- 1State Key Laboratory for Rice Biology, College of Life Sciences, Zhejiang University, Hangzhou, China
- 2Institute of Plant Protection, Jiangsu Academy of Agricultural Sciences, Nanjing, China
- 3State Key Laboratory for Rice Biology, Biotechnology Institute, Zhejiang University, Hangzhou, China
- 4Key Laboratory for Cell and Gene Engineering of Zhejiang Province, Zhejiang University, Hangzhou, China
The glycerol-3-phosphate (G-3-P) shuttle is an important pathway for delivery of cytosolic reducing equivalents into mitochondrial oxidative phosphorylation, and plays essential physiological roles in yeast, plants, and animals. However, its role has been unclear in filamentous and pathogenic fungi. Here, we characterize the function of the G-3-P shuttle in Pyricularia oryzae by genetic and molecular analyses. In P. oryzae, a glycerol-3-phosphate dehydrogenase 1 (PoGpd1) is involved in NO production, conidiation, and utilization of several carbon sources (pyruvate, sodium acetate, glutamate, and glutamine). A glycerol-3-phosphate dehydrogenase 2 (PoGpd2) is essential for glycerol utilization and fungal development. Deletion of PoGPD2 led to delayed aerial hyphal formation, accelerated aerial hyphal collapse, and reduced conidiation on complete medium (CM) under a light–dark cycle. Aerial mycelial surface hydrophobicity to water and Tween 20 was decreased in ΔPogpd2. Melanin synthesis genes required for cell wall construction and two transcription factor genes (COS1 and CONx2) required for conidiation and/or aerial hyphal differentiation were down-regulated in the aerial mycelia of ΔPogpd2 and ΔPogpd1. Culturing under continuous dark could complement the defects of aerial hyphal differentiation of ΔPogpd2 observed in a light–dark cycle. Two light-sensitive protein genes (PoSIR2 encoding an NAD+-dependent deacetylase and TRX2 encoding a thioredoxin 2) were up-regulated in ΔPogpd2 cultured on CM medium in a light–dark cycle. ΔPogpd2 showed an increased intracellular NAD+/NADH ratio and total NAD content, and alteration of intracellular ATP production. Culturing on minimal medium also could restore aerial hyphal differentiation of ΔPogpd2, which is deficient on CM medium in a light–dark cycle. Two glutamate synthesis genes, GDH1 and PoGLT1, which synthesize glutamate coupled with oxidation of NADH to NAD+, were significantly up-regulated in ΔPogpd2 in a light–dark cycle. Moreover, deletion of PoGpd1 or PoGpd2 led to reduced virulence of conidia or hyphae on rice. The glycerol-3-phosphate shuttle is involved in cellular redox, fungal development, and virulence in P. oryzae.
Introduction
The glycerol-3-phosphate shuttle is one of mechanisms channeling cytosolic reducing equivalents to the mitochondrial oxidative phosphorylation pathway (Ansell et al., 1997; Larsson et al., 1998; Rigoulet et al., 2004). In this shuttle, dihydroxyacetone phosphate (DHAP) is converted to glycerol-3-phosphate (G-3-P) by a cytoplasmic glycerol-3-phosphate dehydrogenase 1 (Gpd1 or cGPDH) via oxidizing one molecule of NADH (nicotinamide adenine dinucleotide hydride) to NAD+ (Ansell et al., 1997). G-3-P is then converted back to DHAP by a mitochondrial glycerol-3-phosphate dehydrogenase 2 (Gpd2 or mGPDH) which reduces one molecule of flavin adenine dinucleotide (FAD) to FADH2 (Ronnow and Kielland-Brandt, 1993). FADH2 then enters into mitochondrial respiration by reducing ubiquinone (coenzyme Q) to ubiquinol (QH2) and finally generates adenosine-triphosphate (ATP) (Estabrook and Sacktor, 1958; Lee and Lardy, 1965) (Figure 1A). Gpd1 is a NAD+-dependent dehydrogenase localized in the cytosol or membrane (Gee et al., 1988; Wei et al., 2001; Ou et al., 2006). Gpd2 is a FAD-linked ubiquinone oxidoreductase, with its FAD site in the mitochondrial intermembrane space, and its coenzyme Q-binding site located in the outer leaflet of the mitochondrial inner membrane (Klingenberg, 1970; Cole et al., 1978; Yeh et al., 2008).
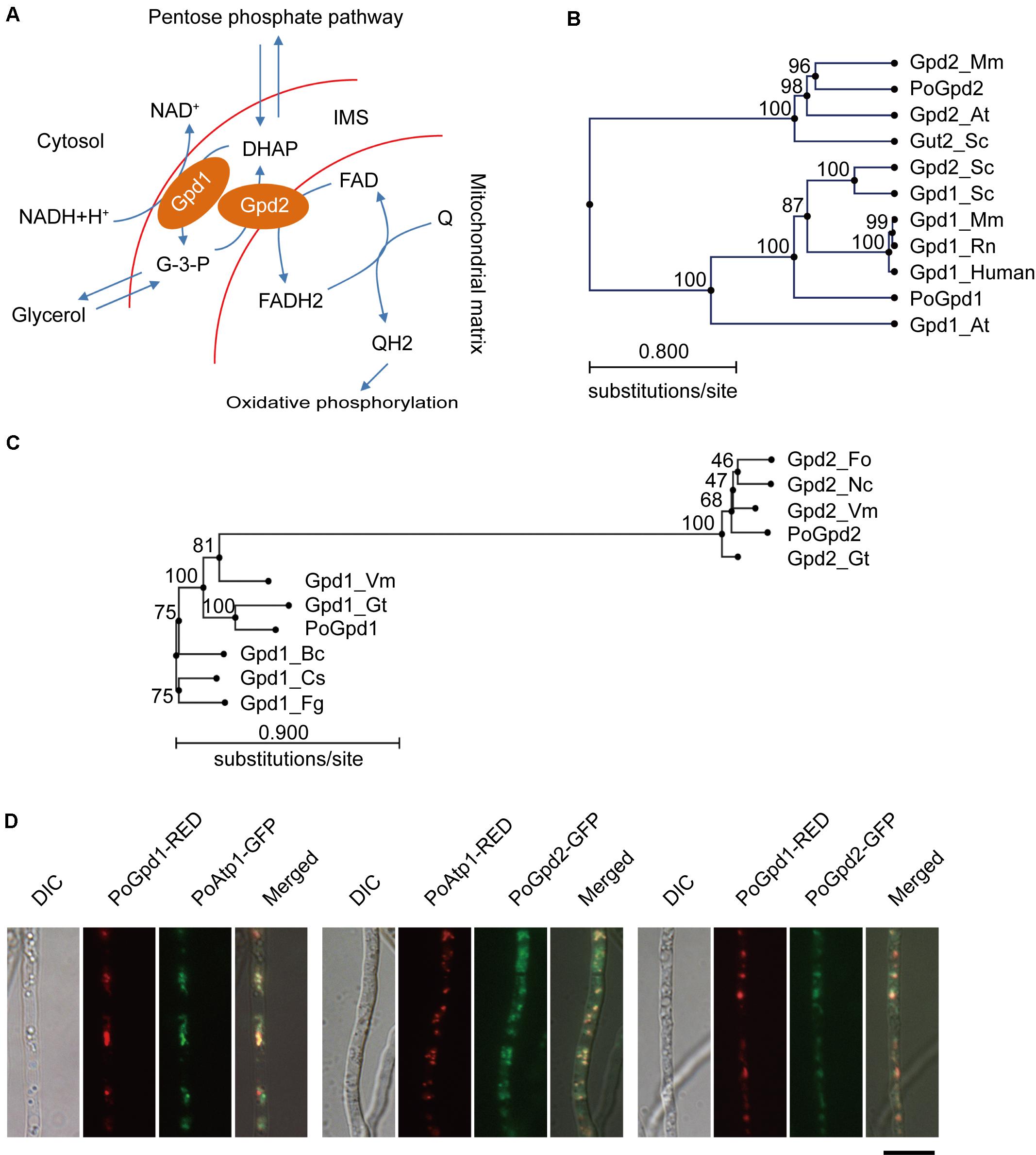
FIGURE 1. Glycerol-3-phosphate dehydrogenases in Pyricularia oryzae. (A) Glycerol-3-phosphate shuttle pathway in P. oryzae. PoGpd1 and PoGpd2 are Glycerol-3-phosphate dehydrogenase 1 and 2. DAHP, dihydroxyacetone phosphate. FAD, flavin adenine dinucleotide. Q, coenzyme Q. (B) Phylogenetic tree (Neighbor-Joining) of PoGpd1 and PoGpd2 homologs in different organisms constructed using CLC Main Workbench 5.5. The PoGpd1 and PoGpd2 homologs were Gpd1_Mm (NP_034401.1) in Mus musculus, Gpd1_Rn (NP_071551.2) in Rattus norvegicus, Gpd1_human (NP_005267.2) in human, Gpd1_At (O22216.1) in Arabidopsis thaliana, Gpd1_Sc (CAA54189.1) in Saccharomyces cerevisiae, Gpd2_Sc (NP_014582.1) in S. cerevisiae, Gpd2_Mm (NP_034404.3) in M. musculus, Gpd2-At (Q9SS48.1) in A. thaliana, and Gut2_Sc (CAA50652.1) in S. cerevisiae. The numbers at branch nodes are the bootstrap values. (C) Phylogenetic tree (Neighbor-Joining) of PoGpd1 and PoGpd2 homologs in different fungi. The PoGpd1 and PoGpd2 homologs were Gpd2_Gt (XP_009227424.1) in Gaeumannomyces tritici, Gpd2_Vm (KUI73636.1) in Valsa mali, Gpd2_Fo (EWZ35447.1) in Fusarium oxysporum, Gpd2_Nc (XP_963445.2) in Neurospora crassa, Gpd1_Fg (XP_011323505.1) in Fusarium graminearum, Gpd1_Gt (XP_009225248.1) in G. tritici, Gpd1_Vm (KUI66172.1) in V. mali, Gpd1_Bc (CCD43082.1) in Botrytis cinerea, Gpd1_Cs (KXH24909.1) in Colletotrichum salicis. (D) Subcellular localization of PoGpd1 and PoGpd2 in mitochondria of P. oryzae hyphae. Both PoGpd1-RED and PoGpd2-GFP fusion protein were co-localized with an alpha subunit of mitochondrial F1F0 ATP synthase PoAtp1. Bar = 10 μm.
The function of the G-3-P shuttle has been extensively studied in animals, plants, and yeast (Ansell et al., 1997; Larsson et al., 1998; Rigoulet et al., 2004; Shen et al., 2006; Mracek et al., 2013). In mammalian mitochondria, the G-3-P shuttle links glycolysis, oxidative phosphorylation, and fatty acid metabolism (Mracek et al., 2013). In mouse tissues, a substantial portion of superoxide and H2O2 on both sides of the mitochondrial inner membrane are generated by mGPDH (Gpd2) (Orr et al., 2012). Mouse pups lacking both Gpd1 and Gpd2 failed to grow and usually died within the first week of age (Brown et al., 2002). The mitochondrial G-3-P shuttle is similarly important in plant cells, where it is involved in redox homeostasis (Shen et al., 2006). The FAD-GPDH gene (GPD2) in Arabidopsis thaliana is expressed at a high level during seed germination, when metabolism of glycerol derived from storage lipids occurs (Shen et al., 2003). In A. thaliana, disruption of GPDHc1, which encodes a cytosolic G-3-P dehydrogenase (Gpd1), led to decreased NAD+/NADH ratios under standard growth conditions and impaired adjustment of NAD+/NADH ratios under stress conditions imposed by abscisic acid (Shen et al., 2006). Accumulation of abscisic acid is associated with a variety of plant responses to stresses (Shen et al., 2006). NAD is an essential coenzyme for many cellular redox reactions and energy metabolism, and is also involved in transcription regulation, phosphate-responsive signaling pathway, and other biological processes (Anderson et al., 2003; Lin and Guarente, 2003; Kato and Lin, 2014). NAD is synthesized through the de novo pathway (de novo synthesis of NAD from tryptophan) and the NAD salvage pathway (regeneration of NAD from its nicotinamide degradation products) (Anderson et al., 2002).
The G-3-P shuttle is required for glycerol metabolism in yeast. Saccharomyces cerevisiae contains one mitochondrial glycerol-3-phosphate dehydrogenase (Gut2) and two cytoplasmic glycerol-3-phosphate dehydrogenases (Gpd1 and Gpd2) which are only partially redundant in functions (Ronnow and Kielland-Brandt, 1993; Albertyn et al., 1994; Ansell et al., 1997; Valadi et al., 2004). Gpd1 occurs in both the cytosol and peroxisomes, while Gpd2 is found both in the cytosol and in mitochondria (Valadi et al., 2004; Jung et al., 2010). A GUT2-deletion mutant cannot utilize glycerol as a carbon source (Ronnow and Kielland-Brandt, 1993), and GPD1-deletion mutants produce little glycerol (Albertyn et al., 1994). Glycerol production is essential for the growth of yeast cells during hyperosmotic stress. The transcription of GPD1 is regulated by the high osmolarity glycerol response (HOG) pathway, and GPD1-deletion mutants are sensitive to osmotic stress (Albertyn et al., 1994). The G-3-P shuttle is also active in maintaining a cellular redox balance, and is required to endure hypoxia stress (Ansell et al., 1997; Bjorkqvist et al., 1997; Larsson et al., 1998). Under anaerobic conditions, transcription of the GPD2 gene increases, and deletion of GPD2 leads to slow growth in yeast (Ansell et al., 1997; Bjorkqvist et al., 1997).
The rice blast fungus Pyricularia oryzae (syn. Magnaporthe oryzae) is a fungal pathogen which causes serious diseases in cultivated rice, and is a primary model to study the interactions between plants and hemibiotrophic fungal pathogens (Talbot, 2003; Dean et al., 2012). The rice blast spreads in fields through three-celled conidia, production of which from aerial hyphae is activated by light. Hydrophobic proteins affect aerial hyphal differentiation and subsequently conidiation (Elliot and Talbot, 2004). Conidial production is regulated by several known transcription factors (TF), such as CNF1, CON7, COS1, PoAP1, and PoHOX2 (Odenbach et al., 2007; Kim et al., 2009; Yang et al., 2009; Zhou et al., 2009; Li et al., 2010; Guo et al., 2011; Lu et al., 2014; Cao et al., 2016). The conidium germinates and forms a specialized infection structure, the appressorium (Tucker and Talbot, 2001). A mature melanized appressorium penetrates the plant cuticle via mechanical force imposed by turgor pressure inside an appressorial cell (Howard et al., 1991; Talbot, 2003). Hyphal and appressorial melanization is regulated by melanin synthesis proteins in P. oryzae (Chumley and Valent, 1990; Thompson et al., 2000). Appressorium turgor is generated by a high concentration of compatible solutes, primarily glycerol, which can reach 4 mol/L (M) in an appressorium (deJong et al., 1997). Glutamine metabolism plays important roles in fungal development and virulence in P. oryzae, such as those performed by PoGlt1 (Zhou et al., 2017), Mgd1/Gdh2 (Oh et al., 2008), Gln1 and Gln2 (Marroquin-Guzman and Wilson, 2015). Glutamate is deaminized to α-ketoglutarate by a glutamate dehydrogenase (Mgd1), or transformed to glutamine by glutamine synthetases (Gln1 and Gln2). Glutamine and α-ketoglutarate are synthesized to two molecules of glutamate by Glt1. Fungal infection in rice requires NAD+ production meditated by a glyoxylate aminotransferase Agt1 (Bhadauria et al., 2012), NADPH production through the non-oxidative pentose phosphate pathway (Wilson et al., 2010), and ATP production by a transketolase Tkl1 (Fernandez et al., 2014b) or by an electron-transferring flavoprotein dehydrogenase EtfB (Li et al., 2016) in P. oryzae. The involvement of the G-3-P shuttle in glycerol metabolism and regulation of NAD+/NADH ratio suggests its potentially important roles in the development and pathogenicity of P. oryzae and other filamentous pathogenic fungi.
