- 1Laboratory of Biotechnology and Plant Pathology, DISTAL, University of Bologna, Bologna, Italy
- 2Plant Protection Research Division of Melkasa Agricultural Research Center, Ethiopian Institute of Agricultural Research (EIAR), Addis Ababa, Ethiopia
- 3Genomics and Biology of Fruit Crops Department, Research and Innovation Centre, Fondazione Edmund Mach, San Michele all’Adige, Italy
- 4Unit of Computational Biology, Research and Innovation Centre, Fondazione Edmund Mach, San Michele all’Adige, Italy
- 5ESAT-ELECTA, Electrical Energy and Computer Architectures, Leuven, Belgium
Gray mold caused by Botrytis cinerea is a major cause of economic losses in strawberry fruit production, limiting fruit shelf life and commercialization. When the fungus infects Fragaria × ananassa strawberry at flowering or unripe fruit stages, symptoms develop after an extended latent phase on ripe fruits before or after harvesting. To elucidate the growth kinetics of B. cinerea on flower/fruit and the molecular responses associated with low susceptibility of unripe fruit stages, woodland strawberry Fragaria vesca flowers and fruits, at unripe white and ripe red stages, were inoculated with B. cinerea. Quantification of fungal genomic DNA within 72 h postinoculation (hpi) showed limited fungal growth on open flower and white fruit, while on red fruit, the growth was exponential starting from 24 hpi and sporulation was observed within 48 hpi. RNA sequencing applied to white and red fruit at 24 hpi showed that a total of 2,141 genes (12.5% of the total expressed genes) were differentially expressed due to B. cinerea infection. A broad transcriptional reprogramming was observed in both unripe and ripe fruits, involving in particular receptor and signaling, secondary metabolites, and defense response pathways. Membrane-localized receptor-like kinases and nucleotide-binding site leucine-rich repeat genes were predominant in the surveillance system of the fruits, most of them being downregulated in white fruits and upregulated in red fruits. In general, unripe fruits exhibited a stronger defense response than red fruits. Genes encoding for pathogenesis-related proteins and flavonoid polyphenols as well as genes involved in cell-wall strengthening were upregulated, while cell-softening genes appeared to be switched off. As a result, B. cinerea remained quiescent in white fruits, while it was able to colonize ripe red fruits.
Introduction
Strawberry ripening is characterized by the simultaneous changes in the physical and chemical structure of the fruit such as the size, color, flavor, aroma, and texture (Schwab and Raab, 2004; Liao et al., 2018). From the inflorescence, the receptacle develops into the edible part of the fruit and increases in size as determined by the volume and expansion of cells (Perkins-Veazie, 1995). The fruit color changes from green to white and ultimately turns red as a function of chlorophyll degradation and increased anthocyanin synthesis. Soluble solids and volatile organic compounds which confer flavor and aroma also increase with ripening (Manning, 1993; Perkins-Veazie, 1995; Montero et al., 1996; Schwab and Raab, 2004). Meanwhile, fruits of Fragaria × ananassa soften with ripening due to the solubilization and depolymerization of cell wall components in strawberry (Rosli et al., 2004; Payasi et al., 2009). As a nonclimacteric crop, the significance of abscisic acid (ABA) as a key hormone in the ripening of F. × ananassa has been demonstrated and recognized (Jia et al., 2011; Symons et al., 2012), and recent studies have suggested that ABA is important in the regulation of ripening-related genes in strawberry (F. × ananassa) via perception and signal transduction (Li et al., 2011).
Along with the changes that occur during ripening is the decline of the innate immunity of crops against pathogen infection (Amil-Ruiz et al., 2011). The disassembly of cell walls that act as a structural barrier against invading organisms during ripening contributes to making strawberry (F. × ananassa) fruits susceptible to pathogen attack (Guidarelli et al., 2011). Meanwhile, secondary plant metabolites which are essential in determining nutritional and sensory characteristics of fruits also functions as preformed defense compounds against several pathogens. Concentrations of specific antifungal compounds have been found to decrease from the unripe to the ripe fruit stages of F. × ananassa (Puhl and Treutter, 2008; Guidarelli et al., 2011; Nagpala et al., 2016). The volatile compounds that are abundantly present in red fruits of F. × ananassa were also found to be beneficial for pathogen growth (Neri et al., 2015).
Botrytis cinerea is one of the most destructive pre- and postharvest strawberry pathogens causing gray mold rot (Feliziani and Romanazzi, 2016), and it is regarded as the second most important fungal pathogen worldwide (Dean et al., 2012). The fungus is a cosmopolitan pathogen able to infect a wide range of plant species (Elad et al., 2016). Moreover, while interacting with resistant unripe fruit, the pathogen is capable of developing quiescent infections, stopping its growth for extended time, and leading to disease symptoms only upon senescence/ripening of host tissues (van Kan, 2005; Prusky et al., 2013; Feliziani and Romanazzi, 2016).
In strawberry (F. × ananassa), early infection of B. cinerea may occur in flowers but remains suppressed until fruit ripening (Bristow et al., 1986; Bulger et al., 1987). It is suggested that the high concentrations of preformed antifungal compounds in the inflorescence of F. × ananassa are responsible for the inactivity of B. cinerea in floral tissues (Terry et al., 2004). Phenotypic evidence of B. cinerea quiescencein F. × ananassa fruits has been demonstrated by artificial inoculation of unripe white and ripe red fruits. Symptoms of infection became evident on red fruits, while white fruits were symptom-free for at least 2 weeks. The advancement of the disease in white fruits 2 weeks after infection is accompanied by evident fruit softening (Nagpala et al., 2016). Meanwhile, elevated concentration of proanthocyanidins, ellagitanins, and catechins were also observed in developing fruits of F. × ananassa inoculated with B. cinerea (Puhl and Treutter, 2008; Nagpala et al., 2016).
The ripening stage of the fruit during which the pathogen established infection largely determines the quiescence of B. cinerea. Unripe strawberry fruits are said to be resistant to infection due to the aforementioned innate immunity that could suppress pathogen growth. On the other hand, ripe strawberry fruits provide favorable conditions for the pathogen to resume activity (Petrasch et al., 2019). While some important physical and biochemical aspects related to the ontogenic resistance of strawberry fruits has been explored, several transcriptional mechanisms remain unknown. In recent years, the application of microarray or RNA-Seq allows for a comprehensive and precise elucidation of the gene expression in a given sample. Transcriptome-wide profiling applied to investigate host–pathogen interactions highlights those transcripts regulated upon pathogen infection. Specific studies focused on the Arabidopsis response to B. cinerea (AbuQamar et al., 2016; Sham et al., 2017) and on the quiescence of fungal pathogens in different development stages of apple leaves (Gusberti et al., 2013), tomato fruits (Alkan et al., 2015), and grapevine fruitlets (Haile et al., 2017) provided stage-specific transcriptome profiles likely related to ontogenic resistance and pathogen activity. A recent RNA-Seq analysis on ripe strawberry (F. × ananassa) fruits infected with B. cinerea reported the time-expression profile of genes from the pathogen and the host during the interaction. Significant changes in the ripe fruit tissues were noted 24 h after B. cinerea inoculation, with emphasis on genes involved in pathogen recognition and signal transduction (Xiong et al., 2018).
In the present study, the transcriptome profiles of the unripe white and ripe red fruits of woodland strawberry (Fragaria vesca) inoculated with B. cinerea were analyzed by RNA-Seq technology in an attempt to elucidate ontogenic resistance mechanisms of strawberry to pathogen attack. The study also provides insights on the gene expression variation between two ripening stages of a nonclimacteric fruit infected with B. cinerea. The findings of this study provide useful information to improve the resistance of strawberry fruits against B. cinerea, an essential factor to minimize losses caused by the gray mold rot along the strawberry supply chain.
Materials and Methods
Pathogen Inoculum and Plant Material
B. cinerea (isolate B05.10) was cultured on potato dextrose agar at 21°C, with a photoperiod of 12 h using near ultraviolet light. Meanwhile, potted plants of F. vesca cv. “Alpine” were grown under controlled conditions at 22°C with natural light (on average 14 h light during April/May). Conventional management practices were observed, and the plants were maintained pesticide free.
To inoculate strawberry flowers and fruits, conidial suspension of B. cinerea was prepared from a 12-day-old culture. The conidia concentration of the suspension was adjusted to 1 × 105 ml−1 using a hemacytometer. Open flowers [7 days after anthesis (DAA)] and white (14 DAA) and red (21 DAA) fruits were used. The flowers were inoculated by dropping 10 μl of the conidial suspension on the base of receptacle, while the fruits were inoculated on the fruit surface, toward the base. The same procedure was followed for the control treatment, where distilled water was used instead. The inoculation was made on flowers and fruits intact to its mother plant. After inoculation, the whole plant was bagged in a water-sprayed, clear plastic bag for 24 h to ensure high humidity.
Growth Kinetics of B. Cinerea on F. Vesca Flower and Fruit
To investigate the growth of B. cinerea in different tissues of strawberry, flower, white fruit, and red fruit were inoculated. Three biological replicates were used for each tissue, where each replicate was composed of 10 open flowers and 5 white and red fruits. Samples were collected at 0, 24, 48, and 72 h postinoculation (hpi), and samples were immediately frozen in liquid nitrogen and kept at −80°C until use. Flower samples, including petals, calyx, stamens, and pistils, were frozen, while fruits were frozen with achenes, without calyx and pedicel.
DNA of B. cinerea was extracted from homogenized strawberry tissue and from 2-week-old mycelium with fungal DNA kit (NucleoSpin Plant II, MN), following the manufacturer’s instructions. On the other hand, DNA extraction from strawberry samples was carried out using a cetyl trimethylammonium bromide (CTAB) method, as described by Porebski et al. (1997), with modifications (Supplemental Information 1). DNA from mycelium and uninoculated red strawberry fruits were used to generate calibration curves to estimate the amount of fungal DNA in inoculated samples.
Using the genomic DNA as a template, quantitative polymerase chain reaction (qPCR) assays were carried out with the MX3000 thermocycler (Stratagene, CA, USA) to amplify target regions B. cinerea and F. vesca. Primers targeting the ribosomal region between 28S and 18S genes (intergenic spacer), Bc3, were used for B. cinerea, while primers targeting the elongation factor gene, EF, were used for F. vesca (Supplemental Table 1). A total of 12.5 µl reaction volume was prepared containing 2.5 µl DNA with Maxima® SYBR Green/ROX qPCR Master Mix (2X; Fermentas) and 200 nM of specific forward and reverse primers. In place of the DNA template, sterile water was used for the negative control. The cycling parameters were as follows: 5 min at 95°C, 40 cycles of 15 s at 95°C, 25 s at 61°C, and 30 s at 72°C. A melting curve was established from 55 to 90°C by changing 0.5°C every 10 s. The efficiency (E) of the PCR assay was calculated using the formula, E = (10 − 1/slope − 1) × 100, where the slope was extracted from the curve Ct = f(log Q0) and Q0 is the initial DNA in the assay. E was expressed as percentage. A fivefold dilution series of genomic DNA of the pathogen and the fruit were used to carry out qPCR reaction to create standard curve by plotting the log value of the starting concentration of DNA (ng) versus the Ct value (Supplemental Figure 1). Eventually, the Ct values of the target DNA from the inoculated samples were used to quantify the initial amount of genomic DNA through extrapolation to its corresponding standard curve. The method was applied for both B. cinerea and F. vesca. The growth of the fungal pathogen in strawberry tissue was analyzed by normalizing the Botrytis DNA concentration to the amount of strawberry genomic DNA in that sample. Fungal growth was expressed as picogram of Botrytis DNA/nanogram of strawberry DNA.
RNA Sequencing, Data Processing, and Data Analysis
Samples of white and red fruits, inoculated or mock-inoculated, in three biological replicates were collected at 24 hpi and subjected to RNA sequencing. Total RNA was extracted from frozen samples with mortar and pestle following the protocol described by Lopez-Gomez and Gomez-Lim (1992).
Next generation sequencing of the RNA samples, including sample quality control, was performed by Genomix4life S.R.L. (Baronissi, Salerno, Italy). Indexed libraries were prepared from 2 μg of each purified RNA with TruSeq RNA Sample Prep Kit (Illumina) according to the manufacturer’s instructions. Libraries were quantified using the Agilent 2100 Bioanalyzer (Agilent Technologies) and pooled such that each index-tagged sample was present in equimolar amounts, with a final concentration of 2 nM for the pooled samples. The pooled samples were subjected to cluster generation and sequencing using an Illumina HiSeq 2500 System (Illumina) in a 2 × 100 paired-end format at a final concentration of 8 pM. All raw RNA‐Seq read data are deposited in the National Center for Biotechnology Information (NCBI) Short Read Archive (http://www.ncbi.nlm.nih.gov/sra/) under the BioProject accession code PRJNA530684.
The raw sequence files generated underwent quality control analysis using FastQC Version 0.10.1 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/), and the Illumina paired-end reads were preprocessed for both quality and adapter trimming with fqtrim Version 0.94 was used with default settings (-A -125 parameters) in order to avoid any read data loss due to false positive (https://ccb.jhu.edu/software/fqtrim/index.shtml). A default setting with the Subread aligner Version 1.4.6 (Liao et al., 2013) was also used for the preprocessed reads to align with the F. vesca genome v1.0 assembly (Shulaev et al., 2011). Raw read counts were extracted from the Subread alignments using the feature Count Version 1.4.6 read summarization program (Liao et al., 2014).
