- 1Australian Export Grains Innovation Centre, South Perth, WA, Australia
- 2Western Barley Genetics Alliance, Murdoch University, Murdoch, WA, Australia
Barley is an important crop for the production of malt and beer. However, crops such as rice and wheat are rarely used for malting. α-amylase is the key enzyme that degrades starch during malting. In this study, we compared the genomic properties, gene copies, and conserved promoter motifs of α-amylase genes in barley, rice, and wheat. In all three crops, α-amylase consists of four subfamilies designated amy1, amy2, amy3, and amy4. In wheat and barley, members of amy1 and amy2 genes are localized on chromosomes 6 and 7, respectively. In rice, members of amy1 genes are found on chromosomes 1 and 2, and amy2 genes on chromosome 6. The barley genome has six amy1 members and three amy2 members. The wheat B genome contains four amy1 members and three amy2 members, while the rice genome has three amy1 members and one amy2 member. The B genome has mostly amy1 and amy2 members among the three wheat genomes. Amy1 promoters from all three crop genomes contain a GA-responsive complex consisting of a GA-responsive element (CAATAAA), pyrimidine box (CCTTTT) and TATCCAT/C box. This study has shown that amy1 and amy2 from both wheat and barley have similar genomic properties, including exon/intron structures and GA-responsive elements on promoters, but these differ in rice. Like barley, wheat should have sufficient amy activity to degrade starch completely during malting. Other factors, such as high protein with haze issues and the lack of husk causing Lauting difficulty, may limit the use of wheat for brewing.
Introduction
The best quality barley grains are used predominantly for making malts and subsequently beer and whiskey. Malting consists of steeping, germination, and kilning (Gupta et al., 2010). Steeping and germination allow production of hydrolyzing enzymes including α-amylase (amy), β-amylase, limit dextrinase, and α-glucosidase for starch degradation (Bak-Jensen et al., 2007; Evans et al., 2010; Fincher, 2010; Shahpiri et al., 2015). Starch comprises an α-D-glucose homo-polymer amylose and branched amylopectin. The former is a linear molecule of α-1,4-linked glucose molecules, while the latter is a larger molecule with α-1,6 branching points (Bahaji et al., 2014). Amy [α-(1,4)-D-glucan glucanohydrolase, EC 3.2.1.1] cleaves α-(1,4) glycosidic linkage internally to produce oligosaccharides and amylopectin. Amy is the most important enzyme for starch degradation during malting and mashing. Barley malts contain sufficient active amy enzymes to almost completely hydrolyze starch during malting and mashing.
Activation of amy expression is strictly controlled by the phytohormones gibberellin and ABA. During grain development, amy expression is repressed by ABA. However, in a genetic defect wheat, a high level of high pI amy genes could be expressed, resulting in poor grain quality during late grain development. This is normally referred to as late maturity α-amylase (LMA) (Barrero et al., 2013). During seed germination, amy expression is induced by elevated GA levels (Lanahan et al., 1992; Gómez-Cadenas et al., 2001; Woodger et al., 2010).
Genetic mapping associated barley malt amy activities with amy1 and amy2 loci on chromosomes 6H and 7H, respectively (Hayes et al., 1993; Oziel et al., 1996; Zale et al., 2000; Gao et al., 2004; Li et al., 2010). Isoelectric focusing electrophoresis identified low and high pI amy isoforms in barley aleurone extracts (Jacobsen and Higgins, 1982; Svensson et al., 1985). However, the number of amy isoforms in the barley genome is unknown but predicted to be from three to eight (Jacobsen and Higgins, 1982; Muthukrishnan et al., 1984; Svensson et al., 1985; Khursheed and Rogers, 1988; Evans et al., 2010). Nomenclatures of amys are complicated. Two families of amy, were named AMY1 and AMY2, referred to low and high pI enzymes, respectively (MacGregor et al., 1971; Jacobsen and Higgins, 1982; Svensson et al., 1985; Evans et al., 2010). A genomic clone and two cDNA clones coding for amy enzymes have been named amy32b, amy6-4, and amy46 (Rogers and Milliman, 1983; Whittier et al., 1987; Khursheed and Rogers, 1988). In a recently published barley genomic sequencing paper, new amy nomenclatures have been proposed. The barley genome contains at least 12 amy genes, grouped into four subfamilies amy1, amy2, amy3, and amy4 (Mascher et al., 2017). Here, we compared gene copy numbers, genomic structures and promoter conserved motifs of amy1 and amy2 subfamilies from barley, wheat, and rice. We hypothesize that the expansion in amy1 members combined with the presence of conserved regulatory motifs on promoters of amy1 and amy2 genes are important determinants for selecting barley as a malting crop.
Materials and Methods
Genome sequences were downloaded to a local computer from ftp://ftp.ensemblgenomes.org/pub/plants/release-35/fasta/hordeum_vulgare/dna/ for barley; ftp://ftp.ensemblgenomes.org/pub/plants/release-35/fasta/oryza_sativa/dna/ for rice, and ftp://ftp.ensemblgenomes.org/pub/plants/release-35/fasta/triticum_aestivum/dna/ for wheat. The identification of barley, wheat, and rice amy genes are described in Mascher et al. (2017). Amy coding and promoter sequences (500 bp upstream of the translation start codon ATG) for all three crops were extracted after being blasted with the amy genes. Briefly, the amy genes were used to blast standalone blastable genomic databases to obtain amy gene nucleotide positions in pseudomolecules. According to these positions, the amy gene coding and promoter sequences were calculated and extracted with a Perl script. The promoter sequences were aligned with a ClustalW program1 and conserved motifs were examined.
