- 1Department of Plant Pathology, University of Minnesota, St. Paul, MN, United States
- 2Cereal Disease Laboratory, Agricultural Research Service, United States Department of Agriculture, St. Paul, MN, United States
Recent stem rust epidemics in eastern Africa and elsewhere demonstrated that wheat stem rust is a re-emerging disease posing a threat to wheat production worldwide. The cultivated wheat gene pool has a narrow genetic base for resistance to virulent races, such as races in the Ug99 race group. Wild relatives of wheat are a tractable source of stem rust resistance genes. Aegilops species in the tertiary genepool have not been exploited to any great extent as a source of stem rust resistance. We evaluated 1,422 accessions of Aegilops spp. for resistance to three highly virulent races (TTKSK, TRTTF, and TTTTF) of Puccinia graminis f. sp. tritici. Species studied include Ae. biuncialis, Ae. caudata, Ae. comosa, Ae. cylindrica, Ae. geniculata, Ae. neglecta, Ae. peregrina, Ae. triuncialis, and Ae. umbellulata that do not share common genomes with cultivated wheat. High frequencies of resistance were observed as 977 (68.8%), 927 (65.2%), and 850 (59.8%) accessions exhibited low infection types to races TTKSK, TTTTF, and TRTTF, respectively. Contingency table analyses showed strong association for resistance to different races in several Aegilops spp., indicating that for a given species, the resistance genes effective against multiple races. Inheritance studies in selected accessions showed that resistance to race TTKSK is simply inherited.
Introduction
Wheat stem rust, caused by Puccinia graminis Pers.:Pers. f. sp. tritici Eriks. & E. Henn. (Pgt), is a devastating disease of durum wheat (Triticum turgidum L. ssp. durum) and common or bread wheat (T. aestivum L.). Severe epidemics have been reported in all major wheat growing areas in the world (Roelfs, 1985; Saari and Prescott, 1985). For decades, stem rust has been under effective control through the use of genetic resistance. The occurrence and spread of Sr31-virulence races in the Ug99 race group in East Africa and other virulent races causing epidemics and localized outbreaks in Ethiopia (Olivera et al., 2015), Europe (Bhattacharya, 2017; Olivera Firpo et al., 2017; Lewis et al., 2018) and Central Asia (Shamanin et al., 2018), indicates that the disease is re-emerging as a threat to wheat production. Races in the Ug99 group have been detected across South, East and northern Africa, and the Middle East (Pretorius et al., 2000; Singh et al., 2015; Newcomb et al., 2016), and have the potential to reach critical wheat growing regions in the world (Park et al., 2011). The Ug99 race group has been rapidly evolving, producing variants with virulence to stem rust resistance genes including Sr24 (Jin et al., 2008), Sr36 (Jin et al., 2009), and SrTmp (Newcomb et al., 2016) that are important in stem rust resistance breeding (Singh et al., 2015).
The cultivated wheat gene pool has a narrow genetic base for resistance to the contemporary virulent races, such as TTKSK (Jin and Singh, 2006; Singh et al., 2006; Newcomb et al., 2016), TRTTF (Olivera et al., 2012); and TKTTF (Olivera Firpo et al., 2017). In order to broaden the basis of stem rust resistance in wheat breeding programs, it is necessary to identify and introgress effective genes from all genepools of wheat. Wild relatives of wheat are a tractable source of stem rust resistance genes. Indeed, a number of resistance genes derived from wild relatives of wheat appeared to be more effective against the races in the Ug99 group than Sr genes of wheat origin (Singh et al., 2006; Jin et al., 2007). Aegilops is the most closely related genus to Triticum (Kimber and Feldman, 1987; Jiang et al., 1994) and comprises 23 species that include diploid, tetraploid, and hexaploid genomes (van Slageren, 1994). Aegilops species are known to be a rich source of stem rust resistance, and several stem rust resistance genes have been transferred into cultivated wheat (Friebe et al., 1996; Schneider et al., 2008; Liu et al., 2011a,b; Olson et al., 2013a,b).
Ease of hybridization and reduced linkage drag make introgression from species in the primary gene pool preferred by wheat breeders to incorporate new alleles in their breeding programs (Feuillet et al., 2008). However, species in the secondary and tertiary gene pools constitute an important reservoir of genetic variability (Qi et al., 2007). Aegilops species in the tertiary genepool have not been exploited to any great extent for wheat improvement, and for resistance to TTKSK and other virulent Pgt races in particular. The objective of this study was to evaluate a collection of nine Aegilops species in the tertiary gene pool of wheat for resistance to race TTKSK and other Pgt races.
Materials and Methods
Germplasm
A total of 1,422 accessions of nine Aegilops species (three diploid and six tetraploid) deposited at the USDA-ARS, National Small Grain Collection (NSGC), Aberdeen, ID, were evaluated in this study. Species, the number of accessions and country of origin of each Aegilops species are given in Table 1.
