- 1Department of Life Sciences, Graduate School of Arts and Sciences, The University of Tokyo, Tokyo, Japan
- 2Department of Biology, Tokyo Gakugei University, Tokyo, Japan
- 3RIKEN Center for Sustainable Resource Science, Yokohama, Japan
- 4National Institute for Basic Biology, Okazaki, Japan
- 5Department of Basic Biology, School of Life Science, Graduate University for Advanced Studies (SOKENDAI), Okazaki, Japan
Plant leaves display abundant morphological richness yet grow to characteristic sizes and shapes. Beginning with a small number of undifferentiated founder cells, leaves evolve via a complex interplay of regulatory factors that ultimately influence cell proliferation and subsequent post-mitotic cell enlargement. During their development, a sequence of key events that shape leaves is both robustly executed spatiotemporally following a genomic molecular network and flexibly tuned by a variety of environmental stimuli. Decades of work on Arabidopsis thaliana have revisited the compensatory phenomena that might reflect a general and primary size-regulatory mechanism in leaves. This review focuses on key molecular and cellular events behind the organ-wide scale regulation of compensatory mechanisms. Lastly, emerging novel mechanisms of metabolic and hormonal regulation are discussed, based on recent advances in the field that have provided insights into, among other phenomena, leaf-size regulation.
1 Introduction
Pioneering studies on Arabidopsis unveiled the basis of genetic control of major plant organs, such as leaves, flowers, and roots (Irish, 2010; Petricka et al., 2012; González et al., 2012; Kalve et al., 2014). Although most developmental events and concomitant cellular processes have been described thoroughly, it is only during the past few decades that we have integrated the molecular pathways (i.e. relating key genes, receptors, sensors, etc.) behind plant morphogenesis.
Leaf emergence and polarity acquisition and the differentiation of the various cell types and tissues have been described in detail (Asnacios and Hamant, 2012; Lau and Bergmann, 2012; Du et al., 2018; Ali et al., 2020; Vercruysse et al., 2020; Gorelova et al., 2021; Liu et al., 2021). Yet, the core molecular framework behind leaf-size regulation remains unclear. We argue that this relates to a conserved feature of growing organs in seed plants: compensation, a phenomenon whereby cell size compensates for cell division in the establishment of organ size. This is what this review focuses on, taking the leaf as a model system.
2 Compensation: One phenotype, several means
Leaves are typically flat. They capture sunlight, and convert carbon dioxide into carbohydrates by photosynthesis to sustain the plant autotrophic lifestyle. Also, plant leaves are polarized, possessing two structurally different sides, the adaxial and abaxial sides, which are specialized for light capture/photosynthesis and gas exchange, respectively (Bowman et al., 2002; Dkhar and Pareek, 2014; Dow and Bergmann, 2014; and the references therein). Hence, plant leaves can be viewed as expandible structural units, which emerge and grow to highly reproducible predetermined sizes and shapes, which are often used as traits in taxonomy (Tsukaya, 2003). Plant leaves, in general, lack a self-renewing meristem and thus cannot grow indefinitely. Leaf development usually proceeds via initiation, growth, and maturation stages (Tsukaya, 2002a; Kalve et al., 2014; Tsukaya, 2014; Tsukaya, 2018).
Leaf cells can divide only for a fixed period of time, which is crucial to determine the number of cells within the leaf. Then, their proliferative ability is gradually lost, as they enter a second stage of post-mitotic cell differentiation marked by a considerable increase in cell size accompanied by increased vacuole volume and cell wall synthesis (Johnson and Lenhard, 2011; González et al., 2012; Powell and Lenhard, 2012). Leaf size is reproducible under controlled light, temperature, and nutritional regimes. In addition, size increase in plants is an irreversible process. That is because morphogenesis in the plant kingdom, unlike the animal kingdom, does not rely on cell contractility, cell migration and cell death. Therefore, leaves are also a simpler model system to decipher what makes organs stop growing upon reaching an appropriate size (Beemster et al., 2006; Micol, 2009; Tsukaya, 2018).
Molecular genetics have identified a plethora of genes contributing to leaf-size control (González et al., 2012; Hepworth and Lenhard, 2014). This has led to draw an overall picture of the gene regulatory network operating during leaf development (González and Inzé, 2015; Wang et al., 2021). The dynamic interactions among key genetic components have recently been uncovered. For instance, DELLA proteins repress gibberellin signaling to modulate cell proliferation and expansion (Sun and Gubler, 2004; de Lucas et al., 2008; Achard et al., 2009). Such DELLA-mediated growth suppression is executed by GROWTH REGULATORY FACTOR (GRF) transcriptional factors at least in cold stress response (Lantzouni et al., 2020). GRFs act within a core module to primarily orchestrate cell proliferation together with GRF INTERACTING FACTOR (GIF) transcriptional co-activators including GIF1/ANGUSTIFOLIA3 (AN3) (Kim and Kende, 2004; Horiguchi et al., 2005; Kim and Tsukaya, 2015; Liebsch and Palatnik, 2020). This module is also connected downstream of TCP transcriptional factors and their targeting microRNA miR319 (Rodriguez et al., 2010; Schommer et al., 2014), which are recognized as important components of leaf lamina growth (Nath et al., 2003; Efroni et al., 2013; Das Gupta and Nath, 2015; Challa et al., 2021; Rath et al., 2022). Although the above findings delineate the hierarchical organization and interconnections among the genetic network governing leaf-size control, this topic has already been covered with an increasing pace. A more exhaustive synthesis of our current knowledge can be found elsewhere (Vercruysse et al., 2020).
In the simplest scenario, leaf size would be defined as the linear function of cell number and cell size. However, in leaf primordia, failure in the proliferative stage to produce sufficient cells triggers excessive cell expansion, the so-called compensation (Tsukaya, 2002b; Beemster et al., 2003; Beemster et al., 2006; Tsukaya, 2008). Because cell division precedes cell differentiation (which also involves cell expansion) in a region confined to the leaf primordia basal part, the proliferative stage likely generates intrinsic signals that affect the final cell size during the following differentiation stage. This poses a question of how the above cellular processes, which occur in distinct regions of leaf primordia, are coordinated during development. Given the importance of cell division and expansion in leaf-size control, compensation has emerged as a key phenomenon to uncover how the interconnection between cell division and expansion is achieved. However the molecular basis behind compensation remains unclear.
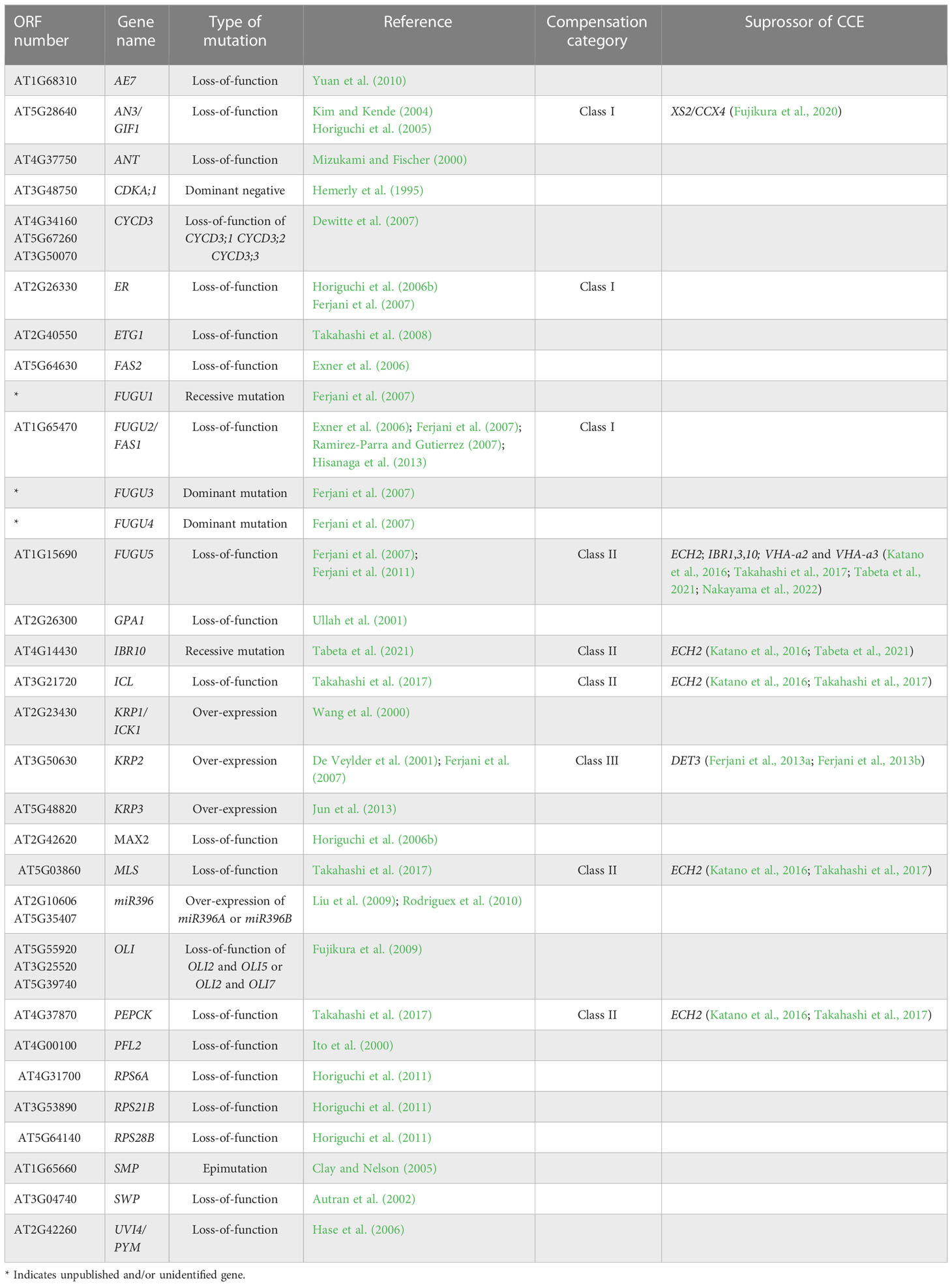
Large scale genetic screening has uncovered some of the compensatory mechanisms through the identification of a large number of mutants and transgenics displaying compensation (Table 1; Horiguchi et al., 2005; Horiguchi et al., 2006a; Horiguchi et al., 2006b; Fujikura et al., 2007a, Fujikura et al., 2007b, Fujikura et al., 2009). For instance, kinematic analyses of cell size dynamism unveiled the presence of three classes of compensation, based on cell expansion mode (Ferjani et al., 2007). More specifically, while class I has an enhanced post-mitotic cell expansion rate (seen in angustifolia [an]3-4, fugu2-1/fasciata [fas]1-6, and erecta[er]-102); class II has an extended post-mitotic cell expansion period (seen in fugu5-1, icl-2, mls-2, pck1-2, and ibr10), and class III has an increased size of dividing cells (seen in KIP-RELATED PROTEIN 2 [KRP2] overexpressing plants) (De Veylder et al., 2001; Ferjani et al., 2007; Ferjani et al., 2008; Ferjani et al., 2010; Ferjani et al., 2013a; Ferjani et al., 2013b; Ferjani et al., 2014a; Ferjani et al., 2014b; Katano et al., 2016; Takahashi et al., 2017; Tabeta et al., 2021).
