- Beijing Key Laboratory of Development and Quality Control of Ornamental Crops, Department of Ornamental Horticulture, China Agricultural University, Beijing, China
The TRICHOME BIREFRINGENCE-LIKE (TBL) family is an important gene family engaged in the O-acetylation of cell wall polysaccharides. There have been a few reports showing that TBL participated in the resistance against phytopathogens in Arabidopsis and rice. However, no relevant studies in rose (Rosa sp.) have been published. In this study, a genome-wide analysis of the TBL gene family in rose was presented, including their phylogenetic relationships, gene structure, chromosomal positioning, and collinearity analysis. The phylogenetic analysis revealed a total of 50 RcTBL genes in the rose genome, and they are unevenly distributed across all seven chromosomes. The occurrence of gene duplication events suggests that both the whole genome duplication and partial duplication may play a role in gene duplication of RcTBLs. The analysis of Ka/Ks showed that the replicated RcTBL genes underwent mainly purifying selection with limited functional differentiation. Gene expression analysis indicated that 12 RcTBLs were down-regulated upon the infection of Botrytis cinerea, the causal agent of the gray mold disease of rose. These RcTBLs may be a sort of candidate genes for regulating the response of rose to B. cinerea. Through virus-induced gene silencing, RcTBL16 was shown to be associated with susceptibility to gray mold in rose. Through this study, meaningful information for further studies on the function of the TBL protein family in rose is provided.
Introduction
The cell wall is particularly important in plant growth and development because it maintains the form of the plant cell, allows intercellular communication, responds to external environmental variables, and interacts with pathogenic microorganisms (Keegstra, 2010; Xin et al., 2010). The plant cell wall has a complicated and dynamic structure, which is mainly composed of polysaccharide polymer, protein and lignin. O-acetylation occurs extensively in the plant cell wall, most notably with hemicelluloses, pectins, and lignins. The replacements of O-acetyl group usually happen on various specific glycosyl residues of cell wall polysaccharides. In addition, cell wall polysaccharides can be either mono- or diacetylated, and the extent of O-acetylation depends on species, tissue type, and growth status of plants.
The biosynthetic pathway and function of O-acetylation of cell wall polysaccharides have not yet been fully understood. Modifications in O-acetylation levels are known to alter plant growth and development, as well as their defense against pathogens, and that this effect is most likely achieved by altering the cell wall structure. The REDUCED WALL ACETYLATION (RWA) protein, as well as the ALTERED XYLOGLUCAN9 protein and TRICHOME BIREFRINGENCE-LIKE (TBL) protein families, has been identified as being involved in the O-acetylation pathway of plant cell walls. RWA proteins could be acetyl donor transporters, transporting acetyl CoA into the Golgi apparatus (Manabe et al., 2011, 2013). AXY9 protein may be required for O-acetylation of cell wall polysaccharides by producing acetylation intermediates (Schultink et al., 2015).
Unlike the RWA and AXY9 proteins, a number of TBLs have been identified as polysaccharide acetyltransferases, catalyzing the O-acetylation of specific cell wall polymers including xylan (Urbanowicz et al., 2015; Zhong et al., 2017a; Zhong et al., 2018a,d), xyloglucan (Zhong et al., 2018c,e; Zhong et al., 2020), mannan (Zhong et al., 2018b), and pectin (Vogel et al., 2004b; Bischoff et al., 2010a; Stranne et al., 2018a; Chiniquy et al., 2019a). TBL proteins have conserved Asp-x-x-His (DxxH) motif and Gly-Asp-Ser (GDS) motif that is required for acetyltransferase activity (Bischoff et al., 2010b), since a mutation in either the GDS or DXXH motif could cause TBL proteins to lose their function completely (Zhong et al., 2017a; Zhong et al., 2018e). Studies of tbl mutants in Arabidopsis have demonstrated that dwarfism, stem weakness, and stunted growth of plants are associated with the lack of the TBL genes (Bischoff et al., 2010a; Xiong et al., 2013; Schultink et al., 2015), implying that TBL is critical for plant development. Besides, TBL proteins are related to abiotic stress in plants. Compared with wild-type Arabidopsis plant, the cold tolerance of esk1 increased significantly (Xin et al., 2010), while tbl10 showed enhanced drought resistance (Stranne et al., 2018a). Furthermore, TBL proteins have also been associated with plant defense against pathogens. The reduction of O-acetyl degree of pmr5 in Arabidopsis may lead to its enhanced resistance to powdery mildew (Vogel et al., 2004b; Chiniquy et al., 2019a). According to a recent research, simultaneous mutation of the OsTBL1 and OsTBL2 genes in rice leads to lower acetylation levels and higher vulnerability to leaf blight disease (Gao et al., 2017).
Rose (Rosa sp.) is the most important cut flower in the world, with 8,500 hectares of cut-flower rose cultivation worldwide, with an annual production of over 15 billion stems (Bendahmane et al., 2013), and sales of more than $11 billion (Zlesak, 2007). Gray mold caused by Botrytis cinerea is the most devastating disease mainly infecting the flower of rose and affects the production of cut rose all over the world (Gleason and Helland, 2003). The cell wall is the initial barrier that pathogens meet when penetrating the plant, and alterations in cell wall structure might affect the plant’s defense against these microbes. O-acetylation is one of the most important modifications of cell wall polymer. TBL proteins, as the main gene family involved in cell wall O-acetylation, may influence the resistance or susceptibility of plants to pathogen by varying the degree of cell wall acetylation. However, no research on the function of the TBL gene at the plant genome-wide level has been conducted thus far. We performed the first genome-wide analysis of the RcTBL family in rose in this study. Furthermore, a virus-induced gene silencing (VIGS) has confirmed that RcTBL16 was involved in B. cinerea susceptibility.
Materials and Methods
Characterization of Putative TBL Proteins in Rose
We downloaded the complete rose genome sequence and CDS sequence from the website https://lipm-browsers.toulouse.inra.fr/pub/RchiOBHm-V2/ to construct a local genome database. In order to identify non-redundant RcTBL genes in rose genome, the HMM profile of the PC-Esterase domain was obtained from Pfam (PF138391). Then, using this HMM profile as a query, by searching the rose genome, all sequences were confirmed to contain a PC-Esterase domain with an E value of <1e-3 in rose. The distribution of all RcTBL genes on chromosomes was mapped by mapchart 2.2 software.
Gene Structure and Phylogenetic Analyses
The gene structure map of RcTBL was completed using TBtools (Chen et al., 2018) by means of the rose genome annotation file and protein sequences. Multiple comparisons of RcTBL amino acids (aa) were performed using the ClustalW default parameters. A phylogenetic analysis of RcTBLs then was carried out in MEGA-6.0 software by the maximum likelihood (ML) method, with 1,000 bootstrap replications, JTT with Freqs (+F) model and 50% partial deletion. Other parameters were set by default. MEGA 6.0 was also applied to construct the unrooted ML trees of TBL proteins from Arabidopsis and rose, the parameter settings being consistent with the separate phylogenetic analysis of RcTBLs.
Collinearity Analyses
For the purpose of identifying the collinearity of RcTBLs, the genome sequence of rose was downloaded on a local server, and then we used a Multiple Collinearity Scan toolkit (Wang et al., 2012) to determine the microsyntenic relationships between RcTBL genes. Furthermore, collinearity scanning (e-value of <1e−10) was used to evaluate the microsynteny relationships.
Calculation of Ratios of Nonsynonymous (Ka) to Synonymous (Ks) Nucleotide Substitutions
We used TBtools (Chen et al., 2018) to calculate Ks and Ka nucleotide substitution rates. The Ka/Ks ratio of duplicated gene pairs was calculated to determine the selection pattern driving the evolution of RcTBL.
Expression of RcTBLs in Response to B. cinerea
RNA-Seq data from rose petals under B. cinerea infection were obtained from the National Center for Biotechnology Information database as accession number PRJNA414570 (Liu et al., 2018). The materials for RNA-seq are rose petal discs infected with B. cinerea at 30h post inoculation (hpi) and 48hpi, with three biological repeats for both infected and control treatments at each time point. Clean sequencing reads were mapped to the Rosa chinensis ‘Old Blush’ reference genome.2 We calculated the gene expression levels of RcTBLs by Reads per kb per million reads and performed a Log2 fold change-based differentially expressed gene analysis by DEseq2.
For validating the RNA-Seq outcomes, RT-qPCR was used to analyze the expression of five RcTBL genes. Total RNA was extracted from rose petals at 30hpi and 48hpi, respectively. As described previously (Wu et al., 2016), the hot borate method was used to extract total RNA. Then, first-strand cDNA was synthesized using HiScript II Q Select RT SuperMix (Vazyme) in a 20-μl reaction volume with 1μg of DNase-treated RNA. RT-qPCR reaction was run using SYBR Green Master Mix (Takara) and detection was achieved in a StepOnePlus real-time PCR system (Thermo Fisher Scientific). We used RcUBI2 as an internal control and conducted expression analysis using the delta–delta-Ct method of calculation. All primers used as RT-qPCR are listed in Supplementary Table S1.
