- 1CAS Key Laboratory of Genome Sciences and Information, Beijing Institute of Genomics, Chinese Academy of Sciences, Beijing, China
- 2College of Life Science and Technology, Huazhong University of Science and Technology, Wuhan, China
- 3University of Chinese Academy of Sciences, Beijing, China
- 4Grail Scientific Co., Ltd., Shenyang, China
- 5Joint Center for Genomics Research, King Abdulaziz City for Science and Technology and Chinese Academy of Sciences, Riyadh, Saudi Arabia
Background: Date palm (Phoenix dactylifera L.) is a cultivated woody plant with agricultural and economic importance in many countries around the world. Wit h the advantages of next generation sequencing technologies, genome sequences for many date palm cultivars have been released recently. Short sequence repeat (SSR) and single nucleotide polymorphism (SNP) can be identified from these genomic data, and have been proven to be very useful biomarkers in plant genome analysis and breeding.
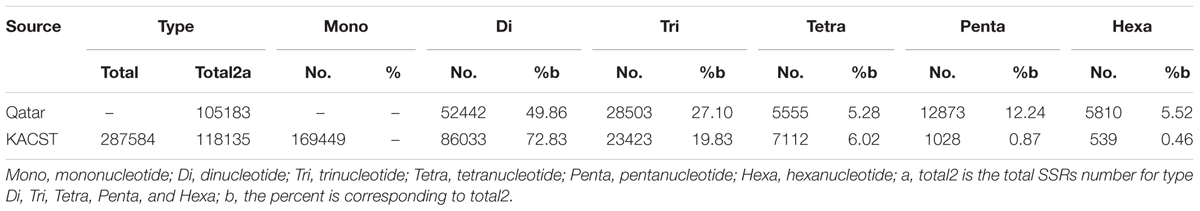
Results: Here, we first improved the date palm genome assembly using 130X of HiSeq data generated in our lab. Then 246,445 SSRs (214,901 SSRs and 31,544 compound SSRs) were annotated in this genome assembly; among the SSRs, mononucleotide SSRs (58.92%) were the most abundant, followed by di- (29.92%), tri- (8.14%), tetra- (2.47%), penta- (0.36%), and hexa-nucleotide SSRs (0.19%). The high-quality PCR primer pairs were designed for most (174,497; 70.81% out of total) SSRs. We also annotated 6,375,806 SNPs with raw read depth≥3 in 90% cultivars. To further reduce false positive SNPs, we only kept 5,572,650 (87.40% out of total) SNPs with at least 20% cultivars support for downstream analyses. The high-quality PCR primer pairs were also obtained for 4,177,778 (65.53%) SNPs. We reconstructed the phylogenetic relationships among the 62 cultivars using these variants and found that they can be divided into three clusters, namely North Africa, Egypt – Sudan, and Middle East – South Asian, with Egypt – Sudan being the admixture of North Africa and Middle East – South Asian cultivars; we further confirmed these clusters using principal component analysis. Moreover, 34,346 SSRs and 4,177,778 SNPs with PCR primers were assigned to shared cultivars for cultivar classification and diversity analysis. All these SSRs, SNPs and their classification are available in our database, and can be used for cultivar identification, comparison, and molecular breeding.
Conclusion: DRDB is a comprehensive genomic resource database of date palm. It can serve as a bioinformatics platform for date palm genomics, genetics, and molecular breeding. DRDB is freely available at http://drdb.big.ac.cn/home.
Introduction
Date palm (Phoenix dactylifera L.) is a widely cultivated plant species with agricultural and economic importance in the world. It is a dioecious plant in palm family Arecaceae with long life cycle, long period of juvenility and various of cultivars (El Hadrami et al., 2011). As the most important crop, Saudi Arabia has an estimated 25 million date palms producing nearly a million tons of dates annually. More than 400 different date palm cultivars are reported to exist in Saudi Arabia (Anonymous, 2006).
Based on the next generation sequencing, date palm genome sequence has been released and many date palm cultivars have been re-sequenced recently (Al-Dous et al., 2011; Al-Mssallem et al., 2013; Hazzouri et al., 2015). Short sequence repeat (SSR) and single nucleotide polymorphism (SNP) are very useful in plant genome analysis and breeding. Billotte et al. (2004) developed the nuclear SSR markers and investigated the genetic diversity for date palm while Hamwieh et al. (2010) analyzed the SSRs across the whole date palm genome. Based on SNPs among 11 date palm varieties, Al-Mssallem et al. (2013) noticed that the northern African varieties are significantly diverged from the Middle Eastern varieties.
To better understand the genetic basis of different date palm cultivars, it is necessary to have a high-quality genome assembly and get insight into genome variations (SSRs and SNPs). Here, based on 130X HiSeq data produced by our lab, we first updated the date palm genome assembly. Then, we annotated SSRs and other types of sequence variants among 62 cultivars. To facilitate the use of these data, we developed a web-based date palm genome database (DRDB). We included in DRDB in total of 6,375,806 SNPs and 246,445 SSRs from 62 cultivars. The main purpose of DRDB is to help the researchers/breeders to distinguish the plethora of date palm cultivars by using well-selected polymorphic markers (SNPs or SSRs). With DRDB, users can search for SNPs or SSRs of interests, and easily find SNP/SSR markers that are unique to user-selected cultivars. In addition, we also provided detailed annotations for the SNPs and SSRs. With DRDB, we not only expanded and enhanced the date palm cultivars data contents, but also incorporated useful tools to mine the data. To our knowledge, DRDB is the first comprehensive genetic variation database for date palm.
Materials and Methods
Plant Materials
Fresh green leaves from an adult date palm plant of Khalas (female) cultivar in Al-Hssa Oasis (24∘∘08′54′′N, 47°18′18′′E) were collected, washed with double distilled water, and frozen immediately in a liquid nitrogen container. After transported to the laboratory, these samples were stored in -80°C freezers until use.
Genomic DNA Isolation and Sequencing
Genomic DNA was isolated from 50 g fresh leaves according to a CTAB-based method as before (Al-Mssallem et al., 2013). Briefly, 5 mg purified DNA was used for constructing the Illumina HiSeq libraries. HiSeq paired-end (180, 300, and 500 bp) and mate-pair libraries (1, 3, 5, and 8 kb) were constructed using the Illumina Simple Paired-End Library and Mate-Pair Library Preparation Protocol, respectively. The libraries were sequenced by Illumina HiSeq 2000 platform.
Genome Assembly
We obtained ∼130X HiSeq data using Illumina HiSeq 2000 platform. The raw data was filtered using an in-house Perl script firstly. Then, SOAPdenovo2 (v2.04) was used to build a new genome assembly based on the newly produced HiSeq data with the option “-k 63” and other default parameters (Luo et al., 2012). Next, the new genome assembly and the previous genome assembly were merged by GAA (v1.1) (Yao et al., 2012). After that, ErrorCorrection and GapCloser modules in SOAPdenovo2 were used sequentially for reads correction and gaps close. Finally, the properly aligned BAC-end sequences (produced before) were used to construct the final scaffolds using an in-house Perl script.
