- 1Center for Viticulture and Enology, College of Food Science and Nutritional Engineering, China Agricultural University, Beijing, China
- 2Key Laboratory of Viticulture and Enology, Ministry of Agriculture and Rural Affairs, Beijing, China
- 3Beijing Advanced Innovation Center for Tree Breeding by Molecular Design, Beijing Forestry University, Beijing, China
Proanthocyanidins (PAs) and anthocyanins are two vital groups of flavonoid compounds for grape berries and red wines. Several transcription factors (TFs) have been identified to be involved in regulating PA and anthocyanin biosynthesis in grape berries. However, research on TFs with different regulatory mechanisms for these two biosynthesis branches in grapes remains limited. In this study, we identified an R2R3-MYB TF, VviMYB86, whose spatiotemporal gene expression pattern in grape berries coincided well with PA accumulation but contrasted with anthocyanin synthesis. Both in vivo and in vitro experiments verified that VviMYB86 positively regulated PA biosynthesis, primarily by upregulating the expression of the two leucoanthocyanidin reductase (LAR) genes in the Arabidopsis protoplast system, as well as in VviMYB86-overexpressing grape callus cultured under 24 h of darkness. Moreover, VviMYB86 was observed to repress the anthocyanin biosynthesis branch in grapes by downregulating the transcript levels of VviANS and VviUFGT. Overall, VviMYB86 is indicated to have a broad effect on flavonoid synthesis in grape berries. The results of this study will help elucidate the regulatory mechanism governing the expression of the two LAR genes in grape berries and provide new insights into the regulation of PA and anthocyanin biosynthesis in grape berries.
Introduction
Flavonoids are an important group of secondary metabolites in plants; they include proanthocyanidins (PAs, also known as condensed tannins), anthocyanins, and flavonols, that largely accumulate in the berries, seeds, leaves, flowers, and bark of many plant species (Dixon et al., 2005). Grape (Vitis vinifera L.) is one of the most widely cultivated fruits worldwide and has important economic value. For grape berries and red wines, PAs and anthocyanins strongly contribute to the astringent taste and red coloration, respectively (Peters and Constabel, 2002; Vidal et al., 2003). Furthermore, both PAs and anthocyanins confer potential beneficial effects on human health due to their antioxidant capacities and radical scavenging functions (Cos et al., 2004; Butelli et al., 2008). Therefore, the biosynthesis and regulation of PAs and anthocyanins in grape berries have attracted considerable amounts of attention (Cavallini et al., 2015; Wei et al., 2020).
PAs, as well as their building blocks and anthocyanins, are synthesized via the flavonoid metabolic pathway, and they share most of the same steps (Supplementary Figure 1; Dixon et al., 2005; Li et al., 2016). Leucoanthocyanidins, which are the first branch point between the PA and anthocyanin biosynthesis pathways, can be converted into 2,3-trans-flavan-3-ols [such as (+)-catechin] by leucoanthocyanidin reductase (LAR) or can be oxidized to generate extremely unstable anthocyanidins by anthocyanidin synthase (ANS). Next, anthocyanidins are utilized to produce 2,3-cis-flavan-3-ols [such as (−)-epicatechin] by anthocyanidin reductase (ANR) or are employed by UDP-glucose: flavonoid-3-O-glucosyltransferase (UFGT) to generate stable anthocyanins (Xie et al., 2003).
LAR and ANR are two key enzymes involved in PA biosynthesis (Tanner et al., 2003; Xie et al., 2003). In grapes, ANR is primarily responsible for the total amount of PAs, and is encoded by the gene VviANR, which is expressed throughout early flower and berry development (Xie et al., 2003; Bogs et al., 2005). Unlike Arabidopsis, which lacks LAR in PA biosynthesis, grape possesses two LAR enzymes (VviLAR1 and VviLAR2) with similar functions—not only producing 2,3-trans-flavan-3-ols from leucoanthocyanidins but also controlling the degree of PA polymerization (Yu et al., 2019). However, the expression of the genes encoding VviLAR1 and VviLAR2 is modulated in a temporal- and tissue-specific manner in grape berries (Bogs et al., 2005). In grape berry skins and seeds, VviLAR2 is highly expressed during the second half of the PA biosynthesis phase, when VviLAR1 is at a low transcript level (Bogs et al., 2005). These results imply that these three PA branch-specific genes might be differentially regulated in grape berries.
Several major PA and anthocyanin biosynthesis genes, such as ANS, LAR, ANR, and UFGT, are strictly regulated at the transcription level, and one of the regulatory mechanisms involves transcription factors (TFs) that regulate the expression of corresponding structural genes (Xu et al., 2014). To date, multiple TFs belonging to the MYB, bHLH, WD, MADS, and WRKY families have been characterized to regulate PA and anthocyanin biosynthesis (Wei et al., 2020). Among these TFs, those from the MYB family have been studied most extensively. Several MYB TFs have been demonstrated to be positively involved in PA and/or anthocyanin synthesis in grape berries. PA synthesis is regulated specifically by VviMYBPA1, VviMYBPA2, and VviMYBPAR, which have been isolated through homologous cloning based on Arabidopsis PA-related genes (Bogs et al., 2007; Terrier et al., 2009; Koyama et al., 2014). In addition, target genes of the three MYB TFs in grape berries are VviLAR1 and VviANR, although the expression pattern of VviMYBPAR is more similar to that of VviLAR2 throughout berry development (Bogs et al., 2007; Terrier et al., 2009; Koyama et al., 2014). These results indicate that additional putative regulators that participate in PA synthesis may exist, primarily by controlling the expression of VviLAR2. With respect to anthocyanins, the TF VviMYBA1 directly controls the expression of VviUFGT in grape berries (Cutanda-Perez et al., 2009). The abovementioned MYB TFs all possess a single regulatory function in flavonoid biosynthesis of grapes—regulate either PA synthesis or anthocyanin synthesis. In contrast with these functionally specialized MYB TFs, some MYB TFs have a broad effect on flavonoid synthesis in planta. For instance, Populus tomentosa PtMYB86, Tartary buckwheat FtMYB15, and Freesia hybrida FhMYB5 can promote both anthocyanin and PA branches at the same time (Luo et al., 2018; Li et al., 2019; Wang et al., 2019). In grape berries, VviMYB5a and VviMYB5b are involved in both anthocyanin and PA synthesis, and their regulation of anthocyanins and PAs is positive (Deluc et al., 2006, 2008).
In addition to MYB activators, several R2R3-MYB TFs have been identified as negative regulators of the flavonoid pathway. Generally, R2R3-MYB repressors can be divided into two clades: AtMYB4-like and FaMYB1-like (Chen et al., 2019). The two clades of R2R3-MYB repressors show variations in their conserved element termed the A2 box or element 3, which is located in the R3 domain (Cavallini et al., 2015). In the AtMYB4-like clade, the sequence of this element is DNEI, whereas the sequence is DNEV in the FaMYB1-like clade (Chen et al., 2019). In addition to the difference in amino acid in the conserved element, these two R2R3-MYB-type repressors are also different in their functions. AtMYB4-type R2R3-MYB repressors negatively affect the gene expressions of lignin and general phenylpropanoid pathway, whereas FaMYB1-type R2R3-MYB repressors downregulate the gene expressions of anthocyanin and/or PA biosynthetic pathway (Cavallini et al., 2015; Jun et al., 2015; Zhou et al., 2019). Unlike most flavonoid R2R3-MYB activators, R2R3-MYB repressors seem to repress more than one end product. Furthermore, MYB TFs with opposite effects on different biosynthesis branches of flavonoids have been observed in other plant species recently, such as Narcissus tazetta NtMYB3, Medicago truncatula MtPAR, and Camellia sinensis CsMYB5a. All of these TFs promote PA biosynthesis but repress other flavonoid biosynthesis branches, such as flavonols, isoflavones, and anthocyanins (Li et al., 2016; Jiang et al., 2018; Anwar et al., 2019). In grape berries, TFs that have opposite regulatory mechanisms to PA and anthocyanin pathways are being identified.
Genetic factors and several abiotic factors such as light, temperature, and water can also influence PA and anthocyanin biosynthesis in planta (Downey et al., 2006). Among these environmental stimuli, light is the major factor associated with flavonoid accumulation and the composition of grapes. The results of several field experiments have confirmed that exposure of grape bunches to light can enhance the expression levels of corresponding structural and regulatory genes involved in the PA and anthocyanin biosynthesis pathways, as well as promote PA and anthocyanin accumulation in berries (Fujita et al., 2007; Matus et al., 2009; Lee, 2017). Recently, our research group investigated the effects of different light conditions on phenolic metabolism in grape berries by applying integrated transcriptomics and pathway-level metabolomics (Sun et al., 2017, 2019). The results of these studies demonstrate that the transcript abundances of VviLAR1 and VviANR have a significant positive correlation with the altered accumulation of PAs in light exclusion-treated grape berries, whereas the expression of VviLAR2 is not sensitive to changes in light (Additional ‘file’ 8: Supplementary Table 6 in Sun et al., 2019). Moreover, to systematically investigate additional putative regulators that may regulate flavonoid biosynthesis in grape berries, a genome-wide co-expression analysis between flavonoid pathway-related synthesis genes and regulatory genes was constructed; notably, TFs belonging to MYB, MYC, WRKY, MADS, and HD-Zip were the most abundant significantly co-expressed TFs (Sun et al., 2017, 2019).
In this study, using both in vivo and in vitro experiments, we investigated the role of VviMYB86 in the regulation of flavonoid biosynthesis in grape berries. We demonstrated that VviMYB86 regulated PA and anthocyanin biosynthesis oppositely in grape berries; this TF significantly increased PA contents, primarily by enhancing the expression levels of the two LAR genes, whereas it reduced anthocyanin contents by downregulating the expression of VviANS and VviUFGT. Taken together, our findings provide insights into the regulatory mechanisms mediating anthocyanin and PA biosynthesis related to R2R3-MYB TFs.
Materials and Methods
Plant Materials and Growth Conditions
Grape materials of V. vinifera L. cv. Cabernet Sauvignon (CS) were collected from the experimental vineyard at the Shangzhuang Experimental Station of China Agricultural University in Haidian district, Beijing (40°14′ N, 116°20′ E, altitude 49 m), during the 2019 growing season. The own-rooted CS vines were planted in 2011 and were spaced at 2.5 m × 1.2 m, arranged in north-south rows. The training system and irrigation method applied in the vineyard were a modified vertical shoot positioning system and drip irrigation, respectively. Pest and nutrition management in the vineyard was undertaken in keeping with the local industry standard. Grapevine organs, including flowers, tendrils, stems, young leaves (10-day aged) and mature leaves (30-day aged) and berries were collected at E-L 23 (full blooming with 50% cap off) from at least nine randomly selected grapevines at distinct developmental time points. The roots were collected at E-L 23 from the underground parts of another nine randomly selected grapevines in the same vineyard. Berries were sampled at six developmental stages, including E-L 29 (peppercorn size), E-L 31 (pea-size), E-L 34, E-L 35 (early version), E-L 37 (end of version) and E-L 38 (commercial maturity), which corresponded to 5, 15, 43, 63, 80, and 98 days after anthesis, respectively. The sampling time was fixed at 8:00 to 9:00 am. After sampling, the grapevine organs and berry samples were placed into an ice box and transported into the laboratory within 2 h (Wen et al., 2015). After the samples arrived at the laboratory, they were immediately frozen in liquid nitrogen and stored at −80°C for further analysis. For the berries corresponding to E-L 31 to E-L 38, skin and seed samples were obtained by peeling fresh flesh, and the dissected skins and seeds were immediately immersed in liquid nitrogen before being stored at −80°C. Three biological replicates were taken for each sample.
The grape callus used in the present study was induced from berry skins of CS following the method described by Wang et al. (2015). The subculture medium comprised solid B5 medium supplemented with 30 g/L sucrose, 2.5 g/L acid-hydrolyzed casein, 3.0 g/L phytagel, 0.1 mg/L α-naphthylacetic acid, and 0.2 mg/L kinetin, and the pH was 5.9–6.0. The culture conditions were 24-h dark at a temperature of 24°C, and the callus was subcultured every 25 days.