Taking the above into account, compensation-exhibiting mutants have altered coordination between cell division and expansion (Ferjani et al., 2008; Nakayama et al., 2022). Furthermore, cell-autonomous and non-cell-autonomous pathways have been demonstrated to be implicated in compensation (Ferjani et al., 2008; Kawade et al., 2010; Ferjani et al., 2013a; Ferjani et al., 2014a; Nozaki et al., 2020). Finally, the fact that compensation occurs in a wide range of plant species other than Arabidopsis—including tobacco, rice, snapdragon, and North American lake cress—suggests that the developmental mechanisms that trigger compensation are widely conserved at least in seed plants (Hemerly et al., 1995; Barrôco et al., 2006; Delgado-Benarroch et al., 2009; Horiguchi and Tsukaya, 2011; Amano et al., 2015). However, despite their similar cellular phenotypes, compensation refers to a group of heterogeneous processes driven by different mechanisms (Ferjani et al., 2007; Ferjani et al., 2008; Hisanaga et al., 2015).
3 Underpinnings of compensatory coordination between cell division and expansion
Cell-to-cell communication is an effective way to coordinate cellular processes in time and space and hence to realize stereotyped leaf size. Besides classical anatomy (Szymkowiak and Sussex, 1996), a series of works on the model plant Arabidopsis identified signaling pathways that facilitate cell-to-cell communication (Sieburth et al., 1998; Kim et al., 2003; Serralbo et al., 2006; Savaldi-Goldstein et al., 2007; Eriksson et al., 2010). However, the compensatory interplay between cell proliferation and post-mitotic cell expansion during the induction of compensatory cell enlargement (CCE) is ill-known. Because this knowledge on cellular dynamics is instrumental in determining future directions aiming to delineate the holistic mechanisms of compensation, it has been recently investigated using leaves chimeric for the key elements of class I, II, or III compensation (Kawade et al., 2010; Gunji et al., 2022) (Figure 1). The above investigations revealed that both cell-autonomous and non-cell autonomous mechanisms are involved in CCE. In the following sections, we summarize the major outcome of these studies.
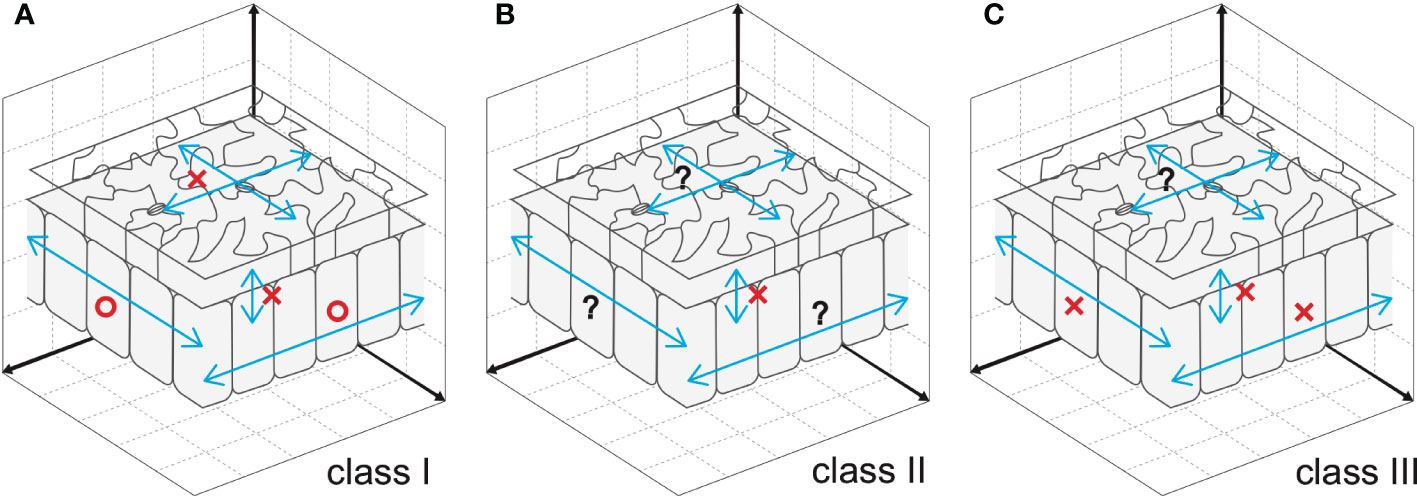
Figure 1 Cellular spatial relationships during induction of CCE. (A) Mesophyll cells exploit cell-to-cell communication to stimulate CCE in response to deficient cell proliferation in an3, which has class I compensation (Kawade et al., 2010). By contrast, epidermal cells trigger CCE in a cell-autonomous manner, preventing cell-to-cell communication across tissue layers (Nozaki et al., 2020). (B) Cell-to-cell communication between epidermal and mesophyll cells is absent in fugu5, which has class II compensation (Gunji et al., 2022). Given that excess PPi inhibits metabolic reactions, fugu5-mediated induction of CCE is likely controlled in a cell-autonomous manner. (C) Direct inhibition of cell cycle progression by KRP overexpression, a representative of class III compensation, induces CCE in a cell- and tissue-autonomous manner. This was determined using chimeric KRP2-overexpressing leaves generated using the Cre/lox-P system (Kawade et al., 2010) and KRP1-overexpressing leaves with a tissue-specific expression system (Bemis and Torii, 2007). Epidermal cell-to-cell communication remains to be explored. Circles and crosses indicate, respectively, the presence and absence of cell-to-cell communication that stimulates CCE. Question marks are added when cell-to-cell communication is still untested.
3.1 an3-mediated class I compensation: contribution of plasmodesmata
The Cre/lox-P system enables heat shock-dependent induction or suppression of AN3 expression in an an3 or wild-type genetic background, respectively (Kawade et al., 2010). When the expression of AN3 is induced in an3 subepidermal cells in very-early-stage leaf primordia, an AN3-expressing sector forms among the an3 cell population during leaf development. This leaf chimera exhibited CCE irrespective of cellular genotype (i.e., an3 and AN3-expressing cells). Similar results were obtained when leaf chimeras were generated by patchy deletion of the expression of AN3 in the wild-type genetic background. These observations indicated that the an3-mediated induction of CCE occurs via cell-to-cell communication in leaf mesophyll tissue. Although the ability to stimulate CCE propagates in a non-cell-autonomous manner, the signaling range is confined to one half of the leaf partitioned by the midrib (Kawade et al., 2010). In contrast to mesophyll tissue, cell-autonomous behavior to stimulate CCE was observed in epidermal tissue of AN3 leaf chimeras generated using the Cre/lox-P system or a tissue-specific expression system (Nozaki et al., 2020). In summary, the cell-to-cell communication that induces CCE in an3 is cell-type dependent (Figure 1A).
It is plausible that this property is attributable to the plasmodesmata aperture size, which varies in time and space to control symplasmic connections (Crawford and Zambryski, 2001; Roberts et al., 2001; Burch-Smith and Zambryski, 2010; Fitzgibbon et al., 2013). Several chaperones and RNA exosomes mediate selective symplasmic transport of signaling molecules, including proteins and mRNAs (Xu et al., 2011; Kitagawa et al., 2022). Apoplastic transport is an alternative pathway for the exchange of signaling molecules between cells. This machinery has been characterized in the context of plant defense, in which bursts of reactive oxygen species (ROS) upon pathogen infection elicit phytohormone signaling (Wendehenne et al., 2014; Noctor et al., 2018). ROS-mediated signaling contributes to the balance between cell proliferation and post-mitotic cell expansion in growing sepals and roots (Tsukagoshi et al., 2010; Tsukagoshi, 2012; Hong et al., 2016; Mabuchi et al., 2018).
In leaf development, the salicylic acid (SA) response is involved in CCE in an3. The extra-small sisters (xs) mutants isolated from a large leaf-size and -shape mutant collection (Horiguchi et al., 2006b) showed normal cell proliferation but compromised post-mitotic cell expansion (Fujikura et al., 2007a). Among them, the xs2 mutant, which harbors a mutation in a gene encoding the putative endomembrane H+-dependent K+ transporter CATION CALCIUM EXCHANGER 4 (CCX4), showed ROS overproduction and an elevated SA response (Fujikura et al., 2020). Importantly, eliminating XS2/CCX4 function in the an3 mutant fully suppressed CCE (Fujikura et al., 2020), indicating crosstalk between CCE and the SA response. This finding provides insight into the mechanism by which cell proliferation and post-mitotic cell expansion are coordinated beyond the cellular level.
Whereas the AN3 protein is capable of moving between cells to promote cell proliferation (Kawade et al., 2013; Kawade et al., 2017), the aforementioned cell-to-cell communication for CCE is observed in the an3 genetic background. The non-cell-autonomous signaling that activates CCE in an3, in addition to the non-cell-autonomous signaling downstream of AN3 protein movement to promote cell proliferation, are both largely unclear. Enhanced knowledge of AN3-related genetic regulatory networks will be instrumental in resolving these issues at the molecular level (Vercruyssen et al., 2014; Nelissen et al., 2015; Zhang et al., 2018; Jun et al., 2019; Fujikura et al., 2020; Hussain et al., 2022).
3.2 fugu5-mediated class II compensation: contribution of metabolic regulation
Although a factor(s) produced in the mesophyll has been proposed as a signal that coordinates leaf size via cell-to-cell communication (class I), compensation is likely mediated by large-scale metabolic reprogramming in class II. Therefore, from the perspective of cell number and size dynamism, a key task is to identify which metabolic changes contribute to class II-mediated compensation.
For instance, class II is observed in the fugu5 Arabidopsis mutant, which has lost H+-PPase activity, the ability to hydrolyze pyrophosphate (PPi), and concomitant vacuolar acidification (Ferjani et al., 2007; Ferjani et al., 2011; Bertoni, 2011; Kriegel et al., 2015; Segami et al., 2018). This mutation leads to excessive accumulation of PPi in the cytosol, partial inhibition of gluconeogenesis, and a reduction in the content of sucrose (Suc) produced from triacylglycerol (TAG), the major seed storage lipid in Arabidopsis (Ferjani et al., 2011). Consequently, Suc deficiency significantly reduced the cell number in fugu5 cotyledons and triggered CCE (Ferjani et al., 2007; Ferjani et al., 2008; Ferjani et al., 2011; Ferjani et al., 2014a; Ferjani et al., 2018). Also, excess PPi triggered major developmental (reduced pavement cell shape complexity) and patterning (stomatal distribution and functioning) defects (Asaoka et al., 2019; Gunji et al., 2020). The above indicates that the impact of PPi is broad but specific. TAG-to-Suc conversion is a major metabolic process that fuels seedling establishment (Bewley and Black, 1994; Graham, 2008). Whereas most studies described a role of Suc in hypocotyl elongation in the dark, its impact on aboveground organ development (in light) was described only recently (Henninger et al., 2022).
PPi may trigger several specific cellular responses (Chen et al., 1990; Lundin et al., 1991; Stitt, 1998; Maeshima, 2000; Heinonen, 2001; Ko et al., 2007; Wang et al., 2016). For example, although excess PPi-related phenotypes in palisade tissue were suppressed by an external supply of carbon (such as Suc) or the removal of PPi in the fugu5 background, Suc supply had no effect on epidermal cell developmental defects (Asaoka et al., 2019; Gunji et al., 2020). Together, the above reports are in line with the assumption that PPi indeed exerts different effects on different plant tissues and cell types and at different developmental stages.
Because PPi is a strong inhibitor of anabolic reactions, several PPases are dedicated to its hydrolysis, preventing accumulation of toxic levels (Segami et al., 2018). H+-PPase (FUGU5) is a master regulator of cytosolic PPi homeostasis (Ferjani et al., 2011; Kriegel et al., 2015; Asaoka et al., 2016; Fukuda et al., 2016; Ferjani et al., 2018; Asaoka et al., 2019). fugu5 provided insight into the effect of PPi metabolism on leaf development and their potential crosstalk. In other words, phenotypic dissection of fugu5 suggested a pivotal role for balanced metabolism during the heterotrophic growth stage, indicating that leaf size is controlled by metabolic networks, with a relatively long-lasting effect. The mechanism is discussed in Section 4.