In this study, we characterize the roles of the G-3-P shuttle in the rice blast fungus by deleting two glycerol-3-phosphate dehydrogenase genes GPD1 and GPD2, and find that the G-3-P shuttle is required for maintaining the NAD+/NADH ratio, ATP production, glycerol catabolism, aerial hyphal differentiation, conidiation, and pathogenicity in P. oryzae.
Materials and Methods
Strains, Culture Conditions, PCR, and Statistical Test
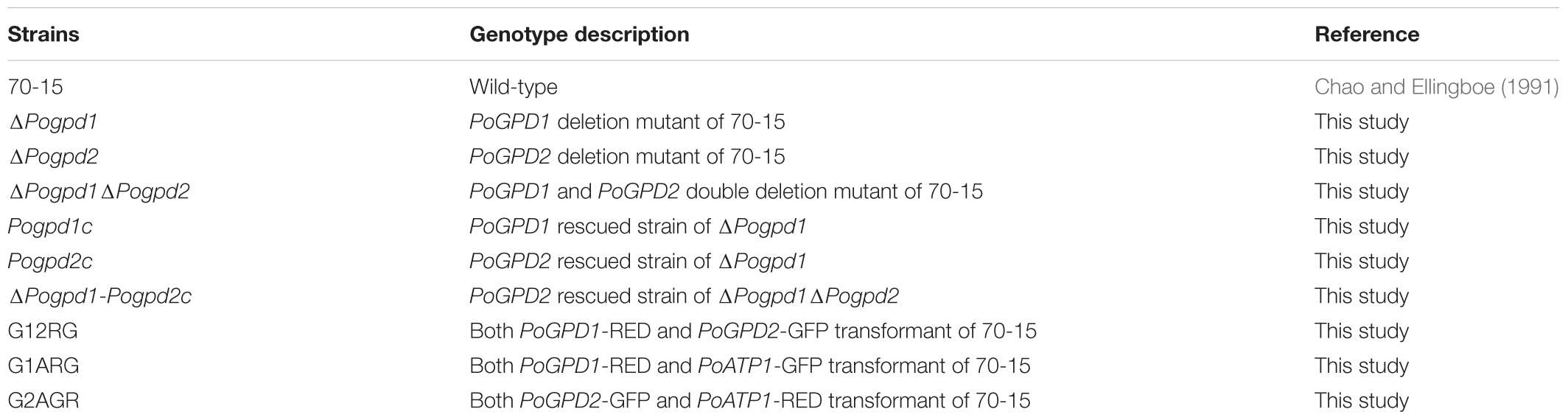
Pyricularia oryzae strains (Table 1) were stored on filter disks at -20°C. Growth tests of P. oryzae were performed in complete medium (CM) (Talbot et al., 1993), minimal medium (MM) (CM medium without peptone, yeast extract, and casamino acid), CM or MM media in which 1% (w/v) glucose was replaced by 1% (v/v) glycerol, 50 mM sodium acetate, 5 mM sodium pyruvate, 1.15% sodium glutamate, 1% glutamine, or 1% olive oil, and CM or MM media supplemented with different chemicals (0.8 M NaCl, 1 M sorbitol, 0.5 mM H2O2 or 0.8 mM Paraquat) at 25°C under a light–dark cycle (16 h–8 h) or under continuous dark. Fungal samples were ground in liquid nitrogen and total RNA was extracted with the Trizol method following the manufacturer’s procedure (TaKaRa, Japan). Total RNA (500 ng) was reverse transcribed into first-strand cDNA using a PrimeScriptTM RT reagent kit with gDNA Eraser (Takara, Japan). Quantitative real time PCR (qPCR) was conducted with five biological replicates using SYBR Premix Ex Taq (Tli RNaseH Plus) kit following the manufacturer’s protocol (TaKaRa, Japan) on a Real-Time PCR Detection System Mastercycler (Eppendorf, Germany). To compare relative abundance of transcripts, average threshold cycle (CT) was normalized to β-TUBULIN and H3 for each strain as 2-ΔCT, where ΔCT = [CTgene – (CTβ-TUBULIN + CTH3)/2]. Fold changes among different strains were calculated as 2-ΔΔCT, where ΔΔCT = ΔCTstrain_1-ΔCTstrain_2 (Livak and Schmittgen, 2001). The PCR primers used in this study are listed in Supplementary Table S1. Tukey’s HSD test was used for all experimental data in this study (Tang and Zhang, 2013).
Generation and Complementation of Null Mutants
The mutants used in this study were generated via a high-throughput gene knockout procedure (Lu et al., 2014). Briefly, 1.0 – 1.2 kb DNA fragments of the 5′ and 3′ flanking sequences of the target gene, the selectable marker gene, and the XbaI/HindIII linearized yeast-Escherichi-Agrobacterium shuttle vector pKO1B were transformed into yeast FY834 competent cells to generate knockout vectors via yeast recombinational cloning. The knockout cassettes were then transformed into the germinating spores of the wild type strain (70-15) via Agrobacterium tumefaciens-mediated transformation (ATMT) (de Groot et al., 1998; Rho et al., 2001). The knockout vector pKO1B contained a GFP gene under the control of the strong promoter of P. oryzae H3 gene. In ectopic transformants, a gene-deletion cassette and a GFP gene were ectopically integrated into the genomic DNA together. GFP was activated in ectopic transformants and could be used as a negative selective marker for null mutants. Null mutants were identified based on the following criteria: a mutant could grow on the selection medium containing 100 μg/ml sulfonylurea or 200 μg/ml hygromycin B (1st positive marker), but did not emit GFP fluorescence (1st negative marker); the specific gene could be identified in the wild type, but not in the mutant, when using double PCR (2nd negative marker) with β-TUBULIN as a positive control; a unique recombinational DNA fragment that indicated a knockout event could be identified in the mutant by PCR, but not in the wild type (2nd positive marker) (Supplementary Figure S1A); only a single copy of the selectable marker gene (SUR or HPH) was confirmed in the mutant by qPCR (single insertion of a knockout cassette) (Lu et al., 2014). The mutants were complemented with their respective native copy of genes in the wild type strain 70-15. The complement strains were confirmed at a transcriptional level (Supplementary Figure S1B).
Observation of Fluorescence Fusion Proteins in Mutants
The coding sequence of PoGPD1, PoGPD2, and PoATP1 (MGG_07752) was amplified from wild type genomic DNA and inserted into selected GFP or DsRED fusion plasmids. PoATP1 was cloned into pKD8-GFP and pKD8-RED containing a G418 resistance gene (NEO) (Li et al., 2012). PoGPD1 was cloned into pKD5-RED which contains a sulfonylurea resistance gene (SUR) (Li et al., 2012). PoGPD2 was cloned into pKD9-GFP which contains a hygromycin B phosphotransferase gene (HPH) (Sun et al., 2017). The fluorescence fusion genes were transformed into the P. oryzae strains (ΔPogpd1, ΔPogpd2 and ΔPogpd1ΔPogpd2) via ATMT. Samples were analyzed via fluorescence microscopy (Filter: FITC and TRITC) (Nikon eclipse 80i, Japan).
Phenotypic Characterization
Phenotypes of the P. oryzae strains were analyzed according to a previously described schema (Lu et al., 2007, 2014). All assays were performed with five biological replicates, and repeated three times. Mycelial growth was evaluated by measuring the diameter of the colony at 8 dpi on CM medium. Conidia of mycelia grown on CM medium were collected and counted at 8 dpi as a metric of conidia production. To measure conidial germination and appressorium formation, 40 μl of a spore suspension (1 × 105 conidia/ml) was inoculated on hydrophobic plastic coverslips and incubated under high humidity at 25°C. The conidial germination rate and the appressorial formation rate were digitized at 4 hr post inoculation (hpi) and 24 hpi respectively. At least 200–300 conidia were counted. Appressorium turgor was evaluated by incipient cytorrhysis (cell collapse) assays as described previously (Howard et al., 1991). A series of glycerol solutions (0.5, 1, and 2 M) replaced water in droplets of spore suspension at 48 hpi and was incubated for 5 min before counting the collapsed appressoria (n = 3, >300 appressoria/experiment). Surface hydrophobicity assays were performed by placing a droplet of 20 μl – 40 μl sterile distilled water or of a solution containing 100 μg/ml Tween 20 onto the surface of aerial mycelia (Kim et al., 2005).
Pathogenicity Assay
The virulence of P. oryzae strains was investigated by inoculation of mycelial blocks on leaf explants of 8-day-old barley (Hordeum vulgare) or 14-day-old rice (Oryza sativa cv CO39), and by conidial spray inoculation on 14-day-old rice seedlings, as previously described (Lu et al., 2007). For the mycelial virulence assays, 5-mm mycelial blocks from agar plates were inoculated on intact leaf explants of rice and barley, and the disease lesions were assayed after culturing at 25°C for 4 days. The conidial virulence assays were conducted by spraying 2 ml of a spore suspension (1 × 105 conidia/ml) in 0.2% (w/v) gelatin onto 25 – 30 rice seedlings using an artist’s airbrush. After incubation at 25°C for 1 d in a humid dark box, the inoculated plants were grown under a 12 h light/dark photoperiod until the wild type developed severe disease lesions (for 7 days). The severity of the disease lesions were assessed according to a previously reported schema (Bonman et al., 1986). Disease lesions were measured for a 5-cm length section of the most severely infected leaf of each plant. Infection assays were repeated at least three times. Host penetration by P. oryzae appressoria was assayed at 24 hpi – 96 hpi after inoculating 20 μl of conidial suspension (1 × 105 conidia/ml) on leaf explants of barley, as previously reported (Lu et al., 2007).
Quantification of Intracellular NAD+, NADH, ATP, NO, and G-3-P Content
For strains cultured under a light–dark cycle, aerial mycelium samples were collected in the daytime. Intracellular NAD+ and NADH were quantified using the NAD/NADH quantification kit according to the technical bulletin (Sigma-Aldrich, United States). 20 mg of mycelium samples were ground in liquid nitrogen, transferred to 1.5 ml tubes which contained 400 μl of NADH/NAD extraction buffer, and incubated 10 min at room temperature. Samples were centrifuged at 12,000 ×g for 5 min and the supernatants were then deproteinized by filtration through a 10 kDa cut-off spin filter (7,500 ×g for 7 min at 4°C). Total NADH and NAD+ were quantified by measuring the absorbance at 450 nm, and NADH was quantified after decomposing NAD+ by incubation at 60°C for 30 min. The concentration of total NAD or NADH was shown in pmol/mg protein. Intracellular ATP content was quantified using the enhanced ATP assay kit according to the user manual (Beyotime, China). 20 mg of mycelium samples were ground in liquid nitrogen, transferred to 1.5 ml tubes which contained 100 μl of lysis buffer, and centrifuged at 12,000 ×g for 5 min in 4°C. ATP concentration in the supernatants was quantified by measuring the intensity of fluorescence emitted by firefly luciferase using a luminometer (Berthold, Germany). The concentration of intracellular ATP was shown in nmol/mg protein. Nitric Oxide (NO) concentration in aerial mycelium samples was measured with the NO assay kit (S0021) following the user manual (Beyotime, China). The concentration of intracellular NO was shown in μmol/g protein. The G-3-P concentration in aerial mycelium was quantified with the Glycerol-3-Phosphate assay kit (MAK207) according to the technical bulletin (Sigma-Aldrich, United States). The concentration of intracellular G-3-P was shown in nmol/μg protein.
Results
Glycerol-3-Phosphate Dehydrogenases in P. oryzae
Mitochondrial glycerol-3-phosphate shuttle consists of two components: a cytoplasmic glycerol-3-phosphate dehydrogenase 1 (Gpd1/cGpdh) and a mitochondrial glycerol-3-phosphate dehydrogenase 2 (Gpd2/mGpdh) (Ronnow and Kielland-Brandt, 1993; Ansell et al., 1997) (Figure 1A). After blasting against Gpd1 (GPDHc1) and Gpd2 (FAD-GADP) proteins in A. thaliana (Shen et al., 2003, 2006) at NCBI, we characterized the homologs of a cytoplasmic glycerol-3-phosphate dehydrogenase 1 and a mitochondrial glycerol-3-phosphate dehydrogenase 2 in P. oryzae: PoGpd1 (MGG_00067) and PoGpd2 (MGG_03147) respectively. The generated phylogenetic tree showed that PoGpd1 was aligned with Gpd1 in Mus musculus (Sato et al., 2014), Rattus norvegicus (Mracek et al., 2009), human (Menaya et al., 1995), A. thaliana (Shen et al., 2006) and S. cerevisiae (Ansell et al., 1997), and with Gpd2 in S. cerevisiae (Ansell et al., 1997). PoGpd2 was aligned with Gpd2 in M. musculus (Koza et al., 1996) and A. thaliana (Shen et al., 2003), and with Gut2 in S. cerevisiae (Ronnow and Kielland-Brandt, 1993) (Figure 1B). In fungi, the homologs of Gpd2 were also aligned into a group; however, relative to Gpd2, Gpd1’s homologs had higher diversity among different fungi (Figure 1C).
The TargetP 1.1 program1 predicted that PoGpd1 and PoGpd2 were mitochondrial proteins, and the CCTOP program2 predicted that PoGpd2 contained two transmembrane segments. We co-localized eGFP- or DsRED2-tagged PoGpd1, PoGpd2 and PoAtp1 to each other in P. oryzae. Three pairs of proteins, PoGpd1-DsRED2 and PoAtp1-eGFP, PoGpd2-eGFP and PoAtp1-DsRED2, and PoGpd1-DsRED2 and PoGpd2-eGFP, were co-localized together (Figure 1D). PoAtp1 is a mitochondrial inner membrane protein (Li et al., 2016). Therefore, PoGpd1 and PoGpd2 are co-localized to a mitochondrion.
Glycerol-3-Phosphate Dehydrogenases Are Not Required for Response to Osmotic Stress
To determine whether the G-3-P shuttle is involved in response to osmotic stress in P. oryzae, the growth of ΔPogpd1 and ΔPogpd2 on hyperosmotic stress media containing 0.8 M NaCl or 1.0 M sorbitol was assayed. ΔPogpd1 and ΔPogpd2 displayed similar responses to high osmotic stresses as that of the wild type (Figure 2). This result suggested that PoGPD1 and PoGPD2 are not involved in response to osmotic stresses in P. oryzae.