VIGS and B. cinerea Inoculation Assays
In order to obtain the pTRV2-RcTBL constructs, a~200bp fragment from the coding region of RcTBLs was amplified with specific primer pairs and subsequently cloned into the pTRV2 vector (Liu et al., 2002). VIGS was established as previously described (Cao et al., 2019). Briefly, separated petals were obtained from the outermost whorls of cut roses in the second stage of flowering. A 15mm disc was then punched from the center of each petal. Agrobacterium tumefaciens consisting of pTRV1 and pTRV2 constructs were mixed in a 1:1 ratio and vacuumed infiltrated into petal discs. Petals were inoculated with B. cinerea on day 6 after TRV infection and photographed 60h later to obtain images with disease lesions, which were statistically analyzed by ImageJ. Each gene was silenced at least three times with 48 petals as a replicate. Student’s t test was performed to determine the significance of lesion size. After photographing, the petal samples were collected for further validating of silencing efficiency by RT-qPCR. The primers used to detect silencing efficiency of RcTBL16 or RcTBL35 are the same primers as those used to detect expression in response to B. cinerea, and listed in Supplementary Table S1.
Results
Identification of RcTBL Genes in Rose
As previously stated, the TBL protein family is distinguished by a conserved GDS signature and DXXH motif, as well as an N-terminal transmembrane domain in most of the cases (Bischoff et al., 2010a,b). A total of 61 candidate RcTBL proteins were obtained in rose by the Hidden Markov model (HMM) profile of PC-Esterase domain (PF13839) contained two conservative motifs of the TBL protein family. All candidate sequences less than 150 amino acids and without the complete conserved motifs were removed; finally, we obtained a total of 50 RcTBLs.
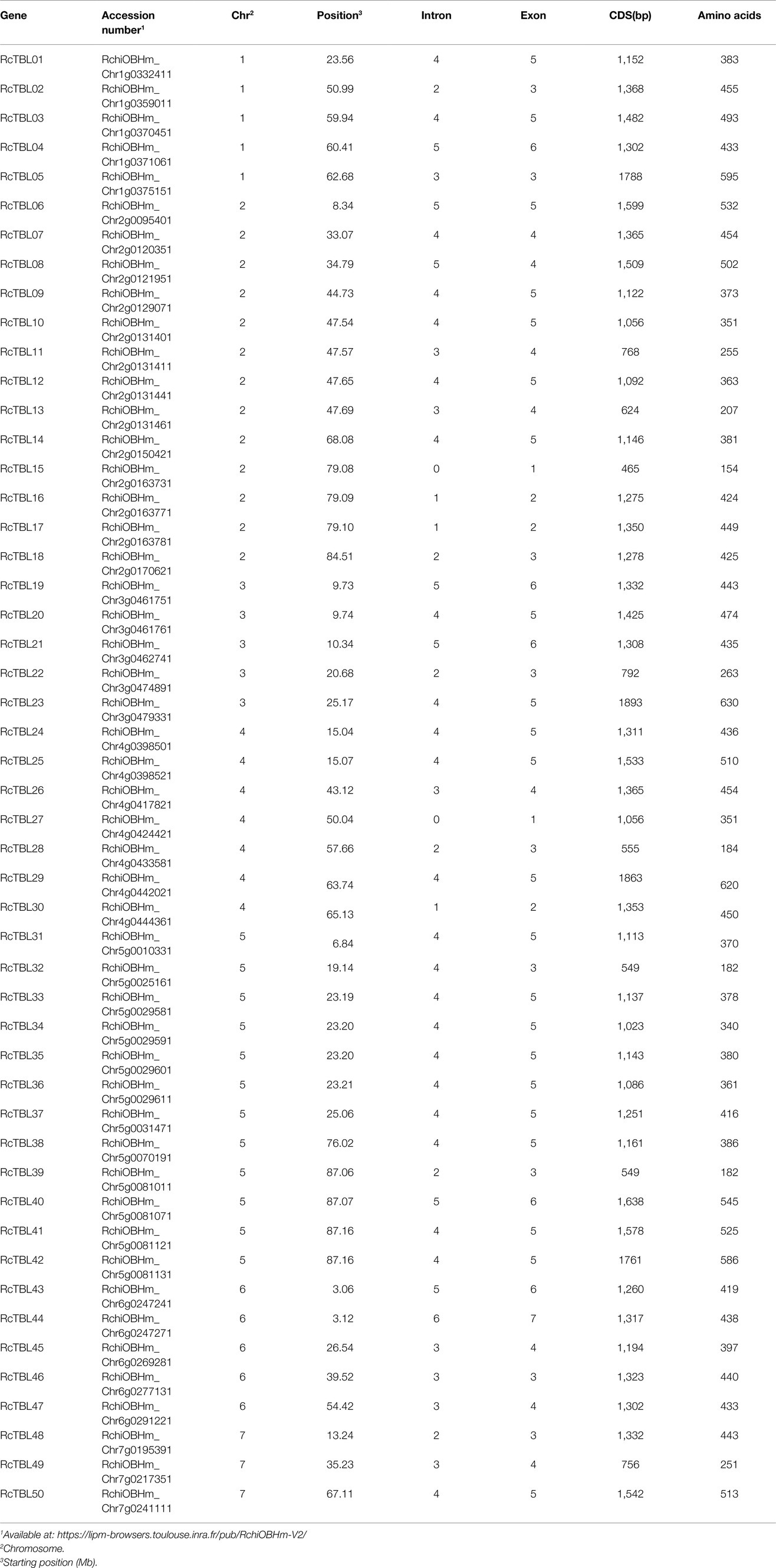
All RcTBLs can be mapped to rose chromosomes; we designated the genes RcTBL01 to RcTBL50 according to their chromosome order. The protein length of RcTBLs varies greatly. Of the 50 RcTBLs, RcTBL23 is the longest protein with 630 aa, while the shortest is RcTBL15 with 154 aa. The average length of proteins in RcTBL family is 409 aa. Details of the RcTBL genes, with gene number, chromosomal location, introns, exons, CDS and aa length are listed in Table 1.
Chromosomal Locations, Whole-Genome Duplication, and Microsynteny
All 50 RcTBL genes were distributed unevenly across seven rose chromosomes (Figure 1), with chromosome 2 having the highest density, gathering 13 RcTBL genes, followed by chromosome 5 with 12 RcTBL genes clustered on it, and chromosome 7 having the fewest RcTBL genes with only three members. We further investigated the duplication events in RcTBLs. Tandemly duplicated genes were defined as arrays of two or more homologous genes in the 100kb range. Three RcTBL gene pairs were discovered in the rose genome, each on a different chromosome, suggesting that segmental duplication occurs within these regions in rose. Collinearity analyses of the RcTBL genes on the chromosome are depicted in Figure 2.