SSRs and Sequence Variants Identification
The updated genome assembly was used for SSR identification by MISA (v1.0) (Thiel et al., 2003) (Table 1). The re-sequencing data of 62 date palm cultivars produced by Hazzouri et al. (2015) were downloaded from the Sequence Read Archive (SRA) of National Center of Biotechnology Information (NCBI) (Supplementary Table S1). The raw data was trimmed by Trimmomatic (v0.33) (Bolger et al., 2014) and aligned to the reference genome using the mem algorithm of BWA (v0.7.12) (Li and Durbin, 2009). The sequence variants were identified by samtools and bcftools (Li and Durbin, 2009; Li, 2011a,b). To reduce the false positives, SNPs presented more than 20% cultivars were kept for subsequent analysis. Primer3 (v3-2.3.7) was used for PCR primer design with default parameters except for “PRIMER_PRODUCT_SIZE_RANGE = 100–280” (Koressaar and Remm, 2007; Untergasser et al., 2012). The putative biological consequences of sequence variants on gene models were predicted by ANNOVAR (v2013-05-20) (Wang et al., 2010).
Phylogenetic Analysis
The phylogenetic tree was constructed using all SNP sites of the varieties by MEGA (NJ method with 1,000 bootstrap, version 6.06) (Tamura et al., 2013). The gaps or missing data were eliminated when the site coverage below 90%. The constructed phylogenetic tree was visualized with EvolView, an online phylogenetic tree visualization tool (Zhang et al., 2012). Principal component analysis (PCA) of SNP genotypes for 62 cultivars was carried out using EIGENSTRAT (Price et al., 2006). The population structure was analyzed using STRUCTURE (Pritchard et al., 2000). The k-value of STRUCTURE means the population can be divided into K categories. Each k-value was repeated five times. CLUMPP (version 1.1.2) was used to permute the clusters generated from independent STRUCTURE runs (Jakobsson and Rosenberg, 2007).
Website Construction
The DRDB contained both SNPs and SSRs data from 62 date palm cultivars. The website was built on LAMP stack (Linux, Apache, MySQL, and PHP). MySQL was used for data storage on the server side. The front-end interface was implemented with Bootstrap and Angular.
Results
The Updated Date Palm Genome Assembly
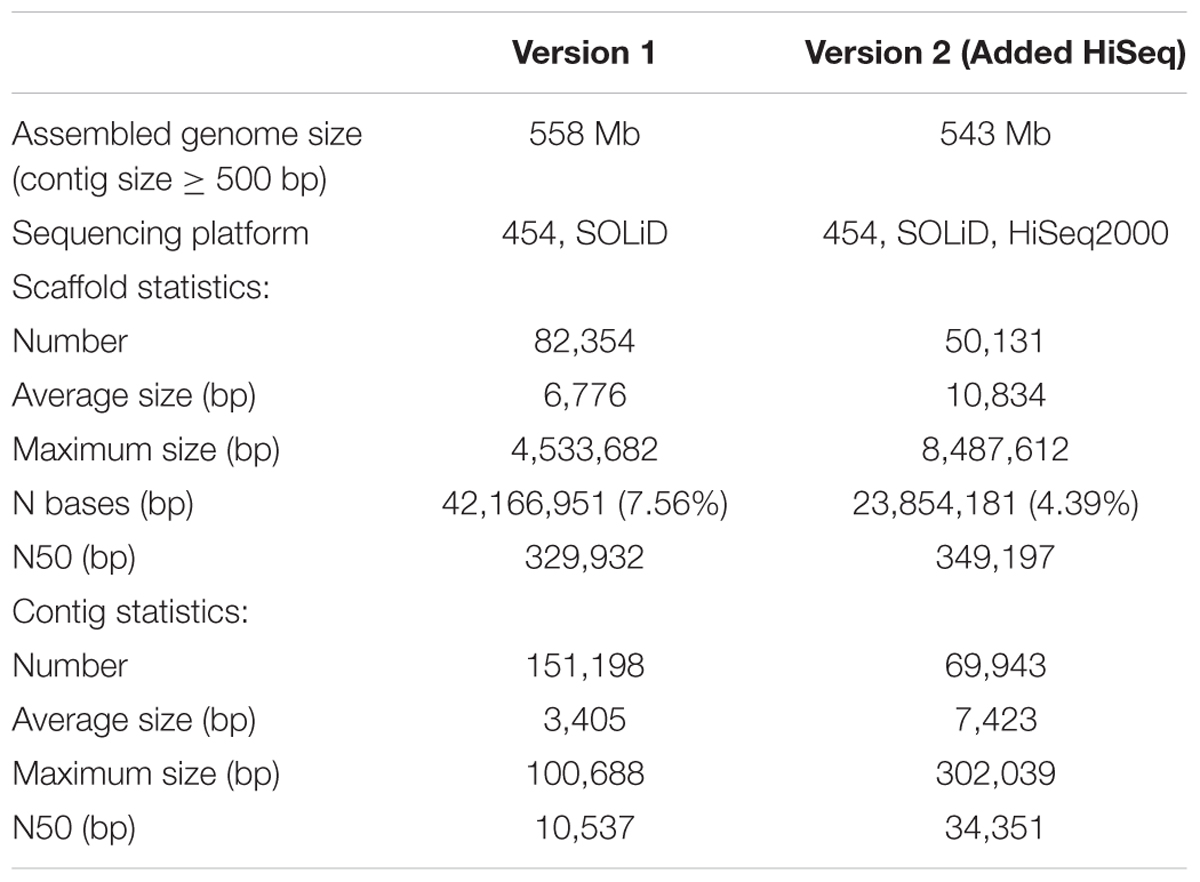
We re-assembled the palm genome by incorporating newly sequenced HiSeq data (∼130X) produced in our lab and obtained a new genome assembly of 602,484,697 bp in length. As summarized in Table 2, comparing the new version (referred as to version 2 below) with the old version (referred as to version 1 below), the number of scaffolds and N bases were reduced to 60.87 and 56.57%, respectively. At contig level, the quality of version 2 was significantly improved; for example, N50 was increased threefold to 34.35 kb, and the number of contigs was decreased ∼50% to 69,963 (Table 2). Our results suggested that the new genome assembly could not be further improved by adding more shot-read data with limited insert size (≤8 kb).
SSRs and SNPs
Using the updated date palm genome sequence, we identified 246,445 SSRs. Among them, 31,544 (12.80%) were compound SSRs (Figure 1) and 26,148 (10.61%) were imperfect SSRs. The imperfect SSRs in date palm are significantly less than those in chickpea (17%) (Winter et al., 1999) and lentils (27.1%) (Hamwieh et al., 2009), similar to those in Elaeis guineensis (10.87%), and more than those in Oryza sativa (9.60%), Aegilops tauschii (7.53%), Zea mays (5.72%), and Brachypodium distachyon (5.47%). Mononucleotide SSRs and dinucleotide SSRs are the most abundant types (58.92 and 29.92%, respectively); among the dinucleotide SSRs, AG/TC (50.50%) is the most abundant, followed by AT/TA (35.50%), AC/TG (13.25%), and CG/GC (0.75%). Trinucleotide SSRs account for 8.14% of the total SSRs, among which AAG/TTC (30.81%) is the most abundant, followed by AAT/TTA (23.20%) and AGG/TCC (17.97%). Using Primer 3, we were able to design high-quality primer pairs for 174,497 (70.81%) SSRs.
