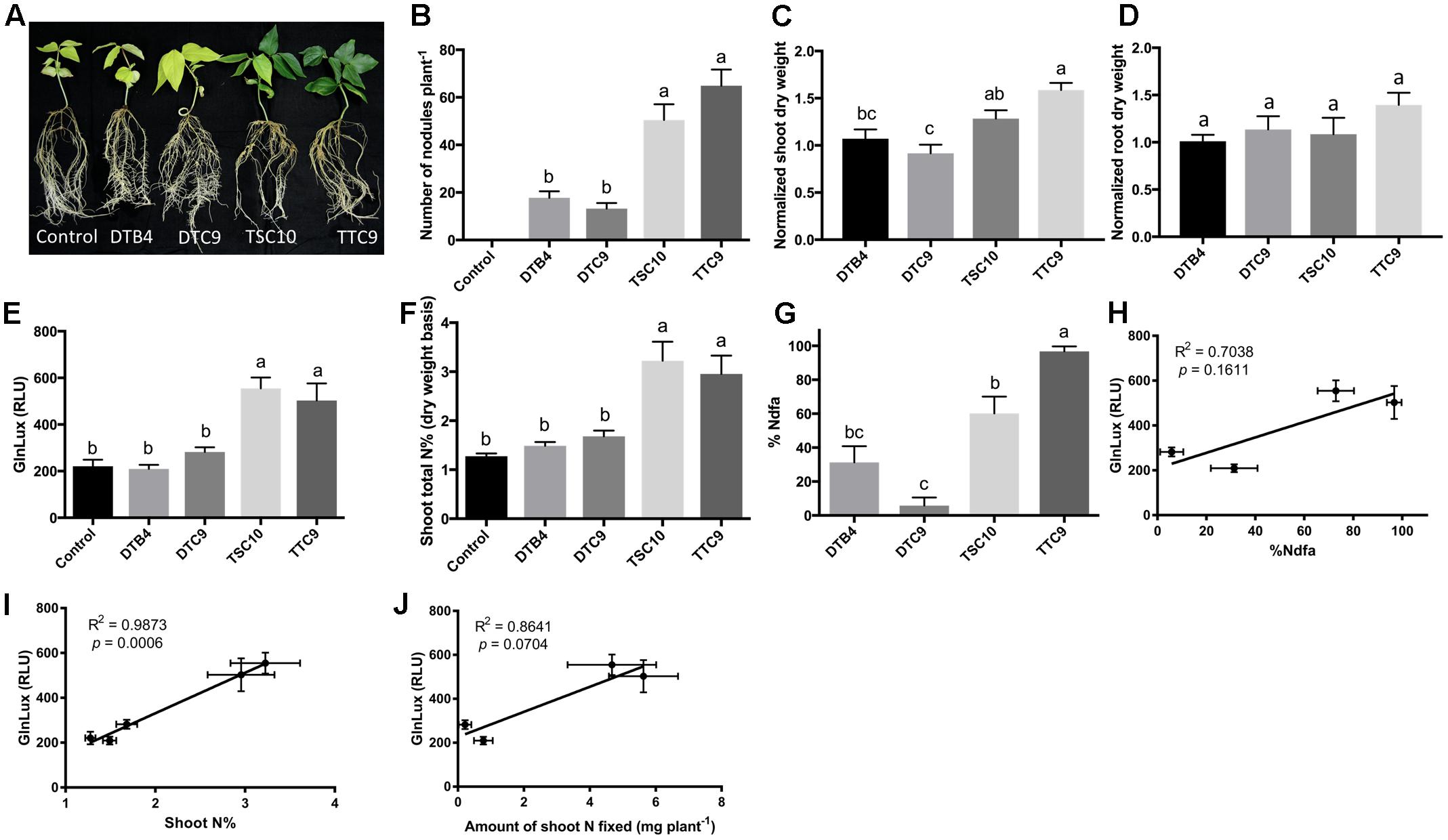
FIGURE 3. The GlnLux leaf punch assay to measure SNF in cowpea. (A–D) Effect of rhizobia inoculants on morphological traits. (A) Representative pictures of cowpea plants inoculated with two strains of B. yuanmingense (TTC9 and TSC10), one strain of B. japonicum (DTB4), one strain of B. elkanii (DTC9), and the non-inoculated control. (B) Number of nodules per plant. The non-inoculated control had no nodules. (C) Normalized shoot dry matter per plant. Normalization of plant dry matter (rhizobia inoculated) was achieved by dividing dry matter of the non-inoculated plants. (D) Normalized root dry matter per plant. (E–J) Validation of the GlnLux luminometer assay as a method to measure SNF in a ureide exporting legume (cowpea) by comparing results to total N% and 15N dilution methods. (E) Corresponding GlnLux leaf punch outputs. A 0 μg/ml Gln standard reading was subtracted from all lux values, which were read in randomized replicates. RLU, relative lux units. (F) Shoot total N% analyzed using a mass spectrometer. (G) Percentage of N derived from the atmosphere (%Ndfa) calculated using the isotope dilution technique (see section Materials and Methods). (H) Correlation between GlnLux and %Ndfa (Pearson R2). (I) Correlation between GlnLux and shoot total N%. (J) Correlation between GlnLux and amount of shoot N fixed. For all graphs, the error bars represent the SEM (N = 8). The different letters on top of each histogram indicate significant differences in the mean between treatments.
