- 1College of Agronomy and Biotechnology, China Agricultural University, Beijing, China
- 2Institute of Crop Sciences, Chinese Academy of Agricultural Sciences, Beijing, China
A corrigendum on
Transcriptome Analysis of Maize Immature Embryos Reveals the Roles of Cysteine in Improving Agrobacterium Infection Efficiency
by Liu, Y., Zhang, Z., Fu, J., Wang, G., Wang, J., and Liu, Y. (2017). Front. Plant Sci. 8:1778. doi: 10.3389/fpls.2017.01778
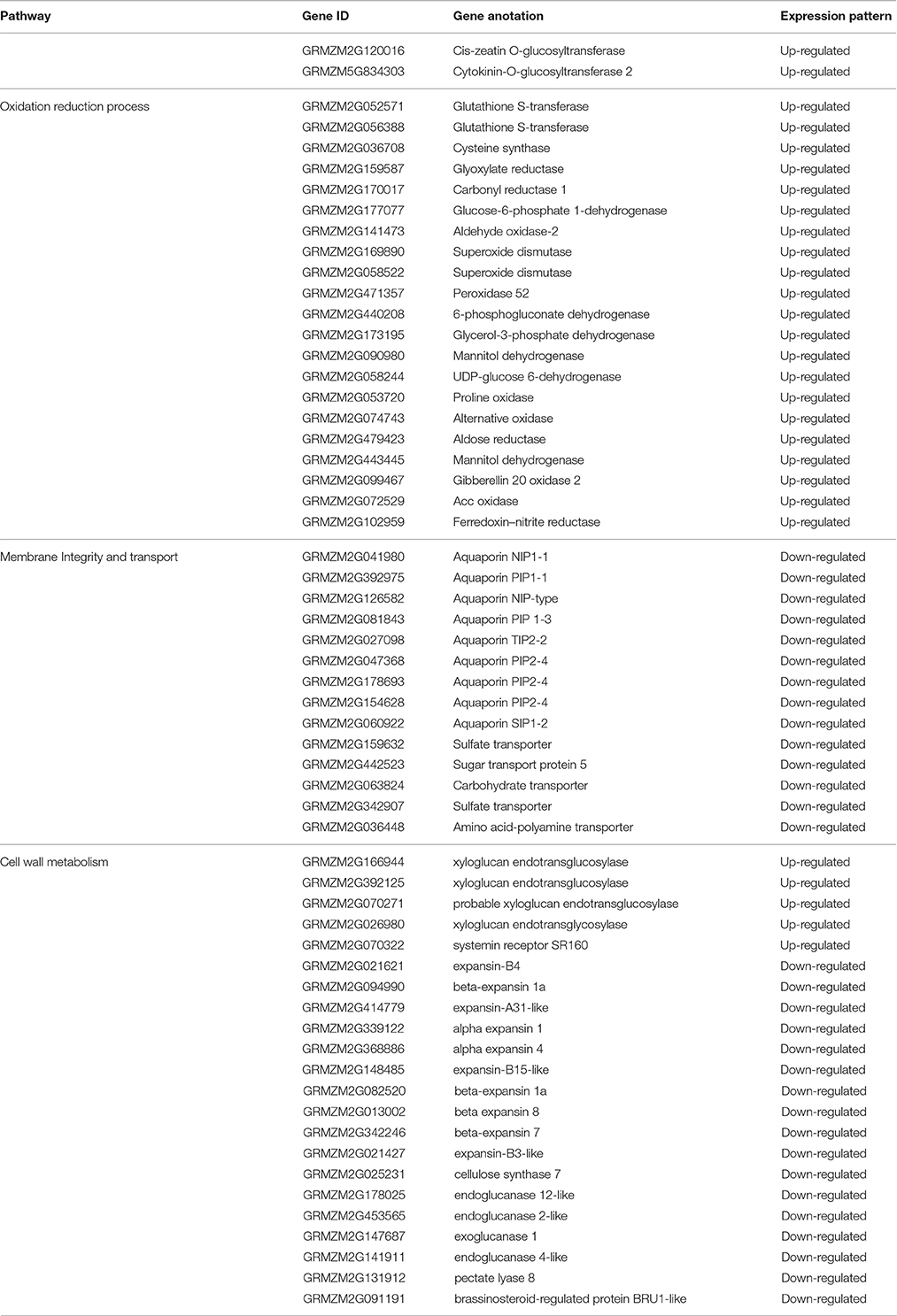
In the original article, there was a mistake in Table 3 as published. In original Table 3, four aquaporin genes (GRMZM2G041980, GRMZM2G392975, GRMZM2G126582, GRMZM2G081843) were repeated two times, and we neglected to include several upregulated glycosyltransferase genes (GRMZM2G120016, GRMZM5G834303). The corrected Table 3 appears below.
In the original article, there was an error. The reads from the HiII embryo samples were incorrectly presented as “7,432,161–15,904,122”. The correct reads from the HiII embryo samples should be “7,432,161–15,881,554” as shown in Table 1.
A correction has been made to (Results), (Maize Embryo Transcriptome Profiling), (Paragraph 1):
To investigate the mechanism of cysteine-related transformation efficiency improvement, we performed transcriptome analysis of the maize embryos cultured on the medium with or without cysteine. The experiment was performed with four independent biological replicates. We obtained 7,432,161–15,881,554 reads from the HiII embryo samples and more than 78.73% of the reads were mapped to the B73 reference genome. We obtained 20,119,176–24,789,278 reads from the inbred line Z31 embryo samples and 73.66% of the reads were mapped to the B73 reference genome (Table 1). Maize has ~ 30,000 genes and the sequencing depth we acquired was enough for subsequent analysis. The authors apologize for this error and state that this does not change the scientific conclusions of the article in any way.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Keywords: Agrobacterium, infection efficiency, cysteine, maize embryo, transcriptome
Citation: Liu Y, Zhang Z, Fu J, Wang G, Wang J and Liu Y (2017) Corrigendum: Transcriptome Analysis of Maize Immature Embryos Reveals the Roles of Cysteine in Improving Agrobacterium Infection Efficiency. Front. Plant Sci. 8:1964. doi: 10.3389/fpls.2017.01964
Received: 24 October 2017; Accepted: 31 October 2017;
Published: 07 November 2017.
Edited and reviewed by: Paulo Arruda, Universidade Estadual de Campinas, Brazil
Copyright © 2017 Liu, Zhang, Fu, Wang, Wang and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jianhua Wang, d2FuZ2poNjNAY2F1LmVkdS5jbg==
Yunjun Liu, bGl1eXVuanVuQGNhYXMuY24=
 Yan Liu1,2
Yan Liu1,2 Zhiqiang Zhang
Zhiqiang Zhang Yunjun Liu
Yunjun Liu