Genetic studies of the TAG-to-Suc pathway identified several key enzymes whose loss-of-function affected seedling establishment with varying levels of penetrance (Hayashi et al., 1998; Hooks et al., 1999; Froman et al., 2000; Eastmond et al., 2000; Germain et al., 2001; Footitt et al., 2002; Rylott et al., 2003; Fulda et al., 2004). To evaluate the link between TAG-to-Suc conversion and CCE, we phenotypically characterized icl-2, mls-2, pck1-2 (Takahashi et al., 2017), and ibr10 mutants (Tabeta et al., 2021; Tabeta et al., 2022), all of which displayed class II CCE (Katano et al., 2016; Takahashi et al., 2017; Tabeta et al., 2021). Hence, producing Suc from TAG during seed germination is crucial for proper cotyledon development. Yet, based on the high mobility of Suc between leaf cells and tissues, as well as its vital role in plant cells, it is technically challenging to pursue the molecular mechanism of class II CCE by simply restricting Suc production and or tracking its dynamic flux.
H+-PPases are implicated in plant growth, development, and PPi homeostasis (Ferjani et al., 2011; Segami et al., 2018). Nonetheless, although the contribution of PPi homeostasis to plant growth and development has been investigated, its effect on different tissues and cell types during the plant lifecycle is unclear (Schilling et al., 2017). Does altered PPi homeostasis act cell-autonomously or non-cell-autonomously to modulate critical cell fates via specific metabolic processes?
The above hypothesis has been formally tested using a spatiotemporal approach by constructing and analyzing transgenic lines in which PPi has been removed from the epidermis, from palisade tissue cells, or during the 4 days following seed imbibition (Gunji et al., 2022). When the yeast PPase IPP1 was expressed in the epidermis or palisade tissue alone, fugu5 phenotypes were independently restored (Gunji et al., 2022) (Figure 1B). Furthermore, the immediate removal of excess PPi after seed imbibition suppressed CCE of palisade cells but failed to totally rescue the epidermal development defects (Gunji et al., 2022). Next, the impacts of spatial and temporal removal of PPi were investigated by capillary electrophoresis time-of-flight mass spectrometry. This analysis revealed that metabolic profiles are differentially affected among transgenic lines, consistent with an axial role in the central metabolism of gluconeogenesis in CCE (Gunji et al., 2022). These findings not only provide a conceptual framework to unveil metabolic fluctuations within leaf tissues with high spatiotemporal resolution, but also suggest that excess PPi exerts its inhibitory effect in planta during the early stages of seedling establishment in a tissue- and cell-autonomous manner. In other words, leaf size is a highly complex trait governed by multiple regulatory layers, in which metabolic regulation represents another fundamental side.
3.3 KRP2 overexpression-mediated class III compensation: contribution of cell cycle regulation
INHIBITOR/INTERACTOR OF CYCLIN-DEPENDENT KINASES/KIP-RELATED PROTEINS (KRPs) encode cyclin-dependent kinase inhibitors, which block cell cycle progression (Wang et al., 1998; Lui et al., 2000; De Veylder et al., 2001). Constitutive overexpression of individual proteins in this family prematurely arrests the mitotic cell cycle and triggers CCE (De Veylder et al., 2001; Verkest et al., 2005; Ferjani et al., 2007; Jun et al., 2013). The activity that stimulates CCE acts in a cell-autonomous manner, because KRP2 overexpressor cells did not stimulate CCE in the adjacent wild-type cells in leaf chimeras for KRP2 overexpression (Kawade et al., 2010). Notably, epidermis-specific expression of KRP1 induced a similar phenomenon (i.e., CCE was detected in pavement cells without affecting the subepidermal cells of palisade tissue) (Bemis and Torii, 2007) (Figure 1C).
The endogenous KRP2 protein is more abundant in post-mitotic cells than in proliferating cells (Ormenese, 2004; Verkest et al., 2005). Strong KRP2-overexpressing lines exhibited a more obvious CCE phenotype compared with their weak-expressing counterparts (Verkest et al., 2005; Ferjani et al., 2013a; Ferjani et al., 2013b). Therefore, KRP2 may enhance post-mitotic cell expansion; however, this was refuted by clonal analysis using the Cre/lox-P system. In brief, CCE is undetectable when KRP2 overexpression is induced after exiting the mitotic cell cycle (Kawade et al., 2010). As mitotic cell size in KRP2 overexpressers is twice that in the wild type (De Veylder et al., 2001; Ferjani et al., 2007; Ferjani et al., 2013a; Ferjani et al., 2013b), the mechanism by which cells sense their default size to set the timing of mitotic entry is likely perturbed. Given that cell-size homeostasis is generally explained by a sizer, timer, or adder model (Roeder et al., 2012; Wood and Nurse, 2015; Pavelescu et al., 2018; D’Ario et al., 2021), it would be of interest to investigate the involvement of the components of these models in CCE. To corroborate this, we need to quantify leaf cellular dynamics, taking into account the contribution of nutritional resource allocation from seeds, which affects seedling growth in particular (Sizani et al., 2019). Future work aims to clarify how mitotic cells integrate information on cell-size homeostasis, and hence cell proliferation, into cell-autonomous stimulation of CCE.
Clonal analyses enabled the dissection of cellular dynamics, i.e., cell-autonomous or non-cell-autonomous, in class I, II, and III compensation. The results suggest that cell-to-cell communication coordinates cell proliferation and post-mitotic cell expansion in class I but is not a prerequisite in classes II and III. How then does deficient cell proliferation trigger post-mitotic cell expansion within a cell or cell population? The molecular mechanism underlying fugu5-mediated compensation is in good agreement with the cell-autonomous nature of cellular metabolic disorders.
4 Hormonal regulation of class II response phase: contribution of the phytohormone auxin
To elucidate the molecular mechanisms of compensation, it is important to understand the molecular framework of the cell-autonomous induction and response phases for which the fugu5 mutant (class II compensation) has been used as a prototype. Although the reduced cell number in the induction phase in fugu5 was attributed to reduced Suc synthesis, CCE in fugu5 is also controlled metabolically and hormonally (Figure 2). In this section, we describe how forward and reverse genetics-based approaches have provided insight into class II CCE (Katano et al., 2016; Tabeta et al., 2021; Nakayama et al., 2022).
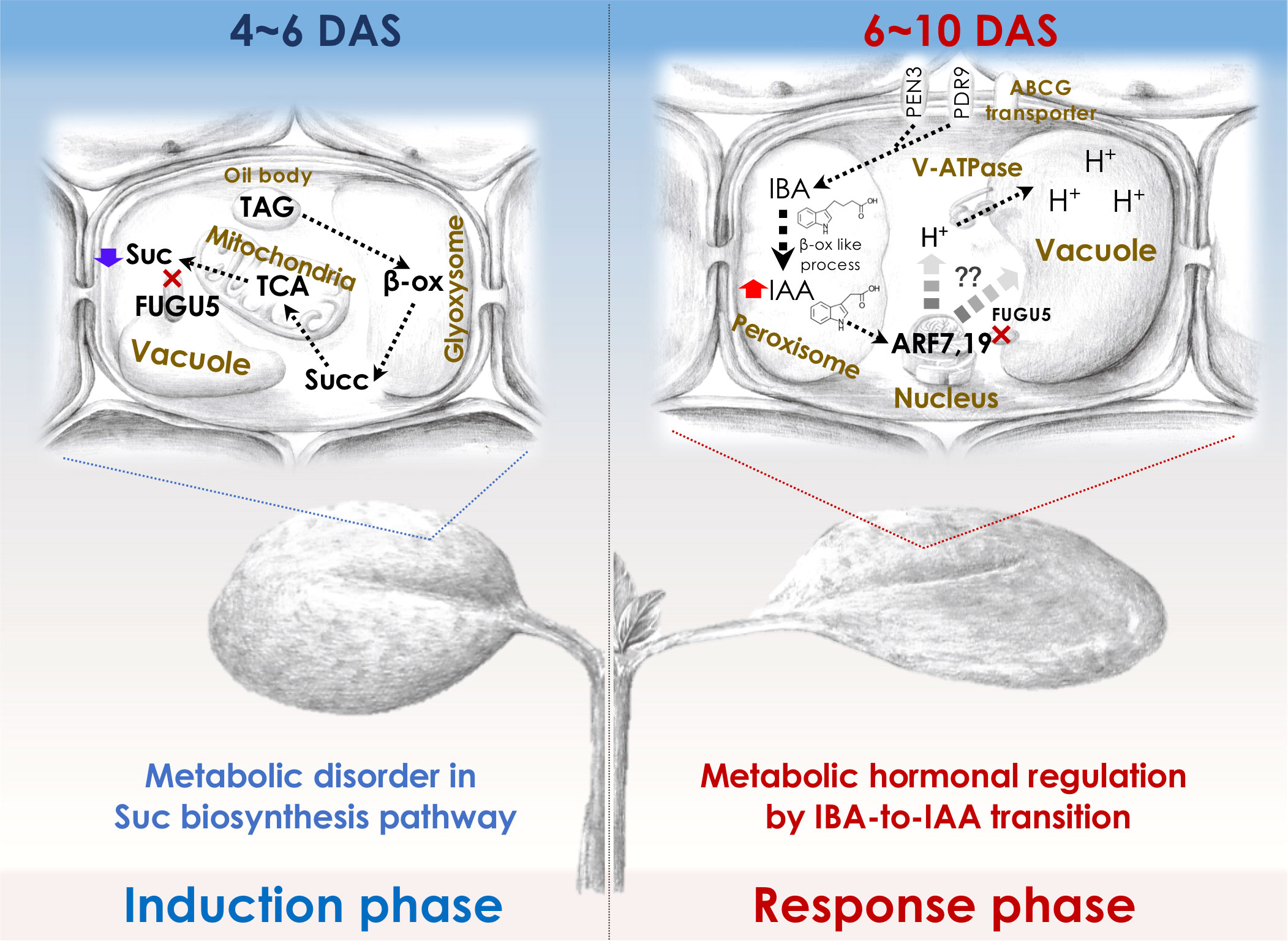
Figure 2 Molecular machinery of the induction and response phases in class II CCE. The decreased cell number (induction phase) in Arabidopsis fugu5-mutant cotyledons has been shown to be exclusively due to a decreased level of TAG-derived Suc, and IBA-derived IAA (Katano et al., 2016; Takahashi et al., 2017) has been suggested to mediate CCE (response phase) as follows: First, upon seed imbibition, excess cytosolic PPi in fugu5 leads to the inhibition of de novo Suc synthesis from TAG, a major seed-storage lipid and substrate for the conversion of fatty acids to acetyl-CoA for glyoxylate bypass that takes place in the glyoxysome (Ferjani et al., 2011). This is owing to inhibition of gluconeogenesis (Ferjani et al., 2018). Second, during seedling establishment, metabolic disorder associated with the reduced Suc content (4-6 DAS; left panel) is converted into an ‘output instructive signal’ (6-10 DAS; right panel) that promotes the conversion of IBA, an auxin precursor, to IAA, the natural phytohormone auxin, leading to an increase in endogenous IAA concentration, which is crucial for CCE (Tabeta et al., 2021). Third, increased endogenous IAA (IAA concentration peaks at 8-10 DAS) triggers the TIR/AFB-dependent auxin signaling pathway through the AUXIN RESPONSE FACTORs ARF7 and ARF19, transcriptional activators of early auxin-response genes. This subsequently activates the vacuolar type V-ATPase, leading to an increase in turgor pressure, which ultimately drives an increase in cell size and CCE (Tabeta et al., 2021; Nakayama et al., 2022). β-ox, β-oxidation; Succ, succinate. Glyoxysome, the single membrane-bound organelles that house most of the biochemical machinery required to convert fatty acids derived from TAG to 4-carbon compounds. TCA, the tricarboxylic acid cycle. PEN3, PENETRATION (PEN) 3 is a membrane-localized ATP-binding cassette (ABC) transporter. PDR9, is a member of the pleiotropic drug resistance (PDR) family of ATP-binding cassette transporters. ABCG transporter, G-type ATP-binding cassette (ABCG) transporter. DAS, days after seed sowing.