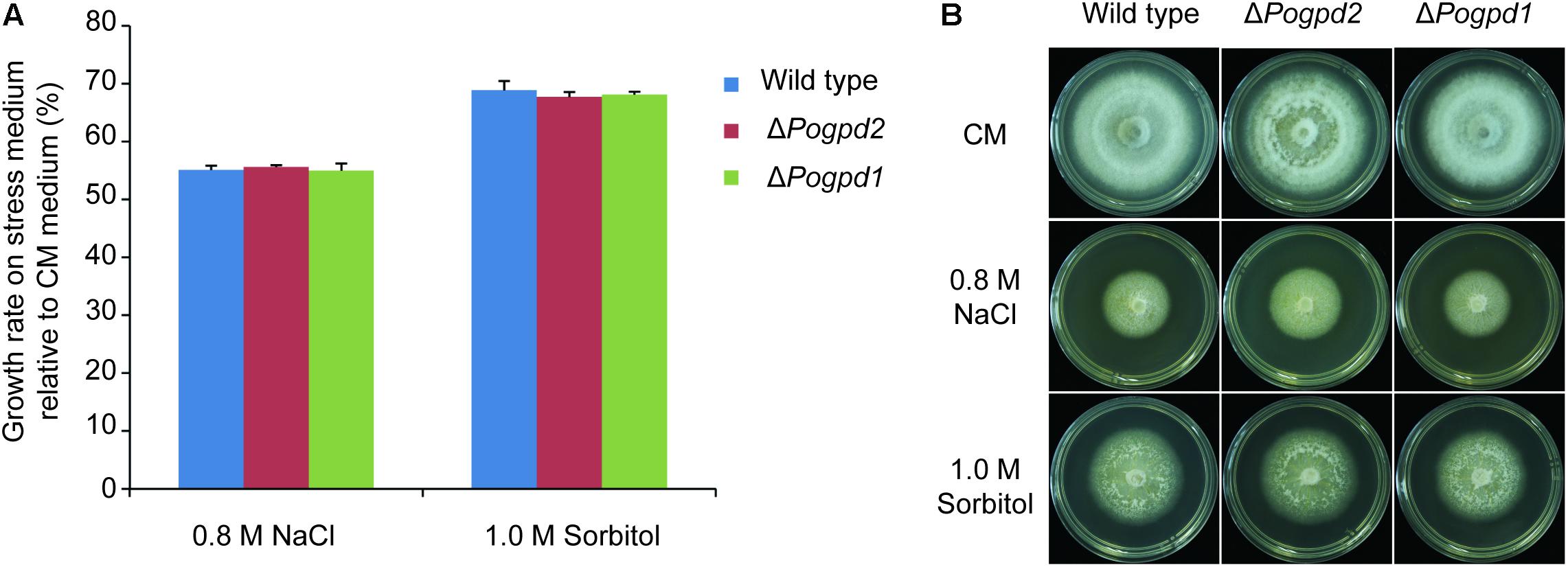
FIGURE 2. Roles of glycerol-3-phosphate dehydrogenases in response to osmotic stress. (A) Relative growth rate on CM medium containing 0.8 M NaCl or 1.0 M sorbitol by ΔPogpd1 and ΔPogpd2. Error bars represent SD. No significant differences were found between the wild type and the mutants as estimated by Tukey’s HSD test (P < 0.05). (B) Mycelial colonies of wild type, ΔPogpd1, and ΔPogpd2 grown on osmotic stress media for 8 days.
Glycerol-3-Phosphate Dehydrogenases Are Required for Utilization of Various Carbon Sources in P. oryzae
To test if PoGpd1 and PoGpd2 are also required for glycerol metabolism, we observed the growth of the mutants on revised CM and MM media in which 1% glucose was replaced by 1% glycerol under continuous dark. ΔPogpd1 grew similarly to the wild type on glycerol media. However, ΔPogpd2 and ΔPogpd1ΔPogpd2 produced much sparser aerial hyphae on glycerol media than did the wild type and ΔPogpd1 (Figures 3A,B). The reintroduction of a native PoGPD2 copy into ΔPogpd2 or ΔPogpd1ΔPogpd2 could recover the mutant’s ability in glycerol utilization, suggesting that PoGPD2 is required for utilization of glycerol in P. oryzae. When glycerol is utilized as a carbon source, it is converted to G-3-P and then DHAP by Gpd2. In the G-3-P shuttle, DHAP is converted back to G-3-P by Gpd1. We measured the G-3-P content in aerial mycelia grown on CM medium, and found 3-fold up-regulation of G-3-P in ΔPogpd2 and ΔPogpd1ΔPogpd2, relative to those in the wild type and ΔPogpd1 (Figure 3C).
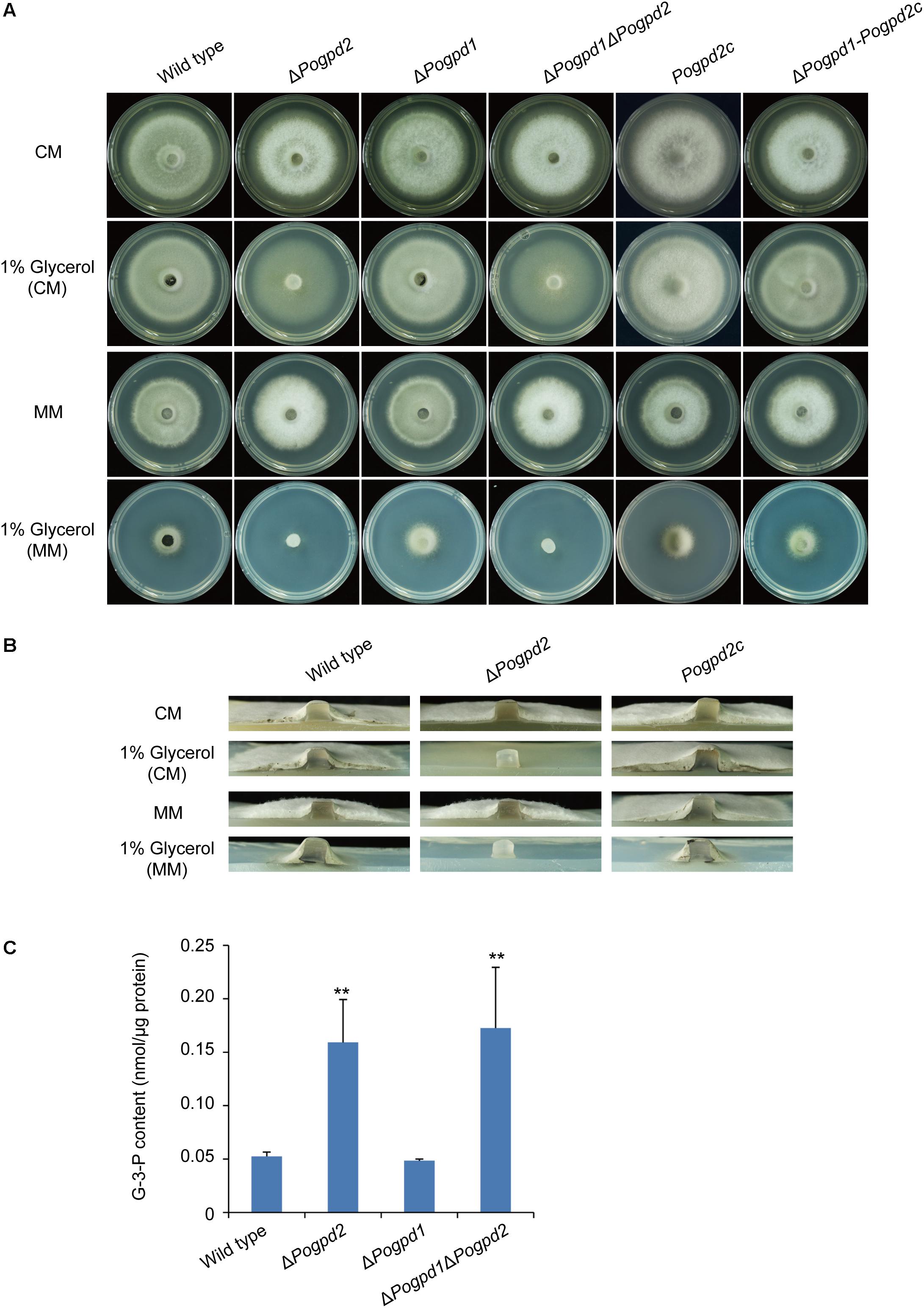
FIGURE 3. Roles of glycerol-3-phosphate dehydrogenases in glycerol catabolism. (A) Mycelial colonies of the wild type, ΔPogpd1, ΔPogpd2, and ΔPogpd1ΔPogpd2, and the complementation strains Pogpd2c and ΔPogpd1-Pogpd2c, grown on glycerol media for 8 days. (B) Aerial mycelia of the wild type, ΔPogpd2, and its complementation strain Pogpd2c grown on glycerol media. (C) The glycerol-3-phosphate (G-3-P) content of aerial hyphal cells in the wild type, ΔPogpd1, ΔPogpd2, and ΔPogpd1ΔPogpd2 cultured on CM medium. Error bars represent SD. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗∗P < 0.01.
We then assayed the growth of the mutants on MM media containing different carbon sources, in which glucose was substituted by non-fermentable carbon sources (pyruvate, sodium acetate, and olive oil) or glycogenic amino acid (glutamate and glutamine). ΔPogpd1 exhibited severe defects in the utilization of pyruvate, acetate, glutamate and glutamine (Figure 4A). Relative to glucose, the growth rates of ΔPogpd1 on media using pyruvate, sodium acetate, glutamine, or glutamate as the sole carbon source were 54.5 ± 2.4, 48.5 ± 2.9, 54.9 ± 4.5, or 44.3 ± 0.2%, respectively, significantly lower than those of the wild type (P < 0.01) (Figure 4B). These results suggested that PoGPD1 is involved in the utilization of pyruvate, sodium acetate, glutamate, and glutamine in P. oryzae.
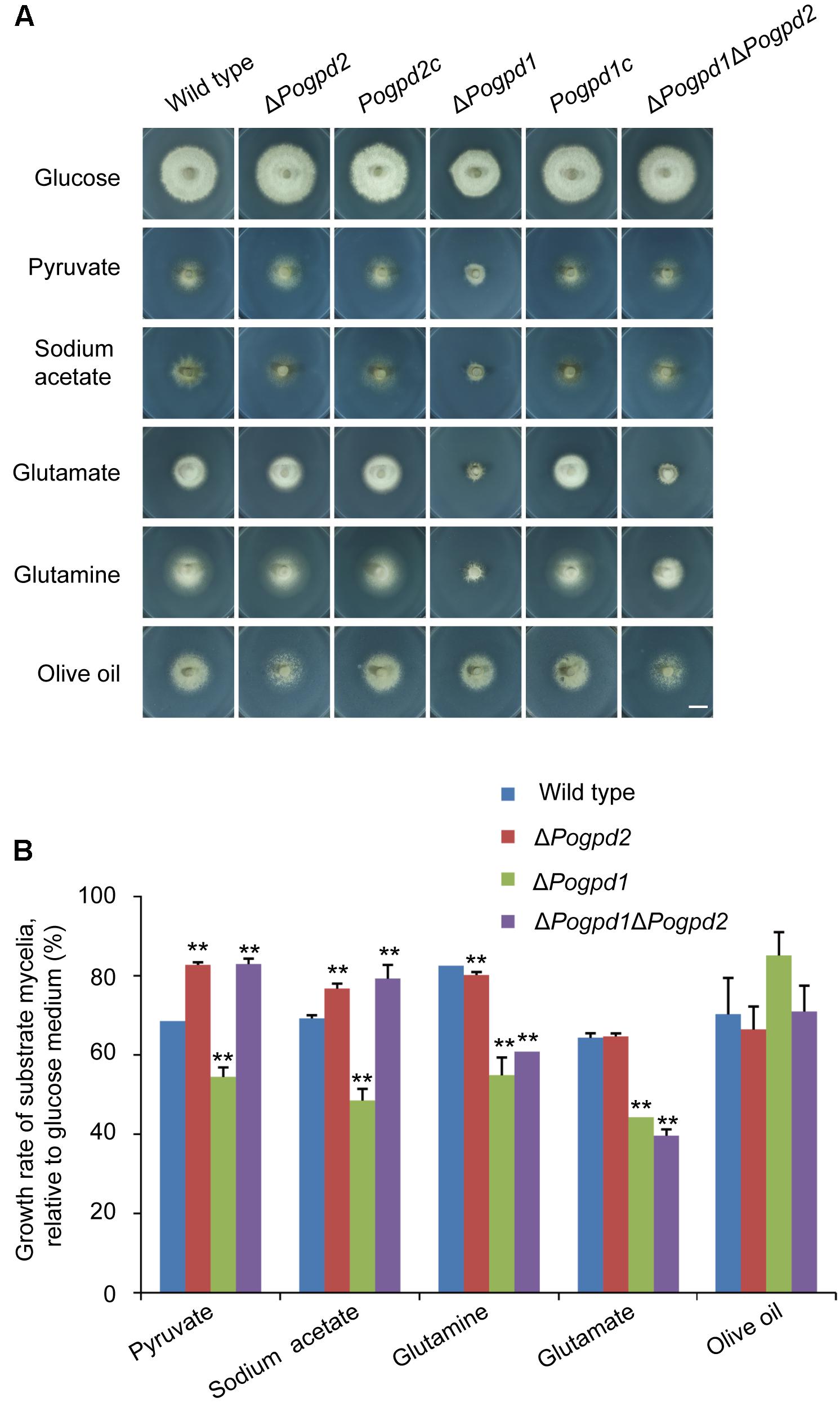
FIGURE 4. Roles of glycerol-3-phosphate dehydrogenases in utilization of carbon sources. (A) Mycelial colonies of the wild type, ΔPogpd1, ΔPogpd2, and ΔPogpd1ΔPogpd2, and the complementation strains Pogpd2c and Pogpd12c, grown at 25°C for 8 days on MM medium (glucose), or MM in which 1% glucose was replaced by 50 mM sodium acetate, 5 mM sodium pyruvate, 1.15% sodium glutamate, 1% glutamine, or 1% olive oil as the sole carbon source. Bar, 10 mm. (B) Growth rate of the wild type, ΔPogpd1, ΔPogpd2, and ΔPogpd1ΔPogpd2, cultured in MM-C medium containing 50 mM sodium acetate, 5 mM sodium pyruvate, 1.15% sodium glutamate, 1% glutamine, or 1% olive oil relative to that in MM medium (1% glucose). Error bars represent SD. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗∗P < 0.01.
Glycerol-3-Phosphate Dehydrogenases Are Required for Aerial Hyphal Growth and Conidiation
When grown on CM medium under a light–dark cycle (16 h: 8 h), the formation of aerial hyphae upward from the medium surface was delayed severely, and the formed aerial hyphae collapsed quickly in ΔPogpd2 (Figure 5A). The aerial mycelium of ΔPogpd2 was also much whiter than that of the wild type. The phenotype of ΔPogpd1 in aerial mycelium was similar to the wild type, while ΔPogpd1ΔPogpd2 was similar to ΔPogpd2 (Figure 5A). The mutants produced fewer conidia than the wild type when grown on CM medium under a light–dark cycle (Figure 5B). ΔPogpd2, ΔPogpd1, and ΔPogpd1ΔPogpd2 produced 9.4, 48.2, and 6.4%, respectively, of the output of the wild type. The complementation of PoGPD2 in ΔPogpd2 and ΔPogpd1ΔPogpd2 could restore normal aerial hyphal differentiation (Figure 5A) and conidiation (Figure 5B) in the mutants. Therefore, PoGPD2 is involved in aerial hyphal differentiation and conidiation under a light–dark cycle in P. oryzae.