Leaf Punch Screening of Inoculants Applied to Soybean
To test the GlnLux leaf punch assay with soybean, another ureide exporter, a Nepalese subsistence farmer variety of soybean was inoculated with three strains of B. japonicum (USDA 110, USDA 123, 510) and two strains of B. elkanii (USDA 94 and USDA 76), which differ in their N-fixation capacities (Figure 4A). The different rhizobia strains produced a significantly different number of root nodules (P < 0.0001) (Figure 4B and Supplementary Figure S4A). The GlnLux output showed a similar trend (Figure 4E) as the shoot N% (Figure 4F), and the %Ndfa (Figure 4G). GlnLux output strongly correlated with SNF measured using the 15N stable isotope assay (%Ndfa) (Pearson correlation coefficient R2 = 0.9772) (Figure 4H), plant shoot N% (R2 = 0.9805) (Figure 4I), and amount of shoot N fixed (R2 = 0.9747) (Figure 4J). Significant differences were not observed among the different rhizobia treatments for root length, root surface area, or volume (Supplementary Figures S4B–D). However, slight differences were found among the rhizobia strains for normalized shoot (Figure 4C), root (Figure 4D), and total dry weight (Supplementary Figure S4E). Plants inoculated with USDA 110 (high fixer) had the highest shoot biomass, total biomass, and shoot/root ratio (Supplementary Figure S4F).
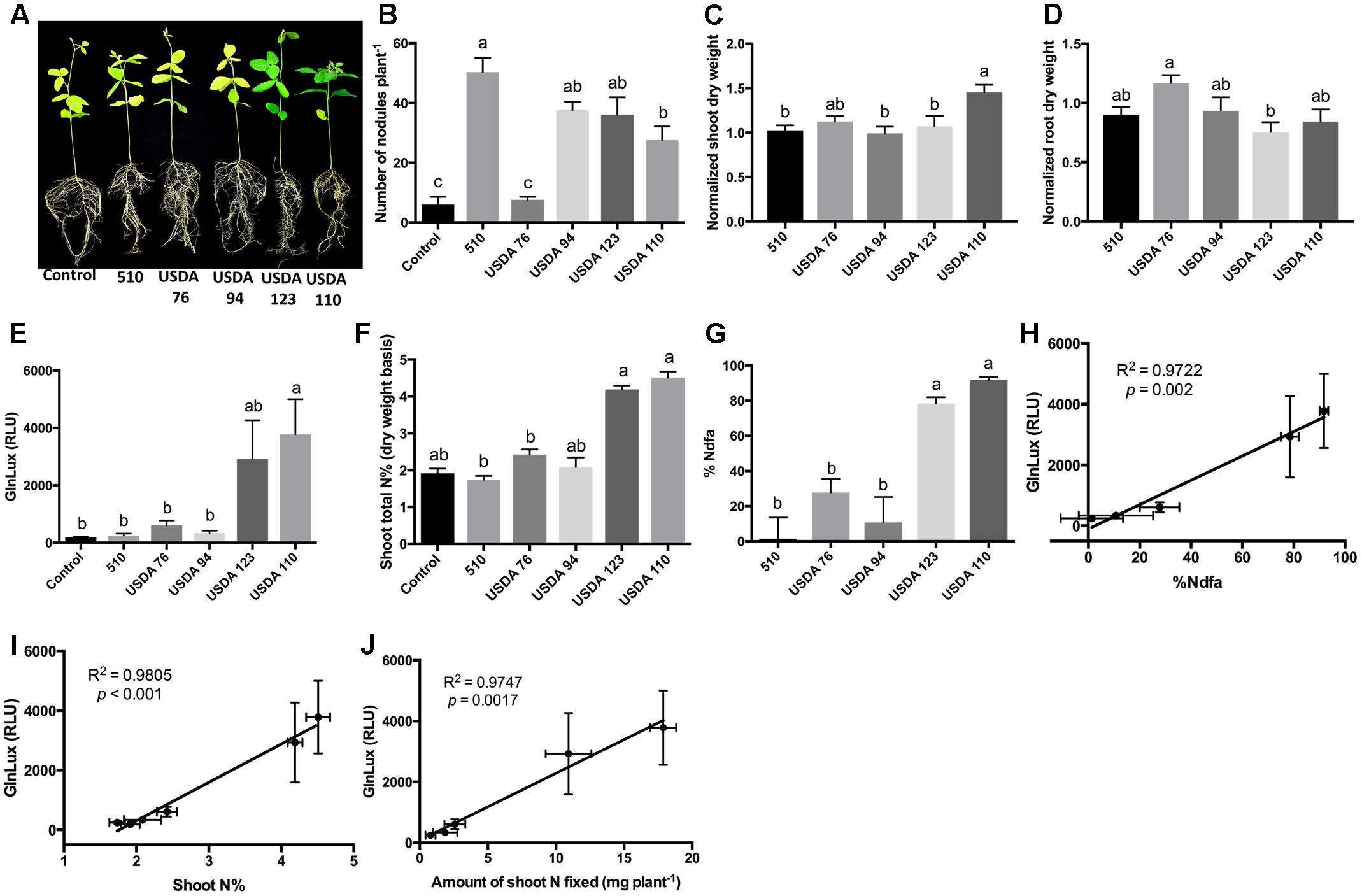
FIGURE 4. The GlnLux leaf punch assay to measure SNF in soybean. (A–D) Effect of rhizobia inoculants on morphological traits. (A) Representative pictures of soybean plants inoculated with three strains of B. japonicum (USDA 110, USDA 123, 510) and two strains of B. elkanii (USDA 94, USDA 76) and the non-inoculated control. (B) Number of nodules per plant. (C) Normalized shoot dry matter per plant. Normalization of plant dry matter (rhizobia inoculated) was achieved by dividing dry matter of the non-inoculated plants. (D) Normalized root dry matter per plant. (E–J) Validation of the GlnLux luminometer assay as a method to measure SNF in a ureide exporting legume (soybean) by comparing results to total N% and 15N dilution methods. (E) Corresponding GlnLux leaf punch outputs. A 0 μg/ml Gln standard reading was subtracted from all lux values, which were read in randomized replicates. RLU, relative lux units. (F) Shoot total N% analyzed using a mass spectrometer. (G) Percentage of N derived from the atmosphere (%Ndfa) calculated using the isotope dilution technique (see section Materials and Methods). (H) Correlation between GlnLux and %Ndfa (Pearson R2). (I) Correlation between GlnLux and shoot total N%. (J) Correlation between GlnLux and amount of shoot N fixed. For all graphs, the error bars represent the SEM (N = 8). The different letters on top of each histogram indicate significant differences in the mean between treatments.