The differential expression analysis was performed with the voom method (Law et al., 2014), which estimates the mean–variance relationship of the log counts, generating a precision weight for each observation that is fed into the limma Version 3.23 empirical Bayes analysis pipeline (Smyth, 2004). Differentially expressed genes (DEGs) of F. vesca were filtered using a threshold of absolute fold change of log2 ≥ 0.8 with p ≤ 0.05. The thresholds were selected based on the expression distribution of each gene, as depicted in volcano plots (Supplemental Figure 2). Gene ontology (GO) and functional annotations were assigned based on F. vesca version 1.0 hybrid gene models (https://www.rosaceae.org/species/fragaria/fragaria_vesca/genome_v1.0; Shulaev et al., 2011) and Blast2GO (e-value 10−3) (Conesa et al., 2005). Functional enrichment was analyzed with AgriGO analysis tool (false discovery rate ≤0.05; Du et al., 2010), using customized annotation and annotated reference of GO terms. MapMan tool (Thimm et al., 2004) was used to visualize DEGs in the context of biotic stress pathway category.
Quantitative Real-Time PCR Validation of RNA-Seq Data
The expression level of eight randomly selected genes was analyzed to validate RNA-Seq results (Supplemental Table 1). cDNA of F. vesca was generated from 1 μg of RNA using ImProm-II Reverse TranscriptaseTM (Promega, USA), following the protocol. Amplification of the cDNA was performed with MX3000 thermocycler (Stratagene, CA, USA), utilizing the same proportions of the mix and cycling parameters described in the growth kinetics trial. Quantification was carried out using the relative standard curve method (Applied Biosystems, 1997 updated 2001). Resulting expression of target genes was normalized with elongation factor (EF) housekeeping gene.
Results and Discussion
Botrytis Cinerea Infection of Strawberry Flower and Fruit
Strawberry plants (F. vesca) at the stage of open flower, white fruit, and red fruit were inoculated with 1,000 conidia of B. cinerea strain B05.10 (Figure 1). Infection symptoms were already clearly visible on red fruit at 2 days postinoculation (dpi) and became very severe at 5 dpi (Figure 1A). No symptoms of Botrytis infection were detected on open flowers(except for petals where brown decay was visible) and white fruits up to 5 dpi (Figure 1A). A time-course real-time PCR quantification of Botrytis genomic DNA was also performed on the inoculated flowers and fruits to measure the real spread of the fungus also at the early stages after inoculation, in the absence of visible symptoms. Consistent with the phenotype observations, very limited increase in fungal DNA was detected on open flowers and white fruits, and only a very minor growth occurred up to 48 hpi. Conversely, in red fruits, Botrytis showed an exponential growth starting at 24 hpi (Figure 1B). This indicates that i) receptacles in open flowers or white stages are much less susceptible to B. cinerea infection and ii) possibly the pathogen infecting at these stages may become quiescent. Similarly, resistance of flowers and fruitlets of grapevine to B. cinerea was also observed where Botrytis, inoculated at cap-off stage, remained quiescent for 12 weeks before egression took place at ripening (Haile et al., 2017).
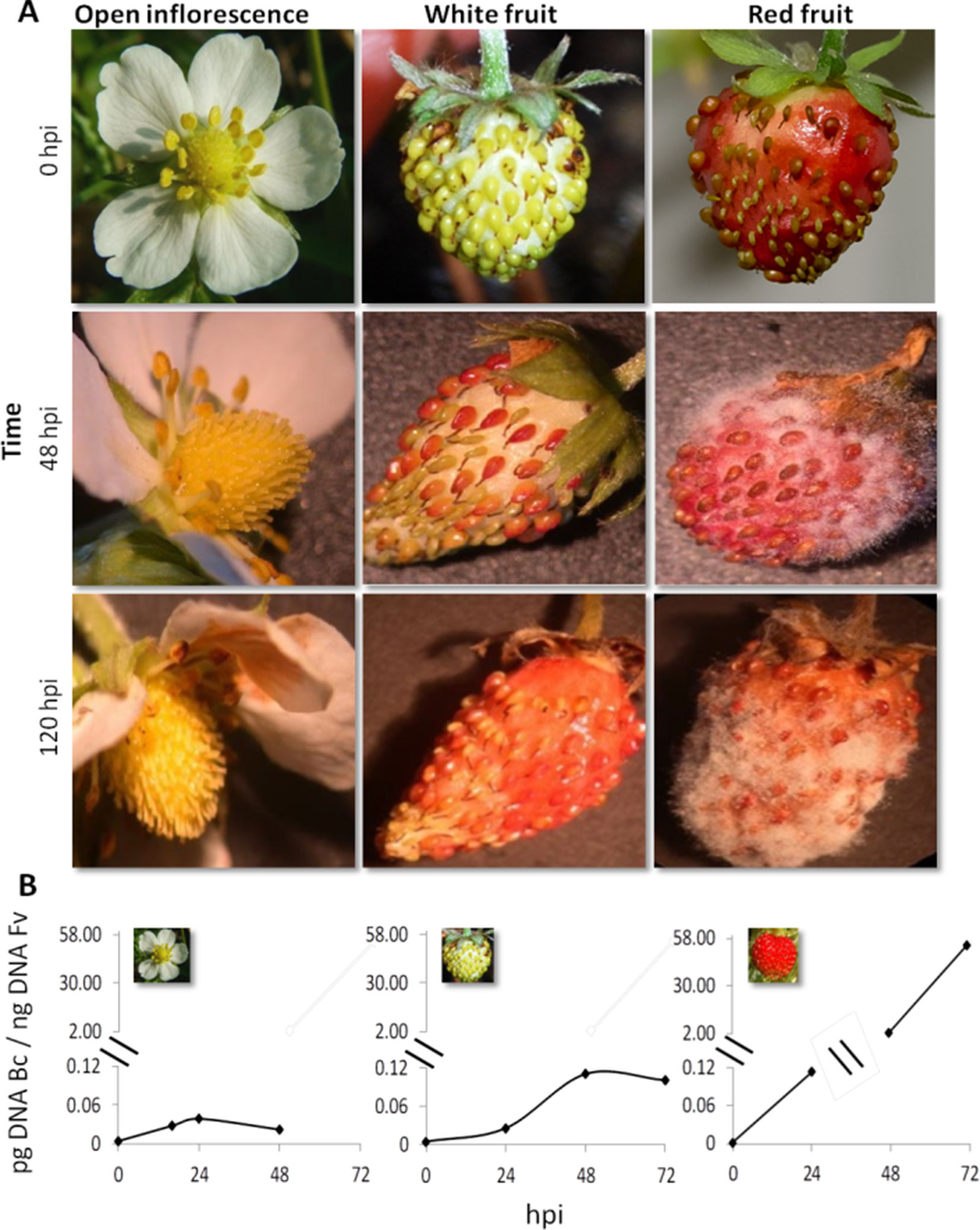
Figure 1 Botrytis cinerea growth on Fragaria vesca flower and fruits. (A) Progress of B. cinerea infection on flower, white fruit, and red fruit up to 120 h postinoculation (hpi). Inoculation was made by dropping 10 μl of 1 × 105 ml−1 conidial suspension. (B) Growth kinetics of B. cinerea on flower, white fruit, and red fruit of F. vesca. Bc, B. cinerea; Fv, F. vesca.
So far, ontogenic variations of polyphenol biosynthesis in strawberry during fruit ontogenesis have been indicated as the basis for quiescence/infection of B. cinerea (Jersch et al., 1989; Puhl and Treutter, 2008; Nagpala et al., 2016). This study provides a molecular understanding to the response of unripe and ripe strawberry fruits to B. cinerea infection, at the whole transcriptome level.
RNA-Seq Analysis of Botrytis-Inoculated White and Red Strawberry Fruits
The transcriptome of white and red woodland strawberry fruits was analyzed in three biological replicates at 24 hpi. Twenty-four hours postinoculation was chosen as sampling time since the qPCR analysis showed that, at this point, the pathogen growth becomes active, especially in red fruits (Figure 1B). The 24 hpi was also reported previously as the most transcriptionally responsive time point in F. × ananassa strawberry ripe fruits in response to B. cinerea (Xiong et al., 2018). The number of reads mapped on F. vesca genome is provided in Supplemental Table 2. Based on principal component analysis, samples were separated by their infection state along the second component and by their ripening stage along the first component (Figure 2A).
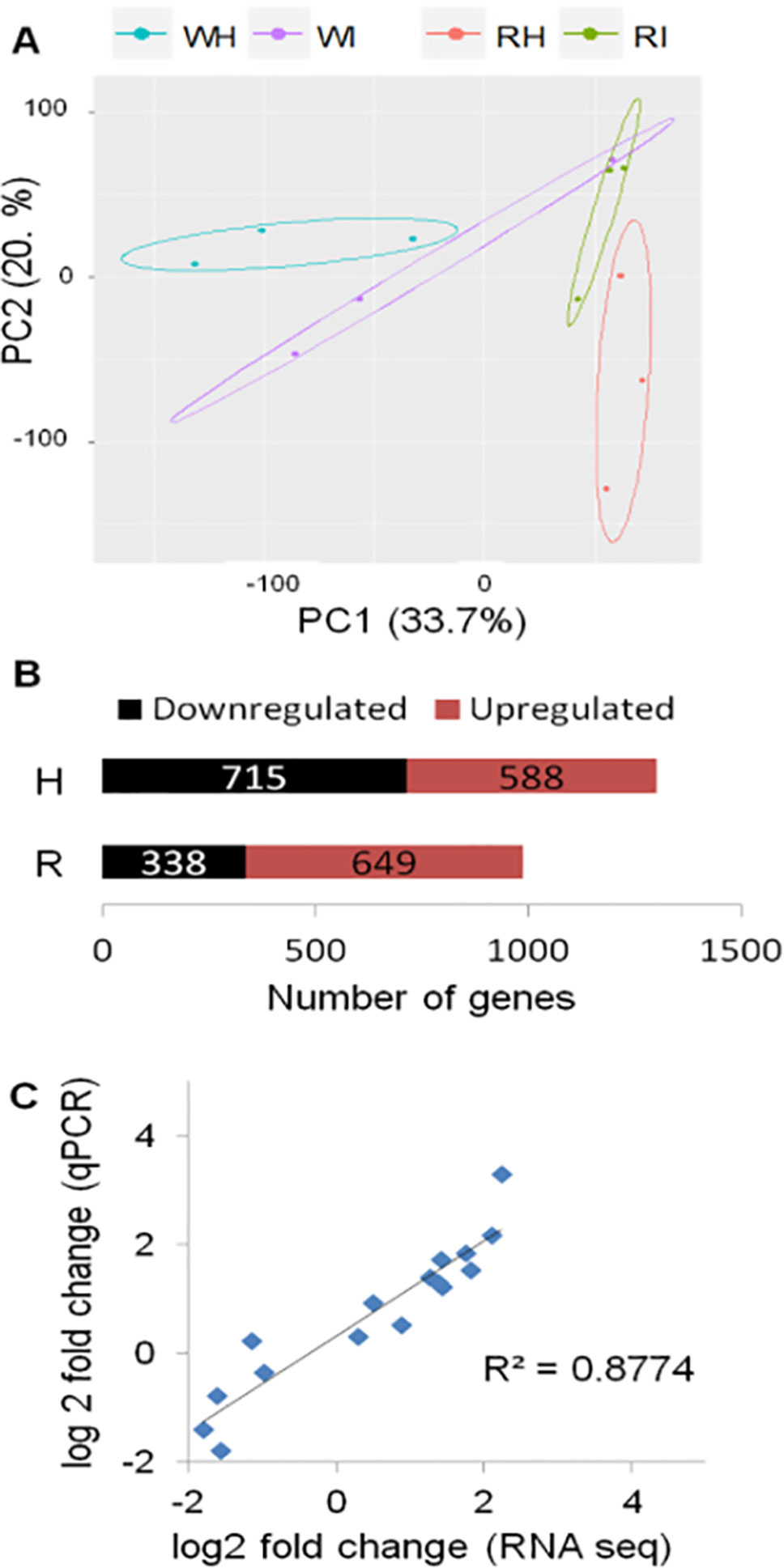
Figure 2 Global evaluation of the RNA-seq data and of the differentially expressed (DE) genes at 24 h postinoculation (hpi). (A) Principal component analysis displaying the biological variations of Fragaria vesca genes among samples. W, white fruit; R, red fruit; H, healthy (mock inoculated); I, Botrytis cinerea inoculated. Raw count data were used after precision weight was calculated by the voom method (Law et al., 2014). (B) Number of DE genes (P < 0.05, absolute fold change >1.74) upon B. cinerea inoculation at 24 hpi; downregulated genes (black) and upregulated genes (red). (C) Comparison of gene expression values of RNA-Seq and qRT-PCR: correlation of fold change values for eight F. vesca genes obtained by RNA-Seq and qRT-PCR.
The DEGs between Botrytis- versus mock-inoculated samples, in both white and in red fruits, were computed. Accordingly, in white fruits, the DEGs were 1,303, out of which 588 were upregulated, whereas in red fruits, 649 of the 987 DEGs were upregulated (Figure 2B; Supplemental Table 4). The fold-change values from RNA-Seq analysis were validated using quantitative real-time PCR (qRT-PCR) assay. The expression measurement of eight F. vesca genes (Supplemental Table 3) performed by qRT-PCR showed similar expression pattern to those of the RNA-Seq values (R2 > 0.88; Figure 2C).
DEGs were functionally categorized using MapMan (Supplemental Table 5). Among the total DEGs, only 149 were common to both growth stages (Supplemental Table 3). These common genes regulate a number of functions (Supplemental Figure 3). Interestingly, the common genes in protein and RNA modifications (pentatricopeptide repeat-containing proteins genes of the “Defense/Stress” class and genes of “RNA processing” classes) and biotic stress (genes of “Recognition and Signaling” class) functional classes were mostly downregulated in white fruits (Supplemental Figure 3).
With regard to the Botrytis transcriptome, the number of reads mapped on Botrytis genome from the white or red inoculated fruits was very low (Supplemental Table 2), prohibiting an in-depth analysis of fungal transcriptional activity during early infection.
