- 1National Glycoengineering Research Center and School of Life Science, Shandong University, Qingdao, China
- 2Key Laboratory of Marine Eco-Environmental Science and Technology, First Institute of Oceanography, Ministry of Natural Resources, Qingdao, China
- 3Shandong Key Laboratory of Carbohydrate Chemistry and Glycobiology, Shandong University, Qingdao, China
Flavonoids, the largest group of polyphenolic secondary metabolites present in all land plants, play essential roles in many biological processes and defense against abiotic stresses. In the flavonoid biosynthesis pathway, flavones synthase I (FNSI), flavanone 3-hydroxylase (F3H), flavonol synthase (FLS), and anthocyanidin synthase (ANS) all belong to 2-oxoglutarate/Fe(II)-dependent dioxygenases (2-ODDs) family, which catalyzes the critical oxidative reactions to form different flavonoid subgroups. Here, a novel 2-ODD gene was cloned from Antarctic moss Pohlia nutans (Pn2-ODD1) and its functions were investigated both in two model plants, Physcomitrella patens and Arabidopsis thaliana. Heterologous expression of Pn2-ODD1 increased the accumulation of anthocyanins and flavonol in Arabidopsis. Meanwhile, the transgenic P. patens and Arabidopsis with expressing Pn2-ODD1 exhibited enhanced tolerance to salinity and drought stresses, with larger gametophyte sizes, better seed germination, and longer root growth. Heterologous expression of Pn2-ODD1 in Arabidopsis also conferred the tolerance to UV-B radiation and oxidative stress by increasing antioxidant capacity. Therefore, we showed that Pn2-ODD1 participated in the accumulation of anthocyanins and flavonol in transgenic plants, and regulated the tolerance to abiotic stresses in plants, contributing to the adaptation of P. nutans to the polar environment.
Introduction
The 2-oxoglutarate/Fe(II)-dependent dioxygenases (2-ODDs) are non-heme iron-containing soluble proteins, which catalyze the oxidation reactions of diverse substrates, including hydroxylation, demethylation, desaturation, halogenation, and epimerization (Wang et al., 2021c; Wei et al., 2021). 2-ODDs mainly participate in the biosynthesis of various metabolic pathways, such as flavonoids, benzylisoquinoline alkaloids, glucosinolates, tropane alkaloids, and plant hormones (Kawai et al., 2014; Wang et al., 2019). Flavonoids are the largest class of specialized metabolites in plants, which have been isolated and identified more than 10,000 species. Despite their diversity, they all share the basic 15-carbon phenylpropanoid core (C6–C3–C6 skeleton structure) (Li C. et al., 2017). Flavonoids biosynthesis starts with the general phenylpropanoid pathway to produce p-coumaroyl-CoA. Then, p-coumaroyl-CoA and malonyl-CoA were catalyzed by chalcone synthase (CHS) and chalcone isomerase (CHI) to form naringenin, a key precursor of flavonoids biosynthesis (Jiang et al., 2016; Busche et al., 2021). Naringenin can be either converted by flavone synthase I and/or II (FNSI or FNSII) to produce flavone, or by flavanone-3-hydroxylase (F3H) to form dihydrokaempferol (DHK). Consequently, DHK can be converted to flavonols or anthocyanins by flavonol synthase (FLS) or dihydroflavonol reductase (DFR) and anthocyanin synthase (ANS) (Ahn et al., 2015; Wang et al., 2021b). Moreover, the modifications and derivatization of the carbon atoms of the basic skeleton, catalyzed by glycosyltransferase and O-methyltransferases, result in the diverse flavonoid groups in nature, including isoflavones, flavones, flavanones, flavanols, flavonols, and anthocyanins (Farrow and Facchini, 2014). Among the enzymes in the flavonoid biosynthesis pathway, flavones synthase I (FNSI), flavanone 3-hydroxylase (F3H), flavonol synthase (FLS), and anthocyanidin synthase (ANS)/leucoanthocyanidin dioxygenase (LDOX) belong to the 2-ODDs family, which requires 2-oxoglutarate, Fe2+, and ascorbate as co-factors to catalyze a series of oxidation reactions (Park et al., 2019; Busche et al., 2021).
FNSIs and F3Hs exhibit a relatively limited substrate selectivity, whereas FLSs and ANSs can accept a wide range of flavonoid compounds as possible substrates (Li et al., 2020a). FNSIs from primitive land plants (Plagiochasma appendiculatum, Physcomitrella patens, and Selaginella moellendorffii) displayed FNS and F3H dual activities, catalyzing naringenin into apigenin and DHK (Han et al., 2014; Li et al., 2020a). PcFNSI from Petroselinum crispum converted (2R,3S)-cis-DHK to kaempferol in vitro (Martens et al., 2003). F3Hs share high similarity with FNSIs and convert (2S)-flavanones into dihydroflavonols. FLSs are responsible for flavonol biosynthesis, in competition with DFR in the anthocyanin pathway for their common substrates DHK (Martens et al., 2010). Overexpression of FLSs could lead to the accumulation of flavonol and the reduction of anthocyanin in plant tissues (Luo et al., 2015; Jiang et al., 2020). In vitro, recombinant FLS protein from Oryza sativa not only converted DHK and dihydroquercetin (DHQ) into kaempferol and quercetin, but also catalyzed eriodictyol and naringenin to form DHQ and DHK, exhibiting FLS and F3H activities (Park et al., 2019). FLS from Citrus unshiu also catalyzed the unnatural (2R)-naringenin and natural (2S)-naringenin to yield dihydrokaempferol and kaempferol, respectively (Lukačin et al., 2003). FLSs share identity with ANSs at the polypeptide level and accept (2R,3S,4R)-leucoanthocyanidins as substrate (Turnbull et al., 2004). ANS can react with leucoanthocyanidins, flavanones, and dihydroflavonols, as well as (+)-Catechin, displaying a broad variety of substrate selectivity (Park et al., 2019). Functional defects or silencing of ANS affect the formation of plant color, thus resulting in the colorless or white organs (Rafique et al., 2016).
Flavonoids are extensively distributed in extant land plants, including bryophytes and vascular plants, which is thought to be crucial for plants land colonization and adaptation to terrestrial ecosystems (Jiang et al., 2016; Li et al., 2020a; Piatkowski et al., 2020). During plant evolution, different classes of flavonoids appeared sequentially (Koes et al., 1994). First, chalcones, flavanones, and flavones were found in the basal land plants liverworts due to the existence of functional FNSI, whereas proanthocyanidins were present in lycophytes (ferns). Furthermore, in addition to the generation of dominant flavone, both ferns and gymnosperms began to synthesize flavonols. In P. patens, flavonols were also detected (Wolf et al., 2010). Finally, flavonols and anthocyanins appeared with the emergence of angiosperms, harboring the true F3H genes (Li et al., 2020a). Flavonols participate in UV-B protection, male fertility, and regulating plant growth and development (Hamamouch et al., 2020). Anthocyanins, a water-soluble plant pigments, accountable for red to purple colors, are abundant in many plant tissues in seed plants (Tanaka et al., 2008; Pervaiz et al., 2017), which are involved in many critical biological functions, including attracting pollinators for pollination, providing resistance of herbivory, and absorbing UV-B radiation. (Ahn et al., 2015; Pervaiz et al., 2017). However, the downstream branch pathway of flavonoids biosynthesis in the early terrestrial plant bryophytes is not clear, and whether bryophytes can synthesize anthocyanins is still controversial.
Non-vascular plant liverworts can produce cell wall-localized red flavonoid pigment riccionidins (an auronidin), which is formed by a branch of phenylpropanoid metabolism pathway distinct from anthocyanins biosynthesis (Kunz et al., 1993; Berland et al., 2019). And moss Sphagnum capillifolium had been reported to generate sphagnorubins (Mues, 2000), described as “anthocyanin-like” pigments, which contributes to alleviating abiotic stresses, much like the anthocyanins in seed plants (Albert et al., 2018; Piatkowski et al., 2020). In P. patens, several probable 2-oxoglutarate-dependent dioxygenase genes were from its genome, and 2-ODD1 from P. patens exhibited FNSI/F3H activity and catalyzed naringenin to produce apigenin and dihydrokaempferol (Li et al., 2020a). However, there were no detectable anthocyanin pigments in stressed P. patens by high-performance thin-layer chromatography (Wolf et al., 2010). A phylogenetic analysis revealed that investigated seedless plants' liverworts, mosses, and ferns possessed no orthologs of ANS, and orthologs representing the complete anthocyanin biosynthetic pathway only existed in the seed plants (Piatkowski et al., 2020). However, some 2-ODD genes from S. moellendorffii and P. patens were also annotated as probable ANS. Furthermore, six anthocyanin compounds including peonidin 3-O-glucoside chloride, peonidin O-hexoside, pelargonidin, malvidin 3-O-glucoside, cyanidin 3-O-galactoside, and cyanidin 3-O-rutinoside were recently confirmed to be present in Antarctic moss Leptobryum pyriforme by a widely targeted metabolomics, and cyanidin 3-O-rutinoside was the significantly up-regulated metabolite with log2(Fold change) 14.68 under ultraviolet-B radiation (Liu et al., 2021).
In Antarctica, extreme aridity, cold, and high UV-B radiation severely restrict the growth of terrestrial plants (Singh et al., 2010). The Antarctic plants have evolved independently for millions of years due to the separation of Antarctica from other continents. Mosses, one of the dominant terrestrial plants in the limited ice-free ground, have evolved a series of special physiological mechanisms to adapt tough environments (Convey et al., 2014; Alavilli et al., 2017). For example, in three East Antarctic mosses, the accumulation of cell wall UV-B-absorbing compounds and red pigments acted as a photoprotective mechanism against UV-B radiation (Waterman et al., 2018). Antarctic moss Andreaea regularis displayed high levels of carotenoids and UV-B screening pigments in foliage in response to UV-B exposure (Newsham, 2003). Bioflavonoids extracted from Antarctic mosses Ceratodon purpureus exhibited UV screening and antioxidant activity, which may be accountable for the high resistance to UV-B radiation (Waterman et al., 2017). The transcriptome profiling of Antarctic moss Pohlia nutans under UV-B treatment indicated that the antioxidant system, DNA-repairing system, and flavonoids biosynthesis pathway contributed to the adaptation and survival of moss to UV-B radiation (Li et al., 2019). Antarctic moss Sanionia uncinata exhibited tolerance to desiccation due to the activation of protective mechanisms that are involved in increased activity of antioxidant enzymes, accumulation of osmotic adjustment compounds like proline, glycine betaine, and dehydrins proteins (Pizarro et al., 2019). In this study, we isolated a 2-ODD1 gene from the Antarctic moss P. nutans (Pn2-ODD1) and investigated its functions in P. patens and Arabidopsis. These results showed that heterologous expression of Pn2-ODD1 promoted the accumulation of anthocyanins and flavonol in transgenic plants, and conferred plant tolerance to salt, drought, and UV-B stress by increasing antioxidant capacity.
Materials and Methods
Plant Materials and Growth Conditions
Arabidopsis thaliana (Col-0) was used as the WT plant and for the generation of transgenic lines. Arabidopsis was grown in a greenhouse at 22°C and 60% relative humidity, with 8-h light/16-h dark (light intensity, 100 μmol·m−2·s−1) for vegetative growth. After about 4 weeks, they were cultured in a growth chamber with 16-h light for subsequent reproductive maturation. Physcomitrella patens were cultivated on BCD solid medium at 25°C, in 16-h light/8-h dark photoperiod (60 μmol·m−2·s−1). For the production of protonema cells, P. patens were homogenized by a polytron homogenizer and cultivated on agar medium (BCD, supplemented with 5 mM diammonium tartrate) overlaid cellophane for convenient subculture.
Bioinformatics Analysis
The sequence of Pn2-ODD1 gene was obtained from the transcriptome of Antarctic moss Pohlia nutans using HMMER program (Li et al., 2019). Subsequently, the multiple sequence alignments of Pn2-ODD1 and other plant 2-ODDs were performed using DNAMAN software. The neighbor-joining method was used to construct the phylogenetic tree using MEGA 5.0 software.
RNA Isolation and Quantitative Real-Time PCR Analysis
Total RNA was extracted from Arabidopsis and P. patens by TRIzol reagent and CTAB method. cDNA was synthesized from 2 μg RNA by 5 × All-In-One RT MasterMix (abm, Canada), according to the manufacturer's instructions. Then, quantitative real-time PCR analysis was performed in an LC480 Thermal Cycler instrument by using the SYBR qPCR Master Mix (Nuoweizan, Nanjing, China). AtTubulin and PpActin were used as an internal control. The expression levels of genes were presented by the comparative 2−ΔΔCt method (Livak and Schmittgen, 2001). All used primers were listed in Supplementary Table 1.
Plasmid Constructions and Plant Transformation
The full length of Pn2-ODD1 was amplified by PCR from cDNA of Pohlia nutans with the specific primers. And the PCR fragment was inserted into the SmaI restriction site of the pTFH15.3 vector by one-step cloning (Nuoweizan, Nanjing, China). Then, the constructed Pn2-ODD1-pTFH15.3 plasmid was linearized by NotI to introduce into P. patens protoplasts by PEG-mediated DNA uptake, with a slight modification (Cove et al., 2009). Different concentrations of antibiotic G418 (25, 50, or 100 μg·mL−1) were used to select surviving transformants. Finally, stable transformants were screened by PCR amplification for phenotypic analysis.
For expressed-Pn2-ODD1 Arabidopsis, Pn2-ODD1 gene was cloned into the pRI101 vector to obtain Pn2-ODD1-pRI101. The recombinant construct was then introduced into Arabidopsis (Col-0) plants via Agrobacterium-mediated infiltration method (Zhang et al., 2006). Positive transgenic seedlings were screened on 1/2 MS medium supplemented with 50 μg·mL−1 kanamycin. Two independent T3 Pn2-ODD1 transgenic lines were obtained for subsequent analysis.