Discussion
During the 1980s, the close relationship between xylem ureide composition and SNF activity was used to develop the well-known ureide assay, a technique to measure SNF in ureide exporting legumes (Herridge, 1982; Herridge and Peoples, 1990). 15N-methods have been used to validate the ureide method for SNF measurement (Herridge and Peoples, 1990). Similarly, here we have developed a new leaf punch-based technique to screen rhizobia inoculants for SNF activity following inoculation of both amide exporting (lentil) and ureide exporting (cowpea, soybean) legumes. The assay is based on measuring Gln output using the GlnLux biosensor, with minimal external N under controlled conditions. SNF-output as inferred from the GlnLux assay strongly correlated with the proportion of N derived from SNF measured using the 15N dilution method in both lentil (Figures 2H,J) and soybean (Figures 4H,J); the correlation for cowpea was lower but still strong (R2 = 0.7038) (Figures 3H,J). The cowpea result may have been due to the four selected rhizobia strains which represented extremes in SNF activity, either high (TSC-10 and TTC-9) or low (DTB-4 and DTC-9), with no intermediate strain(s) (Figure 3B). The SNF capacity of a legume can vary due to the nodule number and/or N fixation activity of a particular rhizobia strain – host genotype combination (Thilakarathna and Raizada, 2017). In lentil, differences in SNF output by the rhizobia were caused by differential SNF activity rather than nodulation (Figures 2B,G). In cowpea, SNF likely varied due to the dissimilar number of nodules observed between inoculants (Figures 3B,G), but we cannot rule out differences in SNF activity. In soybean, differences in SNF output were caused by differences in rhizobia activity and/or nodulation ability, as some inoculants (e.g., USDA 94) had high nodulation but apparent low fixation (Figures 4B,G).
We have compared the efficacy of the GlnLux method against the currently available methods to quantify SNF (Table 1). The GlnLux leaf punch test could be performed on juvenile plants, and bypassed the need for 15N or tissue-N analyses under the conditions used. As the biosensor and resulting assay are highly sensitive (<1 nM Gln), measurements of Gln concentrations require only leaf punches (Tessaro et al., 2012; Goron and Raizada, 2016), and in fact, in this study, single leaf punch extracts were diluted 1/100th prior to the assay. The leaf punch test is thus relatively non-destructive, and furthermore the space requirement to store samples is minimal. Hundreds of leaf punches can be assayed in a single day by one individual with minimal training, especially with automated bead-based grinding of leaf tissue. In our experience, leaf punches can be stored in -80°C for more than 1 year for later GlnLux analysis. All reagent costs are minimal, resulting in an assay cost of only ∼$1 USD per sample, though there is an upfront major equipment cost (for the luminometer). Finally, the assay does not require a non-N-fixing reference plant.
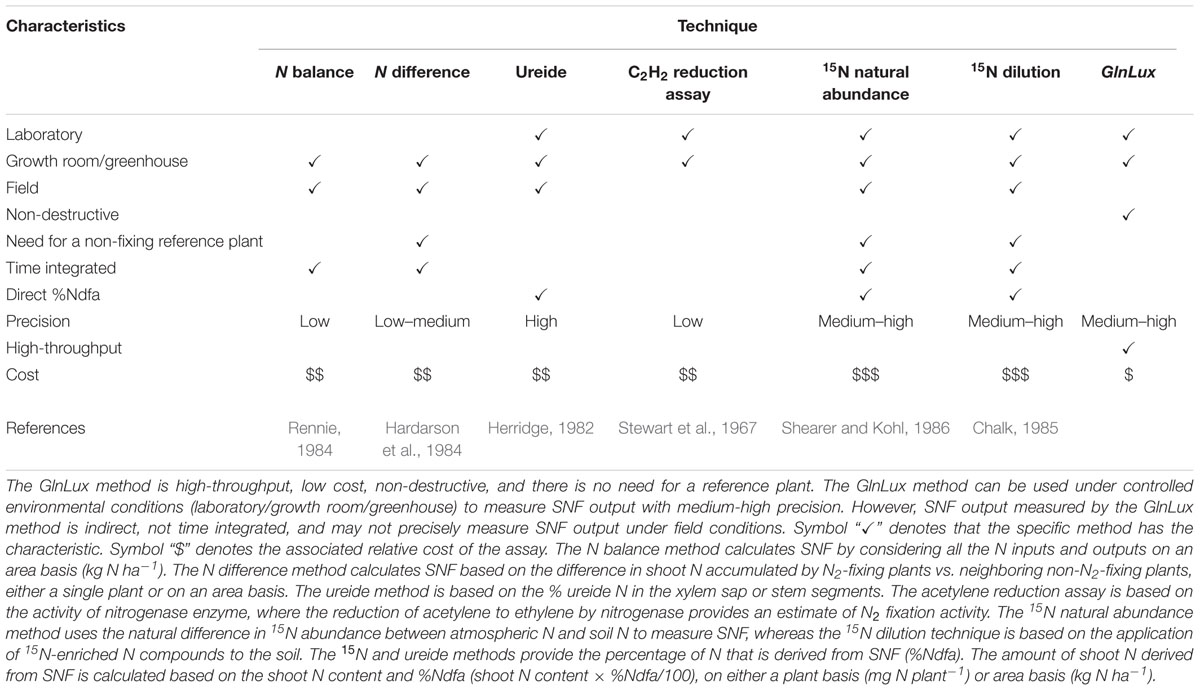
TABLE 1. Characteristics of methods to measure symbiotic nitrogen fixation (SNF) output (adapted from Howieson and Dilworth, 2016).