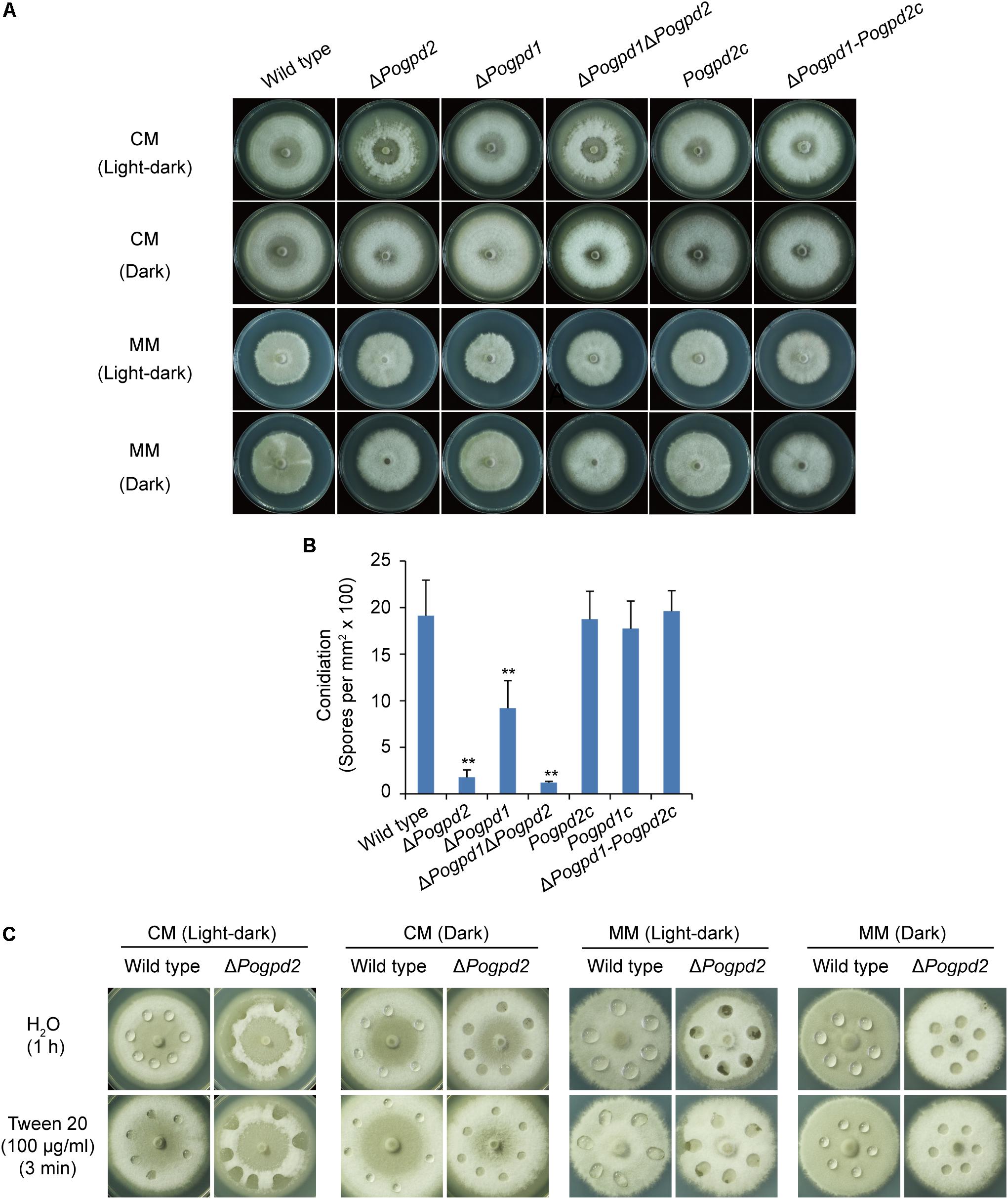
FIGURE 5. Roles of glycerol-3-phosphate dehydrogenases in aerial hyphal differentiation and conidiation. (A) Colonies of the wild type, ΔPogpd1, ΔPogpd2, and ΔPogpd1ΔPogpd2, and the complementation strains Pogpd2c and ΔPogpd1-Pogpd2c, grown on CM and MM media under a light–dark cycle or a continuous dark condition for 8 days. (B) Conidiation in P. oryzae strains (the wild type, ΔPogpd1, ΔPogpd2, and ΔPogpd1ΔPogpd2) cultured on CM medium under a light–dark cycle for 8 days. (C) Surface hydrophobicity assay for the aerial mycelia of the wild type and ΔPogpd2. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗∗P < 0.01.
To see if hyphal hydrophobicity is affected by Pogpd2, we assayed the surface hydrophobicity of the mutants. The aerial mycelia of ΔPogpd2, but not the wild type, cultured on CM and MM medium were wettable within 60 min by water or in 3 min by a detergent solution (100 μg/ml Tween 20) (Figure 5C). We then measured the expression level of four genes encoding hydrophobic proteins (MGG_09134, MPG1, MGG_10105, and MHP1) (Talbot et al., 1993; Kim et al., 2005) in the mutants by qPCR. Three genes (MGG_09134, MPG1 and MGG_10105) were significantly down-regulated (decreased 3.4-, 6.6-, and 7.3-fold, respectively) in ΔPogpd2 under the light–dark cycle. However, only one gene (MPG1) was significantly down-regulated in ΔPogpd2 (decreased to 7.4-fold) under dark conditions (Figure 6A). Hyphal color is determined by melanin content in P. oryzae. Four melanin synthesis genes (4HNR, AIB1, RSY1, and BUF1) (Chumley and Valent, 1990; Thompson et al., 2000) were down-regulated in the aerial mycelia of ΔPogpd2 (decreased 5.0-, 5.6-, 2.0-, and 5.0-fold, respectively) and ΔPogpd1 (decreased 2.8-, 2.3-, 1.6-, and 2.5-fold, respectively); and down-regulated much more in ΔPogpd2 than in ΔPogpd1 (Figure 6B). Several TF had been found to regulate conidial production. Among ten tested transcription factor genes required for conidiation (CNF1, COM1, CON7, CONx1, CONx2, COS1, GCC1, PoAP1, PoHOX2, PoLDB1) (Odenbach et al., 2007; Kim et al., 2009; Yang et al., 2009; Zhou et al., 2009; Li et al., 2010; Guo et al., 2011; Lu et al., 2014; Cao et al., 2016) (Supplementary Figure S2), four TF genes (COS1, CONx2, CON7, and GCC1) were down-regulated in the aerial mycelia of ΔPogpd2 (decreased 2.9-, 5.2-, 1.3-, and 1.2-fold, respectively), and two TF genes (COS1 and CONx2) were down-regulated in ΔPogpd1 (decreased to 1.9- and 2.0-fold) (Figure 6B).
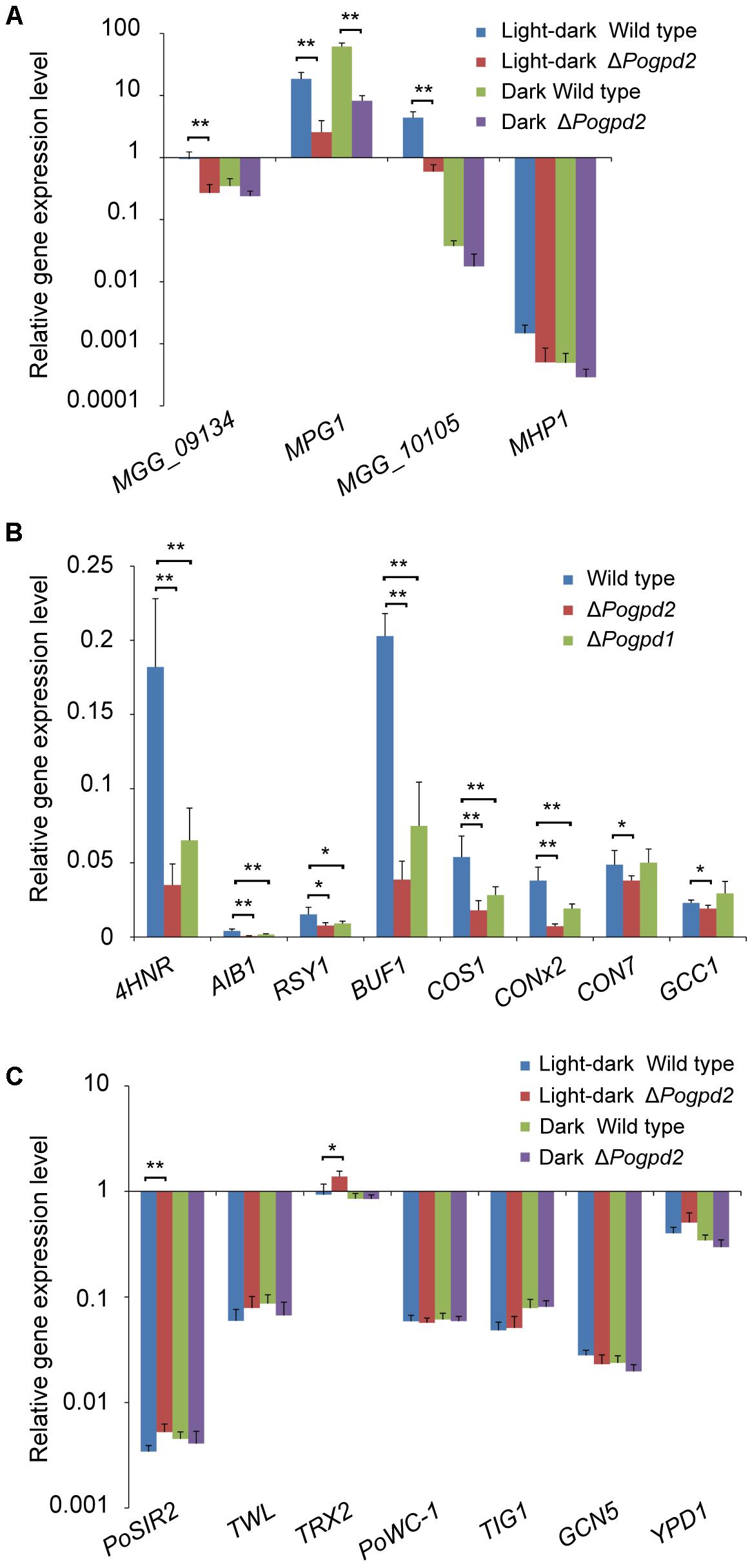
FIGURE 6. Expression level analysis of genes involved in aerial hyphal differentiation and conidiation in the mutants. (A) Relative expression level of four hydrophobic protein genes (MGG_09134, MPG1, MGG_10105, and MHP1) in the aerial mycelia of ΔPogpd2 cultured on CM medium under a light–dark cycle or a continuous dark condition. (B) Relative expression level of four melanin synthesis genes (4HNR, AIB1, RSY1, and BUF1) and four conidiation-required transcription factor genes (COS1, CONx2, CON7, and GCC1) in the aerial mycelia of ΔPogpd1 and ΔPogpd2 cultured on CM medium under a light–dark cycle. (C) Relative expression level of seven light-sensitive protein genes (PoWC-1, TWL, TIG1, GCN5, YPD1, PoSIR2, and TRX2) in the aerial mycelia of ΔPogpd2 cultured on CM medium under a light–dark cycle and a continuous dark condition. β-TUBULIN and H3 were selected as reference genes. Error bars represent SD. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗P < 0.05 and ∗∗P < 0.01.
Light Affects the Aerial Hyphae Differentiation and Maintenance of ΔPogpd2
After comparing aerial hyphae produced by ΔPogpd2 and ΔPogpd1ΔPogpd2 on CM medium, we found that the defects of aerial hyphae (delayed formation and advanced collapse) in ΔPogpd2 and ΔPogpd1ΔPogpd2 cultured under a light–dark cycle (Figures 2B, 5A) disappeared when cultured under continuous dark (Figures 3A, 5A). Aerial hyphal differentiation and conidial development were controlled by a light–dark cycle in P. oryzae. Of seven genes encoding light-sensitive proteins (PoWC-1, TWL, TIG1, GCN5, YPD1, PoSIR2, and TRX2) (Lee et al., 2006; Ding et al., 2010; Fernandez et al., 2014a; Deng et al., 2015; Zhang et al., 2016, 2017; Mohanan et al., 2017), two genes (PoSIR2 and TRX2) were found to be significantly up-regulated in ΔPogpd2 under a light–dark cycle (increased to 1.5- and 1.6-fold), but not in a continuous dark condition (Figure 6C).
Inorganic Nitrogen Restores Aerial Hyphal Differentiation of ΔPogpd2 Cultured Under a Light-Dark Cycle
Whether under a light–dark cycle or in continuous dark, the aerial mycelium of ΔPogpd2 or ΔPogpd1ΔPogpd2 cultured on MM medium, in which NaNO3 was used as the sole nitrogen source, was similar to that of the wild type and of the complementation strains (Figure 5A). We measured the expression level of seven genes assimilating NO3- in ΔPogpd2 by qPCR (Figure 7). NIA1, encoding a nitrate reductase (Samalova et al., 2012), was up-regulated significantly (2.0-fold) in ΔPogpd2 cultured on MM medium (Figure 7B). GDH1 (MGG_08074), a glutamate dehydrogenase gene, and PoGLT1, a glutamate synthase gene (Zhou et al., 2017), were up-regulated significantly in ΔPogpd2 cultured on both CM medium (8.3- and 2.6-fold) and MM medium (6.8- and 2.3-fold) in a light–dark cycle. However, only GDH1 was up-regulated significantly (4.5-fold) in ΔPogpd2 on CM medium in the dark (Figure 7B). GLN2, a glutamine synthetase gene (Marroquin-Guzman and Wilson, 2015), was up-regulated significantly (3.0-fold) in ΔPogpd2 on CM medium under a light–dark cycle.