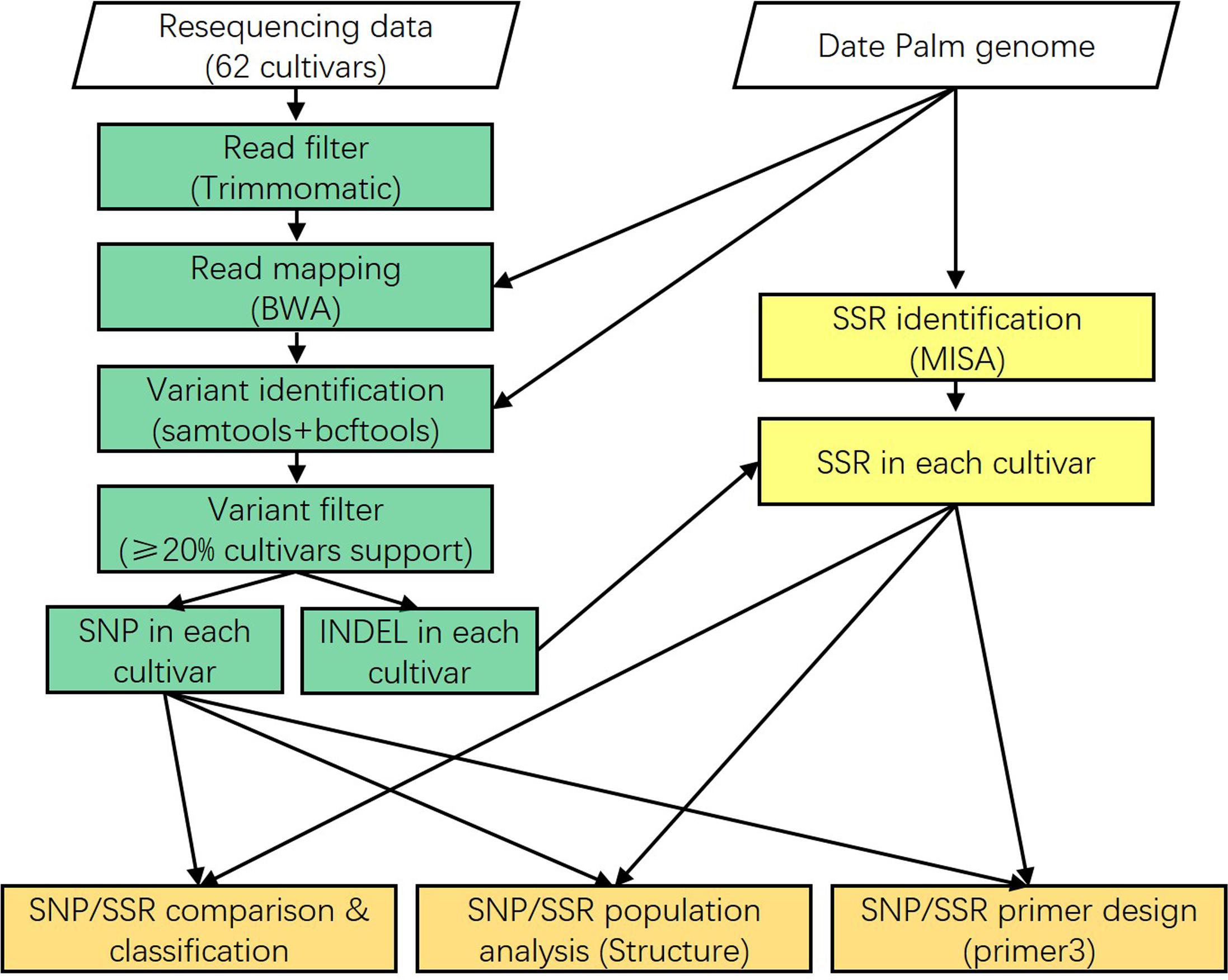
FIGURE 1. Single nucleotide polymorphisms (SNPs) and short sequence repeats (SSRs) identification pipeline. The 62 date palm cultivars were used to detect SNPs and SSRs. Classification, population structure, and primer design were performed for these SNPs and SSRs.
For the published SSRs that we examined, most of them were covered by our dataset. For example, the SSRs that we inferred here contained 9 of 16 ones proposed previously Billotte et al. (2004) and 11 of 17 SSR loci Akkak et al. (2009); we also found 8 of 19 SSR primers Hamwieh et al. (2010) and 13 of 30 SSR primers by Elmeer et al. (2011) in our data (Supplementary Table S2).
We identified 6,375,806 SNPs (raw read depth≥3 in 90% cultivars) among 62 date palm cultivars. To reduce false positives, we only kept those (5,572,650, 87.40%) found in more than 20% cultivars for downstream analysis. We designed high-quality PCR primers for 4,177,778 (65.53%) SNPs.
These genetic variants and pre-designed PCR primers could be a valuable resource for researchers for genetic and breeding studies.
Population Structure
To elucidate the population genetic structure, we selected 34,346 SSRs with length polymorphism and 57,310 SNPs in the largest scaffold of data palm genome for further analysis with STRUCTURE. We run STRUCTURE with k = 1–7 for SSRs and k = 1–6 for SNPs; we run STRUCTURE HARVESTER program to determine the best k-values for SSRs (k = 4) and SNPs (k = 3) (Supplementary Figure S1) (Earl, 2012). Our results showed that date palm can be divided into three genetically differentiated clusters, North Africa, Egypt – Sudan and Middle East – South Asian, with Egypt – Sudan being the admixture of North Africa and Middle East – South Asian cultivars (Figure 2). We further confirmed the clustering using PCA using all SNPs (Figure 3).
