- 1Department of Life Science, College of Science, Rikkyo University, Tokyo, Japan
- 2Graduate School of Science, The University of Tokyo, Tokyo, Japan
- 3Okazaki Institute for Integrative Bioscience, Okazaki, Japan
- 4Research Center for Life Science, College of Science, Rikkyo University, Tokyo, Japan
Leaf abaxial–adaxial patterning is dependent on the mutual repression of leaf polarity genes expressed either adaxially or abaxially. In Arabidopsis thaliana, this process is strongly affected by mutations in ribosomal protein genes and in ribosome biogenesis genes in a sensitized genetic background, such as asymmetric leaves2 (as2). Most ribosome-related mutants by themselves do not show leaf abaxialization, and one of their typical phenotypes is the formation of pointed rather than rounded leaves. In this study, we characterized two ribosome-related mutants to understand how ribosome biogenesis is linked to several aspects of leaf development. Previously, we isolated oligocellula2 (oli2) which exhibits the pointed-leaf phenotype and has a cell proliferation defect. OLI2 encodes a homolog of Nop2 in Saccharomyces cerevisiae, a ribosome biogenesis factor involved in pre-60S subunit maturation. In this study, we found another pointed-leaf mutant that carries a mutation in a gene encoding an uncharacterized protein with a G-patch domain. Similar to oli2, this mutant, named g-patch domain protein1 (gdp1), has a reduced number of leaf cells. In addition, gdp1 oli2 double mutants showed a strong genetic interaction such that they synergistically impaired cell proliferation in leaves and produced markedly larger cells. On the other hand, they showed additive phenotypes when combined with several known ribosomal protein mutants. Furthermore, these mutants have a defect in pre-rRNA processing. GDP1 and OLI2 are strongly expressed in tissues with high cell proliferation activity, and GDP1-GFP and GFP-OLI2 are localized in the nucleolus. These results suggest that OLI2 and GDP1 are involved in ribosome biogenesis. We then examined the effects of gdp1 and oli2 on adaxial–abaxial patterning by crossing them with as2. Interestingly, neither gdp1 nor oli2 strongly enhanced the leaf polarity defect of as2. Similar results were obtained with as2 gdp1 oli2 triple mutants although they showed severe growth defects. These results suggest that the leaf abaxialization phenotype induced by ribosome-related mutations is not merely the result of a general growth defect and that there may be a sensitive process in the ribosome biogenesis pathway that affects adaxial–abaxial patterning when compromised by a mutation.
Introduction
Ribosome biogenesis drives cellular growth, and, in principle, individual cells must grow twofold before division during the cell cycle. To do so, a large proportion of the gene expression machinery in a cell is devoted to ribosome biogenesis (Warner, 1999). In multicellular organisms, behavior of individual cells is under the control of developmental programs to establish the appropriate shape and function of tissues and organs. Therefore, ribosome biogenesis is expected to be important to developmental pattering. However, the details of this putative connection are not well understood.
In eukaryotes, cytosolic ribosomes consist of a 60S subunit and a 40S subunit. Ribosome biosynthesis initiates in a specialized membraneless nuclear subcompartment, the nucleolus. Pre-ribosomal RNA [pre-rRNA: 45S rRNA in Arabidopsis thaliana (Arabidopsis) and 35S rRNA in Saccharomyces cerevisiae (yeast)] is transcribed by RNA polymerase I (Pol I) as a polycistronic transcript from rDNA repeats and contains 25–28S, 5.8S, and 18S rRNA sequences that are flanked by a 5′ external transcribed spacer (5′-ETS) and a 3′-ETS and separated by internal transcribed spacer 1 (ITS1) and ITS2 (Layat et al., 2012). Ribosome biogenesis requires more than 200 ribosome biogenesis factors (RBFs) and small nucleolar RNA (snoRNA) species in addition to ribosomal proteins (r-proteins), and has been best characterized in yeast. The 90S pre-ribosome, also designated as the small subunit (SSU) processome, is a huge ribonucleoprotein complex in which a nascent pre-rRNA and a subset of r-proteins for SSU are encapsulated by U three protein (UTP) complexes and U3 small nucleolar ribonucleoprotein (snoRNP). Numerous RBFs dynamically join and dissociate from the 90S pre-ribosome in a hierarchical manner and carry out folding, cleavage, and trimming of rRNA precursors, as well as assembly of r-proteins with rRNAs to produce pre-40S ribosomes (Grandi et al., 2002; Pérez-Fernández et al., 2007; Kornprobst et al., 2016; Chaker-Margot et al., 2017; Sun et al., 2017). The remaining 3′ part of the pre-rRNA forms the large subunit (LSU) processome and produces pre-60S ribosomes (Konikkat and Woolford, 2017). The 5S rRNA is transcribed separately by Pol III and forms a complex with RPL5 and RPL11, then joins to a pre-60S particle (Zhang et al., 2007). Final maturation of pre-ribosomal subunits takes place after export from the nucleus into the cytoplasm (Nerurkar et al., 2015).
Approximately 250 ribosome biogenesis factors have been identified in yeast by genetic and proteomics analyses. Similarly, 286 ribosome biogenesis factors have been identified in human cells, but among them, 74 do not have a yeast ortholog (Tafforeau et al., 2013). For land plants, orthologs to about 75% of yeast ribosome biogenesis factors were identified by bioinformatics methods (Ebersberger et al., 2013). However, only a small fraction of these molecules have been functionally characterized in Arabidopsis (Weis et al., 2015a). These studies suggest that molecular mechanisms for ribosome biogenesis are largely conserved, but that some processes are mediated by species-specific factors. A recent nucleolar proteomics analysis of Arabidopsis supported this interpretation (Palm et al., 2016).
A plant-specific feature of ribosome biogenesis is also seen in two routes of rRNA processing (Weis et al., 2015a). In Arabidopsis, transcription of 45S rRNA is terminated by endonucleolytic cleavage at site B0 in the 3′-ETS by RIBONUCLEASE THREE LIKE2 (atRTL2; Comella et al., 2007). Then endonucleolytic cleavage at site P within the 5′-ETS takes place by the action of EXORIBONUCLEASE2 (XRN2) and U3 snoRNP to yield 35S rRNA (Sáez-Vasquez et al., 2004; Zakrzewska-Placzek et al., 2010). There are two alternative routes for further processing of 35S rRNA accompanied by simultaneous removal of the 3′-ETS (Missbach et al., 2013). In one pathway, the 5′-ETS is removed before cleavage within ITS1 (5′-ETS-first pathway) while in the other, cleavage within ITS1 of 35S takes place prior to 5′-ETS cleavage (ITS1-first pathway). In yeast, processing of pre-rRNA strictly follows the 5′-ETS-first pathway, while the ITS1-first pathway is the major route in mammals (Fernández-Pevida et al., 2015; Henras et al., 2015).
In Arabidopsis, impaired function of RBFs affects normal processing of pre-rRNAs as well as diverse developmental processes, such as auxin response, cell proliferation, root epidermal patterning, vascular patterning, leaf shape regulation, callus formation, and development of the gynoecium, embryo, and female gametophyte (Shi et al., 2005; Griffith et al., 2007; Petricka and Nelson, 2007; Lange et al., 2008, 2011; Fujikura et al., 2009; Li et al., 2009; Abbasi et al., 2010; Huang et al., 2010; Liu et al., 2010; Ohbayashi et al., 2011; Wieckowski and Schiefelbein, 2012; Cho et al., 2013; Kumakura et al., 2013; Missbach et al., 2013; Hang et al., 2014; Weis et al., 2014, 2015b). Many of these phenotypes are also shared by mutants defective in a gene for an r-protein (for review, see Byrne, 2009; Horiguchi et al., 2012; Machida et al., 2015). These findings suggest that production and/or function of ribosomes may be associated with developmental regulation. Recently, impaired ribosome biogenesis in mutants, such as root initiation defective2 (rid2), was shown to induce ribosomal stress (Ohbayashi et al., 2017). Explants from rid2 are unable to form callus at high temperature and produce pointed leaves, but these phenotypes are suppressed by mutations in the NO APICAL MERISTEM, ARABIDOPSIS TRANSCRIPTION ACTIVATION FACTOR1/2 and, CUP-SHAPED COTYLEDON2 (NAC) transcription factor gene SUPPRESSOR OF RID TWO1 (SRIW1)/ANAC082 (Ohbayashi et al., 2017). Notably, rid2 mutants are defective in the processing of rRNAs, but this phenotype is not suppressed by sriw1, indicating that SRIW1 mediates ribosomal stress to induce developmental alterations (Ohbayashi et al., 2017).
The potential developmental roles of ribosomes have been further expanded by observations that mutants defective in r-protein significantly enhance the leaf polarity defect of asymmetric leaves1 (as1) and as2 (Pinon et al., 2008; Yao et al., 2008; Horiguchi et al., 2011; Szakonyi and Byrne, 2011; Casanova-Sáez et al., 2014) and revoluta (rev) (Pinon et al., 2008). In the double mutants, leaves fail to expand into a flat laminar structure, but form in a trumpet- or needle-like shape. Normally, the vascular tissues are organized such that xylem and phloem tissues face the adaxial (upper) and abaxial (lower) sides of the leaves, respectively (Waites and Hudson, 1995). In contrast, vascular tissues in radialized leaves are organized with the phloem tissues surrounding the vascular tissues, or in extreme cases the vascular tissues are absent (Pinon et al., 2008; Yao et al., 2008; Horiguchi et al., 2011). These double mutants have increased or expanded expression of abaxially expressed genes (Pinon et al., 2008; Yao et al., 2008; Horiguchi et al., 2011), suggesting that mutations in r-protein genes enhance leaf abaxialization in the as mutant backgrounds. A similar leaf polarity defect was also found in mutants defective in Arabidopsis PUMILIO23 (APUM23), which encodes a member of the Nop9 family involved in regulation of rRNA processing (Thomson et al., 2007; Abbasi et al., 2010; Droll et al., 2010; Huang et al., 2014; Zhang and Muench, 2015; Zhang et al., 2016). More recently, mutations in RNA HELICASE10 (RH10), NUCLEOLIN-LIKE1 (NUC-L1), and RID2 were also shown to enhance the leaf polarity defect of as2 (Matsumura et al., 2016). RH10 is a member of the DEAD-box RNA helicases, putative orthologs of which are Rrp3 in yeast and DDX47 in human (O’Day et al., 1996; Matsumura et al., 2016). NUC-L1 is an ortholog of nucleolin in human and plays multiple roles in ribosome biogenesis (Sáez-Vasquez et al., 2004). RID2 is a homolog of Bud23 in yeast, which is a methyltransferase that catalyzes the methylation of G1575 in 18S rRNA (Ohbayashi et al., 2011; Létoquart et al., 2014). Yeast and human orthologs of RH10, NUC1, and RID2 are components of, or are associated with, the SSU processome (Turner et al., 2009; Phipps et al., 2011; Sardana et al., 2013). These findings imply that correct ribosome production and/or function are required for leaf adaxial/abaxial patterning (Matsumura et al., 2016).
To understand the mechanisms of ribosome biogenesis in plants and how it is connected to developmental programs, we have been trying to genetically identify individual RBFs and have characterized their molecular and developmental phenotypes. Although ribosome biogenesis is highly complicated, comparison of phenotypes among different RBF mutants would provide information concerning the general and specific functions of each factor. The purpose of this study is to find differential requirement of RBFs in two major leaf developmental processes, namely cell proliferation and abaxial–adaxial pattering. In this study, we found a novel gene for a nucleolar protein, G-PATCH DOMAIN PROTEIN1 (GDP1), and characterized it together with previously identified OLIGOCELLULA2 (OLI2; Fujikura et al., 2009), which is a homolog of yeast nucleolar protein Nop2 involved in formation of 5-methylcytosine (m5C) at C2870 in 25S rRNA (Sharma et al., 2013). Loss-of-function mutants of GDP1 and OLI2 exhibited a range of phenotypes frequently found in mutants defective in r-proteins or RBFs. Interestingly, gdp1, oli2, and double mutants between them did not show strongly enhanced as2 leaf polarity defects despite their association with significantly impaired cell proliferation in leaves. These results suggest that the roles of ribosome biogenesis/function in leaf adaxial/abaxial patterning and other developmental processes are at least partially separate.
Materials and Methods
Plant Materials
The wild type accession of Arabidopsis used in this study was Columbia-0 (Col-0). T-DNA insertion lines (Salk_065904 [gdp1-1], Salk_041661 [gdp1-2]) were obtained from the Arabidopsis Biological Resource Center (ABRC). gdp1-3 was previously identified as #416 (Horiguchi et al., 2006a,b), and oli2-1 and oli2-2 (Salk_129648) were reported previously (Fujikura et al., 2009). Seeds were sown on rockwool and grown at 22°C under a 16-h light/8-h dark photoperiod for quantitative characterization of leaf phenotypes and RNA preparation. Seedlings were watered daily with 0.5 g L-1 of Hyponex (Hyponex Japan). For fluorescence imaging of root, seedlings were grown for 5 days on half-strength Murashige and Skoog (MS) medium supplemented with Gamborg B5 vitamins and 3% (w/v) sucrose, and solidified with 0.5% (w/v) gellan gum at 22°C under a 16-h light/8-h dark photoperiod. Oligonucleotide pairs for genotyping of gdp1 alleles are listed in Supplementary Table S1.