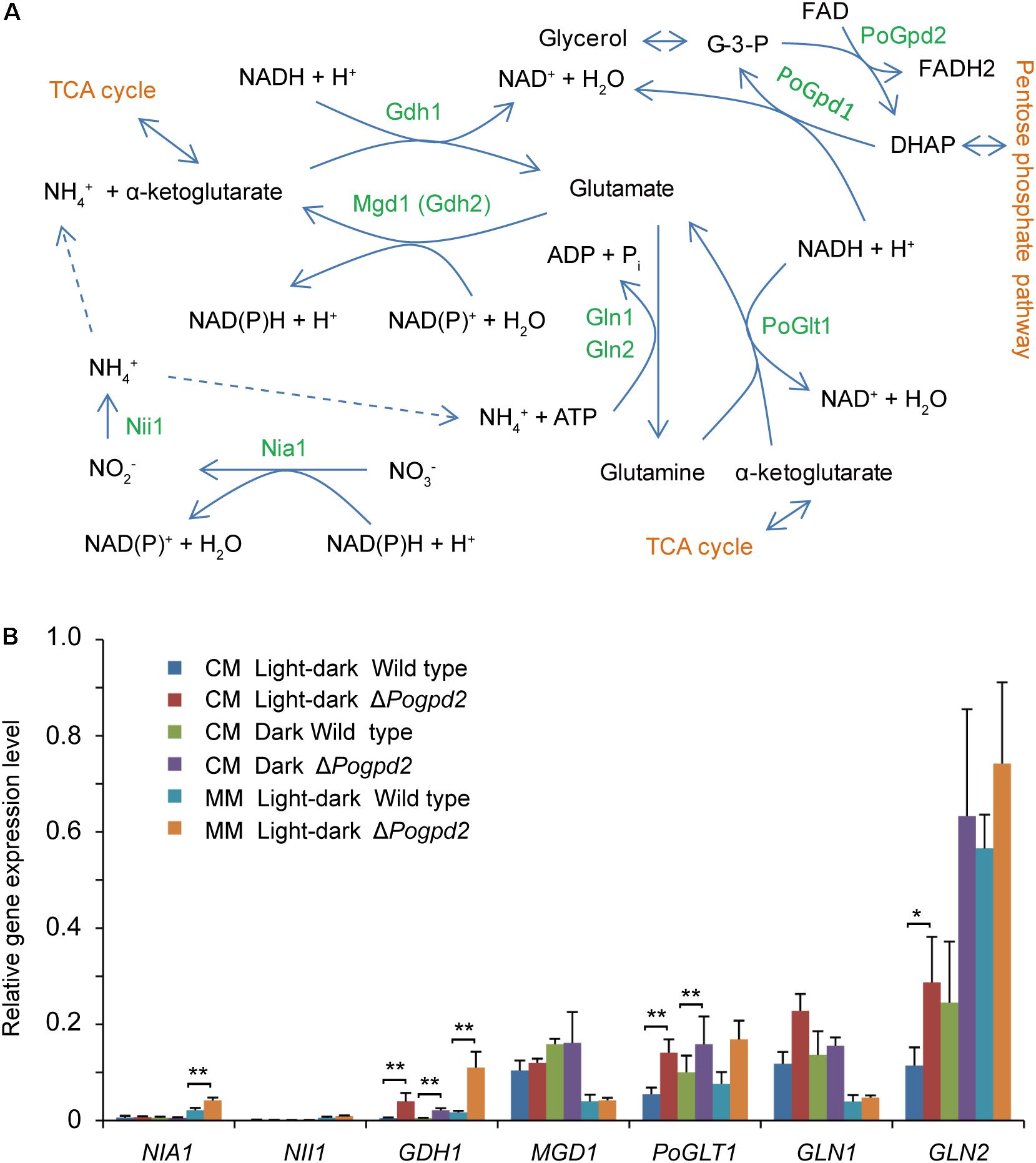
FIGURE 7. Nitrogen assimilation in Pyricularia oryzae. (A) A pathway to assimilate nitrogen in P. oryzae. (B) Relative expression level of seven genes involved in nitrate assimilation in ΔPogpd2 cultured on CM and MM media under a light–dark cycle or a continuous dark condition. β-TUBULIN and H3 were selected as reference genes. Error bars represent SD. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗P < 0.05 and ∗∗P < 0.01. TCA, Tricarboxylic acid cycle; Nia1, a nitrate reductase; Nii1, a nitrite reductase; Gdh1, a glutamate dehydrogenase; Mgd1 (Gdh2), a glutamate dehydrogenase; Gln1 and Gln2, glutamine synthetases; PoGlt1, a glutamate synthase.
Glycerol-3-Phosphate Dehydrogenase 1 Is Involved in NO Production in P. oryzae
To examine the roles of PoGpd1 and PoGpd2 in resistance to reactive oxygen species (ROS), resistance to paraquat and H2O2 by the mutants was tested. Paraquat, [(C6H7N)2]Cl2, is an oxidant that produces superoxide anions by interfering with electron transfer and that could be reduced by an electron donor (such as NADPH) in vivo (Bus and Gibson, 1984). The growth of substrate mycelia in ΔPogpd1 showed increased sensitivity to paraquat (P < 0.01), while the growth of aerial mycelia in both ΔPogpd1 and ΔPogpd2 responded to paraquat and H2O2 in a manner similar to the wild type (Figures 8A,B). We measured the NO content in aerial hyphal cells cultured on CM medium, and found that the NO contents in ΔPogpd1 and ΔPogpd1ΔPogpd2 were 1.4 ± 0.3 and 1.2 ± 0.6 μmol/g protein, significantly lower than those in the wild type and ΔPogpd2 (2.0 ± 0.2 and 1.9 ± 0.4 μmol/g protein) (P < 0.01) (Figure 8C).
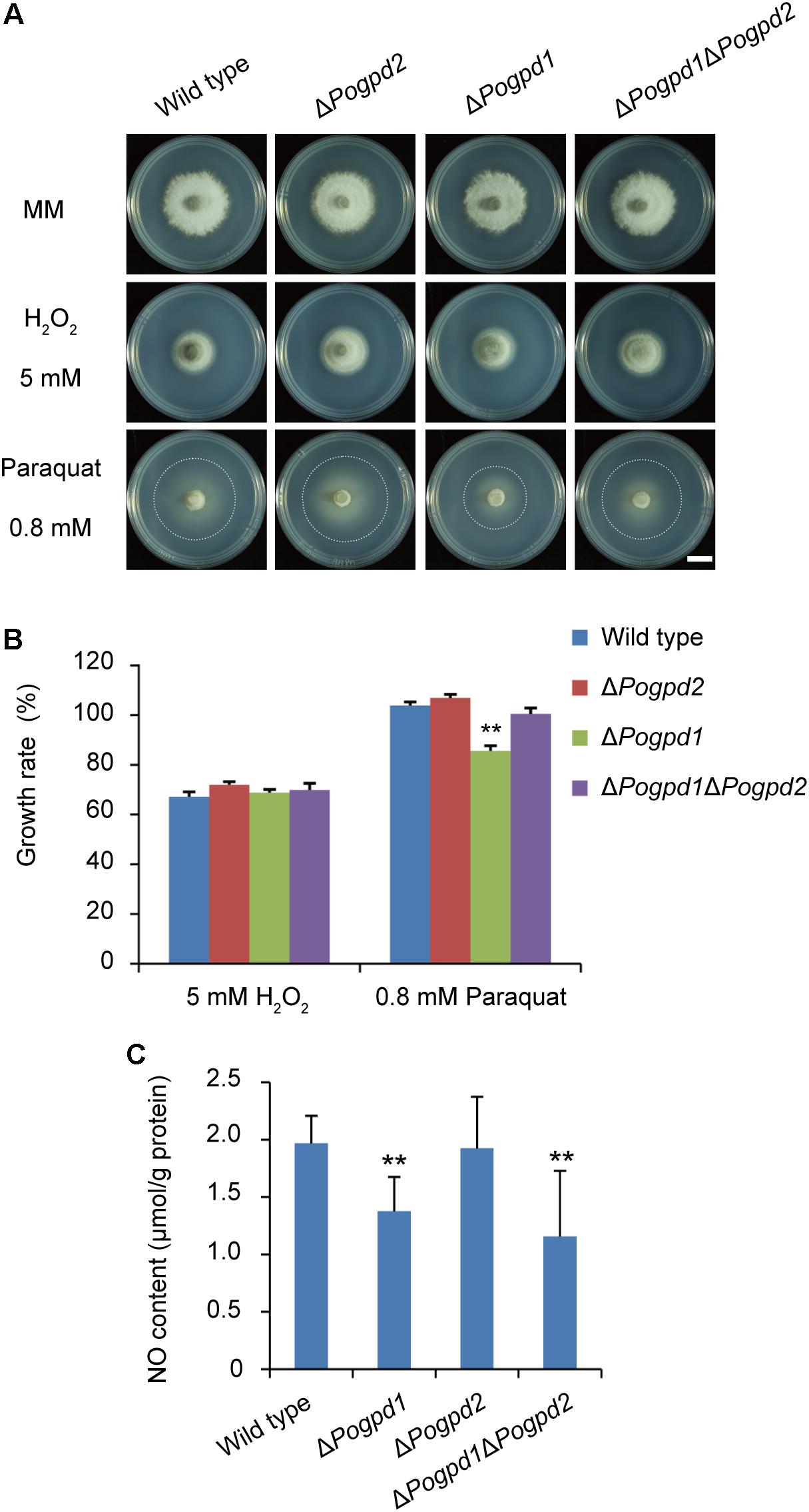
FIGURE 8. Roles of glycerol-3-phosphate dehydrogenases in response to ROS stress. (A) Colonies of the wild type, ΔPogpd1, ΔPogpd1ΔPogpd2, and ΔPogpd2 cultured on MM media with 5 mM H2O2 or 0.8 mM paraquat for 8 days. Bar = 5 mm. (B) Relative growth rate of substrate mycelia in Pyricularia oryzae strains on media with 5 mM H2O2 or 0.8 mM paraquat. Pogpd1c is a complementation strain of ΔPogpd1. (C) The NO content of aerial hyphal cells in the wild type, ΔPogpd1, ΔPogpd2, and ΔPogpd1ΔPogpd2 cultured on CM medium. Error bars represent SD. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗∗P < 0.01.
Glycerol-3-Phosphate Dehydrogenase 2 Is Involved in the NAD+/NADH Ratio and Intracellular ATP Content in P. oryzae
To evaluate the diverse roles of PoGPD2 and PoGPD1 on the glycerol-3-phosphate shuttle, we measured the NAD+/NADH ratio of the mutant aerial mycelia. The NAD+/NADH ratio of the aerial mycelia in ΔPogpd2 was higher than that of the wild type, while the value in ΔPogpd1 was similar to that of the wild type on CM and MM media (Figure 9A). NAD+/NADH ratio in ΔPogpd2 was elevated 2.6-, 2.9-, and 2.0-fold for CM medium/light–dark, CM medium/dark, and MM medium/light–dark cycles, respectively, when compared to the wild type. The NAD+/NADH ratio of the aerial mycelia was greatly elevated on MM medium when compared to CM medium in tested strains (increased 11.2-, 8.4-, 7.7-fold in the wild type, ΔPogpd2, and ΔPogpd1, respectively) (Figure 9A). The total intracellular NAD content (including both NAD+ and NADH) in the aerial mycelia of ΔPogpd2 increased 2.2- and 1.4-fold relative to the wild type cultured on CM medium (2.3 ± 0.1 pmol/μg protein) and on MM medium (3.2 ± 0.5 pmol/μg protein) under a light–dark cycle (Figure 9B).
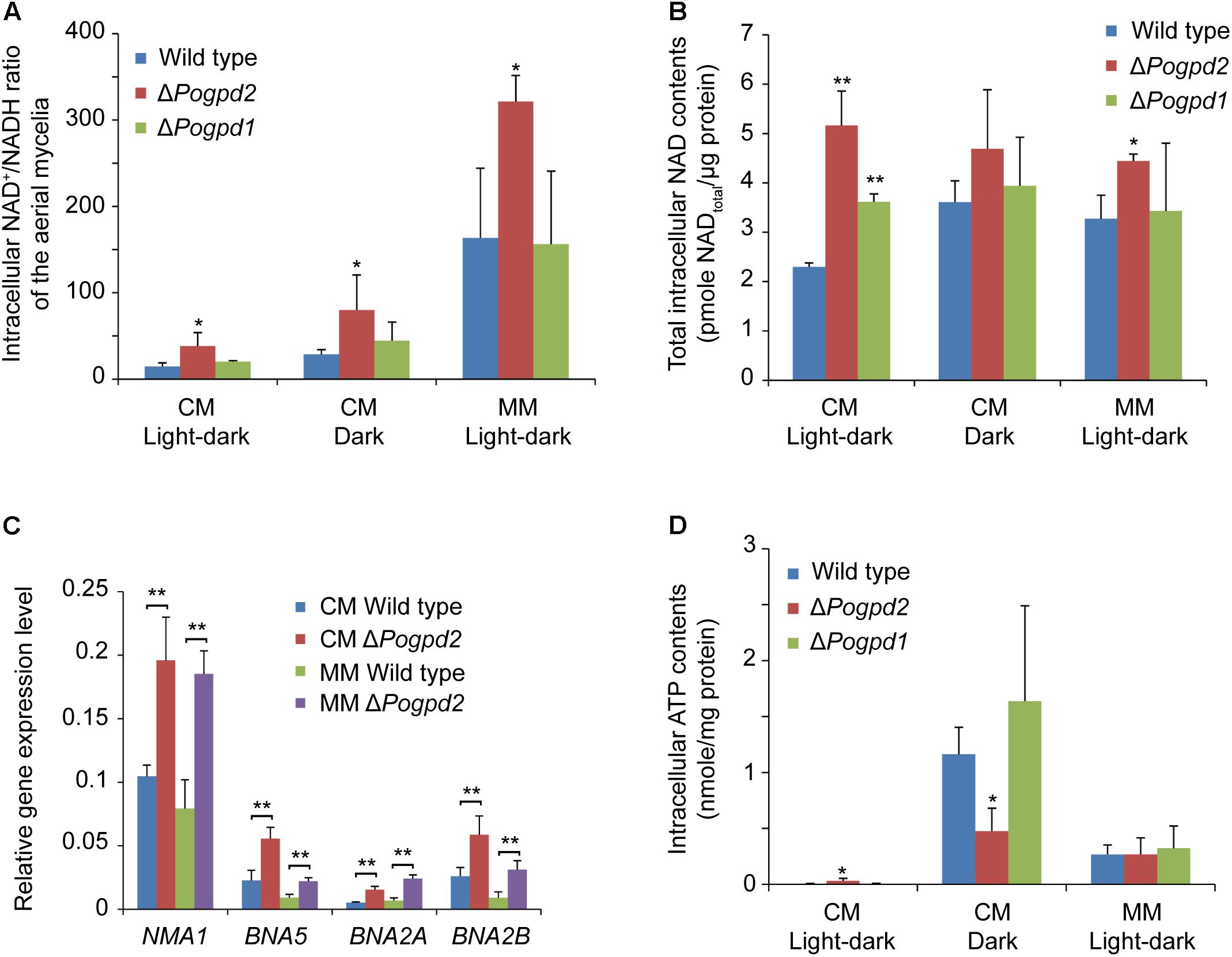
FIGURE 9. Roles of glycerol-3-phosphate dehydrogenases in NAD+/NADH ratio and intracellular ATP content in Pyricularia oryzae. (A) Intracellular NAD+/NADH ratio in the aerial mycelia of the wild type, ΔPogpd1, and ΔPogpd2 cultured on CM and MM media under a light–dark cycle or a continuous dark condition. (B) Total intracellular NAD content (pmol/μg protein) in the aerial mycelia of the wild type, ΔPogpd1, and ΔPogpd2 cultured on CM and MM media under a light–dark cycle or a continuous dark condition. (C) Relative expression level of four NAD biosynthesis genes (NMA1, BNA5, BNA2A, and BNA2B) in ΔPogpd2 cultured on CM and MM media under a light–dark cycle. β-TUBULIN and H3 were selected as reference genes. (D) Intracellular ATP content in the aerial mycelia of the wild type, ΔPogpd1, and ΔPogpd2 cultured on CM and MM media under a light–dark cycle or a continuous dark condition. Error bars represent SD. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗P < 0.05 and ∗∗P < 0.01.