Measurement of Anthocyanin and Flavonols Contents
For the observation of anthocyanin enrichment in 5-day-old Arabidopsis seedlings, the seeds were cultivated on 1/2 MS agar medium, with 24-h light for 5 days at 22°C (Li P. et al., 2017). For the accumulation of anthocyanins in 17-day-old seedlings, sterilized seeds were germinated on 1/2 MS solid medium containing 3% (w/v) sucrose in a culture room with 16-h light for 14 days, then transferred on 1/2 MS medium supplemented with 12% (w/v) sucrose for another 3 days (Yonekura-Sakakibara et al., 2012). The purple coloration of seedlings was recorded, and plant samples were collected for the spectrophotometry measurement of anthocyanin levels and the analysis of flavonoids or anthocyanins metabolomics.
Anthocyanin extraction from Arabidopsis was carried out as described in Xu et al. (2017). Briefly, homogenized samples were incubated with methanol-HCl (1%, v/v) at 4°C for 24 h in the dark. Then, the extracts were mixed with chloroform and distilled water to remove chlorophyll. After centrifugation at 12,000 g for 15 min, the supernatant was collected and measured the absorbance at 657 and 530 nm. The anthocyanin level was expressed as (A530-0.25*A657)/fresh weight (g).
The total flavonols were extracted from Arabidopsis according to the method of Wang et al. (2020), with a slight modification. 0.2 g of fresh samples were ground with liquid nitrogen and extracted using 50% methanol. After centrifugation at 13,000 g for 15 min, the supernatant was transferred into a new tube, followed by adding an equal volume of 2 N HCl and hydrolyzing at 70°C for 40 min. Then, 100% methanol was added to prevent the degradation of the aglycones. The mixture was centrifuged for 15 min at 13,000 g to collect the supernatants for HPLC analysis (Shimadzu LC-20A, Japan), equipped with an Agilent C18 column (4.6 ×250 mm, 5 μm). The mobile phase was 0.1% (v/v) formic acid (A) and 0.1% (v/v) formic acid in acetonitrile (B). The linear gradient conditions were as follows: 0 min, 5% B; 30 min, 55% B; 45 min, 65% B; 50 min, 100% B; 52-60 min, 5% B, at a flow rate of 1 mL·min−1. The column temperature was set at 30°C, and the flavonols were detected at 365 nm (Park et al., 2019).
Targeted Metabolomics Analysis
The flavonoids and anthocyanins metabolomics analyses in Arabidopsis plants were performed by Wuhan Metware Biotechnology Co., Ltd. (Wuhan, China). In general, the freeze-dried sample was homogenized into powder. For widely targeted flavonoids metabolomics, 0.1 g powder was dissolved with 70% aqueous methanol at 4°C overnight. After centrifugation, the filtered supernatant was analyzed using LC-MS/MS (UPLC, Shim-pack UFLC SHIMADZU CBM30A system; MS, Applied Biosystems 6500 Q TRAP). Based on a self-built database and a public metabolite database, the metabolites were qualitatively identified by secondary spectral properties. Differential metabolites with VIP (variable importance in the project) ≥1, fold change ≥1.2, or fold change ≤ 0.83 were filtered as significantly changed metabolites.
For targeted anthocyanins metabolomics analysis, 50 mg homogenized powder was incubated with an extraction solvent containing 0.5 mL methanol/water/hydrochloric acid (500:500:1, V/V/V) by vortex and ultrasound for 5 min. After that, the sample was centrifuged at 12,000 g at 4°C for 3 min. Finally, the supernatants were collected and filtrated to analyze by Ultra Performance Liquid Chromatography and ESItriple quadrupole-linear ion trap mass spectrometer (QTRAP). For the absolute quantitative analysis of anthocyanins, commercial standard substances were diluted to different concentrations to obtain the corresponding mass spectrum peak intensity. Then, the absolute contents of anthocyanins in extracted samples were calculated by the standard curves of different substances. The differential metabolites were selected according to the P values (≤ 0.05) and fold change (≥1.2 or ≤ 0.83).
Plant Stress Treatments
For stress treatments of transgenic P. paten, the stem tips with the same size of transgenic Pn2-ODD1 and WT gametophytes were placed on a BCD medium supplemented with different concentrations of NaCl and D-Mannitol for 5 weeks at 25°C. The visual phenotypes were photographed, and the colony diameters were measured.
For drought stress assay, 3-week-old Arabidopsis seedlings grown in soil containers were subjected to drought stress by depriving water for 21 days, and then rewatered for 3 days to calculate the survival rates. For the seed germination of Arabidopsis, the sterilized seeds were sown on the 1/2 MS solid medium with or without NaCl, D-mannitol, H2O2, and 3-amino-1,2,4-triazole (3-AT, a CAT inhibitor, triggering the accumulation of H2O2). Then plates were placed in the dark at 4°C for 2 days and then cultured in a greenhouse at 22°C for 4–8 days. The germination rate was represented by counting the proportion of cotyledon greening. For root length assay, seedlings were vertically grown on a 1/2 MS medium containing 75 or 100 mM NaCl, 0.2 or 0.3 M D-mannitol, and 0.75 or 1 mM H2O2. After 5–9 days, the root length was calculated using Image J software. For the expression analysis of genes under stress treatment, 2-week-old Arabidopsis were sprayed with 200 mM NaCl, 16% PEG, or 20 mM H2O2 for 2 h, and all plant samples were harvested and frozen at −80°C until use.
For UV-B treatment, 3-week-old WT and transgenic Pn2-ODD1 Arabidopsis seedlings were treated for 12 h with 0.25 mW·cm−2 UV-B intensity. After being recovered for 3 days, the seedlings were collected and immediately frozen in liquid nitrogen and stored at −80°C for further analysis. Arabidopsis cultivated under the normal light conditions was used as the control.
Chlorophyll Content Quantification
For the measurement of chlorophyll content, homogenized samples were extracted with 80% acetone at 4°C in the dark. After centrifugation, the supernatant was collected to detect the absorbance at 663 and 645 nm by spectrophotometry analysis. Total chlorophyll content was calculated using the following formula: (8.05 × A663 + 20.29 × A645) × mL acetone mg−1 fresh weight (Porra et al., 1989).
3,3′-Diaminobenzidine (DAB) and Nitrobluetetrazolium Blue Chloride (NBT) Staining
DAB and NBT staining were used to detect the accumulation of hydrogen peroxide (H2O2) and superoxide anion () as described previously (Wang et al., 2015), with a slight modification. Whole seedlings were incubated with 1 mg·mL−1 DAB solution or 1 mg·mL−1 NBT solution in 50 mM potassium phosphate buffer (pH7.8) for 12 h in the dark. Then, chlorophyll was completely removed by decolorization with a bleaching solution (ethanol:acetic acid:glycerol = 3:1:1) in a boiling water bath.
Statistical Analysis
All experiments were performed at least three times for biological replicates. All data were expressed as mean ± standard deviation (SD).
Results
Characterization and Sequence Analysis of Pn2-ODD1 Gene
The full-length cDNA sequence of Pn2-ODD1 was amplified by PCR, which contains an open reading frame of 1,086 bp, encoding a 40.3 kDa-polypeptide with 361 amino acids and a predicted isoelectric point of 5.53. Multiple alignment analysis showed that Pn2-ODD1 shared only 35.3% similarity with the 2-ODD1 from P. paten and 36.7% similarity with FNSI from P. nutans, whereas about 25.0% identity with other known species 2-ODDs. However, Pn2-ODD1 contained the conserved domains: Fe2+ binding site HxDxnH (His228, Asp230, and His285) and the 2-oxoglutarate (2-OG) binding domain RxS (Tyr213, Arg295, and Ser297) (Figure 1). Phylogenetic analysis showed that in the members of a 2-ODDs family in flavonoids biosynthesis pathway, ANSs, and FLSs belonged to one subclade, while Apiaceae FNSIs and F3Hs constituted another independent branch. Pn2-ODD1 clustered with FLSs and ANSs branch and was closely related to S. moellendorffii and P. patens 2-ODDs proteins (Figure 2).
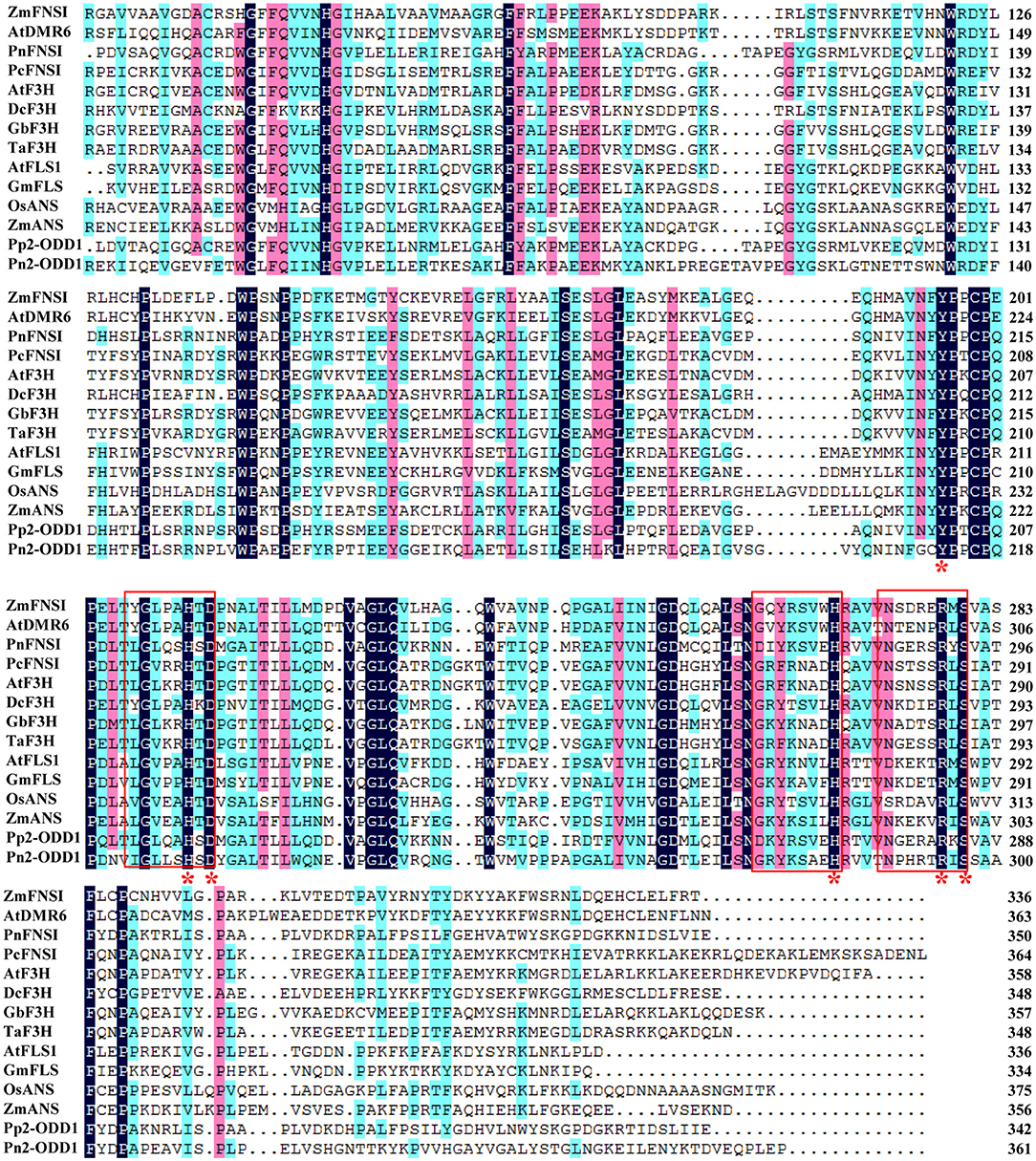

Figure 1. Pn2-ODD1 shared homology with other known species 2-ODDs. The amino acid sequence alignment of Pn2-ODD1 with 2-ODDs protein from other species. Red frames indicated ferrous iron binding site HXDX55H and 2-oxoglutarate (2-ODD) binding site RXS. Red asterisks (*) indicated the conserved amino acid residues of 2OG-FeII_Oxy conserved domain. Zea mays, ONL99521.1_ZmFNSI1; Arabidopsis thaliana, Q9FLV0.1_AtDMR6; Pohlia nutans, QCP71067.1_PnFNSI; Petroselinum crispum, AAP57393.1_PcFNSI; Arabidopsis thaliana, NP_190692.1_AtF3H; Dendrobium catenatum, PKU66857.1_DcF3H; Ginkgo biloba, AAU93347.1_GbF3H; Triticum aestivum, AFJ38181.1_TaF3H; Arabidopsis thaliana, NP_001190266.1_AtFLS1; Glycine max, NP_001237419.1_GmFLS; Oryza sativa, CAA69252.1_OsANS; Zea mays, and NP_001106074.1_ZmANS; Physcomitrella patens, XP_001780809.1_Pp2-ODD1.