Generation of Transgenic Plants
An approximately 1.6-kb GDP1 promoter region with or without the GDP1 transcribed region was amplified and cloned into pDONR201 using BP clonase (Thermo) followed by transfer of the insert into pBGGUS (Horiguchi et al., 2005) or pHWG (Horiguchi and Tsukaya, unpublished) with LR clonase II (Thermo) to yield pGDP1::GUS and pGDP1::GDP1-GFP constructs, respectively. For construction of pOLI2::GUS, an OLI2 promoter DNA fragment and GUS cDNA were cloned into pDONR P4P1R and pENTR/D-TOPO (Thermo), respectively, and combined into the Gateway binary vector, pGWB501 (Nakagawa et al., 2007), with LR clonase II plus (Thermo). For construction of pOLI2::GFP-OLI2, a 3.6-kb promoter region of OLI2, GFP cDNA, and a transcribed region plus a 4.1-kb 3′ untranscribed region of OLI2 were amplified and cloned into pSMAH621 digested with HindIII and SacI using an In-Fusion HD cloning kit (Clontech). These constructs were introduced into Arabidopsis by the floral dip method (Clough and Bent, 1998). Transgenic plants with a single T-DNA insertion were selected and homozygous T3 plants were used. Oligonucleotides used in the construction of these vectors are listed in Supplementary Table S1.
RNA Analyses
Total RNA was prepared from shoots of wild type, gdp1, oli2, and gdp1 oli2 using Trizol reagent (Thermo). Isolated RNA was treated with DNase I (Thermo) followed by first-strand cDNA synthesis with SuperScript III reverse transcriptase (Thermo) primed either with oligo (dT) (for messenger RNA) or random hexamers (for rRNAs). The cDNAs were subjected to semi-quantitative and quantitative RT-PCR (RT-qPCR). RT-qPCR was carried out using GoTaq qPCR master mix (Promega) with an ABI7500 real-time PCR system (Thermo) by the ΔΔCt method. The expression level of ACTIN2 (ACT2) was used as an endogenous control for mRNAs, while 18S rRNA was used as an endogenous control to determine rRNA intermediate levels. Oligonucleotide pairs used to detect AUXIN RESPONSE FACTOR3 (ARF3), KANADI2 (KAN2), YABBY5 (YAB5), PHABOLUSA (PHB), PHAVOLUTA (PHV), REV, and ACT2 were described previously (Iwakawa et al., 2007). Other oligonucleotide pairs are listed in Supplementary Table S1.
For detection of rRNA intermediates by Northern hybridization, aliquots of 3 μg of total RNAs isolated from 12-day-old shoots were separated by formaldehyde-agarose gel electrophoresis (1.2%, w/v) and transferred onto nylon membranes by downward capillary transfer. After ultraviolet crosslinking of RNA to the nylon membranes, hybridization was carried out using a DIG Northern starter kit (Sigma–Aldrich) and hybridization signals were detected using a digital imaging system (LAS 4000 mini; GE Healthcare). RNA probes were generated using 5′-ETS, ITS1, and ITS2 DNA fragments amplified with the appropriate oligonucleotide pairs (Supplementary Table S1) and T7 RNA polymerase.
Microscopic Observation
Quantitative analyses of leaves were carried out as described (Horiguchi et al., 2005). Plants were gron for 25 days and first leaves were fixed in a formalin/acetic acid/alcohol [FAA, formalin: acetic acid: 70% (v/v) ethanol = 1:1:18] solution, cleared using a chloral hydrate solution (200 g chloral hydrate, 20 g glycerol, and 50 ml dH2O) and observed by stereomicroscope (M165FC; Leica) and differential contrast interference microscope (DM2500; Leica), respectively. Leaf blade area and the projection area of palisade cells in adaxial subepidermal layer was determined using Image J1. For each leaf, the palisade cell area was determined by mearuing at least 20 cells. Average palisade cell size was determined using data from 10 leaves. Cell density in adaxial subepidermal layer was manuary determined by counting cells in a unit area. The number of cells in adaxial subepidermal layer per leaf was estimated by dividing the leaf blade area with the cell density. For fluorescence imaging of pOLI2::GFP-OLI2/oli2-1 and pGDP1::GDP1-GFP/gdp1-1 lines, roots of 5-day-old seedlings were fixed in phosphate buffered saline (PBS, pH 7.0) containing 4% (w/v) paraformaldehyde, washed twice with PBS, and stained with 4′,6-diamidine-2′-phenylindole dihydrochloride (DAPI; Merck) or Calcofluor White M2R (Merck) and cleared with TOMEI-II (Hasegawa et al., 2016). For fluorescence imaging of leaves, 14-day-old shoots grown on rock wool were fixed in PBS [pH 7.0 containing 4% (w/v) paraformaldehyde, washed twice with PBS, and treated with TOMEI-II]. Fluorescent signals were observed with a confocal laser scanning microscope (LSM710 or LSM800; Zeiss). Histochemical staining of promoter::GUS lines was carried out according to Donnelly et al. (1999).
Results
Identification of gdp1 Mutants and Characterization of Their Vegetative Phenotypes
During the course of studying mutants with altered leaf size and shape, we found that two T-DNA insertion lines of At1g63980 (Salk_065904 and Salk_041661) exhibited the “pointed-leaves” phenotype that is typically observed in mutants defective in r-protein genes and ribosome biogenesis genes (Figures 1A–C,E). An additional allele was also found in our mutant collection reported previously (#416; Horiguchi et al., 2006a,b) that had a 26-bp deletion in the third exon and a point mutation in the third intron (Figures 1A–C). As At1g63980 was not a characterized gene and it was apparent that At1g63980 does not encode an r-protein, we decided to characterize these mutants in relation to ribosome biogenesis.
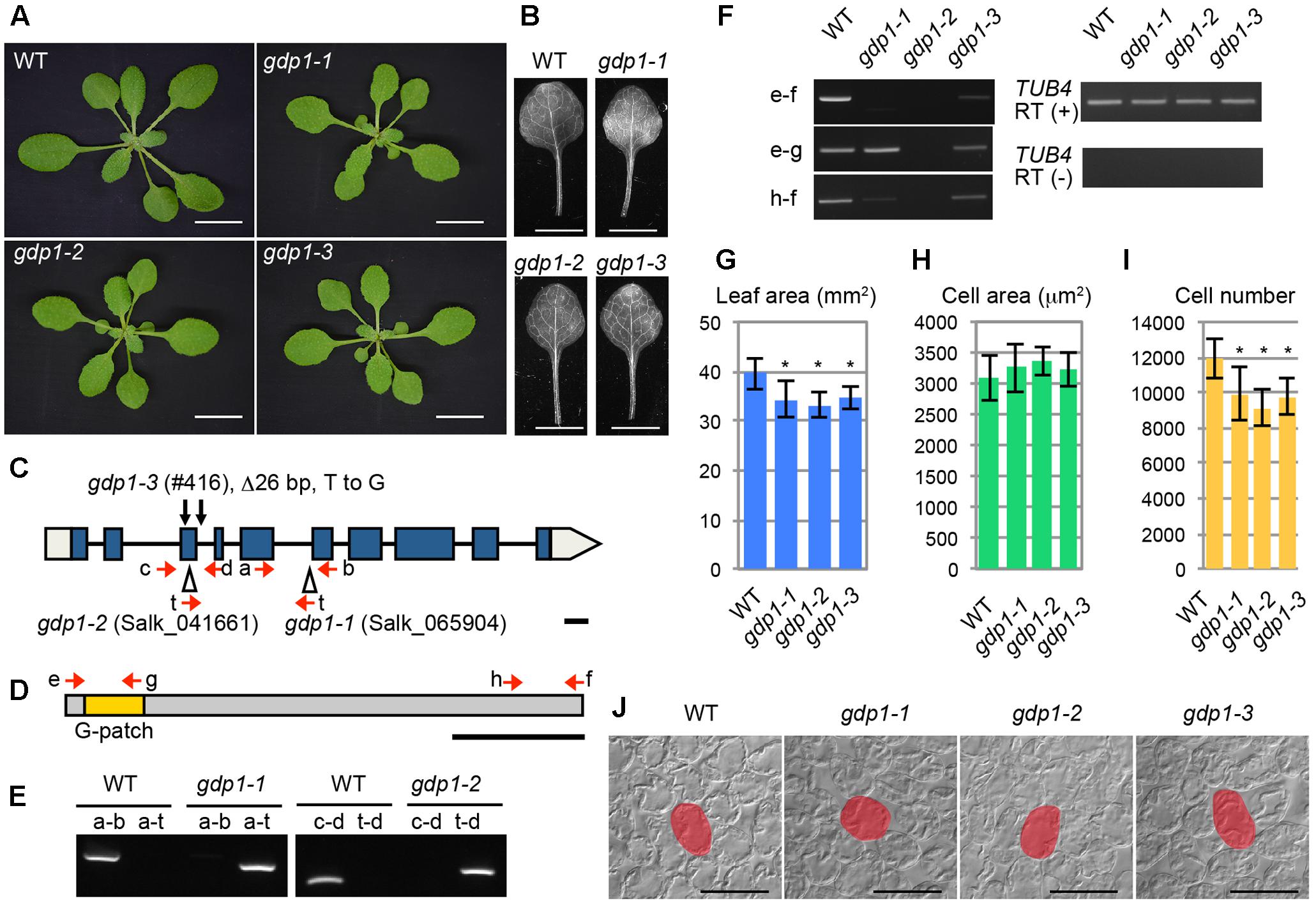
FIGURE 1. Genetic analyses of gdp1 alleles. (A) Shoots of wild type (WT) and gdp1-1, gdp1-2, and gdp1-3 alleles grown for 25 days. Bars, 1 cm. (B) The first leaves of WT and gdp1-1, gdp1-2, and gdp1-3 alleles. Bar, 5 mm. (C) Schematic diagram of mutation points of gdp1 alleles. Red arrows indicate oligonucleotides used for genotyping of gdp1 alleles. Black arrows indicate the mutation points in gdp1-3. T-DNA insertions in gdp1-1 and gdp1-2 are indicated by triangles. Bar, 100 bp. (D) Schematic diagram of GDP1 protein. Arrows indicate the positions of oligonucleotides on the corresponding GDP1 transcript used for RT-PCR analysis. Bar, 100 amino acid residues. (E) Genotyping of gdp1-1 and gdp1-2 alleles. Oligonucleotide pairs indicated by letters were used to amplify genomic DNA fragments from the WT and the two gdp1 alleles. (F) RT-PCR analysis of GDP1 transcripts. Total RNAs were isolated from 10-day-old shoots and subjected to reverse transcription followed by PCR with oligonucleotide pairs indicated by letters. (G–I) Quantitative analyses of leaf phenotypes. The leaf blade area (G), the projection area of leaf mesophyll cells (H), and the number of leaf mesophyll cells (I) in the adaxial subepidermal layer of palisade tissues of first leaves are shown. First leaves were harvested from 25-day-old plants. n = 10, mean ± SD. Asterisks indicate statistically significant differences compared to WT (P < 0.05, Student’s t-test). (J) Leaf palisade cells observed from a paradermal view. Bar, 100 μm. Representative cells are highlighted.
The At1g63980 gene product has a G-patch domain near its amino-terminal region (Figure 1D). The G-patch domain is about 48 amino acid residues in length and contains several conserved glycine residues (Supplementary Figure S1). This domain is often found in RNA processing proteins with or without known RNA-binding motifs (Aravind and Koonin, 1999). Therefore, we named At1g63980 G-PATCH DOMAIN PROTEIN1 (GDP1), and Salk_065904, Salk_041661, and #416 were designated as gdp1-1, gdp1-2, and gdp1-3, respectively (Figure 1C). The Arabidopsis genome includes at least 15 G-patch domain-containing protein genes. The structures of individual G-patch domain proteins differ in the location and number of G-patch domains, combination of RNA-binding motifs, and amino acid lengths (Supplementary Figure S1). Among them, GDP1 appears to be a single-copy gene.
The two T-DNA insertion alleles of gdp1 did not accumulate GDP1 transcripts at a detectable level, while gdp1-3 showed GDP1 transcript accumulation at a lower level than wild type, as determined by RT-PCR using an oligonucleotide pair that amplifies the whole coding region (Figure 1F). When oligonucleotide pairs designed to amplify partial GDP1 cDNA fragments corresponding to the 5′ and 3′ regions of the coding sequence were used, the 5′ fragment was amplified in gdp1-1 at a level similar to the wild type, but neither the 5′ nor 3′ regions were detectable in gdp1-2 (Figure 1F). In gdp1-3, both fragments were detected at lower levels than in wild type (Figure 1F). These results suggest that gdp1-2 is a null allele. We also considered both gdp1-1 and gdp1-3 to be strong loss-of-function alleles as their phenotypes were almost identical to gdp1-2, as shown below.