To clarify the role of PoGpd2 in transcription of NAD synthesis genes, we measured the expression level of four genes (NMA1, BNA5, BNA2A, and BNA2B) involved in NAD biosynthesis. NMA1 (MGG_01290) encodes a nicotinic acid mononucleotide adenylyltransferase, BNA5 (MGG_10969) encodes a kynureninase, and BNA2A (MGG_13773) and BNA2B (MGG_14348) encodes two homologs of Bna2 (Tryptophan 2,3-dioxygenase or indoleamine 2,3-dioxygenase) in P. oryzae. NMA1, BNA5, BNA2A and BNA2B were significantly up-regulated in the aerial mycelia of ΔPogpd2 on CM medium (1.9-, 2.5-, 3.0-, and 2.3-fold, respectively) and MM medium (2.4-, 2.5-, 3.6-, and 3.7-fold, respectively) under a light–dark cycle, relative to the wild type (Figure 9C).
We then quantified the intracellular ATP content in the aerial mycelia cultured on CM and MM media. On CM medium, P. oryzae strains had much lower ATP levels when cultured under a light–dark cycle than those under continuous dark (for the wild type, ΔPogpd2, and ΔPogpd1: 3.5 ± 5.1, 32.0 ± 23.1, 5.0 ± 5.0 pmol/mg protein in light/dark vs. 1164.1 ± 241.4, 475.3 ± 206.0, 1638.6 ± 851.9 pmol/mg protein in dark, respectively). The intracellular ATP content in ΔPogpd2 was higher than that in the wild type or in ΔPogpd1 when cultured on CM medium under a light–dark cycle, whereas it is was lower than the wild type or ΔPogpd1 under dark (Figure 9D).
Glycerol-3-Phosphate Shuttle Is Involved in Virulence in P. oryzae
Virulence of the mutants on barley and rice was tested to assess the function of G-3-P shuttle on pathogenicity. As ΔPogpd2 produced very few conidia, we first tested its hyphal virulence using an excised-leaf inoculation technique. In cut leaf assays, both barley and rice leaves inoculated with mycelial plugs for 4 days showed severe blast lesions from the wild type and PoGPD2-rescued strains, whereas ΔPogpd2 and ΔPogpd1ΔPogpd2 caused very mild disease lesions (Figure 10A). We then confirmed the virulence of ΔPogpd1 on rice by spraying a conidial suspension (1 × 105 spores/ml) on rice seedlings, and found that the virulence of spores was greatly reduced in ΔPogpd1 (Figures 10B,C). When sprayed on rice, the wild type and the PoGPD1-rescued strain caused gray-centered, sporulating, and coalesced lesions on leaves, whereas ΔPogpd1 caused only small and isolated lesions (Figure 10B). Average percentage (± standard deviation) of lesion areas in 5-cm length sections of rice leaves was 14.9 ± 4.9 for ΔPogpd1, 38.1 ± 3.6 for the wild type, and 36.5 ± 3.8 for the PoGPD1-rescued strain (P < 0.01) (Figure 10C). Therefore, PoGPD2 and PoGPD1 are required for fungal hyphal or conidial virulence on rice, suggesting that the glycerol-3-phosphate shuttle is involved in the pathogenesis in P. oryzae.
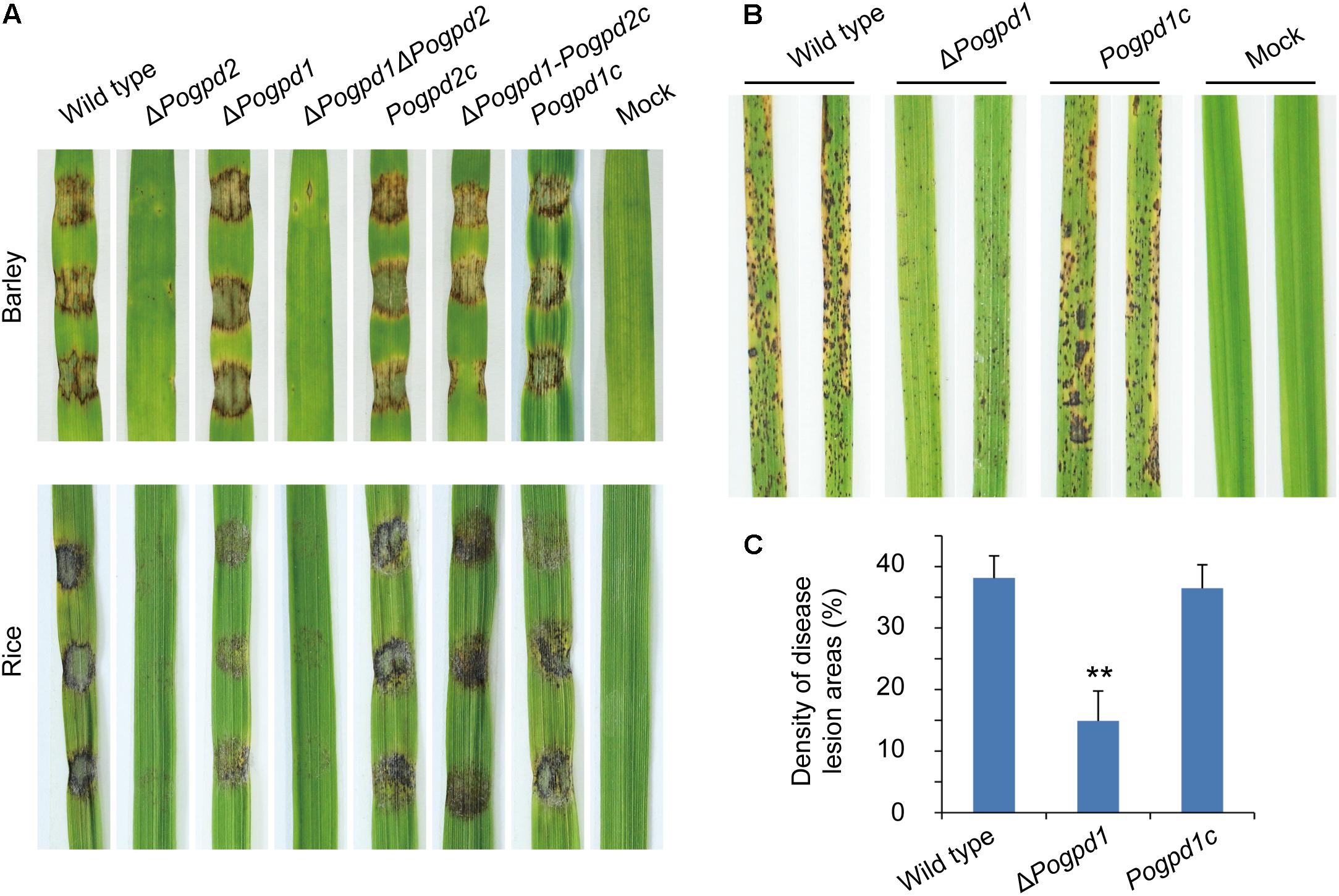
FIGURE 10. Roles of glycerol-3-phosphate dehydrogenases on fungal virulence in Pyricularia oryzae. (A) Pathogenicity assays on excised-leaf of rice and barley inoculated with mycelial plugs. (B) Pathogenicity assays on rice seedling sprayed with conidial suspension. (C) Disease lesion severity assays for tests in (B). Proportion of lesion areas in 5-cm-length leaves (%) was counted. Error bars represent SD. Significant difference compared with the wild type as estimated by Tukey’s HSD test: ∗∗P < 0.01.
Discussion
The glycerol-3-phosphate shuttle is a pathway that translocates electrons produced during glycolysis across the inner membrane of the mitochondrion for oxidative phosphorylation by oxidizing cytoplasmic NADH to NAD+. Gpd1 and Gpd2 in the G-3-P shuttle are important enzymes for the production and utilization of glycerol in yeasts and other organisms (Ronnow and Kielland-Brandt, 1993; Albertyn et al., 1994). We found that G-3-P shuttle is required for aerial hyphal differentiation, conidiation, and pathogenicity in P. oryzae.
In S. cerevisiae, a cytoplasmic glycerol-3-phosphate dehydrogenase Gpd1 is required for glycerol synthesis and resistance to osmotic stress (Albertyn et al., 1994). However, in P. oryzae, PoGPD1 is not involved in fungal resistance to hyperosmotic stresses caused by both salt and sugar (Figure 2). This discrepancy in osmotic stress response may be due to the fact that accumulated glycerol is the primary compatible solute as a response to high-osmolarity in yeast (Albertyn et al., 1994), while in P. oryzae (strain Guy11, (Chao and Ellingboe, 1991)) arabitol is used (Dixon et al., 1999). In P. oryzae appressoria, glycerol is a primary compatible solute to generate turgor (deJong et al., 1997). ΔPogpd1 displays similar appressorium turgor as that in the wild type 70-15 (Supplementary Figure S3), suggesting that PoGPD1 is not required for the glycerol synthesis in appressoria. The expression of FAD-GPDH (GPD2) in A. thaliana is highly coupled with glycerol catabolism in germinated seed (Shen et al., 2003). And Gut2, a mitochondrial glycerol-3-phosphate dehydrogenase in yeast, is a key enzyme to utilize glycerol (Ronnow and Kielland-Brandt, 1993). ΔPogpd2 could not efficiently utilize glycerol as a carbon source and accumulated G-3-P in cells (Figure 3), suggesting the involvement of PoGPD2 in glycerol utilization in P. oryzae.
Nicotinamide adenine dinucleotides (NAD+ and NADH) are key regulators of cellular redox state. The G-3-P shuttle transfers cytosolic reducing equivalents into mitochondria and maintains a balanced NAD+/NADH ratio in eukaryotic cells (Larsson et al., 1998). In P. oryzae, the deletion of PoGPD2, but not PoGPD1, led to an increase in the intracellular NAD+/NADH ratio and to alteration of intracellular ATP content (Figure 9). In A. thaliana, loss of GPDHc1 (GPD1) decreased the cytoplasmic NAD+/NADH ratio (Shen et al., 2006). The gpdhc1 mutants displayed elevated intracellular ROS levels and a correlated modified ratio of metabolites involved in redox exchange between the mitochondria and cytosol (Shen et al., 2006). The muscle tissue of mice lacking Gdc-1 (Gpd1) showed a lowered lactate/pyruvate ratio which signified a lowered NAD+/NADH ratio. When exercised, these knockout mice were unable to maintain normal ATP levels in skeletal muscle (MacDonald and Marshall, 2000). Discrepancies in alteration of the intracellular NAD+/NADH ratio after destruction of GPD1 or GPD2 appear between P. oryzae and other organisms, which could originate from the diversity of metabolism pathways among different organisms.
ΔPogpd2 and ΔPogpd1ΔPogpd2 are sensitive to light (Figure 5). In mammals, circadian rhythms are controlled by two pairs of heterodimeric TF (Clock:BMAL1 and NPAS2:BMAL1) whose activity fluctuates in response to the light–dark cycle (Gekakis et al., 1998). DNA binding of the Clock:BMAL1 and NPAS2:BMAL1 heterodimers is strongly enhanced by NADH, whereas it is inhibited by NAD+ (Rutter et al., 2001), suggesting the importance of the NAD+/NADH ratio during regulation of the light–dark cycle. Although homology of Clock, Bmal1, and Npas2 had not been identified in P. oryzae, several light-sensing proteins had been characterized, such as Trx2 and PoSir2 (Fernandez et al., 2014a; Zhang et al., 2016). Trx2 is a thioredoxin and is required for sulfite assimilation, growth, asexual and sexual differentiation, scavenging of ROS during host cell invasion, invasive hyphal growth, and pathogenicity in P. oryzae (Wang et al., 2016; Zhang et al., 2016). As conidiation is induced by light in P. oryzae, the reduced conidiation in Δtrx2 was considered to be related to the role of Trx2 in light sensing (Zhang et al., 2016). Deletion of TRX2 led to reduced expression of CON7, which is a transcription factor required for conidium and appressorium differentiation, and of COM1, which is a transcription factor important for conidiophore differentiation (Zhang et al., 2016). In yeasts and in animals, sirtuins (NAD+-dependent deacetylases) function in metabolic and nutrient regulation, transcription regulation, and oxidative stress by serving as energy sensors via the sensitivity of their catalytic activity to the metabolite NAD+, and as transcriptional effectors by controlling the acetylation state of histones and transcriptional regulators (Houtkooper et al., 2012; Wierman and Smith, 2014). In S. cerevisiae, Sir2 is a NAD+-dependent histone deacetylase functioning in transcriptional silencing (Wierman and Smith, 2014). An increased NAD+/NADH ratio is considered to stimulate Sir2 activity (Lin et al., 2004; Wierman and Smith, 2014). In P. oryzae, PoSir2 is a fungal sirtuin required for biotrophic growth (Fernandez et al., 2014a). During the early stages of in planta growth, PoSir2 deacetylates a cupin-like JmjC domain-containing protein PoJmjC, which then alleviates PoSOD1 transcript repression, and PoSod1 detoxifies host-derived ROS to the benefit of the fungus (Fernandez et al., 2014a). In ΔPogpd2 grown on CM medium, two light-sensing protein genes TRX2 and PoSIR2 were up-regulated under a light–dark cycle, but not under continuous dark. Light affected conidiation, and the differentiation and death of aerial hyphae in ΔPogpd2. Therefore, light, along with the NAD+/NADH ratio (cellular redox state), affects fungal development and pathogenicity, possibly through light-sensing proteins, such as Trx2 and PoSir2 in ΔPogpd2.
The role of light on the aerial hypha differentiation of ΔPogpd2 is reliant on medium components. On MM medium, ΔPogpd2 was not sensitive to light and its aerial hyphae differentiated like those in the wild type (Figure 5A). None of light-sensitive protein genes were up-regulated in ΔPogpd2 grown on MM medium under a light–dark cycle (Figure 6C). NAD+/NADH ratio of P. oryzae strains grown on MM medium was higher than those on CM medium (Figure 9). In MM medium, NO3- is a sole nitrogen source. NO3- is reduced to NH4+, and NH4+ is incorporated into glutamate and glutamine via glutamate metabolism (Figure 7A). Glutamate metabolism is an important pathway in the rice blast fungus (Marroquin-Guzman and Wilson, 2015; Zhou et al., 2017). MGD1, which encodes a dehydrogenase, is required for appressorium formation and virulence. Δmgd1 lacks aerial hyphae when grown on CM media (Oh et al., 2008). PoGLT1, a glutamate synthase gene, is required for autophagy, conidiation and virulence (Zhou et al., 2017). In the biochemical reactions catalyzed by Nia1, Gdh1 and Glt1, but not by Nii1, Mgd1 and Gln1/Gln2, NADH was oxidized to NAD+ (Figure 7A). Interestingly, the expression levels of NIA1, GDH1 and PoGLT1, but not NII1, MGD1 and GLN1/GLN2, were up-regulated in ΔPogpd2 grown on MM medium under a light–dark cycle (Figure 7B). On CM medium in which NO3-, yeast extract and casamino acid are mixed nitrogen sources, GDH1 and/or PoGLT1 were also up-regulated in ΔPogpd2 under a light–dark cycle or a continuous dark (Figure 7B). However, none of GDH1 and PoGLT1 were up-regulated significantly in ΔPogpd1 (Supplementary Figure S4). Therefore, the synthesis of glutamate by Gdh1 and PoGlt1 in ΔPogpd2 seems have a role in promoting the elevated NAD+/NADH ratio in ΔPogpd2 on CM and MM media.