By mutagenizing fugu5 dry seeds using 12C6+ heavy-ion irradiation, Katano and colleagues performed a large-scale screening using forward genetics to identify key genes in CCE. This phenotypic screening identified ENOYL-CoA HYDRATASE 2 (ECH2) activity as a prerequisite for CCE to occur in the fugu5 background. Because ECH2 has dual functions (β-oxidation of TAG-to-Suc conversion (Graham, 2008; Li et al., 2019) and conversion of indole-3-butyric acid (IBA) to the auxin 3-indole acetic acid (IAA) (Strader et al., 2011)), these metabolic processes have been postulated to play a major role in CCE. IAA levels are tightly regulated by de novo biosynthesis, transportation, and inactive conversion, and IBA has been implicated in cotyledon and root development (Strader et al., 2010; Frick and Strader, 2018). Hence, the suppression of CCE in fugu5 ech2 was a result of defective IAA biosynthesis from IBA caused by loss of ECH2 activity.
Reverse genetics in the fugu5 background showed that CCE in the ibr1 ibr3 ibr10 fugu5 quadruple mutant, which is defective in IBA-to-IAA conversion (Strader et al., 2010; Strader et al., 2011), was totally suppressed (Tabeta et al., 2021). In contrast, pen3 fugu5 and pdr9 fugu5, in which IBA efflux is compromised, exhibited a high-IAA phenotype (Strader and Bartel, 2009; Ruzicka et al., 2010; Aryal et al., 2019) and enhanced CCE (Tabeta et al., 2021). Consistently, endogenous IAA levels were twofold higher in fugu5 (in 8-10-day-old seedlings). These findings indicate that IAA converted from IBA is essential for CCE, in agreement with the finding that the degree of CCE reflects the intracellular IAA level (Tabeta et al., 2021). How is high auxin sensed, interpreted, and transduced into CCE in fugu5?
AUXIN RESPONSE FACTORS (ARF) 7 and 19 and V-ATPase activity are essential for CCE in fugu5. More specifically, arf7-1 arf19-1 fugu5-1 and vha-a2 vha-a3 fugu5-1 triple mutants did not exhibit CCE, despite having a significantly reduced number of cells (Tabeta et al., 2021). The vacuole accounts for the majority of the cell volume and is essential for cell enlargement, in class II CCE, proton translocation via V-ATPase contributes to vacuole enlargement and promotes cell-size control via the ARF7 ARF19 module. The above findings are valid for all other mutants with class II CCE, namely isocitrate lyase (icl; Eastmond et al., 2000), malate synthase (mls; Cornah et al., 2004), phosphoenolpyruvate carboxykinase1 (pck1; Penfield et al., 2004), and ibr10 (Zolman et al., 2008), but not an3 and fas1 (class I compensation) (Katano et al., 2016; Takahashi et al., 2017; Tabeta et al., 2021). Therefore, IBA-related hormonal signaling is likely activated in response to metabolic disorders of the central metabolism. If so, one major question arises: Could this rather indicate that auxin integrates more signals (metabolism, cell cycle, vacuole volume, cell wall remodeling etc.) and thus is at the same hierarchical level as compensation (i.e., integrating heterogeneous pathways)?
Finally, although auxin signaling plays a major role in class II, the phytohormone SA has been proposed to relate to class I CCE in the an3-4 mutant (Fujikura et al., 2020). These findings indicate that the response phase in classes I and II is under phytohormonal control. Importantly, hormonal cross-talk controlling leaf development has also been discussed. As mentioned above, gibberellins together with DELLA proteins, which are downstream of the IAA response, modulate cell proliferation and expansion (Sun and Gubler, 2004; de Lucas et al., 2008; Achard et al., 2009). Also, some TCPs have been reported to regulate auxin homeostasis and cytokinin by altering the expression of auxin biosynthetic enzymes (Lucero et al., 2015). Since IBA-derived IAA provides NO signaling and promotes other hormonal responses (Fattorini et al., 2017), hormonal cross-talk regulation might also have a role in leaf size regulation in compensation-exhibiting mutant background. To this end, compensation-mediated leaf size control may represent a good starting point for further studies aiming to unveil the broader picture of phytohormonal regulation.
5 Environmentally-induced compensation
Growth and development of plants are greatly affected by environmental changes. Because plants cannot move, altering their own tissue structure and concomitant organ size and/or shape to adapt to the ambient environment is critical for their survival. For instance, heterophylly, which is defined as a leaf-form alteration triggered by the surrounding environment, is commonly observed in aquatic and amphibious plants (Li et al., 2019). Rorippa aquatica is an amphibious plant found in riparian environments, such as the bank of a natural watercourse including lakes, ponds, and streams, in North America. R. aquatica shows a remarkable heterophylly, and develops deeply dissected narrow leaves under submerged conditions whereas it develops simple shaped leaves on terrestrial conditions. The leaf shape changes also in response to varying ambient temperature, in which lower temperature induces dissected narrow leaves. Recently, it was shown that both submergence (Sakamoto et al., 2022) and low-temperature (Amano et al., 2015) treatments caused an increase in cell size in the sub-epidermal palisade tissue layer in mature leaves, along with a decrease in leaf blade area. This phenotype in which cell size increase occurs in the background of tissue-size reduction resembles compensation. Indeed, the expression of some of the compensation-related genes is altered under low-temperature or submergence in R. aquatica.
Because phytohormones usually reflect the environmental status, in some ways they represent the secondary messengers of environmental cues. Therefore, it is not surprising to see compensation being dependent on the environment. In other words, in the case of R. aquatica, environmental cues may have triggered compensation via a yet unidentified hormonal regulatory pathway. These observations also indicate that compensation could be induced not only in mutants, transgenics, and γ-ray–treated plants, but also as an adaptive response to a wide range of environmental stimuli. Together, these findings provide evidence that compensation is a universal phenomenon that is also seen in nature, whereby hormones are acting as downstream instructive signal of the environmental status.
6 Outstanding questions and future prospects
Large-scale genetic screening identified a number of genes whose loss- or gain-of-function alters final leaf size. For decades, this prompted work on the morphogenesis of Arabidopsis. Subsequent functional analyses of the above genetic factors revealed the molecular events governing leaf development. Furthermore, bioinformatics techniques identified several key transcription factors (TFs) relevant to core genetic modules (Ichihashi et al., 2014; Sinha et al., 2016). However, our understanding of leaf-size control is incomplete.
Most developmental events are interpreted based on TFs (Kaufmann and Airoldi, 2018; and references therein). Although TFs are crucial in orchestrating organogenesis, other regulatory factors with more indirect effects have been overlooked. This is a result of the inherent bias of most genetic screening approaches, which identify major non-redundant players, overlooking more indirect and diffuse properties that build on more complex interactions (Nakayama et al., 2022). This includes understudied players (e.g., metabolic molecules) and understudied systemic interactions (e.g., feedback loops). Thus, the compensatory mechanism of leaf-size control cannot be explained by transcriptional regulation processes only (Ferjani et al., 2008; Ferjani et al., 2014a; Ferjani et al., 2018). Multi-omics enables identification of novel small molecules (including metabolic components and regulators) critical in major developmental transitions during plant growth (Omidbakhshfard et al., 2021) and leaf development (Tabeta et al., 2021). Systems biology may also help to identify, in the topology of the molecular network, key events and interactions, to explain how compensation emerges.
Beyond molecule identity and interactions, the mechanical properties of plant cells and tissues should be regarded as an integral part of developmental signaling pathways. Advances in quantitative plant biology have allowed experimentation and modeling using plants (Autran et al., 2021). Hence, the regulatory mechanisms at the crossroads of metabolism, morphogenesis, and mechanics should be integrated with genetic mechanisms to understand the multimodal drivers of leaf-size control (Trinh et al., 2021; Nakayama et al., 2022).
Finally, if compensation is a general and primary size-regulatory mechanism in plant leaves, does it involve, by default, a proprioceptive sensing step? Does CCE generate an additional instructive mechanical signal? If so, how is such a signal perceived, resolved, integrated, and executed to guarantee proper size? To address these questions, leaf development should be reexamined from the perspectives of biomechanics and metabolism.
Author contributions
HT, SG, KK and AF drafted and wrote the paper. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by Grant-in-Aid for Scientific Research (B) (JP16H04803 to AF); Grant-in-Aid for Scientific Research on Innovative Areas (JP25113002 and JP18H05487 to AF); The Naito Foundation. HT is a recipient of a Research Fellowship for Young Scientists (20J20901).
Acknowledgments
We thank Prof. Olivier Hamant (ENS de Lyon) and Prof. Seisuke Kimura (Kyoto Sangyo University) for their critical reading of the manuscript. AF along with co-authors wrote this review in celebration of the 20th anniversary of compensation, and dedicates this article to Prof. Hirokazu Tsukaya (The University of Tokyo) and Prof. Gorou Horiguchi (Rikkyo University), the ones who helped to make amazing discoveries, and brought attention to crucial issues in leaf development.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Abbreviations
an3, angustifolia3; ARF, AUXIN RESPONSE FACTOR; CCE, Compensated cell enlargement; CCX4, CATION CALCIUM EXCHANGER 4; H+-PPase, H+-translocating inorganic pyrophosphatase; IAA, Indole-3-acetic acid; IBA, Indole-3-butyric acid; IBR, INDOLE-3-BUTYRIC ACID RESPONSE; KRP, INHIBITOR/INTERACTOR OF CYCLIN-DEPENDENT KINASE /KIP-RELATED PROTEIN; ECH2, ENOYL-COA HYDRATASE 2; PPi, Pyrophosphate; SAM, Shoot apical meristem; TAG, Triacylglycerol; ROS, Reactive oxygen species; SA, Salicylic acid; Suc, Sucrose; V-ATPases, Vacuolar-type H+-ATPases; xs, extra-small sisters.