Wild-type Arabidopsis thaliana (ecotype Col-0) was grown in soil in a growth chamber under cool-white fluorescent lamps providing a 16/8-h light/dark photoperiod at 25 ± 1°C.
Cloning and Bioinformatic Analysis of VviMYB86
On the basis of the predicted cDNA sequence of VviMYB86 (GenBank accession No. XM_002280991) in the NCBI reference sequence database and the genomic sequence of CS (VIT_207s0005g02480)1, the open-reading frame (ORF) of VviMYB86 was cloned via gene-specific primers (VviMYB86-F/R) from a CS cDNA library previously constructed by our lab (Mu et al., 2014). The resulting amplicons were cloned into a pMD19-T vector (Tsingke, China) for sequencing. The primers employed in the present study are listed in Supplementary Table 1.
Phylogenetic analysis was conducted using the neighbor-joining method in MEGA version 7.0 (Kumar et al., 2016). The conserved domains of the VviMYB86 protein were scanned by the Inter-ProScan program2. The theoretical molecular weight and isoelectronic point were calculated using the ProtParam tool3. For multiple sequence alignment analysis, the amino acid sequence of VviMYB86 and those of other MYB homologs from different plant species retrieved from the NCBI database were aligned using DNAMAN 6.0 software (Lynnon Corporation, United States).
Isolation and Bioinformatic Analysis of the VviMYB86 Promoter (pVviMYB86) Region
Grape genomic DNA was extracted from CS berries using a New Rapid Plant DNA Extraction Kit (BioTeke, China) and then used as template for cloning. The specific primers (Supplementary Table 1) were designed for amplification based on the predicted sequence from the NCBI reference sequence (NC_012013.3) and genomic sequence of CS4. The PCR products that were approximately 1,300 bp from the start codon, which were obtained using the Pfu DNA polymerase (Tiangen, China), were inserted into the pMD19-T vector (Tsingke, China) for sequencing validation. The cis-acting elements on pVviMYB86 were predicted by the PlantCARE website5. Sequence data from this article have been deposited in GenBank under accession number MW046258.
Total RNA Extraction, Reverse Transcription, and Quantitative Real Time PCR (qRT-PCR) Analysis of Gene Expression
All frozen plant samples (i.e., grapevine organs, berry skins, berry seeds, and grape calluses) were ground in liquid nitrogen, and total RNA was subsequently extracted by using a Universal Plant Total RNA Extraction Kit (BioTeke, China). The quality and concentration of the obtained RNA were detected using agarose gel electrophoresis and a NanoDrop 2000 spectrophotometer (Thermo Fisher, United States), respectively. The RNA employed for further analysis appeared as clear and bright bands in the agarose gel, the value of OD260/OD230 was more than 1.8, and the value of OD260/OD280 was between 1.8 and 2.1 (Meng et al., 2020). A total amount of 1 μg RNA was utilized for reverse transcription reaction using HiScript® IIQ RT SuperMix for qPCR (+ gDNA wiper) (Vazyme, China). qRT-PCR was performed using ChamQ Universal SYBR qPCR Master Mix (Vazyme, China) and an Applied Biosystems 7300 Real Time PCR System (Thermo Fisher, United States). Each qRT-PCR reaction (20 μL) consisted of 10 μL ChamQ Universal SYBR qPCR Master Mix, 2 μL of cDNA, 7.2 μL of ddH2O, and 0.8 μL of a primer mixture (equal amounts of forward primer and reverse primer, 10 mM). The specific PCR procedure was performed as described by Sun et al. (2015). Each RNA sample was subjected to three independent reactions in qRT-PCR analysis. The analysis followed the method described in a previous report (Ruijter et al., 2009). VviUbiquitin1 was employed as the reference gene (Bogs et al., 2005). The gene-specific primers used in this study are listed in Supplementary Table 1.
Subcellular Localization of VviMYB86
The ORF of VviMYB86 without the stop codon was PCR-amplified with specific primers (Supplementary Table 1) and subsequently inserted into a pEZS-NL expression vector containing the green fluorescent protein (GFP) reporter gene. Next, the specific procedure followed the method described previously (Meng et al., 2020). An empty vector was not used as a control because the “NL” vector could not express GFP well in the absence of a coding sequence added to the 5′ end of the ORF6.
Transactivation Assay of VviMYB86 in Yeast
A transactivation assay of VviMYB86 in yeast (Saccharomyces cerevisiae) was performed following the method described by Wang et al. (2016). The ORFs of VviMYB86, VviMYBPAR (GenBank No. AB911341), and VviMYBC2-L1 (GenBank No. EU181425) were cloned with specific primer pairs (Supplementary Table 1) and integrated into the yeast expression vector pGBKT7 (Takara, Japan). The expression vectors pGBKT7-VviMYB86, pGBKT7-VviMYBPAR (positive control), pGBKT7-VviMYBC2-L1 (negative control), and pGBKT7 (negative control) were transferred into a yeast host strain AH109 according to the manufacturer’s protocol (PT4087-1, Clontech, Japan). Successfully transformed yeast strains were applied onto the corresponding medium (SD/Trp-, SD/Trp-/His-, and SD/Trp-/His-/Ade-) and subsequently observed after being incubated at 30°C for 3–5 days.
Transient Expression Using Arabidopsis Protoplasts and Dual Luciferase Assays
To test the transcription function of VviMYB86, dual luciferase assays were conducted using polyethylene glycol-mediated transformation of Arabidopsis protoplasts. The effector was generated by inserting the ORF region of VviMYB86 into a pCAMBIA 1301 vector driven by the CaMV 35S promoter via an In-fusion HD cloning procedure (Takara, Japan). The empty vector was employed as the negative control. The promoter regions of VviLAR1 (pVviLAR1), VviLAR2 (pVviLAR2), VviANR (pVviANR), VviANS (pVviANS), VviUFGT (pVviUFGT), VviMYBPA1 (pVviMYBPA1), VviMYBPA2 (pVviMYBPA2), VviMYBPAR (pVviMYBPAR), VviMYBC2-L1 (pVviMYBC2-L1), and VviMYBA1 (pVviMYBA1) were amplified by PCR from the genomic DNA of CS by using gene-specific primers (Supplementary Table 1). Then, the promoter fragments were subcloned into a pGreenII 0800-LUC vector to serve as reporter plasmids. Protoplasts were extracted from young leaves of Arabidopsis, and then the effector and corresponding reporter genes were co-transformed into protoplasts as described previously (Mathur and Koncz, 1998). After being cultured in the dark at 23°C for 16 h, the transfected protoplasts were collected for dual luciferase assays using the Dual-Luciferase® Reporter Assay System (Promega, United States) following the manufacturer’s protocol (VPE1910, Promega, United States). The relative luciferase activity was calculated as the ratio of firefly luciferase activity to Renilla luciferase activity. All the transfection experiments were performed in triplicate.
Construction of the VviMYB86 Overexpression Vector and Grape Callus Transformation
The ORF of VviMYB86 was inserted into a pCXSN vector under the control of the CaMV 35S promoter. Subsequently, the successfully constructed expression cassette was introduced into A. tumefaciens strain GV3101 by using the freeze-thaw method. Three-week-old wild-type (WT) callus was chosen for genetic transformation following the method described by Van Eck et al. (2006) and Meng et al. (2020). The subculture medium was solid B5 medium supplemented with 30 g/L sucrose, 2.5 g/L acid-hydrolyzed casein, 3.0 g/L phytagel, 0.1 mg/L α-naphthylacetic acid, 0.2 mg/L kinetin, and 5 mg/L hygromycin, and the pH was 5.9–6.0. The culture conditions and the subculturing period were the same as those applied to the WT callus. VviMYB86 transgenic callus was confirmed by hygromycin-based gene detection and target gene expression quantification. The primers utilized in this section are listed in Supplementary Table 1.
Light Treatment of WT Callus and VviMYB86 Transgenic Grape Callus
Subcultured WT and VviMYB86 transgenic callus were employed to investigate the effects of light on PA and anthocyanin production, as well as the expression of VviMYB86, flavonoid-specific structural genes and related regulatory genes. For light treatment, the dark-cultured callus was subjected to a 16/8-h light/dark photoperiod with a light intensity of 2,000 lux for a subculturing period. The temperature was set at 24°C. After treatment, the callus was collected immediately for RNA extraction and production extraction. Three independent biological replicates were performed. The RNA extraction and qRT-PCR procedures were consistent with those described in section 2.3. VviUbiquitin1 was selected as the reference gene, and the gene-specific primers used in this study are listed in Supplementary Table 1. The extraction and determination procedures for the PAs and anthocyanins are described in sections “4-Dimethylaminocinnamaldehyde (DMACA) Staining and PA Content Determination” and “Determination of the Total Anthocyanin Content”, respectively.
4-Dimethylaminocinnamaldehyde (DMACA) Staining and PA Content Determination
The presence of PAs in grape callus was detected by DMACA staining (Li et al., 1996). Fresh callus was stained with DMACA reagent [1% (w/v) in methanol: 6 M HCl (1:1, v/v)] for 1 h and then observed. Three biological replicates were performed for each set of samples. Approximately 1.0 g of frozen sample was ground into powder in liquid nitrogen for PA extraction. The specific procedure for the extraction of soluble and insoluble PAs followed the methods described by Yu et al. (2019). After extraction, soluble PAs were lyophilized and redissolved in 300 μL of 50% (v/v) methanol solution. The soluble PAs were quantified by the DMACA method. Soluble PA fractions (20 μL) were mixed with 100 μL of DMACA reagent on a 96-well plate. After samples were incubated at room temperature for 4 min, spectrophotometric quantification was performed at 640 nm using a SpectraMax 190 Microplate Reader (Molecular Devices, United States). (+)-Catechin (Sigma, United States) was used as a standard and processed in parallel with experimental samples. The insoluble PA content was analyzed by the butanol/HCl method. For quantification, one hundred microliters of supernatant were added to a 96-well plate, and then the A550 value was measured via a SpectraMax 190 Microplate Reader. Procyanidin B1 (Sigma, United States) was used as a standard.
Determination of the Total Anthocyanin Content
Total anthocyanins were extracted with the HCl-methanol method (Liu et al., 2016), with slight modifications. Approximately 1.0 g of frozen sample was ground into powder in liquid nitrogen. Next, the powder was extracted with 1.0 mL of a hydrochloric acid: methanol solution (60% methanol; 0.1% hydrochloric acid) by sonicating in ice water for 30 min in the dark. The mixture was centrifuged at 8,000 rpm for 5 min at 4°C. The supernatant was collected and the residue was extracted twice. All the supernatants were pooled. The resulting supernatant was lyophilized and redissolved in 400 μL of hydrochloric acid: methanol solution. The extraction solution was finally stored at −80°C. The total anthocyanin content was determined by using the pH-differential method (Stojanovic and Silva, 2006). The extraction was diluted with KCl buffer (pH 1.0) and CH3CO2Na⋅3H2O buffer (pH 4.5). Next, spectrophotometric quantification of the mixture was performed at 520 and 700 nm using a SpectraMax 190 Microplate Reader. The total anthocyanin content was calculated using the following formula: total anthocyanin content = (A × MW × DF × Ve × 1,000)/(ε × M), where A = (A520 − A700)pH1.0 − (A520 − A700)pH4.5, MW is the molecular weight of malvidin-3-glucoside (493.5), DF is the dilution factor, Ve is the extraction volume, ε is the molar extinction coefficient of malvidin-3-glucoside (28,000), and M is the mass of the extracted samples.
Statistical Analysis
The data are presented as the means ± standard deviations (SDs). One-way analysis of variance (ANOVA) was conducted using Microsoft Excel 2016 (Microsoft, United States). String diagrams and bar charts were created by GraphPad Prism 8 (GraphPad Software). Information regarding the experimental protocol, data analysis of RNA sequencing (RNA-seq) and submission to the NCBI Gene Expression Omnibus were recorded in the publication of our research group (Sun et al., 2019). In the present study, we selected the data of the control group at the E-L 29, E-L 31, and E-L 35 to E-L 38 stages of berry development for Pearson’s correlation analysis. Pearson’s correlation analysis was conducted using the fragments per kilobase of exon per million fragments value of each gene in R environmental language (version 3.5.1). TFs with correlation coefficients ≥0.7 or ≤0.7 and P < 0.05 were considered to have a significant expression correlation with the target gene.