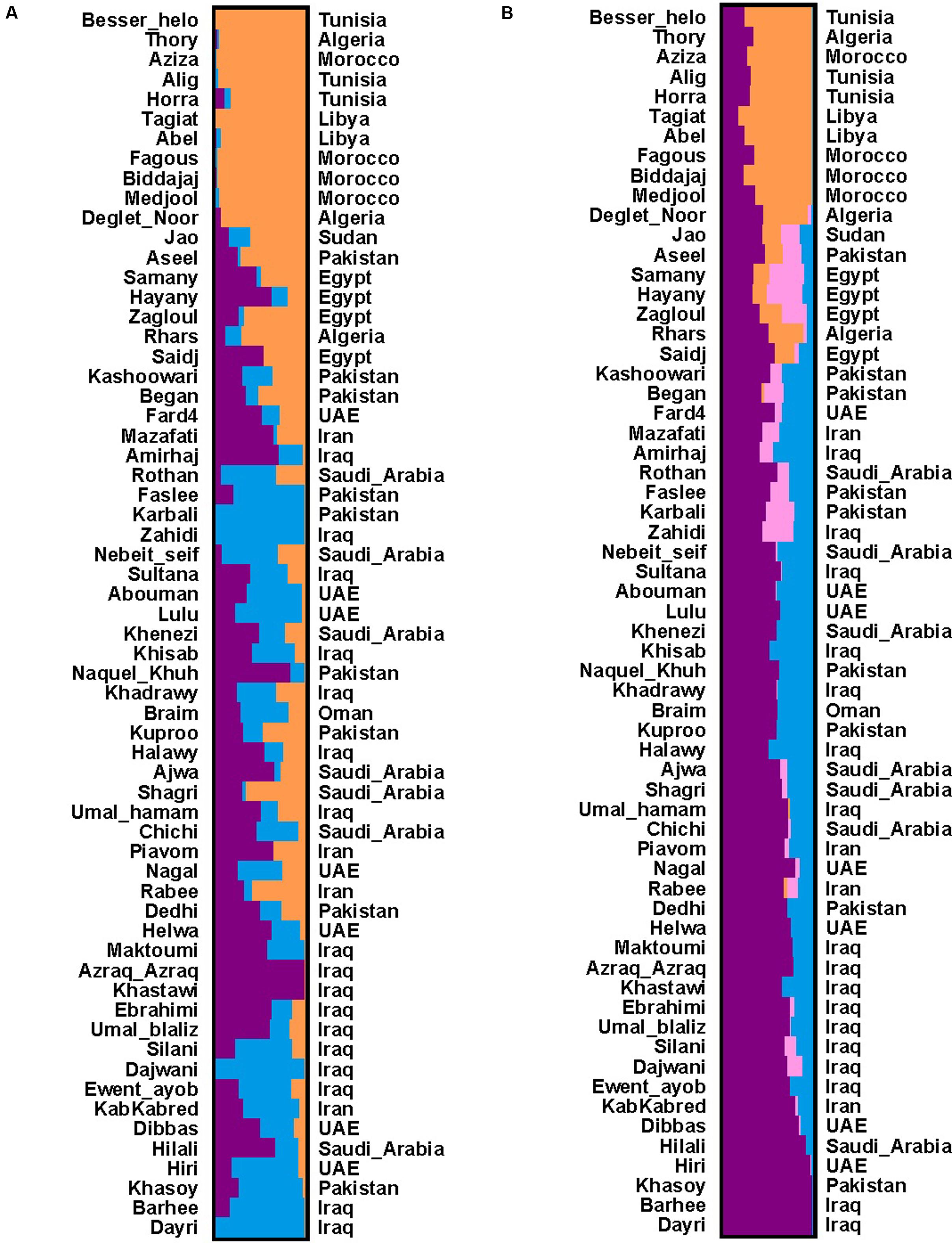
FIGURE 2. (A) Population structure based on SNPs (k = 3). The 62 date palm cultivars were grouped into three clusters. The color bar represents the component of ancestral origin. (B) Population structure based on SSRs (k = 4). The 62 date palm cultivars were grouped into three clusters. The color bar represents the component of ancestral origin. The two clusters (Middle East and South Asia) are nearly same according to the geography locations among four SSR-based clusters. Thus, the SSR-based clusters are considered as three clusters.
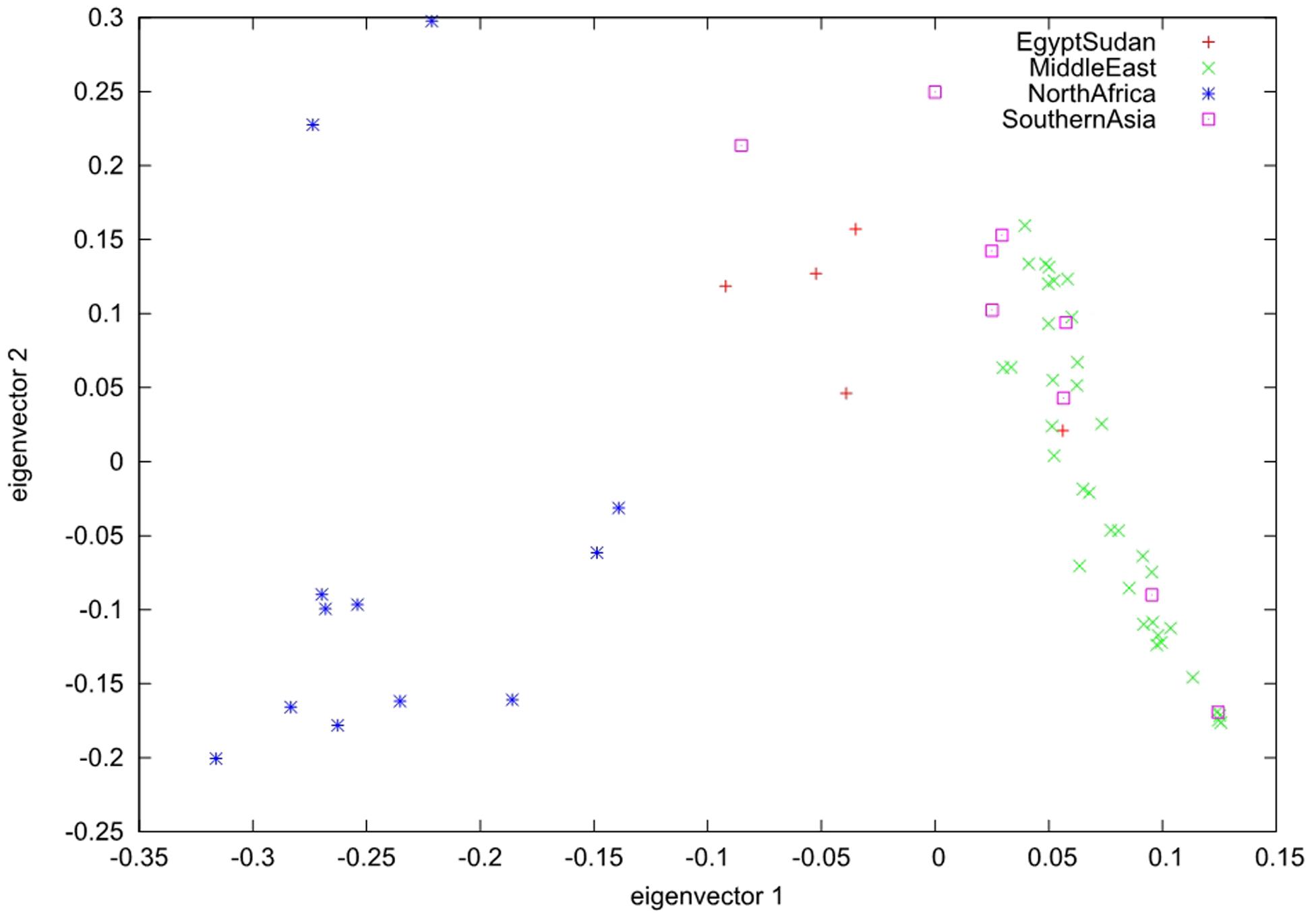
FIGURE 3. Principal component analysis (PCA) analyses of SNPs shows three distinct clusters of date palm cultivars. The scatter plot shows the PCA analysis of SNPs of 62 date palm cultivars. Each shape of dot represents one distinct geographic location. The plot shows three distinct clusters.