References
Achard, P., Gusti, A., Cheminant, S., Alioua, M., Dhondt, S., Coppens, F., et al. (2009). Gibberellin signaling controls cell proliferation rate in Arabidopsis. Curr. Biol. 19, 1188–1193. doi: 10.1016/j.cub.2009.05.059
Ali, S., Khan, N., Xie, L. (2020). Molecular and hormonal regulation of leaf morphogenesis in Arabidopsis. Int. J. Mol. Sci. 21, 5132. doi: 10.3390/ijms21145132
Amano, R., Nakayamam, H., Morohoshim, Y., Kawakatsu, Y., Ferjani, A., Kimura, S. (2015). A decrease in ambient temperature induces post-mitotic enlargement of palisade cells in north american lake cress. PloS One 10, e0141247. doi: 10.1371/journal.pone.0141247
Aryal, B., Huynh, J., Schneuwly, J., Siffert, A., Liu, J., Alejandro, S., et al. (2019). ABCG36/PEN3/PDR8 is an exporter of the auxin precursor, indole-3-Butyric acid, and involved in auxin-controlled development. Front. Plant Sci. 10, 899. doi: 10.3389/fpls.2019.00899
Asaoka, M., Inoue, S. I., Gunji, S., Kinoshita, T., Maeshima, M., Tsukaya, H., et al. (2019). Excess pyrophosphate within guard cells delays stomatal closure. Plant Cell Physiol. 60, 875–887. doi: 10.1093/pcp/pcz002
Asaoka, M., Segami, S., Ferjani, A., Maeshima, M. (2016). Contribution of PPi-hydrolyzing function of vacuolar H+-pyrophosphatase in vegetative growth of Arabidopsis: evidenced by expression of uncoupling mutated enzymes. Front. Plant Sci. 7, 415. doi: 10.3389/fpls.2016.00415
Asnacios, A., Hamant, O. (2012). The mechanics behind cell polarity. Trends Cell Biol. 11, 584–591. doi: 10.1016/j.tcb.2012.08.005
Autran, D., Bassel, G. W., Chae, E., Ezer, D., Ferjani, A., Fleck, C., et al. (2021). What is quantitative plant biology? Quantitative Plant Biol. 2, e10. doi: 10.1017/qpb.2021.8
Autran, D., Jonak, C., Belcram, K., Beemster, G. T., Kronenberger, J., Grandjean, O., et al. (2002). Cell numbers and leaf development in Arabidopsis: A functional analysis of the STRUWWELPETER gene. EMBO J. 21, 6036–6049. doi: 10.1093/emboj/cdf614
Barrôco, R. M., Peres, A., Droual, A. M., De Veylder, L., Nguyen le, S. L., De Wolf, J., et al. (2006). The cyclin-dependent kinase inhibitor Orysa;KRP1 plays an important role in seed development of rice. Plant Physiol. 142, 1053–1064. doi: 10.1104/pp.106.087056
Beemster, G. T., Fiorani, F., Inzé, D. (2003). Cell cycle: the key to plant growth control? Trends Plant Sci. 8, 154–158. doi: 10.1016/S1360-1385(03)00046-3
Beemster, G. T., Vercruysse, S., De Veylder, L., Kuiper, M., Inzé, D. (2006). The Arabidopsis leaf as a model system for investigating the role of cell cycle regulation in organ growth. J. Plant Res. 119, 43–50. doi: 10.1007/s10265-005-0234-2
Bemis, S. M., Torii, K. U. (2007). Autonomy of cell proliferation and developmental programs during Arabidopsis aboveground organ morphogenesis. Dev. Biol. 304, 367–381. doi: 10.1016/j.ydbio.2006.12.049
Bertoni, G. (2011). A surprising role for vacuolar pyrophosphatase. Plant Cell 23, 2808. doi: 10.1105/tpc.111.230813
Bewley, J. D., Black, M. (1994). Seeds: in Physiology of development and germination, Ed. 2 (New York: Plenum Press).
Bowman, J. L., Eshed, Y., Baum, S. F. (2002). Establishment of polarity in angiosperm lateral organs. Trends Genet. 18, 134–141. doi: 10.1016/S0168-9525(01)02601-4
Burch-Smith, T. M., Zambryski, P. C. (2010). Loss of INCREASED SIZE EXCLUSION LIMIT (ISE)1 or ISE2 increases the formation of secondary plasmodesmata. Curr. Biol. 20, 989–993. doi: 10.1016/j.cub.2010.03.064
Challa, K. R., Rath, M., Sharma, A. N., Bajpai, A. K., Davuluri, S., Acharya, K. K., et al. (2021). Active suppression of leaflet emergence as a mechanism of simple leaf development. Nat. Plants 7, 1264–1275. doi: 10.1038/s41477-021-00965-3
Chen, J., Brevet, A., Fromant, M., Lévêque, F., Schmitter, J. M., Blanquet, S., et al. (1990). Pyrophosphatase is essential for growth of Escherichia coli. J. Bacteriol. 172, 5686–5689. doi: 10.1128/jb.172.10.5686-5689.1990
Clay, N. K., Nelson, T. (2005). The recessive epigenetic swellmap mutation affects the expression of two step II splicing factors required for the transcription of the cell proliferation gene STRUWWELPETER and for the timing of cell cycle arrest in the Arabidopsis leaf. Plant Cell 17, 1994–2008. doi: 10.1105/tpc.105.032771
Cornah, J. E., Germain, V., Ward, J. L., Beale, M. H., Smith, S. M. (2004). Lipid utilization, gluconeogenesis, and seedling growth in Arabidopsis mutants lacking the glyoxylate cycle enzyme malate synthase. J. Biol. Chem. 279, 42916–42923. doi: 10.1074/jbc.M407380200
Crawford, K. M., Zambryski, P. C. (2001). Non-targeted and targeted protein movement through plasmodesmata in leaves in different developmental and physiological states. Plant Physiol. 125, 1802–1812. doi: 10.1104/pp.125.4.1802
D’Ario, M., Tavares, R., Schiessl, K., Desvoyes, B., Gutierrez, C., Howard, M., et al. (2021). Cell size controlled in plants using DNA content as an internal scale. Science 372, 1176–1181. doi: 10.1126/science.abb4348
Das Gupta, M., Nath, U. (2015). Divergence in patterns of leaf growth polarity is associated with the expression divergence of miR396. Plant Cell 27, 2785–2799. doi: 10.1105/tpc.15.00196
Delgado-Benarroch, L., Weiss, J., Egea-Cortines, M. (2009). The mutants compacta ähnlich, nitida and grandiflora define developmental compartments and a compensation mechanism in floral development in Antirrhinum majus. J. Plant Res. 122, 559–569. doi: 10.1007/s10265-009-0236-6
Dewitte, W., Scofield, S., Alcasabas, A. A., Maughan, S. C., Menges, M., Braun, N., et al. (2007). Arabidopsis CYCD3 d-type cyclins link cell proliferation and endocycles and are rate-limiting for cytokinin responses. Proc. Natl. Acad. Sci. U. S. A. 104, 14537–14542. doi: 10.1073/pnas.0704166104
de Lucas, M., Davière, J. M., Rodríguez-Falcón, M., Pontin, M., Iglesias-Pedraz, J. M., Lorrain, S., et al. (2008). A molecular framework for light and gibberellin control of cell elongation. Nature 451, 480–484. doi: 10.1038/nature06520
De Veylder, L., Beeckman, T., Beemster, G. T., Krols, L., Terras, F., Landrieu, I., et al. (2001). Functional analysis of cyclin-dependent kinase inhibitors of Arabidopsis. Plant Cell 13, 1653–1668. doi: 10.1105/TPC.010087
Dkhar, J., Pareek, A. (2014). What determines a leaf’s shape? EvoDevo 5, 47. doi: 10.1186/2041-9139-5-47
Dow, G. J., Bergmann, D. C. (2014). Patterning and processes: how stomatal development defines physiological potential. Curr. Opin. Plant Biol. 21, 67–74. doi: 10.1016/j.pbi.2014.06.007
Du, F., Guan, C., Jiao, Y. (2018). Molecular mechanisms of leaf morphogenesis. Mol. Plant 11, 1117–1134. doi: 10.1016/j.molp.2018.06.006
Eastmond, P. J., Germain, V., Lange, P. R., Bryce, J. H., Smith, S. M., Graham, I. A. (2000). Postgerminative growth and lipid catabolism in oilseeds lacking the glyoxylate cycle. Proc. Natl. Acad. Sci. U.S.A. 97, 5669–5674. doi: 10.1073/pnas.97.10.5669
Efroni, I., Han, S. K., Kim, H. J., Wu, M. F., Steiner, E., Birnbaum, K. D., et al. (2013). Regulation of leaf maturation by chromatin-mediated modulation of cytokinin responses. Dev. Cell 24, 438–445. doi: 10.1016/j.devcel.2013.01.019
Eriksson, S., Stransfeld, L., Adamski, N. M., Breuninger, H., Lenhard, M. (2010). KLUH/CYP78A5-dependent growth signaling coordinates floral organ growth in Arabidopsis. Curr. Biol. 20, 527–532. doi: 10.1016/j.cub.2010.01.039
Exner, V., Taranto, P., Schönrock, N., Gruissem, W., Hennig, L. (2006). Chromatin assembly factor CAF-1 is required for cellular differentiation during plant development. Development 133, 4163–4172. doi: 10.1242/dev.02599
Fattorini, L., Veloccia, A., Della Rovere, F., D’Angeli, S., Falasca, G., Altamura, M. M. (2017). Indole-3-butyric acid promotes adventitious rooting in Arabidopsis thaliana thin cell layers by conversion into indole-3-acetic acid and stimulation of anthranilate synthase activity. BMC Plant Biol. 17, 121. doi: 10.1186/s12870-017-1071-x
Ferjani, A., Horiguchi, G., Tsukaya, H. (2010). Organ size control in Arabidopsis: Insights from compensation studies. Plant Morphol. 22, 65–71. doi: 10.5685/plmorphol.22.65
Ferjani, A., Horiguchi, G., Yano, S., Tsukaya, H. (2007). Analysis of leaf development in fugu mutants of Arabidopsis reveals three compensation modes that modulate cell expansion in determinate organs. Plant Physiol. 144, 988–999. doi: 10.1104/pp.107.099325
Ferjani, A., Ishikawa, K., Asaoka, M., Ishida, M., Horiguchi, G., Maeshima, M., et al. (2013a). Enhanced cell expansion in a KRP2 overexpressor is mediated by increased V-ATPase activity. Plant Cell Physiol. 54, 1989–1998. doi: 10.1093/pcp/pct138
Ferjani, A., Ishikawa, K., Asaoka, M., Ishida, M., Horiguchi, G., Maeshima, M., et al. (2013b). Class III compensation, represented by KRP2 overexpression, depends on V-ATPase activity in proliferative cells. Plant Signal. Behav. 8, e27204. doi: 10.4161/psb.27204
Ferjani, A., Kawade, K., Asaoka, M., Oikawa, A., Okada, T., Mochizuki, A., et al. (2018). Pyrophosphate inhibits gluconeogenesis by restricting UDP-glucose formation in vivo. Sci. Rep. 8, 14696. doi: 10.1038/s41598-018-32894-1
Ferjani, A., Segami, S., Asaoka, M., Maeshima, M. (2014a). Regulation of PPi levels through the vacuolar membrane H+-pyrophosphatase. in Progress in botany, vol. 75 . Eds. Lüttge, U., Beyschlag, W., Cushman, J. (Berlin: Springer-Verlag), 145–165. doi: 10.1007/978-3-642-38797-5_5
Ferjani, A., Segami, S., Horiguchi, G., Muto, Y., Maeshima, M., Tsukaya, H. (2011). Keep an eye on PPi: the vacuolar-type H+-pyrophosphatase regulates postgerminative development in Arabidopsis. Plant Cell 23, 2895–2908. doi: 10.1105/tpc.111.085415