Fixed Nitrogen Assimilation and Translocation in Legumes
Nitrogen fixed by rhizobia inside legume nodules is secreted as ammonium into the host plant cell, where it rapidly assimilated into Gln (Atkins, 1987). Gln is further metabolized into asparagine (Asn) in amide exporting legumes or as the ureide compounds, allantoin and allantoic acid, in ureide exporting legumes. Amide exporting legumes then transfer the majority of their fixed N through xylem as Asn (80%) and Gln (10%), whereas in ureide exporting legumes, the major export compounds are allantoin and allantoic acid (90%) (Schubert, 1986; Unkovich et al., 2008; Aranjuelo et al., 2011; Howieson and Dilworth, 2016). In ureide exporting legumes, aside from the ureides, Gln and Asn are the major amino acids found in xylem sap, as well as minor proportions of aspartic acid, lysine, valine, leucine, and glutamic acid (Pate et al., 1980). Indeed, xylem Gln as an immediate product of N fixation was shown to be a useful indicator of SNF activity across various legumes (Amarante et al., 2006; Amarante and Sodek, 2006; Justino and Sodek, 2013). Xylem sap can be obtained either from the stump of intact roots following decapitation of the shoot (root bleeding sap), or from freshly harvested shoots by applying a mild vacuum (vacuum-extracted sap) (Unkovich et al., 2008). In the former method, root xylem is sampled, whereas in the latter, shoot xylem is sampled. Different amide exporting legumes were shown to transfer around 10% of fixed N as Gln in the absence of external N (i.e., when SNF dependent), and low genotypic variability was shown to exist for the concentration of Gln in xylem sap across legumes (e.g., comparing Pisum sativum, Lupinus albus, and Crotalaria juncea) (Amarante et al., 2006). Similarly, low genotypic variability was found for the Asn:Gln ratio among different ureide exporting legumes (Vigna angularis, G. max, V. unguiculata, Cyamopsis tetragonoloba, Macrotyloma uniflorum, Psophocarpus tetragonolobus, Vigna umbellata, Vigna mungo) when plants were grown without external N (Pate et al., 1980). Nevertheless, there is variation in the type and amount of organic N exported from nodules and transferred through xylem in different legumes (Udvardi and Poole, 2013). Furthermore, complex amino acid cycling takes place in legumes in order to maintain SNF while allowing the host plant to regulate the symbiotic process (Lodwig et al., 2003). Interestingly, SNF output measured using the GlnLux biosensor method was highly correlated to relative Gln output in both the amide and ureide exporting legumes tested. Therefore, the GlnLux results suggest that Gln can be a useful indicator of symbiotic N fixation across various legumes under controlled conditions.
Various Factors Can Affect the Glutamine Concentration in Legumes
In general, the N transport form in xylem sap changes based on soil and environmental factors (rainfall/water availability, soil nutrient availability, salinity) (Neo and Layzell, 1997; Cramer et al., 2002; Amarante et al., 2006; Bai et al., 2007; Ladrera et al., 2007; Gil-Quintana et al., 2012, 2013). The experiments in this study were undertaken under optimal, controlled conditions with minimal external N. It is important to note that external N (e.g., soil N) is also assimilated into Gln, which has the potential to confound interpretations of GlnLux data with respect to SNF. For example, in soybean, a ureide exporter, exposure to NH3 was shown to result in increased phloem Gln (Neo and Layzell, 1997) whereas another study showed similar xylem Gln levels when plants were supplied with NO3 (20 mM KNO3) compared to non-fertilized plants (McClure et al., 1979). A higher Asn:Gln ratio was observed in eight ureide exporters exposed to NO3, shown to be as a result of decreased Gln relative to Asn in cowpea (Pate et al., 1980). Xylem sap Gln was shown to change less significantly in amide exporting legumes in response to the source of N (SNF-dependent or external N) compared to ureide exporting legumes (Atkins et al., 1979; Amarante et al., 2006), likely because there is a larger relative background pool of Gln in the former.
In addition to external N, drought stress reduces SNF in legumes (Zahran, 1999; Giller, 2001; Kunert et al., 2016). Generally ureide exporting legumes (soybean, common bean) were shown to be more sensitive to drought stress compared to amide exporters (lentil, lupin) (Arrese-Igor et al., 2011). Therefore, the GlnLux assay may work more reliably with amide exporters than ureide exporters under drought stress conditions. Drought stress was shown to change Gln concentrations in amide exporters (Gil-Quintana et al., 2012) and ureide exporters (Silvente et al., 2012). The alteration in Gln concentration under drought stress varied by plant tissue (leaf/stem/root/nodule) (Ladrera et al., 2007; Gil-Quintana et al., 2012, 2013). It was found that under drought stress, leaf Gln content either decreased or remained close to the control (well-watered plants) in Medicago truncatula (amide exporter) (Gil-Quintana et al., 2012) and soybean (Silvente et al., 2012). Reallocation of leaf N compounds in legumes was shown to occur under drought conditions due to up-regulation of Gln synthetase (Aranjuelo et al., 2011). Further accumulation of total amino acids (Gil-Quintana et al., 2012) and proline during drought stress is a common scenario, where proline has been shown to act as an osmoregulant (Aranjuelo et al., 2011).