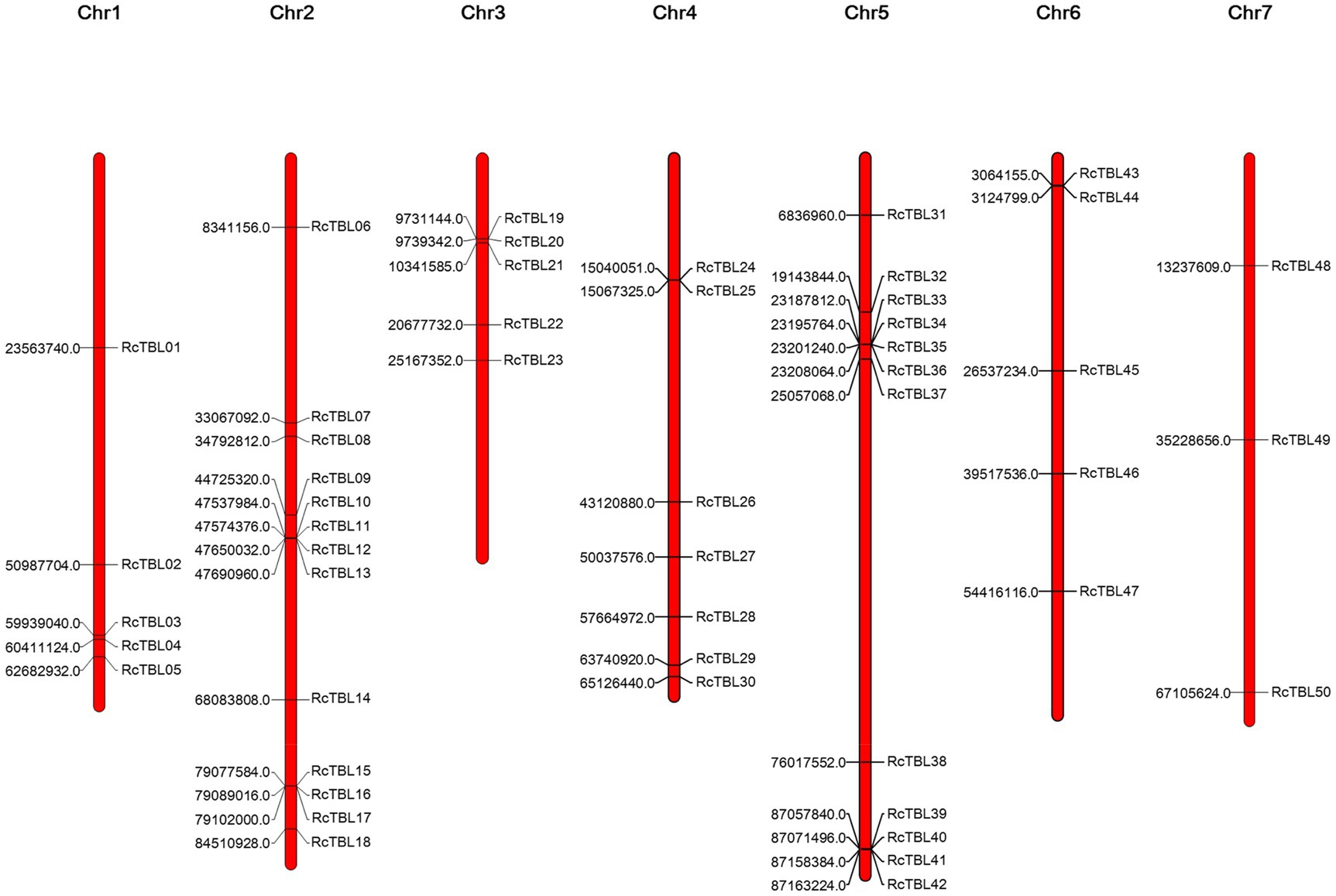
Figure 1. Chromosome localization of rose TBL family members. The distribution of RcTBLs is shown on the seven chromosomes of Rose chinensis. The starting position (bp) of each RcTBL is listed on the left side of the chromosomes.
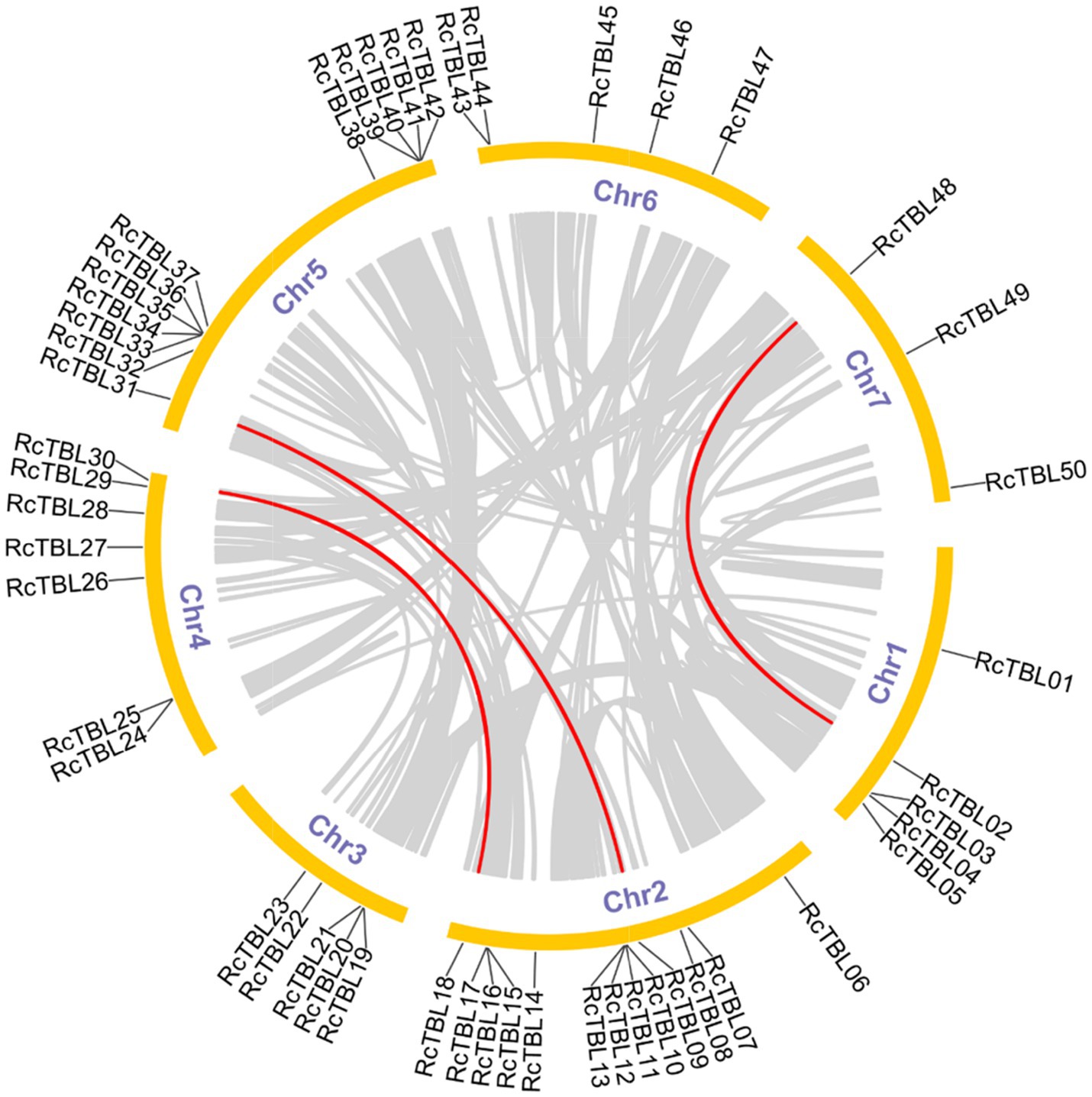
Figure 2. Microsyntenic analyses of the rose TBL protein family members in the R. chinensis genome. Circular visualization of rose TBL protein family members is mapped onto different chromosomes by using Circos. The red lines indicate rose TBL genes with a syntenic relationship. The gray lines represent all syntenic blocks in the genome of R. chinensis.
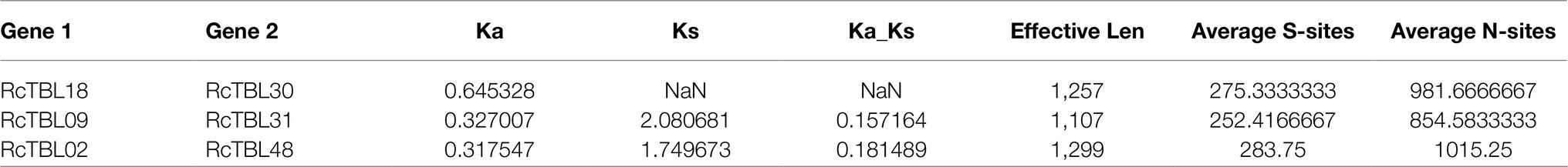
To explore the selective constraints among duplicated RcTBL genes, the ratio of nonsynonymous (Ka) to synonymous (Ks) nucleotide substitutions (Ka/Ks ratio) of three pairs of duplicated genes (Table 2) was calculated. Typically, Ka/Ks >1 is consistent with positive selection, while Ka/Ks <1 indicates purifying selection. Ka/Ks <1 for all three duplicated gene pairs (Table 2) suggested that the primary driver of gene evolution in the RcTBL family was purifying selection.