ΔPogpd2 displayed defects in aerial hyphae differentiation and conidiation. Fungal hydrophobins play important roles in aerial hyphae formation and spore production in fungi (Elliot and Talbot, 2004). When aerial hyphae erect from the aqueous-air interface, they secrete nomomers of hydrophobins which aggregate spontaneously to create a hydrophobic sheath to overcome this barrier. The protective hydrophobic coating on the surface of the hyphae also prevents hyphal dehydration (Elliot and Talbot, 2004). Δmpg1, a mutant in which a hydrophobin gene MGP1 was deleted, does not sporulate efficiently and shows reduced virulence (Talbot et al., 1993, 1996; Beckerman and Ebbole, 1996). In ΔPogpd2, the surface hydrophobicity of aerial mycelia against water and a detergent solution was lower than that of the wild type. This phenomenon was supported by the significantly down-regulated expression of three hydrophobin genes (MPG1, MGG_09134, and MGG_10105) in the mutant. Melanin consists of an important layer of cell wall in aerial hyphae, conidia, and appressoria, which protects fungal cells against environmental stresses and is required for fungal virulence. 4HNR, AIB1, RSY1, and BUF1 are four melanin synthesis genes involved in appressorium formation and pathogenicity in the rice blast fungus (Chumley and Valent, 1990; Thompson et al., 2000). 4HNR, AIB1, RSY1, and BUF1 were significantly down-regulated in ΔPogpd2, and these are consistent with the white aerial hyphae in the mutant. Therefore, the phenotype of tardily differentiated and early collapsed aerial hyphae in ΔPogpd2 cultured on CM medium under a light–dark cycle could be caused, at least partly, by down-regulation of hydrophobin and melanin synthesis genes.
Several TF to regulate aerial hyphal differentiation and conidiation have been previously identified. The deletion of transcription factor genes COS1, CONx2, GCC1, and CON7 led to loss or nearly loss of ability to produce spores in P. oryaze (Zhou et al., 2009; Li et al., 2013; Lu et al., 2014; Cao et al., 2016). COS1 and CONx2 are involved in conidiophore differentiation (Zhou et al., 2009; Cao et al., 2016), CON7 in conidial morphology (Odenbach et al., 2007; Cao et al., 2016), and GCC1 in conidial differentiation (Lu et al., 2014). CON7 and CONx2 are also required for fungal virulence (Odenbach et al., 2007; Cao et al., 2016), and Δcos1, Δconx2, Δcon7, and Δgcc1 showed defects in melanin synthesis (Zhou et al., 2009; Lu et al., 2014; Cao et al., 2016). RNA-seq revealed that 4HNR, AIB1, and BUF1 were down-regulated in Δcos1 (Li et al., 2013). Four TF genes (COS1, CONx2, CON7, and GCC1) were down-regulated in the aerial mycelia of ΔPogpd2. ΔPogpd1 also produced fewer conidia, and expressed a lower level of four melanin synthesis genes (4HNR, AIB1, RSY1, BUF1) and of two TF genes (COS1 and CONx2) in the aerial mycelia of the mutant. The G-3-P shuttle may influence fungal development through altering the expression of transcription factor genes, and subsequently of hydrophobin genes and melanin synthesis genes.
Melanin is not only involved in hyphal differentiation and conidiation, but also required for appressorial turgor, plant penetration, and virulence in P. oryzae (Chumley and Valent, 1990; Thompson et al., 2000). Hydrophobins are also involved in hyphal differentiation, conidiation, appressorium formation, and virulence (Talbot et al., 1993, 1996; Beckerman and Ebbole, 1996). The reduced virulence of mycelium or conidia on rice caused by deletion of PoGPD1 or PoGPD2 is possibly, at least partly, owing to decreased melanin and hydrophobin synthesis in P. oryzae. ΔPogpd1’s conidia showed reduced virulence on rice, but its aerial mycelia still displayed strong virulence on barley (Figure 10). There are two possible causes responsible for these differences in virulence: barley and rice have different proteins and other components participated in innate immunity (PTI/PAMP-triggered immunity and ETI/effector-triggered immunity); hyphae and conidia have different regulative mechanisms in appressorium formation (Kong et al., 2013). In our previous works, we also found that Δcca1’s hyphae could but its conidia could not infect rice (Lu et al., 2014), and ΔPocapn7’s conidia strongly but its hyphae weakly infected rice (Liu et al., 2016).
In summary, the G-3-P shuttle is involved in cellular redox state, development, and virulence of P. oryzae. PoGpd1 is required for utilization of several carbon sources (pyruvate, sodium acetate, glutamate, and glutamine), NO production, as well as conidiation in P. oryzae, whereas PoGpd2 is required for maintenance of intracellular NAD+/NADH ratio, ATP production, glycerol utilization, light sensing, aerial hyphal differentiation, and conidiation.
Author Contributions
JL and YS contributed to experimental design. YS, HW, YY, HC, JL, and XL contributed to experiments. YS, HW, YY, and JL contributed to data analysis and scripts. FL and JL supplied experimental conditions. YS, JL, HW, and FL wrote the manuscript.
Funding
This research was supported by National Natural Science Foundation of China (Grant Nos: 31671975 and 31371891) and National Basic Research Program of China (Grant No: 2014CB541702).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2018.00687/full#supplementary-material
Footnotes
References
Albertyn, J., Hohmann, S., Thevelein, J. M., and Prior, B. A. (1994). GPD1, which encodes glycerol-3-phosphate dehydrogenase, is essential for growth under osmotic stress in Saccharomyces cerevisiae, and its expression is regulated by the high-osmolarity glycerol response pathway. Mol. Cell. Biol. 14, 4135–4144. doi: 10.1128/MCB.14.6.4135
Anderson, R. M., Bitterman, K. J., Wood, J. G., Medvedik, O., Cohen, H., Lin, S. S., et al. (2002). Manipulation of a nuclear NAD+ salvage pathway delays aging without altering steady-state NAD+ levels. J. Biol. Chem. 277, 18881–18890. doi: 10.1074/jbc.M111773200
Anderson, R. M., Bitterman, K. J., Wood, J. G., Medvedik, O., and Sinclair, D. A. (2003). Nicotinamide and PNC1 govern lifespan extension by calorie restriction in Saccharomyces cerevisiae. Nature 423, 181–185. doi: 10.1038/nature01578
Ansell, R., Granath, K., Hohmann, S., Thevelein, J. M., and Adler, L. (1997). The two isoenzymes for yeast NAD+-dependent glycerol 3-phosphate dehydrogenase encoded by GPD1 and GPD2 have distinct roles in osmoadaptation and redox regulation. EMBO J. 16, 2179–2187. doi: 10.1093/emboj/16.9.2179
Beckerman, J. L., and Ebbole, D. J. (1996). MPG1, a gene encoding a fungal hydrophobin of Magnaporthe grisea, is involved in surface recognition. Mol. Plant Microbe Interact. 9, 450–456. doi: 10.1094/Mpmi-9-0450
Bhadauria, V., Banniza, S., Vandenberg, A., Selvaraj, G., and Wei, Y. (2012). Peroxisomal alanine: glyoxylate aminotransferase AGT1 is indispensable for appressorium function of the rice blast pathogen, Magnaporthe oryzae. PLoS One 7:e36266. doi: 10.1371/journal.pone.0036266
Bjorkqvist, S., Ansell, R., Adler, L., and Liden, G. (1997). Physiological response to anaerobicity of glycerol-3-phosphate dehydrogenase mutants of Saccharomyces cerevisiae. Appl. Environ. Microbiol. 63, 128–132.
Bonman, J. M., Dedios, T. I. V., and Khin, M. M. (1986). Physiological specialization of Pyricularia oryzae in the philippines. Plant Dis. 70, 767–769. doi: 10.1094/PD-70-767
Brown, L. J., Koza, R. A., Marshall, L., Kozak, L. P., and MacDonald, M. J. (2002). Lethal hypoglycemic ketosis and glyceroluria in mice lacking both the mitochondrial and the cytosolic glycerol phosphate dehydrogenases. J. Biol. Chem. 277, 32899–32904. doi: 10.1074/jbc.M202409200
Bus, J. S., and Gibson, J. E. (1984). Paraquat: model for oxidant-initiated toxicity. Environ. Health Perspect. 55, 37–46. doi: 10.1289/ehp.845537
Cao, H., Huang, P., Zhang, L., Shi, Y., Sun, D., Yan, Y., et al. (2016). Characterization of 47 Cys2-His2 zinc finger proteins required for the development and pathogenicity of the rice blast fungus Magnaporthe oryzae. New Phytol. 211, 1035–1051. doi: 10.1111/nph.13948
Chao, C., and Ellingboe, A. H. (1991). Selection for mating competence in Magnaporthe grisea pathogenic to rice. Can. J. Bot. 69, 2130–2134. doi: 10.1139/b91-267
Chumley, F. G., and Valent, B. (1990). Genetic-analysis of melanin-deficient, nonpathogenic mutants of Magnaporthe grisea. Mol. Plant Microbe Interact. 3, 135–143. doi: 10.1094/Mpmi-3-135
Cole, E. S., Lepp, C. A., Holohan, P. D., and Fondy, T. P. (1978). Isolation and characterization of flavin-linked glycerol-3-phosphate dehydrogenase from rabbit skeletal muscle mitochondria and comparison with the enzyme from rabbit brain. J. Biol. Chem. 253, 7952–7959.
de Groot, M. J., Bundock, P., Hooykaas, P. J., and Beijersbergen, A. G. (1998). Agrobacterium tumefaciens-mediated transformation of flamentous fungi. Nat. Biotechnol. 16, 839–842. doi: 10.1038/nbt0998-839
Dean, R., Van Kan, J. A., Pretorius, Z. A., Hammond-Kosack, K. E., Di Pietro, A., Spanu, P. D., et al. (2012). The Top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414–430. doi: 10.1111/j.1364-3703.2011.00783.x
deJong, J. C., MCCormack, B. J., Smirnoff, N., and Talbot, N. J. (1997). Glycerol generates turgor in rice blast. Nature 389, 244–245. doi: 10.1038/38418
Deng, Y. Z., Qu, Z., and Naqvi, N. I. (2015). Twilight, a novel circadian-regulated gene, integrates phototropism with nutrient and redox homeostasis during fungal development. PLoS Pathog. 11:e1004972. doi: 10.1371/journal.ppat.1004972
Ding, S. L., Liu, W., Iliuk, A., Ribot, C., Vallet, J., Tao, A., et al. (2010). The tig1 histone deacetylase complex regulates infectious growth in the rice blast fungus Magnaporthe oryzae. Plant Cell 22, 2495–2508. doi: 10.1105/tpc.110.074302
Dixon, K. P., Xu, J. R., Smirnoff, N., and Talbot, N. J. (1999). Independent signaling pathways regulate cellular turgor during hyperosmotic stress and appressorium-mediated plant infection by Magnaporthe grisea. Plant Cell 11, 2045–2058. doi: 10.1105/tpc.11.10.2045
Elliot, M. A., and Talbot, N. J. (2004). Building filaments in the air: aerial morphogenesis in bacteria and fungi. Curr. Opin. Microbiol. 7, 594–601. doi: 10.1016/j.mib.2004.10.013
Estabrook, R. W., and Sacktor, B. (1958). alpha-Glycerophosphate oxidase of flight muscle mitochondria. J. Biol. Chem. 233, 1014–1019.
Fernandez, J., Marroquin-Guzman, M., Nandakumar, R., Shijo, S., Cornwell, K. M., Li, G., et al. (2014a). Plant defence suppression is mediated by a fungal sirtuin during rice infection by Magnaporthe oryzae. Mol. Microbiol. 94, 70–88. doi: 10.1111/mmi.12743
Fernandez, J., Marroquin-Guzman, M., and Wilson, R. A. (2014b). Evidence for a transketolase-mediated metabolic checkpoint governing biotrophic growth in rice cells by the blast fungus Magnaporthe oryzae. PLoS Pathog. 10:e1004354. doi: 10.1371/journal.ppat.1004354
Gee, R. W., Byerrum, R. U., Gerber, D. W., and Tolbert, N. E. (1988). Dihydroxyacetone phosphate reductase in plants. Plant Physiol. 86, 98–103. doi: 10.1104/pp.86.1.98
Gekakis, N., Staknis, D., Nguyen, H. B., Davis, F. C., Wilsbacher, L. D., King, D. P., et al. (1998). Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569. doi: 10.1126/science.280.5369.1564
Guo, M., Chen, Y., Du, Y., Dong, Y., Guo, W., Zhai, S., et al. (2011). The bZIP transcription factor MoAP1 mediates the oxidative stress response and is critical for pathogenicity of the rice blast fungus Magnaporthe oryzae. PLoS Pathog. 7:e1001302. doi: 10.1371/journal.ppat.1001302
Houtkooper, R. H., Pirinen, E., and Auwerx, J. (2012). Sirtuins as regulators of metabolism and healthspan. Nat. Rev. Mol. Cell Biol. 13, 225–238. doi: 10.1038/nrm3293
Howard, R. J., Ferrari, M. A., Roach, D. H., and Money, N. P. (1991). Penetration of hard substrates by a fungus employing enormous turgor pressures. Proc. Natl. Acad. Sci. U.S.A. 88, 11281–11284. doi: 10.1073/pnas.88.24.11281
Jung, S., Marelli, M., Rachubinski, R. A., Goodlett, D. R., and Aitchison, J. D. (2010). Dynamic changes in the subcellular distribution of Gpd1p in response to cell stress. J. Biol. Chem. 285, 6739–6749. doi: 10.1074/jbc.M109.058552
Kato, M., and Lin, S. J. (2014). Regulation of NAD+ metabolism, signaling and compartmentalization in the yeast Saccharomyces cerevisiae. DNA Repair 23, 49–58. doi: 10.1016/j.dnarep.2014.07.009
Kim, S., Ahn, I. P., Rho, H. S., and Lee, Y. H. (2005). MHP1, a Magnaporthe grisea hydrophobin gene, is required for fungal development and plant colonization. Mol. Microbiol. 57, 1224–1237. doi: 10.1111/j.1365-2958.2005.04750.x
Kim, S., Park, S. Y., Kim, K. S., Rho, H. S., Chi, M. H., Choi, J., et al. (2009). Homeobox transcription factors are required for conidiation and appressorium development in the rice blast fungus Magnaporthe oryzae. PLoS Genet. 5:e1000757. doi: 10.1371/journal.pgen.1000757
Klingenberg, M. (1970). Localization of the glycerol-phosphate dehydrogenase in the outer phase of the mitochondrial inner membrane. Eur. J. Biochem. 13, 247–252. doi: 10.1111/j.1432-1033.1970.tb00924.x
Kong, L. A., Li, G. T., Liu, Y., Liu, M. G., Zhang, S. J., Yang, J., et al. (2013). Differences between appressoria formed by germ tubes and appressorium-like structures developed by hyphal tips in Magnaporthe oryzae. Fungal Genet. Biol. 56, 33–41. doi: 10.1016/j.fgb.2013.03.006
Koza, R. A., Kozak, U. C., Brown, L. J., Leiter, E. H., MacDonald, M. J., and Kozak, L. P. (1996). Sequence and tissue-dependent RNA expression of mouse FAD-linked glycerol-3-phosphate dehydrogenase. Arch. Biochem. Biophys. 336, 97–104. doi: 10.1006/abbi.1996.0536
Larsson, C., Pahlman, I. L., Ansell, R., Rigoulet, M., Adler, L., and Gustafsson, L. (1998). The importance of the glycerol 3-phosphate shuttle during aerobic growth of Saccharomyces cerevisiae. Yeast 14, 347–357. doi: 10.1002/(SICI)1097-0061(19980315)14:4<347::AID-YEA226>3.0.CO;2-9
Lee, K., Singh, P., Chung, W. C., Ash, J., Kim, T. S., Hang, L., et al. (2006). Light regulation of asexual development in the rice blast fungus, Magnaporthe oryzae. Fungal Genet. Biol. 43, 694–706. doi: 10.1016/j.fgb.2006.04.005
Lee, Y. P., and Lardy, H. A. (1965). Influence of thyroid hormones on l-alpha-glycerophosphate dehydrogenases and other dehydrogenases in various organs of the rat. J. Biol. Chem. 240, 1427–1436.