Ferjani, A., Segami, S., Horiguchi, G., Muto, Y., Maeshima, M., Tsukaya, H. (2014b). Roles of the vacuolar H+-PPase in seed storage oil mobilization and plant development. Plant Morphol. 26, 45–51. doi: 10.5685/plmorphol.26.45
Ferjani, A., Yano, S., Horiguchi, G., Tsukaya, H. (2008). Control of leaf morphogenesis by long- and short-distance signaling: Differentiation of leaves into sun or shade types and compensated cell enlargement. in Plant cell monographs: Plant growth signaling. Eds. Bögre, L., Beemster, G. T. S. (Berlin, Heidelberg, Germany: Springer Berlin Heidelberg), 47–62. doi: 10.1007/7089_2007_148
Fitzgibbon, J., Beck, M., Zhou, J., Faulkner, C., Robatzek, S., Oparka, K. (2013). A developmental framework for complex plasmodesmata formation revealed by large-scale imaging of the Arabidopsis leaf epidermis. Plant Cell 25, 57–70. doi: 10.1105/tpc.112.105890
Footitt, S., Slocombe, S. P., Larner, V., Kurup, S., Wu, Y., Larson, T., et al. (2002). Control of germination and lipid mobilization by COMATOSE, the Arabidopsis homologue of human ALDP. EMBO J. 21, 2912–2922. doi: 10.1093/emboj/cdf300
Frick, E. M., Strader, L. C. (2018). Roles for IBA-derived auxin in plant development. J. Exp. Bot. 69, 169–177. doi: 10.1093/jxb/erx298
Froman, B. E., Edwards, P. C., Bursch, A. G., Dehesh, K. (2000). ACX3, a novel medium-chain acyl-coenzyme a oxidase from Arabidopsis. Plant Physiol. 123, 733–742. doi: 10.1104/pp.123.2.733
Fujikura, U., Ezaki, K., Horiguchi, G., Seo, M., Kanno, Y., Kamiya, Y., et al. (2020). Suppression of class I compensated cell enlargement by xs2 mutation is mediated by salicylic acid signaling. PloS Genet. 16, e1008873. doi: 10.1371/journal.pgen.1008873
Fujikura, U., Horiguchi, G., Ponce, M. R., Micol, J. L., Tsukaya, H. (2009). Coordination of cell proliferation and cell expansion mediated by ribosome-related processes in the leaves of Arabidopsis thaliana. Plant J. 59, 499–508. doi: 10.1111/j.1365-313X.2009.03886.x
Fujikura, U, Horiguchi, G, Tsukaya, H (2007a). Dissection of enhanced cell expansion processes in leaves triggered by a defect in cell proliferation, with reference to roles of endoreduplication. Plant Cell Physiol. 48, 278–286. doi: 10.1093/pcp/pcm002
Fujikura, U, Horiguchi, G, Tsukaya, H (2007b). Genetic relationship between angustifolia3 and extra-small sisters highlights novel mechanisms controlling leaf size. Plant Signal. Behav 2, 378–380. doi: 10.4161/psb.2.5.4525
Fukuda, M., Segami, S., Tomoyama, T., Asaoka, M., Nakanishi, Y., Gunji, S., et al. (2016). Lack of H+-pyrophosphatase prompts developmental damage in Arabidopsis leaves on ammonia-free culture medium. Front. Plant Sci. 7, 819. doi: 10.3389/fpls.2016.00819
Fulda, M., Schnurr, J., Abbadi, A., Heinz, E., Browse, J. (2004). Peroxisomal acyl-CoA synthetase activity is essential for seedling development in Arabidopsis thaliana. Plant Cell 16, 394–405. doi: 10.1105/tpc.019646
Germain, V., Rylott, E. L., Larson, T. R., Sherson, S. M., Bechtold, N., Carde, J. P., et al. (2001). Requirement for 3-ketoacyl-CoA thiolase-2 in peroxisome development, fatty acid beta-oxidation and breakdown of triacylglycerol in lipid bodies of Arabidopsis seedlings. Plant J. 28, 1–12. doi: 10.1046/j.1365-313X.2001.01095.x
González, N., Inzé, D. (2015). Molecular systems governing leaf growth: from genes to networks. J. Exp. Bot. 66, 1045–1054. doi: 10.1093/jxb/eru541
González, N., Vanhaeren, H., Inzé, D. (2012). Leaf size control: complex coordination of cell division and expansion. Trends Plant Sci. 6, 332–340. doi: 10.1016/j.tplants.2012.02.003
Gorelova, V., Sprakel, J., Weijers, D. (2021). Plant cell polarity as the nexus of tissue mechanics and morphogenesis. Nat. Plants 12, 1548–1559. doi: 10.1038/s41477-021-01021-w
Graham, I. A. (2008). Seed storage oil mobilization. Annu. Rev. Plant Biol. 59, 115–142. doi: 10.1146/annurev.arplant.59.032607.092938
Gunji, S., Kawade, K., Tabeta, H., Horiguchi, G., Oikawa, A., Asaoka, M., et al. (2022). Tissue-targeted inorganic pyrophosphate hydrolysis in a fugu5 mutant reveals that excess inorganic pyrophosphate triggers developmental defects in a cell-autonomous manner. Front. Plant Sci. 13, 945225. doi: 10.3389/fpls.2022.945225
Gunji, S., Oda, Y., Takigawa-Imamura, H., Tsukaya, H., Ferjani, A. (2020). Excess pyrophosphate restrains pavement cell morphogenesis and alters organ flatness in Arabidopsis thaliana. Front. Plant Sci. 11, 31. doi: 10.3389/fpls.2020.00031
Hase, Y., Trung, K. H., Matsunaga, T., Tanaka, A. (2006). A mutation in the uvi4 gene promotes progression of endo-reduplication and confers increased tolerance towards ultraviolet b light. Plant J. 46, 317–326. doi: 10.1111/j.1365-313X.2006.02696.x
Hayashi, M., Toriyama, K., Kondo, M., Nishimura, M. (1998). 2,4-dichlorophenoxybutyric acid-resistant mutants of Arabidopsis have defects in glyoxysomal fatty acid beta-oxidation. Plant Cell 10, 183–195. doi: 10.1105/tpc.10.2.183
Heinonen, J. K. (2001). Biological role of inorganic pyrophosphate (Boston/Dordrecht/London: Kluwer Academic Publishers).
Hemerly, A., Engler, J. A., Bergounioux, C., Van Montagu, M., Engler, G., Inzé, D., et al. (1995). Dominant negative mutants of the Cdc2 kinase uncouple cell division from iterative plant development. EMBO J. 14, 3925–3936. doi: 10.1002/j.1460-2075.1995.tb00064.x
Henninger, M., Pedrotti, L., Krischke, M., Draken, J., Wildenhain, T., Fekete, A., et al. (2022). The evolutionarily conserved kinase SnRK1 orchestrates resource mobilization during Arabidopsis seedling establishment. Plant Cell 34, 616–632. doi: 10.1093/plcell/koab270
Hepworth, J., Lenhard, M. (2014). Regulation of plant lateral-organ growth by modulating cell number and size. Curr. Opin. Plant Biol. 17, 36–42. doi: 10.1016/j.pbi.2013.11.005
Hisanaga, T., Kawade, K., Tsukaya, H. (2015). Compensation: a key to clarifying the organ-level regulation of lateral organ size in plants. J. Exp. Bot. 66, 1055–1063. doi: 10.1093/jxb/erv028
Hisanaga, T., Ferjani, A., Horiguchi, G., Ishikawa, N., Fujikura, U., Kubo, M., et al. (2013). The ATM-dependent DNA damage response acts as an upstream trigger for compensation in the fas1 mutation during Arabidopsis leaf development. Plant Physiol. 162, 831–841. doi: 10.1104/pp.113.216796
Hong, L., Dumond, M., Tsugawa, S., Sapala, A., Routier-Kierzkowska, A. L., Zhou, Y., et al. (2016). Variable cell growth yields reproducible organ development through spatiotemporal averaging. Dev. Cell 38, 15–32. doi: 10.1016/j.devcel.2016.06.016
Hooks, M. A., Kellas, F., Graham, I. A. (1999). Long-chain acyl-CoA oxidases of Arabidopsis. Plant J. 20, 1–13. doi: 10.1046/j.1365-313X.1999.00559.x
Horiguchi, G., Ferjani, A., Fujikura, U., Tsukaya, H. (2006a). Coordination of cell proliferation and cell expansion in the control of leaf size in Arabidopsis thaliana. J. Plant Res. 119, 37–42. doi: 10.1007/s10265-005-0232-4
Horiguchi, G., Fujikura, U., Ferjani, A., Ishikawa, N., Tsukaya, H. (2006b). Large-Scale histological analysis of leaf mutants using two simple leaf observation methods: Identification of novel genetic pathways governing the size and shape of leaves. Plant J. 48, 638–644. doi: 10.1111/j.1365-313X.2006.02896.x
Horiguchi, G., Kim, G. T., Tsukaya, H. (2005). The transcription factor AtGRF5 and the transcription coactivator AN3 regulate cell proliferation in leaf primordia of Arabidopsis thaliana. Plant J. 43, 68–78. doi: 10.1111/j.1365-313X.2005.02429.x
Horiguchi, G., Tsukaya, H. (2011). Organ size regulation in plants: insights from compensation. Front. Plant Sci. 2, 24. doi: 10.3389/fpls.2011.00024
Hussain, E., Romanowski, A., Halliday, K. J. (2022). PIF7 controls leaf cell proliferation through an AN3 substitution repression mechanism. Proc. Natl. Acad. Sci. U.S.A. 119, e2115682119. doi: 10.1073/pnas.2115682119
Ichihashi, Y., Aguilar-Martínez, J. A., Farhi, M., Chitwood, D. H., Kumar, R., Millon, L. V., et al. (2014). Evolutionary developmental transcriptomics reveals a gene network module regulating interspecific diversity in plant leaf shape. Proc. Natl. Acad. Sci. U.S.A. 111, E2616–21. doi: 10.1073/pnas.1402835111
Irish, V. F. (2010). The flowering of Arabidopsis flower development. Plant J. 61, 1014–1028. doi: 10.1111/j.1365-313X.2009.04065.x
Ito, T., Kim, G. T., Shinozaki, K. (2000). Disruption of an Arabidopsis cytoplasmic ribosomal protein S13-homologous gene by transposon-mediated mutagenesis causes aberrant growth and development. Plant J. 22, 257–264. doi: 10.1046/j.1365-313x.2000.00728.x
Johnson, K., Lenhard, M. (2011). Genetic control of plant organ growth. New Phytol. 191, 319–333. doi: 10.1111/j.1469-8137.2011.03737.x
Jun, S. E., Kim, J. H., Hwang, J. Y., Huynh Le, T. T., Kim, G. T. (2019). ORESARA15 acts synergistically with ANGUSTIFOLIA3 and separately from AINTEGUMENTA to promote cell proliferation during leaf growth. Int. J. Mol. Sci. 21, 241. doi: 10.3390/ijms21010241
Jun, S. E., Okushima, Y., Nam, J., Umeda, M., Kim, G. T. (2013). Kip-related protein 3 is required for control of endoreduplication in the shoot apical meristem and leaves of Arabidopsis. Mol. Cells 35, 47–53. doi: 10.1007/s10059-013-2270-4
Kalve, S., De Vos, D., Beemster, G. T. (2014). Leaf development: a cellular perspective. Front. Plant Sci. 5, 362. doi: 10.3389/fpls.2014.00362
Katano, M., Takahashi, K., Hirano, T., Kazama, Y., Abe, T., Tsukaya, H., et al. (2016). Suppressor screen and phenotype analyses revealed an emerging role of the monofunctional peroxisomal enoyl-CoA hydratase 2 in compensated cell enlargement. Front. Plant Sci. 7, 132. doi: 10.3389/fpls.2016.00132