We examined the leaf phenotypes of gdp1 at the cellular level. Both the area of the leaf blade and the number of palisade cells in the subepidermal layer of the first leaves were reduced by about 20% in gdp1 compared to the wild type (Figures 1G,I). On the other hand, the projection area of palisade cells was similar in both gdp1 and wild type (Figures 1H,J). These phenotypes were similar to those observed in oli2, oli7, and oli5, the latter two of which are defective in paralogous r-protein genes, RPL5A and RPL5B (Fujikura et al., 2009). These oli mutants showed strongly enhanced cell enlargement in leaves of angustifolia3 (an3), which is defective in a transcription coactivator (Horiguchi et al., 2005) by further reducing the leaf cell number (Fujikura et al., 2009). Similar to these mutants, when gpd1-1 was crossed with an3-4, the resultant double mutant showed a further decrease in number of leaf palisade cells in the subepidermal layer (86% fewer than wild type) and triggered excessive cell enlargement (Figure 2; 237% larger than wild type), which is known as compensated cell enlargement (Horiguchi and Tsukaya, 2011; Hisanaga et al., 2015).
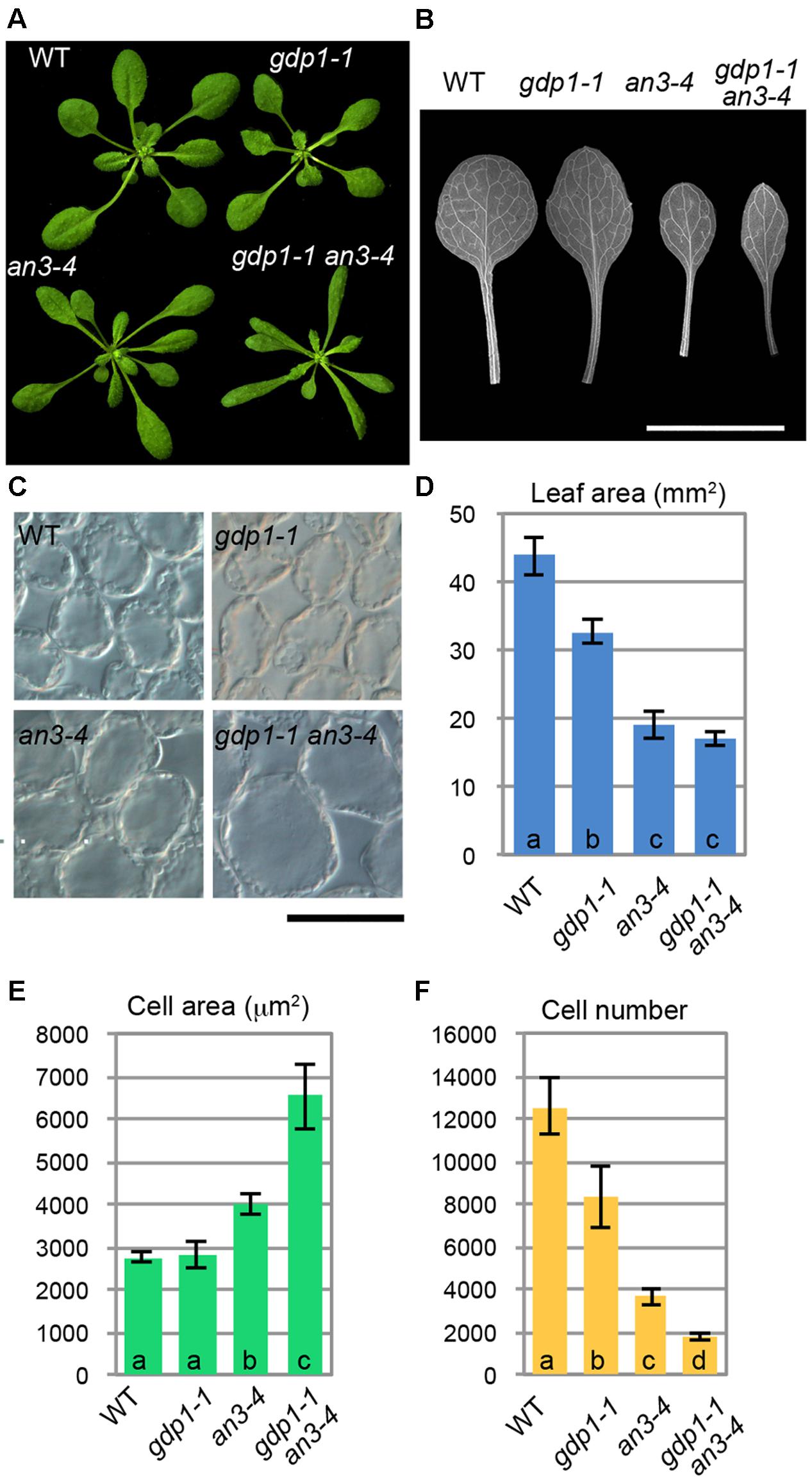
FIGURE 2. Genetic interaction between gdp1 and an3. (A) Shoots of 25-day-old WT, gdp1-1, an3-4, and gdp1-1 an3-4. (B) The first leaves of WT, gdp1-1, an3-4, and gdp1-1 an3-4. Bar, 1 cm. (C) Leaf palisade cells observed from a paradermal view. Bar, 100 μm. (D–F) Quantitative analyses of leaf phenotypes. Leaf blade area (D), the projection area of leaf mesophyll cells (E), and the number of leaf mesophyll cells (F) in the adaxial subepidermal layer of palisade tissues of first leaves are shown. First leaves were harvested from 25-day-old plants. n = 10, mean ± SD. Statistically significant differences were indicated by different letters (one-way ANOVA with Tukey HSD test, p < 0.05).
The above developmental phenotypes are quite similar to those observed in mutants defective in r-protein genes and ribosome biosynthesis genes. To examine potential genetic interactions between gdp1 and these ribosome-related mutants, gdp1-1 was crossed with various r-protein defective mutants reported previously (Figure 3; Horiguchi et al., 2011). Generally, these double mutants showed an additive phenotype as determined from the number of leaf palisade cells (Figure 3D). The double mutants showed reduction of leaf palisade cells in the subepidermal layer by 13–39% when compared with respective parental r-protein mutants (Figure 3D). In relation to the projection area of palisade cells, the double mutants tended to have a larger cell size than parental r-protein mutants, showing the occurrence of compensated cell enlargement (Figure 3C). Consequently, the overall shoot size and first leaf size were less significantly reduced from single r-protein mutants (Figures 3A,B). A unique exception was found in rps6a-2. Strikingly, gdp1-1 rps6a-2 had 66% fewer leaf palisade cells in the subepidermal layer when compared with rps6a-2, suggesting a synergistic interaction between these two mutations (Figure 3D).
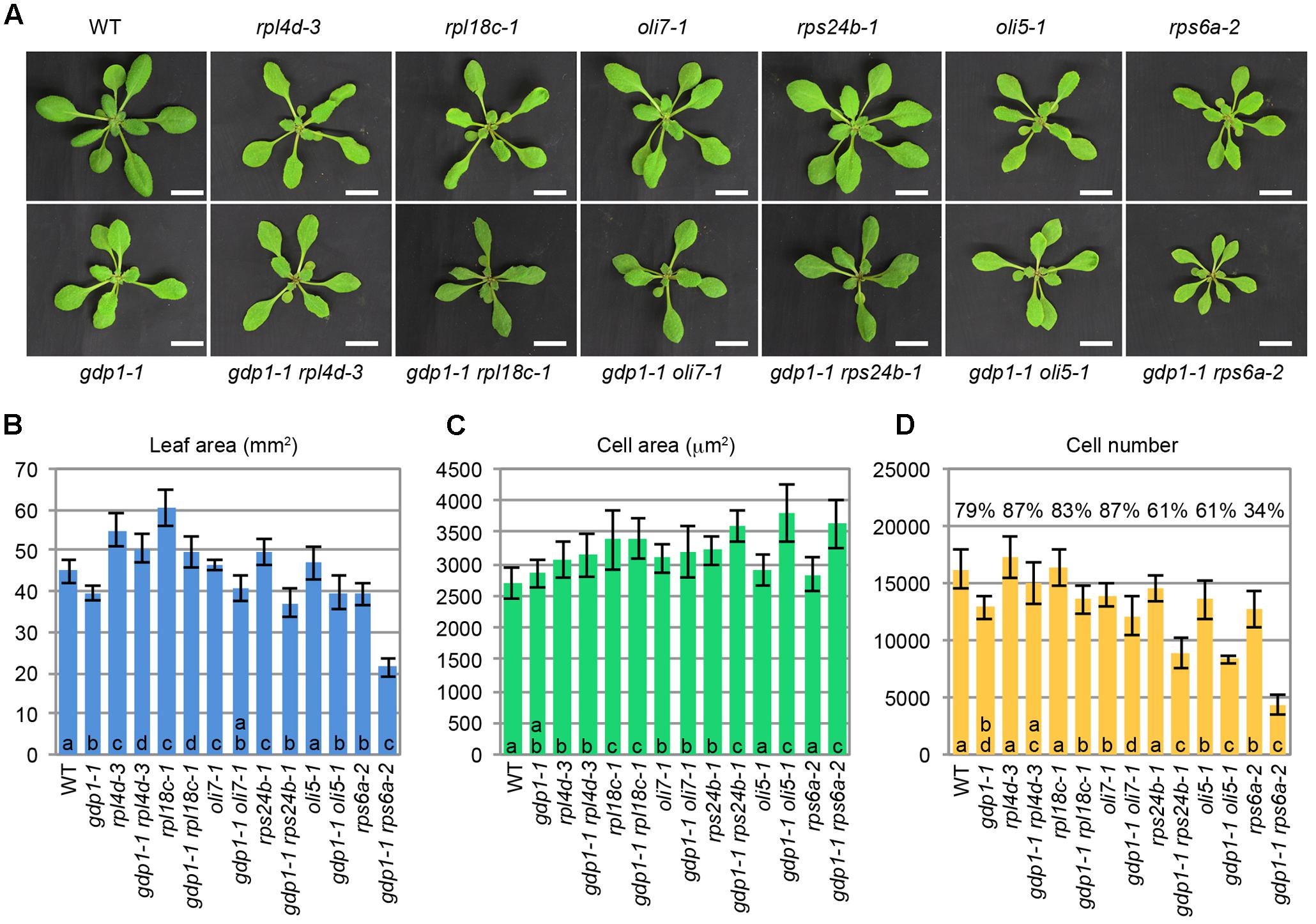
FIGURE 3. Genetic interactions between gdp1 and various r-protein mutants. (A) Shoots of 25-day-old WT, gdp1, r-protein mutants, and double mutants between them. Bar, 1 cm. (B–D) Quantitative analyses of leaf phenotypes. Leaf blade area (B), the projection area of leaf mesophyll cells (C), and the number of leaf mesophyll cells (D) in the adaxial subepidermal layer of palisade tissues of first leaves are shown. First leaves were harvested from 25-day-old plants. n = 10, mean ± SD. Statistically significant differences among four plant lines namely, wild type, gdp1-1, single r-protein mutants, and double mutants between gdp1 and the r-protein mutants were indicated by different letters (one-way ANOVA with Tukey HSD test, p < 0.05). In (D), relative cell number in gdp1 and double mutants on the basis of WT and parental single r-protein mutants, respectively, are shown.
We also examined genetic interactions between gdp1-1 and oli2-1 (Figure 4). The double mutants between gdp1-1 and oli2-1 produced smaller shoots and narrower leaves than the respective parental mutants (Figures 4A,B,D). The double mutants frequently produced monocots or tricots that were rarely observed in the single mutants (Supplementary Figure S2). Although the effect of each single mutation on the number of leaf palisade cells in the subepidermal layer was relatively weak (only 10–30% reduction), the double mutants showed a reduction by about 65% compared to the wild type level (Figure 4F). The projection area of leaf palisade cells was more than 83% larger in gdp1-1 oli2-1 than the wild type, indicating compensated cell enlargement (Figures 4C,E). These phenotypes were found in different allelic combinations, gdp1-3 oli2-2 (Figure 4). As OLI2 encodes a putative m5C methyltransferase and likely participates in ribosome biogenesis (Fujikura et al., 2009), the strong genetic interaction between gdp1 and oli2 suggests a role of GDP1 in ribosome biogenesis.