Results and Discussion
Barley amy Copy Numbers – Historical and Genomic Evidence
Barley amy genes were initially mapped to chromosomes 6H and 7H with wheat–barley addition lines (Brown and Jacobsen, 1982; Muthukrishnan et al., 1984). Southern blot analysis of two different amy gene DNA probes detected at least six and three hybridization bands from addition lines containing barley chromosomes 6H and 7H, respectively (Muthukrishnan et al., 1984; Rogers and Milliman, 1984). There were multiple amy protein bands on SDS PAGE purified from the barley aleurone using cycloheptaamylose-sepharose affinity chromatography and at least four amy activity peaks separated by DEAE cellulose chromatography (Jacobsen and Higgins, 1982). These offered early experimental evidence of the amy multigene family. Isoelectric focusing (IEF) electrophoresis showed that purified amy proteins could be divided into two distinct groups, a low pI group with an isoelectric point of 4.5–5.1 and a high pI group with an isoelectric point of 5.0–6.6 (Jacobsen and Higgins, 1982). Due to its commercial and biological importance, a significant effort was made to clone amy genes. A genomic clone was identified as an amy gene (amy32b) and belongs to a low pI amy protein (Rogers and Milliman, 1983; Whittier et al., 1987). Two cDNA clones were also characterized as amy genes (amy6_4 and amy46) that belong to high pI enzymes (Khursheed and Rogers, 1988). Furthermore, 3D structures have been resolved for two barley amy proteins; one belonging to a low pI amy protein (1AMY) and the other to a high pI amy protein (1HT6) (Kadziola et al., 1994; Robert et al., 2003).
While experimental data has shown that amy proteins are coded by multigene families, the exact numbers of genes are unknown. Barley genome sequencing is a useful resource for identifying the number of amy genes and discovering their genomic features. The barley genome contained 12 amy genes (Mascher et al., 2017), which were grouped into four subfamilies (Table 1). Subfamily 1 consists of six members—four on chromosome 6H (533880485–542858990 bp) and two on the unsorted chromosome (195047130–196261798 bp, Table 1)—designated amy1_1a to amy1_1e and amy1_2. Four of which (amy1_1a to amy1_1d) have almost 100% sequence identity among members (Additional File 1: Supplementary Table S1A). One member (amy1_1e) is a truncated protein missing the carbohydrate-binding domain. Sequence identity analysis showed that the five amy1_1 proteins matched the cloned gene amy6_4 (Khursheed and Rogers, 1988), while the amy1_2 protein, with 95% sequence identity with amy1_1a, matched the cloned gene amy46 in both promoter and coding regions (Khursheed and Rogers, 1988). All amy1 members belong to high pI enzymes (Table 1) and have high sequence identity with a 3D structure-resolved protein 1AMY (Kadziola et al., 1994). Subfamily amy2 has three members on chromosome 7H (556169683–557427021 bp, Table 1), and are designated amy2_1 to amy2_3. They have >92% sequence identity among the members, and >72% when compared to amy1_1a. Amy2_3 had a high sequence identity with cloned gene amy32b (Rogers, 1985; Whittier et al., 1987) and 3D structure-resolved protein 1HT6 (Robert et al., 2003). They belong to genes coding for low pI enzymes (Table 1). Amy3 has one member localized on chromosome 5H (designated amy3), while amy4 has two members localized on chromosomes 2H and 3H (designated amy4_1 and amy4_2). The amy4 members have about 48% sequence identities compared between the members, or 43–46% sequence identity when compared to amy1_1a (Supplementary Table S1A).
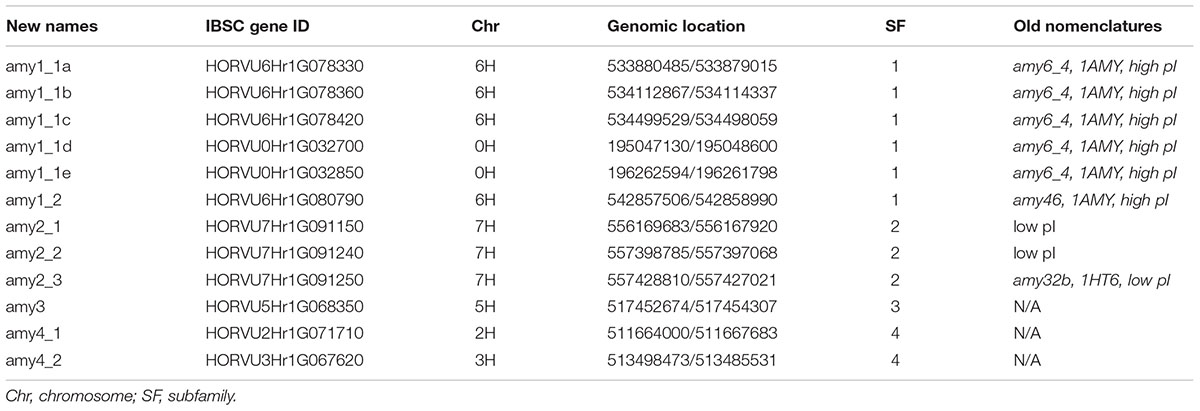
TABLE 1. Barley amy nomenclatures, gene ID, locations, and the association of old and new nomenclatures.
Since amy1 and amy2 were located on chromosomes 6H and 7H, respectively, and many important malt quality QTLs were associated with these genetic loci, we believed that they were the most important members in relation to barley malt qualities (Hayes et al., 1993; Oziel et al., 1996; Zale et al., 2000; Gao et al., 2004; Li et al., 2010), we decided to investigate and compare gene and promoter structures for these two subfamilies in barley, rice, and wheat.
Barley amy Protein Secondary Structure
All amy proteins from amy1 and amy2 had the catalytic amino acid residues Asp203, Glu228, and Asp310 (amy1_1a positions), apart from amy1_1e, which was a truncated protein missing Asp310 (Figure 1). The near full-length proteins consisted of a central domain A forming (α/β)8 barrel, a structural loop domain B and a carbohydrate-binding domain C (Figure 1). Domain C formed five anti-parallel sheets (Kadziola et al., 1994; Robert et al., 2003). Barley amy proteins from amy3 and amy4 also contain the catalytic amino acids and a carbohydrate-binding module as discovered in the Domain Database at NCBI2. However, Asp310 on the active site was replaced with Glu310 for the two amy4 proteins (Figure 1).