Phylogenetic and Gene Structural Analysis of Rose TBL Genes
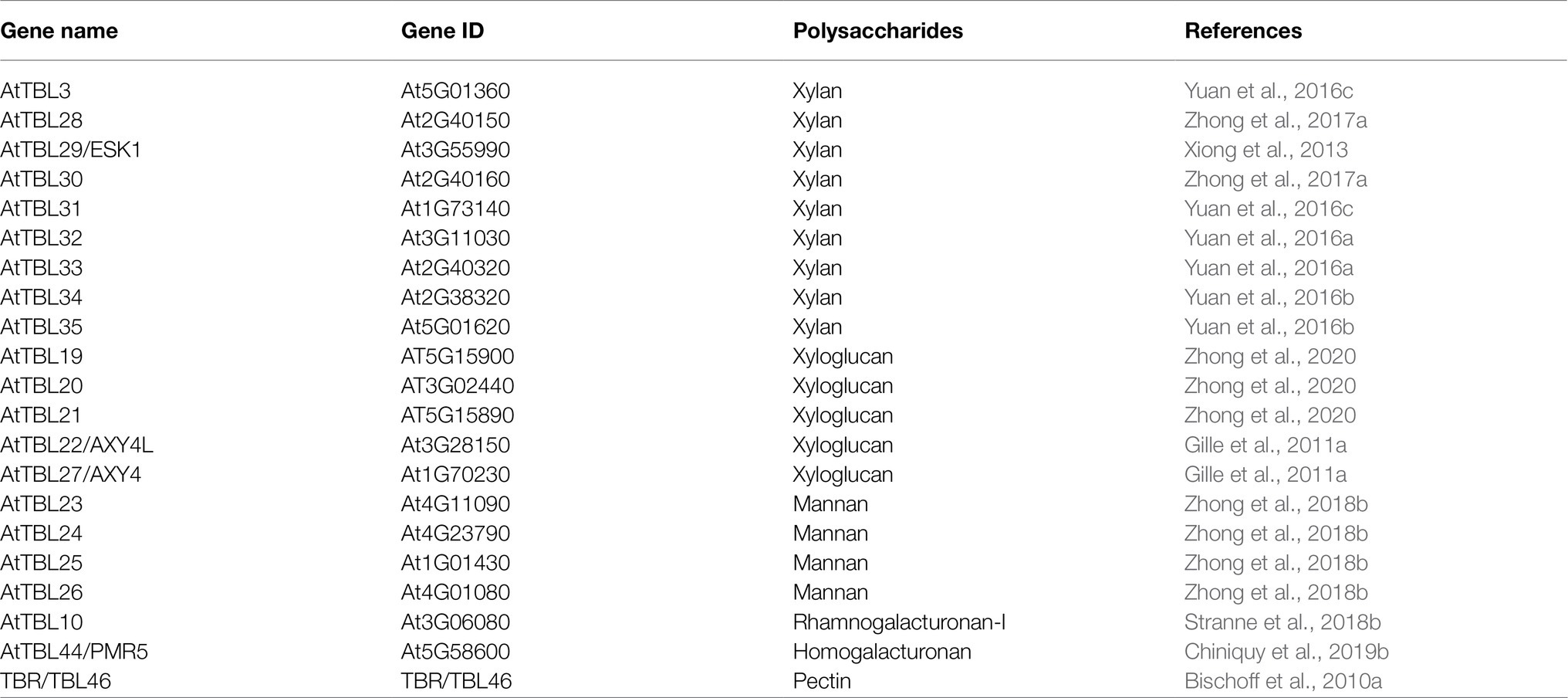
A total of 46 TBLs were identified on Arabidopsis thaliana and many of them have been established as O-acetyltransferases or potential O-acetyltransferase genes (Table 3). To evaluate the relationship between the TBL proteins of rose and A. thaliana, a compound phylogenetic tree was constructed using the full-length protein sequences of 46 Arabidopsis and 50 rose TBLs by the ML method. We found that most of the Arabidopsis TBL proteins had at least one rose homologue. AtTBL members identified as affecting the O-acetylation of xyloglucan, xylan, mannan, and pectin, respectively, were clustered in different branches, suggesting the correctness of our evolutionary tree (Figure 3).
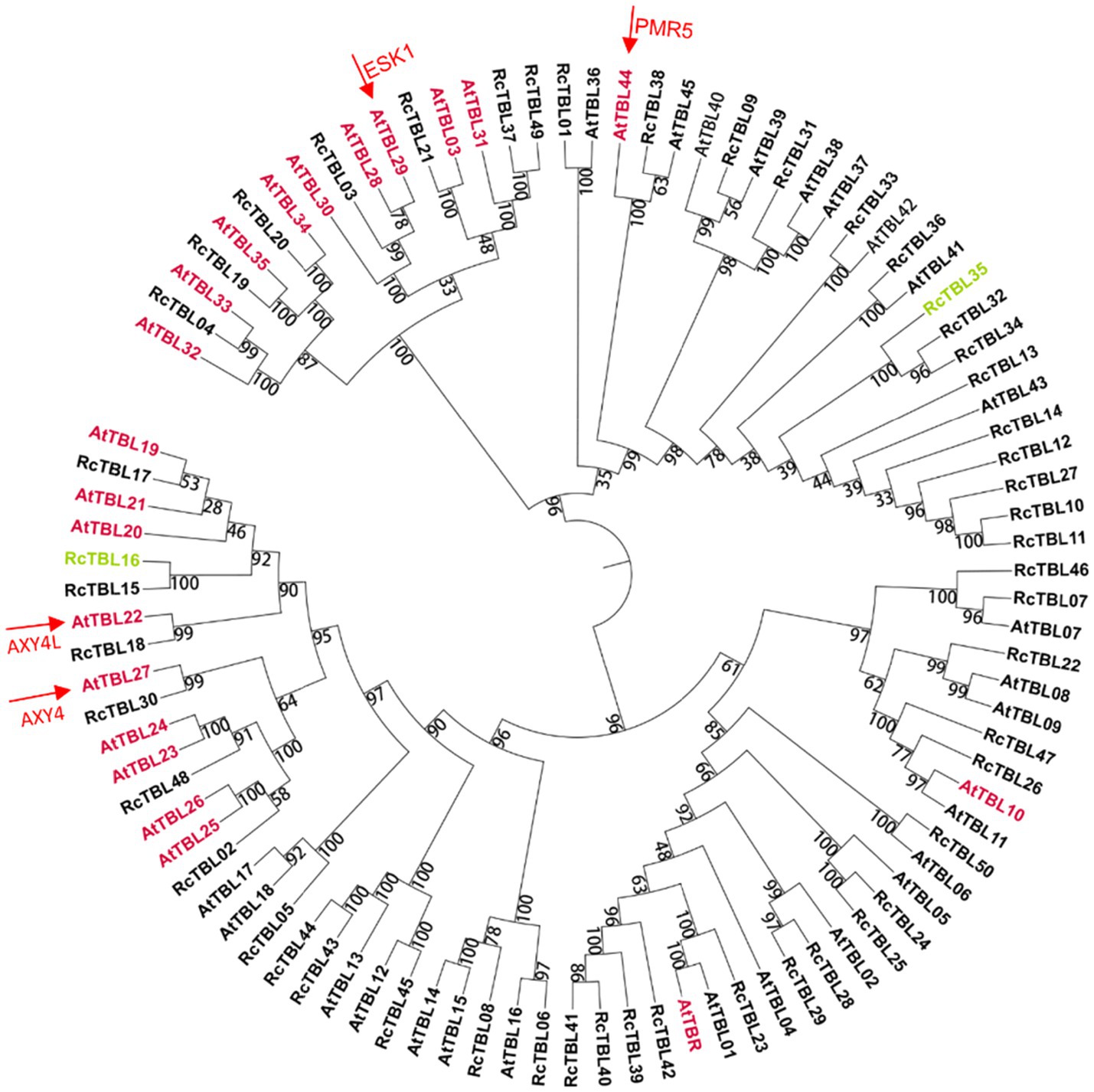
Figure 3. Phylogenetic analyses of TBL protein family in rose with Arabidopsis thaliana TBL protein family. Genes marked with pink have been shown to be involved in O-acetylation of cell wall polysaccharides in A. thaliana (Table 3); genes marked with green may be involved in the defense of roses against B. cinerea and were knocked down by virus-induced gene silencing (VIGS) in rose petals. Arrows refer to the well-known TBL proteins in Arabidopsis. The bootstrap values are indicated on the nodes of the branches.
Analysis of the exon–intron structure revealed that the intron structure of RcTBLs were highly variable, ranging from 1 to 6, with the largest number (23) of RcTBLs containing four introns. In addition, the length of RcTBL introns was extremely varied, ranging from tens to thousands of nucleotides. RcTBL35 contained the longest intron (4,191bp), while the shortest intron (69bp) was present on RcTBL36 (Figure 4).