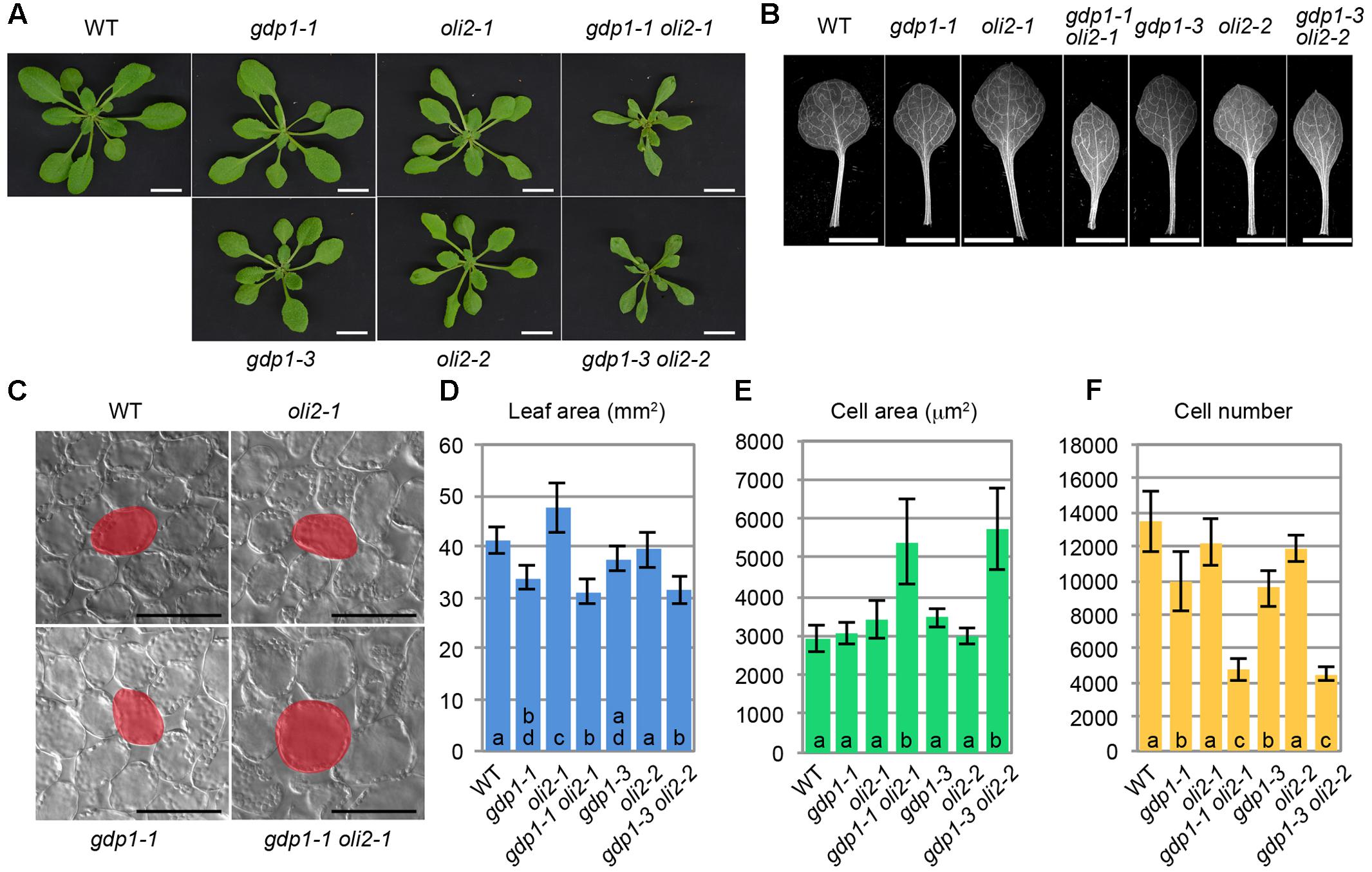
FIGURE 4. Genetic interaction between gdp1 and oli2. (A) Shoots of 25-day-old WT, gdp1 alleles, oli2 alleles, and double mutants between them. Bars, 1 cm. (B) First leaves. Bars, 5 mm. (C) Leaf palisade cells observed from a paradermal view. Bar, 100 μm. (D–F) Quantitative analyses of leaf phenotypes. Leaf blade area (D), the projection area of leaf mesophyll cells (E), and the number of leaf mesophyll cells (F) in the adaxial subepidermal layer of palisade tissues of first leaves are shown. First leaves were harvested from 25-day-old plants. n = 10, mean ± SD. Statistically significant differences were indicated by different letters (one-way ANOVA with Tukey HSD test, p < 0.05).
Tissue-Specific GDP1 and OLI2 Expression and Subcellular Localization of Their Gene Products
When GDP1 expression was observed in a pGDP1::GUS transgenic line, relatively strong GUS staining was detected in the shoot tip, young leaf primordium, root tip, and floral buds (Figures 5A–D). Active ribosome biogenesis takes place in the proliferating cell population. Therefore, GDP1 expression in actively developing tissues is consistent with the expected function of GDP1 in ribosome biogenesis. On the other hand, pOLI2::GUS transgenic lines showed GUS staining in guard cells and the basal parts of lateral roots rather than in these tissues (Figures 5E,F). We considered this result to indicate that the promoter region used in the transgenic line was insufficient to show the authentic expression pattern of OLI2. To overcome this problem, we generated pOLI2::GFP-OLI2/oli2-1 lines. These transgenic lines fully complemented the leaf shape of oli2-1 (Figures 6D–F). In relation to cell proliferation, the two pOLI2::GFP-OLI2/oli2-1 lines had even greater numbers of leaf palisade cells in the subepidermal layer than the wild type (Figure 6F). In contrast to the pOLI2::GUS lines, strong GFP-OLI2 signals were observed in leaf primordia and root apical meristem (Figures 5H,J). We also generated two pGDP1::GDP1-GFP/gdp1-1 lines that also complemented the gdp1 leaf phenotypes (Figures 6A–C). GDP1-GFP signals were also found in root tips and leaf primordia (Figures 5G,I). In addition, strong expression levels of GDP1 and OLI2 were found using the electronic Fluorescent Pictograph (eFP) Browser (Winter et al., 2007). These results suggest that both GDP1 and OLI2 are strongly expressed in actively growing tissues that have a high demand for ribosome production.

FIGURE 5. Tissue-specific expression analysis of GDP1 and OLI2. (A–D) Histochemical staining of pGDP1::GUS plants. (A) A 4-day-old seedling. (B) Primary root tip. (C) A 12-day-old seedling. (D) Inflorescence. (E,F) Histochemical staining of pOLI2::GUS plants. (E) A 5-day-old seedling. The insert shows a close-up view of guard cells in a cotyledon. (F) A lateral root. (G,H) Confocal images of 5-day-old pGDP1::GDP1-GFP/gdp1-1 (G,I) and pOLI2::GFP-OLI2/oli2-1 (H,J). (G,H) Leaf primordia. (I,J) Primary root tips. Calcofluor fluorescence, GFP fluorescence, and merged images are shown from left to right. Bars in (A,C–E): 1 mm, (B,F,I,J): 100 μm, insert in (E): 50 μm, (G,H): 0.5 mm.
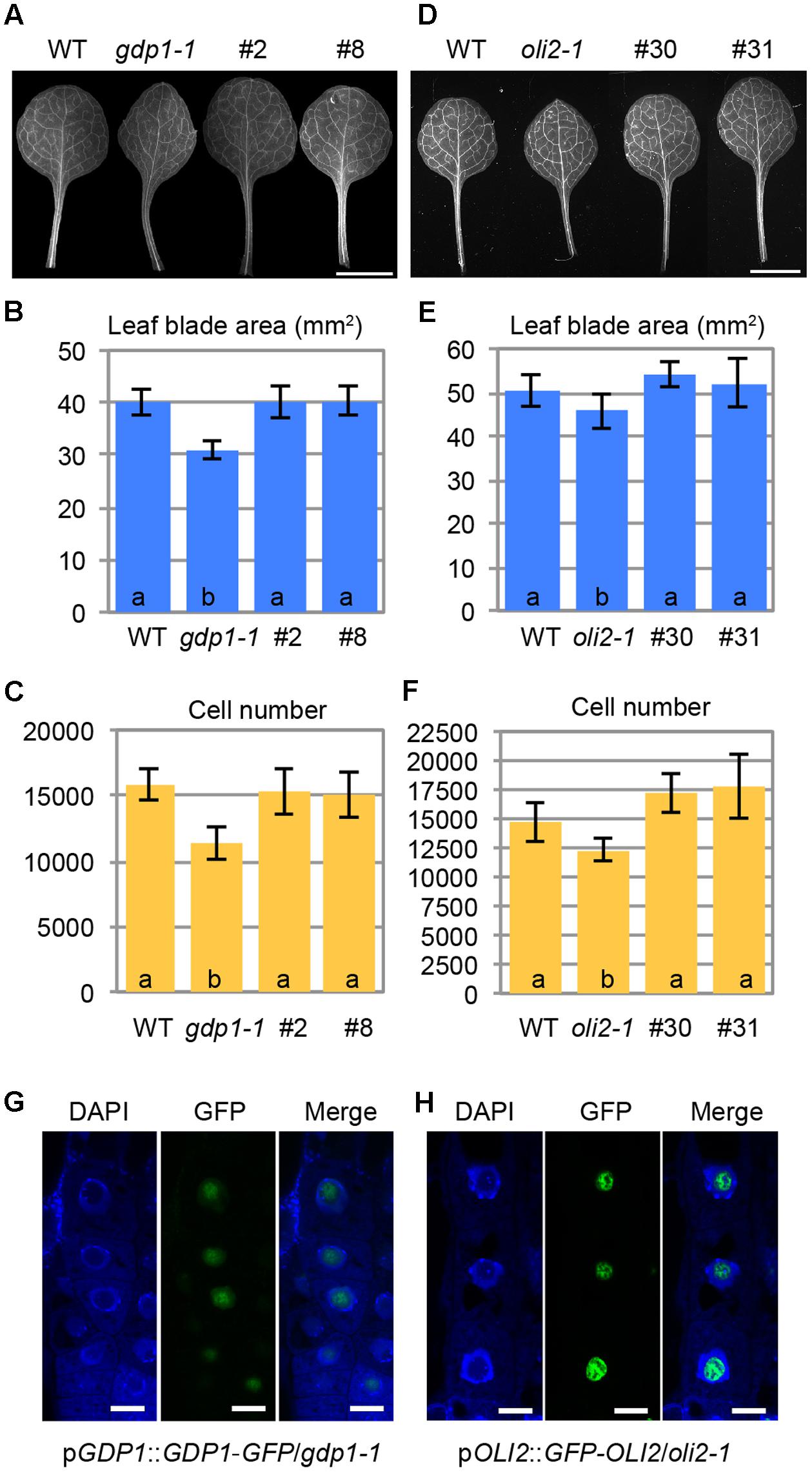
FIGURE 6. Intracellular localization of GDP1-GFP and GFP-OLI2. (A,D) First leaves of transgenic gdp1-1 lines harboring a pGDP1::GDP1-GFP construct (#2 and #8) (A) and a pOLI2::GFP-OLI2 construct (#30 and #31) (D) grown for 25 days. (B,C,E,F) Quantitative analyses of leaf phenotypes of the gdp1 (B,C) and oli2 (E,F) transgenic lines. Leaf blade area (B,E), and the number of leaf mesophyll cells (C,F) in the adaxial subepidermal layer of palisade tissues of first leaves are shown. In (B,C,E,F), data are means ± SD (n = 10). Statistically significant differences were indicated by different letters (one-way ANOVA with Tukey HSD test, p < 0.05). (G,H) Subcellular localization of GDP1-GFP (G) and GFP-OLI2 (H) in cells stained with DAPI. Fluorescence images of DAPI, GFP, and merged images are shown from left to right. Bars, 10 μm.
No information is available regarding the subcellular localization of GDP1 even in nucleolar proteomics analyses (Brown et al., 2005; Pendle et al., 2005; Palm et al., 2016), while OLI2 was detected as a nuclear/nucleolar protein (Palm et al., 2016). We examined the subcellular localization of GDP1-GFP and GFP-OLI2 in root tips stained with DAPI. DAPI stains nuclear chromosomal DNA, and the nucleolus is recognized as a round and dark region in the center of the nucleus. Both GDP1-GFP and GFP-OLI2 signals were found in DAPI-negative nuclear regions (Figures 6G,H). As the nucleolus is the center of ribosome biogenesis, the nucleolar localization of GDP1 and OLI2 further supported their roles in ribosome biogenesis.
Processing of rRNAs in gdp1, oli2, and gdp1 oli2
We next examined the effects of gdp1 and oli2 mutations on rRNA processing. A brief overview of rRNA processing intermediates relevant to this experiment is shown in Figure 7A. We first examined whether the levels of rRNA intermediates were altered in these mutants. RT-qPCR analysis suggested that rRNA intermediates containing 5′-ETS, ITS1, or ITS2 accumulated at higher levels in gdp1-1, oli2-1, and gdp1-1 oli2-1 than in the wild type (Figure 7B). The oli2-1 mutation seemed to have a stronger negative effect on rRNA processing of ITS2-containing intermediates than those containing 5′-ETS or ITS1. On the other hand, gdp1-1 had a broader impact on the accumulation of rRNA intermediates compared to oli2-1 as the levels of 5′-ETS-, ITS1-, or ITS2-containing rRNA intermediates accumulated by more than twofold compared to the wild type. Unexpectedly, the 5′-ETS-, ITS1-, and ITS2-containing intermediates accumulated at similar levels in gdp1-1 oli2-1 compared to those found in gdp1-1, despite their synergistic negative effect on cell proliferation (Figure 7B). Next we examined the patterns of rRNA intermediate accumulation by Northern hybridization (Figure 7C). In gdp1-1, oli2-1, and gdp1-1 oli2-1, 35S rRNA accumulated at higher levels than in the wild type. These mutants also accumulated 27SA, 27SB, P-A3, and 18SA3 rRNAs at higher levels than those seen in the wild type. Similar to the results of RT-qPCR analyses (Figure 7B), gdp1-1 oli2-1 showed only modest increases, if any, in levels of these rRNA intermediates compared to their parental mutants (Figure 7C). These results indicate that OLI2 and GDP1 are required for normal progression of rRNA processing.