Construction of the Online Database
The Web Interface
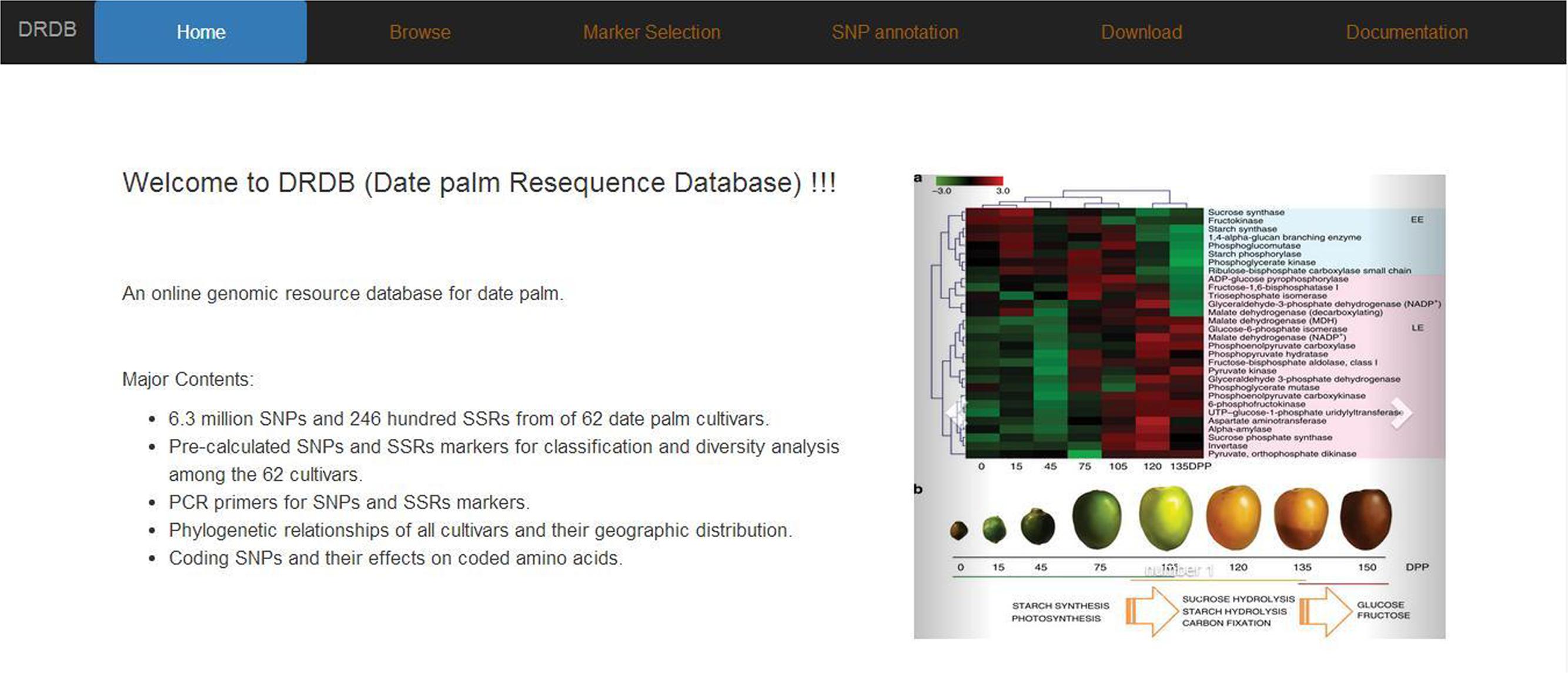
We designed DRDB with a user-friendly web interface, incorporated visualization tools for genomic features and provided powerful searching functionality. DRDB comes with several pages including Home, Browse, Marker selection, SNP annotation, Download, and Documentation. Below we introduce selected pages in details (Figure 4).
The Browse Page
In browse page, the phylogenetic tree that was derived from the SNP data is shown; the topology of this tree suggests that there are at least three sub-groups of date palm cultivars, each features distinct geographic locations: Middle East, Egypt Sudan, and North Africa. Users can click on any cultivar they interested in, and view a list of its available SNPs and SSRs in a new page (Figure 5), which lists all available SNPs and SSRs for user-selected cultivars. Users can search for SNPs and SSRs they are interested by changing different parameters (such as “DEPTH,” “QUALITY” in SNP data or “SSR type,” “Product size” in SSR data) (Figure 6).
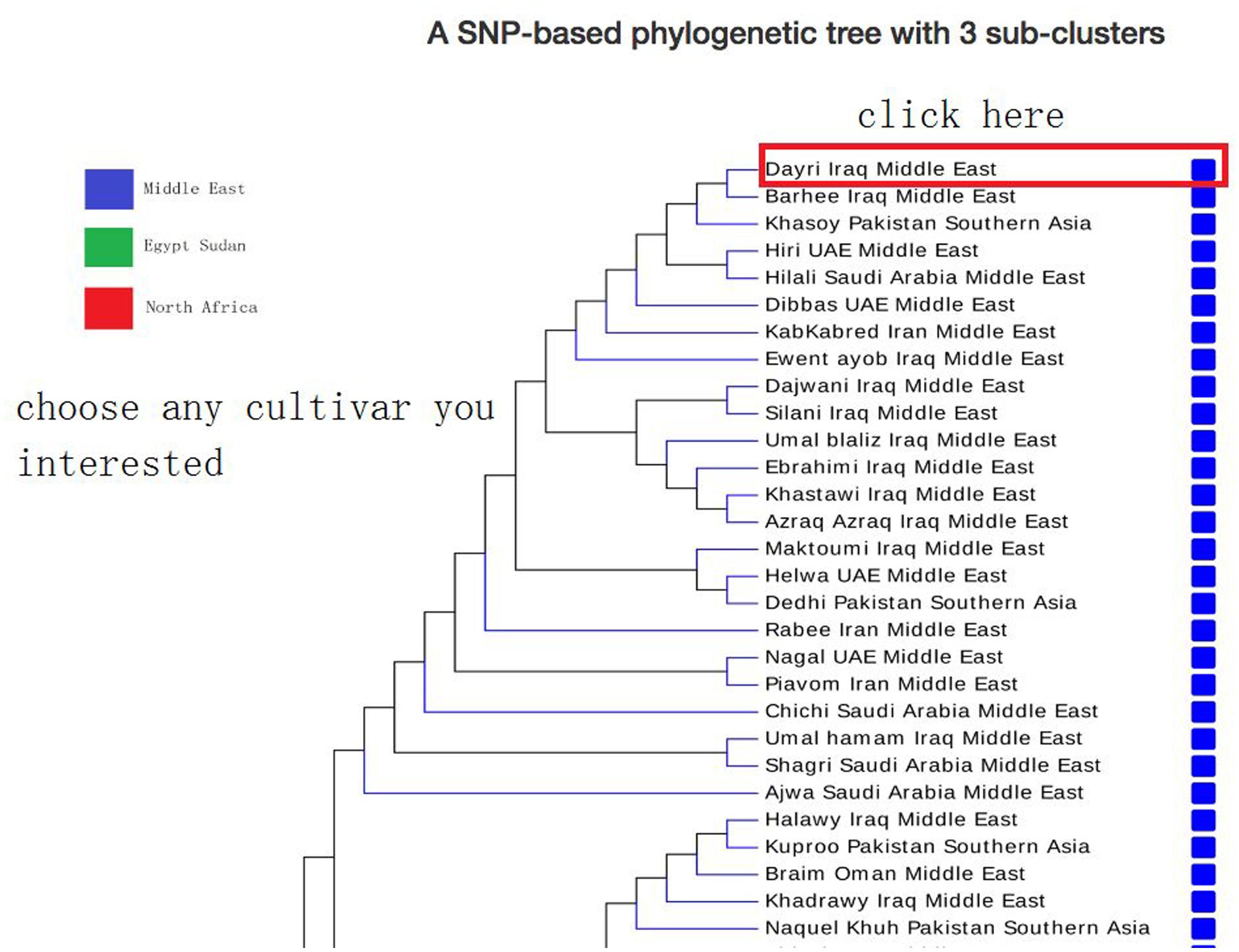

FIGURE 5. Date palm genome database browse page. A SNP-based phylogenetic tree was shown in browse page. Users can choose any cultivar they interested and then jump to the next level page to view SNP/SSR detailed information.