Results
Isolation and Characterization of VviMYB86
To explore additional putative TFs involved in regulating PA biosynthesis, especially the expression of the genes encoding LAR in grape berries, we reanalyzed the RNA-seq database of the whole developmental period of CS berries deposited by Sun et al.7. An R2R3-MYB gene, annotated as VviMYB86, whose expression was strongly positively correlated with the expression of PA-specific structural and regulatory genes, including VviLAR1, VviLAR2, VviANR, and VviMYBPAR, was screened; the Pearson’s coefficients were 0.733, 0.923, 0.796, and 0.961, respectively (Supplementary Table 2). In addition, a co-expression analysis between VviMYB86 and the genes involved in the anthocyanin biosynthesis branch was also performed. The Pearson’s coefficients between the expression levels of VviMYB86 and VviANS and between those of VviMYB86 and VviUFGT were −0.201 and −0.811, respectively (Supplementary Table 2). Therefore, we hypothesized that VviMYB86 might participate in the regulation of both PA and anthocyanin biosynthesis in grape berries.
Next, the 1,356-bp ORF of VviMYB86 was amplified from the cDNA library of CS berries previously constructed by our lab (Mu et al., 2014; Supplementary Figure 2). Bioinformatic analysis indicated that VviMYB86 encoded a predicted hydrophilic protein containing 451 amino acid residues and had a molecular mass of 50.23 kD and a pI of 6.47. Amino acid sequence analysis showed that, similar to other R2R3-MYB members, VviMYB86 processed two adjacent MYB repeats in the conserved DNA-binding domain, in which the R2 region was located between residues 14–61, and R3 was located between residues 67–111 (Figure 1A). A conserved C1 motif (lsrGIDPxT/NHR) (Chen et al., 2019) was found between residues117–128, which also existed in VviMYB5a, VviMYB5b, VviMYBPA1, VviMYBC2-L1, and VviMYBC2-L2. Moreover, VviMYB86 contained a conserved motif 1 [DNEI(A/S/G)N(D/A/N)V], which has been proven to bind to a specific site in bHLH proteins (Zimmermann et al., 2004; Wang et al., 2018), suggesting that VviMYB86 may interact with bHLH-type proteins. Phylogenetic analysis demonstrated that VviMYB86 was clustered with MdMYB6 and AtMYB60, and these TFs all had a negative effect on anthocyanin biosynthesis in planta (Figure 1B). The above results suggested that VviMYB86 was likely a candidate repressor of flavonoid biosynthesis.
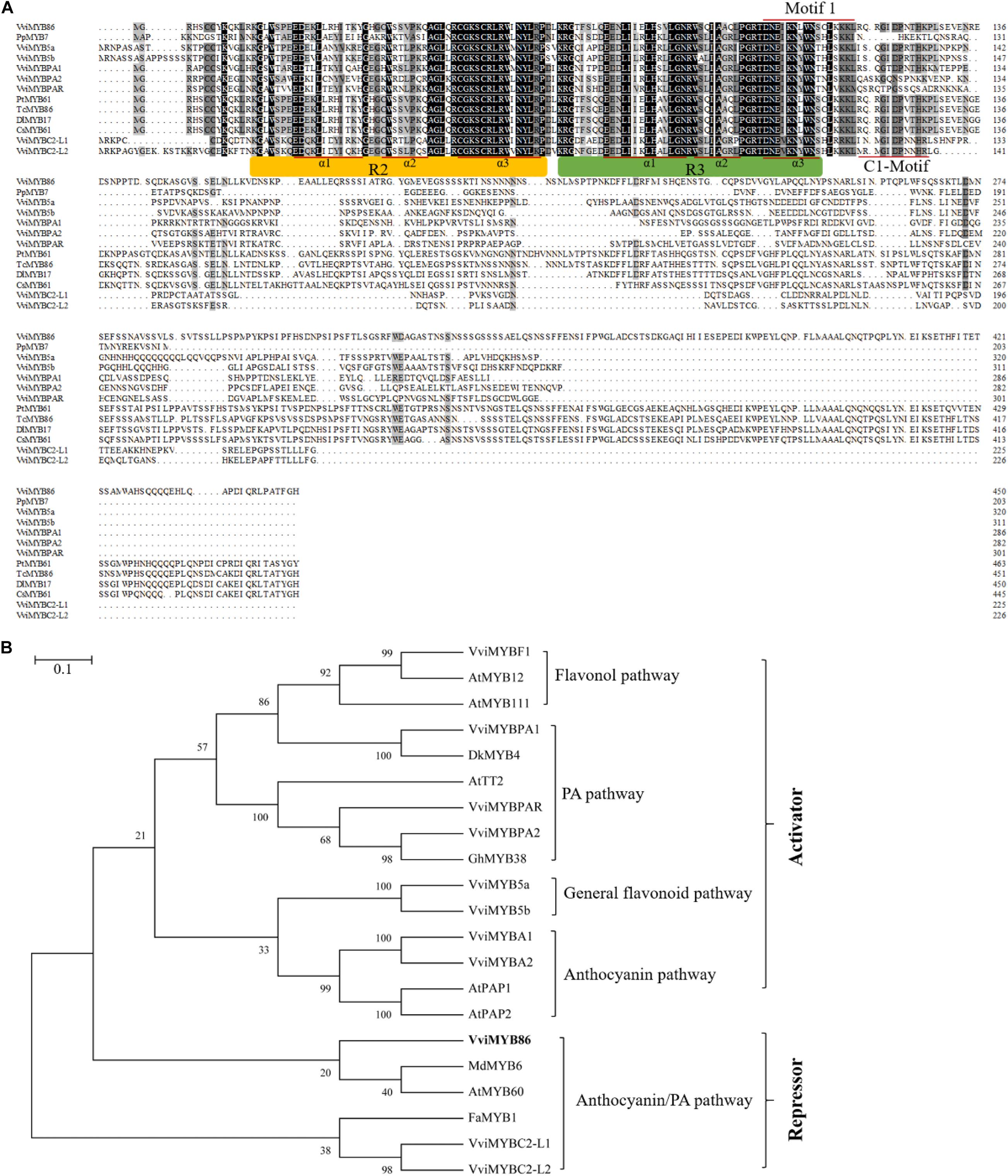
Figure 1. The deduced peptide sequences of VviMYB86 and related MYB’s. (A) Polypeptide alignment analysis of the full-length amino acid sequences of VviMYB86 and other PA-related R2R3 MYB transcription factors (TFs) as well as TFs with high homology. Conserved residues were highlighted in black, and partial conservation was indicated in gray. The R2 and R3 MYB DNA binding domains were indicated above the alignment. The alpha helices of R2 and R3 repeats were indicated with red lines. Motif 1 [DNEI(A/S/G)N(D/A/N)V] and C1-Motif (lsrGIDPxT/NHR) were indicated by underlining with red line. (B) Phylogenetic tree of MYB TFs related to VviMYB86. The phylogenetic tree was constructed using Neighbor-Joining method of MEGA 7.0 software. The scale bar represents the number of substitutions per site. The putative regulatory functions of most of the proteins in the control of flavonoid biosynthesis are indicated. GenBank accession numbers are as follows (in parentheses): AtTT2 (Q9FJA2), AtMYB12 (NP_182268), AtMYB111 (NP_199744), AtMYB60 (AAC83617), AtPAP1 (AAG42001), AtPAP2 (NP_176813), VviMYB5a (AAS68190), VviMYB5b (AAX51291), VviMYBPA1 (CAJ90831), VviMYBPA2 (ACK56131), VviMYBPAR (XP_003633091), VviMYBF1 (ACT88298), VviMYBA1 (BAD18977), VviMYBA2 (BAD18978), VviMYBC2-L1 (ABW34393), VviMYBC2-L2 (ACX50288), DkMYB4 (BAI49721), FaMYB1 (AAK84064), GhMYB38 (AAK19618), GhMYB10 (ABV53918), and MdMYB6 (AAZ20429).
Subcellular Localization and Transcriptional Activity of VviMYB86
To examine the subcellular localization of the VviMYB86 protein, a VviMYB86-GFP expression cassette was generated and transformed into onion (Allium cepa L.) epidermal cells via particle bombardment. Confocal microscopy results revealed that cells expressing the VviMYB86-GFP fusion gene exhibited fluorescence restricted to the cell nucleus (Figure 2A), indicating that VviMYB86 was localized in the nucleus. In a yeast-based transactivation assay, like the positive control cells expressing GAL4 BD-VviMYBPAR, the yeast cells expressing the fusion protein GAL4 BD-VviMYB86 were able to grow on SD medium lacking leucine, histidine, and adenine, whereas the negative control yeast cells containing pGBKT7-VviMYBC2-L1 or the empty vector were able to grow only on SD medium lacking tryptophan (Figure 2B). These results indicated that VviMYB86 possessed transcriptional activity.
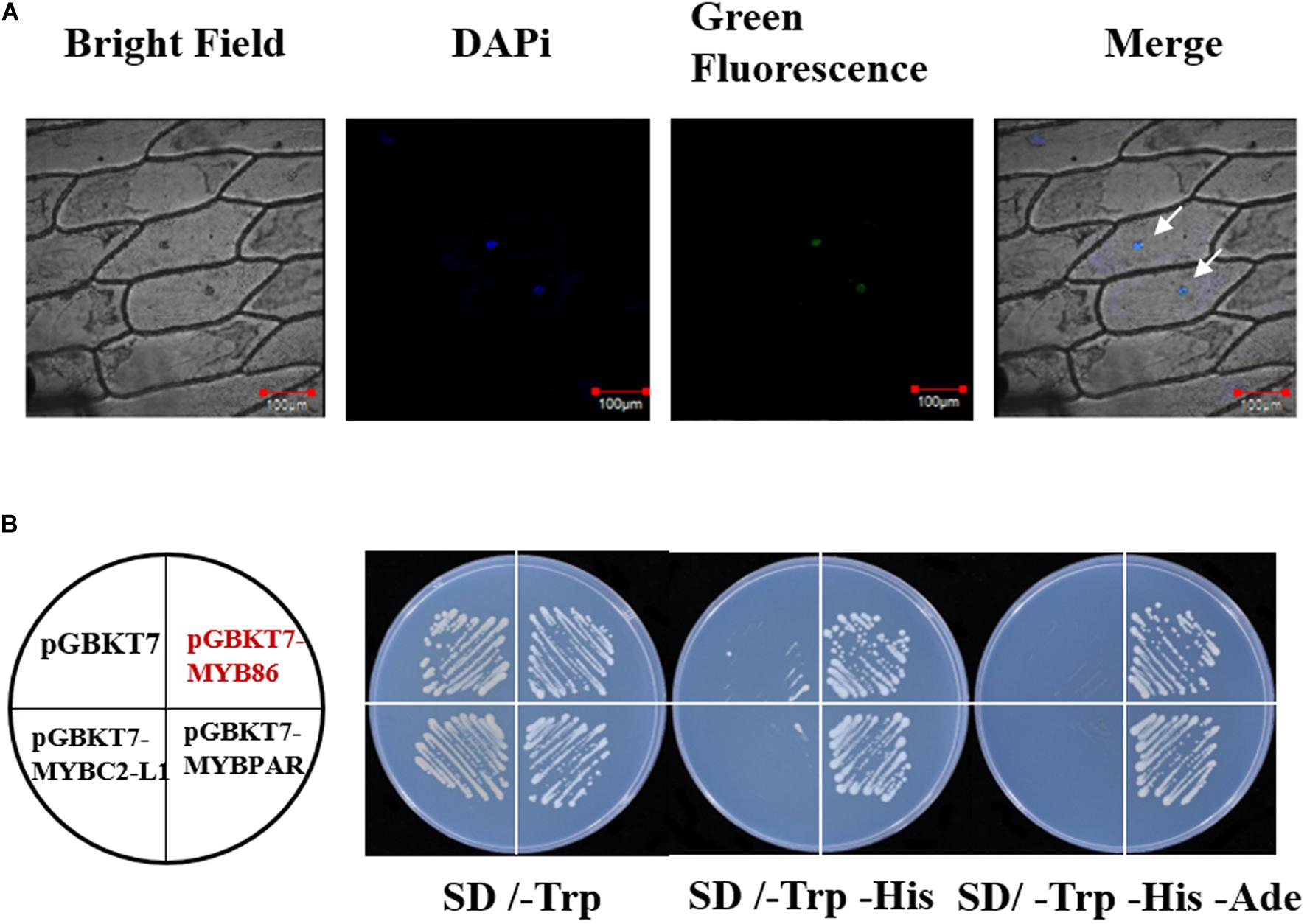
Figure 2. Subcellular localization of VviMYB86 and its transcription activation. (A) GFP activity in onion epidermal cells transiently expressing a VviMYB86-GFP construct. Bar: 100 μm. GFP, green fluorescence channel; Marker, blue fluorescence channel; DIC, bright light channel; Merge, the GFP and DAPi overlap. Arrows were the sites of GFP and DAPi overlap. (B) Transcription activation by VviMYB86 in yeast. The pGBKT7-VviMYBPAR plasmid was employed as a positive control. The pGBKT7-VviMYBC2L1 and pGBKT7 plasmids were employed as the negative controls. SD/-Trp, SD medium lacking tryptophan; SD/-Trp –His, SD medium lacking both tryptophan and histidine; SD/-Trp –His -Ade, SD medium lacking tryptophan, histidine, and adenine.