In addition, stress conditions that impair SNF can change the Gln concentration in xylem sap. For example, xylem Gln concentrations were shown to be reduced under water logging conditions in both amide and ureide exporting legumes (Amarante et al., 2006; Amarante and Sodek, 2006; Justino and Sodek, 2013). Interestingly, a similar pattern of reduction in xylem Gln was observed in both amide exporting (P. sativum, L. albus, C. juncea) and ureide exporting legumes (G. max, Phaseolus vulgaris, V. unguiculata) under water logging conditions similar to the reduction in SNF activity (Amarante et al., 2006). Therefore, in this study, Gln measurements were shown to be a good indicator of reductions in SNF under waterlogging stress.
Finally, Gln concentration can vary during the growth stage of legume plants. For example, Pate et al. (1980) found that the Asn:Gln ratio increased as cowpea and mung bean plants matured. However, the study did not show whether changes in the Asn:Gln ratio were due to decreased Gln or increased Asn.
Limitations and Future Applications
The GlnLux leaf punch assay does not provide a direct measurement of SNF (%Ndfa) compared to 15N-based methods (Table 1). Another disadvantage is that GlnLux reports only the relative amount of Gln, not absolute amount. However, a concentration gradient of pure Gln standards can be used along with the leaf punch assay (Supplementary Figure S1), which was highly replicable. Although we found very strong correlation between GlnLux and 15N based %Ndfa, the GlnLux method is a temporal snapshot and does not provide an integrated measurement of accumulated SNF output over time (Table 1). When collecting samples, leaf punches have to be flash-frozen in liquid N and kept frozen until processing, which limits its applicability to field conditions. Finally, as noted above, stress conditions such as drought can alter the leaf Gln concentration in legumes, which challenges the use of GlnLux to infer SNF under unpredictable conditions. Given this caveat, and since the leaf punch method cannot distinguish between N derived from SNF versus soil (Table 1), it may be most suited for pre-screening rhizobia inoculants under controlled N supply conditions prior to field experiments. We also expect that the GlnLux leaf punch method will assist with basic research especially to understand the unsolved mechanisms involved in the maintenance of SNF by facilitating high-throughput bacteria/plant mutant screens.
Author Contributions
Both MT and MR conceived of the manuscript. MT and MR designed the study and MT and NM conducted all experiments. MT wrote the manuscript and MR edited the manuscript. All authors discussed the results and commented on the manuscript. All authors read and approved the final manuscript.
Funding
This research was supported by CIFSRF grant 107791 to MR from the International Food Development Centre (IDRC, Ottawa) and Global Affairs Canada.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Special thanks to Krzysztof Szczyglowski (AAFC, London, ON, Canada) and Jeremy Murray (John Innes Centre, United Kingdom) for their valuable comments on the paper. We thank Jaclyn Clark and Caleb Niemeyer (University of Guelph) for their significant help in data collection, and Hanan Shehata and Travis Goron (University of Guelph) for training MT in the use of the GlnLux methodologies. We acknowledge Christopher Yost and Dinah Tambalo (University of Regina), Michael Hynes (University of Calgary), David Kuykendall (USDA), Timothy Porch (USDA), Papa Saliou (Kyoto University, Japan), and Takeo Yamakawa (Kyushu University, Japan) for providing rhizobia strains. We thank Kamal Khadka (LI-BIRD, Nepal) for providing seeds. We thank Lisa Smith for preparing graphic illustrations. We thank Myles Stocki (University of Saskatchewan) and Alice Chang (University of British Columbia) for 15N analysis.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2017.01714/full#supplementary-material
FIGURE S1 | Luminescence measurement of Gln standards using the GlnLux 96-well luminometer bioassay to demonstrate the linearity of the assay. Luminescence was measured using a concentration gradient of pure Gln standards (0, 125 × 10-8, 25 × 10-7, 5 × 10-6, and 1 × 10-5 M) using the GlnLux bioassay. A 0 μg/ml Gln standard reading was subtracted from all lux values, which were read in randomized replicates in an endpoint assay set to the “integrate” function. RLU, relative lux units. Error bars represent the standard error of the mean (SEM) (N = 3).
FIGURE S2 | Nodulation, root, and shoot morphological parameters of the lentil plants inoculated with different strains of rhizobia. (A) Lentil roots images. (B) Total root length (cm plant-1). (C) Total root surface area (cm2 plant-1). (D) Total root volume (cm3 plant-1). (E) Normalized total plant dry matter per plant. (F) Shoot/root ratio. Error bars represent the SEM (N = 8). The different letters on top of each histogram indicate significant differences in the mean between treatments.
FIGURE S3 | Nodulation, root, and shoot morphological parameters of the cowpea plants inoculated with different strains of rhizobia. (A) Cowpea roots images. (B) Total root length (cm plant-1). (C) Total root surface area (cm2 plant-1). (D) Total root volume (cm3 plant-1). (E) Normalized total plant dry matter per plant. (F) Shoot/root ratio. Error bars represent the SEM (N = 8). The different letters on top of each histogram indicate significant differences in the mean between treatments.
FIGURE S4 | Nodulation, root, and shoot morphological parameters of the soybean plants inoculated with different strains of rhizobia. (A) Soybean roots images. (B) Total root length (cm plant-1). (C) Total root surface area (cm2 plant-1). (D) Total root volume (cm3 plant-1). (E) Normalized total plant dry matter per plant. (F) Shoot/root ratio. Error bars represent the SEM (N = 8). The different letters on top of each histogram indicate significant differences in the mean between treatments.