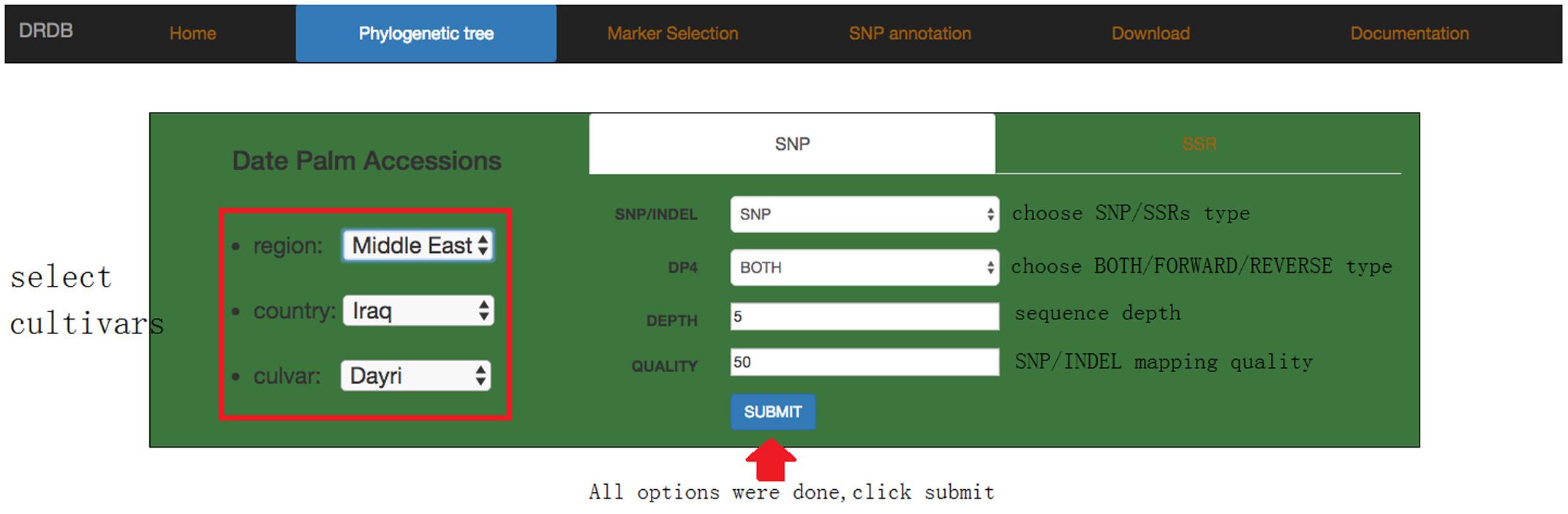
FIGURE 6. Date palm genome database cultivar information page. The next level page of browse is cultivar information page. Users can select any interested cultivar and marker type (SNP or SSR).
The Marker Selection Page
In marker selection page, we divided SNP and SSR markers into three levels (Region, Country, and Cultivars). Users can select any combination of maker types (SNP or SSR) and levels (Region, Country or Cultivars) they are interested, and view the results in a new page (Figure 7), which contains also the allele frequency and detailed information of selected markers, including “position, ref, alt, dp, qual” in SNP marker or “SSR_type, size, type, cov” in SSR marker (Figure 8).
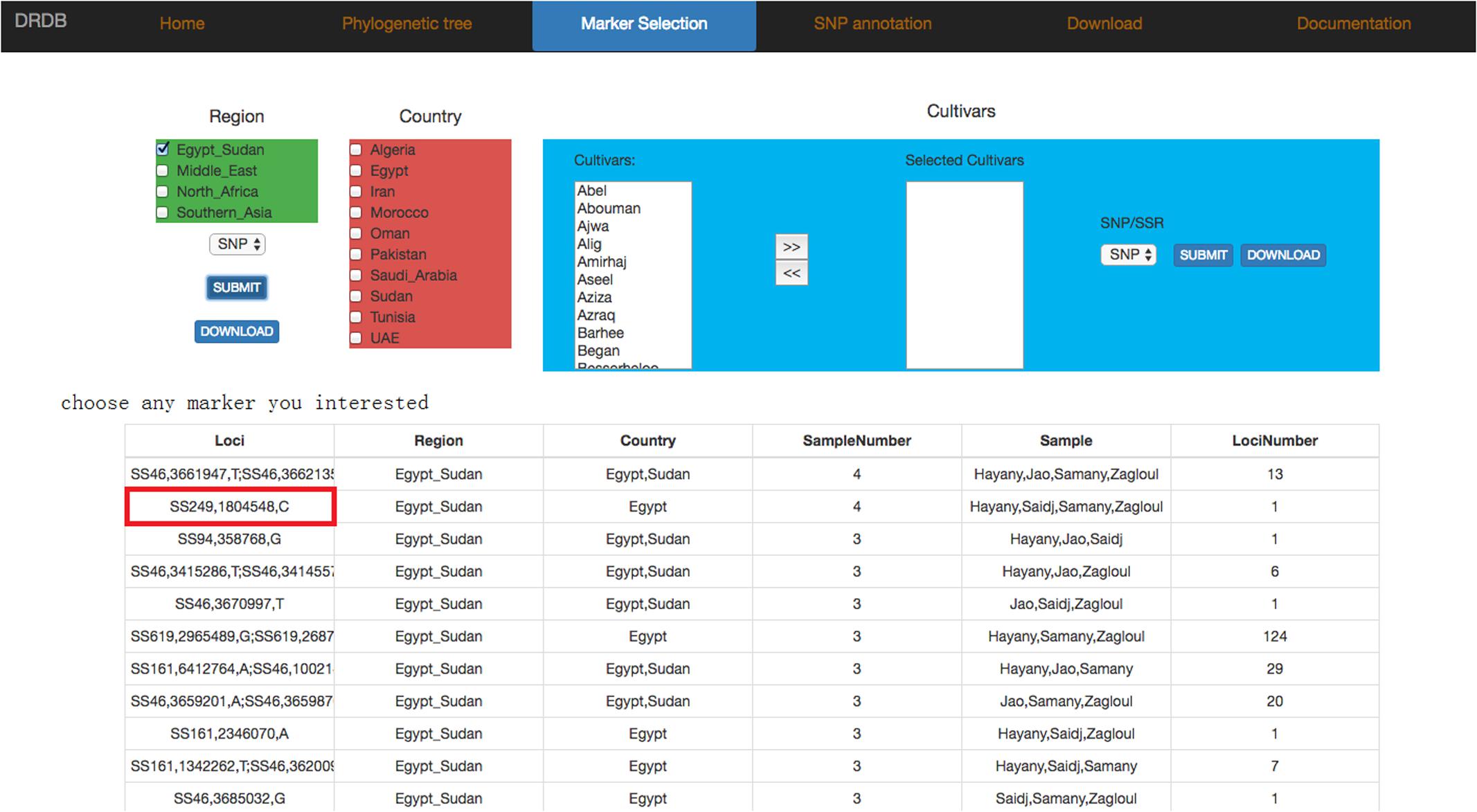
FIGURE 7. Date palm genome database marker selection page. Marker selection page provides SNP/SSR markers of different cultivars. Users can pick any marker to view marker frequency and detailed information in the next level page.