Biological Responses of F. vesca Strawberry Fruits to B. cinerea Inoculation
Global analyses of infected fruits DEGs were computed to have an overview of transcriptional regulation in response to Botrytis infection (Figure 3). Enrichment of GO terms was evaluated to know the biological processes, molecular functions, and cellular components mostly affected by B. cinerea infection. From the GO terms corresponding to biological processes, “response to stress” and “response to stimulus” were overrepresented in both white and red fruits; additional overrepresented GO terms in white fruit were those related to “responses to biotic stimulus,” “response to abiotic stimulus,” and “carbohydrate metabolic process,” whereas in ripe fruits, the “cellular protein metabolic process” functional class was the most represented one (Figure 3A; Supplemental Tables 6, 7). As depicted in Figure 3A, differently from ripe fruits, in white fruits, downregulated genes dominated “response to stimulus” and “response to stress” GO terms, and among the genes those involved in hypersensitive response, such as TMV N resistance (Whitham et al., 1994; Marathe et al., 2002), were prominent (Supplemental Table 7). On the other hand, in red fruits, besides the category “response to stimulus,” upregulated genes dominated functional categories linked to pathogen defense, such as “receptor activity,” “transporter activity,” and “membrane” (Figure 3A).
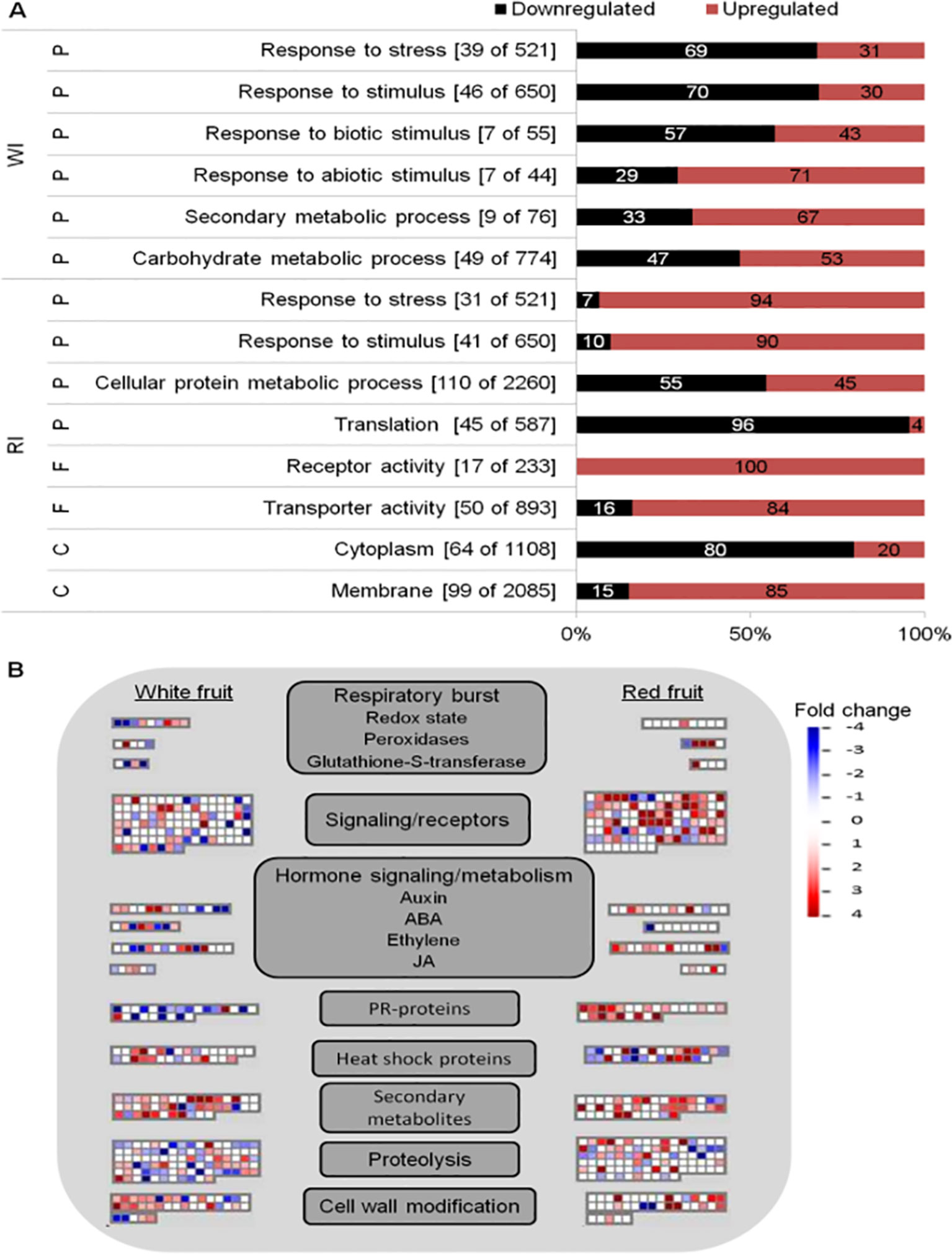
Figure 3 Functional enrichment and biotic stress overview of the differentially expressed (DE) genes of Fragaria vesca in Botrytis cinerea inoculated white and red fruits, 24 h postinoculation. (A) Functionally enriched classes in the DE genes of white inoculated (WI) and red inoculated (RI) fruits, using AgriGO analysis tool. The number of DE genes out of the total F. vesca genes in each category is shown in parenthesis. The proportion of down- and upregulated genes within a category is represented by black and red bars, respectively. C, cellular component; F, molecular function; P, biological process. (B) Overview of biotic stress changes in infected fruits, visualized by MapMan. Up- and downregulated genes are shown in red and blue, respectively. The scale bar displays fold change values; fold change value “0” is given for genes which are not DE. ABA, abscisic acid; JA, jasmonic acid.
The biotic stress pathway of the infected fruits was visualized via the MapMan tool (Thimm et al., 2004). Comparing the white with the red fruit transcriptomic response, a clear difference in biotic stress gene response is apparent, in both expression level and identity of genes with functions varying from recognition to defense responses to the pathogen (Figure 3B). Moreover, the number of differentially regulated genes (mainly upregulated) in “secondary metabolism” and “cell wall modification” was higher in white than in red fruits, but the opposite was true for signal transduction and proteolysis (Figure 3B). In addition,the defense response appeared stronger at the ripening stage, as PR coding genes were more represented and upregulated in red than white fruits (Figure 3B). However, it should be noted that the genes categorized as PR proteins encoding are TMV N proteins (Supplemental Table 5), according to the MapMan annotation used—F. vesca (Fvesca_226) mapping. However, these TMV N proteins are not considered members of the PR protein family (van Loon et al., 2006; Sudisha et al., 2012; Sinha et al., 2014).
Besides regulating developmental processes, phytohormones are involved in plant defense-signaling pathways (Bari and Jones, 2009). As expected, the involvement of hormonal signaling in Botrytis–strawberry fruit interaction was shown in the MapMan visualization. The involvement of auxin, ABA, ethylene (ET), and jasmonic acid (JA) phytohormones was highlighted (Figure 3B). Genes involved in their biosynthesis or signaling, such as 1-aminocyclopropane-1-carboxylate (ACC) oxidase, ethylene response transcription factors, allene oxide cyclase (AOC), 12-oxophytodienoate (OPDA) reductase, lipoxygenase, 9-cis-epoxycarotenoid dioxygenase (NCED), and zeaxanthin epoxidase (ZEP) were differentially regulated (Supplemental Table 5). Auxin, ABA, ET, and JA have been reported to play a central role in plant defense against B. cinerea (Audenaert et al., 2002; El Oirdi et al., 2011; Osorio et al., 2011).
Pathogen Perception and Signal Transduction Genes Are Modulated
Recognition is one of the earliest events in plant–pathogen interaction, before triggering other cascades of the defense response. Membrane-localized receptor-like kinases (RLKs) mediate cell wall–plasma membrane cross-communication, helping plants to respond to various stimuli, an essential prerequisite for pathogen recognition and activation of defense responses (Brutus et al., 2010; Singh and Zimmerli, 2013; Burdiak et al., 2015). In total, about 60 membrane-localized RLK genes were differentially expressed in white and red fruits; only three RLK genes (17154, 22229, and 24962) were common for both fruit growth stages (Table 1 and Supplemental Table 8). More than 70% of the differentially expressed RLK genes encode for proteins containing a leucine-rich repeat domain. The other RLK genes comprised cysteine-rich receptor kinases, lectin receptor kinases, and wall-associated receptor kinases (WAK) domains. Almost all RLK genes were upregulated in red fruits and only half of them in the white ones. It is known that repression of RLK genes that negatively regulate plant basal defense could lead to a reinforcement of the defense response (Delteil et al., 2016). Likewise, in the white fruit, Cysteine-rich RLK28 and Cysteine-rich RLK29 genes (17514 and 21516), involved in cell-death-mediated resistance (Yadeta et al., 2017), were downregulated (Table 1). Unexpectedly, RLK genes involved in immune response to necrotrophs, such as Somatic embryogenesis receptor kinases (16992 and 26498) and ERECTA genes (19894) (Godiard et al., 2003; Llorente et al., 2005; Heese et al., 2007), were repressed (Table 1). With regard to WAK genes, WAK9 and WAK14 in red fruits (34321 and 15062) and WAK4 in white fruits (34339) were upregulated (Table 1). Although reports on the involvement of WAK4 in plant defense are scanty (Lally et al., 2001), in white fruit, its induction was the highest (fourfold) among the RLKs. WAKs are known to serve as a sensor for monitoring cell wall integrity (Kohorn, 2015).
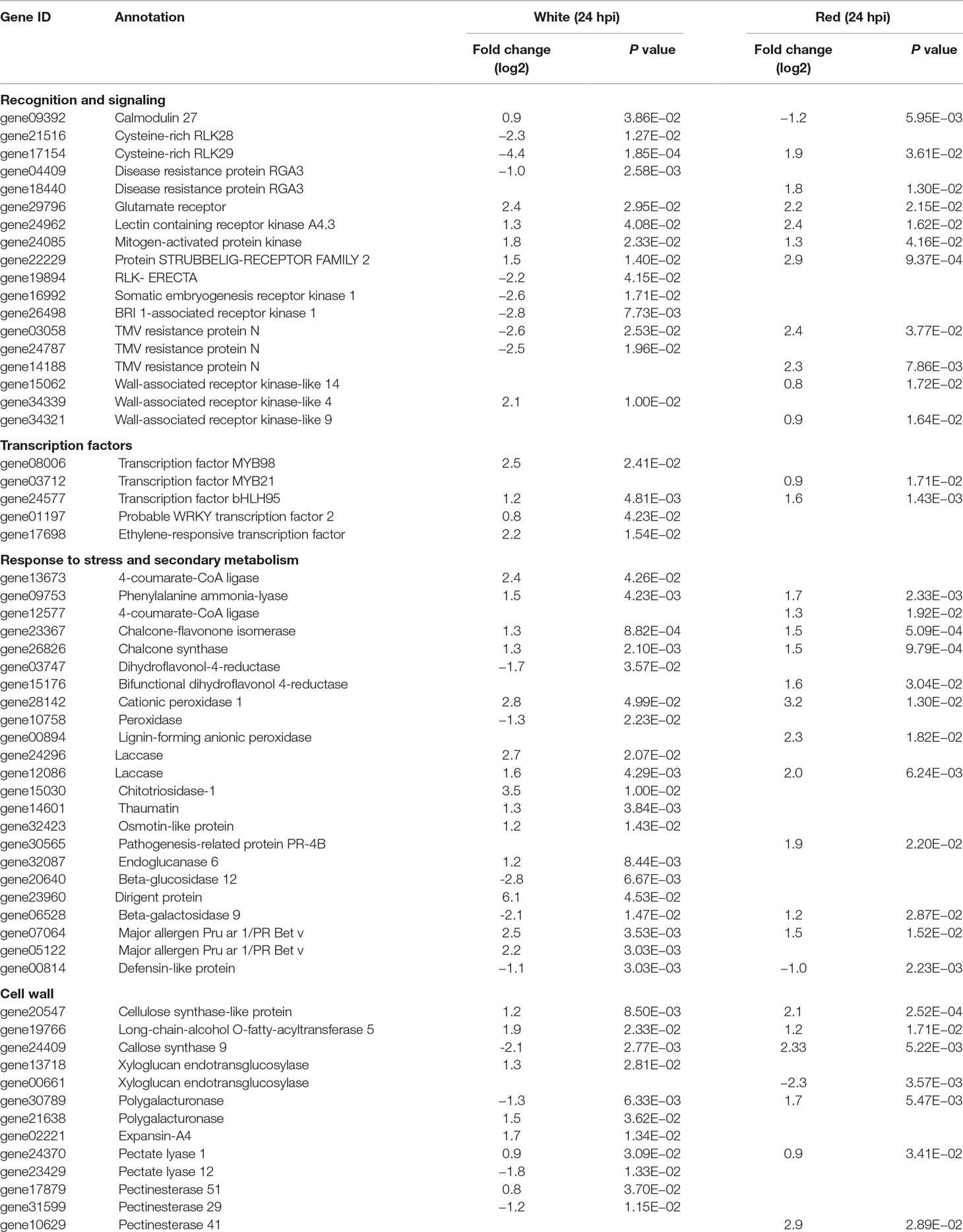
Table 1 Selected Botrytis-induced genes in Fragaria vesca fruits at 24 hpi with P ≤ 0.05 and absolute fold change (log2) of ≥0.8.