Kaufmann, K., Airoldi, C. A. (2018). Master regulatory transcription factors in plant development: a blooming perspective. Methods Mol. Biol. 1830, 3–22. doi: 10.1007/978-1-4939-8657-6_1
Kawade, K., Horiguchi, G., Tsukaya, H. (2010). Non-cell-autonomously coordinated organ size regulation in leaf development. Development 137, 4221–4227. doi: 10.1242/dev.057117
Kawade, K., Horiguchi, G., Usami, T., Hirai, M. Y., Tsukaya, H. (2013). ANGUSTIFOLIA3 signaling coordinates proliferation between clonally distinct cells in leaves. Curr. Biol. 23, 788–792. doi: 10.1016/j.cub.2013.03.044
Kawade, K., Tanimoto, H., Horiguchi, G., Tsukaya, H. (2017). Spatially different tissue-scale diffusivity shapes ANGUSTIFOLIA3 gradient in growing leaves. Biophys. J. 113, 1109–1120. doi: 10.1016/j.bpj.2017.06.072
Kim, J. H., Kende, H. (2004). A transcriptional coactivator, AtGIF1, is involved in regulating leaf growth and morphology in Arabidopsis. Proc. Natl. Acad. Sci. U. S. A. 101, 13374–13379. doi: 10.1073/pnas.0405450101
Kim, J. H., Tsukaya, H. (2015). Regulation of plant growth and development by the GROWTH-REGULATING FACTOR and GRF-INTERACTING FACTOR duo. J. Exp. Bot. 66, 6093–6107. doi: 10.1093/jxb/erv349
Kim, J. Y., Yuan, Z., Jackson, D. (2003). Developmental regulation and significance of KNOX protein trafficking in Arabidopsis. Development 130, 4351–4362. doi: 10.1242/dev.00618
Kitagawa, M., Wu, P., Balkunde, R., Cunniff, P., Jackson, D. (2022). An RNA exosome subunit mediates cell-to-cell trafficking of a homeobox mRNA via plasmodesmata. Science 375, 177–182. doi: 10.1126/science.abm0840
Ko, K. M., Lee, W., Yu, J. R., Ahnn, J. (2007). PYP-1, inorganic pyrophosphatase, is required for larval development and intestinal function in C. elegans. FEBS Lett. 581, 5445–5453. doi: 10.1016/j.febslet.2007.10.047
Kriegel, A., Andrés, Z., Medzihradszky, A., Krüger, F., Scholl, S., Delang, S., et al. (2015). Job sharing in the endomembrane system: Vacuolar acidification requires the combined activity of V-ATPase and V-PPase. Plant Cell 27, 3383–3396. doi: 10.1105/tpc.15.00733
Lantzouni, O., Alkofer, A., Falter-Braun, P., Schwechheimer, C. (2020). GROWTH-REGULATING FACTORS interact with DELLAs and regulate growth in cold stress. Plant Cell 32, 1018–1034. doi: 10.1105/tpc.19.00784
Lau, O. S., Bergmann, D. C. (2012). Stomatal development: a plant's perspective on cell polarity, cell fate transitions and intercellular communication. Development 139, 3683–3692. doi: 10.1242/dev.080523
Liebsch, D., Palatnik, J. F. (2020). MicroRNA miR396, GRF transcription factors and GIF co-regulators: a conserved plant growth regulatory module with potential for breeding and biotechnology. Curr. Opin. Plant Biol. 53, 31–42. doi: 10.1016/j.pbi.2019.09.008
Li, G., Hu, S., Hou, H., Kimura, S. (2019). Heterophylly: Phenotypic plasticity of leaf shape in aquatic and amphibious plants. Plants 8, 420–1–13. doi: 10.3390/plants8100420
Li, Y., Liu, Y., Zolman, B. K. (2019). Metabolic alterations in the enoyl-CoA hydratase 2 mutant disrupt peroxisomal pathways in seedlings. Plant Physiol. 180, 1860–1876. doi: 10.1104/pp.19.00300
Liu, S., Jobert, F., Rahneshan, Z., Doyle, S. M., Robert, S. (2021). Solving the puzzle of shape regulation in plant epidermal pavement cells. Annu. Rev. Plant Biol. 72, 525–550. doi: 10.1146/annurev-arplant-080720-081920
Liu, D., Song, Y., Chen, Z., Yu, D. (2009). Ectopic expression of miR396 suppresses GRF target gene expression and alters leaf growth in Arabidopsis. Physiol. Plant 136, 223–236. doi: 10.1111/j.1399-3054.2009.01229.x
Lucero, L. E., Uberti-Manassero, N. G., Arce, A. L., Colombatti, F., Alemano, S. G., Gonzalez, D. H. (2015). TCP15 modulates cytokinin and auxin responses during gynoecium development in Arabidopsis. Plant J. 8, 267–282. doi: 10.1111/tpj.12992
Lui, H., Wang, H., Delong, C., Fowke, L. C., Crosby, W. L., Fobert, P. R. (2000). The Arabidopsis Cdc2a-interacting protein ICK2 is structurally related to ICK1 and is a potent inhibitor of cyclin-dependent kinase activity in vitro. Plant J. 21, 379–385. doi: 10.1046/j.1365-313x.2000.00688.x
Lundin, M., Baltscheffsky, H., Ronne, H. (1991). Yeast PPA2 gene encodes a mitochondrial inorganic pyrophosphatase that is essential for mitochondrial function. J. Biol. Chem. 266, 12168–12172. doi: 10.1016/S0021-9258(18)98875-7
Mabuchi, K., Maki, H., Itaya, T., Suzuki, T., Nomoto, M., Sakaoka, S., et al. (2018). MYB30 links ROS signaling, root cell elongation, and plant immune responses. Proc. Natl. Acad. Sci. U.S.A. 115, E4710–E4719. doi: 10.1073/pnas.1804233115
Maeshima, M. (2000). Vacuolar H+-pyrophosphatase. Biochim. Biophys. Acta 1465, 37–51. doi: 10.1016/S0005-2736(00)00130-9
Micol, J. L. (2009). Leaf development: time to turn over a new leaf? Curr. Opin. Plant Biol. 12, 9–16. doi: 10.1016/j.pbi.2008.11.001
Mizukami, Y., Fischer, R. L. (2000). Plant organ size control: AINTEGUMENTA regulates growth and cell numbers during organogenesis. Proc. Natl. Acad. Sci. U. S. A. 97, 942–947. doi: 10.1073/pnas.97.2.942
Nakayama, H., Koga, H., Long, Y., Hamant, O., Ferjani, A. (2022). Looking beyond the gene network - metabolic and mechanical cell drivers of leaf morphogenesis. J. Cell Sci. 135, jcs259611. doi: 10.1242/jcs.259611
Nath, U., Crawford, B. C. W., Carpenter, R., Coen, E. (2003). Genetic control of surface curvature. Science 299, 1404–1407. doi: 10.1126/science.1079354
Nelissen, H., Eeckhout, D., Demuynck, K., Persiau, G., Walton, A., van Bel, M., et al. (2015). Dynamic changes in ANGUSTIFOLIA3 complex composition reveal a growth regulatory mechanism in the maize leaf. Plant Cell 27, 1605–1619. doi: 10.1105/tpc.15.00269
Noctor, G., Reichheld, J.-P., Foyer, C. H. (2018). ROS-related redox regulation and signaling in plants. Semin. Cell Dev. Biol. 80, 3–12. doi: 10.1016/j.semcdb.2017.07.013
Nozaki, M., Kawade, K., Horiguchi, G., Tsukaya, H. (2020). an3-mediated compensation is dependent on a cell-autonomous mechanism in leaf epidermal tissue. Plant Cell Physiol. 61, 1181–1190. doi: 10.1093/pcp/pcaa048
Omidbakhshfard, M. A., Sokolowska, E. M., Di Vittori, V., Perez de Souza, L., Kuhalskaya, A., Brotman, Y., et al. (2021). Multi-omics analysis of early leaf development in Arabidopsis thaliana. Patterns (N Y). 2, 100235. doi: 10.1016/j.patter.2021.100235
Ormenese, S. (2004). Analysis of the spatial expression pattern of seven kip related proteins (KRPs) in the shoot apex of Arabidopsis thaliana. Ann. Bot. 93, 575–580. doi: 10.1093/aob/mch077
Pavelescu, I., Vilarrasa-Blasi, J., Planas-Riverola, A., González-García, M., Caño-Delgado, A. I., Ibañes, M. (2018). A sizer model for cell differentiation in Arabidopsis thaliana root growth. Mol. Syst. Biol. 14, e7687. doi: 10.15252/msb.20177687
Penfield, S., Rylott, E. L., Gilday, A. D., Graham, S., Larson, T. R., Graham, I. A. (2004). Reserve mobilization in the Arabidopsis endosperm fuels hypocotyl elongation in the dark, is independent of abscisic acid, and requires PHOSPHOENOLPYRUVATE CARBOXYKINASE1. Plant Cell 16, 2705–2718. doi: 10.1105/tpc.104.024711
Petricka, J. J., Winter, C. M., Benfey, P. N. (2012). Control of Arabidopsis root development. Annu. Rev. Plant Biol. 63, 563–590. doi: 10.1146/annurev-arplant-042811-105501
Powell, A. E., Lenhard, M. (2012). Control of organ size in plants. Curr. Biol. 22, 360–367. doi: 10.1016/j.cub.2012.02.010
Ramirez-Parra, E., Gutierrez, C. (2007). E2F regulates FASCIATA1, a chromatin assembly gene whose loss switches on the endocycle and activates gene expression by changing the epigenetic status. Plant Physiol. 144, 105–120. doi: 10.1104/pp.106.094979
Rath, M., Challa, K. R., Sarvepalli, K., Nath, U. (2022). CINCINNATA-like TCP transcription factors in cell growth - an expanding portfolio. Front. Plant Sci. 13, 825341. doi: 10.3389/fpls.2022.825341
Roberts, I. M., Boevink, P., Roberts, A. G., Sauer, N., Reichel, C., Oparka, K. J. (2001). Dynamic changes in the frequency and architecture of plasmodesmata during the sink-source transition in tobacco leaves. Protoplasma 218, 31–44. doi: 10.1007/BF01288358
Rodriguez, R. E., Mecchia, M. A., Debernardi, J. M., Schommer, C., Weigel, D., Palatnik, J. F. (2010). Control of cell proliferation in Arabidopsis thaliana by microRNA miR396. Development 137, 103–112. doi: 10.1242/dev.043067
Roeder, A. H. K., Cunha, A., Ohno, C. K., Meyerowitz, E. M. (2012). Cell cycle regulates cell type in the Arabidopsis sepal. Development 139, 4416–4427. doi: 10.1242/dev.082925
Ruzicka, K., Strader, L. C., Bailly, A., Yang, H., Blakeslee, J., Langowski, L., et al. (2010). Arabidopsis PIS1 encodes the ABCG37 transporter of auxinic compounds including the auxin precursor indole-3-butyric acid. Proc. Natl. Acad. Sci. U.S.A. 107, 10749–10753. doi: 10.1073/pnas.1005878107
Rylott, E. L., Gilday, A. D., Graham, I. A. (2003). The gluconeogenic enzyme phosphoenolpyruvate carboxykinase in Arabidopsis is essential for seedling establishment. Plant Physiol. 131, 1834–1842. doi: 10.1104/pp.102.019174
Sakamoto, T., Ikematsu, S., Namie, K., Hou, H., Li, G., Kimura, S. (2022). Leaf cell morphology alternation in response to environmental signals in Rorippa aquatica. Int. J. Mol. Sci. 23, 10401–1–10. doi: 10.3390/ijms231810401
Savaldi-Goldstein, S., Peto, C., Chory, J. (2007). The epidermis both drives and restricts plant shoot growth. Nature 446, 199–202. doi: 10.1038/nature05618