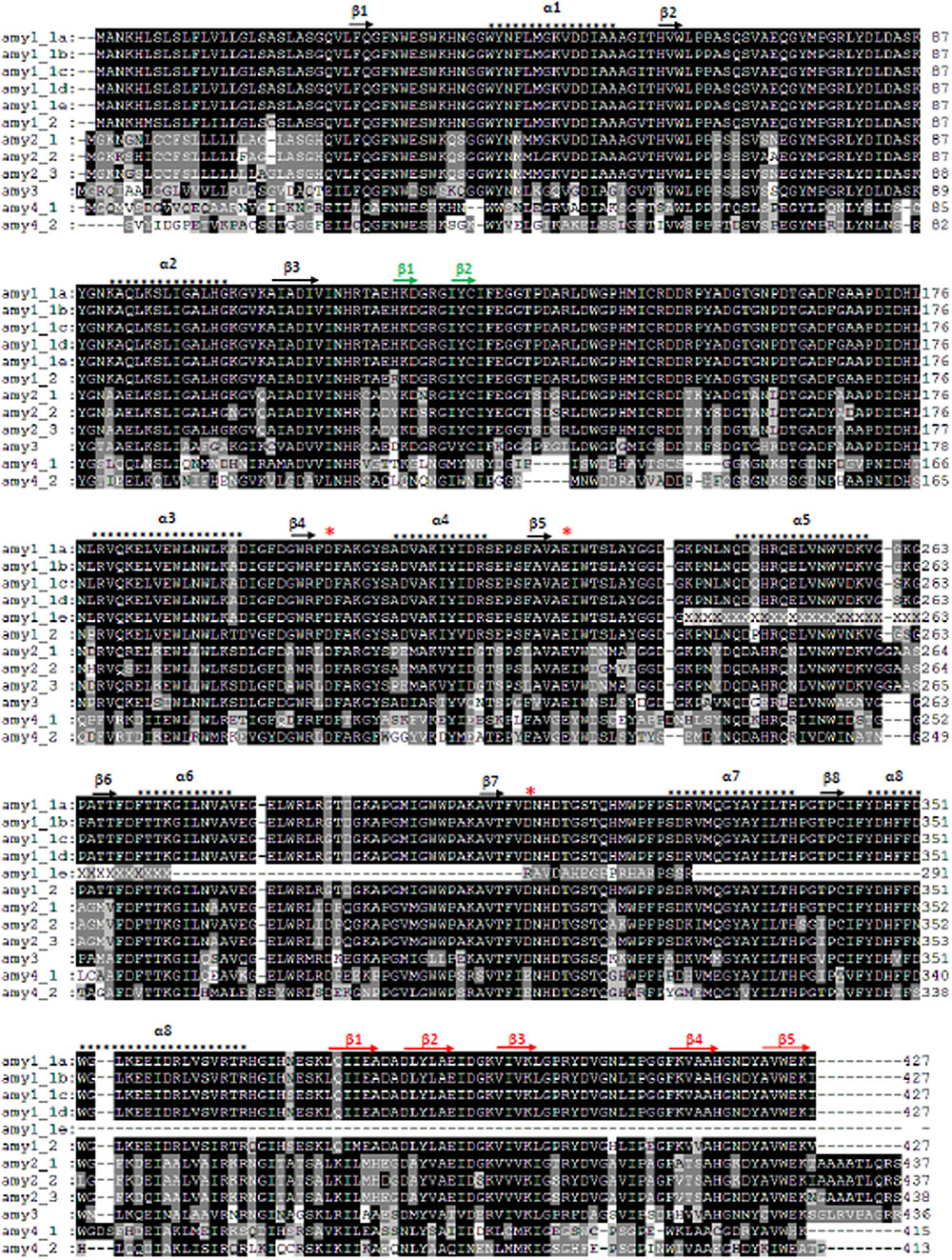

FIGURE 1. Alignments and secondary structures of barley amy proteins. Barley amy1_1a and amy2_3 had 100% sequence identities with the two 3D structure-resolved barley amy proteins (1AMY and 1HT6), respectively (Kadziola et al., 1994; Robert et al., 2003). Their secondary structure features are shown on top of the alignments. There are three domains: domain A (black arrows and asterisks), domain B (green arrows), and domain C (red arrows). Domain A consists of a (α/β)8 barrel, while domain C has five β-sheets. The three catalytic amino acids Asp203, Glu228, and Asp310 are indicated by red asterisks (amy1_1a position).
Barley amy Gene Genomic Arrangement
Four barley amy1 genes (amy1_1a to amy1_1d) had the same genomic arrangements as two introns and three exons. The nucleotide numbers for the introns and exons were the same as amy1_1a to amy1_1d, being 23 and 107 bp for the introns and 87, 1002, and 252 bp for the exons (Supplementary Table S2). While amy1_2 had two introns and three exons, the nucleotide numbers differed from the amy1_1a to amy1_1d members, being 95 and 106 bp for the introns and 87, 945, and 252 bp for the exons. Amy2-1 to amy2_3 had three introns and four exons, but the nucleotide numbers for all introns and exons differed among the three amy genes (Supplementary Table S2). Amy3 had three introns and four exons like amy2, but the nucleotide numbers differed from amy2. Amy4 had more than five introns and six exons.
Barley amy Gene Promoter Conserved Motifs
Promoters of all barley amy1 genes contained a conserved GA response complex (GARC) consisting of GARE (TAACAAA), pyrimidine (CCTTTT) and TATCCAC(T) boxes (Supplementary Table S3A and Figure 2) (Skriver et al., 1991; Gubler and Jacobsen, 1992; Rogers et al., 1994). There was also a cAMP-like response element (TGAGCTC) on amy1 promoters (Gubler and Jacobsen, 1992), which represses gibberellin action. Pyrimidine and TATCCAC boxes enhanced the expression of amy1 proteins. The conserved motifs on subfamily 2 members differed from those on subfamily 1 genes and also among subfamily 2 members (Supplementary Table S3B and Figure 3). All three amy2 genes had GARE (TAACAGAG) and pyrimidine (CCTTTT) boxes. The pyrimidine box was much closer to the translation start site (-17 bp) for amy2_2, but further away for amy2_1, and amy2_3 at -211 and -236 bp, respectively. The original pyrimidine box of the amy2_2 gene, at a similar position to amy2_1, and amy2_3, was mutated to CCATTT on amy2_2 (Figure 3). A TATCCAT box was found in two amy2 genes (amy2_1 and amy2_3), but it was replaced with TACCCAT in the amy2_2 gene. Furthermore, the amy2_2 promoter had a conserved O2S box (CTTGxxTCATC) and cAMP-like box (TGAGCTC). Genomic sequence analysis showed that both amy1 and amy2 genes had a GARC, where GARE was required for GA induction of amy expression with pyrimidine and TA(T/C)CCAT box controlling gene expression levels (Lanahan et al., 1992).