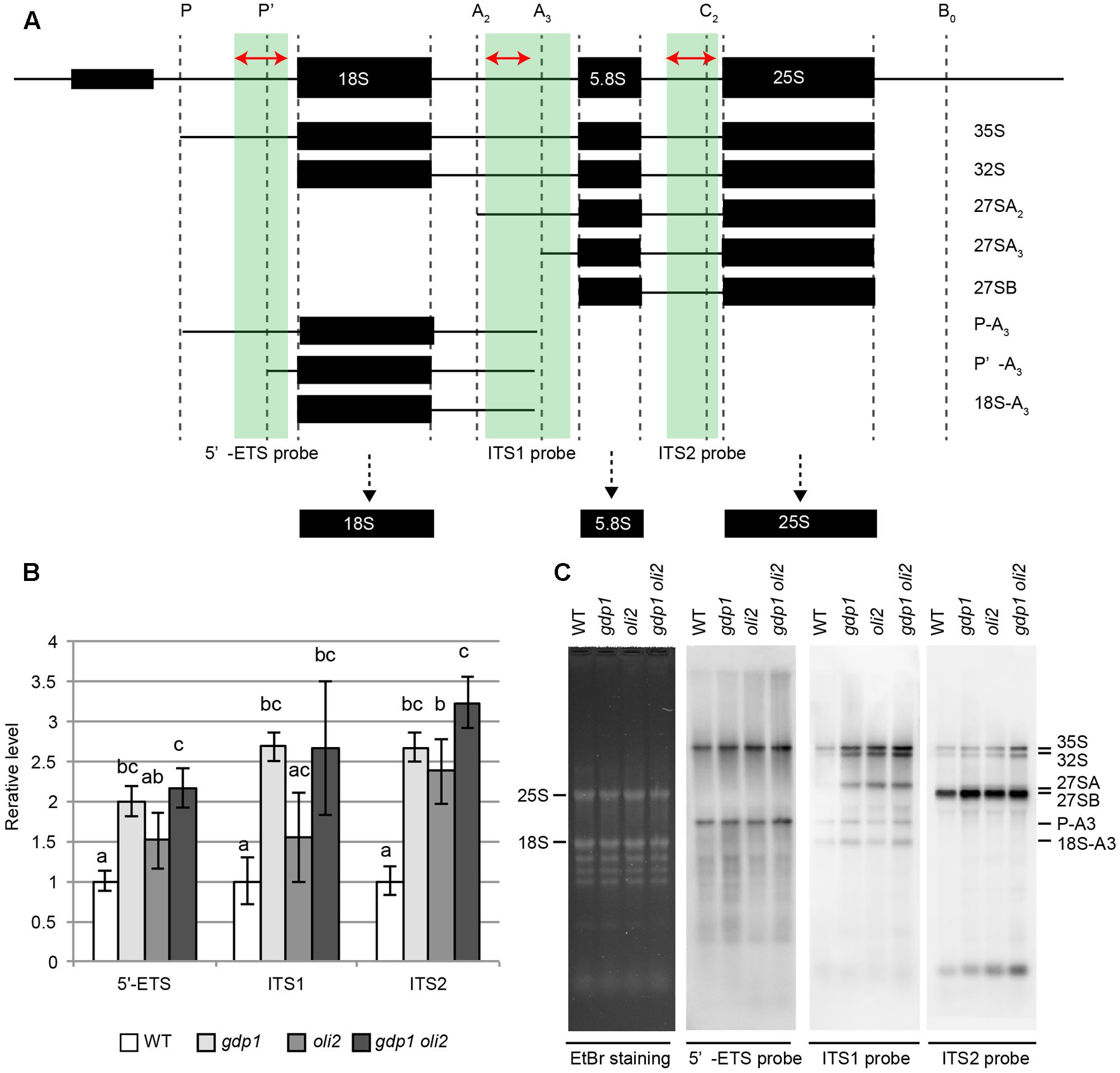
FIGURE 7. Accumulation of rRNA intermediates. (A) Brief overview of rRNA intermediates detected by either RT-qPCR or Northern hybridization. Parts of rRNA intermediates amplified by RT-qPCR and those detected by Northern hybridization are indicated by red arrows and green boxes, respectively. (B) RT-qPCR analysis of rRNA intermediates. n = 3, mean ± SD. Statistically significant differences were indicated by different letters (one-way ANOVA with Tukey HSD test, p < 0.05). (C) Northern hybridization of total RNAs. Shoots of 12-day-old WT, gdp1-1, oli2-1, and gdp1-1 oli2-1 were used.
Effects of gdp1 and oli2 on the Leaf Polarity Defects of as2
As many mutants defective in an r-protein gene show the enhanced leaf abaxialization phenotype of as2, we examined whether gdp1 and oli2 also have similar developmental effects. In this experiment, we included as2-1 rpl4d-3, which was reported previously to show very strong leaf abaxialization (Horiguchi et al., 2011). The as2-1 oli2-1 plants showed only moderately enhanced leaf abaxialization, as determined from the frequencies of formation of needle and trumpet-like leaves (Figures 8A,B). On the other hand, gdp1-1 mutation had an even weaker effect on the leaf polarity defect of as2 than oli2-1; as2-1 gdp1-1 only occasionally produced trumpet- and needle-like leaves (Figures 8A,B). We also generated as2-1 gdp1-1 oli2-1 triple mutants. In contrast to the synergistic negative effect of gdp1-1 and oli2-1 on cell proliferation (Figure 4), the triple mutant showed only slight enhancement of the leaf polarity defect compared to as2-1 oli2-1 (Figures 8A,B). Next, we examined the expression levels of leaf polarity genes. Abaxially expressed genes, such as ARF3, KAN2, and YAB5, were slightly upregulated in gdp1-1, oli2-1, and as2-1, and the degree of upregulation increased progressively in double mutant combinations among as2-1, gdp1-1, and oli2-1 and as2-1 gdp1-1 oli2-1 triple mutants (Figure 8C). On the other hand, the expression levels of adaxially expressed genes, such as PHB, PHV, and REV, were relatively constant between wild type and both single and multiple mutants examined (Figure 8C). These results suggest that gdp1 and oli2 upregulate the expression levels of ARF3, KAN2, and YAB5, and slightly enhance leaf abaxialization of as2. However, the effects of gdp1-1 and oli2-1 on leaf abaxialization were much weaker than those of rpl4d-3 (Figures 8A,B). One possible explanation for this observation is that a ribosome biosynthesis defect has the potential to induce leaf abaxialization in as2-1, but at the same time, GDP1 and OLI2 also play roles in the promotion of leaf abaxialization. To examine this possibility, we generated as2-1 rpl4d-3 gdp1-1 and as2 rpl4d-3 oli2-1. These triple mutants did not show alleviation of the leaf polarity defect of as2-1 rpl4d-3, but had more severe developmental defects judging from their smaller shoot size and formation of filamentous first and/or second leaves (Figure 8D). This result suggests that GDP1 and OLI2 are dispensable for leaf polarity establishment and/or maintenance even in the as2 background.
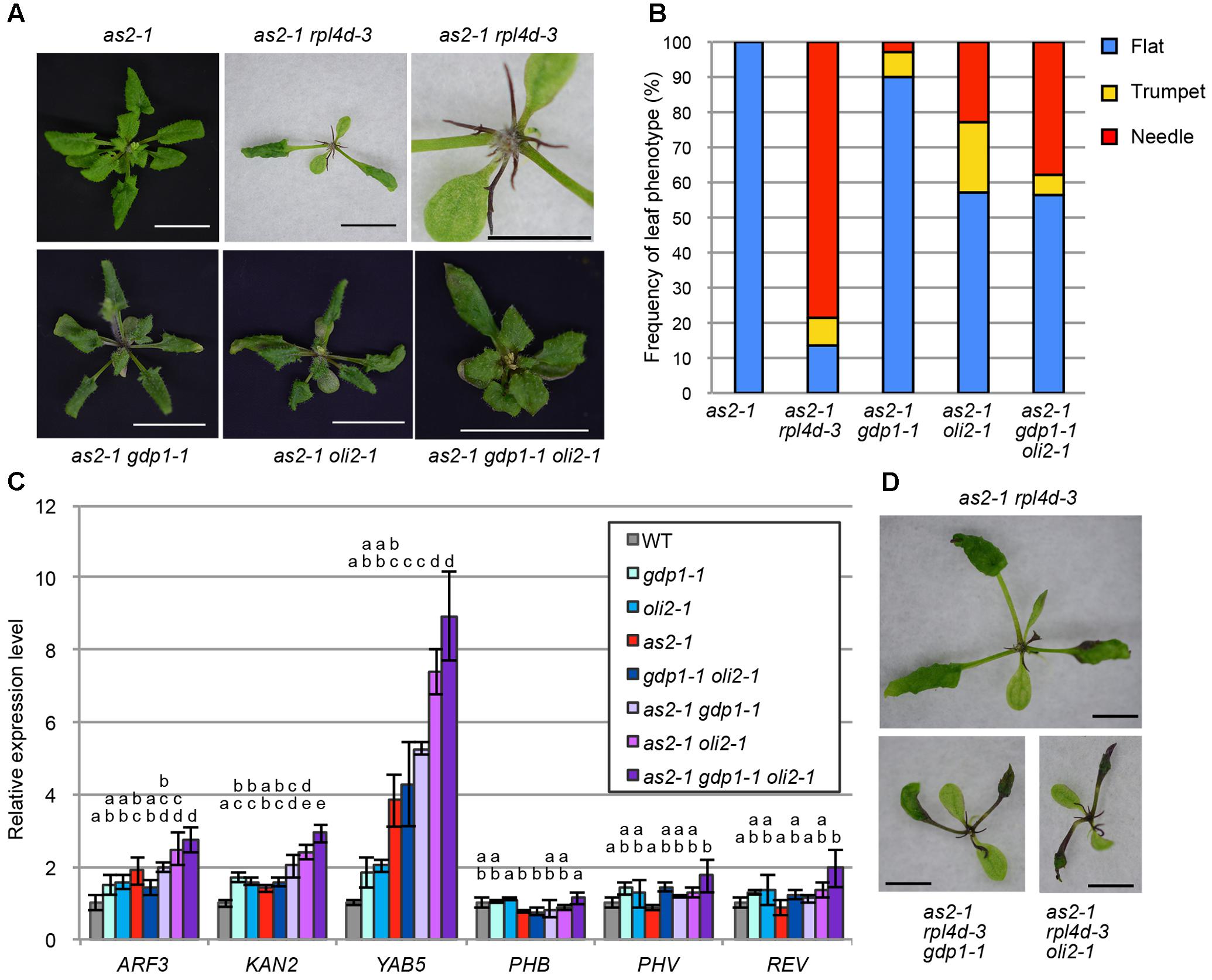
FIGURE 8. The effects of gdp1 and oli2 on leaf development in the as2 background. (A) Shoots of as2-1, as2-1 rpl4d-3, as2-1 gdp1-1, as2-1 oli2-1, and as2-1 gdp1-1 oli2-1 grown for 25 days. Bars indicate 1 cm except for the close-up view of as2-1 rpl4d-3 where it shows 5 mm. (B) Frequencies of flat, trumpet-like, and needle-like leaves. More than 13 plants were examined and all rosette and cauline leaves were scored. (C) Expression levels of leaf polarity genes. Shoots of 10-day-old seedlings were used for RNA extraction. n = 3, mean ± SD. (D) Shoots of as2-1 rpl4d-3, as2 rpl4d-3 gdp1-1, and as2 rpl4d-3 oli2-1 grown for 25 days. Bars, 1 mm. Statistically significant differences were indicated by different letters (one-way ANOVA with Tukey HSD test, p < 0.05).
Discussion
Involvement of GDP1 and OLI2 in Ribosome Biogenesis
In this study, we found that GDP1 and OLI2 are strongly expressed in developing tissues and encode different nucleolar proteins. Their loss-of-function mutations cause developmental phenotypes that are often observed in RBF-defective mutants and affect rRNA processing. These results suggest that the two nucleolar proteins are involved in ribosome biogenesis.
OLI2 is a homolog of yeast Nop2, which has methyltransferase activity and modifies 25S rRNA at a specific cytosine residue (m5C2870; Sharma et al., 2013). This modification is evolutionarily conserved at an equivalent site in animals (Maden, 1988) and plants (Burgess et al., 2015). However, m5C2870 is dispensable in yeast as catalytically inactive Nop2 is able to complement a nop2 deletion mutant (Bourgeois et al., 2015). Nop2 is a component of the LSU processome (McCann et al., 2015) and acts as one of the B factors necessary to carry out the processing of 27S rRNA into 25S and 5.8S rRNAs (Hong et al., 1997; Talkish et al., 2012). Depletion of Nop2 results in increased accumulation of 27S rRNA and corresponding decreases in levels of mature 25S and 5.8S rRNAs (Hong et al., 1997).
The conserved m5C methylation site in 25S rRNA in plants indicates the presence of a functional ortholog of Nop2 in Arabidopsis. However, this modification normally presents in oli2 mutant alleles (named nop2a in Burgess et al., 2015). This may be due to functional redundancy among the OLI2 gene family; double mutants between nop2a and nop2b are lethal (Burgess et al., 2015). Therefore, direct evidence for OLI2 as an m5C methyltransferase is lacking at present. In addition, while both nop2 in yeast and oli2 show overaccumulation of precursors that retain the intact ITS2, and nop2 reduces the 25S rRNA level, oli2 does not detectably reduce the 25S rRNA level (Figure 7C). This suggests that the overall processing mechanisms involving OLI2 and Nop2 are conserved in eukaryotes, but their details may differ. Taken together, these results indicate that OLI2 plays a role as an RBF; however, whether it is a genuine Nop2 ortholog remains to be determined.