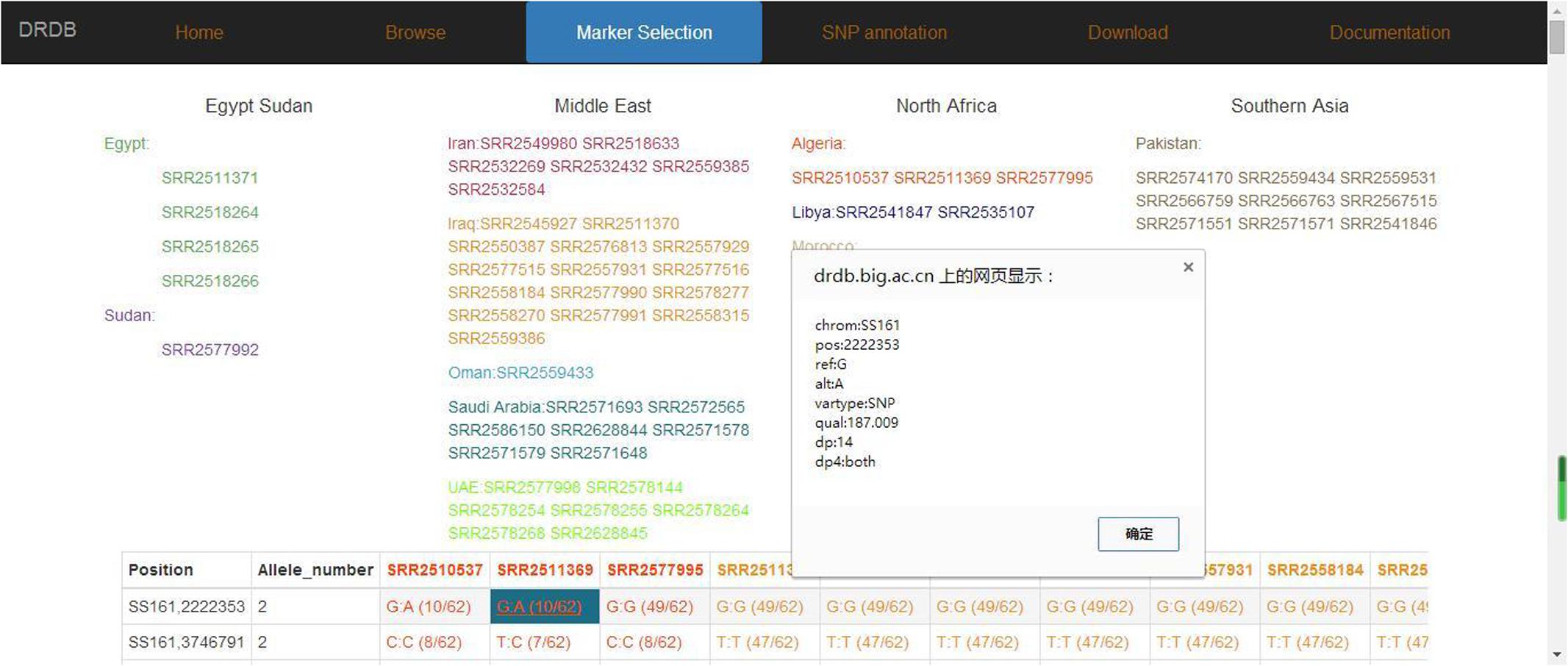
FIGURE 8. Date palm genome database marker frequency page. Marker frequency and detail information are shown in this page.
Other Pages
In addition to “Browse” and “Marker selection” pages, we also developed “SNP annotation” page. Users can search by the names of SNPs or their putative functional consequences such as “non-synonymous SNV,” “synonymous SNV,” “frameshift substitution,” and “non-frameshift substitution.” We also included gene annotation from external data sources including UNIPROT. All the SNPs, SSRs and SNP effect data can be downloaded in “Download” page. The “Documentation” page shows the detail introductions of DRDB.
Discussion
Date palm genome database provides users with a comprehensive genomic resources for date palm. The updated date palm genome assembly reduces the numbers of scaffolds and ambiguous bases significantly (60.87 and 56.57%) comparing with the previous genome assembly. Large (≥20 kb) insert size sequencing data, genetic map and physical map are needed for improving genome assembly in the future.
Date palm has unique genetic diversity due to long-lived life cycle, vegetative propagation, and complex origin. Study data palm genetic diversity and classification is an important research aspect. Hamwieh et al. (2010) identified previously ∼1000 SSR markers in an early version of the draft genome with genome size about 50% of our current assembly; they did not consider the SSRs variability among date palm cultivars, nor provided any SSR markers. Hazzouri et al. (2015) identified SNPs in 62 date palm cultivars, but they didn’t provide convenient tools for SNPs comparison nor designed primers for maker development.
In this study, based on an improved genome assembly, we identified 246,445 SSRs and 6,375,806 SNPs in 62 cultivars and designed high-quality PCR primers for most of variations (70.81% in SSRs and 65.53% in SNPs). These genetic variants and pre-designed PCR primers will be a valuable resource for researchers to perform genetic and breeding studies.
The origin of date palm is controversial because date palm cultivation spread out many countries starting in ancient times. Based on SSRs and SNPs, we elucidated the population genetic structure of date palm and the results showed that date palm can be divided into three genetically differentiated clusters, North Africa, Egypt – Sudan and Middle East – South Asian, with Egypt – Sudan being the admixture of North Africa and Middle East – South Asian cultivars.
To facilitate the use of our data, we developed DRDB for distinguishing the different sub-types of date palms and mining the genetic variation of interest. DRDB provides visualization tools for genomic features and powerful searching functionality. Users can download all these data from download page.
In summary, DRDB provided an updated date palm genome assembly and enabled genetic variation searching for 62 date palm cultivars rapidly and conveniently, which sets the stage for further genetic and breeding studies.
Author Contributions
ZH, CZ, and WL contributed equally to this work. SH, W-HC, and HA led the research. ZH and CZ constructed the database. WL, QL, and TW analyzed the data. ZH, W-HC, and WL wrote and revised the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This study is supported by grants from National Natural Science Foundation of China (Grant no. 31501042, 31271385, and 31200957), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant no. XDA08020102), and KACST grant 1035-35 from King Abdulaziz City for Science and Technology (KACST), Kingdom of Saudi Arabia.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2017.01889/full#supplementary-material
FIGURE S1 |(A) K-value selection of population structure for SNP. The picture of SNP-based k-value selection shows that 3 is the proper k-value, which is the highest value of the y-axis. (B) K-value selection of population structure for SSR. The picture of SSR-based k-value selection shows that 4 is the proper k-value, which is the highest value of the y-axis.