Inoculation, Incubation, and Disease Assessment
With the objective of identifying multiple and diverse resistance genes in individual accessions, we evaluated this Aegilops collection against multiple races with different virulence spectrum and origin. All accessions were characterized for reaction to three virulent Pgt races: TTKSK (Kenya), TRTTF (Yemen), and TTTTF (United States). Accessions resistant to the three races were further evaluated for their reaction to four additional US races (TPMKC, RKRQC, QTHJC, and QFCSC). The race designations are based on the letter code nomenclature system (Roelfs and Martens, 1988; Roelfs et al., 1993; Jin et al., 2008). Avirulence/virulence profile of the Pgt isolates used in the disease assessments is summarized in Table 2. Disease evaluations were conducted in two independent experiments. In each experiment, five seedlings per accession were inoculated with each race on fully expanded primary leaves 8–9 days after planting. Details on inoculation procedures and disease assessment were described by Jin et al. (2007). Disease reactions were classified according to Stakman et al. (1962). Infection types (ITs) 0, 1, and 2 were considered as resistant reactions and ITs 3 and 4 were considered as susceptible. Wheat cultivar McNair 701 (Cltr 15288) was included as susceptible check. Analyses of association via contingency tables were conducted to assess potential relationships of resistance to different Pgt races.

TABLE 2. Isolate designation, origin, and virulence phenotype of Puccinia graminis f. sp. tritici races used to evaluate resistance in Aegilops spp.
Inheritance Study
Bi-parental crosses between selected resistant accessions and a susceptible accession in five Aegilops species were made and F2 progeny were produced by selfing F1 plants. Seventeen F2 populations (four from Ae. cylindrica, four from Ae. peregrina, six from Ae. triuncialis, two from Ae. umbellulata, and one from Ae. comosa) were evaluated for reaction to race TTKSK to determine the inheritance of resistance based on phenotypic ratios. Chi-square (χ2) test was used to determine the goodness of fit to expected genetic ratios in the F2 generation.
Results
A wide array of infection types was observed across the Aegilops spp. and ranged from highly resistant (IT 0) to highly susceptible (ITs 3+ and 4). Low ITs (; or ;1-) were frequently observed in Ae. caudata, Ae. cylindrica, Ae. neglecta, Ae. peregrina and Ae. triuncialis, whereas ITs 2- and 2-; were predominant in Ae. biuncialis and Ae. geniculata. We observed a high percentage of resistance in this Aegilops collection as 977 (68.8%), 927 (65.2%), and 850 (59.8%) accessions produced low infection types to races TTKSK, TTTTF, and TRTTF, respectively (Table 3). Five hundred and fifty one (38.8%) accessions were resistant to the three races evaluated. The frequencies of accessions resistant to race TTKSK varied among the species: over 80% in six Aegilops species (Ae. caudata, Ae. cylindrica, Ae. geniculata, Ae. neglecta, Ae. peregrina, and Ae. triuncialis), and below 30% in three species (Ae. biuncialis, Ae. comosa, and Ae. umbellulata) (Table 3).
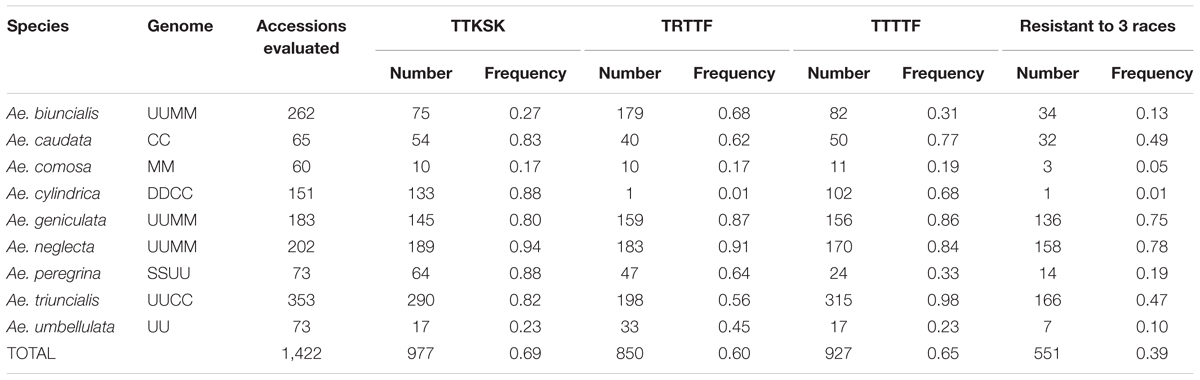
TABLE 3. Number and frequency of Aegilops accessions resistant to Puccinia graminis f. sp. tritici races TTKSK, TRTTF, and TTTTF at the seedling stage.
Pairwise association for resistance to races TTKSK, TRTTF, and TTTTF exhibited variation among species and pathogen races. Over 75% of the accessions of Ae. geniculata and Ae. neglecta were resistant to races TTKSK, TRTTF, and TTTTF (Table 3). Resistance to pairs of the three Pgt races in Ae. geniculata and Ae. neglecta were highly associated (Table 4), suggesting that accessions resistant to race TTKSK are likely to be resistant to races TRTTF and TTTTF. Association for the reaction to races TTSKS, TRTTF, and TTTTF was also observed in Ae. triuncialis (Table 4). Resistance with race specificity was observed in accessions of the remaining species, most noticeably in Ae. cylindrica, where only one accession exhibited resistance to race TRTTF.