Li, H. J., Lu, J. P., Liu, X. H., Zhang, L. L., and Lin, F. C. (2012). Vectors building and usage for gene knockout, protein expression and fluorescent fusion protein in the rice blast fungus. J. Agric. Biotechnol. 20, 94–104.
Li, X., Han, X., Liu, Z., and He, C. (2013). The function and properties of the transcriptional regulator COS1 in Magnaporthe oryzae. Fungal Biol. 117, 239–249. doi: 10.1016/j.funbio.2013.01.010
Li, Y., Liang, S., Yan, X., Wang, H., Li, D., Soanes, D. M., et al. (2010). Characterization of MoLDB1 required for vegetative growth, infection-related morphogenesis, and pathogenicity in the rice blast fungus Magnaporthe oryzae. Mol. Plant Microbe Interact. 23, 1260–1274. doi: 10.1094/MPMI-03-10-0052
Li, Y., Zhu, J., Hu, J., Meng, X., Zhang, Q., Zhu, K., et al. (2016). Functional characterization of electron-transferring flavoprotein and its dehydrogenase required for fungal development and plant infection by the rice blast fungus. Sci. Rep. 6:24911. doi: 10.1038/srep24911
Lin, S. J., Ford, E., Haigis, M., Liszt, G., and Guarente, L. (2004). Calorie restriction extends yeast life span by lowering the level of NADH. Genes Dev. 18, 12–16. doi: 10.1101/gad.1164804
Lin, S. J., and Guarente, L. (2003). Nicotinamide adenine dinucleotide, a metabolic regulator of transcription, longevity and disease. Curr. Opin. Cell Biol. 15, 241–246. doi: 10.1016/S0955-0674(03)00006-1
Liu, X. H., Ning, G. A., Huang, L. Y., Zhao, Y. H., Dong, B., Lu, J. P., et al. (2016). Calpains are involved in asexual and sexual development, cell wall integrity and pathogenicity of the rice blast fungus. Sci. Rep. 6:31204. doi: 10.1038/srep31204
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Lu, J., Cao, H., Zhang, L., Huang, P., and Lin, F. (2014). Systematic analysis of Zn2Cys6 transcription factors required for development and pathogenicity by high-throughput gene knockout in the rice blast fungus. PLoS Pathog. 10:e1004432. doi: 10.1371/journal.ppat.1004432
Lu, J. P., Feng, X. X., Liu, X. H., Lu, Q., Wang, H. K., and Lin, F. C. (2007). Mnh6, a nonhistone protein, is required for fungal development and pathogenicity of Magnaporthe grisea. Fungal Genet. Biol. 44, 819–829. doi: 10.1016/j.fgb.2007.06.003
MacDonald, M. J., and Marshall, L. K. (2000). Mouse lacking NAD+-linked glycerol phosphate dehydrogenase has normal pancreatic beta cell function but abnormal metabolite pattern in skeletal muscle. Arch. Biochem. Biophys. 384, 143–153. doi: 10.1006/abbi.2000.2107
Marroquin-Guzman, M., and Wilson, R. A. (2015). GATA-dependent glutaminolysis drives appressorium formation in Magnaporthe oryzae by suppressing tor inhibition of cAMP/PKA signaling. PLoS Pathog. 11:e1004851. doi: 10.1371/journal.ppat.1004851
Menaya, J., Gonzalez-Manchon, C., Parrilla, R., and Ayuso, M. S. (1995). Molecular cloning, sequencing and expression of a cDNA encoding a human liver NAD-dependent alpha-glycerol-3-phosphate dehydrogenase. Biochim. Biophys. Acta 1262, 91–94. doi: 10.1016/0167-4781(95)00069-S
Mohanan, V. C., Chandarana, P. M., Chattoo, B. B., Patkar, R. N., and Manjrekar, J. (2017). Fungal histidine phosphotransferase plays a crucial role in photomorphogenesis and pathogenesis in Magnaporthe oryzae. Front. Chem. 5:31. doi: 10.3389/fchem.2017.00031
Mracek, T., Drahota, Z., and Houstek, J. (2013). The function and the role of the mitochondrial glycerol-3-phosphate dehydrogenase in mammalian tissues. Biochim. Biophys. Acta 1827, 401–410. doi: 10.1016/j.bbabio.2012.11.014
Mracek, T., Pecinova, A., Vrbacky, M., Drahota, Z., and Houstek, J. (2009). High efficiency of ROS production by glycerophosphate dehydrogenase in mammalian mitochondria. Arch. Biochem. Biophys. 481, 30–36. doi: 10.1016/j.abb.2008.10.011
Odenbach, D., Breth, B., Thines, E., Weber, R. W., Anke, H., and Foster, A. J. (2007). The transcription factor Con7p is a central regulator of infection-related morphogenesis in the rice blast fungus Magnaporthe grisea. Mol. Microbiol. 64, 293–307. doi: 10.1111/j.1365-2958.2007.05643.x
Oh, Y., Donofrio, N., Pan, H., Coughlan, S., Brown, D. E., Meng, S., et al. (2008). Transcriptome analysis reveals new insight into appressorium formation and function in the rice blast fungus Magnaporthe oryzae. Genome Biol. 9:R85. doi: 10.1186/gb-2008-9-5-r85
Orr, A. L., Quinlan, C. L., Perevoshchikova, I. V., and Brand, M. D. (2012). A refined analysis of superoxide production by mitochondrial sn-glycerol 3-phosphate dehydrogenase. J. Biol. Chem. 287, 42921–42935. doi: 10.1074/jbc.M112.397828
Ou, X., Ji, C., Han, X., Zhao, X., Li, X., Mao, Y., et al. (2006). Crystal structures of human glycerol 3-phosphate dehydrogenase 1 (GPD1). J. Mol. Biol. 357, 858–869. doi: 10.1016/j.jmb.2005.12.074
Rho, H. S., Kang, S., and Lee, Y. H. (2001). Agrobacterium tumefaciens-mediated transformation of the plant pathogenic fungus, Magnaporthe grisea. Mol. Cells 12, 407–411.
Rigoulet, M., Aguilaniu, H., Averet, N., Bunoust, O., Camougrand, N., Grandier-Vazeille, X., et al. (2004). Organization and regulation of the cytosolic NADH metabolism in the yeast Saccharomyces cerevisiae. Mol. Cell. Biochem. 256, 73–81. doi: 10.1023/B:MCBI.0000009888.79484.fd
Ronnow, B., and Kielland-Brandt, M. C. (1993). GUT2, a gene for mitochondrial glycerol 3-phosphate dehydrogenase of Saccharomyces cerevisiae. Yeast 9, 1121–1130. doi: 10.1002/yea.320091013
Rutter, J., Reick, M., Wu, L. C., and McKnight, S. L. (2001). Regulation of clock and NPAS2 DNA binding by the redox state of NAD cofactors. Science 293, 510–514. doi: 10.1126/science.1060698
Samalova, M., Johnson, J., Illes, M., Kelly, S., Fricker, M., and Gurr, S. (2012). Nitric oxide generated by the rice blast fungus Magnaporthe oryzae drives plant infection. New Phytol. 197, 207–222. doi: 10.1111/j.1469-8137.2012.04368.x
Sato, T., Morita, A., Mori, N., and Miura, S. (2014). The role of glycerol-3-phosphate dehydrogenase 1 in the progression of fatty liver after acute ethanol administration in mice. Biochem. Biophys. Res. Commun. 444, 525–530. doi: 10.1016/j.bbrc.2014.01.096
Shen, W., Wei, Y., Dauk, M., Tan, Y., Taylor, D. C., Selvaraj, G., et al. (2006). Involvement of a glycerol-3-phosphate dehydrogenase in modulating the NADH/NAD+ ratio provides evidence of a mitochondrial glycerol-3-phosphate shuttle in Arabidopsis. Plant Cell 18, 422–441. doi: 10.1105/tpc.105.039750
Shen, W., Wei, Y., Dauk, M., Zheng, Z., and Zou, J. (2003). Identification of a mitochondrial glycerol-3-phosphate dehydrogenase from Arabidopsis thaliana: evidence for a mitochondrial glycerol-3-phosphate shuttle in plants. FEBS Lett. 536, 92–96. doi: 10.1016/S0014-5793(03)00033-4
Sun, D., Cao, H., Shi, Y., Huang, P., Dong, B., Liu, X., et al. (2017). The regulatory factor X protein MoRfx1 is required for development and pathogenicity in the rice blast fungus Magnaporthe oryzae. Mol. Plant Pathol. 18, 1075–1088. doi: 10.1111/mpp.12461
Talbot, N. J. (2003). On the trail of a cereal killer: exploring the biology of Magnaporthe grisea. Annu. Rev. Microbiol. 57, 177–202. doi: 10.1146/annurev.micro.57.030502.090957
Talbot, N. J., Ebbole, D. J., and Hamer, J. E. (1993). Identification and characterization of MPG1, a gene involved in pathogenicity from the rice blast fungus Magnaporthe grisea. Plant Cell 5, 1575–1590. doi: 10.1105/tpc.5.11.1575
Talbot, N. J., Kershaw, M. J., Wakley, G. E., deVries, O. M. H., Wessels, J. G. H., and Hamer, J. E. (1996). MPG1 encodes a fungal hydrophobin involved in surface interactions during infection-related development of Magnaporthe grisea. Plant Cell 8, 985–999. doi: 10.1105/tpc.8.6.985
Tang, Q. Y., and Zhang, C. X. (2013). Data processing system (DPS) software with experimental design, statistical analysis and data mining developed for use in entomological research. Insect Sci. 20, 254–260. doi: 10.1111/j.1744-7917.2012.01519.x
Thompson, J. E., Fahnestock, S., Farrall, L., Liao, D. I., Valent, B., and Jordan, D. B. (2000). The second naphthol reductase of fungal melanin biosynthesis in Magnaporthe grisea: tetrahydroxynaphthalene reductase. J. Biol. Chem. 275, 34867–34872. doi: 10.1074/jbc.M006659200
Tucker, S. L., and Talbot, N. J. (2001). Surface attachment and pre-penetration stage development by plant pathogenic fungi. Annu. Rev. Phytopathol. 39, 385–417. doi: 10.1146/annurev.phyto.39.1.385
Valadi, A., Granath, K., Gustafsson, L., and Adler, L. (2004). Distinct intracellular localization of Gpd1p and Gpd2p, the two yeast isoforms of NAD+-dependent glycerol-3-phosphate dehydrogenase, explains their different contributions to redox-driven glycerol production. J. Biol. Chem. 279, 39677–39685. doi: 10.1074/jbc.M403310200
Wang, J., Yin, Z., Tang, W., Cai, X., Gao, C., Zhang, H., et al. (2016). The thioredoxin MoTrx2 protein mediates reactive oxygen species (ROS) balance and controls pathogenicity as a target of the transcription factor MoAP1 in Magnaporthe oryzae. Mol. Plant Pathol. 18, 1199–1209. doi: 10.1111/mpp.12484
Wei, Y. D., Periappuram, C., Datla, R., Selvaraj, G., and Zou, J. T. (2001). Molecular and biochemical characterizations of a plastidic glycerol-3-phosphate dehydrogenase from Arabidopsis. Plant Physiol. Biochem. 39, 841–848. doi: 10.1016/S0981-9428(01)01308-0
Wierman, M. B., and Smith, J. S. (2014). Yeast sirtuins and the regulation of aging. FEMS Yeast Res. 14, 73–88. doi: 10.1111/1567-1364.12115
Wilson, R. A., Gibson, R. P., Quispe, C. F., Littlechild, J. A., and Talbot, N. J. (2010). An NADPH-dependent genetic switch regulates plant infection by the rice blast fungus. Proc. Natl. Acad. Sci. U.S.A. 107, 21902–21907. doi: 10.1073/pnas.1006839107
Yang, J., Zhao, X., Sun, J., Kang, Z., Ding, S., Xu, J. R., et al. (2009). A novel protein Com1 is required for normal conidium morphology and full virulence in Magnaporthe oryzae. Mol. Plant Microbe Interact. 23, 112–123. doi: 10.1094/MPMI-23-1-0112
Yeh, J. I., Chinte, U., and Du, S. (2008). Structure of glycerol-3-phosphate dehydrogenase, an essential monotopic membrane enzyme involved in respiration and metabolism. Proc. Natl. Acad. Sci. U.S.A. 105, 3280–3285. doi: 10.1073/pnas.0712331105
Zhang, S., Jiang, C., Zhang, Q., Qi, L., Li, C., and Xu, J. R. (2016). Thioredoxins are involved in the activation of the PMK1 MAP kinase pathway during appressorium penetration and invasive growth in Magnaporthe oryzae. Environ. Microbiol. 18, 3768–3784. doi: 10.1111/1462-2920.13315
Zhang, S., Liang, M., Naqvi, I. N., Lin, C., Qian, W., Zhang, L. H., et al. (2017). Phototrophy and starvation-based induction of autophagy upon removal of Gcn5-catalyzed acetylation of Atg7 in Magnaporthe oryzae. Autophagy 13, 1318–1330. doi: 10.1080/15548627.2017.1327103
Zhou, W., Shi, W., Xu, X. W., Li, Z. G., Yin, C. F., Peng, J. B., et al. (2017). Glutamate synthase MoGlt1-mediated glutamate homeostasis is important for autophagy, virulence and conidiation in the rice blast fungus. Mol Plant Pathol. 19, 564–578. doi: 10.1111/mpp.12541
Keywords: rice blast, Pyricularia oryzae, glycerol-3-phosphate dehydrogenase, NAD, redox, light sensing, aerial hypha, pathogenicity
Citation: Shi Y, Wang H, Yan Y, Cao H, Liu X, Lin F and Lu J (2018) Glycerol-3-Phosphate Shuttle Is Involved in Development and Virulence in the Rice Blast Fungus Pyricularia oryzae. Front. Plant Sci. 9:687. doi: 10.3389/fpls.2018.00687
Received: 26 November 2017; Accepted: 04 May 2018;
Published: 23 May 2018.
Edited by:
Devin Coleman-Derr, United States Department of Agriculture, United StatesReviewed by:
Prasanna Kankanala, Noble Research Institute, LLC, United StatesOswaldo Valdes-Lopez, Universidad Nacional Autónoma de México, Mexico
Vicente Mariscal, Instituto de Bioquímica Vegetal y Fotosíntesis (IBVF), Spain
Copyright © 2018 Shi, Wang, Yan, Cao, Liu, Lin and Lu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jianping Lu, anBsdUB6anUuZWR1LmNu
† These authors have contributed equally to this work.
 Yongkai Shi
Yongkai Shi Huan Wang
Huan Wang Yuxin Yan
Yuxin Yan Huijuan Cao
Huijuan Cao Xiaohong Liu
Xiaohong Liu Fucheng Lin
Fucheng Lin Jianping Lu
Jianping Lu