References
Amarante, L., Lima, J. D., and Sodek, L. (2006). Growth and stress conditions cause similar changes in xylem amino acids for different legume species. Environ. Exp. Bot. 58, 123–129. doi: 10.1016/j.envexpbot.2005.07.002
Amarante, L., and Sodek, L. (2006). Waterlogging effect on xylem sap glutamine of nodulated soybean. Biol. Plant. 50, 405–410. doi: 10.1007/s10535-006-0057-6
Aranjuelo, I., Molero, G., Erice, G., Avice, J. C., and Nogue, S. (2011). Plant physiology and proteomics reveals the leaf response to drought in alfalfa (Medicago sativa L.). J. Exp. Bot. 62, 111–123. doi: 10.1093/jxb/erq249
Arrese-Igor, C., González, E. M., Marino, D., Ladrera, R., Larrainzar, E., and Gil-Quintana, E. (2011). Physiological responses of legume nodules to drought. Plant Stress 5, 24–31.
Atkins, C. A. (1987). Metabolism and translocation of fixed nitrogen in the nodulated legume. Plant Soil 169, 157–169. doi: 10.1016/j.jplph.2016.01.013
Atkins, C. A., Pate, J. S., and Layzell, D. B. (1979). Assimilation and transport of nitrogen in nonnodulated (NO3-grown) Lupinus albus L. Plant Physiol. 64, 1078–1082. doi: 10.1104/pp.64.6.1078
Baba, T., Ara, T., Hasegawa, M., Takai, Y., Okumura, Y., Baba, M., et al. (2006). Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008. doi: 10.1038/msb4100050
Bai, C., Reilly, C. C., and Wood, B. W. (2007). Identification and quantitation of asparagine and citrulline using high-performance liquid chromatography (HPLC). Anal. Chem. Insights 2, 31–36.
Bernard, S. M., and Habash, D. Z. (2009). The importance of cytosolic glutamine synthetase in nitrogen assimilation and recycling. New Phytol. 182, 608–620. doi: 10.1111/j.1469-8137.2009.02823.x
Betti, M., García-Calderón, M., Pérez-Delgado, C. M., Credali, A., Estivill, G., Galván, F., et al. (2012). Glutamine synthetase in legumes: recent advances in enzyme structure and functional genomics. Int. J. Mol. Sci. 13, 7994–8024. doi: 10.3390/ijms13077994
Bhagwat, A. A., Tully, R. E., and Keister, D. L. (1991). Isolation and characterization of a competition-defective Bradyrhizobium japonicum mutant. Appl. Environ. Microbiol. 57, 3496–3501.
Bourcy, M., Brocard, L., Pislariu, C. I., Cosson, V., Mergaert, P., Tadege, M., et al. (2013). Medicago truncatula DNF2 is a PI-PLC-XD-containing protein required for bacteroid persistence and prevention of nodule early senescence and defense-like reactions. New Phytol. 197, 1250–1261. doi: 10.1111/nph.12091
Broughton, W. J., Hern’andez, G., Blair, M., Beebe, S., Gepts, P., and Vanderleyden, J. (2003). Beans (Phaseolus spp.) - model food legumes. Plant Soil 252, 55–128. doi: 10.1023/A:1024146710611
Cabeza, R. A., Liese, R., Fischinger, S. A., Sulieman, S., Avenhaus, U., Lingner, A., et al. (2015). Long-term non-invasive and continuous measurements of legume nodule activity. Plant J. 81, 637–648. doi: 10.1111/tpj.12751
Chalk, P. M. (1985). Estimation of N2 fixation by isotope dilution: an appraisal of techniques involving 15N enrichment and their application. Soil Biol. Biochem. 17, 389–410. doi: 10.1016/0038-0717(85)90001-X
Cramer, V. A., Schmidt, S., Stewart, G. R., and Thorburn, P. J. (2002). Can the nitrogenous composition of xylem sap be used to assess salinity stress in Casuarina glauca? Tree Physiol. 22, 1019–1026.
Ehlers, J. D., and Hall, A. E. (1997). Cowpea (Vigna unguiculata L. Walp.). Field Crops Res. 53, 187–204. doi: 10.1016/S0378-4290(97)00031-2
Foyer, C. H., Lam, H., Nguyen, H. T., Siddique, K. H. M., Varshney, R. K., Colmer, T. D., et al. (2016). Neglecting legumes has compromised human health and sustainable food production. Nat. Plants 2:16112. doi: 10.1038/nplants.2016.112
Gil-Quintana, E., Larrainzar, E., Arrese-Igor, C., and González, E. M. (2012). Is N-feedback involved in the inhibition of nitrogen fixation in drought-stressed Medicago truncatula? J. Exp. Bot. 63, 281–292. doi: 10.1093/jxb/err313
Gil-Quintana, E., Larrainzar, E., Seminario, A., Díaz-Leal, J. L., Alamillo, J. M., Pineda, M., et al. (2013). Local inhibition of nitrogen fixation and nodule metabolism in drought-stressed soybean. J. Exp. Bot. 64, 2171–2182. doi: 10.1093/jxb/ert074
Goron, T. L., Nederend, J., Stewart, G., Deen, B., and Raizada, M. N. (2017). Mid-season leaf glutamine predicts end-season maize grain yield and nitrogen content in response to nitrogen fertilization under field conditions. Agronomy 7:41. doi: 10.3390/agronomy7020041
Goron, T. L., and Raizada, M. N. (2014). Current and future transgenic whole-cell biosensors for plant macro- and micronutrients. Crit. Rev. Plant Sci. 33, 392–413. doi: 10.1080/07352689.2014.885733
Goron, T. L., and Raizada, M. N. (2016). Biosensor-based spatial and developmental mapping of maize leaf glutamine at vein- level resolution in response to different nitrogen rates and uptake/assimilation durations. BMC Plant Biol. 16:230. doi: 10.1186/s12870-016-0918-x
Graham, P. H., and Vance, C. P. (2003). Legumes: importance and constraints to greater use. Plant Physiol. 131, 872–877. doi: 10.1104/pp.017004.872
Hardarson, G., Zapata, F., and Danso, S. K. A. (1984). Effect of plant genotype and nitrogen fertilizer on symbiotic nitrogen fixation by soybean cultivars. Plant Soil 82, 397–405. doi: 10.1007/BF02184277
Herridge, D. F. (1982). Relative abundance of ureides and nitrate in plant tissues of soybean as a quantitative assay of nitrogen fixation. Plant Physiol. 70, 1–6. doi: 10.1104/pp.70.1.1
Herridge, D. F., and Peoples, M. B. (1990). Ureide assay for measuring nitrogen fixation by nodulated soybean calibrated by 15N methods. Plant Physiol. 93, 495–503. doi: 10.1104/pp.93.2.495
Herridge, D. F., Peoples, M. B., and Boddey, R. M. (2008). Global inputs of biological nitrogen fixation in agricultural systems. Plant Soil 311, 1–18. doi: 10.1007/s11104-008-9668-3
Howieson, J. G., and Dilworth, M. J. (2016). Working with Rhizobia. Bruce, ACT: Australian Centre for International Agricultural Research.