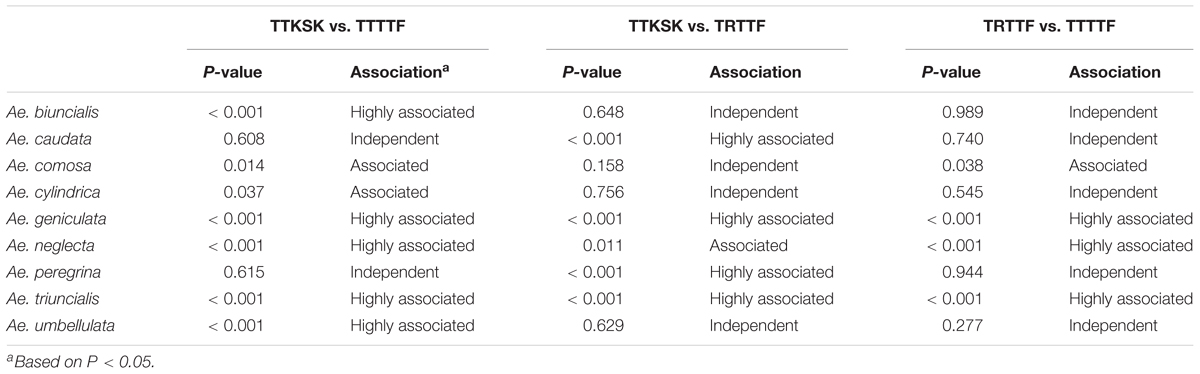
TABLE 4. Probability from contingency tables for association analysis of the reactions of accessions of Aegilops ssp. to races TTKSK, TRTTF, and TTTTF of Puccinia graminis f. sp. tritici.
A group of 408 accessions resistant to races TTKSK, TRTTF, and TTTTF, were evaluated against US races TPMKC, RKRQC, QTHJC, and QFCSC. Three hundred ninety-six accessions remained resistant to all the races evaluated (Supplementary Table 1), indicating these accessions possess genes with broad spectrum resistance.
Sixty-five percent of the accessions evaluated in this study are native to Turkey or Greece. The frequencies of accessions resistant to all races from both countries were similar (30.6%) (Table 5). Higher frequencies of resistance were obtained in accessions from Macedonia (35.8%), France (38.9%), Iraq (46.7%), and Montenegro (75.0%), but the numbers of accessions evaluated from these countries were significantly smaller.
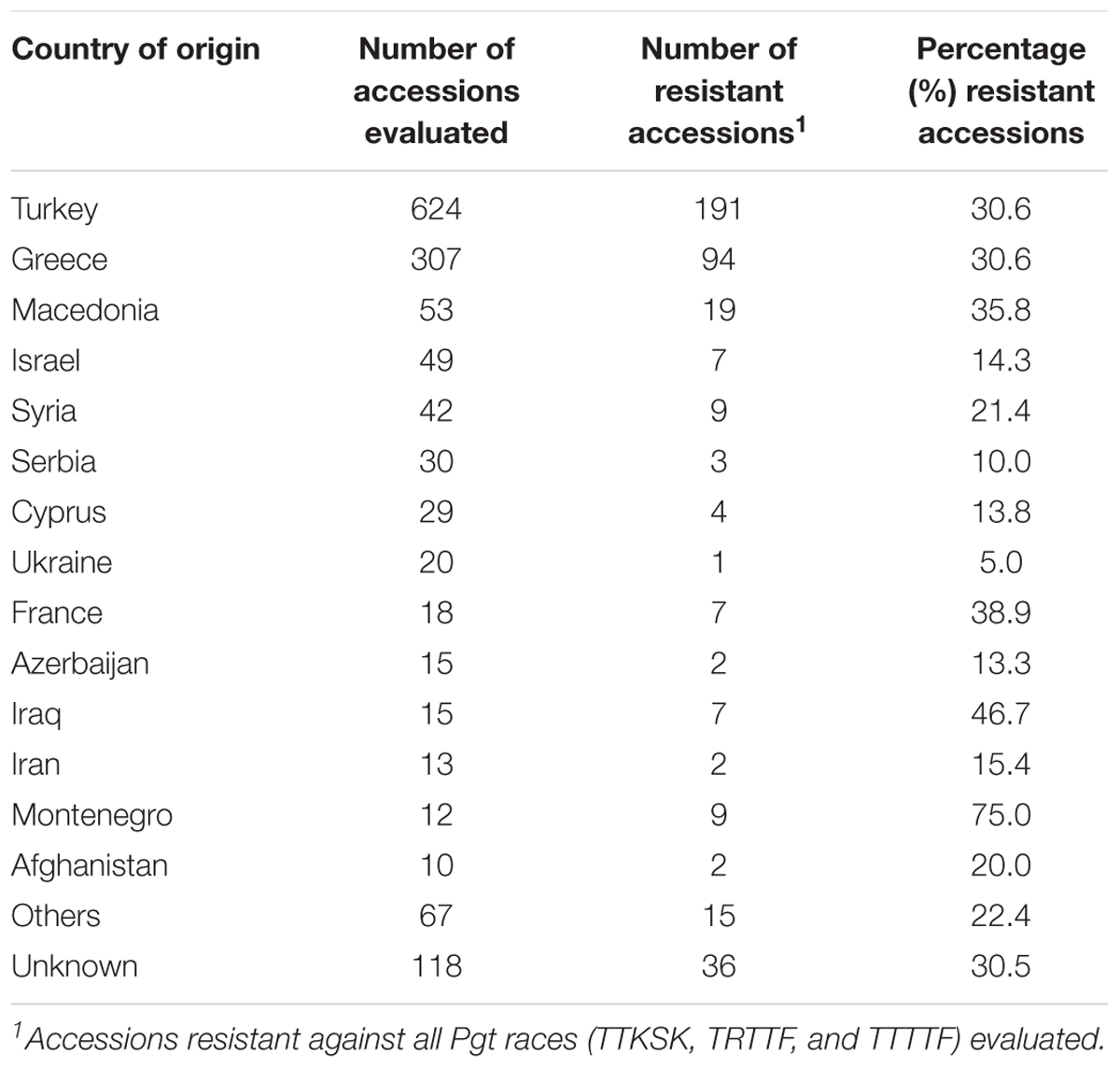
TABLE 5. Number and percentage of resistant Aegilops species accessions according to country of origin.