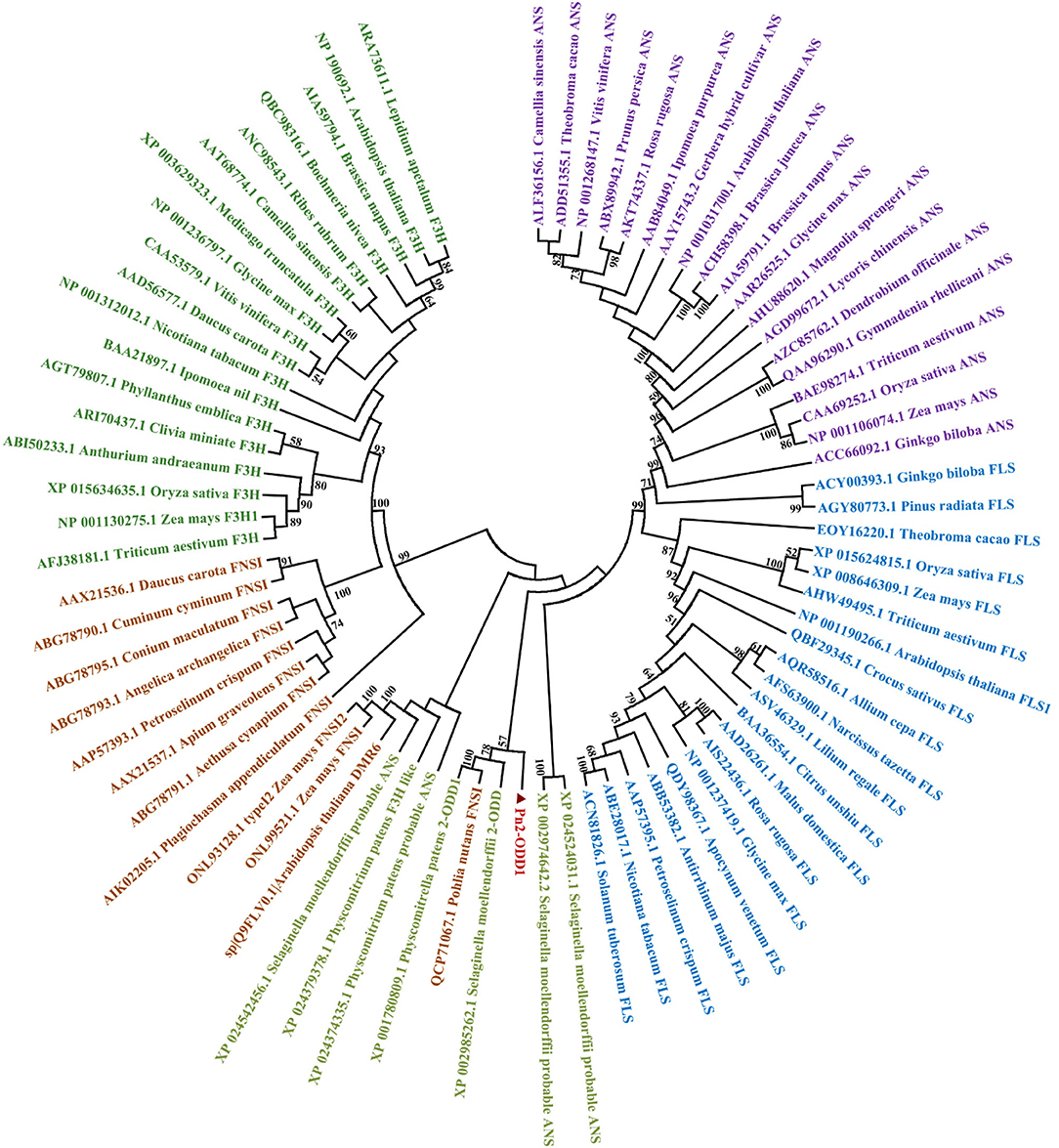
Figure 2. Phylogenetic relationship of Pn2-ODD1 and other known representative 2-ODDs (mosses Physcomitrella patens and Pohlia nutans, ferns Selaginella moellendorffii, gymnosperms, monocotyledons, and dicotyledons).
Pn2-ODD1 Contributed to the Accumulation of Anthocyanin and Flavonols in Transgenic Arabidopsis
Arabidopsis is a model organism in the plant kingdom, due to the easily quantitative stress-resistance indicators, clear flavonoids biosynthesis pathway, and regulatory networks, which is beneficial to promote the substantial development and application of genetic resources in higher plants. To explore the biological function of Pn2-ODD1, two independently expressed Pn2-ODD1 Arabidopsis were identified by genomic PCR (Supplementary Figure 1). Constant light can induce the accumulation of flavonoids in plants. After the sterilized seeds were sown on the 1/2 MS medium and placed vertically with the constant light for 5 days, both WT and Pn2-ODD1-expressed Arabidopsis displayed the accumulation of flavonoids, with the purple hypocotyls. However, AtOE lines accumulated higher total flavonoids and anthocyanin levels (spectrophotometry analysis), with the deeper hypocotyls, which were 1.30- and 1.35-fold in contrast with WT plants, respectively (Figures 3A–D). The flavonoids metabolomics of 5-day-old WT and AtOE lines were detected based on UPLC-MS/MS method. A total of 156 metabolites were identified, including 59 flavones, 71 flavonols, two isoflavones, 10 flavanols, three chalcones, three anthocyanins, and eight flavone C-glycosides (Figure 3E). According to VIP value ≥ 1 and fold-change threshold ≥ 1.2 or ≤ 0.83, 48 differential metabolites (36 up-regulate and 12 down-regulate) were screened (Figure 3F and Supplementary Table 2). Among them, flavonols were the most abundant compounds (16 up-regulated and six down-regulated) (Figure 3G). Notably, three detected anthocyanins were all up-regulated. KEGG classification analysis showed that the differential metabolites dominantly focused on the flavonoid and anthocyanin biosynthesis metabolism (Figure 3H).
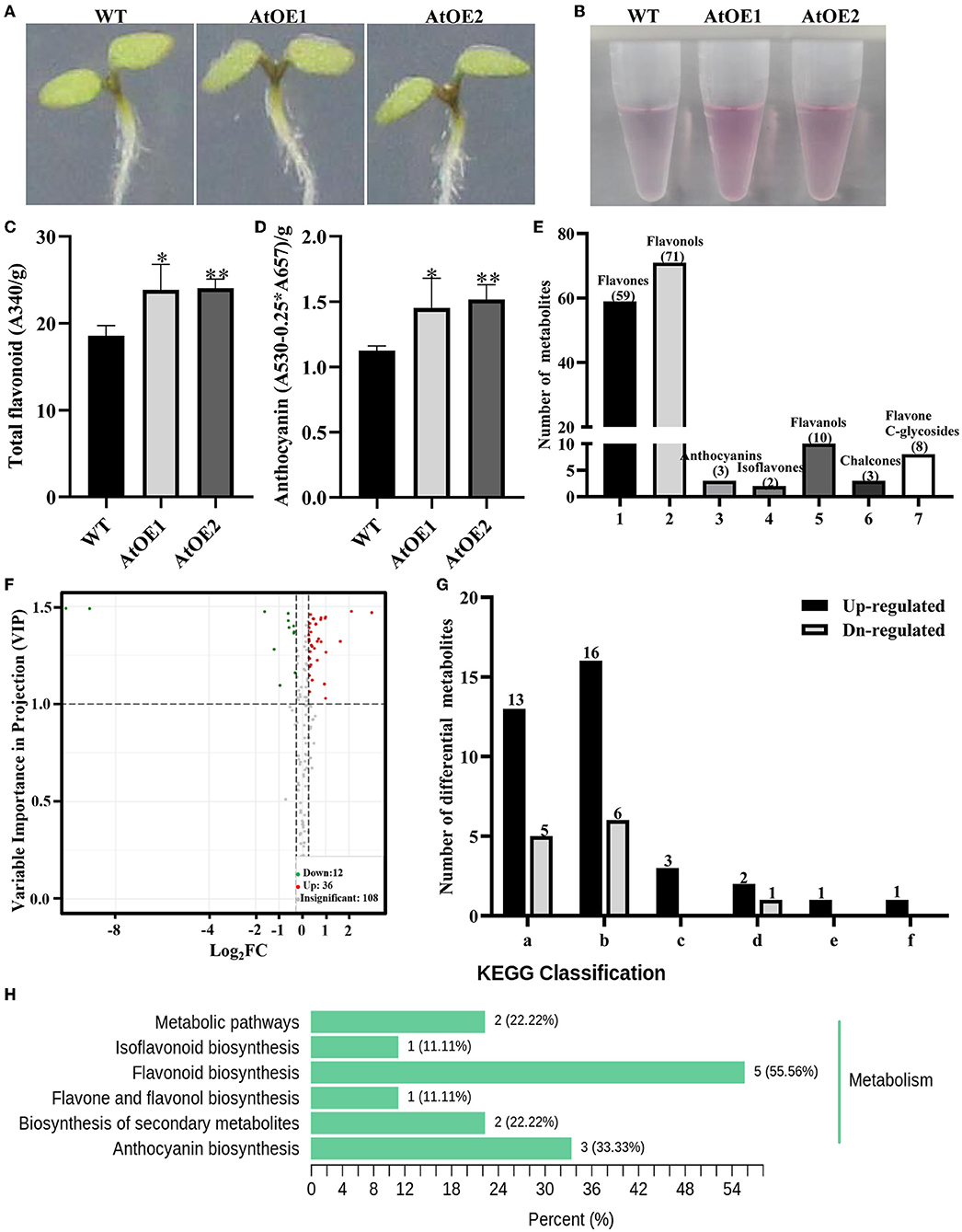
Figure 3. Pn2-ODD1 increased the levels of flavonoids in 5-day-old Arabidopsis. (A) Anthocyanin accumulation visualized by the purple coloration in 5-day-old seedlings with constant light. (B) Anthocyanin extraction solutions for WT and AtOE lines. (C,D) Contents of total flavonoids and anthocyanins in 5-day-old seedlings detected by spectrophotometry analysis. (E) Number of different classes of flavonoids in 5-day-old WT and transgenic Arabidopsis detected by the flavonoids metabolomics. (F) Volcano plot of samples. The volcano plot showed the levels of flavonoids metabolites and the statistical significance. Each point represents a metabolite. Horizontal ordinate indicates the fold change of flavonoids metabolites between WT and OE1, while VIP value means a significant difference in statistical analysis. (G) Statistical analysis of the classes of differential flavonoid metabolites. (a–f) Flavones, flavonols, anthocyanins, flavanols, chalcones, and flavone C-glycosides. (H) Differential metabolites KEGG classification. Asterisk (*) represents a significant difference between the WT plants and AtOE lines (Student's t-test, *P < 0.05, **P < 0.01).
Also, when cultivated on 1/2 MS medium containing sucrose, transgenic Pn2-ODD1 Arabidopsis exhibited greater anthocyanin accumulation, with deeper purple leaves, which was about 1.60-fold in contrast with WT plants (Figures 4A–C). Meanwhile, the expression levels of anthocyanin synthesis pathway genes AtPAL, AtDFR, and AtUFGT were increased in AtOE lines (Figure 4D). To further determine the difference in anthocyanin metabolism profiles, we performed the targeted anthocyanin metabolomics of AtOE lines and WT plants induced by sucrose. As shown in Figure 4E, the OE1 line had a 17.6% increase in the levels of total anthocyanins than that of the WT plants, which was consistent with the result measured by spectrophotometry analysis (anthocyanins at A530 nm). Totally, 31 anthocyanin metabolites were detected, including cyanidin (14), delphinidin (5), petunidin (3), pelargonidin (3), and peonidin (6). Based on a fold-change threshold ≥ 1.2 or ≤ 0.83, 7 up-regulated differential metabolites were identified (Figure 4F). Among them, cyanidin-3-O-(6-O-malonyl-beta-D-glucoside) and peonidin-3,5-O-diglucoside were the most significantly differential metabolites with log2(fold change), which were 1.65 and 1.56 folds compared with WT plants, respectively (Figure 4G). Then, the content of flavonol (including quercetin and kaempferol) in the WT plants and expressed-Pn2-ODD1 Arabidopsis grown with sucrose was analyzed using HPLC. The results showed that there was no significant difference in flavonol components (mainly quercetin and kaempferol) between WT plants and AtOE lines. However, the content of total flavonols in transgenic Arabidopsis was increased by 50.0%, in contrast with WT plants (Figures 4H,I). Among them, the levels of quercetin and kaempferol were significantly enhanced, which were about 65.0% and 50.0% higher than that of the WT plants, respectively (Figure 4J). These results suggested that the heterologous expression of Pn2-ODD1 resulted in the enrichment of anthocyanin and flavonols in Arabidopsis.