Spatiotemporal Expression Patterns of VviMYB86 in Grapevines
It has been found that grape flavonoids preferentially localize in both grape berry skins and seeds but are present in only negligible amounts in the mesocarp (Adams, 2006; Braidot et al., 2008). Therefore, the expression patterns of VviMYB86 during grape berry development were assessed by qRT-PCR with RNA isolated from the skins and seeds of CS berries (Figure 3). In both berry skins and seeds, the expression level of VviMYB86 was high in the early developmental stage, from E-L 31 to E-L 34 (Figure 3). However, the expression of this gene progressively decreased throughout the entire developmental stage of berry skins and seeds. The expression pattern of VviMYB86 in whole berry development coincided well with the expression of PA-specific structural genes in grape berries, whereas it contrasted with that of VviUFGT in berry skins (Figure 3).
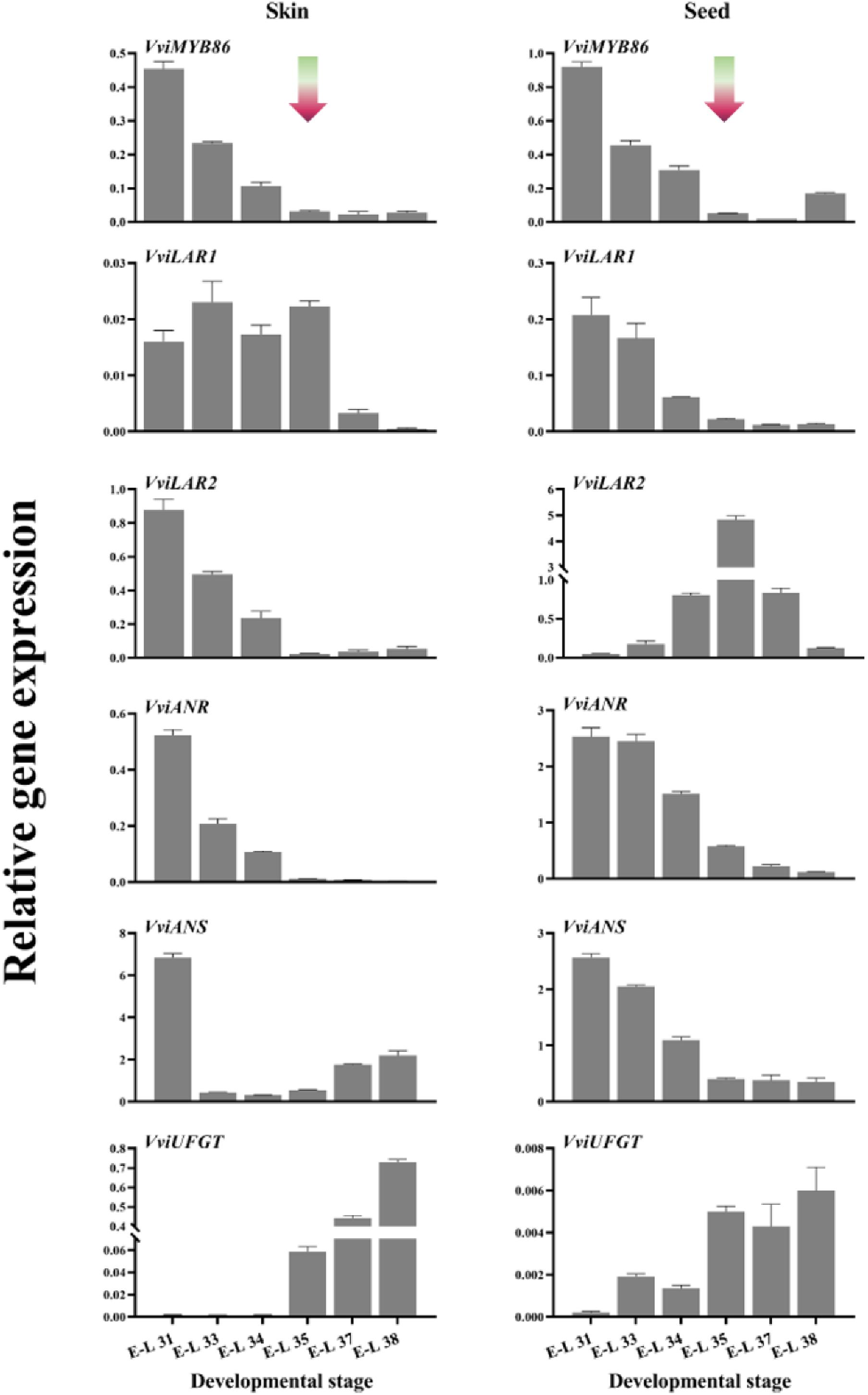
Figure 3. Spatiotemporal expression profile of VviMYB86 and some PA-related and anthocyanin-related genes in grape skins and seeds. The gradient arrows indicated the time-point of véraison. Gene expression was determined by real-time quantitative PCR and normalized with the expression of VviUbiquitin1. Each sample was individually assayed in triplicate. Error bars indicated the standard error of the mean.
In addition, the transcript levels of VviMYB86 in different grapevine organs, including roots, tendrils, flowers, stems, and young and mature leaves, were quantified by qRT-PCR. VviMYB86 was expressed in all the examined grapevine organs with different transcript levels. Specifically, VviMYB86 was most abundantly expressed in tendrils followed by flowers, stems, and then young and mature leaves, with a relatively low level of expression in the roots (Supplementary Figure 3).
The above results indicated that the expression of VviMYB86 was regulated in a tissue- and temporal-specific manner in grapevine.
Promoter Activation of PA and Anthocyanin Biosynthetic Pathway Genes in the Arabidopsis Protoplast System
To identify the target PA and anthocyanin pathway genes of VviMYB86 in grape berries, dual luciferase assays were performed using the Arabidopsis protoplast system. First, the promoters of the structural and regulatory genes in the PA and anthocyanin biosynthesis branches, including VviLAR1, VviLAR2, VviANR, VviANS, VviUFGT, VviMYBPA1, VviMYBPA2, VviMYBPAR, VviMYBC2-L1, and VviMYBA1, were cloned from the genomic DNA of grape berries; the lengths were 1,528, 1,587, 1,369, 973, 1,169, 712, 895, 1,128, 856, and 755 bp, respectively. Next, the cis-elements existing on these gene promoters were predicted by the PlantCARE database (see text footnote 5). As shown in Figure 4A, all ten gene promoters contained at least one of the following types of predicted MYB-binding element: the AC-rich element [(A/C)CC(A/T)A(A/C)], the MYB core element (CNGTTR), the MBSI (TTTTTACGGTTA), the MYBST1 element (GGATA), the MYB1AT element (WAACCA), and the MYBPZM element (CCWACC) (Xu et al., 2015; Zhou et al., 2019), indicating that all ten genes could be regulated by MYB TFs. In addition, the predicted bHLH-binding site G box (CACGTG) was observed in pVviLAR1, pVviANR, pVviANS, pVviMYBPA2, pVviMYBPAR, pVviMYBC2-L1, and pVviMYBA1 (Hichri et al., 2011), which suggested that the seven genes could also be regulated by TFs from the bHLH family. Except for pVviUFGT, pVviMYBPA2, pVviMYBPAR, pVviMYBC2-L1, and pVviMYBA1, five other gene promoters contained a WRKY-binding site W box (TTGACC) (Shang et al., 2010), implying that WRKY family TFs might participate in regulating these genes, which still need to be proven experimentally.
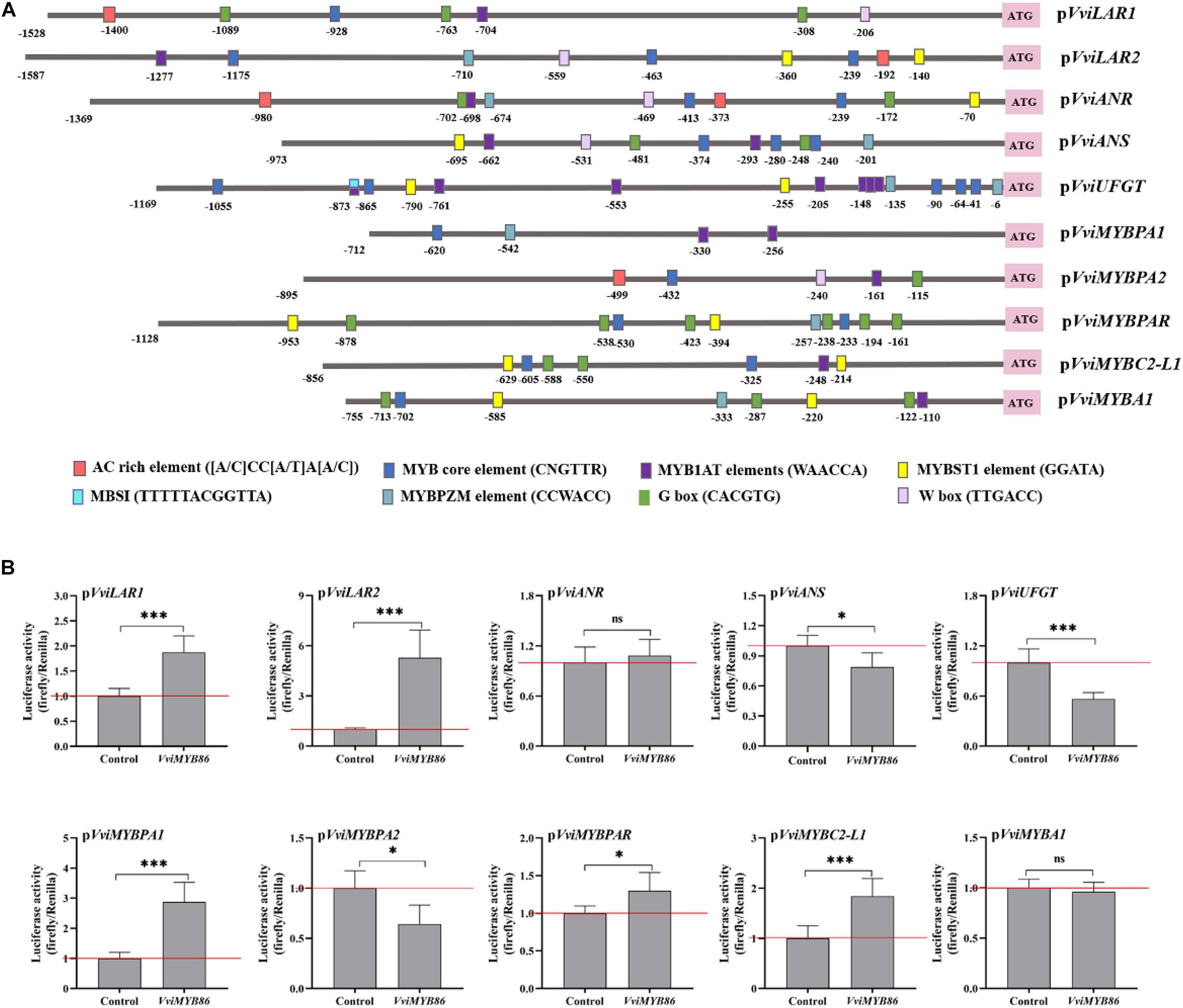
Figure 4. The influence of VviMYB86 on the promoters of structural and regulatory genes in PA and anthocyanin synthesis. (A) The characteristics of cis-elements for binding of MYB, MYC and other proteins in structural and regulatory gene promoters of PA and anthocyanin synthesis. (B) Effects of VviMYB86 on the activities of gene promoters of PA and anthocyanin synthesis as determined by dual luciferase assay. Control indicated the activity of the promoter transfected with the empty vector (pCAMBIA 1301). Normalized luciferase activity was calculated as the ratio between firefly and Renilla reniformis luciferase activities. Each column represents means ± SD from four biological replicates.