Segregation ratios of the F2 progeny from biparental crosses between resistant and susceptible accessions indicated that resistance to race TTKSK in selected accessions is mostly conferred by single genes (Table 6). Eight resistant Aegilops accessions carry a single gene with dominant effect, whereas two resistant accessions carry a single gene with recessive effect. Two genes conferring resistance to race TTKSK were observed in three accessions of Ae. triuncialis. Inheritance with epistatic effect between two genes was also observed in four resistant parents. Segregation ratios of the F2 progeny of one Ae. triuncialis and one Ae. umbellulata resistant parent fit to a 9:7 ratio indicating the presence of a complementary gene action with duplicate recessive epistasis. Epistatic effect between two dominant genes was also observed in two Ae. peregrina resistant parents (Table 6), where the F2 progenies fit to a 11:5 ratio.
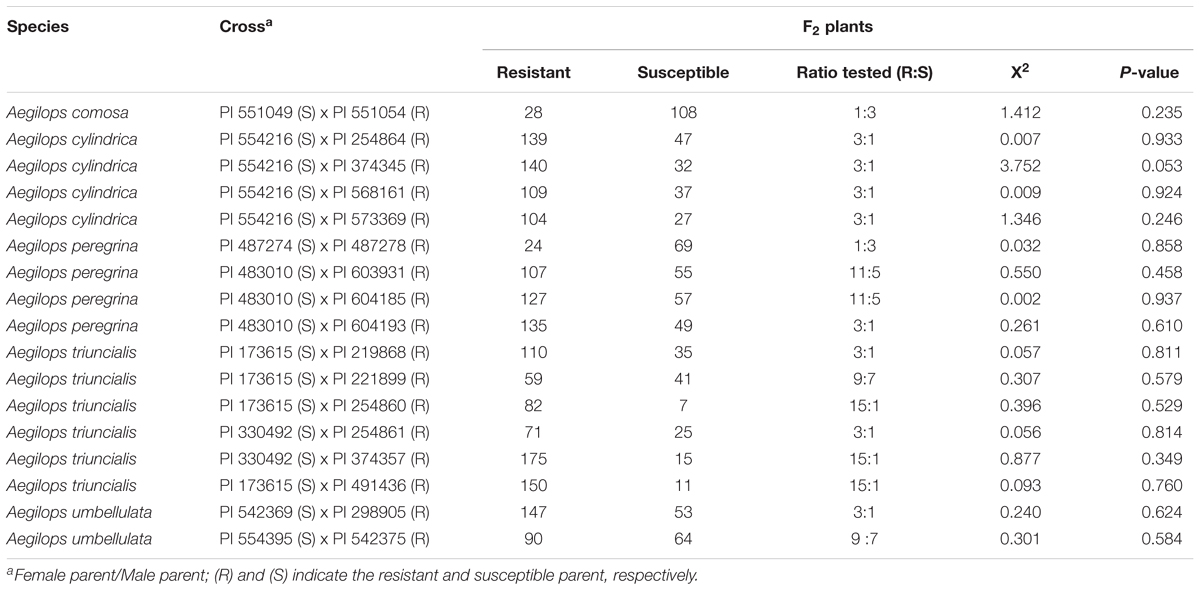
TABLE 6. Segregation of F2 populations of bi-parental crosses of Aegilops spp. to race TTKSK of P. graminis f. sp. tritici.