FIGURE 2. Barley amy1 promoter sequence alignments and conserved motifs. ALL amy1 promoters (except amy1_1e, which is truncated) contain a GA-responsive element (GARE) TAACAAA (red box). It requires for GA induction, They also contain a pyrimidine box (CCTTTT) and a TATCCA(C/T) box (blue boxes), which enhance gene expression after GA responses (Gubler and Jacobsen, 1992). A cAMP-like responsive element (TGAGCTC) is conserved (green box).
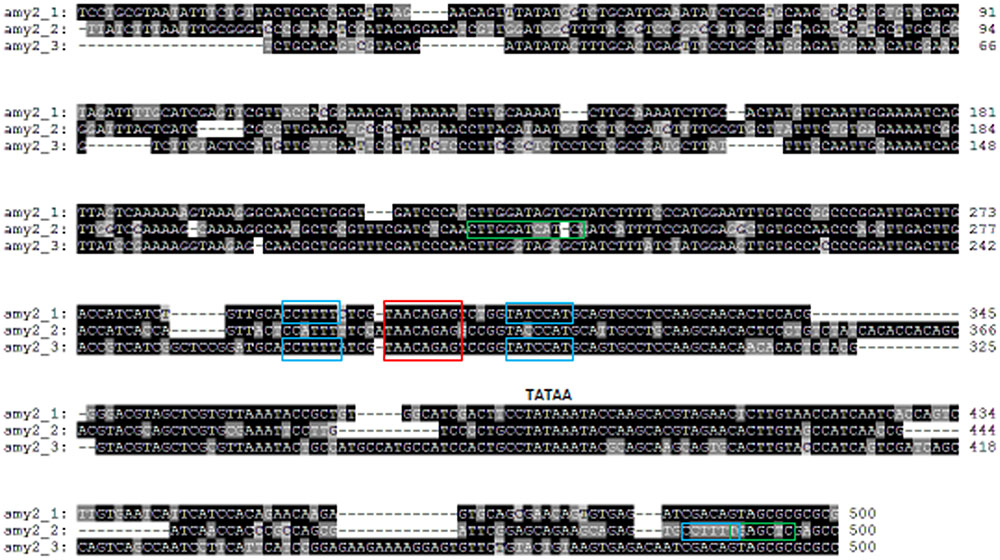
FIGURE 3. Barley amy2 promoter sequence alignments and conserved motif analysis. All three amy2 promoters contain a GA-responsive element (GARE) TAACAGAG (red box) required for GA induction. Promoters of amy2_1 and amy2_3 contained a pyrimidine box (CCTTTT) and a TATCCA(C/T) box (blue boxes), which enhance gene expression (Gubler and Jacobsen, 1992). A pyrimidine box (blue box), a cAMP-like responsive element (TGAGCTC) (green box) and an O2S (CTTGXXTCATC) (green box) were present on amy2_2 promoter.
When promoter regions (-500 bp) of subfamilies 1 and 2 were analyzed, the sequence identities were high (>99%) among amy1_1a, amy1_1b, and amy1_1d (Supplementary Table S4A). However, the amy1_1e promoter region was truncated to -151 bp, despite being 100% identical to the promoter sequence of amy1_1a gene. The promoter region of amy1_1c had high sequence identity within -350 bp, but low sequence identity beyond -350 bp, compared to amy1_1a (Figure 2). The sequence of the amy1_2 promoter (-500 bp) was 64% identical to the amy1_1a promoter. The genomic locations of the conserved motifs in amy1 were the same for all amy1_1 members at -180, -199, -220, and -225 bp for TATCCA, GARE, cAMY-like and pyrimidine boxes, respectively (Supplementary Table S3A). However, the locations of these motifs from the amy1_2 gene was a nucleotide closer to the ATG translation start site compared to the locations of amy1_1 members (Supplementary Table S3A). The sequences of amy2 promoters had low sequence identity (59–68%) compared to their members (Supplementary Table S4B).