The infection of Botrytis affected the regulation of 25 genes putatively encoding for heat shock proteins (HSP) (Supplemental Table 8). HSPs can act as molecular chaperones to control functionality of plasma-membrane-resident receptors and intracellular resistance (R) proteins against pathogens (for review, see Park and Seo, 2015). In F. × ananassa strawberry leaves, HSP showed a high response to Colletotrichum fragariae infection (Fang et al., 2012). In our case, more than 10 HSP genes were upregulated, and in the red fruit, the upregulation level of some of these HSP genes was among the highest: about 34-fold increase for 07771 gene and 23-fold for 20877 gene (Supplemental Table 8).
The signal from cell membrane is downstreamed to cytoplasm via serine/threonine-protein kinase receptor (SRK) and further relayed by mitogen-activated protein kinases (MAPKs) and calcium-dependent protein kinase (CDPK) for transcriptional reprogramming needed in defense (Afzal et al., 2008; Boudsocq et al., 2010). No SRK or CDPK gene was transcriptionally altered in white fruits, but there were three (30638, 30648, and 30753) and two (17503 and 18254) genes, respectively, upregulated in red fruits (Supplemental Table 8), suggesting that in white fruits, downstream signal transduction between membrane-bound receptor kinases and MAPKs was regulated. Of the eight MAPKs DEG, only gene 24085 was common to both fruit-ripening stages, and unlike in white fruits, in red fruits, all of the MAPKs were highly upregulated (Supplemental Table 8). This might be due to the much higher disease pressure on red than white fruits, as can be seen in Figure 1. Some of these genes were found differentially regulated at 24 hpi in susceptible F. × ananassa ripe fruit inoculated with B. cinerea (Xiong et al., 2018), suggesting the existence of a similar signaling cascade to transduce immune signaling in ripe woodland strawberry upon B. cinerea infection.
On the other hand, surprisingly, Botrytis infection stimulated the expression of a number of R genes with a nucleotide binding site leucine-rich repeat (NBS–LRR) and of the pentatricopeptide (PPR) repeat-containing protein types, both in white and red fruits (Table 1 and Supplemental Table 5). This was quite novel because, with the exception of a few R proteins such as Arabidopsis resistance to Leptosphaeria maculans 3 (RLM3) and wheat TaRcR1, known to be associated to necrotroph pathogens (Staal et al., 2008; Zhu et al., 2017), R proteins in general are not associated with a defense response to necrotrophs. On the contrary, in the interaction of host with many biotrophic and hemibiotrophic pathogens, the pathogen effectors cause the induction of R genes to mediate effector-triggered host immunity (Jones and Dangl, 2006). However, in some interactions with necrotrophs, the toxins produced by the pathogen induce R genes to promote infection (Lorang et al., 2007; Wang et al., 2007; Faris et al., 2010). R-gene-mediated susceptibility to necrotrophs is envisaged as ectopic expression of hypersensitive response, which could allow the pathogen to access an initial growth substrate (Laluk and Mengiste, 2010; Mengiste, 2012).
PPR proteins are largely known for their role in modulating RNA-binding proteins, which mediates the expression of genes involved in photosynthesis, respiration, plant development, and environmental responses (Nakamura et al., 2012; Barkan and Small, 2014). There are also many PPR proteins that have been identified to function in defense against necrotrophic pathogens (Laluk et al., 2011; Xing et al., 2018). In this study, 47 genes putatively annotated for PPR proteins were differentially regulated (Supplemental Table 5), suggesting their involvement in F. vesca strawberry–Botrytis interaction. With regard to NBS–LRR, 45 genes putatively annotated for TMV N resistance, recognition of Peronspora parasitica (RPP), and resistance gene analogues (RGAs) were transcriptionally altered both in white and red fruits (Table 1 and Supplemental Table 8). Almost all of these NBS–LRR genes were upregulated in red fruits and downregulated in white ones. In particular, the members of the TMV N gene family were largely represented with several members differently distributed in the two transcriptomes. Of these genes, 03058, 31994, and 16534 were shared by the two ripening stages, but like all the others members, they were oppositely regulated by the pathogen infection (Supplemental Table 8).
TMV N and RPP genes are known to be involved in a hypersensitive response at the site of pathogen entry (Whitham et al., 1994; Cooley et al., 2000; Marathe et al., 2002; Takahashi et al., 2002; Bent and Mackey, 2007). It is also known that the hypersensitive response facilitates the colonization of necrotroph pathogens such as B. cinerea (Govrin and Levine, 2000). Studies show that B. cinerea elicits the expression of hypersensitivity-related genes in its hosts to facilitate infection (Govrin et al., 2006; Rossi et al., 2011); however, with regard to TMV, RPP, and RGA genes, there is no report that the fungus manipulates these genes to establish infection. Nonetheless, their upregulation in susceptible ripe fruits and suppression in unripe white fruits suggest that they might assist Botrytis colonization of the ripe fruits.
In addition, TMV N components of the NBS–LRR repertoire are known to function through signaling cascades that involve ubiquitination for protein degradation (Marathe et al., 2002).
The functional category “protein degradation” was one of the most represented in the DEGs: 61 in white and 42 in red fruits; and about one-third of which are ubiquitin genes where E3 ubiquitin ligases dominated (Figure 3B and Supplemental Table 5). Ubiquitins are a diverse family of proteins involved in posttranslation modification with different biological roles, including plant immunity (Zeng et al., 2006; Cheng and Li, 2012; Li et al., 2016). Many of these ubiquitin ligases have not yet been characterized in Fragaria species. However, it is known that E3 ubiquitin ligases are associated with pathogen and abiotic stress responses (Luo et al., 2010; Zhou and Zeng, 2017). The intricate regulatory network of E3 ligases might be involved in determining the different response of strawberry white and red fruits. Further studies are needed to clarify their regulatory mechanisms influencing defense pathways and immunity responses during strawberry–Botrytis interaction.
Phytohormone Biosynthesis and Metabolism
Genes involved in the biosynthesis and metabolism of phytohormones, except salicylic acid (SA), were differentially regulated, following B. cinerea inoculation (Supplemental Table 5). According to the putative functions of the F. vesca DEGs, ACC-oxidase, AOC, OPDA-reductase, and lipoxygenase genes involved in the important steps of ET and JA biosynthesis were upregulated at both fruit ripening stages. F. × ananassa, achenes of red strawberry fruits were reported to produce ET at a low concentration (Iannetta et al., 2006), while fruits were found responsive to ET when applied externally (Villarreal et al., 2010). In these studies, fruits that received ET externally were prone to B. cinerea. In this regard, the pathogen itself could trigger the production of ET in host tissue. A transcriptomic study on tomato suggested that B. cinerea would induce the expression of genes involved in ripening to favor its development (Cantu et al., 2009). ET and JA are usually associated with defense against necrotrophic pathogens, but their pathways can also functionally interact with SA signaling pathway to fine-tune plant defense (Kunkel and Brooks, 2002; Beckers and Spoel, 2006). However, no SA marker gene was differentially modulated in both ripening stages.
Apart from ET and JA, Botrytis infection affected the expression level of genes involved in the biosynthesis and metabolism of auxin, ABA, and gibberrelic acid, more in white fruits than in red ones (Supplemental Table 5). Auxins have mainly been implicated as key regulators for growth and fruit ripening in strawberry (Aharoni et al., 2002; Mezzetti et al., 2004); however, there is evidence that this growth regulator is also involved in the strawberry defense response (Osorio et al., 2011). Most related DEGs that encode auxin-induced and responsive proteins were downregulated in the resistant white fruit (as low as −42-fold change; Supplemental Table 5). Consistent to this, decrease in auxin content as well as enhanced expression of some auxin-repressed genes in fruit of transgenic F. vesca FaPE1 lines were found to be correlated with B. cinerea resistance (Osorio et al., 2011). ABA also showed a stronger response to infection in white fruits compared to red fruits (Supplemental Table 5). According to the RNA-Seq data, NCED1 and ZEP genes involved in ABA biosynthesis were upregulated but only in white fruit (Supplemental Table 5). ABA is known to promote strawberry ripening, also through the upregulation of WRKY genes (Jia et al., 2011; Li et al., 2019), and to be involved in defense responses to pathogens, where its deficiency made tomato and Arabidopsis more resistant to B. cinerea through its effects on reactive oxygen intermediates and cell wall stiffening (Audenaert et al., 2002; AbuQamar et al., 2006; Asselbergh et al., 2007; Curvers et al., 2010). Moreover, in F. × ananassa strawberry, different WRKY transcription factors were recently shown to be upregulated by ABA. Here, the transcription level of FaNCED1 genes were congruent with fruit ABA content, promoting ripening (Jia et al., 2011). Interestingly, two NECD1-encoding genes (31335 and 30616), which could be taken as a “susceptible factor,” were upregulated in the resistant white fruits only. It is known that B. cinerea synthesizes plant hormone analogues and elicitors (Chagué et al., 2002; Siewers et al., 2004; Siewers et al., 2006) able to induce hormone biosynthesis pathways in order to affect the host’s susceptibility by altering the hormone balance in infected host tissues (Sharon et al., 2004; El Oirdi et al., 2011).
Overall, such complex interplay among the phytohormones was also observed during B. cinerea infection of grapevine inflorescence as well as ripe and unripe tomato fruits (Blanco-Ulate et al., 2013; Haile et al., 2017). This suggests that modulation of phytohormones plays a role in regulating the interaction between strawberry fruit and B. cinerea, most likely in a ripening-stage specific manner. Nevertheless, it is difficult to conclude whether the combined changes in phytohormone-related processes promoted or inhibited immunity in F. vesca since much of the DEGs are involved in the biosynthesis pathway.
Defense-Related Genes of White and Red Fruits Are Differentially Regulated
Following recognition of Botrytis infection and queued signaling cascades, the expression level of a number of defense-related genes was altered both in white and red fruits, including the related transcription factors (TF) (Supplemental Table 5). In this experiment, the expression levels of more than 100 genes putatively encoding TFs belonging to different families were modulated in both white and red fruits (Table 1 and Supplemental Table 9). Besides TFs regulating hormone biosynthesis, bHLH, MYB, WRKY, and zinc fingers TF domains were the predominant ones. TFs play a role in biotic stress by regulating genes involved in innate immunity, hormone signaling pathways, and phytoalexin synthesis (Mzid et al., 2007; Gao et al., 2011; Höll et al., 2013). For example, ectopic expression of grapevine VvWRKY2 in tobacco enhances resistance to necrotrophic fungi such as B. cinerea (Mzid et al., 2007), and MYB98 regulates the activation of genes required for defense (Punwani et al., 2008). In line with this, genes putatively annotated for these two TFs (01197 for WRKY2 and 08006 for MYB98) were upregulated only in white fruit.
With respect to pathogenesis-related proteins (PRs), 28 putative PR genes were differentially regulated, mainly in white fruits as shown in Supplemental Table 10. Additionally, out of the 17 categories of PR protein families (van Loon et al., 2006; Sudisha et al., 2012; Sinha et al., 2014), genes belonging to 9 of them were transcriptionally altered. For genes like chitinase (15030), the upregulation was more than 10-fold in white fruits. In the case of β-1,3-glucanase, two genes in white fruits and one in red fruits were upregulated following Botrytis infection. Glucanases and chitinases are the most abundant classes of strawberry PR genes having hydrolytic activity (reviewed in Amil-Ruiz et al., 2011). Moreover, osmotin and thaumatin PRs were also induced in white fruits only. These proteins belong to PR-5 protein family, well known to inhibit fungal growth including B. cinerea (Monteiro et al., 2003; González et al., 2017). Notably, we found defensin and lipid transfer proteins (belonging to PR-12 and 14 protein family, in respective order), functioning as antimicrobial peptides (Garcia-Olmedo et al., 1995; Ganz, 2003; Nanni et al., 2014), downregulated in white fruits.
Other PR proteins whose expression was altered due to Botrytis infection were those homolog to the major allergen pru ar 1 protein, a PR-10 protein. Among five upregulated genes encoding for major allergens pru ar 1 proteins (Supplemental Table 10), only one gene (07064) was common to white and red fruits, while the others were upregulated in white fruits. PR-10 proteins are known to be induced in F. × ananassa strawberry fruits challenged by B. cinerea and Colletotrichum acutatum (Guidarelli et al., 2011; Xiong et al., 2018) as well as in leaves infected with Podosphaera aphanis (Jambagi and Dunwell, 2015) and in roots infected with Phytophthora cactorum (Toljamo et al., 2016). These proteins are important for defense against pathogens in strawberry, and they are expressed faster and/or stronger in resistant genotypes or resistant fruit phenological stages (unripe ones) compared to susceptible genotypes or susceptible fruit stages (Guidarelli et al., 2011; González et al., 2013). PR-10 proteins bind to different biologically important ligands, including sterols and flavonoids, and in strawberry, they were observed to interact with metabolic intermediates of flavonoid biosynthesis to regulate the biosynthesis pathway (Koistinen et al., 2005; Zubini et al., 2009; Muñoz et al., 2010; Casañal et al., 2013). This entails that the upregulation of genes encoding these proteins in response to Botrytis infection might occur to enhance flavonoid biosynthesis. The expression profiles of TFs and PR-protein genes suggest that defense against B. cinerea infection was elicited earlier in white fruits.
Secondary Metabolites Biosyntheses Pathways Are Triggered in Infected Fruits
In response to Botrytis infection, a number of genes involved in the synthesis of secondary metabolites, in particular those in flavonoid biosynthesis, were differentially regulated (Supplemental Table 5). Genes involved in terpenoid biosynthesis, according to KEGG Pathway database (Kanehisa and Goto, 2000), were also induced. In white fruits, three genes putatively encoding geranylgeranyl transferase, gibberellin 3-beta-dioxygenase, and ent-copalyl diphosphate synthase (12147, 14841, and 20295, respectively), which are involved in di- and tetra-terpenoid biosynthesis, were upregulated. Conversely, in red fruits, one gene (12609) putatively encoding (−)-germacrene D synthase, which catalyzes the first committed step in the synthesis of sesquiterpenoids, and two genes (18138 and 31674) putatively encoding carotenoid isomerase and phytoene synthase, respectively, involved in tetra-terpenoid biosynthesis, were upregulated. Isoprenoids appear to be part of the general defense responses in strawberry leaves and fruits, as genes involved in their biosynthesis were also induced during Colletotrichum spp. and P. cactorum infections (Hirai et al., 2000; Guidarelli et al., 2011; Toljamo et al., 2016).