Discussion
Races of P. graminis f. sp. tritici, such as the Ug99 race group, TKTTF and others detected from the contemporary Pgt populations worldwide, are a serious threat to bread and durum wheat production worldwide because of their broad virulence to many cultivars and rapid geographic spread. The limited number of stem rust resistance genes effective against these virulent races requires the identification of new sources of resistance. Different Aegilops species have contributed several stem rust resistance genes effective against race TTKSK including Sr32, 33, 45, 46, 47, 51, 53, SrTA10187 and SrTA10171 (Friebe et al., 1996; Schneider et al., 2008; Liu et al., 2011a,b; Olson et al., 2013a,b). However, only one gene, Sr53, is derived from Ae. geniculata in the tertiary genepool. Results from this study demonstrated that Aegilops species in the tertiary genepool of wheat are a rich source of resistance to race TTKSK and other Pgt races with broad virulence.
Although the overall frequency of resistant accessions in the entire Aegilops collection evaluated against races TTKSK, TRTTF, and TTTTF in this study was over 60%, we observed significant variation among species. Only two species (Ae. geniculata and Ae. neglecta) exhibited a high frequency of resistance (over 80%) against the three races. Interestingly in Ae. biuncialis, a species that also shares the same genome constitution as Ae. geniculata and Ae. neglecta (UUMM), the frequencies of resistance varied, exhibiting a high level of race specificity. Differences in the frequencies of resistance to stem, stripe, and leaf rust in species carrying the same genome have been also reported in the Section Sitopsis (SS genome) of Aegilops (Anikster et al., 2005; Scott et al., 2014). In species such as Ae. geniculata and Ae. neglecta where there is a high degree of association of the reactions to races TTKSK, TRTTF, and TTTTF, it is highly likely that the genes that confer resistance to one race is also effective against the other races. The progeny populations via bi-parental crosses initiated through this study will be further developed and analyzed to understand the genetic relationships for resistance to different races in these selected accessions.
Race specificity was a common feature observed in this Aegilops collection, as five species exhibited a percentage of accessions resistant to all three races TTKSK, TRTTF, and TTTTF below 20% (Table 3), and have no association of the reaction of two out of three races. Previous studies also report race specificity in Aegilops species (Olivera et al., 2007; Scott et al., 2014). Since gene introgression from Aegilops species in the tertiary genepool is a long and laborious process, it is preferable to use accessions that carry stem rust resistance that is effective against multiple races. About 30% (396 accessions) were resistant against all the races evaluated, indicating the availability of potential sources of new and diverse stem rust resistance genes that could be very useful in wheat breeding programs. Most of these resistant accessions (84%) were from the tetraploid species Ae. geniculata, Ae. neglecta, and Ae. triuncialis. Additional studies are required to assess the diversity in these resistant accessions to allow the identification of donor accessions that are likely to contribute non-redundant stem rust resistance genes. Choosing resistant accessions from geographically diverse countries of origin and exhibiting different infection types for gene introgression is a first step to maximize the chances of capturing new and unique resistance genes (Anikster et al., 2005).
Sixty-five percent of the accessions evaluated in this study originated from Turkey or Greece, two countries having the largest numbers of Aegilops species. Turkey is known to be the center of diversity for Aegilops (Eig, 1929), and 17 out of the 23 Aegilops species have been identified in its territory (van Slageren, 1994). The nine species evaluated in this study are present in Turkey. The number and frequency of resistant accessions from Turkey and Greece (n = 285, 30.6%) from this study demonstrated that valuable sources of new genetic variation for stem rust resistance are present in these countries.
A prior knowledge on the inheritance of resistance in wild wheat relatives will facilitate alien gene introgression into wheat. We produced 17 biparental crosses to investigate the inheritance of TTKSK resistance. These populations will be further developed to map resistance genes and to develop closely linked markers within the wild species. Simple inheritance of stem rust resistance was found in most selected resistant accessions. Our result of a single dominant gene segregating in the Ae. umbellulata biparental F2 population from a cross between PI 542369 and PI 298905 was confirmed in an F3 population and mapped to chromosome 2U (Edae et al., 2016). A similar approach will be followed to characterize the resistance identified in this study. Two stem rust resistance genes were identified in three Ae. triuncialis resistant parents. Further studies are needed to characterize the effectiveness of each resistance gene. Multiple stem rust resistance genes with different resistance profile were reported in Ae. sharonensis (Olivera et al., 2008; Yu et al., 2017). A more complex inheritance of stem rust resistance with genes exhibiting epistatic effects was also observed in three Aegilops species. These results highlight the value of studying the genetics of stem rust resistance in the wild relative before attempting wide crosses for gene transferring.