Rice amy Gene Numbers and Conserved Motifs on Promoters
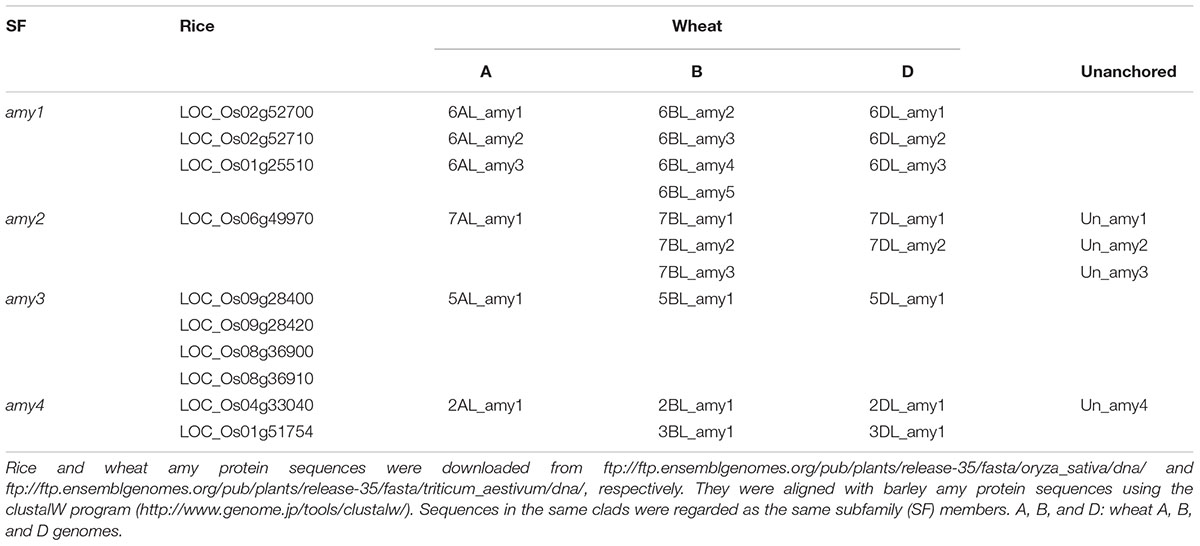
The rice genome contained 10 amy genes with three, one, four, and two members in subfamilies 1, 2, 3, and 4, respectively (Table 2). The number of amy1 and amy2 genes in rice was four, much less than the sum of amy1 and amy2 in barley. Rice amy3 had the most gene members. Two of the amy1 genes (LOC_Os02g52700 and LOC_OS02g52710) had three introns (Supplementary Table S2), unlike the barley amy1 genes, which all had two introns. The other amy1 gene (LOC_Os01g25510) had two introns. Alignment of the three amy1 protein sequences identified one protein (LOC_Os01g25510) with a very low sequence identity (24–30%) compared to the other two amy proteins (LOC_Os02g52700 and LOC_Os02g52710) (Supplementary Table S1B and Figure S1). The two amy1 proteins and one amy2 protein had similar secondary structures to the barley amy proteins. However, the amino acid compositions differed substantially on most of the β-strains and α-helices (Supplementary Figure S1). Rice had the same catalytic amino acids (Aps, Glu, and Asp) as barley. The promoters of the three rice amy1 genes had >65% sequence identities (Supplementary Table S4C). They contained the GARE (TAACAAA), pyrimidine (CCTTTT) and TATCCAT boxes (Supplementary Table S3C and Figure S2A) but not the cAMY-like box. The rice amy2 protein (LOC_Os06g49970) had 72% sequence identity compared to the two amy1 proteins (LOC_OS02g527100 and LOC_Os02g52710). However, the promoter of the rice amy2 gene only contained GARE (TAACAGAG), but not pyrimidine, TATCCAT or TATCCAC boxes (Supplementary Table S3C and Figure S2B).
Wheat amy Gene Numbers and Conserved Motifs on Promoters
The number of amy genes in the wheat A, B, and D genomes was 6, 10, and 8, respectively. The other four amy genes are located in unsorted chromosomes (Table 2). The wheat B genome had the most amy genes with four, three, one and two members in subfamilies amy1, amy2, amy3, and amy4, respectively. The number of amy1 and amy2 genes in each wheat genome did not exceed those in the barley genome (Table 2). All of the wheat amy2 genes had the same genomic arrangement as the barley amy2 genes with three introns and four exons (Supplementary Table S2). Most of the wheat amy1 genes had two introns and three exons except for 6BL4, Un1 and Un2, which had three or four introns. The protein sequence identities were high within the wheat amy1 or amy2, being >80% (Supplementary Tables S1C,D and Figures S3, S4). The sequence identities for promoters of amy1 genes were 50–100%, but much lower for amy2 gene promoters (Supplementary Tables S4D,E). The promoters of all wheat amy1 genes contained GARE (TAACAAA), pyrimidine, TATCCAT or TATCCAC boxes (Supplementary Figure S5). They also had a cAMP-like motif (TGAGCTC) box as per the barley amy1 gene promoters. Five of the wheat amy2 gene promoters contained a GARE (TAACAGAG) box, six contained pyrimidine and TATCCAT boxes, and seven had O2S motifs. The O2S motifs in the wheat genomes contained four variable nucleotides between the conserved CTTC and TCATC (Supplementary Figure S6), while the O2S in the barley amy2 promoters had two variable nucleotides (Figure 3).
A Comparison of amy Gene Copy Numbers and Sequence Properties from Barley, Rice, and Wheat
Barley had the highest number of amy1 genes (six), while wheat had four in the B genome and rice had three (Table 2). Both barley and wheat had the same number of amy2 genes (B genome only), while rice had one. Rice contained the most amy3 genes (four), while barley and wheat had one each. All the barley, wheat and rice genomes contained two amy4 genes (Table 2). The intron numbers for amy2 genes were the same for barley, wheat and rice, but differed for the amy1 genes: barley had two, wheat had two or four, and rice had two or three (Supplementary Table S2). The intron numbers for amy3 and amy4 genes differed, ranging from two to nine (Supplementary Table S2). Barley amy1 genes had high sequence identities with wheat amy1 genes ranging from 83 to 97% (Supplementary Table S5). Barley amy2 genes also had high sequence identities with wheat amy2 genes (80–96%) (Supplementary Table S5). The barley and wheat amy1 and amy2 genes had similar promoter regions with sequence identities ranging from 50 to 76% (Supplementary Table S6). However, there was no similarity between barley and rice promoter sequences.
Expanded amy1 and amy2 Genes Are Important for Barley Malting Qualities
Barley amy proteins are grouped into four subfamilies according to their sequence properties (Mascher et al., 2017). The biological functions for each subfamily are unclear. Genetic mapping using molecular marker technologies showed that the regions associated with the genetic markers amy1 and amy2 on chromosomes 6 and 7 are important for malting qualities including amy enzyme activities and malt extracts (Hayes et al., 1993; Han and Ullrich, 1994; Oziel et al., 1996; Marquez-Cedillo et al., 2000; Emebiri et al., 2004). Thus, we conclude that amy1 and amy2 are the major genes responsible for starch degradation during seed germination. Other indirect evidence includes the induced expression of amy1 and amy2 genes during seed germination by GA (Khursheed and Rogers, 1988; Karrer et al., 1991). The abundance of mRNA levels ranks amy32b (amy2_3) > amy6-4 (amy1_1a to amy1_1e, possibly sum) > amy46 (amy1_2) about 24 h after GA induction (Khursheed and Rogers, 1988; Karrer et al., 1991). Barley amy1 have expanded members due to genome duplication, which may play a key role in barley becoming a malting commodity. Furthermore, many amy genomic and cDNA clones had been deposited on Genbank. Their relationships for some of the clones with amy1 and amy2 was shown by a phylogenetic tree at Supplementary Figure S7.