Then, VviMYB86 and the different aforementioned gene promoters were co-transformed into Arabidopsis protoplasts, and dual luciferase assays were conducted (Figure 4B). The results demonstrated that VviMYB86 strongly enhanced the activities of pVviLAR1 (by approximately 1.87-fold) and pVviLAR2 (by approximately 5.29-fold) but not pVviANR (by approximately 1.08-fold). In contrast, VviMYB86 repressed the activities of pVviANS and pVviUFGT, causing 21% and 44% decreases in the activity of pVviANS and pVviUFGT, respectively, compared with that of the control. Furthermore, VviMYB86 could also have an effect on the activity of the promoters of several regulatory genes related to PA and anthocyanin biosynthesis. In detail, VviMYB86 enhanced the activities of pVviMYBPA1 (by approximately 2.88-fold) and pVviMYBPAR (by approximately 1.30-fold), whereas repressed the activity of pVviMYBA1 but not significantly.
In summary, VviMYB86 might regulate the expression of structural genes encoding LAR and consequently regulate PA biosynthesis in grape berries, and the VviMYB86 signaling might act upstream of VviMYBPA1 and VviMYBPAR. In terms of anthocyanin synthesis, VviMYB86 might exert a negative effect by downregulating the expression of VviANS and VviUFGT.
Promotion of PA Biosynthesis in Grape Callus by the Overexpression of VviMYB86
To demonstrate the function of VviMYB86 on PA biosynthesis in grape berries, we generated VviMYB86-overexpressing transgenic grape callus. WT callus was used as the control. The culture conditions for both the WT and transgenic calluses were 24 h of darkness at a temperature of 24°C. The relative expression levels of VviMYB86 in the WT callus and VviMYB86 transgenic lines were determined by qRT-PCR. Three independent transgenic lines (L1, L3, and L5) with different ectopic expression levels of VviMYB86 (Figure 5A and Supplementary Figure 4) were selected for further analysis. The morphological features of the transgenic lines were similar to those of the WT callus, and both were white and loose (Figure 5B and Supplementary Figure 5A). After being stained with DMACA for 1 h, all the transgenic lines exhibited a notable dark-blue color, whereas the color of the WT callus was pink, suggesting that the VviMYB86 transgenic lines accumulated more PAs than the WT callus (Figure 5B and Supplementary Figure 5A). The determination of both PA and insoluble contents through the DMACA and butanol/HCl methods, respectively, also confirmed that, compared with the WT callus, the three VviMYB86 transgenic lines produced large amounts of both soluble and insoluble PAs (Figure 5C and Supplementary Figure 5B).
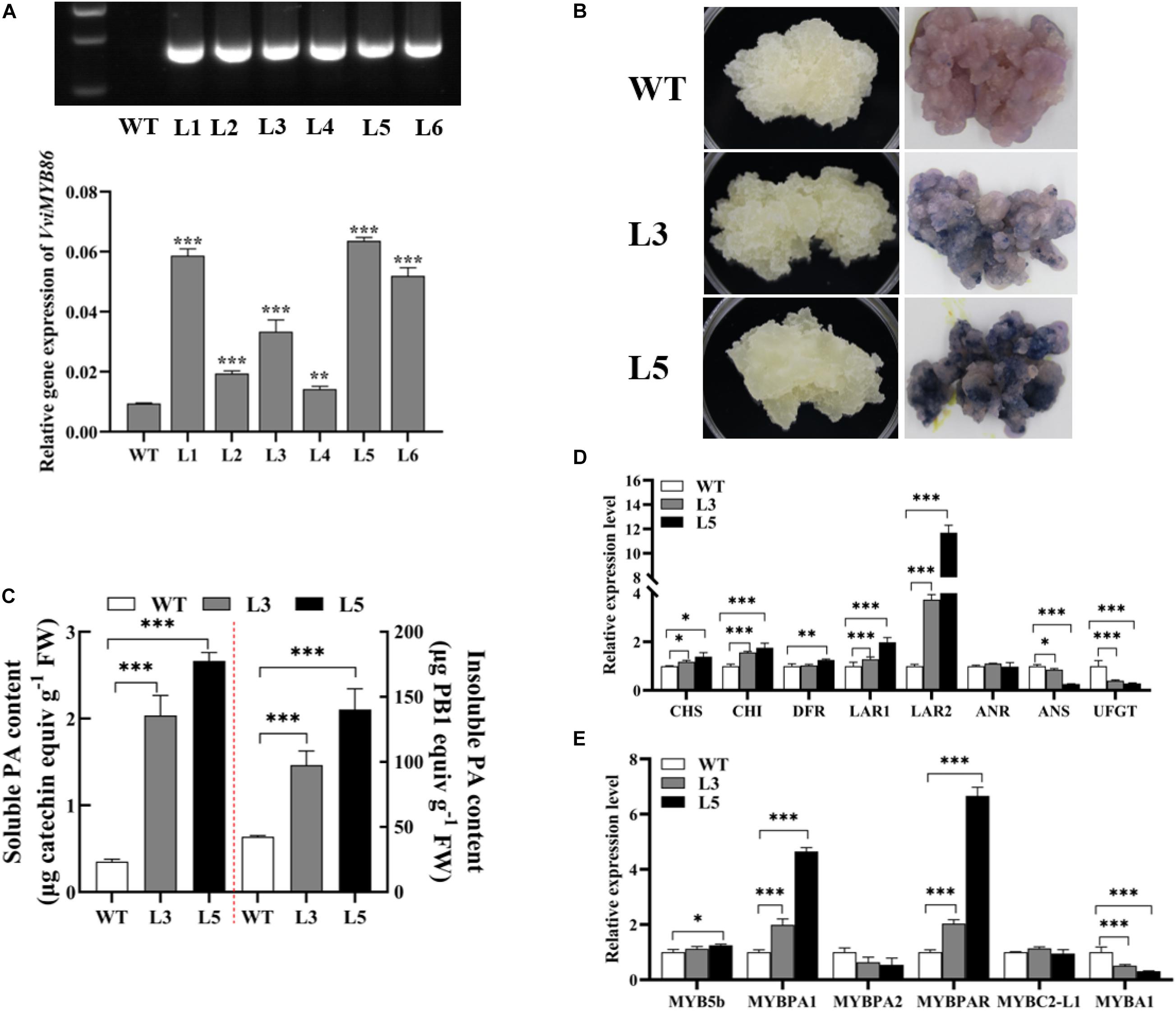
Figure 5. VviMYB86 positively regulated proanthocyanidin (PA) synthesis in grape callus. (A) Identification of transgenic callus. The top was hygromycin gene expression detected by PCR and analyzed by gel electrophoresis. The below was the expression level of VviMYB86 in wild type (WT) callus and different transgenic lines (L1-L6). The value above the column referred to the transcripts of VviMYB86 in transgenic and WT calluses cultured under dark conditions. (B) The photos of WT callus and two independent VviMYB86 transgenic lines (L3 and L5). The left was WT callus and transgenic lines in their natural growth states cultured in darkness. The right was WT callus and transgenic lines after 4-Dimethylaminocinnamaldehyde (DMACA) staining. (C) The soluble and insoluble PA content in the WT callus and transgenic lines. FW, fresh weight. Data was expressed as means ± SD of three replicates. Asterisks indicated significant differences relative to the control by one-way ANOVA test (∗∗∗p < 0.001). (D) The relative expression of flavonoid pathway related structural genes. After several successive rounds of subculture, stable transgenic callus lines were established on selectable medium. Callus grown for 25 days was collected for each assay. CHS, chalcone synthase; CHI, chalcone isomerase; DFR, dihydroflavonol-4-reductase; LAR, leucoanthocyanidin reductase; ANR, anthocyanidin reductase; ANS, anthocyanidin synthase; UFGT; UDP-glucose: flavonoid-3-O-glucosyltransferase. (E) The relative expression of known flavonoid regulators. Data was expressed as means ± SD of three replicates. Asterisks indicated significant differences relative to the control by one-way ANOVA test (∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001).
To confirm the target genes of VviMYB86, the expression levels of genes participating in the flavonoid biosynthesis pathway were determined by qRT-PCR. The results demonstrated that the expression of most genes related to the PA biosynthesis pathway was upregulated in the VviMYB86 transgenic calluses (Figure 5D and Supplementary Figure 5C). Moreover, in the VviMYB86 transgenic calluses, the expression of all the upstream genes including VviCHS, VviCHI, VviFLS, and VviDFR, was all upregulated to different degrees. With respect to PA-specific genes, the expression of VviLAR1 and VviLAR2 was enhanced, especially VviLAR2, whose expression was upregulated 3.16-fold in L1, 2.73-fold in L3 and 10.68-fold in L5. However, in all the transgenic lines, the expression level of VviANR remained unchanged compared with that in the WT callus (Figure 5D and Supplementary Figure 5C). These results suggested that VviMYB86 tended to primarily modulate the expression of the two LAR genes, subsequently leading to increased PA contents. Furthermore, the expression levels of known PA regulators, including VviMYB5a, VviMYB5b, VviMYBPA1, and VviMYBPAR, increased to different levels in the VviMYB86 transgenic calluses relative to those in the WT callus (Figure 5E and Supplementary Figure 5D).
VviMYB86 Represses the Anthocyanin Biosynthesis Branch in Transgenic Grape Callus Under Light Conditions
We observed that the expression levels of anthocyanin pathway genes, including VviANS and VviUFGT, were repressed in the VviMYB86 transgenic lines, compared with those in the WT callus (Figure 5D and Supplementary Figure 5C). However, anthocyanins were detected at very low levels in both WT callus and VviMYB86 transgenic lines cultured under dark conditions (Supplementary Figure 6). However, the effect of overexpressing VviMYB86 on anthocyanin accumulation has not been fully determined.
Then, the promoter region of VviMYB86 was isolated from the genomic DNA of CS berries (Supplementary Figure 7). To elucidate the regulatory mechanisms that might control the expression of VviMYB86, pVviMYB86 was subsequently analyzed by predicting the cis-elements though the PlantCARE database (see text footnote 5), and the results were shown in Figure 6A. Several cis-elements related to light response, environmental response, hormonal response and developmental regulation were found to be widely present in pVviMYB86, indicating that VviMYB86 participated in multiple physiological processes in grape berries. Among these cis-elements, light-responsive elements were the most frequent and most variable, suggesting that the expression of VviMYB86 may be regulated by light.