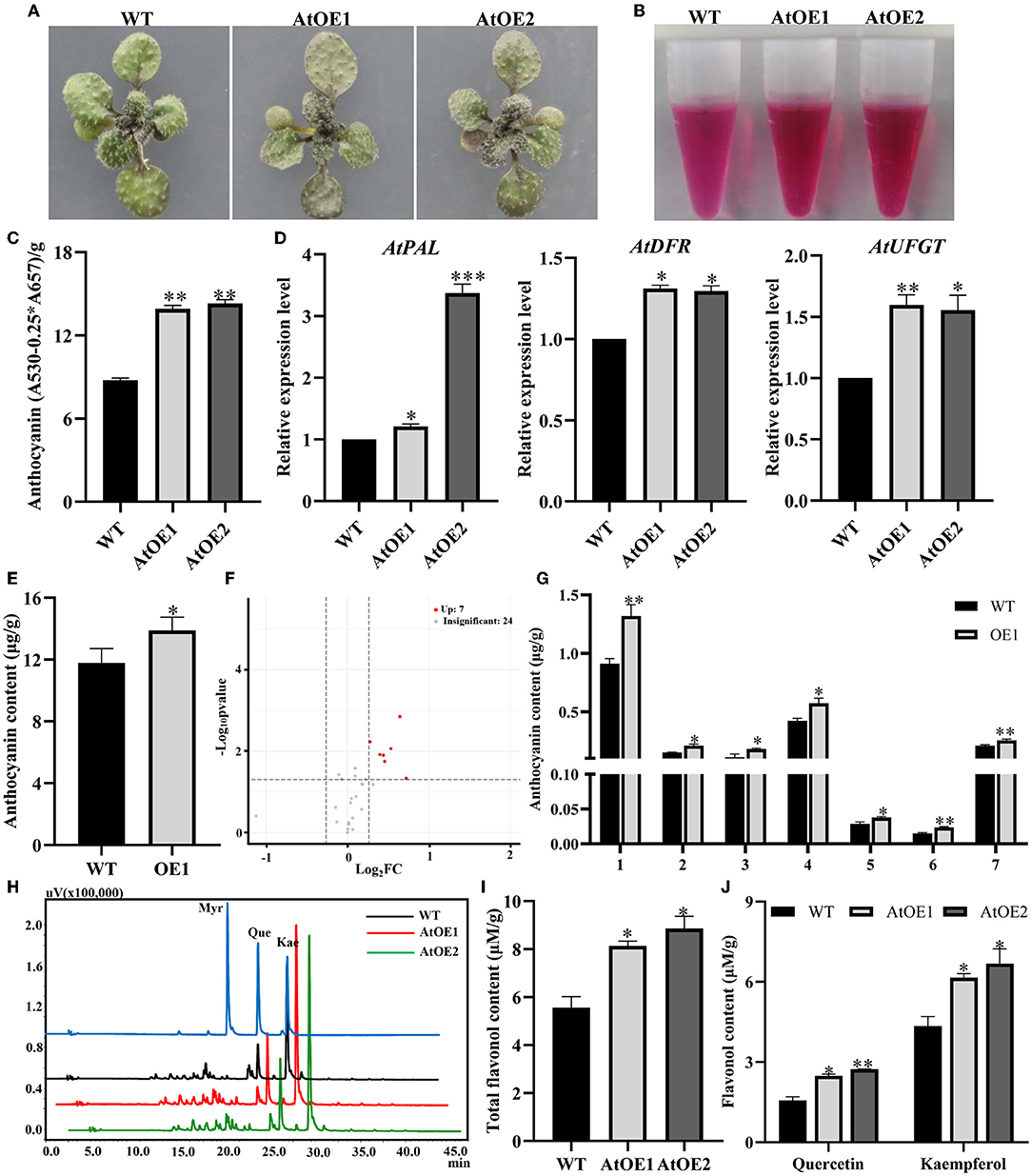
Figure 4. Pn2-ODD1 enhanced the accumulation of anthocyanins and flavonols in 17-day-old Arabidopsis seedlings induced by sucrose. (A) Anthocyanin accumulation visualized by the purple coloration in 17-day-old seedlings induced by sucrose (cultured on a medium containing 3% sucrose for 14 days, then transferred to 12% sucrose medium for 3 days). (B,C) Anthocyanin extraction solutions and anthocyanins contents (spectrophotometry analysis). (D) Expression levels of genes in anthocyanin biosynthesis pathway in WT and AtOE lines induced by sucrose. (E) The absolute quantitative analyses of total anthocyanins in 17-day-old Arabidopsis seedlings induced by sucrose detected by targeted metabolomics strategy. (F) Volcano plot of samples. It showed the levels of anthocyanins metabolites and the statistical significance of the difference. Each point represents a metabolite. Horizontal ordinate represents the fold change of anthocyanins metabolites between two samples, while p-value means significant difference in statistical analysis. (G) 7 up-regulated anthocyanins metabolites identified in the transgenic Pn2-ODD1 Arabidopsis. 1–7: Cyanidin-3,5-O-diglucoside, cyanidin-3-O-rutinoside, cyanidin-3-O-(6-O-malonyl-beta-D-glucoside), cyanidin-3-O-5-O-(6-O-coumaryl)-diglucoside, delphinidin-3-O-rutinoside, peonidin-3,5-O-diglucoside, and peonidin-3-O-(6-O-p-coumaryl)-glucoside. (H) HPLC profiles of flavonols standards (myricetin, quercetin, and kaempferol) and the flavonols extracts from WT and expressed-Pn2-ODD1 Arabidopsis (365 nm). (I,J) Contents of total flavonol, quercetin, and kaempferol in WT and AtOE lines induced by sucrose. Asterisk (*) represents a significant difference between the WT plants and AtOE lines (Student's t-test, *P < 0.05, **P < 0.01, ***P < 0.001).
Pn2-ODD1 Conferred the Tolerance to Salt Stress in Transgenic Physcomitrella patens and Arabidopsis
P. patens has become an important model plant for functional gene research due to its rapid growth cycle and mature genetic transformation methods (He et al., 2019; Rensing et al., 2020). Four independent transgenic P. patens (#1, #4, #7 and #8) were obtained and confirmed by genomic PCR analysis to further determine the function of Pn2-ODD1 in response to abiotic stress (Supplementary Figure 1). The stem tips with the same size of transgenic lines and WT plants were grown on BCD solid medium containing 100, 125, and 150 mM NaCl. Under normal medium, there was no obvious difference in growth performance between expressed-Pn2-ODD1 P. patens and WT plants. However, in the presence of NaCl, the transgenic P. patens displayed larger gametophytes than the WT plants. On 100 mM NaCl medium, the clone size of WT plants was 5.03 mm, whereas those of transgenic P. patens were 7.39, 7.50, 7.67, and 7.22 mm. On 125 or 150 mM NaCl, the diameter of the gametophyte of transgenic P. patens was still larger than that of the wild type, which was about an increase of 1.35- and 1.50-fold, respectively (Figures 5A,B). In Arabidopsis, no detectable differences in germination rate and root length were observed between WT and AtOE lines in the absence of NaCl. On 1/2 MS mediums supplemented with 75 mM NaCl, the germination rate of expressed-Pn2-ODD1 plants was 78.0% and 81.7%, which was about 15.0% higher than that of the WT plants. Similarly, at 100 mM NaCl, AtOE lines also displayed better germination, with an increase of 20.0% in comparison with WT plants (Figures 5C,D). For root length assays, the early root length in the transgenic Pn2-ODD1 Arabidopsis was markedly more vigorous, which was about 1.30-fold or 1.55-fold longer than that of the WT plants at 75 mM NaCl or 100 mM NaCl (Figures 5E,F).
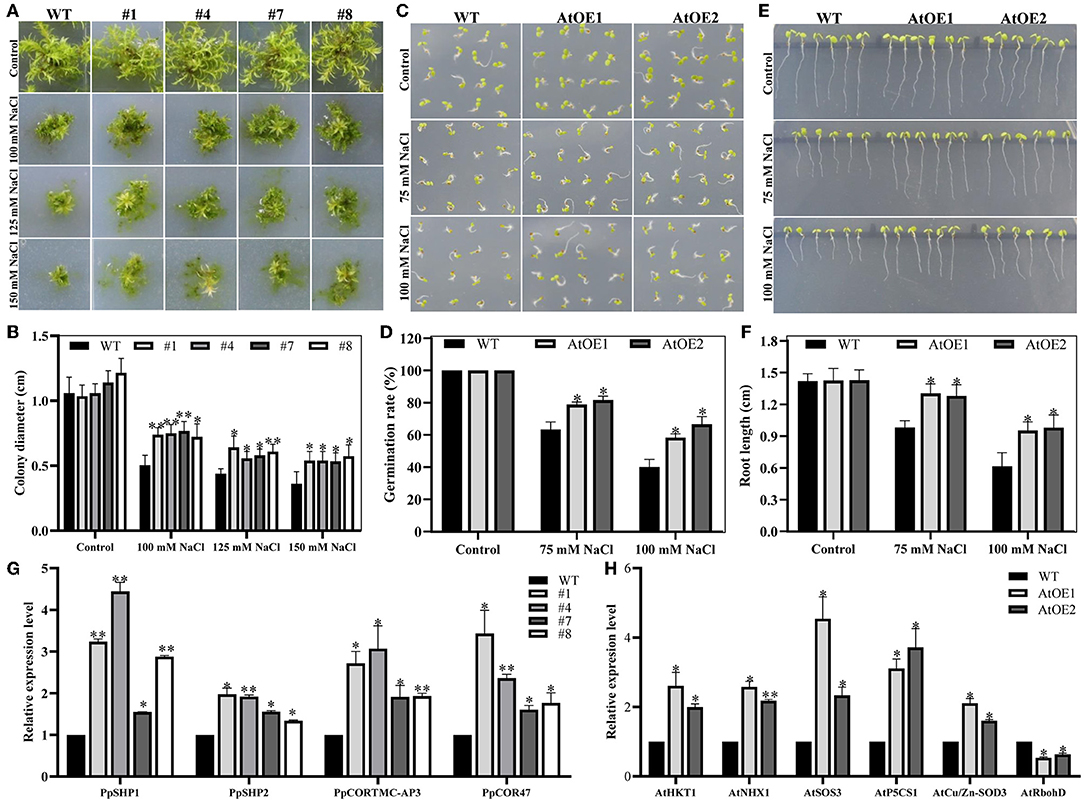
Figure 5. Heterologous expression of Pn2-ODD1 enhanced plant resistance to salt stress. (A) The transgenic P. patens displayed larger gametophytes than the WT plants under NaCl treatment. (B) Statistical analysis of gametophyte size as shown in (A). (C,E) AtOE lines exhibited higher germination rates and longer root length in contrast with WT plants under salt stress. (D,F) Statistical analysis of the seed germination rates and root length as shown in (C,E). (G,H) Expression patterns of stress-responsive genes in Pn2-ODD1 transgenic P. patens and Arabidopsis measured by qRT-PCR. Asterisk (*) represents a significant difference between the WT plants and AtOE lines (Student's t-test, *P < 0.05, **P < 0.01).
To further investigate the possible molecular mechanism of increased tolerance to salt stress in transgenic Pn2-ODD1 lines, the transcript levels of stress-responsive genes were analyzed by qRT-PCR. We found that the expression patterns of salt tolerance genes PpSHP1 and PpSHP2, stress responsible genes PpCOR TMC-AP3 and PpCOR47 were markedly up-regulated in transgenic P. patens, in contrast with WT plants (Figure 5G). Meanwhile, the expression patterns of salt tolerance genes AtHKT1, AtNHX1, AtSOS3, and AtP5CS1, and antioxidant enzymes gene AtCu/Zn-SOD3 were markedly increased in AtOE lines, while ROS generation gene AtRbohD was down-regulated in expressed-Pn2-ODD1 Arabidopsis (Figure 5H). Therefore, these results suggested that enhanced salt tolerance by Pn2-ODD1 might be correlated with upregulating several stress-related genes.
Pn2-ODD1 Enhanced the Resistance to Drought Stress in Transgenic Plants
Salt tolerance is usually associated with osmotic resistance. Thus, the growth performance of WT and expressed-Pn2-ODD1 plants under drought stress was analyzed. As shown in Figures 6A,B, the heterologous expression of Pn2-ODD1 significantly increased the resistance to D-mannitol stress in P. patens. On 0.3 M D-mannitol medium, the clone size of expressed-Pn2-ODD1 P. patens was 8.32, 8.73, 8.44, and 8.31 mm, which was about 40.0% higher than that of WT plants (6.11 mm), following the formation of protonema. When exposed to 0.4 M D-mannitol, the diameter of the protonema of transgenic P. patens was about 6.33 mm, which was 1.50-fold in contrast with WT plants. In Arabidopsis, 3-week-old seedlings with the same growth were treated with water withdrawal for 21 days. As shown in Figure 6G, AtOE lines exhibited better performance with a 65.0% survival rate, whereas WT plants were severely damaged and became wilted, with a 32.5% survival rate (Figure 6H). The sterilized seeds were sown on 1/2 MS medium supplemented with different concentrations of D-mannitol to observe the germination and root growth. On 0.2 M D-mannitol treatment, transgenic Pn2-ODD1 lines displayed 61.6–68.3% germination rates, which was about 20.0% higher than WT plants (Figures 6C,D). The root resistance assays demonstrated that AtOE lines exhibited longer root length, which was about 1.20-fold at 0.2 M D-mannitol and 1.60-fold at 0.3 M D-mannitol, in contrast with WT plants (Figures 6E,F). Drought stress can result in the generation of reactive oxygen species (ROS). The contents of H2O2 in AtOE lines were significantly lower than those of WT plants after 16% PEG treatment (Figure 6I). Antioxidant enzymes, such as CAT and SOD, participate in the reduction process of ROS (Choudhury et al., 2013). qRT-PCR analysis showed that the expression levels of ROS-scavenging related gene AtCAT1, AtFeSOD1, AtCu/Zn-SOD3, and proline biosynthesis-related gene AtP5CS1 were markedly increased in transgenic Pn2-ODD1 Arabidopsis (Figure 6J). These results demonstrated that Pn2-ODD1 contributed to enhanced resistance to drought stress by increasing ROS clearance capacity in transgenic plants.