Aegilops species in the tertiary genepool do not possess genome(s) homologous to the cultivated forms, and gene transfer through homologous recombination cannot be achieved with these species (Harlan and de Wet, 1971). Cytogenetic techniques such as irradiation and chemical treatments, production of synthetic amphiploids, use of gametocidal chromosomes, or Ph1 gene mutants may be required for gene introgression into the cultivated forms (Friebe et al., 1996; Zaharieva and Monneveux, 2006). However, the introgression of alien chromatin to substitute for homoeologous chromosome segments has the potential of a simultaneous introduction of deleterious DNA that can affect agronomic and quality traits of wheat (Feuillet et al., 2008; Wulff and Moscou, 2014). New sequencing technologies, like Genotyping-By-Sequencing, have allowed the development of genetic linkage maps in wild relatives of wheat with non-previous available markers, and the identification of closely linked markers that can facilitate the gene transfer process by reducing the introgressed alien chromatin segment into elite materials (Edae et al., 2016, 2017). The sources of resistance identified from the tertiary genepool will also serve as targets for resistance gene cloning. Cloned genes and their delivery as transgenes in single or multiple resistance gene cassettes will completely resolve the linkage drag problem and ensure the effectiveness and durability of genes derived from more distant relatives of wheat (Wulff and Moscou, 2014). Today, new cloning techniques like mutational genomics (MutRenSeq) (Steuernagel et al., 2016) and association genetics with R gene enrichment sequencing (AgRenSeq) (Arora et al., 2018) allow a rapid and cheaper discovery and cloning of resistance genes. These technologies are opening new doors for fully exploiting the richness and diversity of wild relatives for wheat improvement.
Author Contributions
PO and YJ were involved in the experimental design and manuscript preparation. PO performed the experiments and completed the data analysis. MR was involved in manuscript preparation and revision.
Funding
This research was funded by USDA-ARS and the Durable Rust Resistance in Wheat (OPP1144199) project administrated by Cornell University and funded by the Bill and Melinda Gates Foundation and the United Kingdom Department for International Development.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank H. Bockleman, United States Department of Agriculture-Agricultural Research Service, National Small Grains Collection, Aberdeen, ID, for providing Aegilops spp. seeds.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2018.01719/full#supplementary-material
References
Anikster, Y., Manisterski, J., Long, D. L., and Leonard, K. J. (2005). Resistance to leaf rust, stripe rust, and stem rust in Aegilops spp. in Israel. Plant Dis. 89, 303–308. doi: 10.1094/PD-89-0303
Arora, S., Steuernagel, B., Long, Y., Matny, O., Johnson, R., Enk, J., et al. (2018). Resistance gene discovery and cloning by sequence capture and association genetics. bioRxiv [Preprint]. doi: 10.1101/248146
Bhattacharya, S. (2017). Deadly new wheat disease threatens Europe’s crops. Nature 542, 145–146. doi: 10.1038/nature.2017.21424
Edae, E. A., Olivera, P. D., Jin, Y., Poland, J. A., and Rouse, M. N. (2016). High density linkage map construction with genotype-by-sequencing for Aegilops umbellulata, a wild relative of cultivated wheat. BMC Genomics 17:39966. doi: 10.1534/g3.117.039966
Edae, E. A., Olivera, P. D., Jin, Y., and Rouse, M. N. (2017). Genotyping-by-sequencing facilitates a high-density consensus linkage map for Aegilops umbellulata, a wild relative of cultivated what. G3 7, 1551–1561. doi: 10.1534/g3.117.039966
Eig, A. (1929). Monographisch-kritische Ubersicht der Gattung Aegilops. Feddes Repert., Beih. 55, 1- 228,Tab.I-XVIII. Berlin: Verlag des Repertoriums.
Feuillet, C., Langridge, P., and Waugh, R. (2008). Cereal breeding takes a walk on the wild side. Trends Genet. 24, 24–32. doi: 10.1016/j.tig.2007.11.001
Friebe, B., Jiang, J., Raupp, W. J., McIntosh, R. A., and Gill, B. S. (1996). Characterization of wheat alien translocations conferring resistance to diseases and pests: current status. Euphytica 71, 59–87. doi: 10.1007/BF00035277
Harlan, J. R., and de Wet, J. M. J. (1971). Toward a rational classification of cultivated plants. Taxon 20, 509–517. doi: 10.2307/1218252
Jiang, J., Friebe, B., and Gill, B. S. (1994). Recent advances in alien gene transfer in wheat. Euphytica 73, 199–212. doi: 10.1038/hdy.2012.116
Jin, Y., and Singh, R. P. (2006). Resistance in U.S. wheat to recent eastern African isolates of Puccinia graminis f. sp. tritici with virulence to resistance gene Sr31. Plant Dis. 90, 476–480. doi: 10.1094/PD-90-0476
Jin, Y., Singh, R. P., Ward, R., Wanyera, R., Kinyua, M., Njau, P., et al. (2007). Characterization of seedling infection types and adult plant infection responses of monogenic Sr gene lines to race TTKS of Puccinia graminis f. sp. tritici. Plant Dis. 91, 1096–1099. doi: 10.1094/PDIS-91-9-1096
Jin, Y., Szabo, L. J., Pretorius, Z. A., Singh, R. P., Ward, R. Jr., and Fetch, T. (2008). Detection of virulence to resistance gene Sr24 within race TTKS of Puccinia graminis f. sp. tritici. Plant Dis. 92, 923–926. doi: 10.1094/PDIS-92-6-0923
Jin, Y., Szabo, L. J., Rouse, M., Fetch, T. Jr., Pretorius, Z. A., Wanyera, R., et al. (2009). Detection of virulence to resistance gene Sr36 within race TTKS lineage of Puccinia graminis f. sp. tritici. Plant Dis. 93, 367–370. doi: 10.1094/PDIS-93-4-0367
Kimber, G., and Feldman, M. (1987). Wild Wheat–An Introduction. Special Report 353. Columbia: College of Agriculture, University of Missouri.