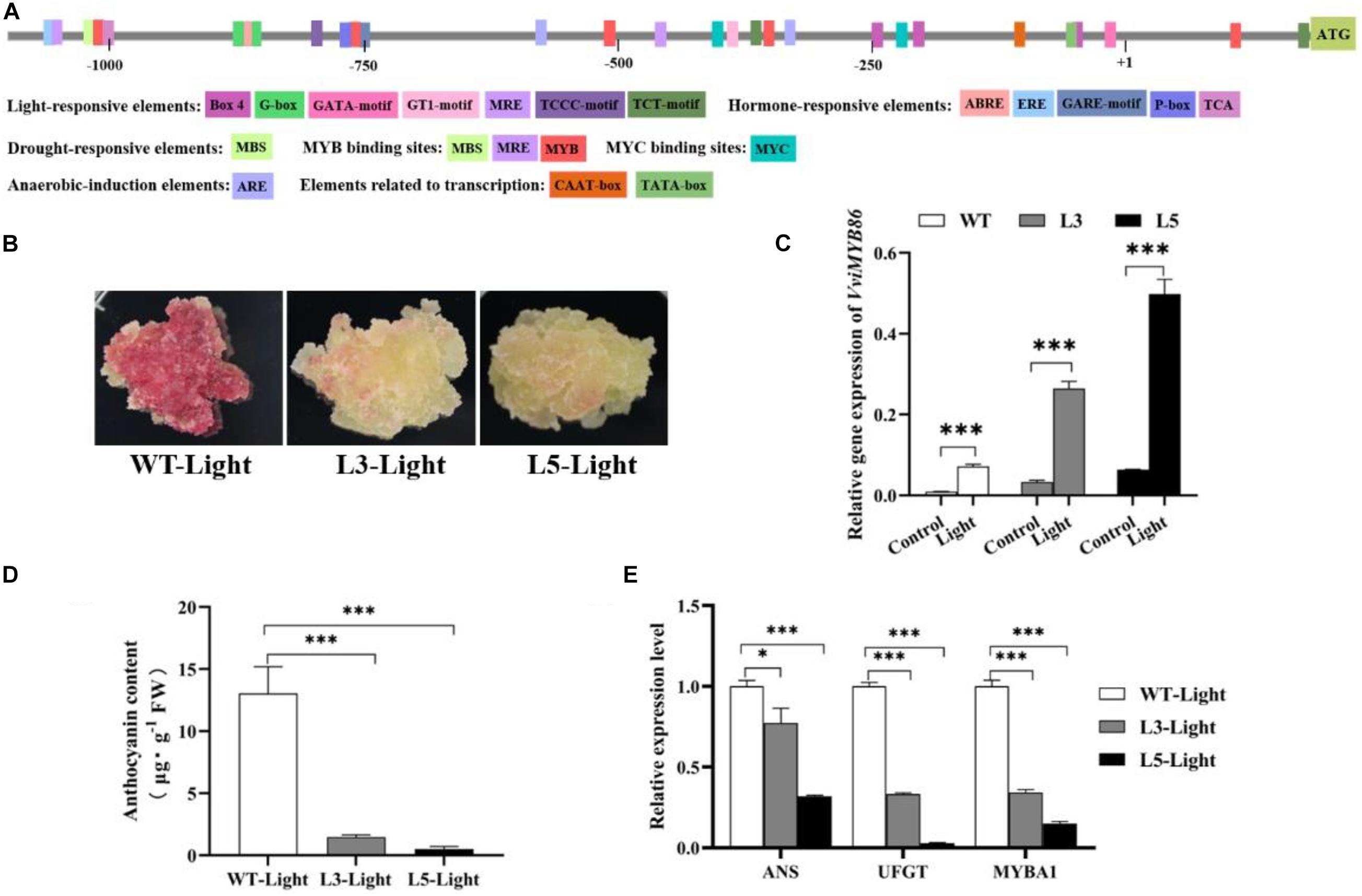
Figure 6. VviMYB86 negatively regulated anthocyanin synthesis in grape callus. WT-Light, wild type callus cultured under light conditions; L3-Light, transgenic line 3 cultured under light conditions; L5-Light, transgenic line 5 cultured under light conditions. (A) The cis-acting elements present in VviMYB86 promoter predicted by PlantCARE website (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/). (B) The nature status of WT callus and transgenic line 3 and line 5 under light conditions. (C) The expression levels of VviMYB86 in WT and transgenic calluses. Control: wild-type callus and transgenic calluses that cultured under dark conditions. Light: wild-type callus and transgenic calluses that cultured under light conditions. The value above the column referred to the transcripts of VviMYB86 in transgenic and WT calluses. (D) Anthocyanin contents in the WT callus and transgenic lines cultured under light conditions. The anthocyanin contents in callus samples measured by using the pH-differential method. FW, fresh weight. (E) Expressions of corresponding genes of anthocyanin biosynthesis pathway in the WT callus and transgenic lines cultured under light conditions. ANS, anthocyanidin synthase; UFGT, UDP-glucose: flavonoid-3-O-glucosyltransferase. Data was expressed as means ± SD of three replicates. Asterisks indicated significant differences relative to the control by one-way ANOVA test (∗p < 0.05; ∗∗∗p < 0.001).
Next, the three independent VviMYB86 transgenic lines cultured under dark conditions were subjected to light conditions (L1-Light, L3-Light, and L5-Light) with a light intensity of approximately 2,000 lux for a subculture period. WT callus subjected to the same light condition (WT-Light) was employed as the control. We observed that the WT-Light turned red, whereas the three transgenic lines were only slightly reddish in color (Figure 6B and Supplementary Figure 8A). The addition of light increased the transcription level of VviMYB86 in the light-exposed transgenic lines as well as in the WT-Light compared with that in the corresponding calluses cultured under normal conditions (Figure 6C and Supplementary Figure 8B). The results revealed that the expression of VviMYB86 could be inducible by light.
In addition, the transcript levels of VviMYB86 in light-exposed transgenic lines were greater than those in the WT-Light (Figure 6C and Supplementary Figure 8B). Anthocyanins were subsequently extracted from both the light-exposed WT and VviMYB86 transgenic calluses. The measurement of anthocyanin contents revealed that anthocyanin production was considerably reduced in the light-exposed transgenic calluses (Figure 6D and Supplementary Figure 8C). The qPCR results showed that the expression of anthocyanin biosynthesis branch genes, including VviANS and VviUFGT, was downregulated in the light-exposed transgenic lines (Figure 6E and Supplementary Figure 8D). In addition, VviMYBA1, a MYB TF, specifically regulates anthocyanin synthesis (Cutanda-Perez et al., 2009); the expression of VviMYBA1 was also repressed (Figure 6E and Supplementary Figure 8D). Taken together, the above results suggested that VviMYB86 repressed anthocyanin biosynthesis in grape callus cultured under light conditions.
Discussion
VviMYB86 Regulates PA Biosynthesis in Grape Berries Primarily by Affecting the Expression of the Two LAR Genes
In this study, we demonstrated that VviMYB86 may participate in the regulation of PA biosynthesis in grape berries. Unlike Arabidopsis, in which PAs are solely detected as monomeric or polymeric (−)-epicatechin forms, PAs are present as both (+)-catechin- and (−)-epicatechin-based PAs in grape skins and seeds (Xie et al., 2003; Bogs et al., 2005). The reason for this discrepancy is that LAR is not encoded in the Arabidopsis genome (Xie et al., 2003). In grapes, there are two LARs (VviLAR1 and VviLAR2) whose function are similar for PA biosynthesis (Yu et al., 2019). The temporal- and tissue-specific expression patterns of genes encoding LAR and VviANR contribute to the continuous accumulation of PAs before véraison in grape berries (Bogs et al., 2005). However, in grape berries, PA-related MYB TFs, including VviMYB5b, VviMYBPA1, VviMYBPA2, and VviMYBC2-L1, significantly influence the expression levels of VviLAR1 and VviANR but not VviLAR2 (Bogs et al., 2007; Deluc et al., 2008; Terrier et al., 2009; Huang et al., 2014). Although VviMYB5a activates the activity of only pVviLAR1, ectopic expression of VviMYB5a also activates the transcription of AtBAN (which encodes ANR) in Arabidopsis and induces the accumulation of (−)-epicatechin-form PAs but not (+)-catechin-form PAs in tobacco (Deluc et al., 2006, 2008). VviMYBPAR, whose gene transcription profile is closely related to that of VviLAR2, significantly activates the activities of pVviLAR1 and pVviANR, whereas it only slightly increases the activity of pVviLAR2 in a transient reporter assay (Koyama et al., 2014). Recently, by calculating the correlations between the expression levels of structural and regulatory genes in the PA biosynthesis branch of grapevine calluses subjected to different light intensities, researchers showed that VviLAR2 was the target gene of VviMYBPA2. However, owing to the low expression level of VviMYBPA2, the effects on the expression level of VviLAR2 are limited (Cheng et al., 2020).
Unlike known PA regulators, VviMYB86 appeared to utilize VviLAR1 and VviLAR2 as target genes to control PA synthesis in grape berries. The spatiotemporal expression profiles of VviMYB86 in grape berry skins and seeds were consistent with those of VviLAR2 and VviLAR1, respectively. The results of our dual luciferase assay indicated that VviMYB86 enhanced the activities of pVviLAR1 and pVviLAR2. Overexpression of VviMYB86 in grape callus that cultured under dark condition also enhanced the transcript levels of VviLAR1 and VviLAR2. Our results may help to elucidate the regulatory mechanism governing the expression of the two LAR genes in grape berries.
In peach, PpMYB7 regulates PA synthesis by activating the transcription of PpLAR1 but not PpANR (Zhou et al., 2015). With respect to members of the R2R3-MYB TF family, the α-helix regions of the R2R3 domain have been shown to play a critical role in directing MYB TFs to bind the promoters of flavonoid structural genes (Williams and Grotewold, 1997; Jia et al., 2004). The first and second α-helices of the R2 domain in both VviMYB86 and PpMYB7 differ from those of the other PA-related R2R3-MYB regulators, whereas the third α-helix of the R2 domain and all the three α-helices of the R3 domain are conserved. Jia et al. (2004) reported that the first two flexible α-helix regions of the R2 domain might be important for the recognition of a specific DNA-binding site, as they afford secondary structure and ternary folding of the R2R3 domain. Therefore, further experiments should be performed to address whether the difference in activation patterns between VviMYB86 as well as PpMYB7 and other PA-related R2R3-MYB regulators is caused by genetic variation in the region encoding the first and second α-helices of the R2 domain. Overall, both VviMYB86 and PpMYB7 represent a new group of R2R3-MYB genes that regulate PA biosynthesis in plants.
Furthermore, VviMYB86 not only controls the expression of PA-specific structural genes in grape berries but also affects the expression of known PA regulators, especially VviMYBPAR and VviMYBPA1. The results of the dual luciferase assay revealed that VviMYB86 enhanced the activities of pVviMYBPA1 and pVviMYBPAR. Overexpression of VviMYB86 in grape callus increased the transcript levels of VviMYBPA1 and VviMYBPAR. It is possible that VviMYB86 signaling acts upstream of VviMYBPA1 and VviMYBPAR. Similar interactions have been suggested between VviMYBPA1 and VviMYBPA2, as well as VviMYBPAR and VviMYBPA1 (Terrier et al., 2009; Koyama et al., 2014).
VviMYB86 Responds to Light and Represses Anthocyanin Biosynthesis
Light is an important environmental factor that influences flavonoid biosynthesis in plants. MYB TFs play pivotal roles in the response to external stimuli and in the regulation of flavonoid synthesis. The majority of MYB TFs associated with flavonoid synthesis have been found to respond to light, such as VviMYBPA1 (Koyama et al., 2012), VviMYBF1 (Azuma et al., 2012), and PyMYB10 (Zhang et al., 2011). The transcript levels of these R2R3-MYB TFs are modulated to regulate the biosynthesis of flavonoid compounds with changing light conditions (Zoratti et al., 2014). In this study, VviMYB86 was also confirmed to be affected by light, since its transcript levels were enhanced when VviMYB86 transgenic callus and the WT callus were subjected to light conditions.
However, anthocyanin production and the expression of corresponding genes were repressed in the light-exposed VviMYB86 transgenic lines compared with those in the WT-Light callus. These results demonstrated that VviMYB86 obviously repressed anthocyanin synthesis under light conditions in grapes. The presence of both positive and negative regulators of anthocyanin-related structural genes keeps the anthocyanin levels in planta in balance (Ricardo Perez-Diaz et al., 2016). The expression pattern of VviMYB86 is opposite that of anthocyanin biosynthesis, as well as that of VviMYBA1, throughout the entire developmental stages of grape berries (Cutanda-Perez et al., 2009). These negative correlations in developmental expression have also been observed with VviMYBC2-L2 and VviMYB4-like (Ricardo Perez-Diaz et al., 2016; Zhu et al., 2019), suggesting that these MYB TFs are expressed in grapevine organs to inhibit ectopic anthocyanin accumulation in grape berries.