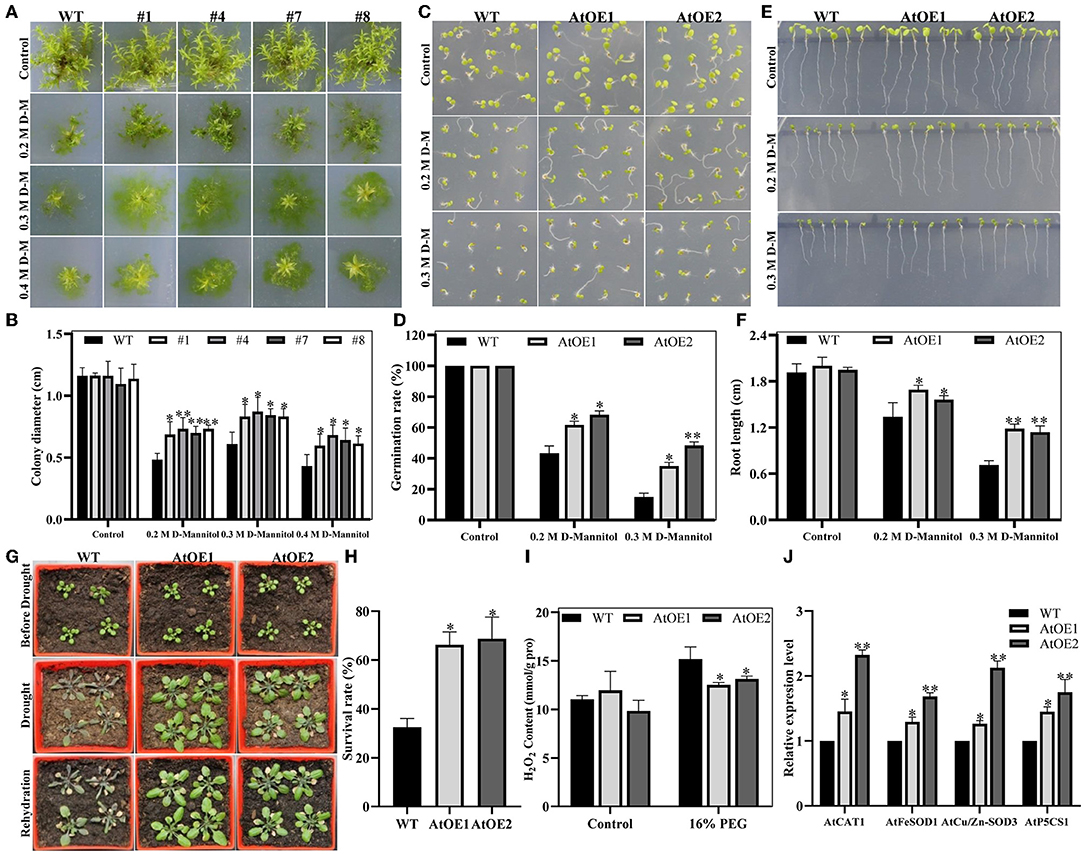
Figure 6. Pn2-ODD1 increased the resistance to drought stress in transgenic P. patens and Arabidopsis. (A) The diameter of transgenic P. patens gametophytes was markedly larger than that of the WT plants under drought stress conditions. (B) Statistical analysis of gametophyte size as shown in (A). (C,E) Expressed-Pn2-ODD1 Arabidopsis exhibited higher germination rates and longer root length compared with WT plants under osmosis stress. (D,F) Statistical analysis of the seed germination rates and root length as shown in (C,E). (G) Three-week-old Arabidopsis seedlings grown in soil were withheld water for 21 days, and then rewatered for 3 days. (H) Statistical analysis of the survival rate, as shown in G. (I) H2O2 content in Arabidopsis under 16% PEG treatment. (J) Gene expression levels of ROS-scavenging related genes in Arabidopsis analyzed by qRT-PCR. Asterisk (*) indicates a significant difference between the WT plants and AtOE lines (Student's t-test, *P < 0.05, **P < 0.01).
Heterologous Expression of Pn2-ODD1 Increased Arabidopsis Tolerance to UV-B Radiation and Oxidative Stress
To investigate whether Pn2-ODD1 could enhance the plant tolerance to UV-B stress, 3-week-old Arabidopsis were subjected to UV-B radiation with 0.25 mW·cm−2 UV-B intensity for 12 h and recovered for 3 days. When compared with WT plants, AtOE lines displayed better growth performance, with greener leaves and less damage (Figure 7A). The chlorophyll degradation ratio of transgenic Pn2-ODD1 Arabidopsis was lower than that of the WT plants under UV-B treatment (Figure 7B). DAB staining suggested that AtOE lines accumulated less endogenous H2O2 levels, in contrast with WT plants (Figure 7C). Furthermore, the transcript patterns of ROS-scavenging genes AtCu/Zn-SOD1, AtCu/Zn-SOD2, AtCu/Zn-SOD3, AtCAT1, and AtCAT3 were up-regulated in expressed-Pn2-ODD1 Arabidopsis (Figure 7D).
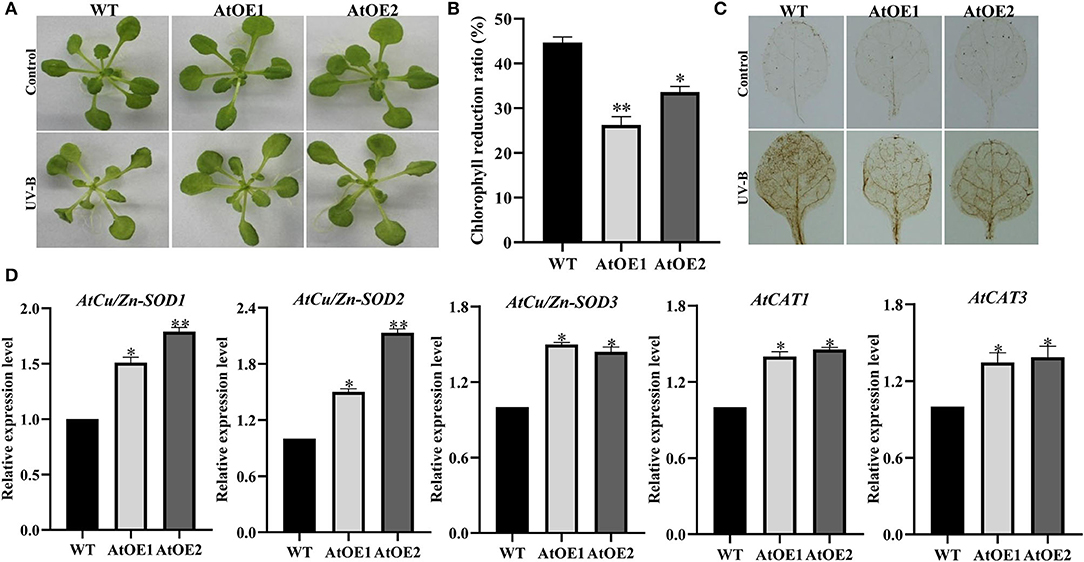
Figure 7. Heterologous expression of Pn2-ODD1 enhanced the Arabidopsis resistance to UV-B radiation. (A) Three-week-old Arabidopsis were treated with 0.25 mW·cm−2 UV-B intensity for 12 h and then recovered for 3 days. (B) Chlorophyll degradation ratio of WT plants and AtOE lines. (C) Levels of hydrogen peroxide (H2O2) measured by 3,3′-diaminobenzidine (DAB) staining under UV-B radiation. (D) Gene expression patterns of antioxidant enzymes under UV-B treatment. Asterisk (*) represents a significant difference between the WT plants and AtOE lines (Student's t-test, *P < 0.05, **P < 0.01).
In general, abiotic stresses lead to a build-up of ROS, which results in oxidative stress (Cruz de Carvalho, 2008; He et al., 2018). H2O2 and 3-amino-1,2,4-triazole (3-AT, a CAT inhibitor, triggering the accumulation of H2O2) were used to simulate exogenous and endogenous oxidative stress for young seedlings, respectively. The results showed that heterologous expression of Pn2-ODD1 conferred the enhanced tolerance to exogenous and endogenous oxidative stress in Arabidopsis. In the presence of 0.75 mM H2O2, the root length of AtOE lines was 1.86 and 1.94 cm, which was an increase of 30.5%, compared with the WT plants (Figures 8A,B). Furthermore, the lateral root numbers of AtOE lines were markedly higher than that of the WT plants, which were about 1.60-fold longer at 0.75 mM H2O2, whereas 1.35-fold longer at 1.0 mM H2O2 (Figure 8C). qRT-PCR analysis demonstrated that the expression levels of AtFeSOD1, AtFeSOD2, and AtCAT1, encoding ROS-scavenging enzymes, were significantly up-regulated in expressed-Pn2-ODD1 Arabidopsis (Figure 8D). Under endogenous oxidative stress, AtOE lines displayed reduced sensitivity to 3-AT, with greener leaves and lower H2O2 and levels, in contrast with WT plants (Figures 8E–G). Previous reports had demonstrated that ROS can trigger the accumulation of anthocyanins (Xu et al., 2017). Consistent with the results, under the treatment of 3-AT for 7 days, the levels of total flavonoid and anthocyanins in AtOE lines were markedly increased by 15.0% and 40.0%, in contrast with WT plants, respectively (Figures 8H,I). Thus, these results illustrated that Pn2-ODD1 improved the resistance to oxidative stress by increasing ROS scavenger and anthocyanins accumulation.
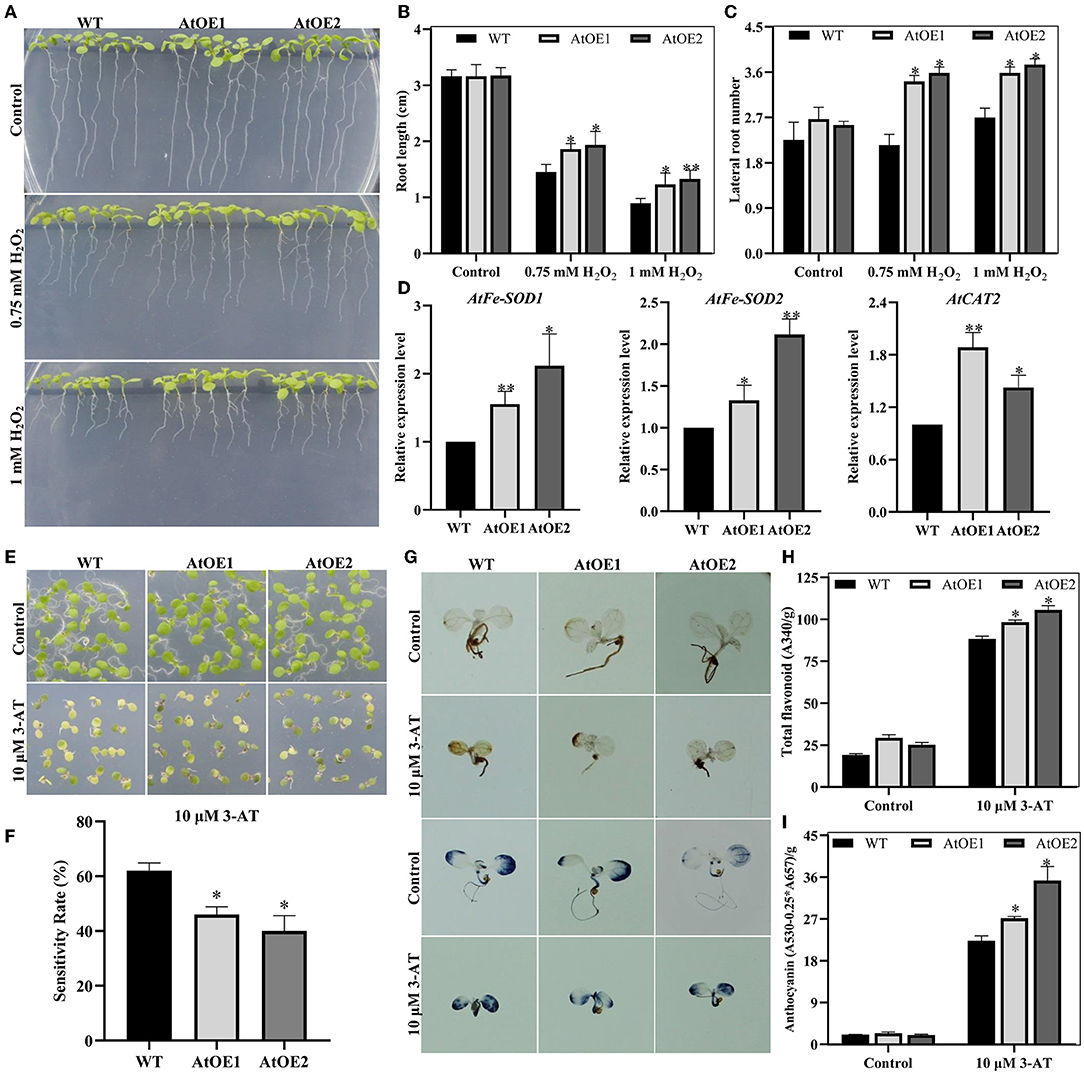
Figure 8. Pn2-ODD1 conferred the tolerance to oxidative stress in Arabidopsis. (A) Primary root length of Arabidopsis seedlings under oxidative stress. (B,C) Statistical analysis of root length and lateral root number as shown in (A). (D) Expression patterns of ROS-scavenging protein (i.e., AtFeSOD1, AtFeSOD2, and AtCAT1) in Arabidopsis under oxidative stress. (E,F) Growth phenotype and statistical analysis of WT and AtOE lines under the treatment of 10 μM 3-AT for 7 days. (G) 3,3′-diaminobenzidine (DAB) and nitrobluetetrazolium blue chloride (NBT) staining. (H,I) Content of total flavonoids and anthocyanins in WT and AtOE lines under 3-AT treatment for 7 days. Asterisk (*) represents a significant difference between the WT plants and AtOE lines (Student's t-test, *P < 0.05, **P < 0.01).
Discussion
2-Oxoglutarate/Fe(II)-dependent dioxygenases (2-ODDs), non-heme iron-containing soluble proteins, participate in the synthesis of important plant hormones and diverse secondary metabolites (Farrow and Facchini, 2014; Wei et al., 2021). Flavonoids, the natural products present in plants, possess enormous chemical diversity due to the organization and modifications of the three-ring structure by 2-ODDs, cytochrome P450-dependent oxygenases (CYPs), glycosyltransferase, and O-methyltransferases (Wang et al., 2019). In flavonoids biosynthesis, the key enzymes of downstream branch pathways FNSI, F3H, FLS, and ANS all belong to the 2-ODDs family, which is involved in the synthesis of key flavonoids compounds, such as flavones, flavonols, and anthocyanins. During the plant land colonization, the acquisition of flavonoids is considered to be an important adaptation for defense against the adverse environmental stresses faced with the transition to a non-aquatic lifestyle (Albert et al., 2018). Bryophytes are thought to be the oldest terrestrial plants, which exhibit the simple morphological organization, but the complex chemical diversification (Wellman et al., 2003; Asakawa et al., 2013; Ludwiczuk and Asakawa, 2019). Although there are several reports on the function of enzymes (4CL, CHS, CHI, and FNSI) in the upstream flavonoids pathway in bryophytes (Han et al., 2014; Gao et al., 2015; Yu et al., 2015; Cheng et al., 2018), the enzymes responsible for the downstream branch pathways of flavanone in bryophytes remain unclear. In particular, whether anthocyanin and anthocyanin synthase are present in bryophytes is still controversial. In this study, a Pn2-ODD1 gene from Antarctic moss Pohlia nutans and its roles in flavonoids metabolism and abiotic stresses were studied. The amino acid sequence of Pn2-ODD1 shared only 35.3% identity with 2-ODD1 of P. patens and 36.8% identity with PnFNSI, whereas below 30.0% sequence identity with 2-ODDs from other plants. But Pn2-ODD1 possessed the conserved Fe2+ binding domains and 2-oxoglutarate (2-OG) binding domains (Figure 1), which was consistent with other members of the 2OG-FeII_Oxy dioxygenase family (Reddy et al., 2007; Han et al., 2014; Zhang et al., 2016; Wang et al., 2020). Phylogenetic analysis demonstrated that Pn2-ODD1 had a closer relationship with ANSs and FLSs, which clustered with the 2-ODDs from P. patens and S. moellendorffii (Figure 2).