In this study we also identified a novel nucleolar RBF named GDP1, which contains a G-patch domain. In yeast, two G-patch domain proteins, Gno1 and Pfa1, are involved in ribosome biogenesis (Guglielmi and Werner, 2002; Lebaron et al., 2005; Chen et al., 2014). According to PANTHER version 12.02 (Mi et al., 2016), GDP1 was classified as a protein family (PTHR23149) that also contains Gno1 and its ortholog in human, PINX1. These three proteins harbor a G-patch domain in their amino-terminal side. On the other hand, Pfa1 has a G-patch domain in its carboxy-terminal side and also has an R3H domain. Gno1 associates with both 90S pre-ribosomes and early pre-60S ribosomes, but only transiently with early pre-40S ribosomes (Chen et al., 2014). The lack of Gno1 leads to processing defects within the 5′-ETS and ITS1 (Chen et al., 2014). Downstream rRNA intermediates, such as 27S rRNAs and mature 18S and 25S rRNAs are reduced in gno1. In gdp1, 35S and 32S rRNAs increased, but in contrast to gno1, 27SA/B rRNAs increased without detectably affecting the steady-state levels of 25S and 18S rRNAs (Figure 7C). Thus, gdp1, like gno1, seems to affect multiple steps of rRNA processing, but it has a partially different effect on rRNA processing. Thus, whether GDP1 has an orthologous function to Gno1 remains to be elucidated. An emerging molecular role of G-patch domain protein is the activation of DEAH/RHA helicases (Robert-Paganin et al., 2015). Both Gno1 and Pfa1 function in ribosome biogenesis in close association with the DEAH/RHA helicase, Prp43. Future characterization of an Arabidopsis homolog of Prp43 may provide additional insight into the function of GDP1.
Relationships between R-Proteins and RBFs
In a series of genetic crosses, we found an additive or synergistic genetic interaction depending on the combination of single mutants examined. The combination between gdp1 and most r-protein mutants examined resulted in an additive phenotype. This suggests that processes of GDP1-dependent ribosome biogenesis and those involving these r-proteins occur largely independently. On the other hand, the synergistic genetic interaction between gdp1 and oli2 suggests the interdependence of GDP1 and OLI2 functions in ribosome biogenesis. In yeast, Gno1 acts at multiple steps of ribosome biogenesis in close association with Prp43 (Lebaron et al., 2005, 2009; Chen et al., 2014). Prp43 is expected to function in rearrangement of pre-ribosomal particles through its RNA helicase activity (Lebaron et al., 2005, 2009). Nop2 is an essential protein and is required for hierarchical recruitment of the B factors (Hong et al., 1997; Talkish et al., 2012). Thus, successive changes in the geometry of pre-ribosomes are important for the progression of subsequent processes. The interdependence of GDP1 and OLI2 functions may have arisen from the dynamic nature of ribosome biogenesis.
Unexpectedly, gdp1 oli2 did not show marked increases in the levels of rRNA intermediates compared to its parental single mutants (Figure 7). This discrepancy may have been due to the nature of the experiments, in that our RNA analyses determined the steady-state levels of rRNA intermediates. Further characterization of GDP1 and OLI2 and measurement of rRNA processing rate could resolve this issue.
Another notable interaction was found between gdp1 and rps6a. Despite being a component of the 40S subunit, RPS6A was shown to interact with the histone deacetylase HD2B and NUCLEOSOME ASSEMBLY PROTEIN1 (NAP1), and to regulate transcription of 45S rRNA (Kim et al., 2014; Son et al., 2015). Thus, the exceptionally strong interaction of rps6a with gdp1 among other r-protein mutants may have been due to the role of RPS6A in ribosome biogenesis. Similar synergistic relationships were also reported between another pair of RBF mutants, apum23 and nuc-l1 (Abbasi et al., 2010). In the same work, double mutants between apum23 and as1/2 enhancer5 (ae5)/rpl28a or ae6/rpl5a were reported not to show more severe phenotypes than the respective single mutants (Abbasi et al., 2010). As ae6 is an allele of oli5, and gdp1 oli5 shows an additive phenotype in relation to the cell proliferation defect (Figure 3), evaluation of genetic interactions between ribosome-related mutants based solely on shoot size could miss a more fundamental interaction. In summary, our genetic experiments suggested that a combination of weak defects in RBFs may cause a marked reduction of the flow of ribosome biogenesis, resulting in a strong cell proliferation defect.
Genetic Defects in RBFs Do Not Always Induce Strong Leaf Abaxialization in as2
Several reports described enhanced leaf abaxialization in as2 by r-protein mutants (Byrne, 2009; Horiguchi et al., 2012; Machida et al., 2015). Recently, mutations in three RBF genes, NUC-L1, RH10, and RID2, were also shown to strongly enhance leaf abaxialization in as2 (Matsumura et al., 2016). In yeast, orthologs of these three proteins are components of, or RBFs associated with, the SSU processome. These findings suggested a tight link between the SSU processome and AS2-dependent cell fate decision (Matsumura et al., 2016). In addition, APUM23 encodes a homolog of yeast Nop9, and apum23 also strongly enhances the as2 leaf polarity phenotype (Huang et al., 2014). Although the precise functions of APUM23 and Nop9 could differ, APUM23 is able to partially complement the nop9 mutant phenotype in yeast (Huang et al., 2014). Interestingly, Nop9 is also a component of the SSU processome (Zhang et al., 2016). In contrast to reports regarding these RBFs (Huang et al., 2014; Matsumura et al., 2016), gdp1, oli2, and even gdp1 oli2 had little or very mild effects on the enhancement of leaf abaxialization in the as2 background (Figure 8). These results suggest that not all of the RBFs are tightly linked with leaf adaxial/abaxial polarity. Gno1 was found in the SSU processome, pre-40S, and pre-60S ribosomes (Chen et al., 2014). If GDP1 is also a component of the SSU processome, this argues against a link between the SSU processome and leaf abaxial/adaxial polarity regulation. A possible explanation for this discrepancy is that there may be a key structure or a subcomplex within the SSU processome, dysfunction of which is linked to leaf abaxialization in as2. This is a likely scenario, as the SSU processome is a 2.2-MDa ribonucleoprotein composed of modular subcomplexes (Pérez-Fernández et al., 2007). On the other hand, Nop2 is a component of the LSU processome in yeast, and OLI2 is a putative ortholog of Nop2. Several scenarios may explain the limited effect of oli2 on leaf abaxialization. First, the LSU processome may not have a link to the regulation of adaxial/abaxial polarity. However, we consider this to be unlikely, because we reported previously that mutations in RPL4D encoding a 60S r-protein have a very strong effect on leaf abaxialization in as2 (Horiguchi et al., 2011), and, in yeast, RPL4 is incorporated during an early stage of pre-60S formation (Gamalinda et al., 2014; Stelter et al., 2015; Chen et al., 2017). Similar to the SSU processome, the LSU processome may also have a link to adaxial/abaxial patterning through its local structures. As ribosome biogenesis is a highly dynamic process, surveillance of ribosome biogenesis defects at every step is probably impossible. Consistently, in yeast cells with specific genetic backgrounds, aberrant ribosomal subunits can escape from the surveillance systems and engage in translation (Rodríguez-Galán et al., 2015). On the other hand, surveillance at multiple key checkpoints to sense local defects of nascent ribosomes represents an easier strategy. The differential contributions of RBFs to leaf abaxial/adaxial polarity regulation may be correlated with their relative importance to such checkpoints.
Our genetic analyses on gdp1 and oli2 illustrated that a genetic defect in ribosome biogenesis leads to cell proliferation defect but it does not have a strong effect to enhance leaf abaxialization when it is introduced into the as2 background. A contrasting example is found in rpl4d where rpl4d single mutants do not have a clear cell proliferation defect yet it causes very strong leaf abaxialization in the as2 background (Horiguchi et al., 2011). These examples suggest there may be a critical point during ribosome biogenesis that is linked to the regulation of leaf adaxial/abaxial polarity. It is noteworthy that oli2 has a statistically significant effect to upregulate YAB5 expression when compared to as2 (Figure 8C) but it did not result in strong leaf abaxialization. In as2 rh10, strong leaf abaxialization is dependent on ARF3 (Matsumura et al., 2016). The expression levels of ARF3 in as2 rh10 and as2 gdp1 oli2 were four and threefold higher than that in the wild type plants, respectively (Matsumura et al., 2016; Figure 8C). Although plant samples used in these studies were different (shoot tips or whole shoots), the differential sensitivities of leaf abaxialization in these mutants might be attributable to the degree of enhanced ARF3 expression. An increasing number of characterized ribosome biogenesis mutants of Arabidopsis should offer an opportunity to localize such a point through quantitative and comparative analyses of these mutants in a future study.
A critical issue concerning the above discussion is whether there is actually such a checkpoint in plants. In mammals, ribosome biogenesis defects lead to activation of the tumor suppressor, p53, which acts as a transcriptional activator and triggers stress responses, such as cell cycle arrest, senescence, and apoptosis (Kastenhuber and Lowe, 2017). Under normal conditions, p53 is ubiquitinated by the E3 ubiquitin ligase, MDM2, and subjected to proteolysis. On the other hand, ribosome biogenesis defects inhibit MDM2 activity through direct interaction between the 5S rRNA-RPL5-RPL11 complex and MDM2 (Sun et al., 2010; Sloan et al., 2013; Bursac et al., 2014). However, plants have orthologs of neither p53 nor MDM2. Recently, Ohbayashi et al. (2017) showed that the NAC transcription factor gene, SRIW1, is upregulated in rid2. Notably, rid2 sriw1 still suffers from the rRNA processing defects, but the developmental phenotypes, such as impaired callus formation and pointed leaf formation, were suppressed (Ohbayashi et al., 2017). These results indicated that the observed developmental phenotypes are indirect consequences of rRNA processing defects, and are more directly mediated by the ribosomal stress response in which SRIW1 plays a central role, similar to p53 in animals.
In this study, we showed that gdp1 oli2 double mutants had a very strong cell proliferation defect, but enhanced the leaf polarity defect of as2 less strongly. How a defect in a general cellular process such as ribosome biogenesis interferes with a specific developmental process is an important emerging issue in plant developmental and cell biology (Tsukaya et al., 2013). We suggest that the cell proliferation defect in leaf primordia and leaf abaxialization triggered by a mutation in RBF or an r-protein gene could be mediated by different mechanisms. It will be interesting to examine whether these developmental phenotypes are mediated by SRIW1.
Author Contributions
GH and HT designed and conducted the experiments. KK, JT, HC, KF, and GH performed the experiments. GH and HT wrote the manuscript.
Funding
This work was supported by Grants-in-Aid for Scientific Research on Innovative Areas (to HT and GH) and Scientific Research (C) (to GH) (JSPS KAKENHI Grant Numbers JP25113002 and JP15K07115).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The handling Editor declared a past co-authorship with several of the authors HT, GH.