VviMYB86 Oppositely Regulates PA and Anthocyanin Biosynthesis in Grapes
Several R2R3-MYB TFs have been verified to have a broad effect on the regulatory function of flavonoid biosynthesis in grape berries (Wei et al., 2020). In contrast to these functionally specialized types of R2R3-MYB TFs, such as VviMYBPA1 (Bogs et al., 2007), VviMYBPA2 (Terrier et al., 2009), and VviMYBPAR (Koyama et al., 2014), VviMYB86 modulates both PA and anthocyanin biosynthesis in grape berries. Unlike VviMYB5a (Deluc et al., 2006) and VviMYB5b (Deluc et al., 2008), both of which positively regulate PA and anthocyanin biosynthesis in grape berries, VviMYB86 had different regulatory mechanisms for PAs and anthocyanins. VviMYB86 positively regulated the PA biosynthesis branch but negatively affected anthocyanin biosynthesis in grape berries. Recently, MYB TFs with different regulatory mechanisms for different biosynthesis branches of flavonoids have also been found in other plant species. In the Chinese narcissus (Narcissus tazetta L. var. chinensis), NtMYB3 induces PA accumulation by repressing flavonol biosynthesis (Anwar et al., 2019). In Medicago truncatula, MtPAR promotes PA accumulation by directly repressing isoflavone biosynthesis and by redirecting the flux of the immediate precursors into the PA biosynthesis branch (Li et al., 2016). In MtMYB14-overexpressing hairy roots, PA accumulation is strongly induced and anthocyanin production is decreased by half (Liu et al., 2014). CsMYB5a-overexpressing tobacco plants exhibit downregulated anthocyanin accumulation but present a high polymeric PA content in the flowers (Jiang et al., 2018). Hence, the results of our study help to elucidate the role and mechanism of VviMYB86 in the regulatory network governing the biosynthesis of anthocyanins and PAs in grape berries.
In summary, the results of this study show that VviMYB86 is a candidate gene implicated in the regulation of different branches of the flavonoid biosynthesis pathway in grape berries. Our results prove that VviMYB86 controls PA biosynthesis, primarily by affecting the expression of the two LAR genes in the Arabidopsis transient transformation system, as well as in grape callus, when overexpressed. Moreover, VviMYB86 is observed to repress anthocyanin biosynthesis branch in grapes by downregulating the transcript levels of VviANS and VviUFGT. The results of this study help to elucidate the regulatory mechanism governing the expression of the two LAR genes in grape berries and provide new insights into the role and mechanism of VviMYB86 in the regulation of the metabolic flux between the PA and anthocyanin biosynthesis branches in grape berries.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary Material.
Author Contributions
CD and JW conceived and guided the experiments. JC performed the research, analyzed the data, and wrote the original manuscript. KY and YS perfected the research scheme. All authors critically revised the article.
Funding
This research was funded by China Agriculture Research System (CARS-29) and National Natural Science Foundation of China, Grant Number U20A2042.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Daqi Fu, Hongliang Zhu, and Xiuqin Wang (all from China Agricultural University) for giving us pCXSN vector, pGreen II 0800-LUC, and pCAMBIA 1301 vectors, and wild type grape callus induced from the skins of Vitis vinifera L. cv. Cabernet Sauvignon as gifts.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2020.613677/full#supplementary-material
Footnotes
- ^ http://genomes.cribi.unipd.it/grape/
- ^ http://www.ebi.ac.uk/Tools/pfa/iprscan/
- ^ http://web.expasy.org/protparam/
- ^ http://genomes.cribi.unipd.it/grape/
- ^ http://bioinformatics.psb.ugent.be/webtools/plantcare/html/
- ^ https://deepgreen.dpb.carnegiescience.edu/
- ^ https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE129916
References
Anwar, M., Yu, W., Yao, H., Zhou, P., Allan, A. C., and Zeng, L. (2019). NtMYB3, an R2R3-MYB from Narcissus, regulates flavonoid biosynthesis. Int. J. Mol. Sci. 20:5456. doi: 10.3390/ijms20215456
Azuma, A., Koshita, Y., and Kobayashi, S. (2012). Flavonoid biosynthesis-related genes in grape skin are differentially regulated by temperature and light conditions. Planta 236, 1067–1080. doi: 10.1007/s00425-012-1650-x
Bogs, J., Downey, M. O., Harvey, J. S., Ashton, A. R., Tanner, G. J., and Robinson, S. P. (2005). Proanthocyanidin synthesis and expression of genes encoding leucoanthocyanidin reductase and anthocyanidin reductase in developing grape berries and grapevine leaves. Plant Physiol. 139, 652–663. doi: 10.1104/pp.105.064238
Bogs, J., Jaffe, F. W., Takos, A. M., Walker, A. R., and Robinson, S. P. (2007). The grapevine transcription factor VvMYBPA1 regulates proanthocyanidin synthesis during fruit development. Plant Physiol. 143, 1347–1361. doi: 10.1104/pp.106.093203
Braidot, E., Zancani, M., Petrussa, E., Peresson, C., Bertolini, A., Patui, S., et al. (2008). Transport and accumulation of flavonoids in grapevine (Vitis vinifera L.). Plant Signal. Behav. 3, 626–632. doi: 10.4161/psb.3.9.6686
Butelli, E., Titta, L., Giorgio, M., Mock, H. P., Matros, A., Peterek, S., et al. (2008). Enrichment of tomato fruit with health-promoting anthocyanins by expression of select transcription factors. Na. Biotechnol. 26, 1301–1308. doi: 10.1038/nbt.1506
Cavallini, E., Matus, J. T., Finezzo, L., Zenoni, S., Loyola, R., Guzzo, F., et al. (2015). The phenylpropanoid pathway is controlled at different branches by a set of R2R3-MYB C2 repressors in grapevine. Plant Physiol. 167, 1448–1470. doi: 10.1104/pp.114.256172
Chen, L., Hu, B., Qin, Y., Hu, G., and Zhao, J. (2019). Advance of the negative regulation of anthocyanin biosynthesis by myb transcription factors. Plant Physiol. Bioch. 136, 178–187. doi: 10.1016/j.plaphy.2019.01.024
Cheng, J., Yu, K., Zhang, M., Shi, Y., Duan, C., and Wang, J. (2020). The effect of light intensity on the expression of leucoanthocyanidin reductase in grapevine calluses and analysis of its promoter activity. Genes 11:1156. doi: 10.3390/genes11101156
Cos, P., De Bruyne, T., Hermans, N., Apers, S., Vanden Berghe, D., and Vlietinck, A. J. (2004). Proanthocyanidins in health care: Current and new trends. Curr. Med. Chem. 11, 1345–1359. doi: 10.2174/0929867043365288
Cutanda-Perez, M. C., Ageorges, A., Gomez, C., Vialet, S., Terrier, N., Romieu, C., et al. (2009). Ectopic expression of VlmybA1 in grapevine activates a narrow set of genes involved in anthocyanin synthesis and transport. Plant Mol. Biol. 69, 633–648. doi: 10.1007/s11103-008-9446-x
Deluc, L., Barrieu, F., Marchive, C., Lauvergeat, V., Decendit, A., Richard, T., et al. (2006). Characterization of a grapevine R2R3-MYB transcription factor that regulates the phenylpropanoid pathway. Plant Physiol. 140, 499–511. doi: 10.1104/pp.105.067231
Deluc, L., Bogs, J., Walker, A. R., Ferrier, T., Decendit, A., Merillon, J. M., et al. (2008). The transcription factor VvMYB5b contributes to the regulation of anthocyanin and proanthocyanidin biosynthesis in developing grape berries. Plant Physiol. 147, 2041–2053. doi: 10.1104/pp.108.118919
Dixon, R. A., Xie, D. Y., and Sharma, S. B. (2005). Proanthocyanidins - a final frontier in flavonoid research? New Phytol. 165, 9–28. doi: 10.1111/j.1469-8137.2004.01217.x
Downey, M. O., Dokoozlian, N. K., and Krstic, M. P. (2006). Cultural practice and environmental impacts on the flavonoid composition of grapes and wine: A review of recent research. Am. J. Enol. Vitic. 57, 257–268.
Fujita, A., Soma, N., Goto-Yamamoto, N., Mizuno, A., Kiso, K., and Hashizume, K. (2007). Effect of shading on proanthocyanidin biosynthesis in the grape berry. J. Jap. Soc.Hortic. Sci. 76, 112–119. doi: 10.2503/jjshs.76.112
Hichri, I., Barrieu, F., Bogs, J., Kappel, C., Delrot, S., and Lauvergeat, V. (2011). Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J. Exp. Bot. 62, 2465–2483. doi: 10.1093/jxb/erq442
Huang, Y. F., Vialet, S., Guiraud, J. L., Torregrosa, L., Bertrand, Y., Cheynier, V., et al. (2014). A negative MYB regulator of proanthocyanidin accumulation, identified through expression quantitative locus mapping in the grape berry. New Phytol. 201, 795–809. doi: 10.1111/nph.12557
Jia, L., Clegg, M. T., and Jiang, T. (2004). Evolutionary dynamics of the DNA-binding domains in putative R2R3-MYB genes identified from rice subspecies indica and japonica genomes. Plant Physiol. 134, 575–585. doi: 10.1104/pp.103.027201
Jiang, X., Huang, K., Zheng, G., Hou, H., Wang, P., Jiang, H., et al. (2018). CsMYB5a and CsMYB5e from Camellia sinensis differentially regulate anthocyanin and proanthocyanidin biosynthesis. Plant Sci. 270, 209–220. doi: 10.1016/j.plantsci.2018.02.009
Jun, J. H., Liu, C., Xiao, X., and Dixon, R. A. (2015). The transcriptional repressor MYB2 regulates both spatial and temporal patterns of proanthocyandin and anthocyanin pigmentation in Medicago truncatula. Plant Cell 27, 2860–2879. doi: 10.1105/tpc.15.00476
Koyama, K., Ikeda, H., Poudel, P. R., and Goto-Yamamoto, N. (2012). Light quality affects flavonoid biosynthesis in young berries of Cabernet Sauvignon grape. Phytochemistry 78, 54–64. doi: 10.1016/j.phytochem.2012.02.026
Koyama, K., Numata, M., Nakajima, I., Goto-Yamamoto, N., Matsumura, H., and Tanaka, N. (2014). Functional characterization of a new grapevine MYB transcription factor and regulation of proanthocyanidin biosynthesis in grapes. J. Exp. Bot. 65, 4433–4449. doi: 10.1093/jxb/eru213
Kumar, S., Stecher, G., and Tamura, K. (2016). MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 33, 1870–1874. doi: 10.1093/molbev/msw054
Lee, J. (2017). Light exclusion influence on grape anthocyanin. Heliyon 3:e00243. doi: 10.1016/j.heliyon.2017.e00243
Li, P. H., Dong, Q., Ge, S. J., He, X. Z., Verdier, J., Li, D. Q., et al. (2016). Metabolic engineering of proanthocyanidin production by repressing the isoflavone pathways and redirecting anthocyanidin precursor flux in legume. Plant Biotechnol. J. 14, 1604–1618. doi: 10.1111/pbi.12524
Li, Y. G., Tanner, G., and Larkin, P. (1996). The DMACA-HCl protocol and the threshold proanthocyanidin content for bloat safety in forage legumes. J. Sci. Food Agr. 70, 89–101. doi: 10.1002/(sici)1097-0010(199601)70:1
Li, Y. Q., Shan, X. T., Zhou, L. D., Gao, R. F., Yang, S., Wang, S. C., et al. (2019). The R2R3-MYB factor FhMYB5 From Freesia hybrida contributes to the regulation of anthocyanin and proanthocyanidin in biosynthesis. Front. Plant Sci. 9:1935. doi: 10.3389/fpls.2018.01935
Liu, C. G., Jun, J. H., and Dixon, R. A. (2014). MYB5 and MYB14 play pivotal roles in seed coat polymer biosynthesis in Medicago truncatula. Plant Physiol. 165, 1424–1439. doi: 10.1104/pp.114.241877
Liu, M. Y., Song, C. Z., Chi, M., Wang, T. M., Zuo, L. L., Li, X. L., et al. (2016). The effects of light and ethylene and their interaction on the regulation of proanthocyanidin and anthocyanin synthesis in the skins of Vitis vinifera berries. Plant Growth Regul. 79, 377–390. doi: 10.1007/s10725-015-0141-z
Luo, X., Zhao, H., Yao, P., Li, Q., Huang, Y., Li, C., et al. (2018). An R2R3-MYB transcription factor FtMYB15 involved in the synthesis of anthocyanin and proanthocyanidins from Tartary buckwheat. J. Plant Growth Regul. 37, 76–84. doi: 10.1007/s00344-017-9709-9703
Mathur, J., and Koncz, C. (1998). PEG-mediated protoplast transformation with naked DNA. Methods Mole. Biol. 82, 267–276. doi: 10.1385/0-89603-391-0, 267
Matus, J. T., Loyola, R. V. A., Pena-Neira, A., Bordeu, E., Arce-Johnson, P., and Alcalde, J. A. (2009). Post-veraison sunlight exposure induces myb-mediated transcriptional regulation of anthocyanin and flavonol synthesis in berry skins of Vitis vinifera. J. Exp. Bot. 60, 853–867. doi: 10.1093/jxb/ern336
Meng, N., Wei, Y., Gao, Y., Yu, K., Cheng, J., Li, X. Y., et al. (2020). Characterization of transcriptional expression and regulation of carotenoid cleavage dioxygenase 4b in grapes. Front. Plant Sci. 11:483. doi: 10.3389/fpls.2020.00483
Mu, L., He, F., Pan, Q. H., Zhou, L., and Duan, C. Q. (2014). Screening and verification of late embryogenesis abundant protein interacting with anthocyanidin reductase in grape berries. Vitis 53, 81–87.