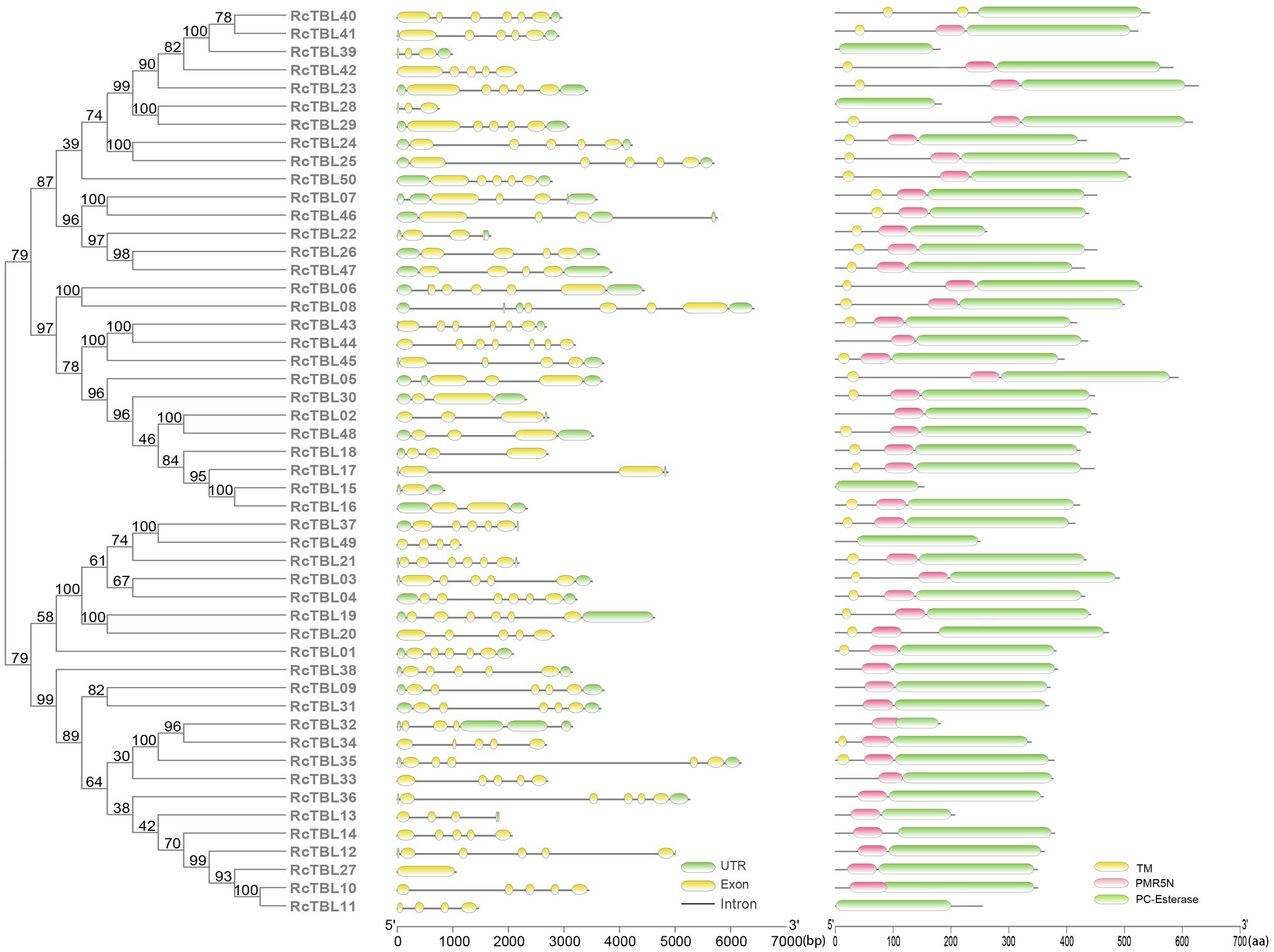
Figure 4. Phylogenetic analyses and gene structures of rose TBL proteins. Complete alignments of all rose TBL proteins were used to construct a phylogenetic tree using the maximum likelihood method. The bootstrap values are indicated on the nodes of the branches. Exon–intron structure of RcTBLs is shown in the middle part of the figure, and the green boxes, yellow boxes, and gray lines represent UTRs, exons, and introns, respectively. Transmembrane and conserved domain of RcTBLs are displayed in the right part of the figure, and the green boxes, yellow boxes, and pink boxes represent PC-Esterase domain, transmembrane domain, and PMR5N domain, respectively. The scale on the bottom is provided as a reference.
The protein sequences of RcTBLs were examined with Pfam and all 50 candidate genes had the PC-esterase domain. Surprisingly, 88% of RcTBLs possessed a cys-rich domain called the PMR5N domain in Pfam (PF14416) that preceded the PC-esterase domain, implying that it could be a crucial structural element of the TBL family. Furthermore, 64% of all 50 RcTBL candidates had at least one transmembrane domain, with RcTBL40 having two transmembrane domains, and 18 RCTBLs had no transmembrane domain (Figure 4).
The Expression of RcTBL Genes in Response to B. cinerea Infection
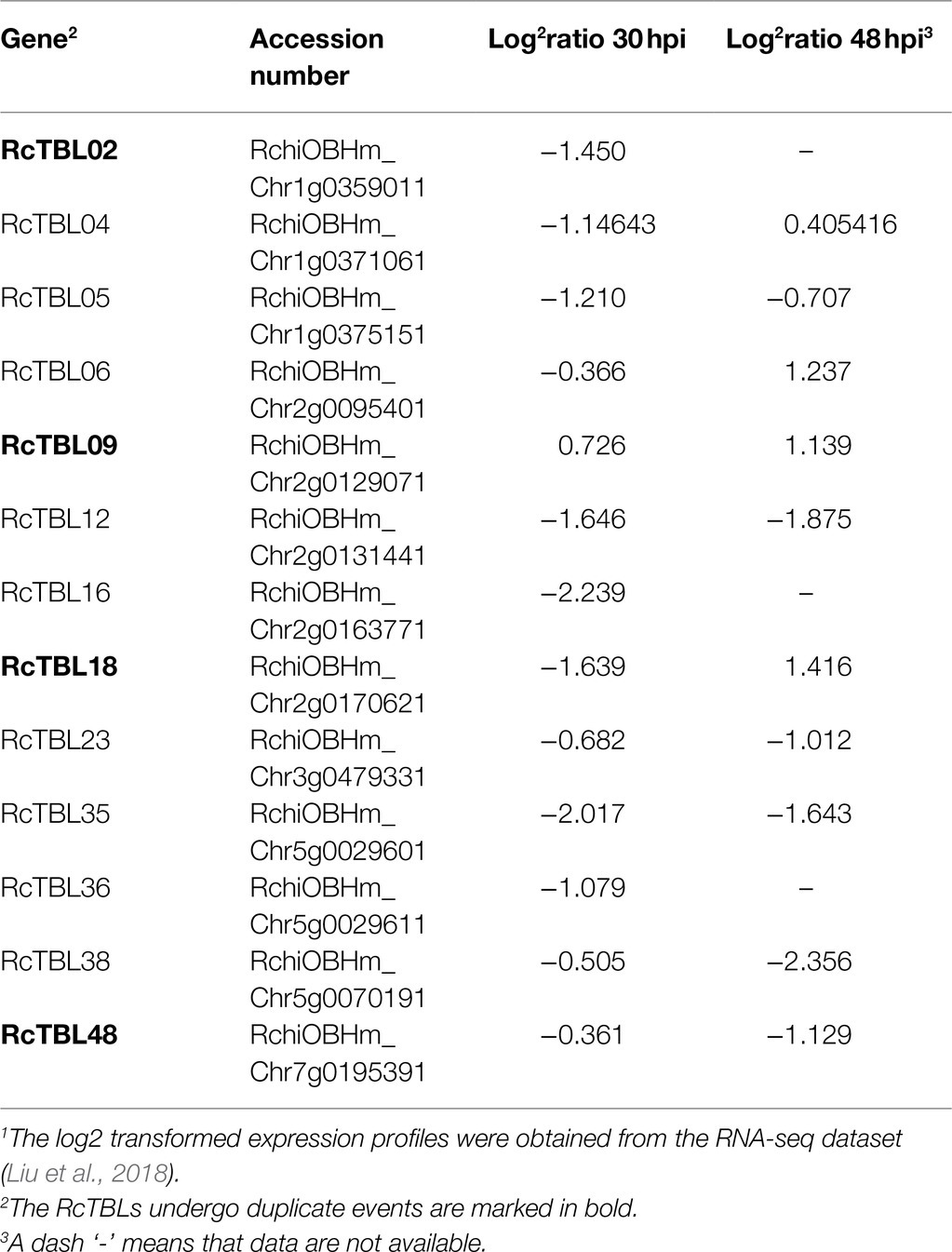
Growing evidence has shown that TBL plays an important role in pathogen defense. In order to investigate the role of RcTBLs in B. cinerea resistance, we obtained RNAseq transcriptomics data from rose petals at 30hpi (hours post-inoculation) and 48hpi (Liu et al., 2018). The 30hpi represents the early response to the infection of B. cinerea, while the 48hpi corresponds to the late response. A total of 13 RcTBL genes showed significant changes in expression and, interestingly, they were mainly down-regulated in expression (Table 4). Among them, RcTBL12 and RcTBL35 were both considerably down-regulated at 30hpi and 48hpi, whereas RcTBL02, RcTBL04, RcTBL05, RcTBL16, and RcTBL36 were only significantly down-regulated at 30hpi, RcTBL23, RcTBL38, and RcTBL48 were only significantly down-regulated at 48hpi. Surprisingly, RcTBL18 expression was dramatically decreased at 30hpi but significantly increased at 48hpi, whereas RcTBL06 and RcTBL09 expression was greatly increased at 48hpi. These genes, which are strongly activated by gray mold, could be crucial in rose resistance to B. cinerea infection. To further validate the RNA-seq expression profile, the expressions of five RcTBLs were examined using RT-qPCR. The RT-qPCR analysis results were found associated with the transcriptome analysis expression patterns (Figure 5).