The upregulation of the major allergen pru ar 1 genes suggests the active involvement of a flavonoid biosynthesis pathway in the interaction between strawberry fruit and B. cinerea. As a result, the expression status of genes involved in the biosynthesis pathway was thoroughly investigated (Figure 4). As depicted in Figure 4, except for cinnamic acid 4-hydroxylase (C4H), genes encoding key enzymes required for flavonoid biosynthesis pathway were differentially altered, at least in one of the ripening stages, albeit on a different scale. Previous studies have shown that flavonoids play an important role in the defense response to pathogen attack. In grapevine flowers, flavonoid concentrations increased following B. cinerea inoculation as pathogen response mechanism (Haile et al., 2017). Similarly, in strawberry, an increased concentration of proanthocyanidins around the penetration site of B. cinerea on immature fruits was reported to keep the pathogen under quiescence (Jersch et al., 1989). Flavan-3-ol derived compounds also seem to be involved in pathogen resistance in strawberry (Yamamoto et al., 2000; Puhl and Treutter, 2008; Nagpala et al., 2016; Toljamo et al., 2016). When white and red F. × ananassa strawberry fruits were challenged with B. cinerea and C. acutatum, a relatively higher accumulation of flavan-3-ols (catechin, procyanidin B1, and procyanidin B3) was observed in less susceptible white fruits with respect to red ones (Nagpala et al., 2016). Interestingly, in our study, gene encoding leucoanthocyanidin reductase (LAR), which converts leucoanthocyanidins to flavan-3-ols, was upregulated only in white fruits (Figure 4). Since Botrytis progress in white fruit was inconspicuous (Figure 1A), flavonoid polyphenols could play a role in determining the susceptibility of strawberry fruit to B. cinerea.
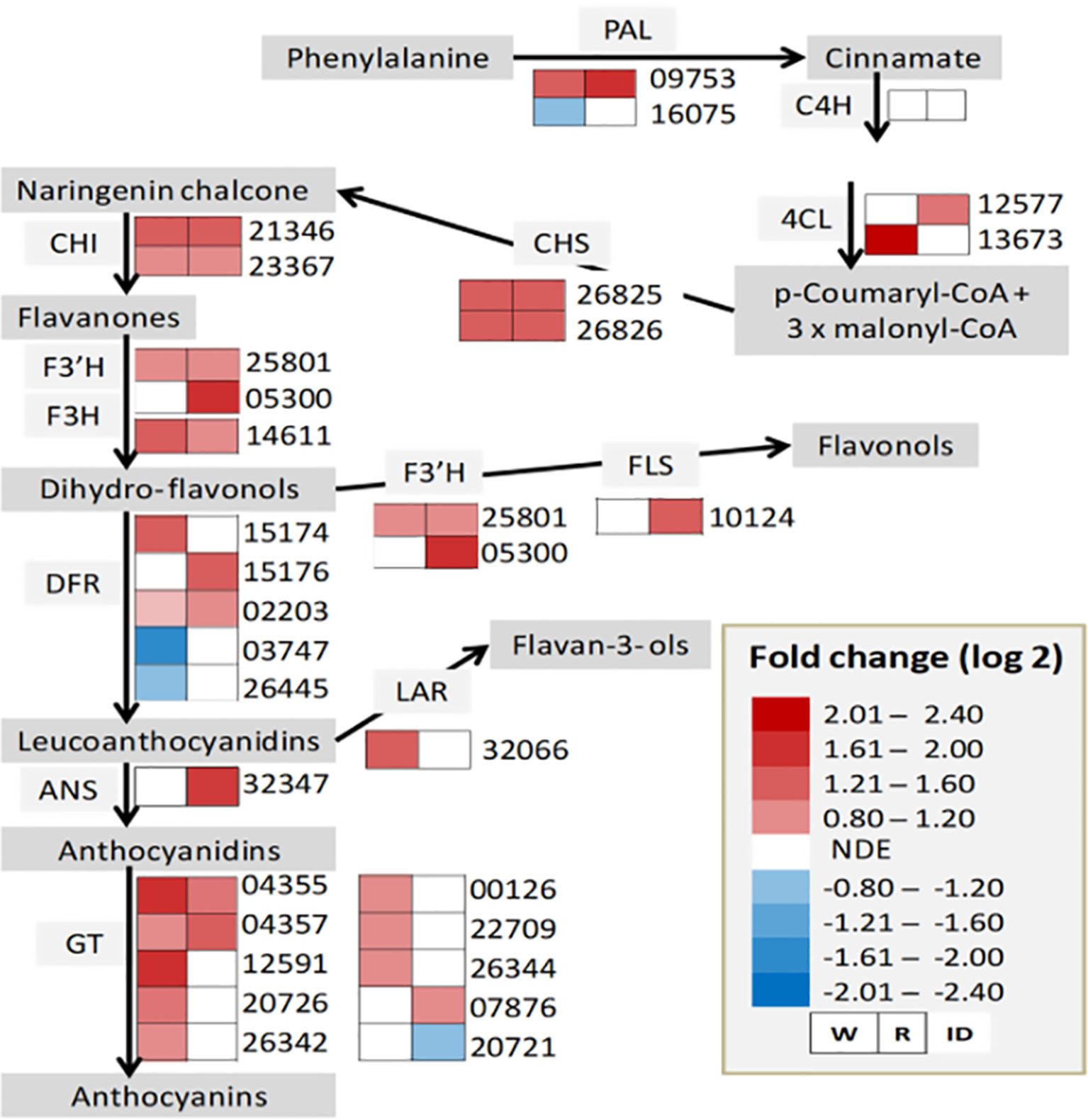
Figure 4 Flavonoid biosynthesis pathway in Botrytis cinerea inoculated Fragaria vesca fruits at 24 h postinoculation. Heatmaps of gene expression (from RNA-Seq result) in inoculated white (W) and red (R) fruits. ANS, anthocyanidin synthase; C4H, cinnamic acid 4-hydroxylase; CHI, chalcone isomerase; CHS, chalcone synthase; DFR, dihydroflavanol 4-reductase; F3’H, flavonoid 3’-monooxygenase; F3H, flavanone 3-hydroxylase; FLS, flavonol synthase GT, glycosyltransferase; LAR, leucoanthocyanidin reductase; PAL, phenylalanine ammonia-lyase; 4CL, 4-coumarate-coenzyme A ligase. NDE, not differentially expressed. Genes such as 02203, 05300, and 10124 are not in the DE genes list, but included for the completeness of the biosynthesis pathway. The expression profile for 02203 in W—log2 FC: 0.63, P value: 0.034; for 05300 in R—log2 FC: 2.0, P–value: 0.069; and for 10124 in R—log2 FC: 1.6, P–value: 0.053.
Cell Wall Modification of Strawberry Fruits Following B. cinerea Infection
The first obstacles that plant pathogens stumble upon are the cuticle and cell wall, the forefront of the plant defense system. Upon contact with a pathogen, the defense system undergoes changes to physically and chemically hinder the penetration via antimicrobial enzymes and secondary metabolites to interfere with the biology of the intruder (reviewed in Malinovsky et al., 2014). In the Botrytis-induced strawberry trancriptome, 56 genes involved in cell wall biosynthesis or modification were differentially regulated (Table 1 and Supplemental Table 11). Endogenous cell wall modifying/degrading proteins were prominent, especially in white fruits. For example, genes encoding expansins, known to endogenously influence cell wall extensibility and necrotroph susceptibility (AbuQamar, 2014), were unexpectedly upregulated in white fruits (Supplemental Table 11). The upregulation of genes encoding enzymes endogenously degrading cell wall such as endoglucanase, rhamnogalacturonate lyase, xyloglucanas endotransglucosylase, polygalacturonase, pectate lyase, and pectinesterase (Supplemental Table 11) during pathogen attacks appears counterintuitive. Nevertheless, for most of the enzymes, downregulation of few redundant genes was also observed. For example, pectinesterases are involved both in cell wall loosening, by making polygalacturonans accessible to degradation by polygalacturonases, and in cell wall strengthening, by increasing the availability of polygalacturonan to Ca2+ binding (Micheli, 2001).Therefore, the induction and suppression of pectinesterases and polygalacturonases suggest that, in white fruits, cell wall stiffening and cell expansion and separation occur as simultaneous processes to counteract the pathogen growth while allowing for cell expansion, since the white fruit itself was also in its active growth phase. On the other hand, genes encoding glucosidase, which degrades cellulose and hemicellulose (Gilbert, 2010; Blanco-Ulate et al., 2014), and galactosidase, a ripening-related cell wall softening enzyme (Nunan et al., 2001), were switched off in infected white fruits.
From the gene expression profiles, processes as the oxidative burst mediated cross-linking of cell wall through germin-like proteins and extensin interaction (Bradley et al., 1992; Godfrey et al., 2007) and callose apposition are suppressed in white fruits, (Supplemental Table 11). On the other hand, both in white and red fruit transcriptome, an increased expression (up to fivefold) of genes involved in cutin and wax biosynthesis was found, as shown by the fivefold induction of gene 03099, which encodes for protein wax 2, involved in cuticle membrane and wax production (Chen et al., 2003). Hemicelluloses and cellulose biosynthesis was also triggered as indicated by the upregulation of 06827, 10606, and 32285 genes and cellulose synthase genes, but mainly in white fruits. Similarly, genes encoding peroxidases and laccases which polymerize monolignols into lignin and are critical for cell wall fortification were also upregulated (Supplemental Table 11). In Arabidopsis, reduced cellulose synthesis leads to ectopic lignin synthesis (Cano-Delgado et al., 2003), implying a compensatory cell wall integrity maintenance mechanism when the integrity is perturbed (Hamann and Denness, 2011; Hamann, 2012). Probably, the suppressed callose synthesis observed in white fruits might be compensated by an enhanced lignin synthesis. In this regard, hemicelluloses are known to play a major role in cell wall toughening by interacting with cellulose and lignin (Scheller and Ulvskov, 2010) to challenge pathogen intrusion. Similarly, the deposition of lignin or lignin-like phenolic polymers in the cell wall has been implicated in plant defense, arresting advancing pathogens and/or limiting their progress (Menden et al., 2007; Miedes et al., 2014; Kelloniemi et al., 2015; Haile et al., 2017). Besides acting as a physical barrier to pathogen invasion, the process of lignin biosynthesis pathway, phenylpropanoid pathway, could also produce compounds with a defense role. Interestingly, the most upregulated (66-fold) gene in infected white fruit was a dirigent protein gene (23960) (Table 1). Dirigent proteins control the stereochemical coupling of monolignol radicals during the biosynthesis of lignans (Davin et al., 1997), which are polyphenols reported to have antipathogenic activity (Davin and Lewis, 1992; Akiyama et al., 2007; Li et al., 2017). Dirigent proteins are often found to be transcriptionally activated under biotic stress conditions (Borges et al., 2013; Thamil Arasan et al., 2013; Haile et al., 2017). Genes encoding dirigent proteins were found upregulated during B. cinerea infection of grapevine flowers. In this work, cell wall reinforcement was addressed as one of the possible mechanisms by which the grapevine flower arrests the advancement of Botrytis (Haile et al., 2017). The strong upregulation of dirigent protein and cell wall modification gene expression in white unripe strawberry fruit suggests a similar mechanism of F. vesca to arrest B. cinerea intrusion.
Conclusion
The transcriptome analysis of strawberry fruits at different ripening stages provided a comprehensive picture of the response of white and red fruits to B. cinerea infection. Genes involved in defense response pathways, from perceiving to combating the pathogen, were differentially regulated within 24 hpi in both white and red fruits, albeit quantitatively different. Yet, Botrytis progress was visible on red fruits only after 48 hpi. Indeed, it has been shown that plant tissues can trigger defense mechanisms even when overwhelmed by pathogen attack (Kelloniemi et al., 2015; Nagpala et al., 2016; Xiong et al., 2018). Even though reprogramming of red fruit transcriptome towards defense was observed, it did not lead to a switch off of cell-death-causing membrane-localized RLK and NBS–LRR genes, such as Cysteine-rich RLK29 and a number of TMV N, RPP, and RGA genes, which could play a role as susceptibility factors. On the other hand, in white fruits, the downregulation of these membrane-localized RLK and NBS–LRR susceptibility genes, the upregulation of chitinase, osmotin, and thaumatin PR protein genes and the triggering of the biosynthesis flavonoid polyphenols are likely correlated with the competence of the white fruit to block B. cinerea growth. With regard to fruit cell wall fortification, crucial to stop Botrytis growth, the RNA-Seq result suggests that, in white fruits responding to B. cinerea infection, hemicellulose-, cellulose-, and lignin-based fortification of the cell wall plays an important role. Nevertheless, callose deposition and oxidative burst mediated cross-linking of the cell wall based on germin-like proteins and extension, another important cell wall stiffening mechanism against Botrytis progress (Kelloniemi et al., 2015; Haile et al., 2017), appeared not to be involved in white fruits.
Data Availability
The datasets generated for this study can be found in National Center for Biotechnology Information (NCBI), BioProject accession code PRJNA530684.
Author Contributions
ZH wrote the manuscript and followed the research. EN-D performed experiments and wrote the first draft of the MS. EN-D is the first co-author together with ZH. MM, PS, and KE made all the bioinformatic analysis from raw RNA-seq to gene mapping, identification, and statistical analysis. LZ initiated the experimental work. CM helped with project supervision and with MS writing. EB supervised the project and MS writing and provided financial support.