There was no direct evidence for the function of amy3 and amy4 proteins in barley. In wheat, amy3 was highly expressed in developing grains, which affected carbon partitioning and diacylglycerol accumulation (Whan et al., 2014), while the amy4 gene may be involved in starch degradation working in partnership with amy1 proteins (Mieog et al., 2017).
Significance of Conserved Motifs on Promoters of amy1 and amy2 in the Induction of amy Gene Expression
Barley amy enzymes are synthesized in barley aleurone layers induced by gibberellin (GA) (Chrispeels and Varner, 1967; Jacobsen et al., 1970). Two groups of proteins (A and B) were detected after GA induction. Their responses to GA induction differed in a time and GA concentration dependent manner (Jacobsen and Higgins, 1982). The group A proteins expressed earlier and required a low GA concentration, while the group B proteins were not detectable till 8 h after GA addition and required a higher GA concentration. However, the synthesis of group B proteins accelerated once expressed, and one of group B isoforms was most abundant at 24 h (Jacobsen and Higgins, 1982). The group A and B proteins were likely to be the products of amy1 and amy2 genes, respectively, as shown by changes in mRNA levels in response to GA (Khursheed and Rogers, 1988). Conserved motifs on amy1 and amy2 promoters played key roles in the induction of gene expression. A comparison of amy1 with amy2 promoters showed substantial differences in nucleotide composition of the conserved motifs. The GA-responsive element is TAACAAA on all amy1 promoters, but TAACAGAG on all amy2 promoters for all three crops. There is a cAMY-like responsive element close to the pyrimidine box on barley and wheat amy1 promoters, but not amy2 promoters. The difference in the conserved motifs may play a key role in GA-induced gene expression.
Gene expression induced by GA may have the same mechanism for all amy1 members, since they contain extract same number of motifs with same nucleotide sequences except Hvamy1_2 gene on which the TATCCAC box was replaced by TATCCAT. In contrast, barley amy2 members may be differentially regulated, particularly for the Hvamy2_2 gene, since the conserved motifs differed substantially from amy1, amy2_1, and amy2_3. Furthermore, no similar motifs were found in the promoters of amy3 and amy4.
Genomic Properties of Wheat amy Genes Did Not Differ from Those of Barley amy Genes
Barley grains are often used for malting. Barley malts contain sufficient diastatic power (enzymatic hydrolytic activities) to completely convert starch to fermentable sugars. Extended numbers of amy genes and the presence of GA-regulatory motifs are important for barley to be used for malts (Table 1 and Figures 2, 3). However, wheat amy genes had similar genomic properties to barley amy genes with similar intron and exon structures (Supplementary Table S2). They also contained GA-regulatory motifs as in barley amy gene promoters (Supplementary Figures S5, S6). This could explain why wheat can also be used for malting (Fleming et al., 1960; Faltermaier et al., 2013). However, there are some problems with using wheat malts for brewing. Wheat grains lack husk, which is a problem for a brewing process called Lautering. Wheat also has higher protein (up to 20%). Wheat proteins promote foam formation, but also enhance haze issues (Faltermaier et al., 2015).
Conclusion
Subfamilies amy1 and amy2 have similar genomic properties in wheat and barley—including the number of exon/intron structures, localized on chromosomes 6 and 7, with GA-responsive elements on promoters—but differ in rice. Interestingly, the barley genome contains at least three more amy1 genes on chromosome 6H. Wheat should contain sufficient amy activity to completely degrade starch during malting. Other factors, such as high protein and the lack of husk, may limit the use of wheat for brewing.
Author Contributions
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2017.01727/full#supplementary-material
FIGURE S1 | Alignment of rice amy1 and amy2 proteins. Rice amy1 and amy2 proteins were aligned using ClustalW software (http://www.genome.jp/tools-bin/clustalw). The secondary structure features are shown on top of the alignments. There are three domains: domain A (black arrows and asterisks), domain B (green arrows) and domain C (red arrows). Domain A consists of a (α/β)8 barrel, while domain C has five β-sheets. The three catalytic amino acids Asp203, Glu228, and Asp310 are indicated by red asterisks.
FIGURE S2 | Rice amy1 and amy2 promoter alignments. (A) ALL amy1 contain a GA-responsive element (GARE) TAACAAA (red box), a pyrimidine box (CCTTTT) and a TATCCA(C/T) box (blue boxes). (B) Rice amy2 promoter (LOC_Os06g49970) within 500 bp from translation start codon (ATG) contained GA-responsive element only. No pyrimidine and TATCCA(C/T) were found.
FIGURE S3 | Alignment of wheat amy1 proteins. Wheat amy1 proteins were aligned using ClustalW software (http://www.genome.jp/tools-bin/clustalw).
FIGURE S4 | Alignment of wheat amy2 proteins. Wheat amy2 proteins were aligned using ClustalW software (http://www.genome.jp/tools-bin/clustalw).
FIGURE S5 | Alignment of wheat amy1 promoter sequences. ALL wheat amy1 contain a GA-responsive element (GARE) TAACAAA (red box), a pyrimidine box (CCTTTT), a TATCCA(C/T) box (blue boxes) and a cAMP-like motif (TGAGCTC) (green box).
FIGURE S6 | Alignment of wheat amy2 promoter sequences. The GA-responsive element (GARE) TAACAAA (red), pyrimidine box (CCTTTT), TATCCA(C/T) box (blue), and OS2 motifs (green) are marked.
FIGURE S7 | A comparison of new and old amy nomenclatures. The phylogenetic tree was generated using the ClustalW program (http://www.genome.jp/tools-bin/clustalw). The nucleotide sequences were collected from GenBank with accession numbers M17125.1, M17126.1, M17127.1, and M17128.1 (Knox et al., 1987); X15226 and X15227 (Rahmatullah et al., 1989); J01236.1 (Rogers and Milliman, 1983); K02637 (Rogers, 1985); J04202 (Khursheed and Rogers, 1988), and X05166 (Whittier et al., 1987).