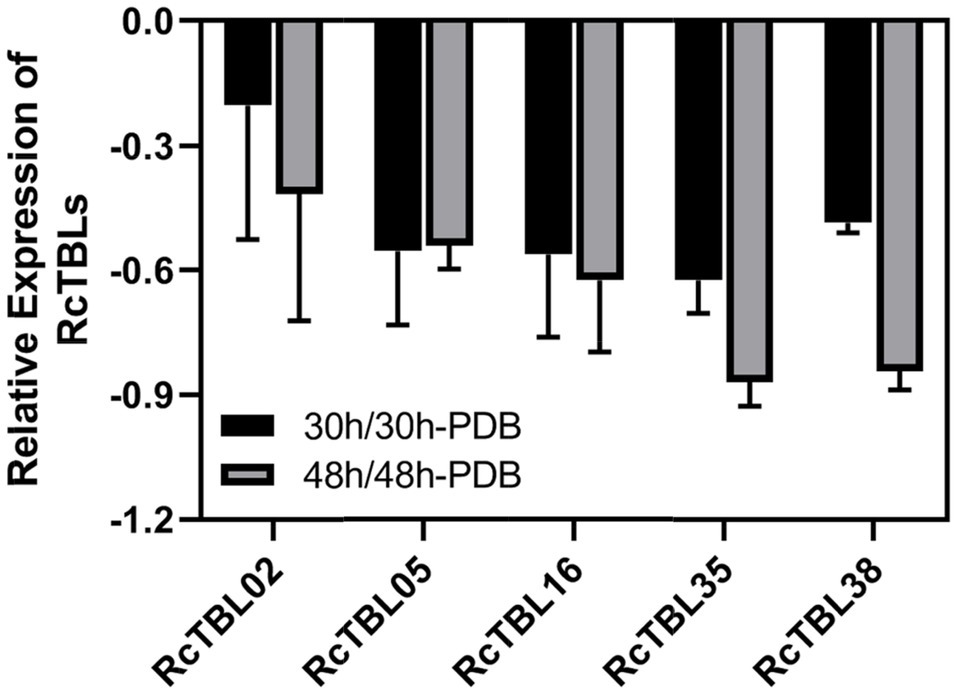
Figure 5. Validation of RNA-Seq results using RT-qPCR. RcUBI2 was used as an internal control. Expression profile data of five RcTBL genes at 30hpi and 48hpi after B. cinerea inoculation were obtained using RT-qPCR. Values are the means of three technical replicates ± SD. The used primers are listed in Supplementary Table S1.
RcTBL16 Involving in the Defense of Rose Against B. cinerea
The elucidation of gene expression patterns can provide important clues about gene function. Based on the expression results in response to B. cinerea, 13 RcTBLs with significant differential expression were considered as potential candidates participating in rose against B. cinerea. Their potential role in resistance to this fungus was further illustrated by the reduced expression in rose petals by VIGS. We selected RcTBL16 and RcTBL35 as candidate genes to explore the relationship between TBL family and rose resistance to B. cinerea, because (1) apart from RcTBL38 (a PMR5 homolog), RcTBL16 and RcTBL35 were the two of maximum down-regulated expressed RcTBL genes in response to B. cinerea (Table 4), and the results of RT-qPCR support this result (Figure 5); (2) both RcTBL16 and RcTBL35 are down-regulated in expression at 30hpi, which represent an early stages of B. cinerea infection (Liu et al., 2018).
In order to silence RcTBL16 and RcTBL35, we cloned approximately 200bp fragments of them into the pTRV2 vector to generate pTRV2-RcTBL16 and pTRV2-RcTBL35. Next, Agrobacterium tumefaciens carrying pTRV2-RcTBL16, pTRV2-RcTBL35 and pTRV1 were co-infiltrated into the rose petals to generate silent rose petals. Then, the infiltrated rose petal discs were put on agar medium and inoculated with B. cinerea. Comparing the control inoculated with TRV-GFP, petals inoculated with TRV-RcTBL16 showed notably reduced disease symptoms, whereas the area of disease spots on petals inoculated with TRV-RcTBL35 had no significantly changes (Figure 6). This result indicated that RcTBL16 is a susceptibility gene involved in Botrytis-rose interaction.
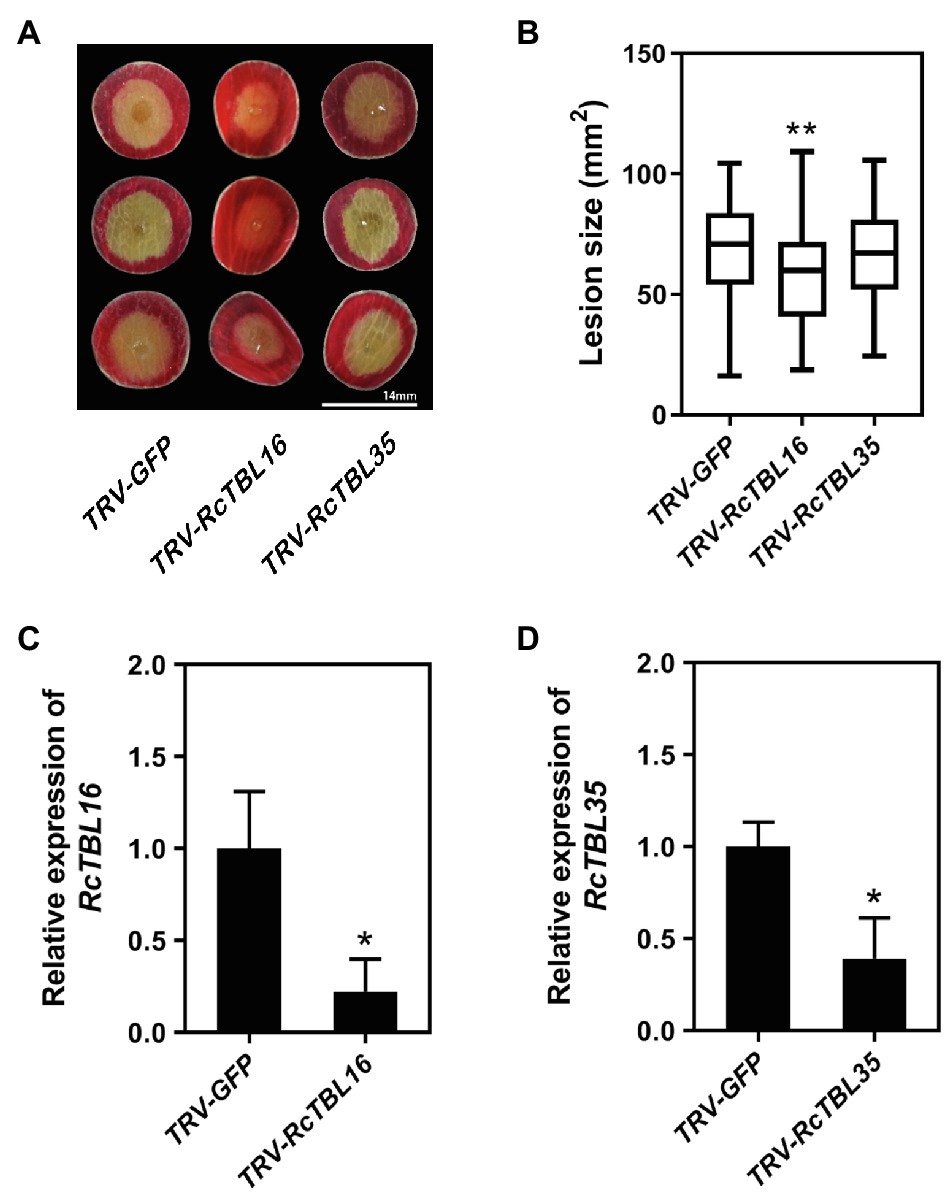
Figure 6. Functional analysis of RcTBL16. (A) Altered B. cinerea resistance upon silencing of RcTBL16 and unaltered B. cinerea resistance upon silenced RcTBL35 at 60hpi post-inoculation. (B) Quantification of B. cinerea disease lesions on TRV-RcTBL16, TRV-RcTBL35, and TRV-GFP-inoculated rose petal discs. The graph shows the lesion size from one replicate out of four (n≥48) with the standard deviation. (C) Quantification of RcTBL16 expression in TRV-RcTBL16 and TRV-GFP-inoculated petal discs. (D) Quantification of RcTBL35 expression in TRV-RcTBL35 and TRV- GFP-inoculated petal discs. Statistical analyses were performed using Student’s t test; *p<0.01, *p<0.05.