References
Akkak, A., Scariot, V., Marinoni, D. T., Boccacci, P., Beltramo, C., and Botta, R. (2009). Development and evaluation of microsatellite markers in Phoenix dactylifera L. and their transferability to other Phoenix species. Biol. Plant. 53, 164–166. doi: 10.1007/s10535-009-0026-y
Al-Dous, E. K., George, B., Al-Mahmoud, M. E., Al-Jaber, M. Y., Wang, H., Salameh, Y. M., et al. (2011). De novo genome sequencing and comparative genomics of date palm (Phoenix dactylifera). Nat. Biotechnol. 29, 521–527. doi: 10.1038/nbt.1860
Al-Mssallem, I. S., Hu, S., Zhang, X., Lin, Q., Liu, W., Tan, J., et al. (2013). Genome sequence of the date palm Phoenix dactylifera L. Nat. Commun. 4:2274. doi: 10.1038/ncomms3274
Anonymous (2006). A Review of the Processing of Dates in the Kingdom of Saudi Arabia: 1997 to 2005. Riyadh: Ministry of Agriculture.
Billotte, N., Marseillac, N., Brottier, P., Noyer, J. L., Jacquemoud-Collet, J. P., Moreau, C., et al. (2004). Nuclear microsatellite markers for the date palm (Phoenix dactylifera L.): characterization and utility across the genus Phoenix and in other palm genera. Mol. Ecol. Resour. 4, 256–258. doi: 10.1111/j.1471-8286.2004.00634.x
Bolger, A. M., Lohse, M., and Usadel, B. (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. doi: 10.1093/bioinformatics/btu170
Earl, D. A. (2012). Structure Harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361. doi: 10.1007/s12686-011-9548-7
El Hadrami, A., Daayf, F., and El Hadrami, I. (2011). Date Palm Biotechnology. Berlin: Springer, 479–512.
Elmeer, K., Sarwath, H., Malek, J., Baum, M., and Hamwieh, A. (2011). New microsatellite markers for assessment of genetic diversity in date palm (Phoenix dactylifera L.). 3 Biotech 1, 91–97. doi: 10.1007/s13205-011-0010-z
Hamwieh, A., Farah, J., Moussally, S., Al-Shamaa, K., Almer, K., Khierallah, H., et al. (2010). Development of 1000 microsatellite markers across the date palm (Phoenix dactylifera L.) genome. Acta Hortic. 882, 269–277. doi: 10.17660/ActaHortic.2010.882.29
Hamwieh, A., Udupa, S. M., Sarker, A., Jung, C., and Baum, M. (2009). Development of new microsatellite markers and their application in the analysis of genetic diversity in lentils. Breed. Sci. 59, 77–86. doi: 10.1270/jsbbs.59.77
Hazzouri, K. M., Flowers, J. M., Visser, H. J., Khierallah, H. S., Rosas, U., Pham, G. M., et al. (2015). Whole genome re-sequencing of date palms yields insights into diversification of a fruit tree crop. Nat. Commun. 6:8824. doi: 10.1038/ncomms9824
Jakobsson, M., and Rosenberg, N. A. (2007). CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806. doi: 10.1093/bioinformatics/btm233
Koressaar, T., and Remm, M. (2007). Enhancements and modifications of primer design program Primer3. Bioinformatics 23, 1289–1291. doi: 10.1093/bioinformatics/btm091
Li, H. (2011a). A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993. doi: 10.1093/bioinformatics/btr509
Li, H. (2011b). Improving SNP discovery by base alignment quality. Bioinformatics 27, 1157–1158. doi: 10.1093/bioinformatics/btr076
Li, H., and Durbin, R. (2009). Fast and Accurate Short Read Alignment with Burrows–Wheeler Transform. Oxford: Oxford University Press.
Luo, R., Liu, B., Xie, Y., Li, Z., Huang, W., Yuan, J., et al. (2012). SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1:18. doi: 10.1186/2047-217X-1-18
Price, A. L., Patterson, N. J., Plenge, R. M., Weinblatt, M. E., Shadick, N. A., and Reich, D. (2006). Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909. doi: 10.1038/ng1847
Pritchard, J. K., Stephens, M., and Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics 155, 945–959.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., and Kumar, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729. doi: 10.1093/molbev/mst197
Thiel, T., Michalek, W., Varshney, R., and Graner, A. (2003). Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor. Appl. Genet. 106, 411–422. doi: 10.1007/s00122-002-1031-0
Untergasser, A., Cutcutache, I., Koressaar, T., Ye, J., Faircloth, B. C., Remm, M., et al. (2012). Primer3—new capabilities and interfaces. Nucleic Acids Res. 40, e115. doi: 10.1093/nar/gks596
Wang, K., Li, M., and Hakonarson, H. (2010). ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164. doi: 10.1093/nar/gkq603
Winter, P., Pfaff, T., Udupa, S., Hüttel, B., Sharma, P., Sahi, S., et al. (1999). Characterization and mapping of sequence-tagged microsatellite sites in the chickpea (Cicer arietinum L.) genome. Mol. Gen. Genet. 262, 90–101. doi: 10.1007/s004380051063
Yao, G., Ye, L., Gao, H., Minx, P., Warren, W. C., and Weinstock, G. M. (2012). Graph accordance of next-generation sequence assemblies. Bioinformatics 28, 13–16. doi: 10.1093/bioinformatics/btr588
Keywords: date palm, short sequence repeat, single nucleotide polymorphism, genome variation, cultivar classification
Citation: He Z, Zhang C, Liu W, Lin Q, Wei T, Aljohi HA, Chen W-H and Hu S (2017) DRDB: An Online Date Palm Genomic Resource Database. Front. Plant Sci. 8:1889. doi: 10.3389/fpls.2017.01889
Received: 15 August 2017; Accepted: 18 October 2017;
Published: 02 November 2017.
Edited by:
Shrikant S. Mantri, National Agri-Food Biotechnology Institute, IndiaReviewed by:
Sumit Kumar Bag, National Botanical Research Institute (CSIR), IndiaXiyin Wang, North China University of Science and Technology, China
Copyright © 2017 He, Zhang, Liu, Lin, Wei, Aljohi, Chen and Hu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hasan A. Aljohi, aGFsam9oaUBrYWNzdC5lZHUuc2E= Wei-Hua Chen, d2VpaHVhY2hlbkBodXN0LmVkdS5jbg== Songnian Hu, aHVzbkBiaWcuYWMuY24=
†These authors have contributed equally to this work.
 Zilong He
Zilong He Chengwei Zhang1,2,3†
Chengwei Zhang1,2,3† Wanfei Liu
Wanfei Liu Qiang Lin
Qiang Lin Ting Wei
Ting Wei Hasan A. Aljohi
Hasan A. Aljohi Wei-Hua Chen
Wei-Hua Chen Songnian Hu
Songnian Hu