Lewis, C. M., Persoons, A., Bebber, D. P., Kigathi, R. N., Maintz, J., Findlay, K., et al. (2018). Potential for re-emergence of wheat stem rust in the United Kingdom. Commun. Biol. 1:13. doi: 10.1038/s42003-018-0013-y
Liu, W., Jin, Y., Rouse, M., Friebe, B., Gill, B., and Pumphrey, M. O. (2011a). Development and characterization of wheat-Ae. searsii Robertsonian translocations and a recombinant chromosome conferring resistance to stem rust. Theor. Appl. Genet. 122, 1537–1545. doi: 10.1007/s00122-011-1553-4
Liu, W., Rouse, M., Friebe, B., Jin, Y., Gill, B., and Pumphrey, M. O. (2011b). Discovery and molecular mapping of a new gene conferring resistance to stem rust, Sr53, derived from Aegilops geniculata and characterization of spontaneous translocation stocks with reduced alien chromatin. Chromosome Res. 19, 669–682. doi: 10.1007/s10577-011-9226-3
Newcomb, M., Olivera, P. O., Rouse, M. N., Szabo, L. J., Johnson, J., Gale, S., et al. (2016). Characterization of Kenyan isolates of Puccinia graminis f. sp. tritici from 2008 to 2014 reveals virulence to SrTmp in the Ug99 race group. Phytopathology 100, 986–996. doi: 10.1094/PHYTO-12-15-0337-R
Olivera, P. D., Anikster, Y., Kolmer, J. A., and Steffenson, B. J. (2007). Resistance of Sharon goat grass (Aegilops sharonensis) to fungal diseases of wheat. Plant Dis. 91, 942–950. doi: 10.1094/PDIS-91-8-0942
Olivera P. D., Jin, Y., Rouse, M., Badebo, A., Fetch, T. Jr., Singh, R. P., et al. (2012). Races of Puccinia graminis f. sp. tritici with combined virulence to Sr13 and Sr9e in a field stem rust screening nursery in Ethiopia. Plant Dis. 96, 623–628. doi: 10.7717/peerj.4146
Olivera, P. D., Millet, E., Anikster, Y., and Steffenson, B. J. (2008). Genetics of resistance to wheat leaf rust, stem rust, and powdery mildew in Aegilops sharonensis. Phytopathology 98, 353–358. doi: 10.1094/PHYTO-98-3-0353
Olivera, P. D., Newcomb, M., Szabo, L. J., Rouse, M., Johnson, J., Gale, S., et al. (2015). Phenotypic and genotypic characterization of race TKTTF of Puccinia graminis f. sp. tritici that caused a wheat stem rust epidemic in southern Ethiopia in 2013/14. Phytopathology 105, 917–928. doi: 10.1094/PHYTO-11-14-0302-FI
Olivera Firpo, P. D., Newcomb, M., Flath, K., Szabo, L. J., Carter, M., Luster, D. G., et al. (2017). Characterization of Puccinia graminis f. sp. tritici isolates derived from an unusual wheat stem rust outbreak in Germany in 2013. Plant Pathol. 66, 1258–1266. doi: 10.1111/ppa.12674
Olson, E. L., Rouse, M. N., Pumphrey, M. O., Bowden, R. L., Gill, B. S., and Poland, J. A. (2013a). Introgression of stem rust resistance genes SrTA10187 and SrTA10171 from Aegilops tauschii to wheat. Theor. Appl. Genet. 126, 2477–2484. doi: 10.1007/s00122-013-2148-z
Olson, E. L., Rouse, M. N., Pumphrey, M. O., Bowden, R. L., Gill, B. S., and Poland, J. A. (2013b). Simultaneous transfer, introgression, and genomic localization of genes for resistance to stem rust race TTKSK (Ug99) from Aegilops tauschii to wheat. Theor. Appl. Genet. 126, 1179–1188. doi: 10.1007/s00122-013-2045-5
Park, R., Fetch, T., Hodson, D., Yin, J., Nazari, K., Prashar, M., et al. (2011). International surveillance of wheat pathogens: progress and challenges. Euphytica 179, 109–117. doi: 10.1007/s10681-011-0375-4
Pretorius, Z. A., Singh, R. P., Wagoire, W. W., and Payne, T. S. (2000). Detection of virulence to wheat stem rust resistance genes Sr31 in Puccinia graminis f. sp. tritici in Uganda. Plant Dis. 84, 2003. doi: 10.1094/PDIS.2000.84.2.203B
Qi, L., Friebe, B., Zhang, P., and Gill, B. S. (2007). Homoeologous recombination, chromosome engendering and crop improvement. Chromosome Res. 15, 3–19. doi: 10.1007/s10577-006-1108-8
Roelfs, A. P. (1985). “Wheat and rye stem rust,” in The Cereal Rusts Vol. II: Diseases, Distribution, Epidemiology and Control, eds W. R. Bushnell and A. P. Roelfs (Orlando, FL: Academic Press), 4–37.