Funding
Institutional funding was received from ‘Alma Mater Studiorum Università degli studi diBologna’.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2019.01131/full#supplementary-material
References
AbuQamar, S. (2014). Expansins: cell wall remodeling proteins with a potential function in plant defense. J. Plant Biochem. Physiol. 2, e118. doi: 10.4172/2329-9029.1000e118
AbuQamar, S. F., Moustafa, K., Tran, L.-S. P. (2016). Omics’ and plant responses to Botrytis cinerea. Front. Plant Sci. 7, 1658. doi: 10.3389/fpls.2016.01658
AbuQamar, S., Chen, X., Dhawan, R., Bluhm, B., Salmeron, J., Lam, S., et al. (2006). Expression profiling and mutant analysis reveals complex regulatory networks involved in Arabidopsis response to Botrytis infection. Plant J. 48, 28–44. doi: 10.1111/j.1365-313X.2006.02849.x
Afzal, A. J., Wood, A. J., Lightfoot, D. A. (2008). Plant receptor-like serine threonine kinases: roles in signaling and plant defense. Mol. Plant Microbe Interact. 21 (5), 507–517. doi: 10.1094/MPMI-21-5-0507
Aharoni, A., Keizer, L. C. P., Van Den Broeck, H. C., Blanco-Portales, R., Munoz-Blanco, J., Bois, et al. (2002). Novel insight into vascular, stress, and auxin-dependent and -independent gene expression programs in strawberry, a non-climacteric fruit. Plant Physiol. 129, 1019–1031. doi: 10.1104/pp.003558
Akiyama, K., Yamauchi, S., Nakato, T., Maruyama, M., Sugahara, T., Kishida, T. (2007). Antifungal activity of tetra-substituted tetrahy-drofuran lignan, (–)-virgatusin, and its structure–activity relationship. Biosci. Biotechnol. Biochem. 71, 1028–1035. doi: 10.1271/bbb.60696
Alkan, N., Friedlander, G., Ment, D., Prusky, D., Fluhr, R. (2015). Simultaneous transcriptome analysis of Colletotrichum gloeosporioides and tomato fruit pathosystem reveals novel fungal pathogenicity and fruit defense strategies. New Phytol. 205 (2), 801–815. doi: 10.1111/nph.13087
Amil-Ruiz, F., Blanco-Portales, R., Muñoz-Blanco, J., Caballero, J. L. (2011). The strawberry plant defense mechanism: a molecular review. Plant Cell Physiol. 52 (11), 1873–1903. doi: 10.1093/pcp/pcr136
Applied Biosystems. (1997 updated 2001). Relative quantification of gene expression: ABI Prism 7700 Sequence Detection System. Appl. Biosyst. User Bull. 2.
Asselbergh, B., Curvers, K., Franca, S. C., Audenaert, K., Vuylsteke, M., Van Breusegem, F., et al. (2007). Resistance to Botrytis cinerea in sitiens, an abscisic acid-deficient tomato mutant, involves timely production of hydrogen peroxide and cell wall modifications in the epidermis. Plant Physiol. 144, 1863–1877. doi: 10.1104/pp.107.099226
Audenaert, K., De Meyer, G. B., Hofte, M. M. (2002). Abscisic acid determines basal susceptibility of tomato to Botrytis cinerea and suppresses salicylic acid-dependent signaling mechanisms. Plant Physiol. 128, 491–501. doi: 10.1104/pp.010605
Barkan, A., Small, I. (2014). Pentatricopeptide repeat proteins in plants. Annu. Rev. Plant Biol. 65, 415–442. doi: 10.1146/annurev-arplant-050213-040159
Bari, R., Jones, J. (2009). Role of plant hormones in plant defence responses. Plant Mol. Biol. 69, 473–488. doi: 10.1007/s11103-008-9435-0
Beckers, G. J., Spoel, S. H. (2006). Fine-tuning plant defence signaling: salicylate versus jasmonate. Plant Biol. (Stuttg.) 8, 1–10. doi: 10.1055/s-2005-872705
Bent, A. F., Mackey, D. (2007). Elicitors, effectors, and R genes: the new paradigm and a lifetime supply of questions. Annu. Rev. Phytopathol. 45, 399–436. doi: 10.1146/annurev.phyto.45.062806.094427
Blanco-Ulate, B., Morales-Cruz, A., Amrine, K. C., Labavitch, J. M., Powell, A. L., Cantu, D. (2014). Genome-wide transcriptional profiling of Botrytis cinerea genes targeting plant cell walls during infections of different hosts. Front. Plant Sci. 5, 435. doi: 10.3389/fpls.2014.00435
Blanco-Ulate, B., Vincenti, E., Powell, A. L. T., Cantu, D. (2013). Tomato transcriptome and mutant analyses suggest a role for plant stress hormones in the interaction between fruit and Botrytis cinerea. Front. Plant Sci. 4, 142. doi: 10.3389/fpls.2013.00142
Borges, A. F., Ferreira, R. B., Monteiro, S. (2013). Transcriptomic changes following the compatible interaction Vitis vinifera–Erysiphe necator. Paving the way towards an enantioselective role in plant defence modulation. Plant Physiol. Biochem. 68, 71–80. doi: 10.1016/j.plaphy.2013.03.024
Boudsocq, M., Willmann, M. R., Mccormack, M., Lee, H., Shan, L., He, P., et al. (2010). Differential innate immune signalling via Ca(2+) sensor protein kinases. Nature 464, 418–422. doi: 10.1038/nature08794
Bradley, D. J., Kjellbom, P., Lamb, C. J. (1992). Elicitor- and wound-induced oxidative cross-linking of a proline-rich plant cell wall protein: a novel, rapid defense response. Cell 70, 21–30. doi: 10.1016/0092-8674(92)90530-P
Bristow, P. R., McNicol, R. J., Williamson, B. (1986). Infection of strawberry flowers by Botrytis cinerea and its relevance to grey mould development. Ann. Appl. Biol. 109 (3), 545–554. doi: 10.1111/j.1744-7348.1986.tb03211.x
Brutus, A., Sicilia, F., Macone, A., Cervone, F., De Lorenzo, G. (2010). A domain swap approach reveals a role of the plant wall-associated kinase 1 (WAK1) as a receptor of oligogalacturonides. Proc. Natl. Acad. Sci. 107, 9452–9457. doi: 10.1073/pnas.1000675107
Bulger, M. A., Ellis, M. A., Madden, L. V. (1987). Influence of temperature and wetness duration on infection of strawberry flowers by Botrytis cinerea and disease incidence of fruit originating from infected flowers. Phytopathology 77 (8), 1225–1230. doi: 10.1094/Phyto-77-1225
Burdiak, P., Rusaczonek, A., Witon, D., Glow, D., Karpinski, S. (2015). Cysteine-rich receptor-like kinase CRK5 as aregulator of growth, development and ultraviolet radiation responses in Arabidopsis thaliana. J. Exp. Bot. 66 (11), 3325–3337. doi: 10.1093/jxb/erv143
Cano-Delgado, A., Penfield, S., Smith, C., Catley, M., Bevan, M. (2003). Reduced cellulose synthesis invokes lignification and defense responses in Arabidopsis thaliana. Plant J. 34, 351–362. doi: 10.1046/j.1365-313X.2003.01729.x
Cantu, D., Blanco-Ulate, B., Yang, L., Labavitch, J. M., Bennett, A. B., Powell, A. L. (2009). Ripening-regulated susceptibility of tomato fruit to Botrytis cinerea requires NOR but not RIN or ethylene. Plant Physiol. 150, 1434–1449. doi: 10.1104/pp.109.138701
Casañal, A., Zander, U., Muñoz, C., Dupeux, F., Luque, I., Botella, M. A., et al. (2013). The strawberry pathogenesis related 10 (PR-10) Fra a proteins control flavonoid biosynthesis by binding to metabolic intermediates. J. Biol. Chem. 288, 35322–35332. doi: 10.1074/jbc.M113.501528
Chagué, V., Elad, Y., Barakat, R., Tudzynski, P., Sharon, A., and (2002). Ethylene biosynthesis in Botrytis cinerea. FEMS Microbiol. Ecol. 40, 143–149. doi: 10.1016/S0168-6496(02)00222-2
Chen, X., Goodwin, S. M., Boroff, V. L., Liu, X., Jenks, M. A. (2003). Cloning and characterization of the WAX2 gene of Arabidopsis involved in cuticle membrane and wax production. Plant Cell 15, 1170–1185. doi: 10.1105/tpc.010926
Cheng, Y. T., Li, X. (2012). Ubiquitination in NB-LRR-mediated immunity. Curr. Opin. Plant Biol. 15, 392–399. doi: 10.1016/j.pbi.2012.03.014
Conesa, A., Götz, S., García-Gómez, J. M., Terol, J., Talón, M., Robles, M. (2005). Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21 (18), 3674–3676. doi: 10.1093/bioinformatics/bti610
Cooley, M. B., Pathirana, S., Wu, H. J., Kachroo, P., Klessig, D. F. (2000). Members of the Arabidopsis HRT/RPP8 family of resistance genes confer resistance to both viral and oomycete pathogens. Plant Cell 12, 663–676. doi: 10.1105/tpc.12.5.663
Curvers, K., Seifi, H., Mouille, G., de Rycke, R., Asselbergh, B., Van Hecke, A., et al. (2010). Abscisic acid deficiency causes changes in cuticle permeability and pectin composition that influence tomato resistance to Botrytis cinerea. Plant Physiol. 154, 847–860. doi: 10.1104/pp.110.158972
Davin, L. B., Lewis, N. G. (1992). Phenylpropanoid metabolism: biosynthesis of monolignols, lignans and neolignans, lignins and suberins. Rec. Adv. Phytochem. 26, 325–375. doi: 10.1007/978-1-4615-3430-3_11
Davin, L. B., Wang, H. B., Crowell, A. L., Bedgar, D. L., Martin, D. M., Sarkanen, S., et al. (1997). Stereoselective bimolecular phenoxy radical coupling by an auxiliary (dirigent) protein without an active center. Science 275, 362–366. doi: 10.1126/science.275.5298.362
Dean, R., Van Kan, J. A., Pretorius, Z. A., Hammond-Kosack, K. E., Di Pietro, A., Spanu, P. D., et al. (2012). The Top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13 (4), 414–430. doi: 10.1111/j.1364-3703.2011.00783.x
Delteil, A., Gobbato, E., Cayrol, B., Estevan, J., Michel-Romiti, C., Dievart, A., et al. (2016). Several wall-associated kinases participate positively and negatively in basal defense against rice blast fungus. BMC Plant Biol. 16, 17. doi: 10.1186/s12870-016-0711-x
Du, Z., Zhou, X., Ling, Y., Zhang, Z., Su, Z. (2010). AgriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res. 38, W64–W70. doi: 10.1093/nar/gkq310
El Oirdi, M., El Rahman, T., Rigano, L., El Hadrami, A., Rodriguez, M. C., Daayf, F., et al. (2011). Botrytis cinerea manipulates the antagonistic effects between immune pathways to promote disease development in tomato. Plant Cell 23, 2405–2421. doi: 10.1105/tpc.111.083394
Elad, Y., Vivier, M., Fillinger, S., (2016). “Botrytis, the good, the bad and the ugly,” in Botrytis the fungus, the pathogen and its management in agricultural systems. (Cham: Springer), 1–15. doi: 10.1007/978-3-319-23371-0_1
Fang, X., Chen, W., Xin, Y., Zhang, H., Yan, Ch., Yu, H., et al. (2012). Proteomic analysis of strawberry leaves infected with Colletotrichum fragariae. J. Proteomics 75, 4074–4090. doi: 10.1016/j.jprot.2012.05.022
Faris, J. D., Zhang, Z., Lu, H., Lu, S., Reddy, L., Cloutier, S., et al. (2010). A unique wheat disease resistance-like gene governs effector-triggered susceptibility to necrotrophic pathogens. Proc. Natl. Acad. Sci. U.S.A. 107, 13544–13549. doi: 10.1073/pnas.1004090107
Feliziani, E.G., Romanazzi. (2016). Postharvest decay of strawberry fruit: etiology, epidemiology, and disease management. J. Berry Res. 6, 47–63. doi: 10.3233/JBR-150113
Ganz, T. (2003). Defensins: antimicrobial peptides of innate immunity. Nat. Rev. 3, 710–720. doi: 10.1038/nri1180
Gao, Q.-M., Venugopal, S., Navarre, D., Kachroo, A. (2011). Low oleic acid-derived repression of jasmonic acid-inducible defense responses requires the WRKY50 and WRKY51 proteins. Plant Physiol. 155, 464–476. doi: 10.1104/pp.110.166876
Garcia-Olmedo, F., Molina, A., Segura, A., Moreno, M. (1995). The defensive role of nonspecific lipid-transfer proteins in plants. Trends Microbiol. 3, 72–74. doi: 10.1016/S0966-842X(00)88879-4
Gilbert, H. J. (2010). The biochemistry and structural biology of plant cell wall deconstruction. Plant Physiol. 153, 444–455. doi: 10.1104/pp.110.156646
Godfrey, D., Able, A. J., Dry, I. B. (2007). Induction of a grapevine germin-like protein (VvGLP3) gene is closely linked to the site of Erysiphe necator infection: a possible role in defense? Mol. Plant Microbe Interact. 20, 1112–1125. doi: 10.1094/MPMI-20-9-1112
Godiard, L., Sauviac, L., Torii, K. U., Grenon, O., Mangin, B., Grimsley, N. H., et al. (2003). ERECTA, an LRR receptor like kinase protein controlling development pleiotropically affects resistance to bacterial wilt. Plant J. 36, 353–365. doi: 10.1046/j.1365-313X.2003.01877.x