Schilling, R. K., Tester, M., Marschner, P., Plett, D. C., Roy, S. J. (2017). AVP1: One protein, many roles. Trends Plant Sci. 22, 154–162. doi: 10.1016/j.tplants.2016.11.012
Schommer, C., Debernardi, J. M., Bresso, E. G., Rodriguez, R. E., Palatnik, J. F. (2014). Repression of cell proliferation by miR319-regulated TCP4. Mol. Plant 7, 1533–1544. doi: 10.1093/mp/ssu084
Segami, S., Tomoyama, T., Sakamoto, S., Gunji, S., Fukuda, M., Kinoshita, S., et al. (2018). Vacuolar H+-pyrophosphatase and cytosolic soluble pyrophosphatases cooperatively regulate pyrophosphate levels in Arabidopsis thaliana. Plant Cell 30, 1040–1061. doi: 10.1105/tpc.17.00911
Serralbo, O., Pérez-Pérez, J. M., Heidstra, R., Scheres, B. (2006). Non-cell-autonomous rescue of anaphase-promoting complex function revealed by mosaic analysis of HOBBIT, an Arabidopsis CDC27 homolog. Proc. Natl. Acad. Sci. U.S.A. 103, 13250–13255. doi: 10.1073/pnas.0602410103
Sieburth, L. E., Drews, G. N., Meyerowitz, E. M. (1998). Non-autonomy of AGAMOUS function in flower development: use of a Cre/loxP method for mosaic analysis in Arabidopsis. Development 125, 4303–4312. doi: 10.1242/dev.125.21.4303
Sinha, N. R., Rowland, S. D., Ichihashi, Y. (2016). Using gene networks in EvoDevo analyses. Curr. Opin. Plant Biol. 33, 133–139. doi: 10.1016/j.pbi.2016.06.016
Sizani, B. L., Kalve, S., Markakis, M. N., Domagalska, M. A., Stelmaszewska, J., AbdElgawad, H., et al. (2019). Multiple mechanisms explain how reduced KRP expression increases leaf size of Arabidopsis thaliana. New Phytol. 221, 1345–1358. doi: 10.1111/nph.15458
Stitt, M. (1998). Pyrophosphate as an energy donor in the cytosol of plant cells: an enigmatic alternative to ATP. Bot. Acta 111, 167–175. doi: 10.1111/j.1438-8677.1998.tb00692.x
Strader, L. C., Bartel, B. (2009). The Arabidopsis PLEIOTROPIC DRUG RESISTANCE8/ABCG36 ATP binding cassette transporter modulates sensitivity to the auxin precursor indole-3-butyric acid. Plant Cell 21, 1992–2007. doi: 10.1105/tpc.109.065821
Strader, L. C., Culler, A. H., Cohen, J. D., Bartel, B. (2010). Conversion of endogenous indole-3-butyric acid to indole-3-acetic acid drives cell expansion in Arabidopsis seedlings. Plant Physiol. 153, 1577–1586. doi: 10.1104/pp.110.157461
Strader, L. C., Wheeler, D. L., Christensen, S. E., Berens, J. C., Cohen, J. D., Rampey, R. A., et al. (2011). Multiple facets of Arabidopsis seedling development require indole-3-butyric acid-derived auxin. Plant Cell 23, 984–999. doi: 10.1105/tpc.111.083071
Sun, T., Gubler, F. (2004). Molecular mechanism of gibberellin signaling in plants. Annu. Rev. Plant Biol. 55, 197–223. doi: 10.1146/annurev.arplant.55.031903.141753
Szymkowiak, E. J., Sussex, I. M. (1996). What chimeras can tell us about plant development. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 351–376. doi: 10.1146/annurev.arplant.47.1.351
Tabeta, H., Higashi, Y., Okazaki, Y., Toyooka, K., Wakazaki, M., Sato, M., et al. (2022). Skotomorphogenesis exploits threonine to promote hypocotyl elongation. Quantitative Plant Biol. 3, E26. doi: 10.1017/qpb.2022.19
Tabeta, H., Watanabe, S., Fukuda, K., Gunji, S., Asaoka, M., Hirai, M. Y., et al. (2021). An auxin signaling network translates low-sugar-state input into compensated cell enlargement in the fugu5 cotyledon. PloS Genet. 17, e1009674. doi: 10.1371/journal.pgen.1009674
Takahashi, K., Morimoto, R., Tabeta, H., Asaoka, M., Ishida, M., Maeshima, M., et al. (2017). Compensated cell enlargement in fugu5 is specifically triggered by lowered sucrose production from seed storage lipids. Plant Cell Physiol. 58, 668–678. doi: 10.1093/pcp/pcx021
Takahashi, N., Lammens, T., Boudolf, V., Maes, S., Yoshizumi, T., De Jaeger, G., et al. (2008). The DNA replication checkpoint aids survival of plants deficient in the novel replisome factor ETG1. EMBO J. 27, 1840–1851. doi: 10.1038/emboj.2008.107
Trinh, D. C., Alonso-Serra, J., Asaoka, M., Colin, L., Cortes, M., Malivert, A., et al. (2021). How mechanical forces shape plant organs. Curr. Biol. 31, R143–R159. doi: 10.1016/j.cub.2020.12.001
Tsukagoshi, H. (2012). Defective root growth triggered by oxidative stress is controlled through the expression of cell cycle-related genes. Plant Sci. 197, 30–39. doi: 10.1016/j.plantsci.2012.08.011
Tsukagoshi, H., Busch, W., Benfey, P. N. (2010). Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 143, 606–616. doi: 10.1016/j.cell.2010.10.020
Tsukaya, H. (2002a). Interpretation of mutants in leaf morphology: Genetic evidence for a compensatory system in leaf morphogenesis that provides a new link between cell and organismal theories. Int. Rev. Cytol. 217, 1–39. doi: 10.1016/S0074-7696(02)17011-2
Tsukaya, H. (2002b). The leaf index : Heteroblasty, natural variation, and the genetic control of polar processes of leaf expansion. Plant Cell Physiol. 43, 372–378. doi: 10.1093/pcp/pcf051
Tsukaya, H. (2003). Organ shape and size : A lesson from studies of leaf morphogenesis. Curr. Opi. Plant Biol. 6, 57–62. doi: 10.1016/S1369526602000055
Tsukaya, H. (2008). Controlling size in multicellular organs: focus on the leaf. PloS Biol. 6, e174. doi: 10.1371/journal.pbio.0060174
Tsukaya, H. (2014). Comparative leaf development in angiosperms. Curr. Opin. Plant Biol. 17, 103–109. doi: 10.1016/j.pbi.2013.11.012
Tsukaya, H. (2018). Leaf shape diversity with an emphasis on leaf contour variation, developmental background, and adaptation. Semin. Cell Dev. Biol. 79, 48–57. doi: 10.1016/j.semcdb.2017.11.035
Ullah, H., Chen, J. G., Young, J. C., Im, K. H., Sussman, M. R., Jones, A. M. (2001). Modulation of cell proliferation by heterotrimeric g protein in Arabidopsis. Science 292, 2066–2069. doi: 10.1126/science.1059040
Vercruysse, J., Baekelandt, A., Gonzalez, N., Inzé, D. (2020). Molecular networks regulating cell division during Arabidopsis leaf growth. J. Exp. Bot. 71, 2365–2378. doi: 10.1093/jxb/erz522
Vercruyssen, L., Verkest, A., Gonzalez, N., Heyndrickx, K. S., Eeckhout, D., Han, S. K., et al. (2014). ANGUSTIFOLIA3 binds to SWI/SNF chromatin remodeling complexes to regulate transcription during Arabidopsis leaf development. Plant Cell 26, 210–229. doi: 10.1105/tpc.113.115907
Verkest, A., Manes, C. L., de, O., Vercruysse, S., Maes, S., van der Schueren, E., et al. (2005). The cyclin-dependent kinase inhibitor KRP2 controls the onset of the endoreduplication cycle during Arabidopsis leaf development through inhibition of mitotic CDKA;1 kinase complexes. Plant Cell 17, 1723–1736. doi: 10.1105/tpc.105.032383
Wang, H., Kong, F., Zhou, C. (2021). From genes to networks: The genetic control of leaf development. J. Integr. Plant Biol. 63, 1181–1196. doi: 10.1111/jipb.13084
Wang, H., Qi, Q., Schorr, P., Cutler, A. J., Crosby, W. L., Fowke, L. C. (1998). ICK1, a cyclin-dependent protein kinase inhibitor from Arabidopsis thaliana interacts with both Cdc2a and CycD3, and its expression is induced by abscisic acid. Plant J. 15, 501–510. doi: 10.1046/j.1365-313X.1998.00231.x
Wang, X., Wang, H., Liu, S., Ferjani, A., Li, J., Yan, J., et al. (2016). Genetic variation in ZmVPP1 contributes to drought tolerance in maize seedlings. Nat. Genet. 48, 1233–1241. doi: 10.1038/ng.3636
Wang, H., Zhou, Y., Gilmer, S., Whitwill, S., Fowke, L. C. (2000). Expression of the plant cyclin-dependent kinase inhibitor ICK1 affects cell division, plant growth and morphology. Plant J. 24, 613–623. doi: 10.1046/j.1365-313x.2000.00899.x
Wendehenne, D., Gao, Q., Kachroo, A., Kachroo, P. (2014). Free radical-mediated systemic immunity in plants. Curr. Opin. Plant Biol. 20, 127–134. doi: 10.1016/j.pbi.2014.05.012
Wood, E., Nurse, P. (2015). Sizing up to divide: Mitotic cell-size control in fission yeast. Annu. Rev. Cell Dev. Biol. 31, 11–29. doi: 10.1146/annurev-cellbio-100814-125601
Xu, X. M., Wang, J., Xuan, Z., Goldshmidt, A., Borrill, P. G. M., Hariharan, N., et al. (2011). Chaperonins facilitate KNOTTED1 cell-to-Cell trafficking and stem cell function. Science 333, 1141–1144. doi: 10.1126/science.1205727
Yuan, Z., Luo, D., Li, G., Yao, X., Wang, H., Zeng, M., et al. (2010). Characterization of the AE7 gene in Arabidopsis suggests that normal cell proliferation is essential for leaf polarity establishment. Plant J. 64, 331–342. doi: 10.1111/j.1365-313X.2010.04326.x
Zhang, D., Sun, W., Singh, R., Zheng, Y., Cao, Z., Li, M., et al. (2018). GRF-interacting factor1 regulates shoot achitecture and meristem determinacy in maize. Plant Cell 30, 360–374. doi: 10.1105/tpc.17.00791
Keywords: Arabidopsis thaliana, leaf morphogenesis, cell proliferation, post-mitotic cell expansion, compensation, cell-autonomous, non-cell-autonomous
Citation: Tabeta H, Gunji S, Kawade K and Ferjani A (2023) Leaf-size control beyond transcription factors: Compensatory mechanisms. Front. Plant Sci. 13:1024945. doi: 10.3389/fpls.2022.1024945
Received: 22 August 2022; Accepted: 28 December 2022;
Published: 23 January 2023.
Edited by:
Mary Byrne, The University of Sydney, AustraliaReviewed by:
Aashish Ranjan, National Institute of Plant Genome Research (NIPGR), IndiaJohn Paul Alvarez, Monash University, Australia
Chihiro Furumizu, Hiroshima University, Japan, in collaboration with reviewer JPA
Copyright © 2023 Tabeta, Gunji, Kawade and Ferjani. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ali Ferjani, ZmVyamFuaUB1LWdha3VnZWkuYWMuanA=
†These authors have contributed equally to this work and share first authorship
 Hiromitsu Tabeta
Hiromitsu Tabeta Shizuka Gunji
Shizuka Gunji Kensuke Kawade
Kensuke Kawade Ali Ferjani
Ali Ferjani