Roelfs, A. P., Long, D. L., and Roberts, J. J. (1993). Races of Puccinia graminis in the United States during 1992. Plant Dis. 77, 1122–1125. doi: 10.1094/PD-77-1122
Roelfs, A. P., and Martens, J. W. (1988). An international system of nomenclature for Puccinia graminis f. sp. tritici. Phytopathology 78, 526–533. doi: 10.1094/Phyto-78-526
Saari, E. E., and Prescott, J. M. (1985). “World distribution in relation to economic losses. Wheat and rye stem rust,” in The Cereal Rusts Vol. II: Diseases, Distribution, Epidemiology and Control, eds W. R. Bushnell and A. P. Roelfs (Orlando, FL: Academic Press), 259–298. doi: 10.1016/B978-0-12-148402-6.50017-1
Schneider, A., Molnar, I., and Molnar-Lang, M. (2008). Utilisation of Aegilops (goatgrass) species to widen the genetic diversity of cultivated wheat. Euphytica 163, 1–9. doi: 10.1007/s10681-007-9624-y
Scott, J. C., Manisterski, J., Sela, H., Ben-Yehuda, P., and Steffenson, B. J. (2014). Resistance of Aegilops species from Israel to widely virulent African and Israeli races of the wheat stem rust pathogen. Plant Dis. 98, 1309–1320. doi: 10.1094/PDIS-01-14-0062-RE
Singh, R. P., Hodson, D. P., Jin, Y., Huerta-Espino, J., Kinyua, M. G., Wanyera, R., et al. (2006). Current status, likely migration and strategies to mitigate the threat to wheat production from race Ug99 (TTKS) of stem rust pathogen. CAB Rev. Perspect. Agric. Vet. Sci. Nutr. Nat. Res. 54, 1–13. doi: 10.1079/PAVSNNR20061054
Singh, R. P., Hodson, D. P., Jin, Y., Lagudah, E. S., Ayliffe, M. A., Bhavani, S., et al. (2015). Emergence and spread of new races of wheat stem rust fungus: continued threat to food security and prospects of genetic control. Phytopathology 105, 872–884. doi: 10.1094/PHYTO-01-15-0030-FI
Shamanin, V., Salina, E., Zelenskiv, Y., Kokhmetova, A., Patpour, M., Holmoller, M., et al. (2018). “Large scale wheat stem rust outbreaks in Western Siberia/Northern Kazakhstan in 2015–2017,” in Proceedings of the BGRI 2018 Technical Workshop. (Omsk: Omsk State Agricultural University).
Stakman, E. C., Stewart, D. M., and Loegering, Q. W. (1962). Identification of Physiologic Races of Puccinia graminis var. tritici. Maryland: United States Department of Agriculture, Agricultural Research Service, 617.
Steuernagel, B., Periyannan, S. K., Hernaìndez-Pinzoìn, I., Witek, K., Rouse, M. N., Yu, G., et al. (2016). Rapid cloning of disease-resistance genes in plants using mutagenesis and sequence capture. Nat. Biotechnol. 34, 652–655. doi: 10.1038/nbt.3543
van Slageren, M. W. (1994). Wild wheats: a monograph of Aegilops L. and Amblyopyrum (Jaub. & Spach) Eig (Poaceae). Wageningen: Wageningen Agriculture University Papers.
Wulff, B. B. H., and Moscou, M. J. (2014). Strategies for transferring resistance into wheat: from wide crosses to GM cassettes. Front. Plant Sci. 5:692. doi: 10.3389/fpls.2014.00692
Yu, G., Champouret, N., Steuernagel, B., Olivera, P. D., Simmons, J., Williams, C., et al. (2017). Discovery and characterization of two new stem rust resistance genes in Aegilops sharonensis. Theor. Appl. Genet. 130, 1207–1222. doi: 10.1007/s00122-017-2882-8
Keywords: wild wheats, disease resistance, Ug99, genetic resources, tertiary genepool
Citation: Olivera PD, Rouse MN and Jin Y (2018) Identification of New Sources of Resistance to Wheat Stem Rust in Aegilops spp. in the Tertiary Genepool of Wheat. Front. Plant Sci. 9:1719. doi: 10.3389/fpls.2018.01719
Received: 26 August 2018; Accepted: 05 November 2018;
Published: 22 November 2018.
Edited by:
István Molnár, Centre for Agricultural Research (MTA), HungaryReviewed by:
Peter Michael Dracatos, University of Sydney, AustraliaHakan Ozkan, Çukurova University, Turkey
Copyright © 2018 Olivera, Rouse and Jin. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yue Jin, WXVlLkppbkBhcnMudXNkYS5nb3Y=
 Pablo D. Olivera
Pablo D. Olivera Matthew N. Rouse2
Matthew N. Rouse2 Yue Jin
Yue Jin