González, G., Fuentes, L., Moya-León, M. A., Sandoval, C., Herrera, R. (2013). Characterization of two PR genes from Fragaria chiloensis in response to Botrytis cinerea infection: a comparison with Fragaria× ananassa. Physiol. Mol. Plant Pathol. 82, 73–80. doi: 10.1016/j.pmpp.2013.02.001
González, M., Brito, N., González, C. (2017). The Botrytis cinerea elicitor protein BcIEB1 interacts with the tobacco PR5-family protein osmotin and protects the fungus against its antifungal activity. New Phytol. 118, 11–14. doi: 10.1111/nph.14588
Govrin, E. M., Levine, A. (2000). The hypersensitive response facilitates plant infection by the necrotrophic pathogen Botrytis cinerea. Curr. Biol. 10, 751–757. doi: 10.1016/S0960-9822(00)00560-1
Govrin, E. M., Rachmilevitch, S., Tiwari, B. S., Solomon, M., Levine, A. (2006). An elicitor from Botrytis cinerea induces the hypersensitive response in Arabidopsis thaliana and other plants and promotes the gray mold disease. Phytopathology 96, 299–307. doi: 10.1094/PHYTO-96-0299
Guidarelli, M., Carbone, F., Mourgues, F., Perrotta, G., Rosati, C., Bertolini, P., et al. (2011). Colletotrichum acutatum interactions with unripe and ripe strawberry fruits and differential responses at histological and transcriptional levels. Plant Pathol. 60, 685–697. doi: 10.1111/j.1365-3059.2010.02423.x
Gusberti, M., Gessler, C., Broggini, G. A. (2013). RNA-Seq analysis reveals candidate genes for ontogenic resistance in Malus–Venturia pathosystem. PLoS One 8 (11), e78457. doi: 10.1371/journal.pone.0078457
Haile, Z. M., Pilati, S., Sonego, P., Malacarne, G., Vrhovsek, U., Engelen, K., et al. (2017). Molecular analysis of the early interaction between the grapevine flower and Botrytis cinerea reveals that prompt activation of specific host pathways leads to fungus quiescence. Plant Cell Environ. 40, 1409–1428. doi: 10.1111/pce.12937
Hamann, T. (2012). Plant cell wall integrity maintenance as an essential component of biotic stress response mechanisms. Front. Plant Sci. 3, 77. doi: 10.3389/fpls.2012.00077
Hamann, T., Denness, L. (2011). Cell wall integrity maintenance in plants: lessons to be learned from yeast? Plant Signal. Behav. 6, 1–5. doi: 10.4161/psb.6.11.17782
Heese, A., Hann, D. R., Gimenez-Ibanez, S., Jones, A. M. E., He, K., Li, J., et al. (2007). The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc. Natl. Acad. Sci. 104, 12217–12222. doi: 10.1073/pnas.0705306104
Hirai, N., Sugie, M., Wada, M., Lahlou, E. H., Kamo, T., Yoshida, R., et al. (2000). Triterpene phytoalexins from strawberry fruit. Biosci. Biotechnol. Biochem. 64, 1707–1712. doi: 10.1271/bbb.64.1707
Höll, J., Vannozzi, A., Czemmel, S., D’Onofrio, C., Walker, A. R., Rausch, T., et al. (2013). The R2R3-MYB transcription factors MYB14 and MYB15 regulate stilbene biosynthesis in Vitis vinifera. Plant Cell 25, 4135–4149. doi: 10.1105/tpc.113.117127
Iannetta, P. P. M., Laarhoven, L.-J., Medina-Escobar, N., James, E. K., McManus, M. T., Davies, H. V., et al. (2006). Ethylene and carbon di-oxide production by developing strawberries show a correlative pattern that is indicative of ripening climacteric fruit. Physiol. Plant 127, 247–259. doi: 10.1111/j.1399-3054.2006.00656.x
Jambagi, S., Dunwell, J. (2015). Global transcriptome analysis and identification of differentially expressed genes after infection of Fragaria vesca with Powdery Mildew (Podosphaeraaphanis). Transcriptomics 3, 1. doi: 10.4172/2329-8936.1000106
Jersch, S., Scherer, C., Huth, G., Schlösser, E. (1989). Proanthocyanidins as basis for quiescence of Botrytis cinerea in immature strawberry fruits. Z. Pflanzenkrankh Pflanzenschutz 96, 365–378.
Jia, H., Chai, Y., Li, C., Lu, D., Luo, J., Qin, L., et al. (2011). Abscisic acid plays an important role in the regulation of strawberry fruit ripening. Plant Physiol. 157, 188–199. doi: 10.1104/pp.111.177311
Jones, J. D., Dangl, J. L. (2006). The plant immune system. Nature 444, 323–329. doi: 10.1038/nature05286
Kanehisa, M., Goto, S. (2000). KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. doi: 10.1093/nar/28.1.27
Kelloniemi, J., Trouvelot, S., Heloir, M. C., Simon, A., Dalmais, B., Frettinger, P., et al. (2015). Analysis of the molecular dialogue between gray mold (Botrytis cinerea) and grapevine (Vitis vinifera) reveals a clear shift in defense mechanisms during berry ripening. Mol. Plant Microbe Interact. 28, 1167–1180. doi: 10.1094/MPMI-02-15-0039-R
Kohorn, B. D. (2015). The state of cell wall pectin monitored by wall associated kinases: a model. Plant Signal. Behav. 10, e1035854. doi: 10.1080/15592324.2015.1035854
Koistinen, K. M., Soininen, P., Venäläinen, T. A., Häyrinen, J., Laatikainen, R., Peräkylä, M., et al. (2005). Birch PR-10c interacts with several biologically important ligands. Phytochemistry 66, 2524–2533. doi: 10.1016/j.phytochem.2005.09.007
Kunkel, B. N., Brooks, D. M. (2002). Cross talk between signaling pathways in pathogen defence. Curr. Opin. Plant Biol. 5, 325–331. doi: 10.1016/S1369-5266(02)00275-3
Lally, D., Ingmire, P., Tong, H. Y., He, Z. H. (2001). Antisense expression of a cell wall- associated protein kinase, WAK4, inhibits cell elongation and alters morphology. Plant Cell 13, 1317–1331. doi: 10.2307/3871298
Laluk, K., Mengiste, T. (2010). Necrotroph attacks on plants: wanton destruction or covert extortion? Arabidopsis Book 8, 2053−2068. doi: 10.1199/tab.0136
Laluk, K., Abuqamar, S., Mengiste, T. (2011). The Arabidopsis mitochondria-localized pentatricopeptide repeat protein PGN functions in defense against necrotrophic fungi and abiotic stress tolerance. Plant Physiol. 156, 2053–2068. doi: 10.1104/pp.111.177501
Law, C. W., Chen, Y., Shi, W., Smyth, G. K. (2014). Voom: precision weights unlock linear model analysis tools for RNA-Seq read counts. Genome Biol. 15 (2), R29. doi: 10.1186/gb-2014-15-2-r29
Liao, X., Li, M., Liu, B., Yan, M., Yu, X., Zi, H., et al. (2018). Interlinked regulatory loops of ABA catabolism and biosynthesis coordinate fruit growth and ripening in woodland strawberry. Proc. Natl. Acad. Sci. 115, 49. doi: 10.1073/pnas.1812575115
Li, J., Chai, Q. Y., Liu, C. H. (2016). The ubiquitin system: a critical regulator of innate immunity and pathogen–host interactions. Cell. Mol. Immunol. 13, 560–576. doi: 10.1038/cmi.2016.40
Li, C., Jia, H., Chai, Y., Shen, Y. (2011). Abscisic acid perception and signaling transduction in strawberry: a model for non-climacteric fruit ripening. Plant Signal. Behav. 6 (12), 1950–1953. doi: 10.4161/psb.6.12.18024
Li, D., Mou, W., Xia, R., Li, L., Zawora, C., Ying, T., et al. (2019). Integrated analysis of high-throughput sequencing data shows abscisic acid-responsive genes and miRNAs in strawberry receptacle fruit ripening. Hortic. Res. 6, 26. doi: 10.1038/s41438-018-0100-8
Li, N., Zhao, M., Liu, T., Dong, L., Cheng, Q., Wu, J., et al. (2017). A novel soybean dirigent gene GmDIR22 contributes to promotion of lignan biosynthesis and enhances resistance to Phytophthora sojae. Front. Plant Sci. 8, 1185. doi: 10.3389/fpls.2017.01185
Liao, Y., Smyth, G. K., Shi, W. (2013). The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108. doi: 10.1093/nar/gkt214
Liao, Y., Smyth, G. K., Shi, W. (2014). FeatureCounts: an efficient general-purpose program for assigning sequence reads to genomic features. Bioinformatics 30 (7), 923–930. doi: 10.1093/bioinformatics/btt656
Llorente, F., Alonsi-Blanco, C., Sánchez-Rodriguez, C., Jorda, L., Molina, A. (2005). ERECTA receptor-like kinase and heterotrimeric G protein from Arabidopsis are required for resistance to the necrotrophic fungus Plectosphaerella cucumerina. Plant J. 43, 165–180. doi: 10.1111/j.1365-313X.2005.02440.x
Lopez-Gomez, R., Gomez-Lim, M. A. (1992). A method for extracting intact RNA from fruits rich in polysaccharides using ripe mango mesocarp. Hort. Science 27, 440–442. doi: 10.21273/HORTSCI.27.5.440
Lorang, J. M., Sweat, T. A., Wolpert, T. J. (2007). Plant disease susceptibility conferred by a “resistance” gene. Proc. Natl. Acad. Sci. U. S. A. 104, 14861–14866. doi: 10.1073/pnas.0702572104
Luo, H., Laluk, K., Lai, Z., Veronese, P., Song, F., Mengiste, T. (2010). The Arabidopsis Botrytis Susceptible1 Interactor defines a subclass of RING E3 ligases that regulate pathogen and stress responses. Plant Physiol. 154, 1766–1782. doi: 10.1104/pp.110.163915
Malinovsky, F. G., Fangel, J. U., Willats, W. G. T. (2014). The role of the cell wall in plant immunity. Front. Plant Sci. 5, 178. doi: 10.3389/fpls.2014.00178
Manning, K. (1993). “Soft fruit,” in Biochemistry of fruit ripening (Dordrecht: Springer), 347–377. doi: 10.1007/978-94-011-1584-1_12
Marathe, R., Anandalakshmi, R., Liu, Y., Dinesh-Kumar, S. P. (2002). The tobacco mosaic virus resistance gene, N. Mol. Plant Pathol. 3, 167–172. doi: 10.1046/j.1364-3703.2002.00110.x
Menden, B., Kohlhoff, M., Moer-schbacher, B. M.. (2007). Wheat cells accumulate asyringyl-rich lignin during the hypersensitive resistance response. Phytochemistry 68, 513–520. doi: 10.1016/j.phytochem.2006.11.011
Mengiste, T. (2012). Plant immunity to necrotrophs. Annu. Rev. Phytopathol. 50, 267–294. doi: 10.1146/annurev-phyto-081211-172955
Mezzetti, B., Landi, L., Pandolfini, T., Spena, A. (2004). The defH9-iaaM auxin-synthesizing gene increases plant fecundity and fruit production in strawberry and raspberry. BMC Biotechnol. 4, 4. doi: 10.1186/1472-6750-4-4
Micheli, F. (2001). Pectin methylesterases: cell wall enzymes with important roles in plant physiology. Trends Plant Sci. 6, 414–419. doi: 10.1016/S1360-1385(01)02045-3
Miedes, E., Vanholme, R., Boerjan, W., Molina, A. (2014). The role of the secondary cell wall in plant resistance to pathogens. Front. Plant Sci. 5, 358. doi: 10.3389/fpls.2014.00358
Montero, T. M., Moll, E. M., Esteban, R. M., Lupez-Andrèu, F. J. (1996). Quality attributes of strawberry during ripening. Sci. Hortic. 65 (4), 239–250. doi: 10.1016/0304-4238(96)00892-8
Monteiro, S., Barakat, M., Piçarra-Pereira, M. A., Teixeira, A. R., Ferreira, R. B. (2003). Osmotin and thaumatin from grape: a putative general defense mechanism against pathogenic fungi. Phytopathology 93 (12), 1505–1512. doi: 10.1094/PHYTO.2003.93.12.1505
Muñoz, C., Hoffmann, T., Escobar, N. M., Ludemann, F., Botella, M. A., Valpuesta, V., et al. (2010). The strawberry fruit Fra a allergen functions in flavonoid biosynthesis. Mol. Plant 3, 113–124. doi: 10.1093/mp/ssp087
Mzid, R., Marchive, C., Blancard, D., Deluc, L., Barrieu, F., Corio-Costet, M. F., et al. (2007). Overexpression of VvWRKY2 in tobacco enhances broad resistance to necrotrophic fungal pathogens. Physiol. Plant 131, 434–447. doi: 10.1111/j.1399-3054.2007.00975.x
Nagpala, E.vG., Guidarelli, M., Gasperotti, M., Masuero, D., Bertolini, P., Vrhovsek, U., et al. (2016). Polyphenols variation in fruits of the susceptible strawberry cultivar Alba during ripening and upon fungal pathogen interaction and possible involvement in unripe fruit tolerance. J. Agric. Food Chem. 64, 1869–1878. doi: 10.1021/acs.jafc.5b06005
Nakamura, T., Yagi, Y., Kobayashi, K. (2012). Mechanistic insight into pentatricopeptide repeat proteins as sequence-specific RNA-binding proteins for organellar RNAs in plants. Plant Cell Physiol. 53 (7), 1171–1179. doi: 10.1093/pcp/pcs069