Footnotes
References
Bahaji, A., Li, J., Sánchez-López, Á. M., Baroja-Fernández, E., Muñoz, F. J., Ovecka, M., et al. (2014). Starch biosynthesis, its regulation and biotechnological approaches to improve crop yields. Biotechnol. Adv. 32, 87–106. doi: 10.1016/j.biotechadv.2013.06.006
Bak-Jensen, K. S., Laugesen, S., Ostergaard, O., Finnie, C., Roepstorff, P., and Svensson, B. (2007). Spatio-temporal profiling and degradation of alpha-amylase isozymes during barley seed germination. Febs J. 274, 2552–2565. doi: 10.1111/j.1742-4658.2007.05790.x
Barrero, J. M., Mrva, K., Talbot, M. J., White, R. G., Taylor, J., Gubler, F., et al. (2013). Genetic, hormonal and physiological analysis of late maturity alpha-amylase (LMA) in wheat. Plant Physiol. 161, 1265–1277. doi: 10.1104/pp.112.209502
Brown, A. H. D., and Jacobsen, J. V. (1982). Genetic basis and natural variation of α-amylase isozymes in barley. Genet. Res. 40, 315–324. doi: 10.1017/s0016672300019182
Chrispeels, M. J., and Varner, J. E. (1967). Gibberellic acid-enhanced synthesis and release of α-amylase and ribonuclease by isolated barley and aleurone layers. Plant Physiol. 42, 398–406. doi: 10.1104/pp.42.3.398
Emebiri, L. C., Moody, D. B., Panozzo, J. F., and Read, B. J. (2004). Mapping of QTL for malting quality attributes in barley based on a cross of parents with low grain protein concentration. Field Crop. Res. 87, 195–205. doi: 10.1016/j.fcr.2003.11.002
Evans, D. E., Li, C., and Eglinton, J. K. (2010). “The properties and genetics of barley malt starch degrading enzymes,” in Genetics and Improvement of Barley Malt Quality, eds G. Zhang and C. Li (Berlin: Springer), 143–189.
Faltermaier, A., Waters, D., Becker, T., Arendt, E., and Gastl, M. (2013). Protein modifications and metabolic changes taking place during the malting of common wheat (Triticum aestivum L.). J. Am. Soc. Brew. Chem. 71, 153–160.
Faltermaier, A., Zarnkow, M., Becker, T., Gastl, M., and Arendt, E. K. (2015). Common wheat (Triticum aestivum L.): evaluating microstructural changes during the malting process by using confocal laser scanning microscopy and scanning electron microscopy. Eur. Food Res. Technol. 241, 239–252. doi: 10.1007/s00217-015-2450-x
Fincher, G. B. (2010). “Biochemistry, physiology, and genetics of endosperm mobilization in germinated barley grain,” in Barley, ed. S. Ullrich (Chichester: Wiley-Blackwell), 449–477. doi: 10.1002/9780470958636.ch14
Fleming, J. R., Johnson, J. A., and Miller, B. S. (1960). Effect of malting procedure and wheat storage conditions on alpha-amylase and protease activities. Cereal Chem. 37, 363–370.
Gao, W., Clancy, J. A., Han, F., Jones, B. L., Budde, A., Wesenberg, D. M., et al. (2004). Fine mapping of a malting-quality QTL complex near the chromosome 4H S telomere in barley. Theor. Appl. Genet. 109, 750–760. doi: 10.1007/s00122-004-1688-7
Gómez-Cadenas, A., Zentella, R., Walker-Simmons, M. K., and Ho, T.-H. D. (2001). Gibberellin/abscisic acid antagonism in barley aleurone cells: site of action of the protein kinase PKABA1 in relation to gibberellin signaling molecules. Plant Cell 13, 667–679. doi: 10.1105/tpc.13.3.667
Gubler, F., and Jacobsen, J. V. (1992). Gibberellin-responsive elements in the promoter of a barley high-pI alpha-amylase gene. Plant Cell 4, 1435–1441.
Gupta, M., Abu-Ghannam, N., and Gallaghar, E. (2010). Barley for brewing: Characteristic changes during malting, brewing and applications of its by-products. Compr. Rev. Food Sci. F. 9, 318–328. doi: 10.1111/j.1541-4337.2010.00112.x
Han, F., and Ullrich, S. E. (1994). Mapping of quantitative trait loci associated with malting quality in barley. Barley Genet. Newsl. 23, 84–97.
Hayes, P. M., Liu, B. H., Knapp, S. J., Chen, F., Jones, B., Blake, T., et al. (1993). Quantitative trait locus effects and environmental interaction in a sample of North American barley germ plasm. Theor. Appl. Genet. 87, 392–401. doi: 10.1007/bf01184929
Jacobsen, J. V., and Higgins, T. J. V. (1982). Characterization of the α-amylases synthesized by aleurone layers of Himalaya barley in response to gibberellic acid. Plant Physiol. 70, 1647–1653. doi: 10.1104/pp.70.6.1647
Jacobsen, J. V., Scandalios, J. G., and Varner, J. E. (1970). Multiple forms of amylase induced by gibberellic acid in isolated barley aleurone layers. Plant Physiol. 45, 367–371. doi: 10.1104/pp.45.4.367
Kadziola, A., Abe, J.-I., Svensson, B., and Haser, R. (1994). Crystal and molecular structure of barley α-amylase. J. Mol. Biol. 239, 104–121. doi: 10.1006/jmbi.1994.1354
Karrer, E. E., Litts, J. C., and Rodriguez, R. L. (1991). Differential expression of alpha-amylase genes in germinating rice and barley seeds. Plant Mol. Biol. 16, 797–805. doi: 10.1007/BF00015072
Khursheed, B., and Rogers, J. C. (1988). Barley alpha-amylase genes. Quantitative comparison of steady-state mRNA levels from individual members of the two different families expressed in aleurone cells. J. Biol. Chem. 263, 18953–18960.
Knox, C. A., Sonthayanon, B., Chandra, G. R., and Muthukrishnan, S. (1987). Structure and organization of two divergent alpha-amylase genes from barley. Plant Mol. Biol. 9, 3–17. doi: 10.1007/bf00017982
Lanahan, M. B., Ho, T. H., Rogers, S. W., and Rogers, J. C. (1992). A gibberellin response complex in cereal alpha-amylase gene promoters. Plant Cell 4, 203–211. doi: 10.1105/tpc.4.2.203
Li, C. D., Cakir, M., and Lance, R. (2010). “Genetic improvement of malting quality through conventional breeding and marker-assisted selection,” in Genetics and Improvement of Barley Malt Quality, eds G. Zhang and C. Li (Berlin: Springer), 260–292.