2-ODDs family of flavonoids synthesis pathway participate in catalyzing the oxidation of the flavonoid “C ring” to form different flavonoid subclasses. F3Hs and FNSIs can both react with flavanones, while FNSIs in primitive land plants also displayed F3H activity (Li et al., 2020a). Both FLSs and ANSs can accept the unnatural (2R)-naringenin and the natural (2S)-naringenin, dihydroflavonols as well as leucoanthocyanidins as substrates (Turnbull et al., 2000, 2004; Lukačin et al., 2003). ANS also catalyzes (+)-Catechin to cyanidin and a dimeric flavan-3-one (Wellmann et al., 2006). Although 2-ODDs displayed the versatility and mutual substitutability in substrate selectivity, they performed their respective major functions of the flavonoid biosynthesis metabolism in plants. The accumulation of flavonoids can be changed by operating the heterologous or homologous expression of genes in the flavonoid biosynthetic pathway to further clarify the function of 2-ODDs in plants. Overexpression of MnFNSI from Morus notabilis in tobacco increased the levels of flavones in leaves and decreased the accumulation of anthocyanin in flowers (Li et al., 2020b). Overexpression of F3Ha and F3Hb from Camellia sinensis significantly enhanced the accumulation of oligomeric proanthocyanidins and flavonol glycosides, whereas the contents of monocatechin derivatives were decreased in Arabidopsis (Han et al., 2017). Antisense down-regulation of ANS in Medicago truncatula caused a reduction of anthocyanins in foliar (Pang et al., 2007). The ans/fls1-2 seedlings complemented ANS from Musa spp. exhibited significantly increased levels of anthocyanins, compared with ans/fls1-2 plants (Busche et al., 2021). In this study, the contents of total flavonoids and anthocyanins in AtOE lines grown for 5 days were significantly increased in contrast with WT plants (Figures 3A–D). Metabolomics is a valuable approach to analyze the chemical complexity and measure the bioaccumulated phytochemicals, which reveals the relationship between the metabolites and the physiological state in response to environmental or genetic changes (Abdelhafez et al., 2020). The widely targeted flavonoids metabolomics analysis showed that although these significantly changed metabolites were mainly flavones and flavonols, three detected anthocyanins were found to be all significantly up-regulated (Figures 3E–H). At present, it is still controversial whether anthocyanin is synthesized in bryophytes and anthocyanidin synthase in lower plants has not been reported. Therefore, we focused on the effect of Pn2-ODD1 on anthocyanin synthesis. Sucrose can induce the accumulation of anthocyanins by increasing the expression of anthocyanins biosynthesis genes in plants (Teng et al., 2005; Solfanelli et al., 2006; Yoon et al., 2021). When induced by sucrose, the spectrophotometry analysis and targeted anthocyanin metabolomics demonstrated that heterologous expression of Pn2-ODD1 significantly enhanced the levels of anthocyanins (Figures 4A–G). Meanwhile, heterologous expression of Pn2-ODD1 also promoted the accumulation of flavonols in Arabidopsis (Figures 4H–J).
Heterologous expression of FLS from Muscari aucheri in tobacco increased the total flavonol level, whereas the total anthocyanin content in the petals was reduced (Liu et al., 2019). The antisense expression of FLS genes led to a reduction in the contents of flavonols and an increase in the accumulation of anthocyanins, when compared with the non-transformed plants (Holton et al., 1993; Nielsen et al., 2002). Also, overexpression of ANS from Theobroma cacao in tobacco led to flower petal color changes, which was consistent with an increased level of anthocyanins in flower petals (Liu et al., 2013). Previously, transgenic rice with ANS accumulated the increased anthocyanins and flavonols, and the reduced proanthocyanin levels, suggesting that rice ANS may be a multifunctional dioxygenase (Reddy et al., 2007). Overexpressed-RtLDOX2 Arabidopsis exhibited the enhanced anthocyanin and flavonol contents, possibly due to the versatility and mutual substitutability of RtLDOX2 in anthocyanin and flavonol biosynthesis (Li et al., 2021). In Arabidopsis, fls1-2 mutant seedlings displayed the decreased flavonol glycosides levels and accumulated the glycosylated forms of dihydrofavonols, and the FLS-like side activity of LDOX in plants resulted in the remaining flavonol glycoside accumulation (Stracke et al., 2009). Therefore, we concluded that Pn2-ODD1 contributed to the accumulation of anthocyanins and flavonol in Arabidopsis.
2-ODDs in the flavonoids biosynthesis pathway are involved in the growth and development, as well as stress responses of plants (Wei et al., 2021). Salinity stress can adversely hamper plant germination and growth, transpiration, and photosynthesis, which is one of the major threats to plant productivity (Wang et al., 2021a). The accumulation of flavonoids can be induced in response to salt stress (Arif et al., 2020). Overexpressed-CsF3H (from Camellia sinensis) tobacco exhibited increased tolerance to salinity stress by improved antioxidant system (Mahajan and Yadav, 2014). Overexpression of FLS1 from Triticum aestivum enhanced the root length of Arabidopsis seedlings under salinity stress (Wang et al., 2014a). Also, overexpression of ANS from Morus alba L. promoted the resistance to NaCl and mannitol stress in transgenic tobacco (Li et al., 2018). Similar resistant phenotypes were also observed in transgenic Pn2-ODD1 plants. Heterologous expression of Pn2-ODD1 enhanced the resistance to salinity stress in P. patens and Arabidopsis, with larger diameters of gametophytes, increasing seed germination and root elongation, when exposed to NaCl stress (Figure 5). Plant can regulate ion homeostasis and compartmentalization by ion influx in response to salinity stress (Gupta and Huang, 2014). PpSHP1 and PpSHP2, encoding a small hydrophobic protein with two transmembrane domains, are involved in the maintenance of membrane structure and function under environmental stress and preventing over-accumulation of K+ and Na+ ions (Wang et al., 2014b). PpCOR47, the homolog of Arabidopsis LEA-like protein, protects cells from water stress and can be induced by salt and osmotic stresses (Gilmour et al., 1992). PpCOR TMC-AP3, homologous to chloroplastic amino acid-selective channel protein from barley, regulates the amino acid transportation between chloroplast and cytoplasm to resynthesize damaged proteins (Frank et al., 2005). In this study, the expression patterns of PpSHP1, PpSHP2, PpCOR47, and PpCOR TMC-AP3 were significantly increased in transgenic Pn2-ODD1 P. patens under NaCl stress (Figure 5G). HKT and NHX, encoding K+ transporters and Na+/H+ exchangers, participate in maintaining ion homeostasis by controlling the transportation of Na+ and K+ during salinity stress (Gupta and Huang, 2014). SOS3 encodes a myristoylated Ca2+ binding protein with three EF hands, which is involved in salt tolerance through mediating Ca2+-dependent microfilament (MF) reorganization (M. Ishitani et al., 2000). P5CS1 catalyzes Glu to form Glu semialdehyde and is the key enzyme in the biosynthesis of proline, which is an osmoprotectant and stabilizes cellular structures, and keeps redox equilibrium in abiotic stresses (Aleksza et al., 2017). Overexpression of AvFLS from Apocynum venetum enhanced salt stress tolerance in tobacco by maintaining Na+/K+ homeostasis and increasing antioxidant enzyme activity (Wang et al., 2021a). We found that the transcript levels of AtHKT1, AtNHX1, AtSOS3, and AtP5CS1 were obviously up-regulated in transgenic Pn2-ODD1 Arabidopsis (Figure 5H).
Drought stress can lead to a decline in quantity and quality of crop production (Naing et al., 2018). The biosynthesis of flavonoids in plants can be up-regulated when subjected to drought stress (Ma et al., 2014). Metabolome and transcriptome profiling in Arabidopsis showed that the overaccumulation of flavonoids was involved in enhanced tolerance to drought and oxidative stress in MYB overexpressors, transparent testa4 (tt4), and WT plants (Nakabayashi et al., 2014). Overexpression of PnFNSI contributed to the increased drought resistance by enhancing the antioxidant capacity in Arabidopsis (Wang et al., 2020). Overexpression of RtLDOX2 from Reaumuria trigyna conferred enhanced tolerance to drought, UV-B, and salt stress in Arabidopsis by promoting the levels of anthocyanins and flavonols (Li et al., 2021). Here, heterologous expression of Pn2-ODD1 contributed to the improved tolerance to drought stress in plants (Figure 6). The generation of ROS can be caused by drought stress. Reducing ROS enrichment in transgenic plants usually increased the resistance to drought stress (Choudhury et al., 2013). Transgenic tobacco overexpressing F3H from Lycium chinense displayed enhanced resistance to drought stress, improving the antioxidant system (Song et al., 2016). In this study, after 16% PEG treatment, Pn2-ODD1 decreased the levels of H2O2 in Arabidopsis (Figure 6I). Superoxide dismutase (SOD) and catalase (CAT) are important antioxidant enzymes, which participate in the process of scavenging ROS in plants. SOD catalyzes the conversion of superoxide anions into H2O2 and O2, and CAT converts H2O2 to produce H2O and O2 (Sharma et al., 2012). qRT-PCR analysis showed that Pn2-ODD1 markedly up-regulated the expression patterns of AtCAT1, AtFeSOD1, and AtCu/Zn-SOD3 under 16% PEG treatment (Figure 6J).
Due to lower stratospheric ozone levels, plants are exposed to increased solar UV-B irradiation, which leads to oxidative damage to DNA, proteins, and lipids (Wolf et al., 2010). Flavonoids compounds, as a non-enzymatic antioxidant system, exhibit high antioxidant activity to protect plants from UV-B and oxidative damage (Buer et al., 2010). In Arabidopsis, anthocyanin-deficient mutants generated more ROS in vivo, accompanied by reduced antioxidant ability (Xu et al., 2017). Overexpression of rice ANS in rice mutant Nootripathu (NP) exhibited enhanced antioxidant activity by promoting the accumulation of anthocyanin and other flavonoids (Reddy et al., 2007). Overexpressing ZmFNSI or ZmFNSII Arabidopsis displayed less UV-B-induced damage due to the accumulation of apigenin, when compared with WT plants (Righini et al., 2019). In our study, heterologous expression of Pn2-ODD1 conferred the tolerance to UV-B stress, exhibiting lower ROS levels and increased expression of antioxidant enzymes genes in Arabidopsis (Figure 7). Consistent with these results, we observed that heterologous expression of Pn2-ODD1 in Arabidopsis conferred the tolerance to exogenous oxidative stress with longer primary roots and up-regulated the transcript levels of ROS scavenging gene, AtFeSOD1, AtFeSOD2, and AtCAT1 (Figures 8A–D). ROS can induce the production of anthocyanins, and anthocyanin-deficient mutants accumulated more endogenous ROS and were hypersensitive to ROS (Xu et al., 2017). 3-AT, a CAT inhibitor, was used to stimulate endogenous oxidative stress. AtOE lines seedlings displayed lower sensitivity to ROS, less ROS accumulation, and markedly increased total flavonoids and anthocyanin levels, when exposed to 3-AT treatment (Figures 8E–I).
In conclusion, our results provided some evidence that a Pn2-ODD1 gene from P. nutans increased the accumulation of anthocyanin and flavonol in transgenic plants. Also, heterologous expression of Pn2-ODD1 conferred the plant resistance to salinity, drought, and UV-B stress, which may play a key role in the adaptation of P. nutans to the polar environment.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author.
Author Contributions
PZ and SL designed and supervised the experiments. HW conducted the experiments and wrote the manuscript. FF and QY assisted in phenotype analysis. HW and PZ analyzed and discussed the results. All authors agreed to publish the manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (41976225), Key Technology Research and Development Program of Shandong Province (2019GSF107064), Scientific Fund for National Public Research Institutes of China (GY0219Q05), and Central Government Guide Local Science and Technology Development Funds (YDZX20203700002579).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher's Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2022.850062/full#supplementary-material
Supplementary Figure 1. Gene transformation process of Physcomitrella patens and Arabidopsis. (A) Undigested P. patens protonema. (B, C) Digested P. patens protoplasts. (D, E) Screening of transgenic Pn2-ODD1 P. patens and Arabidopsis by antibiotic. (F, G) Identification of Pn2-ODD1 in P. patens and Arabidopsis by PCR analysis.
Supplementary Table 1. All primers used for vector construction and gene expression analysis.
Supplementary Table 2. Differential metabolites (36 up-regulate and 12 down-regulate) identified by the flavonoids metabolomics in 5-day-old WT and transgenic Arabidopsis.