Peters, D. J., and Constabel, C. P. (2002). Molecular analysis of herbivore-induced condensed tannin synthesis: cloning and expression of dihydroflavonol reductase from trembling aspen (Populus tremuloides). Plant J. 32, 701–712. doi: 10.1046/j.1365-313x.2002.01458.x
Ricardo Perez-Diaz, J., Perez-Diaz, J., Madrid-Espinoza, J., Gonzalez-Villanueva, E., Moreno, Y., and Ruiz-Lara, S. (2016). New member of the R2R3-MYB transcription factors family in grapevine suppresses the anthocyanin accumulation in the flowers of transgenic tobacco. Plant Mol. Biol. 90, 63–76. doi: 10.1007/s11103-015-0394-y
Ruijter, J. M., Ramakers, C., Hoogaars, W. M. H., Karlen, Y., Bakker, O., Van Den Hoff, M. J. B., et al. (2009). Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data. Nucleic Acids Res. 37:e45. doi: 10.1093/nar/gkp045
Shang, Y., Yan, L., Liu, Z. Q., Cao, Z., Mei, C., Xin, Q., et al. (2010). The Mg-chelatase H subunit of Arabidopsis antagonizes a group of WRKY transcription repressors to relieve ABA-responsive genes of inhibition. Plant Cell 22, 1909–1935. doi: 10.1105/tpc.110.073874
Stojanovic, J., and Silva, J. L. (2006). Influence of osmoconcentration, continuous high-frequency ultrasound and dehydration on properties and microstructure of rabbiteye blueberries. Dry.Technol. 24, 165–171. doi: 10.1080/07373930600558995
Sun, R. Z., Cheng, G., Li, Q., He, Y. N., Wang, Y., Lan, Y. B., et al. (2017). Light-induced variation in phenolic compounds in Cabernet Sauvignon grapes (Vitis vinifera L.) involves extensive transcriptome reprogramming of biosynthetic enzymes, transcription factors, and phytohormonal regulators. Front. Plant Sci. 8:547. doi: 10.3389/fpls.2017.00547
Sun, R. Z., Cheng, G., Li, Q., Zhu, Y. R., Zhang, X., Wang, Y., et al. (2019). Comparative physiological, metabolomic, and transcriptomic analyses reveal developmental stage-dependent effects of cluster bagging on phenolic metabolism in Cabernet Sauvignon grape berries. BMC Plant Biol. 19:583. doi: 10.1186/s12870-019-2186-z
Sun, R. Z., Pan, Q. H., Duan, C. Q., and Wang, J. (2015). Light response and potential interacting proteins of a grape flavonoid 3′ -hydroxylase gene promoter. Plant Physiol. Bioch. 97, 70–81. doi: 10.1016/j.plaphy.2015.09.016
Tanner, G. J., Francki, K. T., Abrahams, S., Watson, J. M., Larkin, P. J., and Ashton, A. R. (2003). Proanthocyanidin biosynthesis in plants - Purification of legume leucoanthocyanidin reductase and molecular cloning of its cDNA. J. Biol. Chem. 278, 31647–31656. doi: 10.1074/jbc.M302783200
Terrier, N., Torregrosa, L., Ageorges, A., Vialet, S., Verries, C., Cheynier, V., et al. (2009). Ectopic expression of VvMybPA2 promotes proanthocyanidin biosynthesis in grapevine and suggests additional targets in the pathway. Plant Physiol. 149, 1028–1041. doi: 10.1104/pp.108.131862
Van Eck, J., Kirk, D. D., and Walmsley, A. M. (2006). Tomato (Lycopersicum esculentum). Methods Mole. Biol. 343, 459–473. doi: 10.1385/1-59745-130-4:459
Vidal, S., Francis, L., Guyot, S., Marnet, N., Kwiatkowski, M., Gawel, R., et al. (2003). The mouth-feel properties of grape and apple proanthocyanidins in a wine-like medium. J. Sci. Food Agr. 83, 564–573. doi: 10.1002/jsfa.1394
Wang, F., Tong, W., Zhu, H., Kong, W., Peng, R., Liu, Q., et al. (2016). A novel Cys(2)/His(2) zinc finger protein gene from sweetpotato, IbZFP1, is involved in salt and drought tolerance in transgenic Arabidopsis. Planta 243, 783–797. doi: 10.1007/s00425-015-2443-2449
Wang, H., Wang, W., Zhan, J., Huang, W., and Xu, H. (2015). An efficient PEG-mediated transient gene expression system in grape protoplasts and its application in subcellular localization studies of flavonoids biosynthesis enzymes. Sci. Hortic. 191, 82–89. doi: 10.1016/j.scienta.2015.04.039
Wang, L. J., Lu, W. X., Ran, L. Y., Dou, L. W., Yao, S., Hu, J., et al. (2019). R2R3-MYB transcription factor MYB6 promotes anthocyanin and proanthocyanidin biosynthesis but inhibits secondary cell wall formation in Populus tomentosa. Plant J. 99, 733–751. doi: 10.1111/tpj.14364
Wang, W. L., Wang, Y. X., Li, H., Liu, Z. W., Cui, X., and Zhuang, J. (2018). Two MYB transcription factors (CsMYB2 and CsMYB26) are involved in flavonoid biosynthesis in tea plant Camellia sinensis (L.) O. Kuntze. BMC Plant Biol. 18:288. doi: 10.1186/s12870-018-1502-1503
Wei, X., Ju, Y., Ma, T., Zhang, J., Fang, Y., and Sun, X. (2020). New perspectives on the biosynthesis, transportation, astringency perception and detection methods of grape proanthocyanidins. Crit. Rev. Food Sci. Nutr. 17, 1–27. doi: 10.1080/10408398.2020.1777527
Wen, Y. Q., Zhong, G. Y., Gao, Y., Lan, Y. B., Duan, C. Q., and Pan, Q. H. (2015). Using the combined analysis of transcripts and metabolites to propose key genes for differential terpene accumulation across two regions. BMC Plant Biol. 15:240. doi: 10.1186/s12870-015-0631-631/
Williams, C. E., and Grotewold, E. (1997). Differences between plant and animal MYB domains are fundamental for DNA binding activity, and chimeric MYB domains have novel DNA binding specificities. J. Biol. Chem. 272, 563–571. doi: 10.1074/jbc.272.1.563
Xie, D. Y., Sharma, S. B., Paiva, N. L., Ferreira, D., and Dixon, R. A. (2003). Role of anthocyanidin reductase, encoded by BANYULS in plant flavonoid biosynthesis. Science 299, 396–399. doi: 10.1126/science.1078540
Xu, W., Grain, D., Bobet, S., Le Gourrierec, J., Thevenin, J., Kelemen, Z., et al. (2014). Complexity and robustness of the flavonoid transcriptional regulatory network revealed by comprehensive analyses of MYB-bHLH-WDR complexes and their targets in Arabidopsis seed. New Phytol. 202, 132–144. doi: 10.1111/nph.12620
Xu, W. J., Dubos, C., and Lepiniec, L. (2015). Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci. 20, 176–185. doi: 10.1016/j.tplants.2014.12.001
Yu, K., Jun, J. H., Duan, C., and Dixon, R. A. (2019). VvLAR1 and VvLAR2 are bifunctional enzymes for proanthocyanidin biosynthesis in grapevine. Plant Physiol. 180, 1362–1374. doi: 10.1104/pp.19.00447
Zhang, X., Allan, A. C., Yi, Q., Chen, L., Li, K., Shu, Q., et al. (2011). Differential gene expression analysis of Yunnan Red Pear, Pyrus pyrifolia, during fruit skin coloration. Plant Mol. Biol. Rep. 29, 305–314. doi: 10.1007/s11105-010-0231-z
Zhou, H., Kui, L. W., Liao, L., Chao, G., Lu, Z. Q., Allan, A. C., et al. (2015). Peach MYB7 activates transcription of the proanthocyanidin pathway gene encoding leucoanthocyanidin reductase, but not anthocyanidin reductase. Front. Plant Sci. 6:908. doi: 10.3389/fpls.2015.00908
Zhou, H., Ling-Wang, K., Wang, F., Espley, R. V., Ren, F., Zhao, J., et al. (2019). Activator−type R2R3 MYB genes induce a repressor−type R2R3 MYB gene to balance anthocyanin and proanthocyanidin accumulation. New Phytol. 221, 1919–1934. doi: 10.1111/nph.15486
Zhu, Z. G., Li, G. R., Liu, L., Zhang, Q. T., Han, Z., Chen, X. S., et al. (2019). A R2R3-MYB transcription factor, VvMYBC2L2, functions as a transcriptional repressor of anthocyanin biosynthesis in grapevine (Vitis vinifera L.). Molecules 24:92. doi: 10.3390/molecules24010092
Zimmermann, I. M., Heim, M. A., Weisshaar, B., and Uhrig, J. F. (2004). Comprehensive identification of Arabidopsis thaliana MYB transcription factors interacting with R/B-like BHLH proteins. Plant J. 40, 22–34. doi: 10.1111/j.1365-313X.2004.02183.x
Keywords: flavonoid, grape (Vitis vinifera L.), leucoanthocyanidin reductase, light, regulation, R2R3-MYB
Citation: Cheng J, Yu K, Shi Y, Wang J and Duan C (2021) Transcription Factor VviMYB86 Oppositely Regulates Proanthocyanidin and Anthocyanin Biosynthesis in Grape Berries. Front. Plant Sci. 11:613677. doi: 10.3389/fpls.2020.613677
Received: 05 October 2020; Accepted: 22 December 2020;
Published: 13 January 2021.
Edited by:
Supaart Sirikantaramas, Chulalongkorn University, ThailandReviewed by:
Yuepeng Han, Wuhan Botanical Garden, Chinese Academy of Sciences, ChinaQian Shen, Shanghai Jiao Tong University, China
Laura Jaakola, Arctic University of Norway, Norway
Copyright © 2021 Cheng, Yu, Shi, Wang and Duan. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Changqing Duan, Y2hxZHVhbkBjYXUuZWR1LmNu
 Jing Cheng
Jing Cheng Keji Yu1,2,3
Keji Yu1,2,3 Jun Wang
Jun Wang Changqing Duan
Changqing Duan