Justino, G. C., and Sodek, L. (2013). Recovery of nitrogen fixation after short-term flooding of the nodulated root system of soybean. J. Plant Physiol. 170, 235–241. doi: 10.1016/j.jplph.2012.10.006
Kaneko, T., Nakamura, Y., Sato, S., Minamisawa, K., Uchiumi, T., Sasamoto, S., et al. (2002). Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res. 9, 189–197. doi: 10.1093/dnares/9.6.189
Kiers, E. T., Rousseau, R. A., West, S. A., and Denison, R. F. (2003). Host sanctions and the legume–rhizobium mutualism. Nature 425, 1095–1098. doi: 10.1038/nature01931
Kunert, K. J., Vorster, B. J., Fenta, B. A., Kibido, T., Dionisio, G., and Foyer, C. H. (2016). Drought stress responses in soybean roots and nodules. Front. Plant Sci. 7:1015. doi: 10.3389/fpls.2016.01015
Kuykendall, L. D., Saxena, B., Devine, T. E., and Udell, S. E. (1992). Genetic diversity in Bradyrhizobium japonicum Jordan 1982 and a proposal for Bradyrhizobium elkanii sp.nov. Can. J. Microbiol. 38, 501–505. doi: 10.1139/m92-082
Ladrera, R., Marino, D., Larrainzar, E., González, E. M., and Arrese-Igor, C. (2007). Reduced carbon availability to bacteroids and elevated ureides in nodules, but not in shoots, are involved in the nitrogen fixation response to early drought in soybean. Plant Physiol. 145, 539–546. doi: 10.1104/pp.107.102491
Leigh, G. J. (2004). The World’s Greatest Fix: A History of Nitrogen and Agriculture. Oxford: Oxford University Press.
Lodwig, E. M., Hosie, A. H. F., Bourde‘s, A., Findlay, K., Allaway, D., Karunakaran, R., et al. (2003). Amino-acid cycling drives nitrogen fixation in the legume – Rhizobium symbiosis. Nature 422, 722–726. doi: 10.1038/nature01549.1
McClure, P. R., Israel, D. W., and Carolina, N. (1979). Transport of nitrogen in the xylem of soybean Plants. Plant Physiol. 64, 411–416. doi: 10.1104/pp.64.3.411
Meighen, E. A., and Szittner, R. B. (1992). Multiple repetitive elements and organization of the lux operons of luminescent terrestrial bacteria. J. Bacteriol. 174, 5371–5381. doi: 10.1128/jb.174.16.5371-5381.1992
Neo, H. H., and Layzell, D. B. (1997). Phloem glutamine and the regulation of O2 diffusion in legume nodules. Plant Physiol. 113, 259–267. doi: 10.1104/pp.113.1.259
Pate, J. S., Atkins, C. A., White, S. T., Rainbird, R. M., and Woo, K. C. (1980). Nitrogen nutrition and xylem transport of nitrogen in ureide- producing grain legumes. Plant Physiol. 65, 961–965. doi: 10.1104/pp.65.5.961
Prell, J., and Poole, P. (2006). Metabolic changes of rhizobia in legume nodules. Trends Microbiol. 14, 161–168. doi: 10.1016/j.tim.2006.02.005
Rennie, R. J. (1984). Comparison of N balance and 15N isotope dilution to quantify N2 fixation in field-grown legumes. Agronomy 76, 785–790. doi: 10.2134/agronj1984.00021962007600050018x
Sadowsky, M. J., Tully, R. E., Cregan, P. B., and Keyser, H. H. (1987). Genetic diversity in Bradyrhizobium japonicum serogroup 123 and its relation to genotype-specific nodulation of soybean. Appl. Environ. Microbiol. 53, 2624–2630.