References
Abdelhafez, O. H., Othman, E. M., Fahim, J. R., Desoukey, S. Y., Pimentel-Elardo, S. M., Nodwell, J. R., et al. (2020). Metabolomics analysis and biological investigation of three Malvaceae plants. Phytochem. Anal. 31, 204–214. doi: 10.1002/pca.2883
Ahn, J. H., Kim, J. S., Kim, S., Soh, H. Y., Shin, H., Jang, H., et al. (2015). De novo transcriptome analysis to identify anthocyanin biosynthesis genes responsible for tissue-specific pigmentation in Zoysiagrass (Zoysia japonica Steud.). PLoS ONE 10, e0124497. doi: 10.1371/journal.pone.0124497
Alavilli, H., Lee, H., Park, M., and Lee, B. H. (2017). Antarctic moss multiprotein bridging factor 1c overexpression in Arabidopsis resulted in enhanced tolerance to salt stress. Front. Plant Sci. 8, 1206. doi: 10.3389/fpls.2017.01206
Albert, N. W., Thrimawithana, A. H., McGhie, T. K., Clayton, W. A., Deroles, S. C., Schwinn, K. E., et al. (2018). Genetic analysis of the liverwort Marchantia polymorpha reveals that R2R3MYB activation of flavonoid production in response to abiotic stress is an ancient character in land plants. New Phytol. 218, 554–566. doi: 10.1111/nph.15002
Aleksza, D., Horvath, G. V., Sandor, G., and Szabados, L. (2017). Proline accumulation is regulated by transcription factors associated with phosphate starvation. Plant Physiol. 175, 555–567. doi: 10.1104/pp.17.00791
Arif, Y., Singh, P., Siddiqui, H., Bajguz, A., and Hayat, S. (2020). Salinity induced physiological and biochemical changes in plants: an omic approach towards salt stress tolerance. Plant Physiol. Biochem. 156, 64–77. doi: 10.1016/j.plaphy.2020.08.042
Asakawa, Y., Ludwiczuk, A., and Nagashima, F. (2013). Phytochemical and biological studies of bryophytes. Phytochemistry 91, 52–80. doi: 10.1016/j.phytochem.2012.04.012
Berland, H., Albert, N. W., Stavland, A., Jordheim, M., McGhie, T. K., Zhou, Y., et al. (2019). Auronidins are a previously unreported class of flavonoid pigments that challenges when anthocyanin biosynthesis evolved in plants. Proc. Natl. Acad. Sci. U. S. A. 116, 20232–20239. doi: 10.1073/pnas.1912741116
Buer, C. S., Imin, N., and Djordjevic, M. A. (2010). Flavonoids: new roles for old molecules. J. Integr. Plant Biol. 52, 98–111. doi: 10.1111/j.1744-7909.2010.00905.x
Busche, M., Acatay, C., Martens, S., Weisshaar, B., and Stracke, R. (2021). Functional characterisation of Banana (Musa spp.) 2-oxoglutaratedependent dioxygenases involved in flavonoid biosynthesis. Front. Plant Sci. 12, 701780. doi: 10.3389/fpls.2021.701780
Cheng, A. X., Zhang, X., Han, X. J., Zhang, Y. Y., Gao, S., Liu, C. J., et al. (2018). Identification of chalcone isomerase in the basal land plants reveals an ancient evolution of enzymatic cyclization activity for synthesis of flavonoids. New Phytol. 217, 909–924. doi: 10.1111/nph.14852
Choudhury, S., Panda, P., Sahoo, L., and Panda, S. K. (2013). Reactive oxygen species signaling in plants under abiotic stress. Plant Signal. Behav. 8, e23681. doi: 10.4161/psb.23681
Convey, P., Chown, S. L., Clarke, A., Barnes, D. K. A., Bokhorst, S., Cummings, V., et al. (2014). The spatial structure of Antarctic biodiversity. Ecol. Monogr. 84, 203–244. doi: 10.1890/12-2216.1
Cove, D. J., Perroud, P. F., Charron, A. J., McDaniel, S. F., Khandelwal, A., and Quatrano, R. S. (2009). Transformation of the moss Physcomitrella patens using direct DNA uptake by protoplasts. Cold Spring Harb. Protoc. 2009, pdb prot5143. doi: 10.1101/pdb.prot5143
Cruz de Carvalho, M. H.. (2008). Drought stress and reactive oxygen species: production, scavenging and signaling. Plant Signal. Behav. 3, 156–165. doi: 10.4161/psb.3.3.5536
Farrow, S. C., and Facchini, P. J. (2014). Functional diversity of 2-oxoglutarate/Fe(II)-dependent dioxygenases in plant metabolism. Front. Plant Sci. 5, 524. doi: 10.3389/fpls.2014.00524
Frank, W., Ratnadewi, D., and Reski, R. (2005). Physcomitrella patens is highly tolerant against drought, salt and osmotic stress. Planta 220, 384–394. doi: 10.1007/s00425-004-1351-1
Gao, S., Yu, H. N., Xu, R. X., Cheng, A. X., and Lou, H. X. (2015). Cloning and functional characterization of a 4-coumarate CoA ligase from liverwort Plagiochasma appendiculatum. Phytochemistry 111, 48–58. doi: 10.1016/j.phytochem.2014.12.017
Gilmour, S. J., Artus, N. N., and Thomashow, M. F. (1992). cDNA sequence analysis and expression of two cold-regulated genes of Arabidopsis thaliana. Plant Mol Biol. 18, 13–21. doi: 10.1007/BF00018452
Gupta, B., and Huang, B. (2014). Mechanism of salinity tolerance in plants: physiological, biochemical, and molecular characterization. Int. J. Genom. 2014, 1–18. doi: 10.1155/2014/701596
Hamamouch, N., Winkel, B. S. J., Li, C., and Davis, E. L. (2020). Modulation of Arabidopsis flavonol biosynthesis genes by cyst and root-knot nematodes. Plants 9, 253. doi: 10.3390/plants9020253
Han, X. J., Wu, Y. F., Gao, S., Yu, H. N., Xu, R. X., Lou, H. X., et al. (2014). Functional characterization of a Plagiochasma appendiculatum flavone synthase I showing flavanone 2-hydroxylase activity. FEBS Lett. 588, 2307–2314. doi: 10.1016/j.febslet.2014.05.023
Han, Y., Huang, K., Liu, Y., Jiao, T., Ma, G., Qian, Y., et al. (2017). Functional analysis of two flavanone-3-hydroxylase genes from Camellia sinensis: a critical role in flavonoid accumulation. Genes 8, 300. doi: 10.3390/genes8110300
He, J., Li, P., Huo, H., Liu, L., Tang, T., and He, M. (2019). Heterologous expression of HpBHY and CrBKT increases heat tolerance in Physcomitrella patens. Plant Divers. 41, 266–274. doi: 10.1016/j.pld.2019.04.001
He, L., Wu, Y. H., Zhao, Q., Wang, B., Liu, Q. L., and Zhang, L. (2018). Chrysanthemum DgWRKY2 gene enhances tolerance to salt stress in transgenic Chrysanthemum. Int. J. Mol. Sci. 19, 2062. doi: 10.3390/ijms19072062
Holton, T. A., Brugliera, F., and Tanaka, Y. (1993). Cloning and expression of flavonol synthase from Petunia hybrida. Plant J. 4, 1003–1010. doi: 10.1046/j.1365-313x.1993.04061003.x
Ishitani, M., Liu, J., Halfter, U., Kim, C. S., Shi, W., and Zhu, J. K. (2000). SOS3 function in plant salt tolerance requires N-myristoylation and calcium binding. Plant Cell 12, 1667–1677. doi: 10.1105/tpc.12.9.1667
Jiang, N., Doseff, A. I., and Grotewold, E. (2016). Flavones: from biosynthesis to health benefits. Plants 5, 27. doi: 10.3390/plants5020027
Jiang, X., Shi, Y., Fu, Z., Li, W. W., Lai, S., Wu, Y., et al. (2020). Functional characterization of three flavonol synthase genes from Camellia sinensis: roles in flavonol accumulation. Plant Sci. 300, 110632. doi: 10.1016/j.plantsci.2020.110632
Kawai, Y., Ono, E., and Mizutani, M. (2014). Evolution and diversity of the 2-oxoglutarate-dependent dioxygenase superfamily in plants. Plant J. 78, 328–343. doi: 10.1111/tpj.12479
Koes, R. E. Q. F, and Mol, J.N.M. (1994). The flavonoid biosynthetic pathway in plants: function and evolution. BioEssays 16, 123–132. doi: 10.1002/bies.950160209
Kunz, S., Burkhardt, G., and Becker, H. (1993). Riccionidins a and b, anthocyanidins from the cell walls of the liverwort Ricciocarpos natans. Phytochemistry 35, 233–235. doi: 10.1016/S0031-9422(00)90540-5
Li, C., Liu, S., Yao, X., Wang, J., Wang, T., Zhang, Z., et al. (2017). PnF3H, a flavanone 3-hydroxylase from the Antarctic moss Pohlia nutans, confers tolerance to salt stress and ABA treatment in transgenic Arabidopsis. Plant Growth Regul. 83, 489–500. doi: 10.1007/s10725-017-0314-z
Li, C., Liu, S., Zhang, W., Chen, K., and Zhang, P. (2019). Transcriptional profiling and physiological analysis reveal the critical roles of ROS-scavenging system in the Antarctic moss Pohlia nutans under ultraviolet-B radiation. Plant Physiol. Biochem. 134, 113–122. doi: 10.1016/j.plaphy.2018.10.034
Li, D. D., Ni, R., Wang, P. P., Zhang, X. S., Wang, P. Y., Zhu, T. T., et al. (2020a). Molecular basis for chemical evolution of flavones to flavonols and anthocyanins in land plants. Plant Physiol. 184, 1731–1743. doi: 10.1104/pp.20.01185
Li, H., Li, D., Yang, Z., Zeng, Q., Luo, Y., and He, N. (2020b). Flavones produced by Mulberry flavone synthase type I constitute a defense line against the ultraviolet-B stress. Plants 9, 215. doi: 10.3390/plants9020215
Li, J., Zhao, A. Z., Yu, M. D., Li, Y. F., Liu, X. Q., and Chen, X. Y. (2018). Function analysis of anthocyanidin synthase from Morus alba L. by expression in bacteria and tobacco. Electron. J. Biotechn. 36, 9–14. doi: 10.1016/j.ejbt.2018.09.001
Li, N., Wang, X., Ma, B., Wu, Z., Zheng, L., Qi, Z., et al. (2021). A leucoanthocyanidin dioxygenase gene (RtLDOX2) from the feral forage plant Reaumuria trigyna promotes the accumulation of flavonoids and improves tolerance to abiotic stresses. J. Plant Res. 134, 1121–1138. doi: 10.1007/s10265-021-01315-2
Li, P., Li, Y. J., Zhang, F. J., Zhang, G. Z., Jiang, X. Y., Yu, H. M., et al. (2017). The Arabidopsis UDP-glycosyltransferases UGT79B2 and UGT79B3, contribute to cold, salt and drought stress tolerance via modulating anthocyanin accumulation. Plant J. 89, 85–103. doi: 10.1111/tpj.13324
Liu, H., Su, B., Zhang, H., Gong, J., Zhang, B., Liu, Y., et al. (2019). Identification and functional analysis of a flavonol synthase gene from Grape Hyacinth. Molecules 24, 1579. doi: 10.3390/molecules24081579
Liu, S., Fang, S., Liu, C., Zhao, L., Cong, B., and Zhang, Z. (2021). Transcriptomics integrated with metabolomics reveal the effects of ultraviolet-B radiation on flavonoid biosynthesis in Antarctic moss. Front. Plant Sci. 12, 788377. doi: 10.3389/fpls.2021.788377
Liu, Y., Shi, Z., Maximova, S., Payne, M. J., and Guiltinan, M. J. (2013). Proanthocyanidin synthesis in Theobroma cacao: genes encoding anthocyanidin synthase, anthocyanidin reductase, and leucoanthocyanidin reductase. BMC Plant Biol. 13, 202. doi: 10.1186/1471-2229-13-202
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Ludwiczuk, A., and Asakawa, Y. (2019). Bryophytes as a source of bioactive volatile terpenoids – a review. Food Chem. Toxicol. 132, 110649. doi: 10.1016/j.fct.2019.110649
Lukačin, R., Wellmann, F., Britsch, L., Martens, S., and Matern, U. (2003). Flavonol synthase from Citrus unshiu is a bifunctional dioxygenase. Phytochemistry 62, 287–292. doi: 10.1016/s0031-9422(02)00567-8
Luo, P., Ning, G., Wang, Z., Shen, Y., Jin, H., Li, P., et al. (2015). Disequilibrium of flavonol synthase and dihydroflavonol-4-reductase expression associated tightly to White vs. Red color flower formation in plants. Front. Plant Sci. 6, 1257. doi: 10.3389/fpls.2015.01257
Ma, D., Sun, D., Wang, C., Li, Y., and Guo, T. (2014). Expression of flavonoid biosynthesis genes and accumulation of flavonoid in wheat leaves in response to drought stress. Plant Physiol Biochem. 80, 60–66. doi: 10.1016/j.plaphy.2014.03.024
Mahajan, M., and Yadav, S. K. (2014). Overexpression of a tea flavanone 3-hydroxylase gene confers tolerance to salt stress and Alternaria solani in transgenic tobacco. Plant Mol. Biol. 85, 551–573. doi: 10.1007/s11103-014-0203-z
Martens, S., Forkmann, G., Britsch, L., Wellmann, F., Matern, U., and Lukačin, R. (2003). Divergent evolution of flavonoid 2-oxoglutarate-dependent dioxygenases in parsley1. FEBS Lett. 544, 93–98. doi: 10.1016/s0014-5793(03)00479-4
Martens, S., Preuss, A., and Matern, U. (2010). Multifunctional flavonoid dioxygenases: flavonol and anthocyanin biosynthesis in Arabidopsis thaliana L. Phytochemistry 71, 1040–1049. doi: 10.1016/j.phytochem.2010.04.016
Mues, R.. (2000). “Chemical constituents and biochemistry,” in Bryophyte Biology, eds A. J. Shaw, and B. Goffinet (Cambridge: Cambridge University Press), 150–181.