MacGregor, A., LaBerge, D., and Meredith, W. (1971). Separation of α-and β-amylase enzymes from barley malt by ion-exchange chromatography. Cereal Chem. 48, 490–498.
Marquez-Cedillo, A. L., Hayes, M. P., Jones, L. B., Kleinhofs, A., Legge, G. W., Rossnagel, G. B., et al. (2000). QTL analysis of malting quality in barley based on the doubled-haploid progeny of two elite North American varieties representing different germplasm groups. Theor. Appl. Genet. 101, 173–184. doi: 10.1007/s001220051466
Mascher, M., Gundlach, H., Himmelbach, A., Beier, S., Twardziok, S. O., Wicker, T., et al. (2017). A chromosome conformation capture ordered sequence of the barley genome. Nature 544, 427–433. doi: 10.1038/nature22043
Mieog, J. C., Janeček, Š, and Ral, J.-P. (2017). New insight in cereal starch degradation: identification and structural characterization of four α-amylases in bread wheat. Amylase 1, 35–49. doi: 10.1515/amylase-2017-0004
Muthukrishnan, S., Gill, B. S., Swegle, M., and Chandra, G. R. (1984). Structural genes for alpha-amylases are located on barley chromosomes 1 and 6. J. Biol. Chem. 259, 13637–13639.
Oziel, A., Hayes, P. M., Chen, F. Q., and Jones, B. (1996). Application of quantitative trait locus mapping to the development of winter-habit malting barley. Plant Breed. 115, 43–51. doi: 10.1111/j.1439-0523.1996.tb00869.x
Rahmatullah, R. J., Huang, J. K., Clark, K. L., Reeck, G. R., Chandra, G. R., and Muthukrishnan, S. (1989). Nucleotide and predicted amino acid sequences of two different genes for high-pI alpha-amylases from barley. Plant Mol. Biol. 12, 119–121. doi: 10.1007/bf00017454
Robert, X., Haser, R., Gottschalk, T. E., Ratajczak, F., Driguez, H., Svensson, B., et al. (2003). The structure of barley α-amylase isozyme 1 reveals a novel role of domain C in substrate recognition and binding: a pair of sugar tongs. Structure 11, 973–984. doi: 10.1016/S0969-2126(03)00151-5
Rogers, J. C. (1985). Two barley alpha-amylase gene families are regulated differently in aleurone cells. J. Biol. Chem. 260, 3731–3738.
Rogers, J. C., Lanahan, M. B., and Rogers, S. W. (1994). The cis-acting gibberellin response complex in high-pl [alpha]-amylase gene promoters (requirement of a coupling element for high-level transcription). Plant Physiol. 105, 151–158. doi: 10.1104/pp.105.1.151
Rogers, J. C., and Milliman, C. (1983). Isolation and sequence analysis of a barley alpha-amylase cDNA clone. J. Biol. Chem. 258, 8169–8174.
Rogers, J. C., and Milliman, C. (1984). Coordinate increase in major transcripts from the high pI alpha-amylase multigene family in barley aleurone cells stimulated with gibberellic acid. J. Biol. Chem. 259, 12234–12240.
Shahpiri, A., Talaei, N., and Finnie, C. (2015). Spatio-temporal appearance of alpha-amylase and limit dextrinase in barley aleurone layer in response to gibberellic acid, abscisic acid and salicylic acid. J. Sci. Food Agric. 95, 141–147. doi: 10.1002/jsfa.6695
Skriver, K., Olsen, F. L., Rogers, J. C., and Mundy, J. (1991). Cis-acting DNA elements responsive to gibberellin and its antagonist abscisic acid. Proc. Natl. Acad. Sci. U.S.A. 88, 7266–7270. doi: 10.1073/pnas.88.16.7266
Svensson, B., Mundy, J., Gibson, R. M., and Svendsen, I. (1985). Partial amino acid sequences of α-amylase isozymes from barley malt. Carlsberg Res. Commun. 50, 15–22. doi: 10.1007/BF02910534
Whan, A., Dielen, A.-S., Mieog, J., Bowerman, A. F., Robinson, H. M., Byrne, K., et al. (2014). Engineering α-amylase levels in wheat grain suggests a highly sophisticated level of carbohydrate regulation during development. J. Exp. Bot. 65, 5443–5457. doi: 10.1093/jxb/eru299
Whittier, R. F., Dean, D. A., and Rogers, J. C. (1987). Nucleotide sequence analysis of alpha-amylase and thiol protease genes that are hormonally regulated in barley aleurone cells. Nucleic Acids Res. 15, 2515–2535. doi: 10.1093/nar/15.6.2515
Woodger, F., Jacobsen, J. V., and Gubler, F. (2010). “Gibberellin action in germinated cereal grains,” in Plant Hormones: Biosynthesis, Signal Transduction, Action!, ed. P. J. Davies (Dordrecht: Springer), 221–240.
Keywords: α-amylase, barley, conserved motif, genome, gibberellin responsive complex, promoter
Citation: Zhang Q and Li C (2017) Comparisons of Copy Number, Genomic Structure, and Conserved Motifs for α-Amylase Genes from Barley, Rice, and Wheat. Front. Plant Sci. 8:1727. doi: 10.3389/fpls.2017.01727
Received: 07 July 2017; Accepted: 21 September 2017;
Published: 05 October 2017.
Edited by:
Jacqueline Batley, University of Western Australia, AustraliaReviewed by:
You-Liang Zheng, Sichuan Agricultural University, ChinaMarcelo Helguera, Instituto Nacional de Tecnología Agropecuaria (INTA), Argentina
Copyright © 2017 Zhang and Li. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Chengdao Li, Yy5saUBtdXJkb2NoLmVkdS5hdQ==
 Qisen Zhang
Qisen Zhang Chengdao Li
Chengdao Li