Discussion
O-acetylation is a common modification on plant cell walls and is essential for the stability of the polysaccharide network, with a small amount of acetylation modification affecting plant growth and susceptibility to pathogens. TBL is the largest protein family involved in O-acetylation in plants. There are 46 TBLs in Arabidopsis (Bischoff et al., 2010a), 18 of which have been identified as O-acetyltransferases, involved in the O-acetylation of xylan, xyloglucan, mannan and pectin respectively (Table 3). Several TBL proteins from rice and poplar have also been shown to be xylan O-acetyltransferases (Zhong et al., 2018a,d). However, there is still a gap in the comprehensive analysis of the rose TBL gene family, and their function remains to be discovered. Rose genome sequencing project was completed in 2018, enabling genome-wide analysis of the RcTBL gene family. In this study, we have comprehensively analyzed the RcTBL family, including phylogeny, gene structure, chromosomal location, gene duplication events, sequence characterization, and analysis of expression profiles.
We identified 50 rose TBLs, more than Arabidopsis (46) (Bischoff et al., 2010a) but less than poplar (64) (Zhong et al., 2018d), rice (66) (Gao et al., 2017), tomatoes (69) (Zhong et al., 2020), which indicates that TBL protein family has expanded to varying degrees in different plants during the evolution. In our structural analysis, we found that in addition to the PC-esterase domain, RcTBLs have a cys-rich domain, namely PMR5N domain, which may be another vital characteristic of the TBL family. Gene duplication plays a very important role in the expansion of gene families. Checking the phylogenetic relationships of TBL between rose and Arabidopsis showed that most evolved branches contained different amounts of AtTBL and RcTBL proteins, suggesting that the two species displayed conserved evolution.
The composition of the cell wall is closely correlated with fungal disease resistance, and altered levels of cell wall polysaccharide O-acetylation modification can lead to altered plant resistance to fungi. In some cases, a reduction in the level of acetylation enhances plant resistance to pathogens. For instance, Arabidopsis rwa2 showed a 20% reduction in the degree of cell wall polysaccharide acetylation modification and was more resistant to B. cinerea than wild type. In this study, we also found RcTBL16 negatively regulating resistance to gray mold in rose. PMR5, a member of the TBL family, has the same performance in resisting powdery mildew in Arabidopsis (Vogel et al., 2004a), by the O-acetylation modification of homogalacturonan (Chiniquy et al., 2019a). Therefore, we consider that partial rose TBL genes may be involved in the susceptibility of rose to B. cinerea through acetylating cell wall.
It is clear that changes in the level of O-acetylation modification of cell walls can affect plant resistance or susceptibility to pathogens, but the exact mechanism remains a mystery at present. One hypothesis suggests that alterations in the polysaccharide composition of the cell wall will modify the cell wall integrity (CWI) system, thereby triggering plant defenses and activating specific defense responses (Bacete et al., 2017). Alteration of cell wall xylan acetylation caused by ESK1 impairment was accompanied by an enhanced accumulation of abscisic acid, the constitutive expression of genes encoding antibiotic peptides and enzymes involved in the biosynthesis of tryptophan-derived metabolites, and the accumulation of disease resistance-related secondary metabolites and different osmolites, implying that the damage to CWI triggers defense response of plant (Lugan et al., 2010; Xin et al., 2010; Xu et al., 2014; Escudero et al., 2017). However, it needs to be substantiated by stronger evidence. Overall, more research remains to be done on the certain mechanisms of TBL participant plant–pathogen interactions, but there is no doubt that TBLs in rose is most possibly as a susceptibility factor for the resistance to gray mold, and this result will also provide clues for rose breeding application (i.e., enhancing persistent plant resistance by silencing or knocking out susceptibility genes).
Conclusion
The study performed a genome-wide analysis of RcTBLs, including phylogenetic relationships, collinearity, and expression analysis. A total of 50 non-redundant RcTBL members were identified. The expression analysis showed that transcription regulation of several RcTBL was reduced by B. cinerea infection in rose petals. Based on these analyses and VIGS, we further demonstrated that RcTBL16 was engaged in susceptibility of rose to gray mold. The information provided by these results can promote the further functional analysis of RcTBL genes and application in rose disease resistance breeding.
Data Availability Statement
Publicly available datasets were analyzed in this study. These data can be found at: https://lipm-browsers.toulouse.inra.fr/pub/RchiOBHm-V2/.
Author Contributions
ZZ and YT conceived and designed the research and wrote the paper. YT performed the experiments. YT, SZ, XL, and ZZ analyzed the data. All the authors have read and approved the final version of the manuscript. All authors contributed to the article and approved the submitted version.
Funding
his study was supported by the National Natural Science Foundation of China (grant number 31772344) to ZZ. It is further supported by the Science and Technology Program of Beijing Municipality (grant number Z181100002518002). The funders played no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2021.738880/full#supplementary-material
Abbreviations
hpi, hours post inoculation; ML, maximum likelihood; RPKM, number of reads per kb per million reads; HMM, hidden Markov model; CWI, cell wall integrity; aa, amino acids; VIGS, virus-induced gene silencing.
Footnotes
References
Bacete, L., Mélida, H., Miedes, E., and Molina, A. (2017). Plant cell wall-mediated immunity: cell wall changes trigger disease resistance responses. Plant J. 93, 614–636. doi: 10.1111/tpj.13807
Bendahmane, M., Dubois, A., Raymond, O., and Bris, M. L. (2013). Genetics and genomics of flower initiation and development in roses. J. Exp. Bot. 64, 847–857. doi: 10.1093/jxb/ers387
Bischoff, V., Nita, S., Neumetzler, L., Schindelasch, D., Urbain, A., Eshed, R., et al. (2010a). TRICHOME BIREFRINGENCE and its homolog AT5G01360 encode plant-specific DUF231 proteins required for cellulose biosynthesis in Arabidopsis. Plant Physiol. 153, 590–602. doi: 10.1104/pp.110.153320
Bischoff, V., Selbig, J., and Scheible, W. R. (2010b). Involvement of TBL/DUF231 proteins into cell wall biology. Plant Signaling 5, 1057–1059.
Cao, X., Yan, H., Liu, X., Li, D., Sui, M., Wu, J., et al. (2019). A detached petal disc assay and virus-induced gene silencing facilitate the study of Botrytis cinerea resistance in rose flowers. Hortic. Res. 6:136. doi: 10.1038/s41438-019-0219-2
Chen, C., Xia, R., Chen, H., and He, Y. (2018). TBtools, a toolkit for biologists integrating various HTS-data handling tools with a user-friendly interface. Mol. Plant 13, 1194–1202. doi: 10.1016/j.molp.2020.06.009
Chiniquy, D., Underwood, W., Corwin, J., Ryan, A., Szemenyei, H., Lim, C. C., et al. (2019a). PMR5, an acetylation protein at the intersection of pectin biosynthesis and defense against fungal pathogens. Plant J. 100, 1022–1035. doi: 10.1111/tpj.14497
Chiniquy, D., Underwood, W., Corwin, J., Ryan, A., Szemenyei, H., Lim, C. C., et al. (2019b). PMR5, an acetylation protein at the intersection of pectin biosynthesis and defense against fungal pathogens. Plant J. 100, 1022–1035. doi: 10.1111/tpj.14497
Escudero, V., Jordá, L., SopeA-Torres, S., Mélida, H., Miedes, E., MuOz-Barrios, A., et al. (2017). Alteration of cell wall xylan acetylation triggers defense responses that counterbalance the immune deficiencies of plants impaired in the β-subunit of the heterotrimeric G-protein. Plant J. 92, 386–399. doi: 10.1111/tpj.13660
Gao, Y., He, C., Zhang, D., Liu, X., and Xu, Z. (2017). Two trichome birefringence-like proteins mediate xylan acetylation, which is essential for leaf blight resistance in rice. Plant Physiol. 173, 470–481. doi: 10.1104/pp.16.01618
Gille, S., de Souza, A., Xiong, G., Benz, M., Cheng, K., Schultink, A., et al. (2011a). O-acetylation of Arabidopsis hemicellulose xyloglucan requires AXY4 or AXY4L, proteins with a TBL and DUF231 domain. Plant Cell 23, 4041–4053. doi: 10.1105/tpc.111.091728
Gille, S., De Souza, A., Xiong, G., Benz, M., Cheng, K., Schultink, A., et al. (2011b). O-acetylation of Arabidopsis hemicellulose xyloglucan requires AXY4 or AXY4L, proteins with a TBL and DUF231 domain. Plant Cell 23, 4041–4053. doi: 10.1105/tpc.111.091728
Gleason, M., and Helland, S. (2003). “DISEASE|Botrytis,” in Encyclopedia of Rose Science. eds. A. Roberts, T. Debener, and S. Gudin (Amsterdam: Elsevier Academic Press), 144–148.