Acknowledgments
We thank Hiroaki Ichikawa and Tsuyoshi Nakagawa for binary vectors pSMAH621 and pGWB501, respectively. We also thank Moeno Kagami for her experimental assistance.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2017.02240/full#supplementary-material
Footnotes
References
Abbasi, N., Kim, H. B., Park, N. I., Kim, H. S., Kim, Y. K., Park, Y. I., et al. (2010). APUM23, a nucleolar Puf domain protein, is involved in pre-ribosomal RNA processing and normal growth patterning in Arabidopsis. Plant J. 64, 960–976. doi: 10.1111/j.1365-313X.2010.04393.x
Aravind, L., and Koonin, E. V. (1999). G-patch: a new conserved domain in eukaryotic RNA-processing proteins and type D retroviral polyproteins. Trends Biochem. Sci. 24, 342–344. doi: 10.1016/S0968-0004(99)01437-1
Bourgeois, G., Ney, M., Gaspar, I., Aigueperse, C., Schaefer, M., Kellner, S., et al. (2015). Eukaryotic rRNA modification by yeast 5-methylcytosine-methyltransferases and human proliferation-associated antigen p120. PLOS ONE 10:e133321. doi: 10.1371/journal.pone.0133321
Brown, J. W. S., Shaw, P. J., Shaw, P., and Marshall, D. F. (2005). Arabidopsis nucleolar protein database (AtNoPDB). Nucleic Acids Res. 33, D633–D636. doi: 10.1093/nar/gki052
Burgess, A. L., David, R., and Searle, I. R. (2015). Conservation of tRNA and rRNA 5-methylcytosine in the kingdom Plantae. BMC Plant Biol. 15:119. doi: 10.1186/s12870-015-0580-8
Bursac, S., Brdovcak, M. C., Donati, G., and Volarevic, S. (2014). Activation of the tumor suppressor p53 upon impairment of ribosome biogenesis. Biochim. Biophys. Acta 1842, 817–830. doi: 10.1016/j.bbadis.2013.08.014
Byrne, M. E. (2009). A role for the ribosome in development. Trends Plant Sci. 14, 512–519. doi: 10.1016/j.tplants.2009.06.009
Casanova-Sáez, R., Candela, H., and Micol, J. L. (2014). Combined haploinsufficiency and purifying selection drive retention of RPL36a paralogs in Arabidopsis. Sci. Rep. 4:4122. doi: 10.1038/srep04122
Chaker-Margot, M., Barandun, J., Hunziker, M., and Klinge, S. (2017). Architecture of the yeast small subunit processome. Science 355:eaal1880. doi: 10.1126/science.aal1880
Chen, W., Xie, Z., Yang, F., and Ye, K. (2017). Stepwise assembly of the earliest precursors of large ribosomal subunits in yeast. Nucleic Acids Res. 45, 6837–6847. doi: 10.1093/nar/gkx254
Chen, Y. L., Capeyrou, R., Humbert, O., Mouffok, S., Kadri, Y. A., Lebaron, S., et al. (2014). The telomerase inhibitor Gno1p/PINX1 activates the helicase Prp43p during ribosome biogenesis. Nucleic Acids Res. 42, 7330–7345. doi: 10.1093/nar/gku357
Cho, H. K., Ahn, C. S., Lee, H. S., Kim, J. K., and Pai, H. S. (2013). Pescadillo plays an essential role in plant cell growth and survival by modulating ribosome biogenesis. Plant J. 76, 393–405. doi: 10.1111/tpj.12302
Clough, S. J., and Bent, A. F. (1998). Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743. doi: 10.1046/j.1365-313x.1998.00343.x
Comella, P., Pontvianne, F., Lahmy, S., Vignols, F., Barbezier, N., DeBures, A., et al. (2007). Characterization of a ribonuclease III-like protein required for cleavage of the pre-rRNA in the 3’ETS in Arabidopsis. Nucleic Acids Res. 36, 1163–1175. doi: 10.1093/nar/gkm1130
Donnelly, P. M., Bonetta, D., Tsukaya, H., Dengler, R. E., and Dengler, N. G. (1999). Cell cycling and cell enlargement in developing leaves of Arabidopsis. Dev. Biol. 215, 407–419. doi: 10.1006/dbio.1999.9443
Droll, D., Archer, S., Fenn, K., Delhi, P., Matthews, K., and Clayton, C. (2010). The trypanosome Pumilio-domain protein PUF7 associates with a nuclear cyclophilin and is involved in ribosomal RNA maturation. FEBS Lett. 584, 1156–1162. doi: 10.1016/j.febslet.2010.02.018
Ebersberger, I., Simm, S., Leisegang, M. S., Schmitzberger, P., Mirus, O., von Haeseler, A., et al. (2013). The evolution of the ribosome biogenesis pathway from a yeast perspective. Nucleic Acids Res. 42, 1509–1523. doi: 10.1093/nar/gkt1137
Fernández-Pevida, A., Kressler, D., and de la Cruz, J. (2015). Processing of preribosomal RNA in Saccharomyces cerevisiae. Wiley Interdiscip. Rev. RNA 6, 191–209. doi: 10.1002/wrna.1267
Fujikura, U., Horiguchi, G., Ponce, M. R., Micol, J. L., and Tsukaya, H. (2009). Coordination of cell proliferation and cell expansion mediated by ribosome-related processes in the leaves of Arabidopsis thaliana. Plant J. 59, 499–508. doi: 10.1111/j.1365-313X.2009.03886.x
Gamalinda, M., Ohmayer, U., Jakovljevic, J., Kumcuoglu, B., Woolford, J., Mbom, B., et al. (2014). A hierarchical model for assembly of eukaryotic 60S ribosomal subunit domains. Genes Dev. 28, 198–210. doi: 10.1101/gad.228825.113
Grandi, P., Rybin, V., Bassler, J., Petfalski, E., Strauss, D., Marzioch, M., et al. (2002). 90S pre-ribosomes include the 35S pre-rRNA, the U3 snoRNP, and 40S subunit processing factors but predominantly lack 60S synthesis factors. Mol. Cell 10, 105–115. doi: 10.1016/S1097-2765(02)00579-8
Griffith, M. E., Mayer, U., Capron, A., Ngo, Q. A., Surendrarao, A., McClinton, R., et al. (2007). The TORMOZ gene encodes a nucleolar protein required for regulated division planes and embryo development in Arabidopsis. Plant Cell 19, 2246–2263. doi: 10.1105/tpc.106.042697
Guglielmi, B., and Werner, M. (2002). The yeast homolog of human PinX1 is involved in rRNA and small nucleolar RNA maturation, not in telomere elongation inhibition. J. Biol. Chem. 277, 35712–35719. doi: 10.1074/jbc.M205526200
Hang, R., Liu, C., Ahmad, A., Zhang, Y., Lu, F., and Cao, X. (2014). Arabidopsis protein arginine methyltransferase 3 is required for ribosome biogenesis by affecting precursor ribosomal RNA processing. Proc. Natl. Acad. Sci. U.S.A. 111, 16190–16195. doi: 10.1073/pnas.1412697111
Hasegawa, J., Sakamoto, Y., Nakagami, S., Aida, M., Sawa, S., and Matsunaga, S. (2016). Three-dimensional imaging of plant organs using a simple and rapid transparency technique. Plant Cell Physiol. 57, 462–472. doi: 10.1093/pcp/pcw027
Henras, A. K., Plisson-Chastang, C., O’Donohue, M.-F., Chakraborty, A., and Gleizes, P. E. (2015). An overview of pre-ribosomal RNA processing in eukaryotes. Wiley Interdiscip. Rev. RNA 6, 225–242. doi: 10.1002/wrna.1269
Hisanaga, T., Kawade, K., and Tsukaya, H. (2015). Compensation: a key to clarifying the organ-level regulation of lateral organ size in plants. J. Exp. Bot. 66, 1055–1063. doi: 10.1093/jxb/erv028
Hong, B., Brockenbrough, J. S., Wu, P., and Aris, J. P. (1997). Nop2p is required for pre-rRNA processing and 60S ribosome subunit synthesis in yeast. Mol. Cell. Biol. 17, 378–388. doi: 10.1128/MCB.17.1.378
Horiguchi, G., Ferjani, A., Fujikura, U., and Tsukaya, H. (2006a). Coordination of cell proliferation and cell expansion in the control of leaf size in Arabidopsis thaliana. J. Plant Res. 119, 37–42. doi: 10.1007/s10265-005-0232-4
Horiguchi, G., Fujikura, U., Ferjani, A., Ishikawa, N., and Tsukaya, H. (2006b). Large-scale histological analysis of leaf mutants using two simple leaf observation methods: identification of novel genetic pathways governing the size and shape of leaves. Plant J. 48, 638–644. doi: 10.1111/j.1365-313X.2006.02896.x
Horiguchi, G., Kim, G. T., and Tsukaya, H. (2005). The transcription factor AtGRF5 and the transcription coactivator AN3 regulate cell proliferation in leaf primordia of Arabidopsis thaliana. Plant J. 43, 68–78. doi: 10.1111/j.1365-313X.2005.02429.x
Horiguchi, G., Mollá-Morales, A., Pérez-Pérez, J. M., Kojima, K., Robles, P., Ponce, M. R., et al. (2011). Differential contributions of ribosomal protein genes to Arabidopsis thaliana leaf development. Plant J. 65, 724–736. doi: 10.1111/j.1365-313X.2010.04457.x
Horiguchi, G., and Tsukaya, H. (2011). Organ size regulation in plants: insights from compensation. Front. Plant Sci. 2:24. doi: 10.3389/fpls.2011.00024
Horiguchi, G., Van Lijsebettens, M., Candela, H., Micol, J. L., and Tsukaya, H. (2012). Ribosomes and translation in plant developmental control. Plant Sci. 19, 24–34. doi: 10.1016/j.plantsci.2012.04.008
Huang, C. K., Huang, L. F., Huang, J. J., Wu, S. J., Yeh, C. H., and Lu, C. A. (2010). A DEAD-box protein, AtRH36, is essential for female gametophyte development and is involved in rRNA biogenesis in Arabidopsis. Plant Cell Physiol. 51, 694–706. doi: 10.1093/pcp/pcq045
Huang, T., Kerstetter, R. A., and Irish, V. F. (2014). APUM23, a PUF family protein, functions in leaf development and organ polarity in Arabidopsis. J. Exp. Bot. 65, 1181–1191. doi: 10.1093/jxb/ert478
Iwakawa, H., Iwasaki, M., Kojima, S., Ueno, Y., Soma, T., Tanaka, H., et al. (2007). Expression of the ASYMMETRIC LEAVES2 gene in the adaxial domain of Arabidopsis leaves represses cell proliferation in this domain and is critical for the development of properly expanded leaves. Plant J. 51, 173–184. doi: 10.1111/j.1365-313X.2007.03132.x
Kastenhuber, E. R., and Lowe, S. W. (2017). Putting p53 in context. Cell 170, 1062–1078. doi: 10.1016/j.cell.2017.08.028
Kim, Y.-K., Kim, S., Shin, Y., Hur, Y.-S., Kim, W.-Y., Lee, M.-S., et al. (2014). Ribosomal Protein S6, a target of rapamycin, is involved in the regulation of rRNA genes by possible epigenetic changes in Arabidopsis. J. Biol. Chem. 289, 3901–3912. doi: 10.1074/jbc.M113.515015
Konikkat, S., and Woolford, J. L. Jr. (2017). Principles of 60S ribosomal subunit assembly emerging from recent studies in yeast. Biochem. J. 474, 195–214. doi: 10.1042/BCJ20160516
Kornprobst, M., Turk, M., Kellner, N., Cheng, J., Flemming, D., Kos-Braun, I., et al. (2016). Architecture of the 90S pre-ribosome: a structural view on the birth of the eukaryotic ribosome. Cell 166, 380–393. doi: 10.1016/j.cell.2016.06.014
Kumakura, N., Otsuki, H., Tsuzuki, M., Takeda, A., and Watanabe, Y. (2013). Arabidopsis AtRRP44A is the functional homolog of Rrp44/Dis3, an exosome component, is essential for viability and is required for RNA processing and degradation. PLOS ONE 8:e79219. doi: 10.1371/journal.pone.0079219
Lange, H., Holec, S., Cognat, V., Pieuchot, L., Le Ret, M., Canaday, J., et al. (2008). Degradation of a polyadenylated rRNA maturation by-product involves one of the three RRP6-like proteins in Arabidopsis thaliana. Mol. Cell. Biol. 28, 3038–3044. doi: 10.1128/MCB.02064-07
Lange, H., Sement, F. M., and Gagliardi, D. (2011). MTR4, a putative RNA helicase and exosome co-factor, is required for proper rRNA biogenesis and development in Arabidopsis thaliana. Plant J. 68, 51–63. doi: 10.1111/j.1365-313X.2011.04675.x
Layat, E., Sáez-Vásquez, J., and Tourmente, S. (2012). Regulation of Pol I-transcribed 45S rDNA and Pol III-transcribed 5S rDNA in Arabidopsis. Plant Cell Physiol. 53, 267–276. doi: 10.1093/pcp/pcr177
Lebaron, S., Froment, C., Fromont-Racine, M., Rain, J., Monsarrat, B., Caizergues-Ferrer, M., et al. (2005). The splicing ATPase Prp43 is a component of multiple preribosomal particles. Mol. Cell. Biol. 25, 9269–9282. doi: 10.1128/MCB.25.21.9269-9282.2005
Lebaron, S., Papin, C., Capeyrou, R., Chen, Y.-L., Froment, C., Monsarrat, B., et al. (2009). The ATPase and helicase activities of Prp43p are stimulated by the G-patch protein Pfa1p during yeast ribosome biogenesis. EMBO J. 28, 3808–3819. doi: 10.1038/emboj.2009.335
Létoquart, J., Huvelle, E., Wacheul, L., Bourgeois, G., Zorbas, C., Graille, M., et al. (2014). Structural and functional studies of Bud23-Trm112 reveal 18S rRNA N7-G1575 methylation occurs on late 40S precursor ribosomes. Proc. Natl. Acad. Sci. U.S.A. 111, E5518–E5526. doi: 10.1073/pnas.1413089111
Li, N., Yuan, L., Liu, N., Shi, D., Li, X., Tang, Z., et al. (2009). SLOW WALKER2, a NOC1/MAK21 homologue, is essential for coordinated cell cycle progression during female gametophyte development in Arabidopsis. Plant Physiol. 151, 1486–1497. doi: 10.1104/pp.109.142414