Sarr, P. S., Yamakawa, T., Fujimoto, S., Saeki, Y., Thao, H. T., and Myint, A. K. (2009). Phylogenetic diversity and symbiotic effectiveness of root-nodulating bacteria associated with cowpea in the south-west area of Japan. Microbes Environ. 24, 105–112. doi: 10.1264/jsme2.ME08558
Schubert, K. R. (1986). Products of biological nitrogen fixation in higher plants: synthesis, transport, and metabolism. Annu. Rev. Plant Physiol. 37, 539–574. doi: 10.1146/annurev.pp.37.060186.002543
Sharpe, A. G., Ramsay, L., Sanderson, L., Fedoruk, M. J., Clarke, W. E., Li, R., et al. (2013). Ancient orphan crop joins modern era: gene-based SNP discovery and mapping in lentil. BMC Genomics 14:192. doi: 10.1186/1471-2164-14-192
Shearer, G., and Kohl, D. H. (1986). N2-fixation in field settings: estimations based on natural 15N abundance. Funct. Plant Biol. 13, 699–756. doi: 10.1071/PP9860699
Silvente, S., Sobolev, A. P., and Lara, M. (2012). Metabolite adjustments in drought tolerant and sensitive soybean genotypes in response to water stress. PLOS ONE 7:e38554. doi: 10.1371/journal.pone.0038554
Starker, C. G., Parra-Colmenares, A. L., Smith, L., Mitra, R. M., and Long, S. R. (2006). Nitrogen fixation mutants of Medicago truncatula fail to support plant and bacterial symbiotic. Plant Physiol. 140, 671–680. doi: 10.1104/pp.105.072132.1
Stewart, W. D., Fitzgerald, G. P., and Burris, R. H. (1967). In situ studies on N2 fixation using the acetylene reduction technique. Proc. Natl. Acad. Sci. U.S.A. 58, 2071–2078. doi: 10.1073/pnas.58.5.2071
Tegeder, M. (2014). Transporters involved in source to sink partitioning of amino acids and ureides: opportunities for crop improvement. J. Exp. Bot. 65, 1865–1878. doi: 10.1093/jxb/eru012
Tessaro, M. J., Soliman, S. S. M., and Raizada, M. N. (2012). Bacterial whole-cell biosensor for glutamine with applications for quantifying and visualizing glutamine in plants. Appl. Environ. Microbiol. 78, 604–606. doi: 10.1128/AEM.05261-11
Thilakarathna, M. S., Mcelroy, M. S., Chapagain, T., Papadopoulos, Y. A., and Raizada, M. N. (2016a). Belowground nitrogen transfer from legumes to non-legumes under managed herbaceous cropping systems. A review. Agron. Sustain. Dev. 36:58. doi: 10.1007/s13593-016-0396-4
Thilakarathna, M. S., Papadopoulos, Y. A., Rodd, A. V., Grimmett, M., Fillmore, S. A. E., Crouse, M., et al. (2016b). Nitrogen fixation and transfer of red clover genotypes under legume–grass forage based production systems. Nutr. Cycl. Agroecosyst. 106, 233–247. doi: 10.1007/s10705-016-9802-1
Thilakarathna, M. S., and Raizada, M. N. (2017). A meta-analysis of the effectiveness of diverse rhizobia inoculants on soybean traits under field conditions. Soil Biol. Biochem. 105, 177–196. doi: 10.1016/j.soilbio.2016.11.022
Thilakarathna, M. S., Serran, S., Lauzon, J., Janovicek, K., and Deen, B. (2015). Management of manure nitrogen using cover crops. Agronomy 107, 1595–1607. doi: 10.2134/agronj14.0634
Thilakarathna, R. M. M. S., Papadopoulos, Y. A., Rodd, A. V., Gunawardena, A. N., Fillmore, S. A. E., and Prithiviraj, B. (2012). Characterizing nitrogen transfer from red clover populations to companion bluegrass under field conditions. Can. J. Plant Sci. 92, 1163–1173. doi: 10.4141/cjps2012-036
Udvardi, M., and Poole, P. S. (2013). Transport and metabolism in lgume-rhizobia symbioses. Annu. Rev. Plant Biol. 64, 781–805. doi: 10.1146/annurev-arplant-050312-120235
Unkovich, M., Herridge, D., Peoples, M., Cadisch, G., Boddey, B., Giller, K., et al. (2008). Measuring Plant-Associated Nitrogen Fixation in Agricultural Systems. Bruce, ACT: Australian Centre for International Agricultural Research.
Unkovich, M. J., and Pate, J. S. (2000). An appraisal of recent field measurements of symbiotic N2 fixation by annual legumes. Field Crops Res. 65, 211–228. doi: 10.1016/S0378-4290(99)00088-X
Vanderlinde, E. M., Magnus, S. A., Tambalo, D. D., Koval, S. F., and Yost, C. K. (2011). Mutation of a broadly conserved operon (RL3499-RL3502) from Rhizobium leguminosarum biovar viciae causes defects in cell morphology and envelope integrity. J. Bacteriol. 193, 2684–2694. doi: 10.1128/JB.01456-10
Yost, C. K., Rath, A. M., Noel, T. C., and Hynes, M. F. (2006). Characterization of genes involved in erythritol catabolism in Rhizobium leguminosarum bv. viciae. Microbiology 152, 2061–2074. doi: 10.1099/mic.0.28938-0
Keywords: legume, symbiotic nitrogen fixation, nodule, rhizobia, biosensor, glutamine
Citation: Thilakarathna MS, Moroz N and Raizada MN (2017) A Biosensor-Based Leaf Punch Assay for Glutamine Correlates to Symbiotic Nitrogen Fixation Measurements in Legumes to Permit Rapid Screening of Rhizobia Inoculants under Controlled Conditions. Front. Plant Sci. 8:1714. doi: 10.3389/fpls.2017.01714
Received: 11 June 2017; Accepted: 20 September 2017;
Published: 09 October 2017.
Edited by:
Benjamin Gourion, UMR2594 Laboratoire des Interactions Plantes Microorganismes (LIPM), FranceReviewed by:
Virginie Bourion, INRA Centre Dijon Bourgogne Franche-Comté, FranceKatharina Pawlowski, Stockholm University, Sweden
Copyright © 2017 Thilakarathna, Moroz and Raizada. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Manish N. Raizada, cmFpemFkYUB1b2d1ZWxwaC5jYQ==
 Malinda S. Thilakarathna
Malinda S. Thilakarathna Nicholas Moroz
Nicholas Moroz Manish N. Raizada
Manish N. Raizada