Liu, X., Cao, X., Shi, S., Zhao, N., Li, D., Fang, P., et al. (2018). Comparative RNA-Seq analysis reveals a critical role for brassinosteroids in rose (Rosa hybrida) petal defense against Botrytis cinerea infection. BMC Genet. 19, 1–10. doi: 10.1186/s12863-018-0668-x
Liu, Y., Schiff, M., and Dinesh-Kumar, S. P. (2002). Virus-induced gene silencing in tomato. Plant J. 31, 777–786. doi: 10.1046/j.1365-313X.2002.01394.x
Lugan, R., Niogret, M.-F., Kervazo, L., Larher, F. R., and Bouchereau, A. (2010). Metabolome and water status phenotyping of Arabidopsis under abiotic stress cues reveals new insight into ESK1 function. Plant Cell Environ. Ecol. 32, 95–108. doi: 10.1111/j.1365-3040.2008.01898.x
Manabe, Y., Nafisi, M., Verhertbruggen, Y., Orfila, C., Gille, S., Rautengarten, C., et al. (2011). Loss-of-function mutation of REDUCED WALL ACETYLATION2 in Arabidopsis leads to reduced cell wall ACETYLATION and increased resistance to Botrytis cinerea. Plant Physiol. 155, 1068–1078. doi: 10.1104/pp.110.168989
Manabe, Y., Verhertbruggen, Y., Gille, S., Harholt, J., Chong, S.-L., Pawar, P. M.-A., et al. (2013). Reduced wall acetylation proteins play vital and distinct roles in cell wall o-acetylation in Arabidopsis. Plant Physiol. 163, 1107–1117. doi: 10.1104/pp.113.225193
Schultink, A., Naylor, D., Dama, M., and Pauly, M. (2015). The role of the plant-specific ALTERED XYLOGLUCAN9 protein in Arabidopsis cell wall polysaccharide O-acetylation. Plant Physiol. 167, 1271–U1243. doi: 10.1104/pp.114.256479
Stranne, M., Ren, Y., Fimognari, L., Birdseye, D., and Sakuragi, Y. (2018a). TBL10 is required for O-acetylation of pectic rhamnogalacturonan-I in Arabidopsis thaliana. Plant J. 96, 772–785. doi: 10.1111/tpj.14067
Stranne, M., Ren, Y., Fimognari, L., Birdseye, D., Yan, J., Bardor, M., et al. (2018b). TBL10 is required for O-acetylation of pectic rhamnogalacturonan-I in Arabidopsis thaliana. Plant J. 96, 772–785. doi: 10.1111/tpj.14067
Urbanowicz, B. R., Pea, M. J., Moniz, H. A., Moremen, K. W., and York, W. S. (2015). Two Arabidopsis proteins synthesize acetylated xylan invitro. Plant J. 80, 197–206. doi: 10.1111/tpj.12643
Vogel, J. P., Raab, T. K., Somerville, C. R., and Somerville, S. C. (2004a). Mutations in PMR5 result in powdery mildew resistance and altered cell wall composition. Plant J. 40:968. doi: 10.1111/j.1365-313X.2004.02264.x
Vogel, J. P., Raab, T. K., Somerville, C. R., and Somerville, S. C. (2004b). Mutations in PMR5 result in powdery mildew resistance and altered cell wall composition. Plant J. 40, 968–978. doi: 10.1111/j.1365-313X.2004.02264.x
Wang, Y., Tang, H., DeBarry, J. D., Tan, X., Li, J., Wang, X., et al. (2012). MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40:e49. doi: 10.1093/nar/gkr1293
Wu, L., Ma, N., Jia, Y., Zhang, Y., Feng, M., Jiang, C. Z., et al. (2016). An ethylene-induced regulatory module delays flower senescence by regulating cytokinin content. Plant Physiol. 173, 853–862. doi: 10.1104/pp.16.01064
Xin, Z., Mandaokar, A., Chen, J., Last, R. L., and Browse, J. (2010). Arabidopsis ESK1 encodes a novel regulator of freezing tolerance. Plant J. 49, 786–799. doi: 10.1111/j.1365-313X.2006.02994.x
Xiong, G., Cheng, K., and Pauly, M. (2013). Xylan O-acetylation impacts xylem development and enzymatic recalcitrance as indicated by the Arabidopsis mutant tbl29. Mol. Plant 6, 1373–1375. doi: 10.1093/mp/sst014
Xu, F., Liu, Z., Xie, H., Zhu, J., and Zhang, J. (2014). Increased drought tolerance through the suppression of ESKMO1 gene and Overexpression of CBF-related genes in Arabidopsis. PLoS One 9:e106509. doi: 10.1371/journal.pone.0115300
Yuan, Y., Teng, Q., Zhong, R., Haghighat, M., Richardson, E. A., and Ye, Z.-H. (2016a). Mutations of Arabidopsis TBL32 and TBL33 affect xylan acetylation and secondary wall deposition. PLoS One 11:e0146460. doi: 10.1371/journal.pone.0146460
Yuan, Y., Teng, Q., Zhong, R., and Ye, Z.-H. (2016b). Roles of Arabidopsis TBL34 and TBL35 in xylan acetylation and plant growth. Plant Sci. 243, 120–130. doi: 10.1016/j.plantsci.2015.12.007
Yuan, Y., Teng, Q., Zhong, R., and Ye, Z.-H. (2016c). TBL3 and TBL31, two Arabidopsis DUF231 domain proteins, are required for 3-O-monoacetylation of xylan. Plant Cell Physiol. 57, 35–45. doi: 10.1093/pcp/pcv172
Zhong, R., Cui, D., Dasher, R. L., and Ye, Z. H. (2018a). Biochemical characterization of rice xylan O-acetyltransferases. Planta 247, 1–10. doi: 10.1007/s00425-018-2882-1
Zhong, R., Cui, D., Phillips, D. R., Richardson, E. A., and Ye, Z.-H. (2020). A group of O-acetyltransferases catalyze xyloglucan backbone acetylation and can alter xyloglucan xylosylation pattern and plant growth when expressed in Arabidopsis. Plant Cell Physiol. 61, 1064–1079. doi: 10.1093/pcp/pcaa031
Zhong, R. Q., Cui, D. T., and Ye, Z. H. (2017a). Regiospecific acetylation of xylan is mediated by a group of DUF231-containing O-acetyltransferases. Plant Cell Physiol. 58, 2126–2138. doi: 10.1093/pcp/pcx147
Zhong, R., Cui, D., and Ye, Z.-H. (2018b). Members of the DUF231 family are O-acetyltransferases catalyzing 2-O- and 3-O-acetylation of mannan. Plant Cell Physiol. 59, 2339–2349. doi: 10.1093/pcp/pcy159
Zhong, R., Cui, D., and Ye, Z.-H. (2018c). Xyloglucan O -acetyltransferases from Arabidopsis thaliana and Populus trichocarpa catalyze acetylation of fucosylated galactose residues on xyloglucan side chains. Planta 248, 1159–1171. doi: 10.1007/s00425-018-2972-0
Zhong, R. Q., Cui, D. T., and Ye, Z. H. (2018e). Xyloglucan O-acetyltransferases from Arabidopsis thaliana and Populus trichocarpa catalyze acetylation of fucosylated galactose residues on xyloglucan side chains. Planta 248, 1159–1171. doi: 10.1007/s00425-018-2972-0
Zhong, R., Cui, D., Ye, Z.-H., and Zabotina, O. A. (2018d). A group of Populus trichocarpa DUF231 proteins exhibit differential O-acetyltransferase activities toward xylan. PLoS One 13:e0194532. doi: 10.1371/journal.pone.0194532
Zhong, R., Dongtao, C., and Zheng-Hua, Y. (2017b). Regiospecific acetylation of xylan is mediated by a group of DUF231-containing O-acetyltransferases. Plant Cell Physiol. 58, 2126–2138. doi: 10.1093/pcp/pcx147
Keywords: Rosa sp., TBL, Botrytis cinerea, gene family, expression, S-gene
Citation: Tian Y, Zhang S, Liu X and Zhang Z (2021) Global Investigation of TBL Gene Family in Rose (Rosa chinensis) Unveils RcTBL16 Is a Susceptibility Gene in Gray Mold Resistance. Front. Plant Sci. 12:738880. doi: 10.3389/fpls.2021.738880
Edited by:
Meixiang Zhang, Nanjing Agricultural University, ChinaReviewed by:
Qiong Zhang, University of California, Berkeley, United StatesBiao Gu, Northwest A and F University, China
Copyright © 2021 Tian, Zhang, Liu and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhao Zhang, emhhbmd6aGFvQGNhdS5lZHUuY24=
 Yu Tian
Yu Tian Shiya Zhang
Shiya Zhang Zhao Zhang
Zhao Zhang