Nanni, V., Schumacher, J., Giacomelli, L., Brazzale, D., Sbolci, L., Moser, C., et al. (2014). Vvamp2, a grapevine flower-specific defensin capable of inhibiting Botrytis cinerea growth: insights into its mode of action. Plant Pathol. 63, 899–910. doi: 10.1111/ppa.12170
Neri, F., Cappellin, L., Spadoni, A., Cameldi, I., Algarra Alarcon, A., Aprea, E., et al. (2015). Role of strawberry volatile organic compounds in the development of Botrytis cinerea infection. Plant Pathol. 64 (3), 709–717. doi: 10.1111/ppa.12287
Nunan, K. J., Davies, C., Robinson, S. P., Fincher, G. B. (2001). Expression patterns of cell wall-modifying enzymes during grape berry development. Planta 214, 257–264. doi: 10.1007/s004250100609
Osorio, S., Bombarely, A., Giavalisco, P., Usadel, B., Stephens, C., Araguez, et al. (2011). Demethylation of oligogalacturonides by FaPE1 in the fruits of the wild strawberry Fragaria vesca triggers metabolic and transcriptional changes associated with defence and development of the fruit. J. Exp. Bot. 62, 2855–2873. doi: 10.1093/jxb/erq465
Park, C. J., Seo, Y. S. (2015). Heat shock proteins: a review of the molecular chaperones for plant immunity. Plant Pathol. J. 31 (4), 323–333. doi: 10.5423/PPJ.RW.08.2015.0150
Payasi, A., Mishra, N. N., Chaves, A. L. S., Singh, R. (2009). Biochemistry of fruit softening: an overview. Physiol. Mol. Biol. Plants 15 (2), 103–113. doi: 10.1007/s12298-009-0012-z
Perkins-Veazie, P. (1995). Growth and ripening of strawberry fruit. Hortic. Rev. 17, 267–297. doi: 10.1002/9780470650585.ch8
Petrasch, S., Knapp, S. J., Van Kan, J. A., Blanco-Ulate, B. (2019). Grey mould of strawberry, a devastating disease caused by the ubiquitous necrotrophic fungal pathogen Botrytis cinerea. Mol. Plant Pathol. 6, 877–892. doi: 10.1111/mpp.12794
Porebski, S., Bailey, L. G., Baum, B. R. (1997). Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol. Biol. Rep. 15 (1), 8–15. doi: 10.1007/BF02772108
Prusky, D., Alkan, N., Mengiste, T., Fluhr, R. (2013). Quiescent and necrotrophic lifestyle choice during postharvest disease development. Annu. Rev. Phytopathol. 51, 155–176. doi: 10.1146/annurev-phyto-082712-102349
Puhl, I., Treutter, D. (2008). Ontogenetic variation of catechin biosynthesis as basis for infection and quiescence of Botrytis cinerea in developing strawberry fruits. J. Plant. Dis. Prot. 115, 247–251. doi: 10.1007/BF03356272
Punwani, J. A., Rabiger, D. S., Lloyd, A., Drews, G. N. (2008). The MYB98 subcircuit of the synergid gene regulatory network includes genes directly and indirectly regulated by MYB98. Plant J. 55, 406–414. doi: 10.1111/j.1365-313X.2008.03514.x
Rosli, H. G., Civello, P. M., Martinez, G. A. (2004). Changes in cell wall composition of three Fragaria× ananassa cultivars with different softening rate during ripening. Plant Physiol. Biochem. 42 (10), 823–831.s. doi: 10.1016/j.plaphy.2004.10.002
Rossi, F. R., Garriz, A., Marina, M., Romero, F. M., Gonzalez, M. E., Collad, I. G., et al. (2011). The sesquiterpene botrydial produced by Botrytis cinerea induces the hypersensitive response on plant tissues and its action is modulated by salicylic acid and jasmonic acid signaling. Mol. Plant Microbe Interact. 24, 888–896. doi: 10.1094/MPMI-10-10-0248
Scheller, H. V., Ulvskov, P. (2010). Hemicelluloses. Annu. Rev. Plant Biol. 61, 263–289. doi: 10.1146/annurev-arplant-042809-112315
Schwab, W., Raab, T. (2004). “Developmental changes during strawberry fruit ripening and physico-chemical changes during postharvest storage,” in Production practices and quality assessment of food crops. Eds. Dris, R., Jain, S. M. (Dordrecht: Springer).
Sham, A., Moustafa, K., Al-Shamisi, S., Alyan, S., Iratni, R., AbuQamar, S. (2017). Microarray analysis of Arabidopsis WRKY33 mutants in response to the necrotrophic fungus Botrytis cinerea. PLoS One 12 (2), e0172343. doi: 10.1371/journal.pone.0172343
Sharon, A., Elad, Y., Barakat, R., Tudzynski, P. (2004). “Phytohormones in Botrytis–plant interactions,” in Botrytis: Biology, pathology and control. Eds. Elad, Y., Williamson, B., Tudzynski, P., Delen, N.. (Dordrecht, the Netherlands: Kluwer Academic Publishers), 163–176. doi: 10.1007/978-1-4020-2626-3_10
Shulaev, V., Sargent, D. J., Crowhurst, R. N., Mockler, T. C., Folkerts, O., et al. (2011). The genome of woodland strawberry (Fragaria vesca). Nat. Genet. 43, 109–116. doi: 10.1038/ng.740
Siewers, V., Kokkelink, L., Smedsgaard, J., Tudzynski, P. (2006). Identification of an abscisic acid gene cluster in the grey mold Botrytis cinerea. Appl. Environ. Microbiol. 7, 4619–4626. doi: 10.1128/AEM.02919-05
Siewers, V., Smedsgaard, J., Tudzynski, P. (2004). The P450 mono-oxygenase BcABA1 is essential for abscisic acid biosynthesis in Botrytis cinerea. Appl. Environ. Microbiol. 70, 3868–3876. doi: 10.1128/AEM.70.7.3868-3876.2004
Singh, P., Zimmerli, L. (2013). Lectin receptor kinases in plant innate immunity. Front. Plant Sci. 4, 124. doi: 10.3389/fpls.2013.00124
Sinha, M., Singh, R. P., Kushwaha, G. S., Iqbal, N., Singh, A., Kaushik, S., et al. (2014). Current overview of allergens of plant pathogenesis related protein families. Sci. World J. 2014, 543195. doi: 10.1155/2014/543195
Smyth, G. K. (2004). Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 3. doi: 10.2202/1544-6115.1027
Staal, J., Kaliff, M., Dewaele, E., Persson, M., Dixelius, C. (2008). RLM3, a TIR domain encoding gene involved in broad-range immunity of Arabidopsis to necrotrophic fungal pathogens. Plant J. 55, 188–200. doi: 10.1111/j.1365-313X.2008.03503.x
Sudisha, J., Sharathchandra, R. G., Amruthesh, K. N., Kumar, A., Shetty, H. S. (2012). “Pathogenesis related proteins in plant defense response,” in Plant defence: biological control. Progress in biological control, vol. 12. Eds. Mérillon, J., Ramawat, K. (Dordrecht: Springer). doi: 10.1007/978-94-007-1933-0_17
Symons, G. M., Chua, Y. J., Ross, J. J., Quittenden, L. J., Davies, N. W., Reid, J. B. (2012). Hormonal changes during non-climacteric ripening in strawberry. J. Exp. Bot. 63 (13), 4741–4750. doi: 10.1093/jxb/ers147
Takahashi, H., Miller, J., Nozaki, Y., Takeda, M., Shah, J., Hase, S., et al. (2002). RCY1, an Arabidopsis thaliana RPP8/HRT family resistance gene, conferring resistance to cucumber mosaic virus requires salicylic acid, ethylene and a novel signal transduction mechanism. Plant J. 32, 655–667. doi: 10.1046/j.1365-313X.2002.01453.x
Terry, L. A., Joyce, D. C., Adikaram, N. K., Khambay, B. P. (2004). Pre-formed antifungal compounds in strawberry fruit and flower tissues. Postharvest Biol. Tec. 31 (2), 201–212. doi: 10.1016/j.postharvbio.2003.08.003
Thamil Arasan, S. K., Park, J. I., Ahmed, N. U., Jung, H. J., Hur, Y., Kang, K. K., et al. (2013). Characterization and expression analysis of dirigent family genes related to stresses in Brassica. Plant Physiol. Biochem. 67, 144–153. doi: 10.1016/j.plaphy.2013.02.030
Thimm, O., Bläsing, O., Gibon, Y., Nagel, A., Meyer, S., Krüger, P., et al. (2004). MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 37, 914–939. doi: 10.1111/j.1365-313X.2004.02016.x
Toljamo, A., Blande, D., Kärenlampi, S., Kokko, H. (2016). Reprogramming of Strawberry (Fragaria vesca) root transcriptome in response to Phytophthora cactorum. PLoS ONE 11 (8), e0161078. doi: 10.1371/journal.pone.0161078
van Kan, J. A. (2005). Infection strategies of Botrytis cinerea. Acta Hortic. 669, 77–90. doi: 10.17660/ActaHortic.2005.669.9
van Loon, L. C., Rep, M., Pieterse, C. M. J. (2006). Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 44, 135–162. doi: 10.1146/annurev.phyto.44.070505.143425
Villarreal, N. M., Bustamante, C. A., Civello, P. M., Martínez, G. A. (2010). Effect of ethylene and 1-MCP treatments on strawberry fruit ripening. J. Sci. Food Agric. 90 (4), 683–689. doi: 10.1002/jsfa.3868
Wang, W., Devoto, A., Turner, J. G., Xiao, S. (2007). Expression of the membrane-associated resistance protein RPW8 enhances basal defense against biotrophic pathogens. Mol. Plant Microbe Interact. 20, 966–976. doi: 10.1094/MPMI-20-8-0966
Whitham, S., Dinesh-Kumar, S. P., Choi, D., Hehl, R., Corr, C., Baker, B. (1994). The product of the tobacco mosaic virus resistance gene N: similarity to toll and the interleukin-1 receptor. Cell 78, 1101–1115. doi: 10.1016/0092-8674(94)90283-6
Xing, H., Fu, X., Yang, C., Tang, X., Guo, L., Li, C., et al. (2018). Genome-wide investigation of pentatricopeptide repeat gene family in poplar and their expression analysis in response to biotic and abiotic stresses. Sci. Rep. 8, 2817. doi: 10.1038/s41598-018-21269-1
Xiong, J. S., Zhu, H. Y., Bai, Y. B., Liu, H., Cheng, Z. M. (2018). RNA sequencing-based transcriptome analysis of mature strawberry fruit infected by necrotrophic fungal pathogen Botrytis cinerea. Physiol. Mol. Plant Pathol. 104, 77–85. doi: 10.1016/j.pmpp.2018.08.005
Yadeta, K. A., Elmore, J. M., Creer, A. Y., Feng, B., Franco, J. Y., Rufian, et al. (2017). A cysteine-rich protein kinase associates with a membrane immune complex and the cysteine residues are required for cell death. Plant Physiol. 173, 771–778. doi: 10.1104/pp.16.01404
Yamamoto, M., Nakatsuka, S., Otani, H., Kohmoto, K., Nishimura, S. (2000). (+)-Catechin acts as an infection inhibiting factor in strawberry leaf. Phytopathology 90, 595–600. doi: 10.1094/PHYTO.2000.90.6.595
Zeng, L. R., Vega-Sanchez, M. E., Zhu, T., Wang, G. L. (2006). Ubiquitination-mediated protein degradation and modification: an emerging theme in plant–microbe interactions. Cell Res. 16, 413–426. doi: 10.1038/sj.cr.7310053
Zhou, B., Zeng, L. (2017). Conventional and unconventional ubiquitination in plant immunity. Mol. Plant Pathol. 18, 1313–1330. doi: 10.1111/mpp.12521
Zhu, X., Lu, C., Du, L., Ye, X., Liu, X., Coules, A., et al. (2017). The wheat NB-LRR gene TaRCR1 is required for host defence response to the necrotrophic fungal pathogen Rhizoctonia cerealis. Plant Biotechnol. J. 15, 674–687. doi: 10.1111/pbi.12665
Zubini, P., Zambelli, B., Musiani, F., Ciurli, S., Bertolini, P., Baraldi, E. (2009). The RNA hydrolysis and the cytokinin binding activities of PR-10 proteins are differently performed by two isoforms of the Pru p 1 peach major allergen and are possibly functionally related. Plant Physiol. 150, 1235–1247. doi: 10.1104/pp.109.139543
Keywords: Botrytis cinerea, defense response, Fragaria vesca, fungal quiescence, transcriptomic analysis
Citation: Haile ZM, Nagpala-De Guzman EG, Moretto M, Sonego P, Engelen K, Zoli L, Moser C and Baraldi E (2019) Transcriptome Profiles of Strawberry (Fragaria vesca) Fruit Interacting With Botrytis cinerea at Different Ripening Stages. Front. Plant Sci. 10:1131. doi: 10.3389/fpls.2019.01131
Received: 30 March 2019; Accepted: 15 August 2019;
Published: 18 September 2019.
Edited by:
Vincenzo Lionetti, Sapienza University of Rome, ItalyReviewed by:
Stefan Petrasch, University of California, United StatesSynan F. AbuQamar, United Arab Emirates University, United Arab Emirates
Copyright © 2019 Haile, Nagpala-De Guzman, Moretto, Sonego, Engelen, Zoli, Moser and Baraldi. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Elena Baraldi, ZWxlbmEuYmFyYWxkaUB1bmliby5pdA==
†These authors have contributed equally to this work
 Zeraye Mehari Haile
Zeraye Mehari Haile Ellaine Grace Nagpala-De Guzman
Ellaine Grace Nagpala-De Guzman Marco Moretto
Marco Moretto Paolo Sonego
Paolo Sonego Kristof Engelen5
Kristof Engelen5 Claudio Moser
Claudio Moser Elena Baraldi
Elena Baraldi