Liu, M., Shi, D. Q., Yuan, L., Liu, J., and Yang, W. C. (2010). SLOW WALKER3, encoding a putative DEAD-box RNA helicase, is essential for female gametogenesis in Arabidopsis. J. Integr. Plant Biol. 52, 817–828. doi: 10.1111/j.1744-7909.2010.00972.x
Machida, C., Nakagawa, A., Kojima, S., Takahashi, H., and Machida, Y. (2015). The complex of ASYMMETRIC LEAVES (AS) proteins plays a central role in antagonistic interactions of gene s for leaf polarity specification in Arabidopsis. Wiley Interdiscip. Rev. Dev. Biol. 4, 655–671. doi: 10.1002/wdev.196
Maden, B. E. (1988). Locations of methyl groups in 28S rRNA of Xenopus laevis and man. Clustering in the conserved core of molecule. J. Mol. Biol. 201, 289–314. doi: 10.1016/0022-2836(88)90139-8
Matsumura, Y., Ohbayashi, I., Takahashi, H., Kojima, S., Ishibashi, N., Keta, S., et al. (2016). A genetic link between epigenetic repressor AS1-AS2 and a putative small subunit processome in leaf polarity establishment of Arabidopsis. Biol. Open 5, 942–954. doi: 10.1242/bio.019109
McCann, K. L., Charette, J. M., Vincent, N. G., and Baserga, S. J. (2015). A protein interaction map of the LSU processome. Genes Dev. 29, 862–875. doi: 10.1101/gad.256370.114
Mi, H., Huang, X., Muruganujan, A., Tang, H., Mills, C., Kang, D., et al. (2016). PANTHER version 11: expanded annotation data from gene ontology and reactome pathways, and data analysis tool enhancements. Nucleic Acids Res. 45, D183–D189. doi: 10.1093/nar/gkw1138
Missbach, S., Weis, B. L., Martin, R., Simm, S., Bohnsack, M. T., and Schleiff, E. (2013). 40S ribosome biogenesis co-factors are essential for gametophyte and embryo development. PLOS ONE 8:e54084. doi: 10.1371/journal.pone.0054084
Nakagawa, T., Suzuki, T., Murata, S., Nakamura, S., Hino, T., Maeo, K., et al. (2007). Improved Gateway binary vectors: high-performance vectors for creation of fusion constructs in transgenic analysis of plants. Biosci. Biotechnol. Biochem. 71, 2095–2100. doi: 10.1271/bbb.70216
Nerurkar, P., Altvater, M., Gerhardy, S., Schütz, S., Fischer, U., Weirich, C., et al. (2015). Eukaryotic ribosome assembly and nuclear export. Int. Rev. Cell Mol. Biol. 319, 107–140. doi: 10.1016/bs.ircmb.2015.07.002
O’Day, C. L., Chavanikamannil, F., and Abelson, J. (1996). 18S rRNA processing requires the RNA helicase-like protein Rrp3. Nucleic Acids Res. 24, 3201–3207. doi: 10.1093/nar/24.16.3201
Ohbayashi, I., Konishi, M., Ebine, K., and Sugiyama, M. (2011). Genetic identification of Arabidopsis RID2 as an essential factor involved in pre-rRNA processing. Plant J. 67, 49–60. doi: 10.1111/j.1365-313X.2011.04574.x
Ohbayashi, I., Lin, C. Y., Shinohara, N., Matsumura, Y., Machida, Y., Horiguchi, G., et al. (2017). Evidence for a role of ANAC082 as a ribosomal stress response mediator leading to growth defects and developmental alterations in Arabidopsis. Plant Cell 29, 2644–2660. doi: 10.1105/tpc.17.00255
Palm, D., Simm, S., Darm, K., Weis, B. L., Ruprecht, M., Schleiff, E., et al. (2016). Proteome distribution between nucleoplasm and nucleolus and its relation to ribosome biogenesis in Arabidopsis thaliana. RNA Biol. 13, 441–454. doi: 10.1080/15476286.2016.1154252
Pendle, A. F., Clark, G. P., Boon, R., Lewandowska, D., Lam, Y. W., Andersen, J., et al. (2005). Proteomic analysis of the Arabidopsis nucleolus suggests novel nucleolar functions. Mol. Biol. Cell 16, 260–269. doi: 10.1091/mbc.E04-09-0791
Pérez-Fernández, J., Román,Á, De Las Rivas, J., Bustelo, X. R., and Dosil, M. (2007). The 90S preribosome is a multimodular structure that is assembled through a hierarchical mechanism. Mol. Cell. Biol. 27, 5414–5429. doi: 10.1128/MCB.00380-07
Petricka, J. J., and Nelson, T. M. (2007). Arabidopsis nucleolin affects plant development and patterning. Plant Physiol. 144, 173–186. doi: 10.1104/pp.106.093575
Phipps, K. R., Charette, J., and Baserga, S. J. (2011). The small subunit processome in ribosome biogenesis – progress and prospects. Wiley Interdiscip. Rev. RNA 2, 1–21. doi: 10.1002/wrna.57
Pinon, V., Etchells, J. P., Rossignol, P., Collier, S. A., Arroyo, J. M., Martienssen, R. A., et al. (2008). Three PIGGYBACK genes that specifically influence leaf patterning encode ribosomal proteins. Development 135, 1315–1324. doi: 10.1242/dev.016469
Robert-Paganin, J., Réty, S., and Leulliot, N. (2015). Regulation of DEAH/RHA helicases by G-patch proteins. Biomed. Res. Int. 2015:931857. doi: 10.1155/2015/931857
Rodríguez-Galán, O., García-Gómez, J. J., Kressler, D., and de la Cruz, J. (2015). Immature large ribosomal subunits containing the 7S pre-rRNA can engage in translation in Saccharomyces cerevisiae. RNA Biol. 12, 838–846. doi: 10.1080/15476286.2015.1058477
Sáez-Vasquez, J., Caparros-Ruiz, D., Barneche, F., and Echeverría, M. (2004). A plant snoRNP complex containing snoRNAs, Fibrillarin, and Nucleolin-like proteins is competent for both rRNA gene binding and pre-rRNA processing in vitro. Mol. Cell. Biol. 24, 7284–7297. doi: 10.1128/MCB.24.16.7284-7297.2004
Sardana, R., White, J. P., and Johnson, A. W. (2013). The rRNA methyltransferase Bud23 shows functional interaction with components of the SSU processome and RNase MRP. RNA 19, 828–840. doi: 10.1261/rna.037671.112
Sharma, S., Yang, J., Watzinger, P., Kötter, P., and Entian, K. D. (2013). Yeast Nop2 and Rcm1 methylate C2870 and C2278 of the 25S rRNA, respectively. Nucleic Acids Res. 41, 9062–9076. doi: 10.1093/nar/gkt679
Shi, D. Q., Liu, J., Xiang, Y. H., Ye, D., Sundaresan, V., and Yang, W. C. (2005). SLOW WALKER1, essential for gametogenesis in Arabidopsis, encodes a WD40 protein involved in 18S ribosomal RNA biogenesis. Plant Cell 17, 2340–2354. doi: 10.1105/tpc.105.033563
Sloan, K. E., Bohnsack, M. T., and Watkins, N. J. (2013). The 5S RNP couples p53 homeostasis to ribosome biogenesis and nucleolar stress. Cell Rep. 5, 237–247. doi: 10.1016/j.celrep.2013.08.049
Son, O., Kim, S., Shin, Y. J., Kim, W. Y., Koh, H., and Cheon, C. I. (2015). Identification of nucleosome assembly protein 1 (NAP1) as an interacting partner of plant ribosomal protein S6 (RPS6) and a positive regulator of rDNA transcription. Biochem. Biophys. Res. Commun. 465, 200–205. doi: 10.1016/j.bbrc.2015.07.150
Stelter, P., Huber, F. M., Kunze, R., Flemming, D., Hoelz, A., and Hurt, E. (2015). Coordinated ribosomal L4 protein assembly into the pre-ribosome is regulated by its eukaryote-specific extension. Mol. Cell 58, 854–862. doi: 10.1016/j.molcel.2015.03.029
Sun, Q., Zhu, X., Qi, J., An, W., Lan, P., Tan, D., et al. (2017). Molecular architecture of the 90S small subunit pre-ribosome. eLife 6:e22086. doi: 10.7554/eLife.22086
Sun, X. X., Wang, Y. G., Xirodimas, D. P., and Dai, M. S. (2010). Perturbation of 60 S ribosomal biogenesis results in ribosomal protein L5- and L11-dependent p53 activation. J. Biol. Chem. 285, 25812–25821. doi: 10.1074/jbc.M109.098442
Szakonyi, D., and Byrne, M. E. (2011). Ribosomal protein L27a is required for growth and patterning in Arabidopsis thaliana. Plant J. 65, 269–281. doi: 10.1111/j.1365-313X.2010.04422.x
Tafforeau, L., Zorbas, C., Langhendries, J. L., Mullineux, S. T., Stamatopoulou, V., Mullier, R., et al. (2013). The complexity of human ribosome biogenesis revealed by systematic nucleolar screening of pre-rRNA processing factors. Mol. Cell 22, 539–551. doi: 10.1016/j.molcel.2013.08.011
Talkish, J., Zhang, J., Jakovljevic, J., Horsey, E. W., and Woolford, J. L. Jr. (2012). Hierarchical recruitment into nascent ribosomes of assembly factors required for 27SB pre-rRNA processing in Saccharomyces cerevisiae. Nucleic Acids Res. 40, 8646–8661. doi: 10.1093/nar/gks609
Thomson, E., Rappsilber, J., and Tollervey, D. (2007). Nop9 is an RNA binding protein present in pre-40S ribosomes and required for 18S rRNA synthesis in yeast. RNA 13, 2165–2174. doi: 10.1261/rna.747607
Tsukaya, H., Byrne, M. E., Horiguchi, G., Sugiyama, M., Van Lijsebettens, M., and Lenhard, M. (2013). How do ‘housekeeping’ genes control organogenesis? –unexpected new findings on the role of housekeeping genes in cell and organ differentiation. J. Plant Res. 126, 3–15. doi: 10.1007/s10265-012-0518-2
Turner, A. J., Knox, A. A., Prieto, J. L., McStay, B., and Watkins, N. J. (2009). A novel small-subunit processome assembly intermediate that contains the U3 snoRNP, Nucleolin, RRP5, and DBP4. Mol. Cell. Biol. 29, 3007–3017. doi: 10.1128/MCB.00029-09
Waites, R., and Hudson, A. (1995). phantastica: a gene required for dorsoventrality of leaves in Antirrhinum majus. Development 121, 2143–2154.
Warner, J. R. (1999). The economics of ribosome biosynthesis in yeast. Trends Biochem. Sci. 24, 437–440. doi: 10.1016/S0968-0004(99)01460-7
Weis, B. L., Kovacevic, J., Missbach, S., and Schleiff, E. (2015a). Plant-specific features of ribosome biogenesis. Trends Plant Sci. 20, 729–740. doi: 10.1016/j.tplants.2015.07.003
Weis, B. L., Palm, D., Missbach, S., Bohnsack, M. T., and Schleiff, E. (2015b). atBRX1-1 and atBRX1-2 are involved in an alternative rRNA processing pathway in Arabidopsis thaliana. RNA 21, 415–425. doi: 10.1261/rna.047563.114
Weis, B. L., Missbach, S., Marzi, J., Bohnsack, M. T., and Schleiff, E. (2014). The 60S associated ribosome biogenesis factor LSG1-2 is required for 40S maturation in Arabidopsis thaliana. Plant J. 80, 1043–1056. doi: 10.1111/tpj.12703
Wieckowski, Y., and Schiefelbein, J. (2012). Nuclear ribosome biogenesis mediated by the DIM1A rRNA dimethylase is required for organized root growth and epidermal patterning in Arabidopsis. Plant Cell 24, 2839–2856. doi: 10.1105/tpc.112.101022
Winter, D., Vinegar, B., Nahal, H., Ammar, R., Wilson, G. V., and Provart, N. J. (2007). An “electronic fluorescent pictograph” browser for exploring and analyzing large-scale biological data set. PLOS ONE 2:e718. doi: 10.1371/journal.pone.0000718
Yao, Y., Ling, Q., Wang, H., and Huang, H. (2008). Ribosomal proteins promote leaf adaxial identity. Development 135, 1325–1334. doi: 10.1242/dev.017913
Zakrzewska-Placzek, M., Souret, F. F., Sobczyk, G. J., Green, P. J., and Kufel, J. (2010). Arabidopsis thaliana XRN2 is required for primary cleavage in the pre-ribosomal RNA. Nucleic Acids Res. 38, 4487–4502. doi: 10.1093/nar/gkq172
Zhang, C., and Muench, D. G. (2015). A nucleolar PUF RNA-binding protein with specificity for a unique RNA sequence. J. Biol. Chem. 290, 30108–30118. doi: 10.1074/jbc.M115.691675
Zhang, J., Harnpicharnchai, P., Jakovljevic, J., Tang, L., Guo, Y., Oeffinger, M., et al. (2007). Assembly factors Rpf2 and Rrs1 recruit 5S rRNA and ribosomal proteins rpL5 and rpL11 into nascent ribosomes. Genes Dev. 21, 2580–2592. doi: 10.1101/gad.1569307
Keywords: OLI2, GDP1, ribosome biogenesis, cell proliferation, adaxial–abaxial polarity regulation, leaf development, Arabidopsis
Citation: Kojima K, Tamura J, Chiba H, Fukada K, Tsukaya H and Horiguchi G (2018) Two Nucleolar Proteins, GDP1 and OLI2, Function As Ribosome Biogenesis Factors and Are Preferentially Involved in Promotion of Leaf Cell Proliferation without Strongly Affecting Leaf Adaxial–Abaxial Patterning in Arabidopsis thaliana. Front. Plant Sci. 8:2240. doi: 10.3389/fpls.2017.02240
Received: 26 October 2017; Accepted: 20 December 2017;
Published: 09 January 2018.
Edited by:
Yasunori Machida, Nagoya University, JapanReviewed by:
Taisuke Nishimura, Nagaoka University of Technology, JapanYoshihisa Ueno, Ryukoku University, Japan
Copyright © 2018 Kojima, Tamura, Chiba, Fukada, Tsukaya and Horiguchi. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Gorou Horiguchi, Z2hvcmlAcmlra3lvLmFjLmpw
†Present address: Koji Kojima, Department of Biomolecular Engineering, Graduate School of Engineering, Tohoku University, Miyagi, Japan
 Koji Kojima
Koji Kojima Junya Tamura1
Junya Tamura1 Hirokazu Tsukaya
Hirokazu Tsukaya Gorou Horiguchi
Gorou Horiguchi