Naing, A. H., Ai, T. N., Lim, K. B., Lee, I. J., and Kim, C. K. (2018). Overexpression of Rosea1 from Snapdragon enhances anthocyanin accumulation and abiotic stress tolerance in transgenic tobacco. Front. Plant Sci. 9, 1070. doi: 10.3389/fpls.2018.01070
Nakabayashi, R., Yonekura-Sakakibara, K., Urano, K., Suzuki, M., Yamada, Y., Nishizawa, T., et al. (2014). Enhancement of oxidative and drought tolerance in Arabidopsis by overaccumulation of antioxidant flavonoids. Plant J. 77, 367–379. doi: 10.1111/tpj.12388
Newsham, K. K.. (2003). UV-B radiation arising from stratospheric ozone depletion influences the pigmentation of the Antarctic moss Andreaea regularis. Oecologia 135, 327–331. doi: 10.1007/s00442-003-1191-x
Nielsen, K., Deroles, S. C., Markham, K. R., Bradley, M. J., Podivinsky, E., and Manson, D. (2002). Antisense flavonol synthase alters copigmentation and flower color in lisianthus. Mol. Breed. 9, 217–229. doi: 10.1023/A:1020320809654
Pang, Y., Peel, G. J., Wright, E., Wang, Z., and Dixon, R. A. (2007). Early steps in proanthocyanidin biosynthesis in the model legume Medicago truncatula. Plant Physiol. 145, 601–615. doi: 10.1104/pp.107.107326
Park, S., Kim, D. H., Park, B. R., Lee, J. Y., and Lim, S. H. (2019). Molecular and functional characterization of Oryza sativa flavonol synthase (OsFLS), a bifunctional dioxygenase. J. Agric. Food Chem. 67, 7399–7409. doi: 10.1021/acs.jafc.9b02142
Pervaiz, T., Songtao, J., Faghihi, F., Haider, M. S., and Fang, J. (2017). Naturally occurring anthocyanin, structure, functions and biosynthetic pathway in fruit plants. J. Plant Biochem. Physiol. 5, 187. doi: 10.4172/2329-9029.1000187
Piatkowski, B. T., Imwattana, K., Tripp, E. A., Weston, D. J., Healey, A., Schmutz, J., et al. (2020). Phylogenomics reveals convergent evolution of red-violet coloration in land plants and the origins of the anthocyanin biosynthetic pathway. Mol. Phylogenet. Evol. 151, 106904. doi: 10.1016/j.ympev.2020.106904
Pizarro, M., Contreras, R. A., Kohler, H., and Zuniga, G. E. (2019). Desiccation tolerance in the Antarctic moss Sanionia uncinata. Biol. Res. 52, 46. doi: 10.1186/s40659-019-0251-6
Porra, R. J., Thompson, W. A., and Kriedemann, P. E. (1989). Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim. Biophys. Acta (BBA) –Bioenerg. 975, 384–394. doi: 10.1016/s0005-2728(89)80347-0
Rafique, M. Z., Carvalho, E., Stracke, R., Palmieri, L., Herrera, L., Feller, A., et al. (2016). Nonsense mutation inside anthocyanidin synthase gene controls pigmentation in yellow raspberry (Rubus idaeus L.). Front. Plant Sci. 7, 1892. doi: 10.3389/fpls.2016.01892
Reddy, A. M., Reddy, V. S., Scheffler, B. E., Wienand, U., and Reddy, A. R. (2007). Novel transgenic rice overexpressing anthocyanidin synthase accumulates a mixture of flavonoids leading to an increased antioxidant potential. Metab. Eng. 9, 95–111.
Rensing, S. A., Goffinet, B., Meyberg, R., Wu, S. Z., and Bezanilla, M. (2020). The moss Physcomitrium (Physcomitrella) patens: a model organism for non-seed plants. Plant Cell. 32, 1361–1376. doi: 10.1105/tpc.19.00828
Righini, S., Rodriguez, E. J., Berosich, C., Grotewold, E., Casati, P., and Falcone Ferreyra, M. L. (2019). Apigenin produced by maize flavone synthase I and II protects plants against UV-B-induced damage. Plant Cell Environ. 42, 495–508. doi: 10.1111/pce.13428
Sharma, P., Jha, A. B., Dubey, R. S., and Pessarakli, M. (2012). Reactive oxygen species, oxidative damage, and antioxidative defense mechanism in plants under stressful conditions. J. Bot. 2012, 1–26. doi: 10.1155/2012/217037
Singh, J., Dubey, A. K., and Singh, R. P. (2010). Antarctic terrestrial ecosystem and role of pigments in enhanced UV-B radiations. Rev. Environ. Sci. Biotechnol. 10, 63–77. doi: 10.1007/s11157-010-9226-3
Solfanelli, C., Poggi, A., Loreti, E., Alpi, A., and Perata, P. (2006). Sucrose-specific induction of the anthocyanin biosynthetic pathway in Arabidopsis. Plant Physiol. 140, 637–646. doi: 10.1104/pp.105.072579
Song, X., Diao, J., Ji, J., Wang, G., Guan, C., Jin, C., et al. (2016). Molecular cloning and identification of a flavanone 3-hydroxylase gene from Lycium chinense, and its overexpression enhances drought stress in tobacco. Plant Physiol. Biochem. 98, 89–100. doi: 10.1016/j.plaphy.2015.11.011
Stracke, R., De Vos, R. C., Bartelniewoehner, L., Ishihara, H., Sagasser, M., Martens, S., et al. (2009). Metabolomic and genetic analyses of flavonol synthesis in Arabidopsis thaliana support the in vivo involvement of leucoanthocyanidin dioxygenase. Planta 229, 427–445. doi: 10.1007/s00425-008-0841-y
Tanaka, Y., Sasaki, N., and Ohmiya, A. (2008). Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids. Plant J. 54, 733–749. doi: 10.1111/j.1365-313X.2008.03447.x
Teng, S., Keurentjes, J., Bentsink, L., Koornneef, M., and Smeekens, S. (2005). Sucrose-specific induction of anthocyanin biosynthesis in Arabidopsis requires the MYB75/PAP1 gene. Plant Physiol. 139, 1840–1852. doi: 10.1104/pp.105.066688
Turnbull, J. J., Nakajima, J., Welford, R. W., Yamazaki, M., Saito, K., and Schofield, C. J. (2004). Mechanistic studies on three 2-oxoglutarate-dependent oxygenases of flavonoid biosynthesis. J. Biol. Chem. 279, 1206–1216. doi: 10.1074/jbc.M309228200
Turnbull, J. J., Sobey, W. J., Aplin, R. T., Hassan, A., Schofield, C. J., Firmin, J. L., et al. (2000). Are anthocyanidins the immediate products of anthocyanidin synthase? Chem. Commun. 24, 2473–2474. doi: 10.1039/b007594i
Wang, H., Liu, S., Wang, T., Liu, H., Xu, X., Chen, K., et al. (2020). The moss flavone synthase I positively regulates the tolerance of plants to drought stress and UV-B radiation. Plant Sci. 298, 110591. doi: 10.1016/j.plantsci.2020.110591
Wang, M., Qin, L., Xie, C., Li, W., Yuan, J., Kong, L., et al. (2014a). Induced and constitutive DNA methylation in a salinity-tolerant wheat introgression line. Plant Cell Physiol. 55, 1354–1365. doi: 10.1093/pcp/pcu059
Wang, M., Ren, T., Huang, R., Li, Y., Zhang, C., and Xu, Z. (2021a). Overexpression of an Apocynum venetum flavonols synthetase gene confers salinity stress tolerance to transgenic tobacco plants. Plant Physiol. Biochem. 162, 667–676. doi: 10.1016/j.plaphy.2021.03.034
Wang, M., Zhang, Y., Zhu, C., Yao, X., Zheng, Z., Tian, Z., et al. (2021b). EkFLS overexpression promotes flavonoid accumulation and abiotic stress tolerance in plant. Physiol. Plant. 172, 1966–1982. doi: 10.1111/ppl.13407
Wang, S. H., Lim, J. H., Kim, S. S., Cho, S. H., Yoo, S. C., Koh, H. J., et al. (2015). Mutation of SPOTTED LEAF3 (SPL3) impairs abscisic acid-responsive signalling and delays leaf senescence in rice. J. Exp. Bot. 66, 7045–7059. doi: 10.1093/jxb/erv401
Wang, X., Liu, Z., and He, Y. (2014b). Responses and tolerance to salt stress in bryophytes. Plant Signal. Behav. 3, 516–518. doi: 10.4161/psb.3.8.6337
Wang, Y., Shi, Y., Li, K., Yang, D., Liu, N., Zhang, L., et al. (2021c). Roles of the 2-oxoglutarate-dependent dioxygenase superfamily in the flavonoid pathway: a review of the functional diversity of F3H, FNSI, FLS, and LDOX/ANS. Molecules 26, 6745. doi: 10.3390/molecules26216745
Wang, Z., Wang, S., Wu, M., Li, Z., Liu, P., Li, F., et al. (2019). Evolutionary and functional analyses of the 2-oxoglutarate-dependent dioxygenase genes involved in the flavonoid biosynthesis pathway in tobacco. Planta 249, 543–561. doi: 10.1007/s00425-018-3019-2
Waterman, M. J., Bramley-Alves, J., Miller, R. E., Keller, P. A., and Robinson, S. A. (2018). Photoprotection enhanced by red cell wall pigments in three East Antarctic mosses. Biol. Res. 51, 49. doi: 10.1186/s40659-018-0196-1
Waterman, M. J., Nugraha, A. S., Hendra, R., Ball, G. E., Robinson, S. A., and Keller, P. A. (2017). Antarctic moss biflavonoids show high antioxidant and ultraviolet-screening activity. J. Nat. Prod. 80, 2224–2231. doi: 10.1021/acs.jnatprod.7b00085
Wei, S., Zhang, W., Fu, R., and Zhang, Y. (2021). Genome-wide characterization of 2-oxoglutarate and Fe(II)-dependent dioxygenase family genes in tomato during growth cycle and their roles in metabolism. BMC Genom. 22, 126. doi: 10.1186/s12864-021-07434-3
Wellman, C. H., Osterloff, P. L., and Mohiuddin, U. (2003). Fragments of the earliest land plants. Nature 425, 282–285. doi: 10.1038/nature01884
Wellmann, F., Griesser, M., Schwab, W., Martens, S., Eisenreich, W., Matern, U., et al. (2006). Anthocyanidin synthase from Gerbera hybrida catalyzes the conversion of (+)-catechin to cyanidin and a novel procyanidin. FEBS Lett. 580, 1642–1648. doi: 10.1016/j.febslet.2006.02.004
Wolf, L., Rizzini, L., Stracke, R., Ulm, R., and Rensing, S. A. (2010). The molecular and physiological responses of Physcomitrella patens to ultraviolet-B radiation. Plant Physiol. 153, 1123–1134. doi: 10.1104/pp.110.154658
Xu, Z., Mahmood, K., and Rothstein, S. J. (2017). ROS induces anthocyanin production via late biosynthetic genes and anthocyanin deficiency confers the hypersensitivity to ROS-generating stresses in Arabidopsis. Plant Cell Physiol. 58, 1364–1377. doi: 10.1093/pcp/pcx073
Yonekura-Sakakibara, K., Fukushima, A., Nakabayashi, R., Hanada, K., Matsuda, F., Sugawara, S., et al. (2012). Two glycosyltransferases involved in anthocyanin modification delineated by transcriptome independent component analysis in Arabidopsis thaliana. Plant J. 69, 154–167. doi: 10.1111/j.1365-313X.2011.04779.x
Yoon, J., Cho, L. H., Tun, W., Jeon, J. S., and An, G. (2021). Sucrose signaling in higher plants. Plant Sci. 302, 110703. doi: 10.1016/j.plantsci.2020.110703
Yu, H. N., Wang, L., Sun, B., Gao, S., Cheng, A. X., and Lou, H. X. (2015). Functional characterization of a chalcone synthase from the liverwort Plagiochasma appendiculatum. Plant Cell Rep. 34, 233–245. doi: 10.1007/s00299-014-1702-8
Zhang, H., Du, C., Wang, Y., Wang, J., Zheng, L., and Wang, Y. (2016). The Reaumuria trigyna leucoanthocyanidin dioxygenase (RtLDOX) gene complements anthocyanidin synthesis and increases the salt tolerance potential of a transgenic Arabidopsis LDOX mutant. Plant Physiol. Biochem. 106, 278–287. doi: 10.1016/j.plaphy.2016.05.005
Keywords: abiotic stress, Antarctic moss, anthocyanin accumulation, 2-oxoglutarate/Fe(II)-dependent dioxygenases (2-ODDs), flavonoids, flavonol
Citation: Wang H, Liu S, Fan F, Yu Q and Zhang P (2022) A Moss 2-Oxoglutarate/Fe(II)-Dependent Dioxygenases (2-ODD) Gene of Flavonoids Biosynthesis Positively Regulates Plants Abiotic Stress Tolerance. Front. Plant Sci. 13:850062. doi: 10.3389/fpls.2022.850062
Received: 07 January 2022; Accepted: 21 June 2022;
Published: 29 July 2022.
Edited by:
Paula Casati, Centro de Estudios Fotosintéticos y Bioquímicos (CEFOBI), ArgentinaReviewed by:
Ashutosh Pandey, National Institute of Plant Genome Research (NIPGR), IndiaJaspreet Kaur Sembi, Panjab University, India
Copyright © 2022 Wang, Liu, Fan, Yu and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Pengying Zhang, emhhbmdweTgwQHNkdS5lZHUuY24=
 Huijuan Wang
Huijuan Wang Shenghao Liu
Shenghao Liu Fenghua Fan
Fenghua Fan Qian Yu
Qian Yu Pengying Zhang
Pengying Zhang